microRNA information: hsa-let-7f-5p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-let-7f-5p | miRbase |
| Accession: | MIMAT0000067 | miRbase |
| Precursor name: | hsa-let-7f-1 | miRbase |
| Precursor accession: | MI0000067 | miRbase |
| Symbol: | MIRLET7F1 | HGNC |
| RefSeq ID: | NR_029483 | GenBank |
| Sequence: | UGAGGUAGUAGAUUGUAUAGUU |
Reported expression in cancers: hsa-let-7f-5p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-let-7f-5p | colon cancer | deregulation | "The expression of 384 miRNAs was determined by usi ......" | 26556872 | Microarray |
| hsa-let-7f-5p | liver cancer | deregulation | "Expression of serum miR 16 let 7f and miR 21 in pa ......" | 24697119 | Reverse transcription PCR; qPCR |
| hsa-let-7f-5p | ovarian cancer | deregulation | "Among them 10 miRNA were down-regulated including ......" | 21122376 | |
| hsa-let-7f-5p | thyroid cancer | downregulation | "Thus reduced expression of let-7f might be an esse ......" | 19956384 | |
| hsa-let-7f-5p | thyroid cancer | downregulation | "MicroRNAs miR 146 5p and let 7f as prognostic tool ......" | 23295297 |
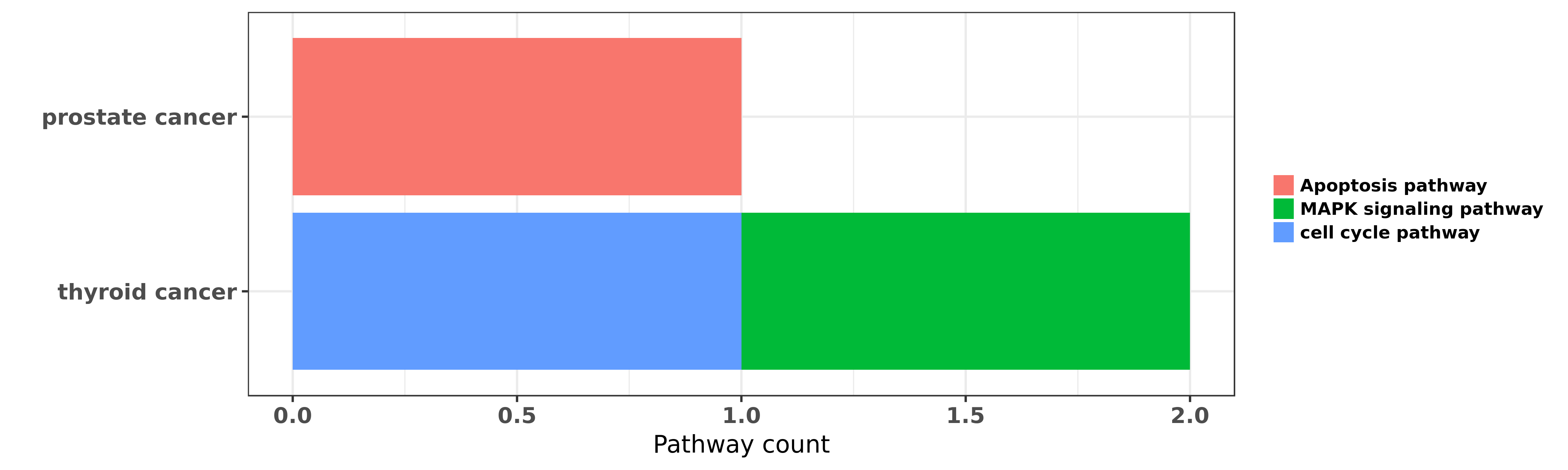
Reported cancer pathway affected by hsa-let-7f-5p
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-let-7f-5p | prostate cancer | Apoptosis pathway | "MicroRNA let 7f 1 is induced by lycopene and inhib ......" | 26847233 | Western blot |
| hsa-let-7f-5p | thyroid cancer | cell cycle pathway; MAPK signaling pathway | "Here we report that RET/PTC3 oncogenic activation ......" | 19956384 |
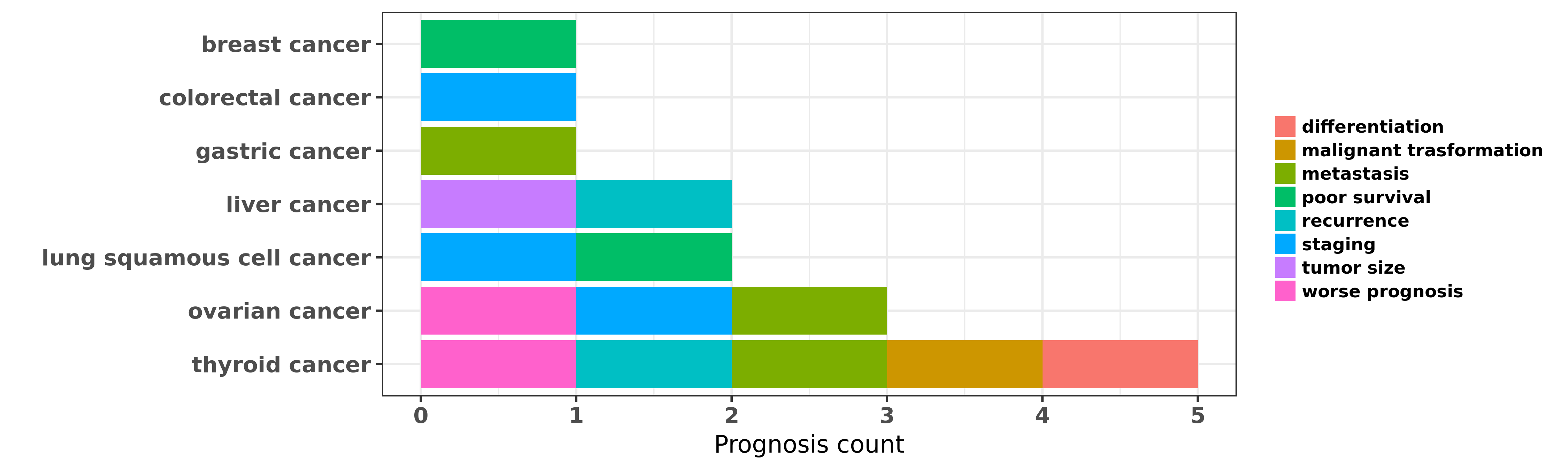
Reported cancer prognosis affected by hsa-let-7f-5p
| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-let-7f-5p | breast cancer | poor survival | "To investigate the global expression profile of mi ......" | 18812439 | |
| hsa-let-7f-5p | colorectal cancer | staging | "Simultaneous Underexpression of let 7a 5p and let ......" | 26793011 | |
| hsa-let-7f-5p | gastric cancer | metastasis | "MicroRNA let 7f inhibits tumor invasion and metast ......" | 21533124 | Western blot; Luciferase |
| hsa-let-7f-5p | liver cancer | recurrence; tumor size | "Expression of serum miR 16 let 7f and miR 21 in pa ......" | 24697119 | |
| hsa-let-7f-5p | lung squamous cell cancer | poor survival; staging | "Five selected miRNAs let-7f miR-20b miR-30e-3p miR ......" | 20595154 | |
| hsa-let-7f-5p | ovarian cancer | metastasis | "miRNA microarray was applied to compare the miRNA ......" | 21122376 | |
| hsa-let-7f-5p | ovarian cancer | staging; worse prognosis | "We scanned the circulating plasma miRNAs by TaqMan ......" | 24223734 | |
| hsa-let-7f-5p | thyroid cancer | malignant trasformation; differentiation | "Here we report that RET/PTC3 oncogenic activation ......" | 19956384 | |
| hsa-let-7f-5p | thyroid cancer | metastasis; recurrence; worse prognosis | "MicroRNAs miR 146 5p and let 7f as prognostic tool ......" | 23295297 |
Reported gene related to hsa-let-7f-5p

| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-let-7f-5p | kidney renal cell cancer | KLK10 | "A luciferase assay demonstrated that hsa-let-7f ca ......" | 20180642 |
| hsa-let-7f-5p | ovarian cancer | KLK10 | "When we co-transfected cells with pMIR-KLK10 and e ......" | 20354523 |
| hsa-let-7f-5p | breast cancer | CYP19A1 | "Aromatase inhibitor treatment of breast cancer cel ......" | 22407818 |
| hsa-let-7f-5p | kidney renal cell cancer | LARP6 | "A luciferase assay demonstrated that hsa-let-7f ca ......" | 20180642 |
| hsa-let-7f-5p | gastric cancer | MYH9 | "MicroRNA let 7f inhibits tumor invasion and metast ......" | 21533124 |
Expression profile in cancer corhorts:
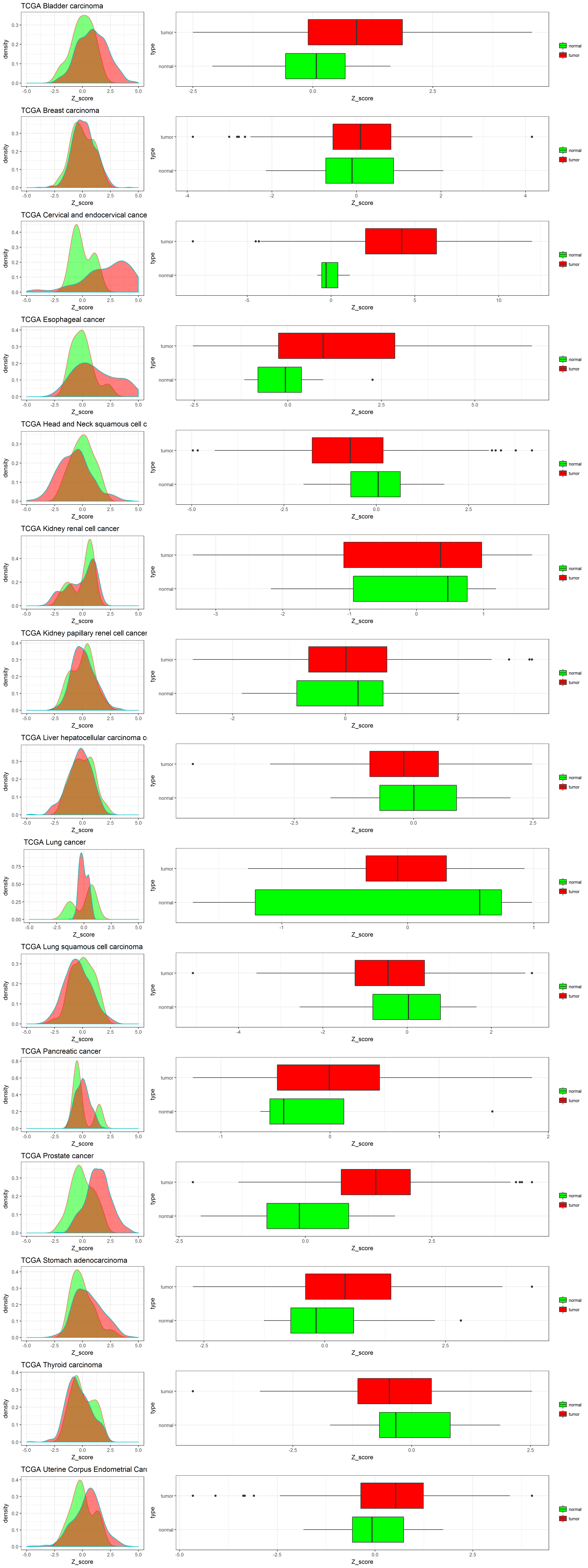
Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-let-7f-5p | CERCAM | 9 cancers: BLCA; CESC; COAD; ESCA; LUAD; PAAD; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.357; TCGA CESC -0.212; TCGA COAD -0.437; TCGA ESCA -0.194; TCGA LUAD -0.206; TCGA PAAD -0.177; TCGA PRAD -0.165; TCGA STAD -0.308; TCGA UCEC -0.175 |
hsa-let-7f-5p | LIN28B | 9 cancers: BLCA; CESC; HNSC; LIHC; LUAD; LUSC; OV; STAD; UCEC | MirTarget; miRNATAP; RAID | TCGA BLCA -0.707; TCGA CESC -0.741; TCGA HNSC -0.601; TCGA LIHC -1.964; TCGA LUAD -0.334; TCGA LUSC -0.76; TCGA OV -1.027; TCGA STAD -0.656; TCGA UCEC -1.167 |
hsa-let-7f-5p | IGLON5 | 9 cancers: BLCA; COAD; ESCA; HNSC; KIRC; LGG; LUAD; OV; STAD | miRNATAP | TCGA BLCA -0.572; TCGA COAD -0.495; TCGA ESCA -0.379; TCGA HNSC -0.266; TCGA KIRC -0.292; TCGA LGG -0.192; TCGA LUAD -0.243; TCGA OV -0.391; TCGA STAD -0.295 |
hsa-let-7f-5p | FBXO32 | 9 cancers: BLCA; COAD; ESCA; LUSC; PRAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.445; TCGA COAD -0.414; TCGA ESCA -0.225; TCGA LUSC -0.408; TCGA PRAD -0.269; TCGA SARC -0.521; TCGA THCA -0.158; TCGA STAD -0.614; TCGA UCEC -0.26 |
hsa-let-7f-5p | DZIP1 | 10 cancers: BLCA; COAD; ESCA; LGG; LUAD; OV; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.443; TCGA COAD -0.524; TCGA ESCA -0.333; TCGA LGG -0.081; TCGA LUAD -0.106; TCGA OV -0.113; TCGA PAAD -0.19; TCGA PRAD -0.258; TCGA STAD -0.466; TCGA UCEC -0.248 |
hsa-let-7f-5p | SPEG | 10 cancers: BLCA; COAD; ESCA; KIRC; KIRP; LUAD; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.917; TCGA COAD -1.472; TCGA ESCA -0.408; TCGA KIRC -0.243; TCGA KIRP -0.412; TCGA LUAD -0.28; TCGA PRAD -0.404; TCGA SARC -0.939; TCGA STAD -0.853; TCGA UCEC -0.754 |
hsa-let-7f-5p | CALD1 | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; PAAD; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.683; TCGA BRCA -0.081; TCGA CESC -0.188; TCGA COAD -0.509; TCGA ESCA -0.225; TCGA PAAD -0.312; TCGA PRAD -0.378; TCGA SARC -0.442; TCGA STAD -0.55; TCGA UCEC -0.257 |
hsa-let-7f-5p | PLEKHO1 | 9 cancers: BLCA; COAD; ESCA; LIHC; LUAD; OV; PRAD; THCA; STAD | miRNATAP | TCGA BLCA -0.457; TCGA COAD -0.491; TCGA ESCA -0.154; TCGA LIHC -0.112; TCGA LUAD -0.09; TCGA OV -0.091; TCGA PRAD -0.162; TCGA THCA -0.138; TCGA STAD -0.289 |
hsa-let-7f-5p | ELOVL4 | 9 cancers: BLCA; COAD; ESCA; HNSC; LUSC; OV; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.518; TCGA COAD -0.395; TCGA ESCA -0.643; TCGA HNSC -0.28; TCGA LUSC -0.309; TCGA OV -0.245; TCGA THCA -0.097; TCGA STAD -0.595; TCGA UCEC -0.593 |
hsa-let-7f-5p | SESTD1 | 9 cancers: BRCA; COAD; KIRP; OV; PAAD; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.077; TCGA COAD -0.135; TCGA KIRP -0.214; TCGA OV -0.152; TCGA PAAD -0.095; TCGA PRAD -0.134; TCGA SARC -0.108; TCGA STAD -0.143; TCGA UCEC -0.153 |
hsa-let-7f-5p | DOCK3 | 10 cancers: CESC; COAD; HNSC; KIRC; LIHC; LUAD; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA CESC -0.326; TCGA COAD -0.338; TCGA HNSC -0.218; TCGA KIRC -0.279; TCGA LIHC -0.202; TCGA LUAD -0.198; TCGA PRAD -0.264; TCGA SARC -0.562; TCGA STAD -0.5; TCGA UCEC -0.312 |
hsa-let-7f-5p | FBXL19 | 9 cancers: HNSC; KIRC; KIRP; LGG; LIHC; LUAD; OV; THCA; UCEC | miRNATAP | TCGA HNSC -0.102; TCGA KIRC -0.061; TCGA KIRP -0.136; TCGA LGG -0.062; TCGA LIHC -0.076; TCGA LUAD -0.237; TCGA OV -0.063; TCGA THCA -0.057; TCGA UCEC -0.108 |
Enriched cancer pathways of putative targets