microRNA information: hsa-let-7e-3p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-let-7e-3p | miRbase |
| Accession: | MIMAT0004485 | miRbase |
| Precursor name: | hsa-let-7e | miRbase |
| Precursor accession: | MI0000066 | miRbase |
| Symbol: | MIRLET7E | HGNC |
| RefSeq ID: | NR_029482 | GenBank |
| Sequence: | CUAUACGGCCUCCUAGCUUUCC |
Reported expression in cancers: hsa-let-7e-3p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-let-7e-3p | breast cancer | deregulation | "Three of these miRNAs were up-regulated miR-141 mi ......" | 25451164 | |
| hsa-let-7e-3p | colorectal cancer | deregulation | "In our CD patients six miRNAs were upregulated fro ......" | 22241525 | |
| hsa-let-7e-3p | esophageal cancer | deregulation | "Differentially-expressed miRNAs were analyzed usin ......" | 23534712 | Reverse transcription PCR |
| hsa-let-7e-3p | lung squamous cell cancer | downregulation | "Differential expression of miR 125a 5p and let 7e ......" | 24945821 | qPCR |
| hsa-let-7e-3p | ovarian cancer | deregulation | "Among them 10 miRNA were down-regulated including ......" | 21122376 | |
| hsa-let-7e-3p | retinoblastoma | upregulation | "Identification of miRNAs associated with tumorigen ......" | 18818933 | Microarray; Northern blot; in situ hybridization |
| hsa-let-7e-3p | sarcoma | upregulation | "There were 21 significantly up-regulated miRNAs in ......" | 21213367 |
Reported cancer pathway affected by hsa-let-7e-3p
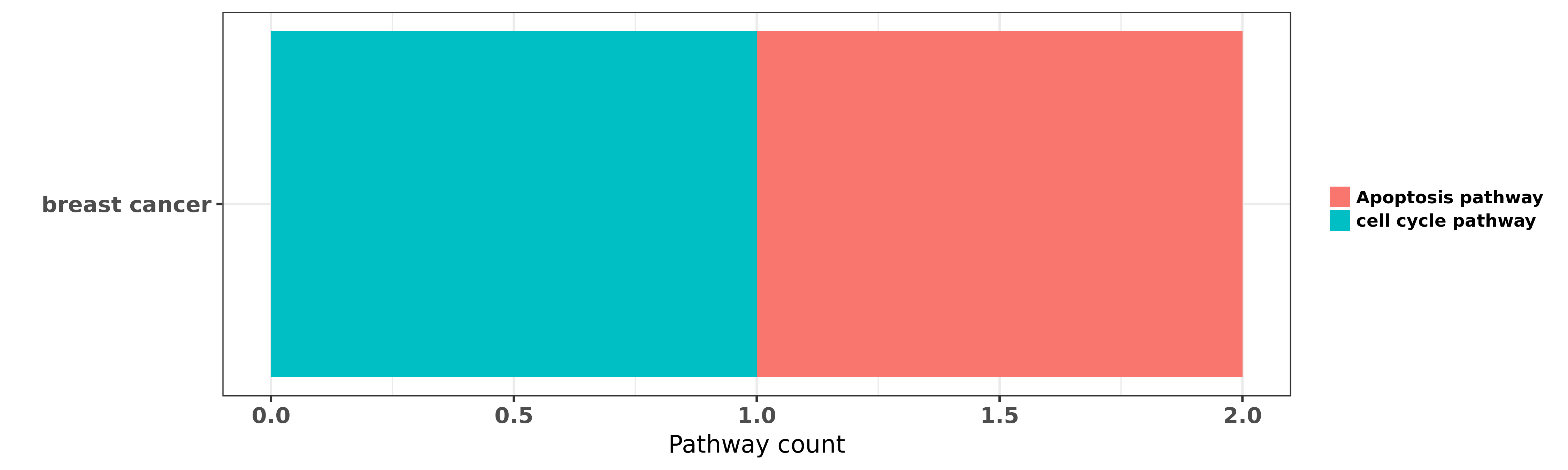
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-let-7e-3p | breast cancer | cell cycle pathway | "Jumonji/ARID1 B JARID1B protein promotes breast tu ......" | 21969366 | RNAi |
| hsa-let-7e-3p | breast cancer | Apoptosis pathway | "In vitro functional experiments were performed in ......" | 24257477 |
Reported cancer prognosis affected by hsa-let-7e-3p
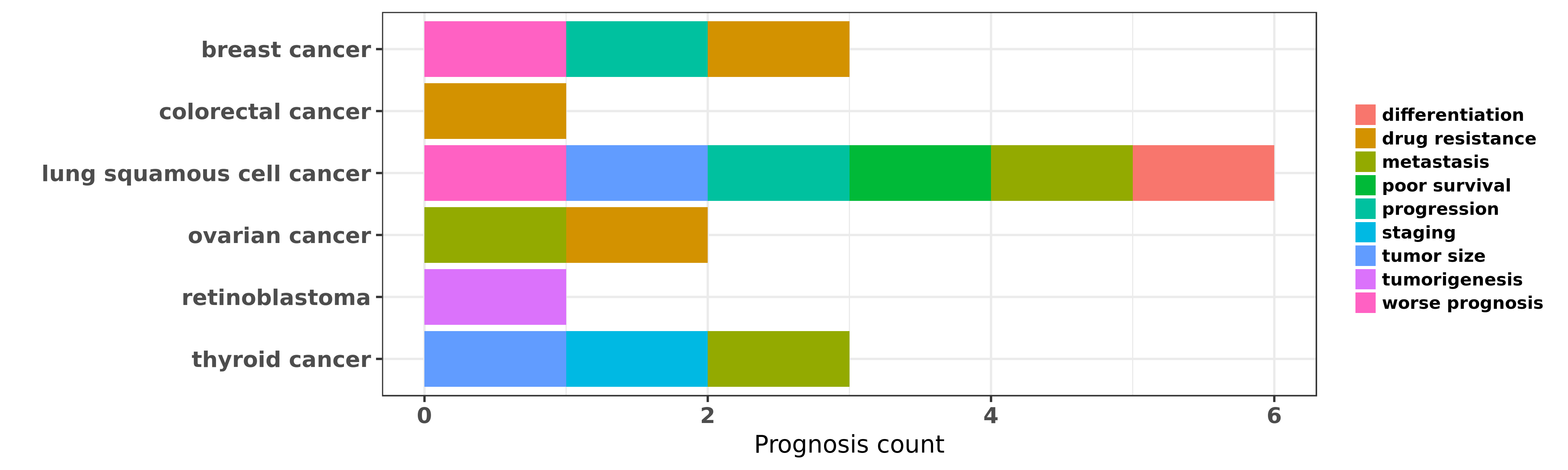
| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-let-7e-3p | breast cancer | progression | "Jumonji/ARID1 B JARID1B protein promotes breast tu ......" | 21969366 | RNAi |
| hsa-let-7e-3p | breast cancer | worse prognosis | "In vitro functional experiments were performed in ......" | 24257477 | |
| hsa-let-7e-3p | breast cancer | drug resistance | "We created a doxorubicin-resistant MCF-7 MCF-/Adr ......" | 25451164 | MTT assay |
| hsa-let-7e-3p | colorectal cancer | drug resistance | "We investigated this issue by profiling the expres ......" | 20881268 | |
| hsa-let-7e-3p | lung squamous cell cancer | metastasis; differentiation; tumor size; poor survival; worse prognosis | "High-throughput microarray was used to measure miR ......" | 22618509 | |
| hsa-let-7e-3p | lung squamous cell cancer | progression; worse prognosis; differentiation; poor survival | "Differential expression of miR 125a 5p and let 7e ......" | 24945821 | |
| hsa-let-7e-3p | ovarian cancer | drug resistance | "High-throughput analysis of the miRNA profile in a ......" | 18823650 | |
| hsa-let-7e-3p | ovarian cancer | metastasis | "miRNA microarray was applied to compare the miRNA ......" | 21122376 | |
| hsa-let-7e-3p | retinoblastoma | tumorigenesis | "Identification of miRNAs associated with tumorigen ......" | 18818933 | |
| hsa-let-7e-3p | thyroid cancer | staging; metastasis; tumor size | "Genome-wide serum miRNA expression profiles were d ......" | 22472564 | |
| hsa-let-7e-3p | thyroid cancer | staging; metastasis; tumor size | "Expression levels of miRs-21 -34b -130b -135b -146 ......" | 26414548 |
Reported gene related to hsa-let-7e-3p
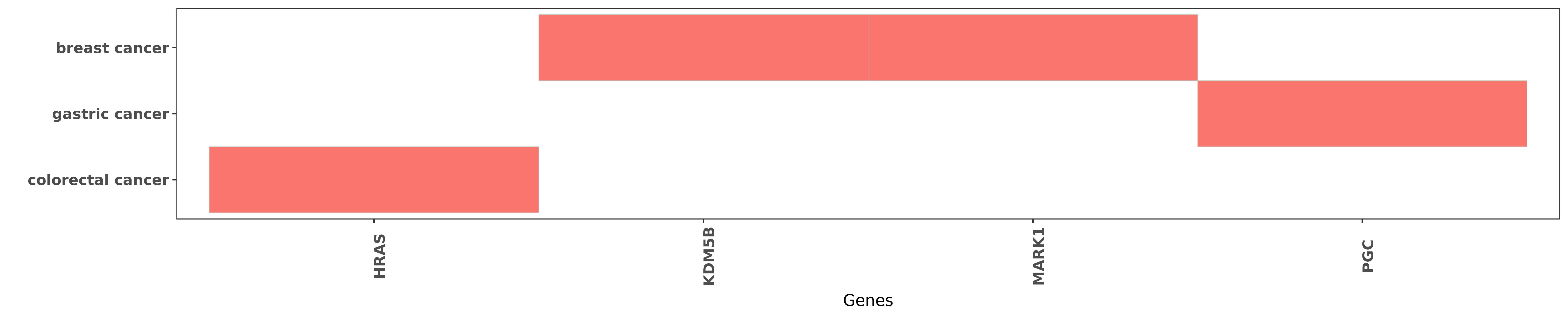
| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-let-7e-3p | colorectal cancer | HRAS | "On the basis of these data we suggest that the dow ......" | 20881268 |
| hsa-let-7e-3p | breast cancer | KDM5B | "Jumonji/ARID1 B JARID1B protein promotes breast tu ......" | 21969366 |
| hsa-let-7e-3p | breast cancer | MARK1 | "Chromatin immunoprecipitation analysis demonstrate ......" | 21969366 |
| hsa-let-7e-3p | gastric cancer | PGC | "Also let-7e rs8111742 PGC rs6458238 and H pylori i ......" | 26988755 |
Expression profile in cancer corhorts:
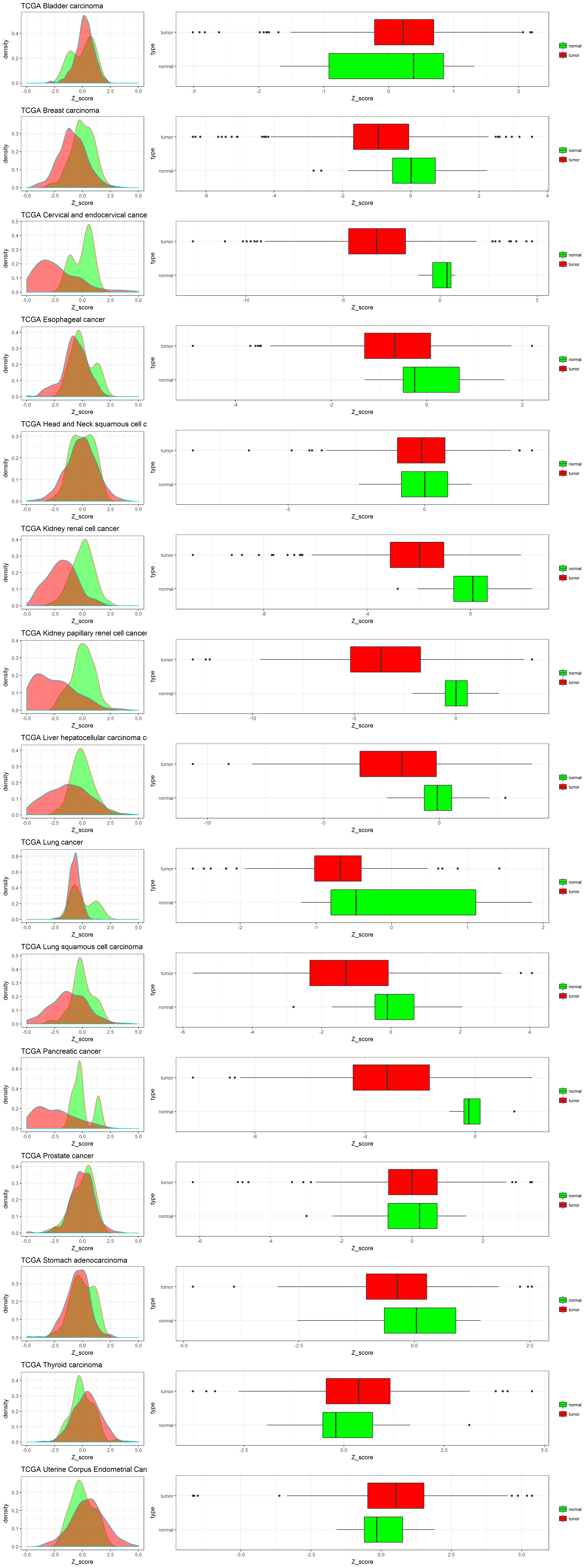
Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-let-7e-3p | ZCCHC6 | 14 cancers: BLCA; CESC; ESCA; HNSC; KIRC; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.124; TCGA CESC -0.204; TCGA ESCA -0.243; TCGA HNSC -0.199; TCGA KIRC -0.087; TCGA LIHC -0.082; TCGA LUAD -0.062; TCGA LUSC -0.13; TCGA OV -0.144; TCGA PAAD -0.159; TCGA PRAD -0.269; TCGA THCA -0.089; TCGA STAD -0.207; TCGA UCEC -0.189 |
hsa-let-7e-3p | COPS2 | 10 cancers: BLCA; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.074; TCGA COAD -0.078; TCGA ESCA -0.079; TCGA HNSC -0.058; TCGA LUAD -0.092; TCGA LUSC -0.11; TCGA PRAD -0.078; TCGA SARC -0.071; TCGA STAD -0.093; TCGA UCEC -0.116 |
Enriched cancer pathways of putative targets