microRNA information: hsa-miR-106a-5p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-106a-5p | miRbase |
| Accession: | MIMAT0000103 | miRbase |
| Precursor name: | hsa-mir-106a | miRbase |
| Precursor accession: | MI0000113 | miRbase |
| Symbol: | MIR106A | HGNC |
| RefSeq ID: | NR_029523 | GenBank |
| Sequence: | AAAAGUGCUUACAGUGCAGGUAG |
Reported expression in cancers: hsa-miR-106a-5p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-106a-5p | bladder cancer | deregulation | "By using high-throughput sequencing technologies w ......" | 23939156 | RNA-Seq |
| hsa-miR-106a-5p | breast cancer | upregulation | "It is based on complete miRNA release from the pro ......" | 26967564 | qPCR |
| hsa-miR-106a-5p | chronic myeloid leukemia | deregulation | "Some onco-miRNAs were found to be downregulated mi ......" | 25833191 | |
| hsa-miR-106a-5p | colon cancer | downregulation | "Deregulated expression of miR 106a predicts surviv ......" | 18521848 | |
| hsa-miR-106a-5p | colon cancer | upregulation | "To present proof-of-principle application for empl ......" | 23741026 | qPCR; Microarray |
| hsa-miR-106a-5p | colorectal cancer | upregulation | "Expression of plasma miR 106a in colorectal cancer ......" | 24670448 | qPCR |
| hsa-miR-106a-5p | colorectal cancer | upregulation | "Plasma miR-106a miR-20a miR-27b miR-92a and miR-29 ......" | 26261602 | qPCR |
| hsa-miR-106a-5p | gastric cancer | upregulation | "The most highly expressed miRNAs in gastric cancer ......" | 19175831 | |
| hsa-miR-106a-5p | gastric cancer | upregulation | "The expression levels of miR-20a miR-21 miR-25 miR ......" | 22996433 | qPCR |
| hsa-miR-106a-5p | gastric cancer | upregulation | "MicroRNAs miRNAs such as miR-17-5p miR-21 miR-106a ......" | 23267156 | |
| hsa-miR-106a-5p | gastric cancer | upregulation | "Multidrug resistance MDR is the main barrier to th ......" | 23932924 | |
| hsa-miR-106a-5p | gastric cancer | upregulation | "Here we report that miR-106a is frequently up-regu ......" | 24440352 | |
| hsa-miR-106a-5p | gastric cancer | upregulation | "Up regulated Circulating miR 106a by DNA Methylati ......" | 26179261 | qPCR |
| hsa-miR-106a-5p | gastric cancer | upregulation | "Diagnostic significance of miR 106a in gastric can ......" | 26722506 | |
| hsa-miR-106a-5p | gastric cancer | upregulation | "In our previous studies we have found that the up- ......" | 27142596 | qPCR |
| hsa-miR-106a-5p | glioblastoma | deregulation | "A number of changes in the levels of microRNAs wer ......" | 24155920 | |
| hsa-miR-106a-5p | kidney renal cell cancer | deregulation | "Exploring the miRNA mRNA regulatory network in cle ......" | 24977165 | RNA-Seq |
| hsa-miR-106a-5p | liver cancer | upregulation | "Upregulated expression of miR 106a by DNA hypometh ......" | 25510666 | |
| hsa-miR-106a-5p | lung squamous cell cancer | upregulation | "In this study we investigated the expression and t ......" | 26097565 | |
| hsa-miR-106a-5p | ovarian cancer | upregulation | "Expression profiling of miRNAs in HGSOC indicated ......" | 24045973 | qPCR; in situ hybridization |
| hsa-miR-106a-5p | pancreatic cancer | upregulation | "Upregulated miR 106a plays an oncogenic role in pa ......" | 24444603 | |
| hsa-miR-106a-5p | prostate cancer | downregulation | "Multiple tumor-suppressive miRNAs were downregulat ......" | 22719071 | |
| hsa-miR-106a-5p | thyroid cancer | upregulation | "Expression levels of six identified miRNAs were as ......" | 27342319 | qPCR |
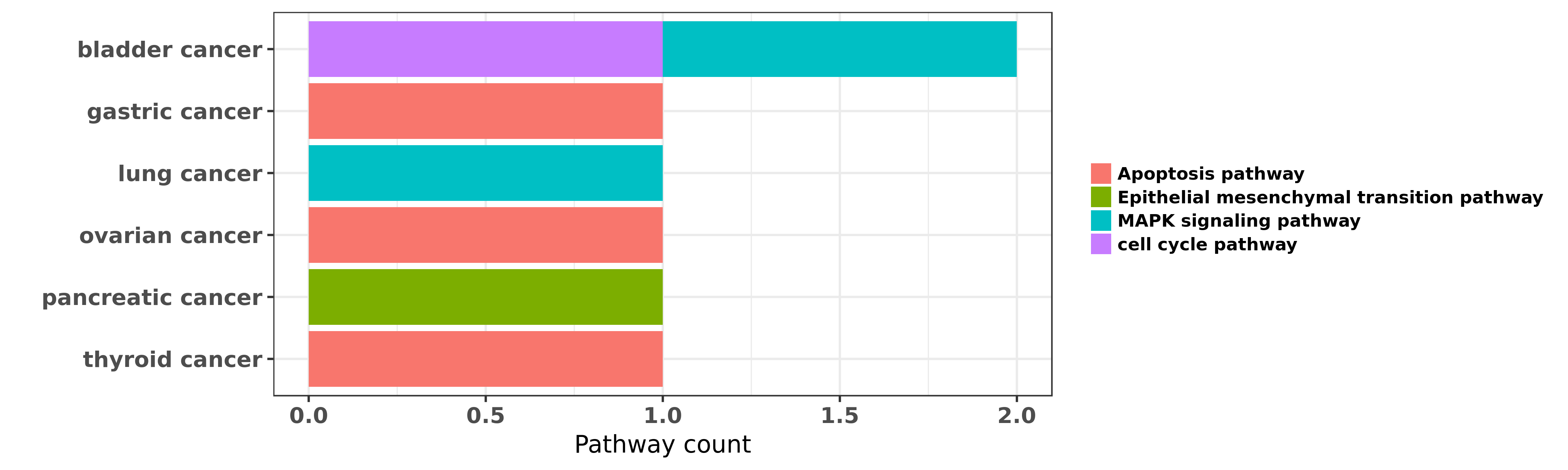
Reported cancer pathway affected by hsa-miR-106a-5p
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-106a-5p | bladder cancer | cell cycle pathway; MAPK signaling pathway | "MicroRNA 106a suppresses proliferation migration a ......" | 27513725 | Flow cytometry |
| hsa-miR-106a-5p | gastric cancer | Apoptosis pathway | "miR 106a is frequently upregulated in gastric canc ......" | 22431000 | |
| hsa-miR-106a-5p | gastric cancer | Apoptosis pathway | "MicroRNA 106a induces multidrug resistance in gast ......" | 23932924 | |
| hsa-miR-106a-5p | gastric cancer | Apoptosis pathway | "MicroRNA 106a functions as an oncogene in human ga ......" | 27142596 | Transwell assay; Western blot; Luciferase |
| hsa-miR-106a-5p | lung cancer | MAPK signaling pathway | "RNA-seq data and miRNA-seq data of lung adenocarci ......" | 27338053 | |
| hsa-miR-106a-5p | ovarian cancer | Apoptosis pathway | "Dysregulation of miR 106a and miR 591 confers pacl ......" | 23807165 | Cell migration assay; Luciferase |
| hsa-miR-106a-5p | ovarian cancer | Apoptosis pathway | "MiR 106a targets Mcl 1 to suppress cisplatin resis ......" | 23904379 | Flow cytometry; MTT assay; Western blot; Luciferase |
| hsa-miR-106a-5p | ovarian cancer | Apoptosis pathway | "microRNA 106a modulates cisplatin sensitivity by t ......" | 24348845 | Luciferase; Western blot |
| hsa-miR-106a-5p | pancreatic cancer | Epithelial mesenchymal transition pathway | "Upregulated miR 106a plays an oncogenic role in pa ......" | 24444603 | |
| hsa-miR-106a-5p | thyroid cancer | Apoptosis pathway | "Expression levels of six identified miRNAs were as ......" | 27342319 | Luciferase; Flow cytometry; MTT assay; Western blot |
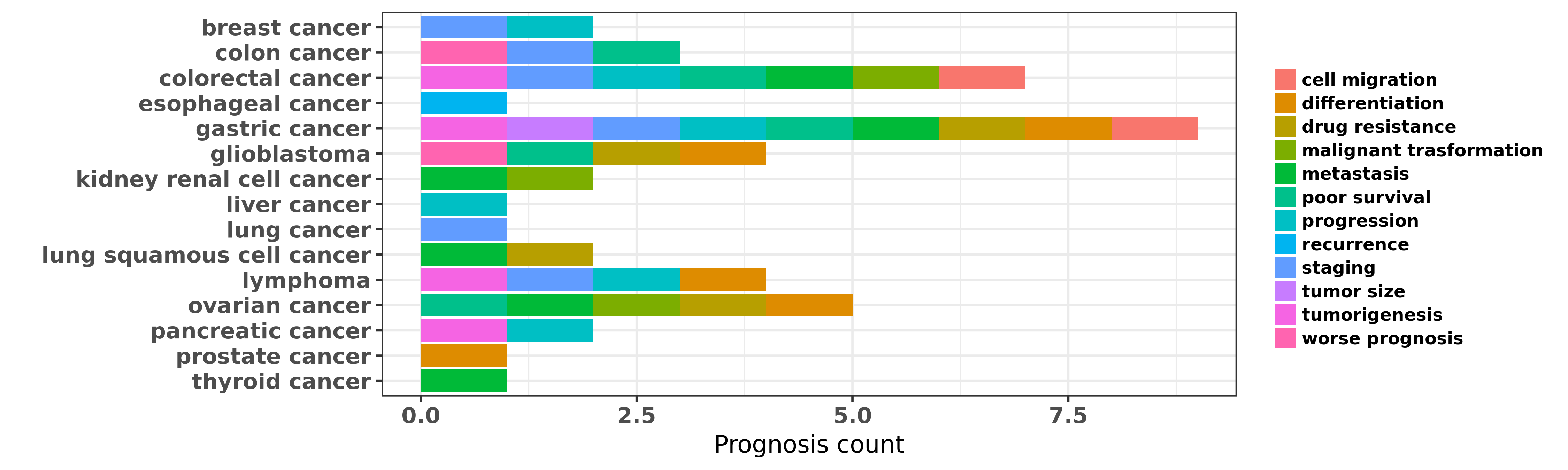
Reported cancer prognosis affected by hsa-miR-106a-5p
| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-106a-5p | breast cancer | progression | "MicroRNA microArray analysis was performed compari ......" | 24626163 | |
| hsa-miR-106a-5p | breast cancer | staging | "Results showed differential pattern of expressions ......" | 26276721 | |
| hsa-miR-106a-5p | colon cancer | poor survival; worse prognosis; staging | "Deregulated expression of miR 106a predicts surviv ......" | 18521848 | |
| hsa-miR-106a-5p | colon cancer | staging | "To present proof-of-principle application for empl ......" | 23741026 | |
| hsa-miR-106a-5p | colon cancer | poor survival; worse prognosis | "Long non coding RNA Fer 1 like protein 4 suppresse ......" | 26224446 | |
| hsa-miR-106a-5p | colorectal cancer | metastasis; cell migration; progression | "Colorectal cancer migration and invasion initiated ......" | 22912877 | |
| hsa-miR-106a-5p | colorectal cancer | tumorigenesis | "miR 106a overexpression and pRB downregulation in ......" | 23178825 | |
| hsa-miR-106a-5p | colorectal cancer | staging | "In the present study we measured the levels of fiv ......" | 23625654 | |
| hsa-miR-106a-5p | colorectal cancer | staging; metastasis | "Expression of plasma miR 106a in colorectal cancer ......" | 24670448 | |
| hsa-miR-106a-5p | colorectal cancer | metastasis | "The expression of miR-106a was found to be upregul ......" | 24746948 | |
| hsa-miR-106a-5p | colorectal cancer | malignant trasformation; staging; poor survival | "Notably deregulation of the key oncogene miR-21 wa ......" | 25788261 | |
| hsa-miR-106a-5p | colorectal cancer | cell migration | "However the role and underlying mechanism of miR-1 ......" | 26223867 | MTT assay; Transwell assay; Wound Healing Assay |
| hsa-miR-106a-5p | colorectal cancer | staging; poor survival | "Using individual RT-qPCR verification in 175 stage ......" | 26250939 | |
| hsa-miR-106a-5p | esophageal cancer | recurrence | "The expression of miR-21 miR-106a miR-148a miR-205 ......" | 20628822 | |
| hsa-miR-106a-5p | gastric cancer | staging; metastasis; differentiation | "Detection of miR 106a in gastric carcinoma and its ......" | 18996365 | |
| hsa-miR-106a-5p | gastric cancer | tumorigenesis | "miR 106a is frequently upregulated in gastric canc ......" | 22431000 | |
| hsa-miR-106a-5p | gastric cancer | metastasis; poor survival | "The expression levels of miR-20a miR-21 miR-25 miR ......" | 22996433 | |
| hsa-miR-106a-5p | gastric cancer | poor survival | "MicroRNAs miRNAs such as miR-17-5p miR-21 miR-106a ......" | 23267156 | |
| hsa-miR-106a-5p | gastric cancer | drug resistance | "MicroRNA 106a induces multidrug resistance in gast ......" | 23932924 | |
| hsa-miR-106a-5p | gastric cancer | drug resistance | "miR 106a confers cisplatin resistance by regulatin ......" | 24108762 | |
| hsa-miR-106a-5p | gastric cancer | metastasis | "MicroRNA 106a targets TIMP2 to regulate invasion a ......" | 24440352 | |
| hsa-miR-106a-5p | gastric cancer | staging; metastasis; progression | "Up regulated Circulating miR 106a by DNA Methylati ......" | 26179261 | |
| hsa-miR-106a-5p | gastric cancer | staging; metastasis; differentiation; tumor size | "Diagnostic significance of miR 106a in gastric can ......" | 26722506 | |
| hsa-miR-106a-5p | gastric cancer | metastasis; progression; cell migration | "MicroRNA 106a functions as an oncogene in human ga ......" | 27142596 | Transwell assay; Western blot; Luciferase |
| hsa-miR-106a-5p | glioblastoma | poor survival; worse prognosis; drug resistance | "MiR 106a is an independent prognostic marker in pa ......" | 23416698 | |
| hsa-miR-106a-5p | glioblastoma | differentiation | "Using this in vitro system a microarray-based high ......" | 24155920 | |
| hsa-miR-106a-5p | kidney renal cell cancer | metastasis; malignant trasformation | "To obtain accurate results in miRNA expression cha ......" | 21741950 | |
| hsa-miR-106a-5p | liver cancer | progression | "Upregulated expression of miR 106a by DNA hypometh ......" | 25510666 | Luciferase |
| hsa-miR-106a-5p | lung cancer | staging | "miRNAs were assessed by quantitative real-time PCR ......" | 19209007 | |
| hsa-miR-106a-5p | lung squamous cell cancer | metastasis | "The miRs were quantified by microarray hybridizati ......" | 22295063 | |
| hsa-miR-106a-5p | lung squamous cell cancer | drug resistance | "MicroRNA 106a confers cisplatin resistance in non ......" | 25339370 | |
| hsa-miR-106a-5p | lung squamous cell cancer | metastasis | "miR 106a promotes growth and metastasis of non sma ......" | 26097565 | |
| hsa-miR-106a-5p | lymphoma | tumorigenesis | "Retroviral activation of the mir 106a microRNA cis ......" | 17442096 | |
| hsa-miR-106a-5p | lymphoma | differentiation | "We also used microarray analysis to identify a dif ......" | 21698185 | |
| hsa-miR-106a-5p | lymphoma | staging; progression | "Herein we used a global quantitative real-time pol ......" | 25503151 | |
| hsa-miR-106a-5p | ovarian cancer | drug resistance | "Dysregulation of miR 106a and miR 591 confers pacl ......" | 23807165 | Cell migration assay; Luciferase |
| hsa-miR-106a-5p | ovarian cancer | drug resistance; poor survival | "MiR 106a targets Mcl 1 to suppress cisplatin resis ......" | 23904379 | Flow cytometry; MTT assay; Western blot; Luciferase |
| hsa-miR-106a-5p | ovarian cancer | differentiation; malignant trasformation | "miR 106a represses the Rb tumor suppressor p130 to ......" | 24045973 | |
| hsa-miR-106a-5p | ovarian cancer | drug resistance | "microRNA 106a modulates cisplatin sensitivity by t ......" | 24348845 | Luciferase; Western blot |
| hsa-miR-106a-5p | ovarian cancer | metastasis | "MicroRNA 106a regulates phosphatase and tensin hom ......" | 27510094 | |
| hsa-miR-106a-5p | pancreatic cancer | progression; tumorigenesis | "Upregulated miR 106a plays an oncogenic role in pa ......" | 24444603 | |
| hsa-miR-106a-5p | prostate cancer | differentiation | "We demonstrate hypoxia promotes neuronal and neuro ......" | 24135225 | |
| hsa-miR-106a-5p | thyroid cancer | metastasis | "Expression levels of six identified miRNAs were as ......" | 27342319 | Luciferase; Flow cytometry; MTT assay; Western blot |
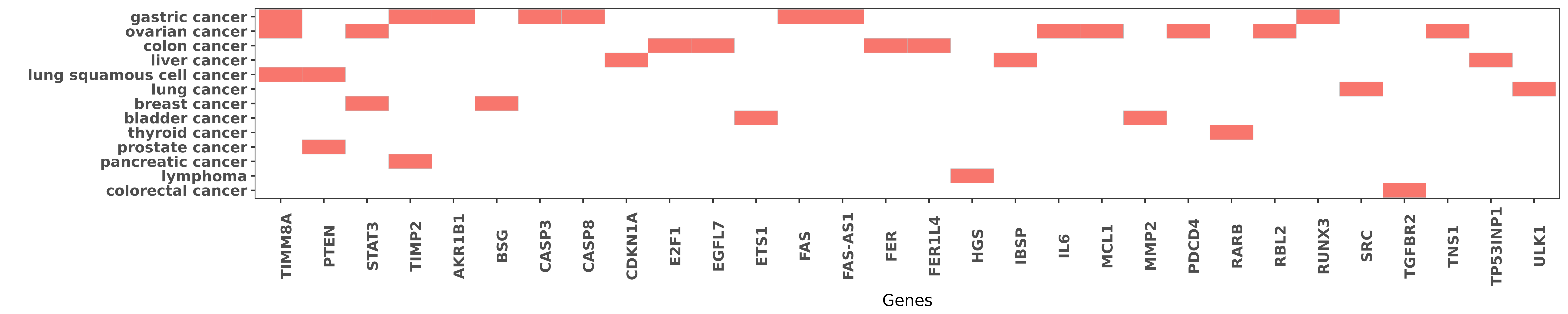
Reported gene related to hsa-miR-106a-5p
| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-miR-106a-5p | gastric cancer | TIMM8A | "In this study we investigated whether miR-106a med ......" | 24108762 |
| hsa-miR-106a-5p | lung squamous cell cancer | TIMM8A | "Fewer studies have explored the roles of miR-106a ......" | 25339370 |
| hsa-miR-106a-5p | ovarian cancer | TIMM8A | "This study was to investigate whether miR-106a med ......" | 23904379 |
| hsa-miR-106a-5p | gastric cancer | TIMP2 | "MicroRNA 106a targets TIMP2 to regulate invasion a ......" | 24440352 |
| hsa-miR-106a-5p | gastric cancer | TIMP2 | "Moreover expression of TIMP2 was inversely associa ......" | 27142596 |
| hsa-miR-106a-5p | pancreatic cancer | TIMP2 | "Furthermore our investigation shows that miR-106a ......" | 24444603 |
| hsa-miR-106a-5p | gastric cancer | FAS | "miR 106a is frequently upregulated in gastric canc ......" | 22431000 |
| hsa-miR-106a-5p | gastric cancer | FAS | "Functional experiment ascertained that miR-106a in ......" | 27142596 |
| hsa-miR-106a-5p | ovarian cancer | PDCD4 | "microRNA 106a modulates cisplatin sensitivity by t ......" | 24348845 |
| hsa-miR-106a-5p | ovarian cancer | PDCD4 | "Tumor volume and weight as well as Ki-67 and progr ......" | 27393101 |
| hsa-miR-106a-5p | lung squamous cell cancer | PTEN | "miR 106a promotes growth and metastasis of non sma ......" | 26097565 |
| hsa-miR-106a-5p | prostate cancer | PTEN | "Further pterostilbene through downregulation of mi ......" | 26318586 |
| hsa-miR-106a-5p | breast cancer | STAT3 | "We also found that EMMPRIN could down-regulate miR ......" | 27325313 |
| hsa-miR-106a-5p | ovarian cancer | STAT3 | "Furthermore the present study revealed that IL-6 i ......" | 27510094 |
| hsa-miR-106a-5p | gastric cancer | AKR1B1 | "miR-106a promotes chemo-resistance of GC cells acc ......" | 23932924 |
| hsa-miR-106a-5p | breast cancer | BSG | "We also found that EMMPRIN could down-regulate miR ......" | 27325313 |
| hsa-miR-106a-5p | gastric cancer | CASP3 | "Rescue experiments and examination of caspase-8 PA ......" | 22431000 |
| hsa-miR-106a-5p | gastric cancer | CASP8 | "Rescue experiments and examination of caspase-8 PA ......" | 22431000 |
| hsa-miR-106a-5p | liver cancer | CDKN1A | "After prediction with online software we further u ......" | 25510666 |
| hsa-miR-106a-5p | colon cancer | E2F1 | "Inverse correlations were found between miR-17-5p ......" | 18521848 |
| hsa-miR-106a-5p | colon cancer | EGFL7 | "Altered expression of miR-17-5p miR-106a and EGFL7 ......" | 18521848 |
| hsa-miR-106a-5p | bladder cancer | ETS1 | "MicroRNA 106a suppresses proliferation migration a ......" | 27513725 |
| hsa-miR-106a-5p | gastric cancer | FAS-AS1 | "Bioinformatic analysis combining with validation e ......" | 22431000 |
| hsa-miR-106a-5p | colon cancer | FER | "Long non coding RNA Fer 1 like protein 4 suppresse ......" | 26224446 |
| hsa-miR-106a-5p | colon cancer | FER1L4 | "The results showed the aberrant expression of FER1 ......" | 26224446 |
| hsa-miR-106a-5p | lymphoma | HGS | "Expression of miR-17 and miR-106a was detected in ......" | 21953646 |
| hsa-miR-106a-5p | liver cancer | IBSP | "Then we used quantitative real-time PCR qPCR and B ......" | 25510666 |
| hsa-miR-106a-5p | ovarian cancer | IL6 | "Furthermore the present study revealed that IL-6 i ......" | 27510094 |
| hsa-miR-106a-5p | ovarian cancer | MCL1 | "MiR 106a targets Mcl 1 to suppress cisplatin resis ......" | 23904379 |
| hsa-miR-106a-5p | bladder cancer | MMP2 | "MicroRNA 106a suppresses proliferation migration a ......" | 27513725 |
| hsa-miR-106a-5p | thyroid cancer | RARB | "The results of dual-luciferase reporter assay demo ......" | 27342319 |
| hsa-miR-106a-5p | ovarian cancer | RBL2 | "Among many miR-106a predicated target genes p130 R ......" | 24045973 |
| hsa-miR-106a-5p | gastric cancer | RUNX3 | "MicroRNA 106a induces multidrug resistance in gast ......" | 23932924 |
| hsa-miR-106a-5p | lung cancer | SRC | "MicroRNA 106a targets autophagy and enhances sensi ......" | 27372519 |
| hsa-miR-106a-5p | colorectal cancer | TGFBR2 | "MiR-106a inhibits the expression of transforming g ......" | 22912877 |
| hsa-miR-106a-5p | ovarian cancer | TNS1 | "MicroRNA 106a regulates phosphatase and tensin hom ......" | 27510094 |
| hsa-miR-106a-5p | liver cancer | TP53INP1 | "After prediction with online software we further u ......" | 25510666 |
| hsa-miR-106a-5p | lung cancer | ULK1 | "This ULK1 upregulation is caused by downregulation ......" | 27372519 |
Expression profile in cancer corhorts:
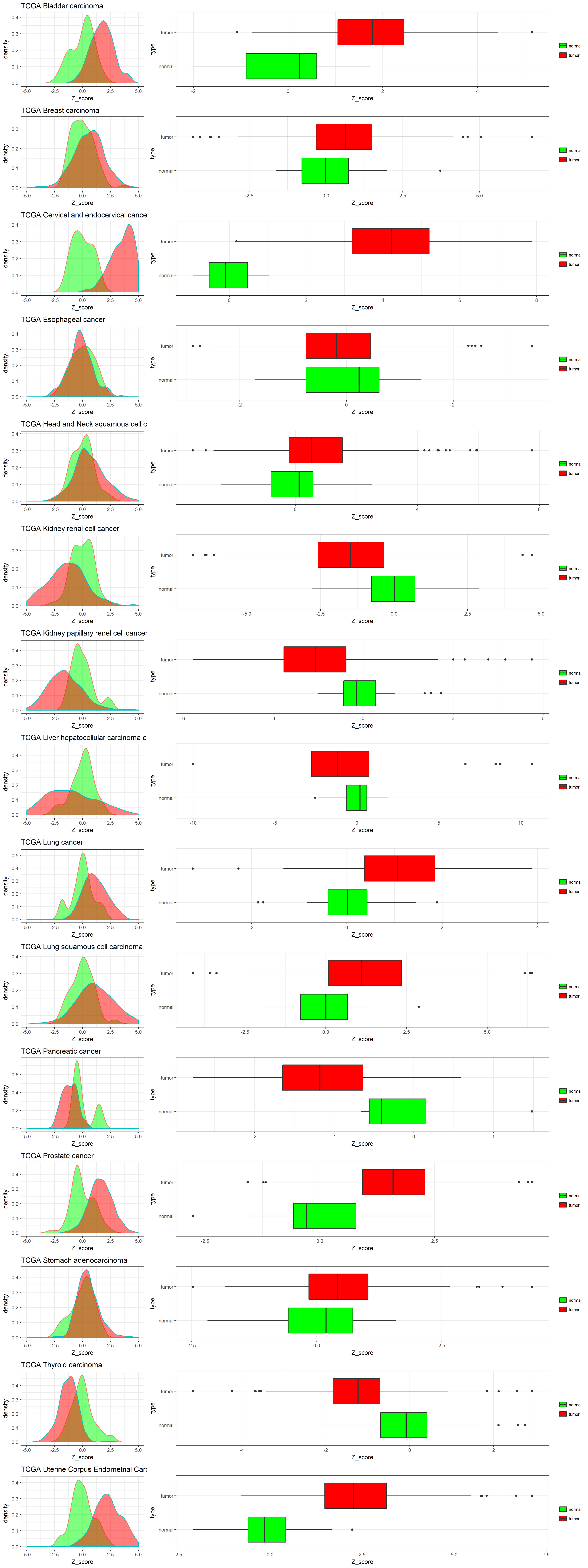
Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-106a-5p | CDKN1A | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; THCA; STAD | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | TCGA BLCA -0.129; TCGA BRCA -0.115; TCGA CESC -0.107; TCGA COAD -0.131; TCGA ESCA -0.15; TCGA HNSC -0.128; TCGA KIRC -0.219; TCGA KIRP -0.192; TCGA LUAD -0.063; TCGA LUSC -0.283; TCGA PAAD -0.132; TCGA THCA -0.212; TCGA STAD -0.174 |
hsa-miR-106a-5p | FAS | 12 cancers: BLCA; CESC; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; LUSC; PRAD; THCA; UCEC | miRNAWalker2 validate; miRTarBase | TCGA BLCA -0.286; TCGA CESC -0.14; TCGA COAD -0.309; TCGA ESCA -0.155; TCGA KIRC -0.176; TCGA KIRP -0.255; TCGA LIHC -0.303; TCGA LUAD -0.275; TCGA LUSC -0.24; TCGA PRAD -0.225; TCGA THCA -0.328; TCGA UCEC -0.19 |
hsa-miR-106a-5p | HIPK3 | 9 cancers: BLCA; COAD; HNSC; LGG; LUAD; LUSC; PRAD; STAD; UCEC | miRNAWalker2 validate; miRTarBase | TCGA BLCA -0.165; TCGA COAD -0.188; TCGA HNSC -0.162; TCGA LGG -0.172; TCGA LUAD -0.21; TCGA LUSC -0.156; TCGA PRAD -0.44; TCGA STAD -0.135; TCGA UCEC -0.189 |
hsa-miR-106a-5p | NOTCH2 | 13 cancers: BLCA; COAD; ESCA; HNSC; KIRP; LGG; LUAD; LUSC; PRAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.066; TCGA COAD -0.198; TCGA ESCA -0.135; TCGA HNSC -0.078; TCGA KIRP -0.215; TCGA LGG -0.204; TCGA LUAD -0.111; TCGA LUSC -0.143; TCGA PRAD -0.14; TCGA SARC -0.069; TCGA THCA -0.181; TCGA STAD -0.14; TCGA UCEC -0.052 |
hsa-miR-106a-5p | RB1 | 9 cancers: BLCA; BRCA; KIRC; KIRP; LUAD; LUSC; OV; PRAD; UCEC | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | TCGA BLCA -0.207; TCGA BRCA -0.077; TCGA KIRC -0.066; TCGA KIRP -0.069; TCGA LUAD -0.115; TCGA LUSC -0.07; TCGA OV -0.091; TCGA PRAD -0.195; TCGA UCEC -0.137 |
hsa-miR-106a-5p | RUNX1 | 9 cancers: BLCA; BRCA; COAD; KIRC; KIRP; LUAD; LUSC; PRAD; THCA | miRNAWalker2 validate; miRTarBase; miRNATAP | TCGA BLCA -0.054; TCGA BRCA -0.063; TCGA COAD -0.165; TCGA KIRC -0.303; TCGA KIRP -0.205; TCGA LUAD -0.103; TCGA LUSC -0.124; TCGA PRAD -0.114; TCGA THCA -0.713 |
hsa-miR-106a-5p | SIRPA | 12 cancers: BLCA; CESC; COAD; HNSC; KIRP; LGG; LUAD; LUSC; PRAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.267; TCGA CESC -0.295; TCGA COAD -0.2; TCGA HNSC -0.253; TCGA KIRP -0.105; TCGA LGG -0.08; TCGA LUAD -0.173; TCGA LUSC -0.357; TCGA PRAD -0.139; TCGA THCA -0.185; TCGA STAD -0.101; TCGA UCEC -0.101 |
hsa-miR-106a-5p | ZBTB47 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LUAD; LUSC; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate; MirTarget | TCGA BLCA -0.15; TCGA BRCA -0.136; TCGA CESC -0.128; TCGA COAD -0.334; TCGA ESCA -0.13; TCGA HNSC -0.126; TCGA LGG -0.089; TCGA LUAD -0.116; TCGA LUSC -0.185; TCGA OV -0.107; TCGA PRAD -0.101; TCGA SARC -0.138; TCGA STAD -0.247; TCGA UCEC -0.119 |
hsa-miR-106a-5p | TGFBR2 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | miRTarBase; miRNATAP | TCGA BLCA -0.174; TCGA BRCA -0.167; TCGA CESC -0.15; TCGA COAD -0.212; TCGA ESCA -0.226; TCGA HNSC -0.255; TCGA LIHC -0.077; TCGA LUAD -0.307; TCGA LUSC -0.514; TCGA OV -0.065; TCGA PRAD -0.258; TCGA STAD -0.138; TCGA UCEC -0.237 |
hsa-miR-106a-5p | FBXO31 | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LIHC; LUSC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.12; TCGA BRCA -0.072; TCGA CESC -0.063; TCGA ESCA -0.072; TCGA HNSC -0.064; TCGA LIHC -0.336; TCGA LUSC -0.053; TCGA THCA -0.106; TCGA STAD -0.095; TCGA UCEC -0.088 |
hsa-miR-106a-5p | ABCD2 | 9 cancers: BLCA; CESC; COAD; ESCA; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.352; TCGA CESC -0.232; TCGA COAD -0.429; TCGA ESCA -0.216; TCGA LUAD -0.153; TCGA LUSC -0.259; TCGA PRAD -0.418; TCGA STAD -0.213; TCGA UCEC -0.303 |
hsa-miR-106a-5p | CRIM1 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.108; TCGA BRCA -0.12; TCGA CESC -0.144; TCGA COAD -0.118; TCGA ESCA -0.126; TCGA HNSC -0.176; TCGA LIHC -0.138; TCGA LUAD -0.166; TCGA LUSC -0.173; TCGA OV -0.066; TCGA PRAD -0.271; TCGA STAD -0.138; TCGA UCEC -0.201 |
hsa-miR-106a-5p | TNKS1BP1 | 13 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; LIHC; LUAD; LUSC; PAAD; SARC; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.092; TCGA CESC -0.141; TCGA COAD -0.104; TCGA ESCA -0.116; TCGA HNSC -0.161; TCGA KIRC -0.087; TCGA LIHC -0.057; TCGA LUAD -0.052; TCGA LUSC -0.148; TCGA PAAD -0.142; TCGA SARC -0.076; TCGA THCA -0.143; TCGA STAD -0.109 |
hsa-miR-106a-5p | TOLLIP | 9 cancers: BLCA; BRCA; CESC; HNSC; LIHC; LUAD; LUSC; THCA; STAD | MirTarget | TCGA BLCA -0.064; TCGA BRCA -0.075; TCGA CESC -0.116; TCGA HNSC -0.107; TCGA LIHC -0.064; TCGA LUAD -0.059; TCGA LUSC -0.106; TCGA THCA -0.056; TCGA STAD -0.078 |
hsa-miR-106a-5p | PIK3R1 | 9 cancers: BLCA; BRCA; LGG; LIHC; LUAD; LUSC; PRAD; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.068; TCGA BRCA -0.061; TCGA LGG -0.142; TCGA LIHC -0.098; TCGA LUAD -0.153; TCGA LUSC -0.103; TCGA PRAD -0.233; TCGA THCA -0.073; TCGA STAD -0.13 |
hsa-miR-106a-5p | PKD2 | 10 cancers: BLCA; BRCA; CESC; COAD; KIRP; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.147; TCGA BRCA -0.103; TCGA CESC -0.07; TCGA COAD -0.38; TCGA KIRP -0.134; TCGA LUAD -0.101; TCGA LUSC -0.088; TCGA PRAD -0.261; TCGA STAD -0.221; TCGA UCEC -0.213 |
hsa-miR-106a-5p | CORO2B | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.197; TCGA BRCA -0.224; TCGA CESC -0.333; TCGA COAD -0.536; TCGA HNSC -0.262; TCGA LUAD -0.231; TCGA LUSC -0.43; TCGA PRAD -0.289; TCGA STAD -0.31; TCGA UCEC -0.186 |
hsa-miR-106a-5p | FGD5 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.212; TCGA BRCA -0.115; TCGA CESC -0.246; TCGA COAD -0.304; TCGA HNSC -0.136; TCGA KIRC -0.168; TCGA LUAD -0.198; TCGA LUSC -0.389; TCGA PAAD -0.143; TCGA PRAD -0.22; TCGA STAD -0.206; TCGA UCEC -0.139 |
hsa-miR-106a-5p | CYBRD1 | 13 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; OV; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.407; TCGA BRCA -0.234; TCGA CESC -0.127; TCGA COAD -0.695; TCGA HNSC -0.197; TCGA LGG -0.135; TCGA LUAD -0.349; TCGA LUSC -0.33; TCGA OV -0.214; TCGA PRAD -0.298; TCGA SARC -0.184; TCGA STAD -0.426; TCGA UCEC -0.281 |
hsa-miR-106a-5p | CHD9 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.111; TCGA BRCA -0.085; TCGA CESC -0.079; TCGA COAD -0.162; TCGA HNSC -0.097; TCGA LGG -0.116; TCGA LIHC -0.126; TCGA LUAD -0.128; TCGA LUSC -0.095; TCGA PRAD -0.221; TCGA STAD -0.09; TCGA UCEC -0.119 |
hsa-miR-106a-5p | JAK1 | 14 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRP; LGG; LIHC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.133; TCGA CESC -0.082; TCGA COAD -0.1; TCGA ESCA -0.102; TCGA HNSC -0.101; TCGA KIRP -0.151; TCGA LGG -0.125; TCGA LIHC -0.083; TCGA LUAD -0.11; TCGA LUSC -0.199; TCGA PRAD -0.182; TCGA THCA -0.116; TCGA STAD -0.06; TCGA UCEC -0.133 |
hsa-miR-106a-5p | FYCO1 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.188; TCGA BRCA -0.102; TCGA CESC -0.066; TCGA COAD -0.198; TCGA ESCA -0.19; TCGA HNSC -0.152; TCGA KIRP -0.092; TCGA LIHC -0.114; TCGA LUAD -0.166; TCGA LUSC -0.188; TCGA PRAD -0.231; TCGA STAD -0.241; TCGA UCEC -0.203 |
hsa-miR-106a-5p | ITPRIPL2 | 13 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.12; TCGA CESC -0.076; TCGA COAD -0.071; TCGA ESCA -0.078; TCGA HNSC -0.056; TCGA KIRC -0.061; TCGA KIRP -0.067; TCGA LUAD -0.081; TCGA LUSC -0.145; TCGA PRAD -0.088; TCGA THCA -0.246; TCGA STAD -0.094; TCGA UCEC -0.096 |
hsa-miR-106a-5p | IL1RAP | 10 cancers: BLCA; CESC; COAD; HNSC; KIRC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.184; TCGA CESC -0.218; TCGA COAD -0.191; TCGA HNSC -0.121; TCGA KIRC -0.123; TCGA LUAD -0.092; TCGA LUSC -0.186; TCGA PRAD -0.195; TCGA THCA -0.661; TCGA UCEC -0.114 |
hsa-miR-106a-5p | STOM | 12 cancers: BLCA; BRCA; CESC; COAD; KIRC; KIRP; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.318; TCGA BRCA -0.076; TCGA CESC -0.113; TCGA COAD -0.419; TCGA KIRC -0.113; TCGA KIRP -0.095; TCGA LUAD -0.15; TCGA LUSC -0.193; TCGA PRAD -0.228; TCGA THCA -0.128; TCGA STAD -0.228; TCGA UCEC -0.186 |
hsa-miR-106a-5p | MAP3K3 | 9 cancers: BLCA; COAD; ESCA; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.099; TCGA COAD -0.211; TCGA ESCA -0.114; TCGA LUAD -0.115; TCGA LUSC -0.135; TCGA PRAD -0.067; TCGA THCA -0.057; TCGA STAD -0.191; TCGA UCEC -0.129 |
hsa-miR-106a-5p | PRRX1 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LUAD; LUSC; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.484; TCGA BRCA -0.075; TCGA CESC -0.342; TCGA COAD -0.789; TCGA ESCA -0.212; TCGA HNSC -0.15; TCGA LGG -0.24; TCGA LUAD -0.147; TCGA LUSC -0.28; TCGA PRAD -0.273; TCGA SARC -0.196; TCGA STAD -0.283; TCGA UCEC -0.106 |
hsa-miR-106a-5p | PLSCR4 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.286; TCGA BRCA -0.178; TCGA CESC -0.139; TCGA COAD -0.301; TCGA ESCA -0.186; TCGA HNSC -0.076; TCGA LUAD -0.193; TCGA LUSC -0.217; TCGA OV -0.186; TCGA PRAD -0.315; TCGA STAD -0.37; TCGA UCEC -0.334 |
hsa-miR-106a-5p | TGM2 | 12 cancers: BLCA; CESC; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.445; TCGA CESC -0.302; TCGA ESCA -0.263; TCGA HNSC -0.116; TCGA KIRC -0.316; TCGA KIRP -0.3; TCGA LUAD -0.095; TCGA LUSC -0.562; TCGA PRAD -0.074; TCGA THCA -0.554; TCGA STAD -0.177; TCGA UCEC -0.336 |
hsa-miR-106a-5p | MAP3K8 | 9 cancers: BLCA; BRCA; CESC; ESCA; KIRP; LIHC; LUSC; PRAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.07; TCGA BRCA -0.076; TCGA CESC -0.158; TCGA ESCA -0.136; TCGA KIRP -0.121; TCGA LIHC -0.154; TCGA LUSC -0.207; TCGA PRAD -0.207; TCGA UCEC -0.094 |
hsa-miR-106a-5p | UBE2Q2 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUSC; OV; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.11; TCGA BRCA -0.052; TCGA CESC -0.083; TCGA COAD -0.186; TCGA ESCA -0.135; TCGA HNSC -0.155; TCGA KIRP -0.07; TCGA LUSC -0.085; TCGA OV -0.075; TCGA PRAD -0.088; TCGA SARC -0.057; TCGA STAD -0.112; TCGA UCEC -0.064 |
hsa-miR-106a-5p | OSTM1 | 11 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.184; TCGA CESC -0.112; TCGA COAD -0.13; TCGA ESCA -0.086; TCGA HNSC -0.122; TCGA LUAD -0.077; TCGA LUSC -0.197; TCGA OV -0.085; TCGA PRAD -0.134; TCGA STAD -0.079; TCGA UCEC -0.118 |
hsa-miR-106a-5p | ABCA1 | 13 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; KIRP; LGG; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.125; TCGA BRCA -0.055; TCGA CESC -0.228; TCGA COAD -0.344; TCGA HNSC -0.155; TCGA KIRC -0.122; TCGA KIRP -0.159; TCGA LGG -0.344; TCGA LUAD -0.144; TCGA LUSC -0.176; TCGA PRAD -0.171; TCGA STAD -0.111; TCGA UCEC -0.078 |
hsa-miR-106a-5p | ELK3 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.184; TCGA BRCA -0.102; TCGA CESC -0.288; TCGA COAD -0.347; TCGA ESCA -0.151; TCGA HNSC -0.195; TCGA KIRC -0.168; TCGA LUAD -0.212; TCGA LUSC -0.296; TCGA PRAD -0.323; TCGA THCA -0.173; TCGA STAD -0.214; TCGA UCEC -0.224 |
hsa-miR-106a-5p | DPYD | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.473; TCGA BRCA -0.064; TCGA CESC -0.188; TCGA COAD -0.632; TCGA ESCA -0.206; TCGA HNSC -0.088; TCGA LUAD -0.284; TCGA LUSC -0.46; TCGA PRAD -0.315; TCGA THCA -0.163; TCGA STAD -0.266; TCGA UCEC -0.29 |
hsa-miR-106a-5p | ADARB1 | 9 cancers: BLCA; CESC; COAD; HNSC; KIRC; LUAD; LUSC; STAD; UCEC | MirTarget | TCGA BLCA -0.282; TCGA CESC -0.124; TCGA COAD -0.221; TCGA HNSC -0.128; TCGA KIRC -0.106; TCGA LUAD -0.131; TCGA LUSC -0.313; TCGA STAD -0.213; TCGA UCEC -0.115 |
hsa-miR-106a-5p | CYP2U1 | 9 cancers: BLCA; BRCA; COAD; LGG; LIHC; LUAD; LUSC; STAD; UCEC | MirTarget | TCGA BLCA -0.074; TCGA BRCA -0.09; TCGA COAD -0.275; TCGA LGG -0.066; TCGA LIHC -0.103; TCGA LUAD -0.153; TCGA LUSC -0.098; TCGA STAD -0.246; TCGA UCEC -0.059 |
hsa-miR-106a-5p | RORA | 11 cancers: BLCA; BRCA; COAD; HNSC; KIRC; KIRP; LIHC; LUAD; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.116; TCGA BRCA -0.073; TCGA COAD -0.388; TCGA HNSC -0.075; TCGA KIRC -0.115; TCGA KIRP -0.14; TCGA LIHC -0.198; TCGA LUAD -0.093; TCGA PRAD -0.183; TCGA STAD -0.232; TCGA UCEC -0.085 |
hsa-miR-106a-5p | CLIP4 | 9 cancers: BLCA; CESC; COAD; HNSC; LUAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.303; TCGA CESC -0.302; TCGA COAD -0.718; TCGA HNSC -0.193; TCGA LUAD -0.101; TCGA PRAD -0.307; TCGA THCA -0.139; TCGA STAD -0.359; TCGA UCEC -0.21 |
hsa-miR-106a-5p | LAPTM4A | 15 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.071; TCGA CESC -0.099; TCGA COAD -0.064; TCGA ESCA -0.062; TCGA HNSC -0.065; TCGA KIRP -0.115; TCGA LIHC -0.101; TCGA LUAD -0.077; TCGA LUSC -0.06; TCGA OV -0.073; TCGA PAAD -0.096; TCGA PRAD -0.099; TCGA SARC -0.095; TCGA STAD -0.07; TCGA UCEC -0.097 |
hsa-miR-106a-5p | ZBTB4 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LIHC; LUAD; LUSC; OV; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.127; TCGA BRCA -0.125; TCGA CESC -0.13; TCGA COAD -0.275; TCGA ESCA -0.115; TCGA HNSC -0.113; TCGA KIRP -0.06; TCGA LIHC -0.06; TCGA LUAD -0.152; TCGA LUSC -0.192; TCGA OV -0.119; TCGA PRAD -0.139; TCGA SARC -0.106; TCGA STAD -0.272; TCGA UCEC -0.187 |
hsa-miR-106a-5p | ITFG1 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LIHC; LUAD; LUSC; OV; PRAD; SARC; UCEC | MirTarget | TCGA BLCA -0.099; TCGA BRCA -0.077; TCGA CESC -0.05; TCGA COAD -0.074; TCGA HNSC -0.098; TCGA LIHC -0.143; TCGA LUAD -0.081; TCGA LUSC -0.09; TCGA OV -0.125; TCGA PRAD -0.113; TCGA SARC -0.053; TCGA UCEC -0.076 |
hsa-miR-106a-5p | EDA2R | 14 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.244; TCGA BRCA -0.287; TCGA CESC -0.39; TCGA COAD -0.349; TCGA HNSC -0.272; TCGA KIRC -0.372; TCGA KIRP -0.225; TCGA LIHC -0.225; TCGA LUAD -0.324; TCGA LUSC -0.377; TCGA PRAD -0.335; TCGA THCA -0.275; TCGA STAD -0.196; TCGA UCEC -0.337 |
hsa-miR-106a-5p | RGL1 | 10 cancers: BLCA; BRCA; COAD; HNSC; LGG; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.148; TCGA BRCA -0.059; TCGA COAD -0.271; TCGA HNSC -0.061; TCGA LGG -0.057; TCGA LUAD -0.151; TCGA LUSC -0.183; TCGA PRAD -0.251; TCGA STAD -0.216; TCGA UCEC -0.062 |
hsa-miR-106a-5p | TXNIP | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; LGG; LUAD; LUSC; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.177; TCGA BRCA -0.104; TCGA CESC -0.127; TCGA COAD -0.365; TCGA ESCA -0.253; TCGA LGG -0.149; TCGA LUAD -0.287; TCGA LUSC -0.282; TCGA PRAD -0.147; TCGA SARC -0.114; TCGA STAD -0.373; TCGA UCEC -0.272 |
hsa-miR-106a-5p | ZNFX1 | 15 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LGG; LIHC; LUAD; LUSC; OV; PRAD; SARC; THCA | MirTarget; miRNATAP | TCGA BLCA -0.083; TCGA CESC -0.1; TCGA COAD -0.088; TCGA ESCA -0.121; TCGA HNSC -0.128; TCGA KIRC -0.076; TCGA KIRP -0.061; TCGA LGG -0.076; TCGA LIHC -0.162; TCGA LUAD -0.067; TCGA LUSC -0.181; TCGA OV -0.092; TCGA PRAD -0.109; TCGA SARC -0.076; TCGA THCA -0.069 |
hsa-miR-106a-5p | SEMA4B | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LGG; PAAD; THCA | MirTarget | TCGA BLCA -0.113; TCGA BRCA -0.071; TCGA CESC -0.16; TCGA ESCA -0.16; TCGA HNSC -0.058; TCGA KIRC -0.101; TCGA KIRP -0.114; TCGA LGG -0.175; TCGA PAAD -0.17; TCGA THCA -0.106 |
hsa-miR-106a-5p | OSR1 | 10 cancers: BLCA; COAD; ESCA; LIHC; LUAD; LUSC; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.307; TCGA COAD -0.836; TCGA ESCA -0.309; TCGA LIHC -0.388; TCGA LUAD -0.126; TCGA LUSC -0.183; TCGA PRAD -0.175; TCGA SARC -0.218; TCGA STAD -0.397; TCGA UCEC -0.127 |
hsa-miR-106a-5p | TSPAN9 | 13 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; LUAD; LUSC; OV; PAAD; SARC; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.073; TCGA BRCA -0.069; TCGA CESC -0.082; TCGA COAD -0.174; TCGA HNSC -0.076; TCGA KIRC -0.081; TCGA LUAD -0.106; TCGA LUSC -0.164; TCGA OV -0.071; TCGA PAAD -0.116; TCGA SARC -0.089; TCGA THCA -0.11; TCGA STAD -0.16 |
hsa-miR-106a-5p | ANO6 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.06; TCGA BRCA -0.057; TCGA CESC -0.124; TCGA COAD -0.112; TCGA ESCA -0.095; TCGA HNSC -0.144; TCGA LUAD -0.133; TCGA LUSC -0.187; TCGA PRAD -0.314; TCGA THCA -0.249; TCGA UCEC -0.168 |
hsa-miR-106a-5p | PTHLH | 9 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.55; TCGA BRCA -0.158; TCGA CESC -0.417; TCGA COAD -0.731; TCGA HNSC -0.436; TCGA KIRC -0.721; TCGA PRAD -0.206; TCGA STAD -0.397; TCGA UCEC -0.094 |
hsa-miR-106a-5p | ARHGAP1 | 14 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRP; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.167; TCGA CESC -0.105; TCGA COAD -0.143; TCGA ESCA -0.154; TCGA HNSC -0.119; TCGA KIRP -0.071; TCGA LUAD -0.057; TCGA LUSC -0.178; TCGA PAAD -0.067; TCGA PRAD -0.081; TCGA SARC -0.078; TCGA THCA -0.156; TCGA STAD -0.124; TCGA UCEC -0.088 |
hsa-miR-106a-5p | DCUN1D3 | 9 cancers: BLCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.061; TCGA CESC -0.103; TCGA COAD -0.082; TCGA HNSC -0.142; TCGA LUAD -0.117; TCGA LUSC -0.172; TCGA PRAD -0.09; TCGA THCA -0.179; TCGA UCEC -0.05 |
hsa-miR-106a-5p | ATP1A2 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; STAD | MirTarget | TCGA BLCA -0.8; TCGA BRCA -0.602; TCGA CESC -0.278; TCGA COAD -1.257; TCGA ESCA -0.679; TCGA HNSC -0.48; TCGA LIHC -0.24; TCGA LUAD -0.42; TCGA LUSC -0.443; TCGA PRAD -0.407; TCGA STAD -1.108 |
hsa-miR-106a-5p | FIBIN | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUAD; LUSC; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.623; TCGA BRCA -0.195; TCGA CESC -0.421; TCGA COAD -0.666; TCGA ESCA -0.3; TCGA HNSC -0.527; TCGA KIRP -0.248; TCGA LUAD -0.27; TCGA LUSC -0.505; TCGA PRAD -0.175; TCGA SARC -0.431; TCGA STAD -0.327; TCGA UCEC -0.216 |
hsa-miR-106a-5p | SGMS1 | 10 cancers: BLCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; SARC; THCA; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.084; TCGA CESC -0.114; TCGA COAD -0.147; TCGA HNSC -0.096; TCGA LUAD -0.096; TCGA LUSC -0.136; TCGA PRAD -0.078; TCGA SARC -0.062; TCGA THCA -0.202; TCGA UCEC -0.122 |
hsa-miR-106a-5p | MCC | 9 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; STAD | MirTarget | TCGA BLCA -0.207; TCGA BRCA -0.128; TCGA CESC -0.326; TCGA COAD -0.533; TCGA HNSC -0.155; TCGA LUAD -0.214; TCGA LUSC -0.139; TCGA PRAD -0.378; TCGA STAD -0.344 |
hsa-miR-106a-5p | APBB2 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PAAD; PRAD | MirTarget; miRNATAP | TCGA BLCA -0.155; TCGA BRCA -0.149; TCGA CESC -0.098; TCGA COAD -0.117; TCGA ESCA -0.11; TCGA HNSC -0.208; TCGA LIHC -0.136; TCGA LUAD -0.093; TCGA LUSC -0.238; TCGA PAAD -0.104; TCGA PRAD -0.077 |
hsa-miR-106a-5p | TSHZ3 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.462; TCGA BRCA -0.184; TCGA CESC -0.127; TCGA COAD -0.581; TCGA ESCA -0.293; TCGA HNSC -0.288; TCGA LUAD -0.173; TCGA LUSC -0.317; TCGA PRAD -0.247; TCGA STAD -0.336; TCGA UCEC -0.299 |
hsa-miR-106a-5p | SH3PXD2A | 10 cancers: BLCA; CESC; COAD; HNSC; KIRC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.277; TCGA CESC -0.14; TCGA COAD -0.215; TCGA HNSC -0.104; TCGA KIRC -0.113; TCGA LUAD -0.098; TCGA LUSC -0.199; TCGA PRAD -0.286; TCGA STAD -0.173; TCGA UCEC -0.085 |
hsa-miR-106a-5p | LMO3 | 9 cancers: BLCA; COAD; ESCA; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.415; TCGA COAD -1.275; TCGA ESCA -0.401; TCGA LUAD -0.252; TCGA LUSC -0.483; TCGA PRAD -0.414; TCGA THCA -0.192; TCGA STAD -0.567; TCGA UCEC -0.417 |
hsa-miR-106a-5p | CCND1 | 9 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LIHC; LUAD; LUSC; THCA | MirTarget; miRNATAP | TCGA BLCA -0.279; TCGA BRCA -0.174; TCGA CESC -0.431; TCGA HNSC -0.29; TCGA KIRC -0.425; TCGA LIHC -0.262; TCGA LUAD -0.247; TCGA LUSC -0.216; TCGA THCA -0.333 |
hsa-miR-106a-5p | AKAP13 | 16 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LGG; LIHC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget; mirMAP; miRNATAP | TCGA BLCA -0.168; TCGA BRCA -0.071; TCGA CESC -0.107; TCGA COAD -0.22; TCGA ESCA -0.133; TCGA HNSC -0.108; TCGA KIRC -0.101; TCGA KIRP -0.113; TCGA LGG -0.103; TCGA LIHC -0.069; TCGA LUAD -0.142; TCGA LUSC -0.22; TCGA PRAD -0.171; TCGA THCA -0.079; TCGA STAD -0.066; TCGA UCEC -0.134 |
hsa-miR-106a-5p | CERCAM | 11 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUSC; OV; PAAD; PRAD; SARC; STAD | MirTarget | TCGA BLCA -0.144; TCGA BRCA -0.13; TCGA CESC -0.109; TCGA COAD -0.573; TCGA HNSC -0.232; TCGA LUSC -0.164; TCGA OV -0.102; TCGA PAAD -0.231; TCGA PRAD -0.101; TCGA SARC -0.31; TCGA STAD -0.176 |
hsa-miR-106a-5p | CSGALNACT1 | 10 cancers: BLCA; CESC; COAD; HNSC; KIRC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.238; TCGA CESC -0.153; TCGA COAD -0.225; TCGA HNSC -0.099; TCGA KIRC -0.108; TCGA LUAD -0.092; TCGA LUSC -0.29; TCGA PRAD -0.087; TCGA STAD -0.104; TCGA UCEC -0.234 |
hsa-miR-106a-5p | SH2B3 | 9 cancers: BLCA; BRCA; CESC; COAD; KIRC; LUAD; LUSC; PRAD; UCEC | MirTarget | TCGA BLCA -0.129; TCGA BRCA -0.068; TCGA CESC -0.15; TCGA COAD -0.136; TCGA KIRC -0.227; TCGA LUAD -0.114; TCGA LUSC -0.29; TCGA PRAD -0.145; TCGA UCEC -0.129 |
hsa-miR-106a-5p | YPEL4 | 13 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; KIRP; LGG; LUSC; SARC; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.097; TCGA BRCA -0.19; TCGA CESC -0.286; TCGA COAD -0.581; TCGA HNSC -0.104; TCGA KIRC -0.47; TCGA KIRP -0.164; TCGA LGG -0.197; TCGA LUSC -0.195; TCGA SARC -0.171; TCGA THCA -0.456; TCGA STAD -0.259; TCGA UCEC -0.295 |
hsa-miR-106a-5p | SYNE1 | 11 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.336; TCGA BRCA -0.157; TCGA CESC -0.166; TCGA COAD -0.587; TCGA HNSC -0.122; TCGA LGG -0.078; TCGA LUAD -0.263; TCGA LUSC -0.502; TCGA PRAD -0.305; TCGA STAD -0.296; TCGA UCEC -0.314 |
hsa-miR-106a-5p | HAS2 | 9 cancers: BLCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; PRAD; THCA | MirTarget | TCGA BLCA -0.26; TCGA CESC -0.488; TCGA COAD -0.259; TCGA HNSC -0.238; TCGA LGG -0.424; TCGA LUAD -0.155; TCGA LUSC -0.439; TCGA PRAD -0.361; TCGA THCA -0.266 |
hsa-miR-106a-5p | SHOC2 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PRAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.084; TCGA CESC -0.063; TCGA COAD -0.075; TCGA ESCA -0.057; TCGA HNSC -0.05; TCGA LUAD -0.096; TCGA LUSC -0.071; TCGA OV -0.08; TCGA PRAD -0.111; TCGA UCEC -0.087 |
hsa-miR-106a-5p | SLC46A3 | 11 cancers: BLCA; BRCA; CESC; HNSC; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.255; TCGA BRCA -0.162; TCGA CESC -0.202; TCGA HNSC -0.092; TCGA LIHC -0.304; TCGA LUAD -0.092; TCGA LUSC -0.278; TCGA OV -0.09; TCGA PRAD -0.267; TCGA STAD -0.129; TCGA UCEC -0.097 |
hsa-miR-106a-5p | FGL2 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.503; TCGA BRCA -0.122; TCGA CESC -0.329; TCGA COAD -0.432; TCGA ESCA -0.347; TCGA HNSC -0.125; TCGA LUAD -0.247; TCGA LUSC -0.348; TCGA PRAD -0.29; TCGA STAD -0.38; TCGA UCEC -0.184 |
hsa-miR-106a-5p | ETV1 | 11 cancers: BLCA; BRCA; COAD; KIRP; LGG; LUAD; LUSC; PAAD; PRAD; THCA; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.26; TCGA BRCA -0.117; TCGA COAD -0.331; TCGA KIRP -0.309; TCGA LGG -0.299; TCGA LUAD -0.321; TCGA LUSC -0.325; TCGA PAAD -0.234; TCGA PRAD -0.228; TCGA THCA -0.253; TCGA UCEC -0.108 |
hsa-miR-106a-5p | DYNC1LI2 | 9 cancers: BLCA; COAD; HNSC; KIRP; LGG; LIHC; LUSC; PRAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.064; TCGA COAD -0.07; TCGA HNSC -0.088; TCGA KIRP -0.074; TCGA LGG -0.094; TCGA LIHC -0.052; TCGA LUSC -0.072; TCGA PRAD -0.096; TCGA UCEC -0.062 |
hsa-miR-106a-5p | NACC2 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.195; TCGA BRCA -0.072; TCGA CESC -0.094; TCGA COAD -0.106; TCGA ESCA -0.193; TCGA HNSC -0.11; TCGA LUAD -0.082; TCGA LUSC -0.122; TCGA PRAD -0.272; TCGA STAD -0.14; TCGA UCEC -0.149 |
hsa-miR-106a-5p | KATNAL1 | 9 cancers: BLCA; COAD; KIRC; KIRP; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.209; TCGA COAD -0.255; TCGA KIRC -0.059; TCGA KIRP -0.094; TCGA LUAD -0.062; TCGA LUSC -0.106; TCGA PRAD -0.213; TCGA STAD -0.184; TCGA UCEC -0.194 |
hsa-miR-106a-5p | FNDC3A | 9 cancers: BLCA; BRCA; HNSC; KIRP; LIHC; LUAD; LUSC; PRAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.116; TCGA BRCA -0.067; TCGA HNSC -0.092; TCGA KIRP -0.106; TCGA LIHC -0.103; TCGA LUAD -0.059; TCGA LUSC -0.164; TCGA PRAD -0.247; TCGA UCEC -0.121 |
hsa-miR-106a-5p | ATL3 | 9 cancers: BLCA; COAD; ESCA; HNSC; LUAD; LUSC; OV; PRAD; STAD | MirTarget; miRNATAP | TCGA BLCA -0.091; TCGA COAD -0.165; TCGA ESCA -0.147; TCGA HNSC -0.146; TCGA LUAD -0.071; TCGA LUSC -0.157; TCGA OV -0.078; TCGA PRAD -0.144; TCGA STAD -0.12 |
hsa-miR-106a-5p | AKTIP | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LIHC; LUAD; LUSC; OV; PRAD; SARC; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.078; TCGA BRCA -0.055; TCGA CESC -0.078; TCGA COAD -0.053; TCGA HNSC -0.06; TCGA LIHC -0.121; TCGA LUAD -0.073; TCGA LUSC -0.073; TCGA OV -0.116; TCGA PRAD -0.095; TCGA SARC -0.079; TCGA UCEC -0.088 |
hsa-miR-106a-5p | RAB11FIP5 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PAAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.125; TCGA BRCA -0.09; TCGA CESC -0.138; TCGA COAD -0.105; TCGA ESCA -0.073; TCGA HNSC -0.195; TCGA LUAD -0.138; TCGA LUSC -0.113; TCGA OV -0.067; TCGA PAAD -0.095; TCGA SARC -0.179; TCGA THCA -0.149; TCGA STAD -0.127; TCGA UCEC -0.187 |
hsa-miR-106a-5p | DAB2 | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.13; TCGA BRCA -0.111; TCGA CESC -0.337; TCGA COAD -0.175; TCGA ESCA -0.194; TCGA HNSC -0.221; TCGA LUAD -0.201; TCGA LUSC -0.444; TCGA STAD -0.179; TCGA UCEC -0.18 |
hsa-miR-106a-5p | ZHX2 | 10 cancers: BLCA; COAD; KIRP; LGG; LIHC; LUAD; LUSC; OV; THCA; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.073; TCGA COAD -0.172; TCGA KIRP -0.143; TCGA LGG -0.239; TCGA LIHC -0.063; TCGA LUAD -0.066; TCGA LUSC -0.07; TCGA OV -0.121; TCGA THCA -0.155; TCGA UCEC -0.159 |
hsa-miR-106a-5p | RHOC | 10 cancers: BLCA; BRCA; COAD; ESCA; LIHC; LUAD; LUSC; OV; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.121; TCGA BRCA -0.082; TCGA COAD -0.111; TCGA ESCA -0.11; TCGA LIHC -0.125; TCGA LUAD -0.08; TCGA LUSC -0.155; TCGA OV -0.066; TCGA THCA -0.182; TCGA STAD -0.103 |
hsa-miR-106a-5p | ST6GALNAC6 | 14 cancers: BLCA; BRCA; COAD; ESCA; HNSC; KIRC; LIHC; LUAD; LUSC; OV; SARC; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.153; TCGA BRCA -0.127; TCGA COAD -0.378; TCGA ESCA -0.144; TCGA HNSC -0.073; TCGA KIRC -0.063; TCGA LIHC -0.084; TCGA LUAD -0.179; TCGA LUSC -0.208; TCGA OV -0.109; TCGA SARC -0.128; TCGA THCA -0.089; TCGA STAD -0.293; TCGA UCEC -0.164 |
hsa-miR-106a-5p | CRY2 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LIHC; LUAD; LUSC; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.142; TCGA BRCA -0.13; TCGA CESC -0.052; TCGA COAD -0.104; TCGA HNSC -0.095; TCGA LIHC -0.169; TCGA LUAD -0.132; TCGA LUSC -0.175; TCGA PRAD -0.102; TCGA SARC -0.14; TCGA STAD -0.146; TCGA UCEC -0.11 |
hsa-miR-106a-5p | PDCD1LG2 | 11 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; SARC; THCA; UCEC | MirTarget | TCGA BLCA -0.364; TCGA CESC -0.378; TCGA COAD -0.465; TCGA ESCA -0.207; TCGA HNSC -0.166; TCGA LUAD -0.183; TCGA LUSC -0.322; TCGA PRAD -0.236; TCGA SARC -0.183; TCGA THCA -0.4; TCGA UCEC -0.179 |
hsa-miR-106a-5p | BMPR2 | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.134; TCGA BRCA -0.052; TCGA CESC -0.055; TCGA COAD -0.229; TCGA HNSC -0.119; TCGA LUAD -0.127; TCGA LUSC -0.215; TCGA PRAD -0.198; TCGA STAD -0.087; TCGA UCEC -0.134 |
hsa-miR-106a-5p | ZDHHC1 | 9 cancers: BLCA; BRCA; ESCA; HNSC; LUAD; LUSC; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.1; TCGA BRCA -0.115; TCGA ESCA -0.156; TCGA HNSC -0.078; TCGA LUAD -0.096; TCGA LUSC -0.08; TCGA SARC -0.162; TCGA STAD -0.159; TCGA UCEC -0.092 |
hsa-miR-106a-5p | ZNF532 | 9 cancers: BLCA; CESC; COAD; KIRC; LGG; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.101; TCGA CESC -0.114; TCGA COAD -0.384; TCGA KIRC -0.17; TCGA LGG -0.078; TCGA PRAD -0.089; TCGA THCA -0.099; TCGA STAD -0.106; TCGA UCEC -0.09 |
hsa-miR-106a-5p | KIAA0513 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.33; TCGA BRCA -0.062; TCGA CESC -0.131; TCGA COAD -0.323; TCGA ESCA -0.208; TCGA HNSC -0.163; TCGA LIHC -0.171; TCGA LUAD -0.119; TCGA LUSC -0.338; TCGA PRAD -0.159; TCGA STAD -0.234; TCGA UCEC -0.175 |
hsa-miR-106a-5p | CTSK | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.456; TCGA BRCA -0.175; TCGA CESC -0.293; TCGA COAD -0.55; TCGA ESCA -0.212; TCGA HNSC -0.331; TCGA LUAD -0.135; TCGA LUSC -0.363; TCGA PRAD -0.115; TCGA SARC -0.391; TCGA THCA -0.189; TCGA STAD -0.18; TCGA UCEC -0.23 |
hsa-miR-106a-5p | ANK2 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LUSC; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.382; TCGA BRCA -0.168; TCGA CESC -0.2; TCGA COAD -0.84; TCGA ESCA -0.249; TCGA HNSC -0.244; TCGA LGG -0.098; TCGA LUSC -0.218; TCGA PRAD -0.42; TCGA SARC -0.15; TCGA STAD -0.446; TCGA UCEC -0.166 |
hsa-miR-106a-5p | CTSA | 9 cancers: BLCA; ESCA; HNSC; LIHC; LUSC; OV; PAAD; SARC; THCA | MirTarget | TCGA BLCA -0.051; TCGA ESCA -0.084; TCGA HNSC -0.111; TCGA LIHC -0.065; TCGA LUSC -0.16; TCGA OV -0.13; TCGA PAAD -0.09; TCGA SARC -0.117; TCGA THCA -0.278 |
hsa-miR-106a-5p | ARHGEF3 | 9 cancers: BLCA; CESC; COAD; ESCA; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.116; TCGA CESC -0.07; TCGA COAD -0.159; TCGA ESCA -0.087; TCGA LUAD -0.122; TCGA LUSC -0.17; TCGA PRAD -0.144; TCGA STAD -0.074; TCGA UCEC -0.072 |
hsa-miR-106a-5p | RNASEL | 10 cancers: BLCA; BRCA; COAD; KIRP; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.088; TCGA BRCA -0.087; TCGA COAD -0.176; TCGA KIRP -0.078; TCGA LIHC -0.198; TCGA LUAD -0.062; TCGA LUSC -0.067; TCGA PRAD -0.164; TCGA STAD -0.113; TCGA UCEC -0.188 |
hsa-miR-106a-5p | JAZF1 | 11 cancers: BLCA; BRCA; CESC; COAD; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.296; TCGA BRCA -0.058; TCGA CESC -0.127; TCGA COAD -0.415; TCGA LIHC -0.128; TCGA LUAD -0.128; TCGA LUSC -0.151; TCGA OV -0.103; TCGA PRAD -0.288; TCGA STAD -0.129; TCGA UCEC -0.147 |
hsa-miR-106a-5p | NDEL1 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUSC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.069; TCGA CESC -0.083; TCGA COAD -0.112; TCGA ESCA -0.112; TCGA HNSC -0.174; TCGA KIRC -0.104; TCGA KIRP -0.092; TCGA LUSC -0.127; TCGA STAD -0.079; TCGA UCEC -0.051 |
hsa-miR-106a-5p | RASGRF2 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.254; TCGA BRCA -0.139; TCGA CESC -0.247; TCGA COAD -0.319; TCGA HNSC -0.228; TCGA KIRC -0.135; TCGA LUAD -0.144; TCGA LUSC -0.388; TCGA OV -0.139; TCGA PRAD -0.289; TCGA STAD -0.128; TCGA UCEC -0.205 |
hsa-miR-106a-5p | ERAP1 | 9 cancers: BLCA; KIRC; LGG; LIHC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.091; TCGA KIRC -0.078; TCGA LGG -0.093; TCGA LIHC -0.124; TCGA LUAD -0.102; TCGA LUSC -0.153; TCGA PRAD -0.232; TCGA THCA -0.084; TCGA UCEC -0.097 |
hsa-miR-106a-5p | SLAIN2 | 9 cancers: BLCA; CESC; HNSC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.073; TCGA CESC -0.085; TCGA HNSC -0.056; TCGA LUAD -0.085; TCGA LUSC -0.066; TCGA OV -0.078; TCGA PRAD -0.098; TCGA STAD -0.06; TCGA UCEC -0.051 |
hsa-miR-106a-5p | F3 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.38; TCGA BRCA -0.165; TCGA CESC -0.257; TCGA COAD -0.199; TCGA ESCA -0.27; TCGA HNSC -0.201; TCGA LUAD -0.195; TCGA LUSC -0.328; TCGA PRAD -0.445; TCGA STAD -0.167; TCGA UCEC -0.113 |
hsa-miR-106a-5p | PDGFRA | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.371; TCGA BRCA -0.114; TCGA CESC -0.26; TCGA COAD -0.461; TCGA HNSC -0.187; TCGA LGG -0.291; TCGA LUAD -0.262; TCGA LUSC -0.319; TCGA PRAD -0.288; TCGA SARC -0.318; TCGA STAD -0.128; TCGA UCEC -0.249 |
hsa-miR-106a-5p | LIMA1 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; PAAD; PRAD; THCA; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.15; TCGA BRCA -0.116; TCGA CESC -0.123; TCGA COAD -0.14; TCGA HNSC -0.165; TCGA LGG -0.102; TCGA LUAD -0.082; TCGA LUSC -0.164; TCGA PAAD -0.135; TCGA PRAD -0.14; TCGA THCA -0.178; TCGA UCEC -0.082 |
hsa-miR-106a-5p | THBS2 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUAD; LUSC; PAAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.453; TCGA BRCA -0.088; TCGA CESC -0.336; TCGA COAD -0.852; TCGA ESCA -0.263; TCGA HNSC -0.498; TCGA KIRP -0.321; TCGA LUAD -0.141; TCGA LUSC -0.385; TCGA PAAD -0.319; TCGA SARC -0.2; TCGA THCA -0.374; TCGA STAD -0.192; TCGA UCEC -0.156 |
hsa-miR-106a-5p | SCN2B | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.556; TCGA BRCA -0.475; TCGA CESC -0.498; TCGA COAD -1.045; TCGA ESCA -0.326; TCGA HNSC -0.577; TCGA LUAD -0.432; TCGA LUSC -0.595; TCGA PRAD -0.38; TCGA SARC -0.383; TCGA THCA -0.205; TCGA STAD -0.96; TCGA UCEC -0.486 |
hsa-miR-106a-5p | KLF9 | 10 cancers: BLCA; BRCA; COAD; HNSC; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.366; TCGA BRCA -0.119; TCGA COAD -0.391; TCGA HNSC -0.088; TCGA LIHC -0.202; TCGA LUAD -0.2; TCGA LUSC -0.296; TCGA PRAD -0.095; TCGA STAD -0.297; TCGA UCEC -0.203 |
hsa-miR-106a-5p | TMEM127 | 10 cancers: BLCA; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.056; TCGA COAD -0.106; TCGA ESCA -0.084; TCGA HNSC -0.072; TCGA LIHC -0.064; TCGA LUAD -0.061; TCGA LUSC -0.067; TCGA PRAD -0.104; TCGA THCA -0.081; TCGA STAD -0.09 |
hsa-miR-106a-5p | ZBTB7A | 13 cancers: BLCA; BRCA; CESC; ESCA; KIRC; KIRP; LIHC; LUAD; LUSC; OV; SARC; THCA; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.071; TCGA BRCA -0.053; TCGA CESC -0.071; TCGA ESCA -0.102; TCGA KIRC -0.084; TCGA KIRP -0.065; TCGA LIHC -0.112; TCGA LUAD -0.099; TCGA LUSC -0.104; TCGA OV -0.09; TCGA SARC -0.055; TCGA THCA -0.064; TCGA UCEC -0.07 |
hsa-miR-106a-5p | TBC1D2 | 13 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.14; TCGA BRCA -0.063; TCGA CESC -0.212; TCGA ESCA -0.145; TCGA HNSC -0.106; TCGA KIRC -0.16; TCGA KIRP -0.15; TCGA LIHC -0.086; TCGA LUAD -0.071; TCGA LUSC -0.162; TCGA PRAD -0.14; TCGA THCA -0.553; TCGA STAD -0.161 |
hsa-miR-106a-5p | S1PR1 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LGG; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.253; TCGA BRCA -0.173; TCGA CESC -0.222; TCGA COAD -0.42; TCGA ESCA -0.165; TCGA HNSC -0.111; TCGA KIRC -0.253; TCGA LGG -0.121; TCGA LIHC -0.116; TCGA LUAD -0.219; TCGA LUSC -0.437; TCGA OV -0.069; TCGA PRAD -0.158; TCGA STAD -0.278; TCGA UCEC -0.205 |
hsa-miR-106a-5p | PLEKHO2 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LGG; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.201; TCGA BRCA -0.068; TCGA CESC -0.153; TCGA COAD -0.323; TCGA ESCA -0.187; TCGA HNSC -0.106; TCGA KIRC -0.175; TCGA LGG -0.087; TCGA LUAD -0.111; TCGA LUSC -0.288; TCGA PRAD -0.127; TCGA STAD -0.131; TCGA UCEC -0.097 |
hsa-miR-106a-5p | TAOK3 | 12 cancers: BLCA; CESC; COAD; ESCA; HNSC; LGG; LUAD; LUSC; OV; PRAD; SARC; UCEC | MirTarget | TCGA BLCA -0.051; TCGA CESC -0.163; TCGA COAD -0.057; TCGA ESCA -0.112; TCGA HNSC -0.091; TCGA LGG -0.119; TCGA LUAD -0.129; TCGA LUSC -0.2; TCGA OV -0.051; TCGA PRAD -0.065; TCGA SARC -0.075; TCGA UCEC -0.151 |
hsa-miR-106a-5p | BNIP3L | 11 cancers: BLCA; BRCA; COAD; KIRC; KIRP; LGG; LUAD; LUSC; OV; PAAD; PRAD | MirTarget | TCGA BLCA -0.068; TCGA BRCA -0.079; TCGA COAD -0.235; TCGA KIRC -0.117; TCGA KIRP -0.096; TCGA LGG -0.053; TCGA LUAD -0.102; TCGA LUSC -0.064; TCGA OV -0.057; TCGA PAAD -0.093; TCGA PRAD -0.176 |
hsa-miR-106a-5p | DOCK4 | 9 cancers: BLCA; CESC; COAD; HNSC; KIRC; KIRP; LUAD; LUSC; UCEC | MirTarget | TCGA BLCA -0.077; TCGA CESC -0.179; TCGA COAD -0.321; TCGA HNSC -0.062; TCGA KIRC -0.165; TCGA KIRP -0.104; TCGA LUAD -0.122; TCGA LUSC -0.319; TCGA UCEC -0.135 |
hsa-miR-106a-5p | ANKRD29 | 10 cancers: BLCA; BRCA; CESC; COAD; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.515; TCGA BRCA -0.249; TCGA CESC -0.21; TCGA COAD -0.4; TCGA LIHC -0.482; TCGA LUAD -0.191; TCGA LUSC -0.393; TCGA PRAD -0.239; TCGA STAD -0.379; TCGA UCEC -0.201 |
hsa-miR-106a-5p | FAT4 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LUAD; LUSC; PRAD; STAD | MirTarget; miRNATAP | TCGA BLCA -0.418; TCGA BRCA -0.217; TCGA CESC -0.243; TCGA COAD -0.507; TCGA ESCA -0.175; TCGA HNSC -0.143; TCGA LGG -0.244; TCGA LUAD -0.383; TCGA LUSC -0.462; TCGA PRAD -0.45; TCGA STAD -0.246 |
hsa-miR-106a-5p | FRMD6 | 9 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.264; TCGA BRCA -0.134; TCGA CESC -0.396; TCGA COAD -0.69; TCGA HNSC -0.156; TCGA LUAD -0.113; TCGA PRAD -0.375; TCGA STAD -0.335; TCGA UCEC -0.202 |
hsa-miR-106a-5p | REEP3 | 11 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PRAD; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.06; TCGA CESC -0.08; TCGA COAD -0.18; TCGA ESCA -0.117; TCGA HNSC -0.146; TCGA LUAD -0.114; TCGA LUSC -0.169; TCGA OV -0.117; TCGA PRAD -0.089; TCGA THCA -0.143; TCGA STAD -0.072 |
hsa-miR-106a-5p | TACC1 | 11 cancers: BLCA; BRCA; CESC; COAD; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.204; TCGA BRCA -0.078; TCGA CESC -0.118; TCGA COAD -0.194; TCGA LIHC -0.11; TCGA LUAD -0.165; TCGA LUSC -0.184; TCGA OV -0.128; TCGA PRAD -0.293; TCGA STAD -0.168; TCGA UCEC -0.235 |
hsa-miR-106a-5p | HIF1A | 9 cancers: BLCA; CESC; COAD; ESCA; HNSC; LGG; LUSC; PRAD; THCA | MirTarget | TCGA BLCA -0.082; TCGA CESC -0.073; TCGA COAD -0.211; TCGA ESCA -0.106; TCGA HNSC -0.137; TCGA LGG -0.102; TCGA LUSC -0.095; TCGA PRAD -0.181; TCGA THCA -0.202 |
hsa-miR-106a-5p | GAS7 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.487; TCGA BRCA -0.18; TCGA CESC -0.306; TCGA COAD -0.626; TCGA ESCA -0.32; TCGA HNSC -0.26; TCGA LUAD -0.247; TCGA LUSC -0.45; TCGA PRAD -0.23; TCGA STAD -0.336; TCGA UCEC -0.146 |
hsa-miR-106a-5p | TRAF3IP2 | 11 cancers: BLCA; CESC; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; PRAD; SARC | mirMAP | TCGA BLCA -0.103; TCGA CESC -0.13; TCGA ESCA -0.113; TCGA HNSC -0.113; TCGA KIRC -0.179; TCGA KIRP -0.121; TCGA LIHC -0.115; TCGA LUAD -0.098; TCGA LUSC -0.093; TCGA PRAD -0.072; TCGA SARC -0.092 |
hsa-miR-106a-5p | PPP2R3A | 11 cancers: BLCA; CESC; COAD; HNSC; KIRP; LGG; LUAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.071; TCGA CESC -0.207; TCGA COAD -0.266; TCGA HNSC -0.176; TCGA KIRP -0.159; TCGA LGG -0.099; TCGA LUAD -0.1; TCGA PRAD -0.133; TCGA THCA -0.16; TCGA STAD -0.211; TCGA UCEC -0.096 |
hsa-miR-106a-5p | NUAK1 | 10 cancers: BLCA; BRCA; COAD; HNSC; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.151; TCGA BRCA -0.11; TCGA COAD -0.462; TCGA HNSC -0.078; TCGA LUAD -0.085; TCGA LUSC -0.246; TCGA PAAD -0.268; TCGA PRAD -0.224; TCGA STAD -0.103; TCGA UCEC -0.122 |
hsa-miR-106a-5p | PGR | 9 cancers: BLCA; BRCA; COAD; ESCA; LUAD; LUSC; PRAD; THCA; STAD | mirMAP | TCGA BLCA -0.482; TCGA BRCA -0.436; TCGA COAD -0.886; TCGA ESCA -0.245; TCGA LUAD -0.391; TCGA LUSC -0.562; TCGA PRAD -0.471; TCGA THCA -0.484; TCGA STAD -0.557 |
hsa-miR-106a-5p | LAT2 | 11 cancers: BLCA; CESC; COAD; ESCA; KIRC; KIRP; LGG; LUAD; LUSC; THCA; STAD | mirMAP | TCGA BLCA -0.213; TCGA CESC -0.17; TCGA COAD -0.225; TCGA ESCA -0.154; TCGA KIRC -0.34; TCGA KIRP -0.146; TCGA LGG -0.143; TCGA LUAD -0.168; TCGA LUSC -0.377; TCGA THCA -0.297; TCGA STAD -0.113 |
hsa-miR-106a-5p | STX1B | 10 cancers: BLCA; BRCA; CESC; ESCA; KIRC; KIRP; LIHC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.166; TCGA BRCA -0.129; TCGA CESC -0.276; TCGA ESCA -0.162; TCGA KIRC -0.167; TCGA KIRP -0.274; TCGA LIHC -0.272; TCGA THCA -0.401; TCGA STAD -0.142; TCGA UCEC -0.186 |
hsa-miR-106a-5p | TTC28 | 9 cancers: BLCA; BRCA; COAD; LGG; LUAD; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.124; TCGA BRCA -0.143; TCGA COAD -0.287; TCGA LGG -0.145; TCGA LUAD -0.062; TCGA PRAD -0.278; TCGA SARC -0.081; TCGA STAD -0.249; TCGA UCEC -0.051 |
hsa-miR-106a-5p | CBX7 | 13 cancers: BLCA; BRCA; COAD; ESCA; KIRC; LIHC; LUAD; LUSC; OV; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.217; TCGA BRCA -0.237; TCGA COAD -0.239; TCGA ESCA -0.129; TCGA KIRC -0.118; TCGA LIHC -0.198; TCGA LUAD -0.174; TCGA LUSC -0.118; TCGA OV -0.14; TCGA PRAD -0.169; TCGA SARC -0.112; TCGA STAD -0.369; TCGA UCEC -0.34 |
hsa-miR-106a-5p | SCN1B | 10 cancers: BLCA; BRCA; COAD; HNSC; KIRC; LUSC; OV; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.374; TCGA BRCA -0.234; TCGA COAD -0.421; TCGA HNSC -0.289; TCGA KIRC -0.287; TCGA LUSC -0.262; TCGA OV -0.11; TCGA THCA -0.645; TCGA STAD -0.265; TCGA UCEC -0.173 |
hsa-miR-106a-5p | ASPA | 9 cancers: BLCA; BRCA; COAD; ESCA; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.257; TCGA BRCA -0.307; TCGA COAD -0.605; TCGA ESCA -0.314; TCGA LUAD -0.227; TCGA LUSC -0.492; TCGA PRAD -0.498; TCGA STAD -0.497; TCGA UCEC -0.41 |
hsa-miR-106a-5p | COL1A1 | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; LUSC; PAAD; SARC; THCA | mirMAP | TCGA BLCA -0.436; TCGA BRCA -0.087; TCGA CESC -0.27; TCGA COAD -0.594; TCGA HNSC -0.379; TCGA KIRC -0.328; TCGA LUSC -0.433; TCGA PAAD -0.267; TCGA SARC -0.244; TCGA THCA -0.648 |
hsa-miR-106a-5p | CPEB4 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.168; TCGA BRCA -0.13; TCGA CESC -0.109; TCGA COAD -0.193; TCGA HNSC -0.166; TCGA LIHC -0.098; TCGA LUAD -0.092; TCGA LUSC -0.239; TCGA OV -0.064; TCGA PRAD -0.22; TCGA STAD -0.103; TCGA UCEC -0.128 |
hsa-miR-106a-5p | FHL2 | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRP; LIHC; LUAD; LUSC; PRAD; STAD | mirMAP | TCGA BLCA -0.156; TCGA BRCA -0.15; TCGA CESC -0.215; TCGA ESCA -0.207; TCGA HNSC -0.225; TCGA KIRP -0.219; TCGA LIHC -0.149; TCGA LUAD -0.104; TCGA LUSC -0.335; TCGA PRAD -0.213; TCGA STAD -0.21 |
hsa-miR-106a-5p | MEF2D | 11 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; LIHC; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.097; TCGA BRCA -0.059; TCGA CESC -0.096; TCGA COAD -0.163; TCGA HNSC -0.069; TCGA KIRC -0.121; TCGA LIHC -0.05; TCGA LUSC -0.123; TCGA PRAD -0.074; TCGA STAD -0.094; TCGA UCEC -0.139 |
hsa-miR-106a-5p | ESYT2 | 11 cancers: BLCA; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.097; TCGA ESCA -0.096; TCGA HNSC -0.057; TCGA KIRC -0.07; TCGA KIRP -0.148; TCGA LIHC -0.073; TCGA LUAD -0.061; TCGA LUSC -0.104; TCGA PRAD -0.107; TCGA STAD -0.074; TCGA UCEC -0.079 |
hsa-miR-106a-5p | ANXA11 | 12 cancers: BLCA; BRCA; ESCA; KIRP; LIHC; LUAD; LUSC; OV; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.119; TCGA BRCA -0.058; TCGA ESCA -0.107; TCGA KIRP -0.07; TCGA LIHC -0.082; TCGA LUAD -0.113; TCGA LUSC -0.15; TCGA OV -0.09; TCGA SARC -0.1; TCGA THCA -0.128; TCGA STAD -0.086; TCGA UCEC -0.053 |
hsa-miR-106a-5p | LDB3 | 11 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.373; TCGA BRCA -0.145; TCGA COAD -0.825; TCGA ESCA -0.403; TCGA HNSC -0.391; TCGA LIHC -0.151; TCGA LUAD -0.159; TCGA LUSC -0.152; TCGA PRAD -0.417; TCGA STAD -0.794; TCGA UCEC -0.113 |
hsa-miR-106a-5p | RAB30 | 10 cancers: BLCA; BRCA; COAD; HNSC; LGG; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP; miRNATAP | TCGA BLCA -0.163; TCGA BRCA -0.093; TCGA COAD -0.142; TCGA HNSC -0.128; TCGA LGG -0.158; TCGA LUAD -0.159; TCGA LUSC -0.085; TCGA PRAD -0.164; TCGA STAD -0.153; TCGA UCEC -0.074 |
hsa-miR-106a-5p | ENTPD1 | 11 cancers: BLCA; CESC; COAD; ESCA; KIRC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.215; TCGA CESC -0.077; TCGA COAD -0.266; TCGA ESCA -0.084; TCGA KIRC -0.105; TCGA LUAD -0.081; TCGA LUSC -0.142; TCGA PRAD -0.12; TCGA THCA -0.377; TCGA STAD -0.149; TCGA UCEC -0.133 |
hsa-miR-106a-5p | NTRK3 | 11 cancers: BLCA; BRCA; CESC; COAD; LGG; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.562; TCGA BRCA -0.158; TCGA CESC -0.271; TCGA COAD -0.928; TCGA LGG -0.178; TCGA LUAD -0.356; TCGA LUSC -0.287; TCGA PRAD -0.256; TCGA THCA -0.262; TCGA STAD -0.692; TCGA UCEC -0.133 |
hsa-miR-106a-5p | FTO | 13 cancers: BLCA; BRCA; COAD; HNSC; KIRC; KIRP; LGG; LIHC; LUAD; LUSC; PRAD; SARC; UCEC | mirMAP | TCGA BLCA -0.097; TCGA BRCA -0.101; TCGA COAD -0.097; TCGA HNSC -0.109; TCGA KIRC -0.094; TCGA KIRP -0.096; TCGA LGG -0.093; TCGA LIHC -0.095; TCGA LUAD -0.125; TCGA LUSC -0.125; TCGA PRAD -0.182; TCGA SARC -0.077; TCGA UCEC -0.071 |
hsa-miR-106a-5p | ZCCHC14 | 10 cancers: BLCA; HNSC; KIRP; LGG; LUAD; LUSC; PAAD; PRAD; THCA; UCEC | mirMAP | TCGA BLCA -0.063; TCGA HNSC -0.052; TCGA KIRP -0.11; TCGA LGG -0.08; TCGA LUAD -0.058; TCGA LUSC -0.062; TCGA PAAD -0.109; TCGA PRAD -0.085; TCGA THCA -0.089; TCGA UCEC -0.069 |
hsa-miR-106a-5p | WTIP | 9 cancers: BLCA; BRCA; COAD; HNSC; LUAD; LUSC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.079; TCGA BRCA -0.062; TCGA COAD -0.321; TCGA HNSC -0.155; TCGA LUAD -0.096; TCGA LUSC -0.131; TCGA THCA -0.349; TCGA STAD -0.252; TCGA UCEC -0.103 |
hsa-miR-106a-5p | SETD7 | 13 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; LIHC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.14; TCGA BRCA -0.052; TCGA CESC -0.074; TCGA ESCA -0.085; TCGA HNSC -0.118; TCGA KIRC -0.086; TCGA LIHC -0.117; TCGA LUAD -0.103; TCGA LUSC -0.176; TCGA PRAD -0.094; TCGA THCA -0.156; TCGA STAD -0.098; TCGA UCEC -0.239 |
hsa-miR-106a-5p | CXorf36 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.182; TCGA BRCA -0.195; TCGA CESC -0.207; TCGA COAD -0.366; TCGA HNSC -0.142; TCGA KIRC -0.425; TCGA LIHC -0.118; TCGA LUAD -0.171; TCGA LUSC -0.383; TCGA PRAD -0.153; TCGA STAD -0.233; TCGA UCEC -0.255 |
hsa-miR-106a-5p | DIXDC1 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.388; TCGA BRCA -0.096; TCGA CESC -0.101; TCGA COAD -0.351; TCGA ESCA -0.208; TCGA HNSC -0.197; TCGA LUAD -0.143; TCGA LUSC -0.214; TCGA PRAD -0.254; TCGA STAD -0.312; TCGA UCEC -0.239 |
hsa-miR-106a-5p | GFRA1 | 11 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.448; TCGA BRCA -0.387; TCGA CESC -0.316; TCGA COAD -0.706; TCGA HNSC -0.213; TCGA LGG -0.214; TCGA LUAD -0.367; TCGA LUSC -0.36; TCGA PRAD -0.33; TCGA STAD -0.658; TCGA UCEC -0.345 |
hsa-miR-106a-5p | DST | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.202; TCGA BRCA -0.134; TCGA CESC -0.292; TCGA COAD -0.259; TCGA HNSC -0.142; TCGA LGG -0.224; TCGA LUAD -0.117; TCGA LUSC -0.179; TCGA PRAD -0.255; TCGA THCA -0.303; TCGA STAD -0.269; TCGA UCEC -0.133 |
hsa-miR-106a-5p | MR1 | 11 cancers: BLCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; THCA; STAD | mirMAP | TCGA BLCA -0.102; TCGA CESC -0.118; TCGA COAD -0.204; TCGA ESCA -0.118; TCGA HNSC -0.092; TCGA LIHC -0.123; TCGA LUAD -0.203; TCGA LUSC -0.215; TCGA PRAD -0.289; TCGA THCA -0.1; TCGA STAD -0.147 |
hsa-miR-106a-5p | GNAQ | 9 cancers: BLCA; COAD; ESCA; KIRP; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.08; TCGA COAD -0.199; TCGA ESCA -0.105; TCGA KIRP -0.084; TCGA LUAD -0.115; TCGA LUSC -0.131; TCGA PRAD -0.093; TCGA STAD -0.063; TCGA UCEC -0.11 |
hsa-miR-106a-5p | MMP16 | 13 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; LGG; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.299; TCGA BRCA -0.195; TCGA CESC -0.355; TCGA COAD -0.528; TCGA HNSC -0.364; TCGA KIRC -0.3; TCGA LGG -0.255; TCGA LUAD -0.232; TCGA LUSC -0.245; TCGA PRAD -0.454; TCGA THCA -0.754; TCGA STAD -0.261; TCGA UCEC -0.114 |
hsa-miR-106a-5p | BACH1 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; LGG; LUSC; PRAD; THCA; UCEC | mirMAP | TCGA BLCA -0.08; TCGA CESC -0.153; TCGA COAD -0.134; TCGA ESCA -0.108; TCGA HNSC -0.088; TCGA LGG -0.123; TCGA LUSC -0.15; TCGA PRAD -0.064; TCGA THCA -0.111; TCGA UCEC -0.124 |
hsa-miR-106a-5p | SHROOM4 | 11 cancers: BLCA; BRCA; CESC; HNSC; LIHC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.15; TCGA BRCA -0.111; TCGA CESC -0.117; TCGA HNSC -0.159; TCGA LIHC -0.132; TCGA LUAD -0.294; TCGA LUSC -0.42; TCGA PRAD -0.304; TCGA THCA -0.617; TCGA STAD -0.18; TCGA UCEC -0.139 |
hsa-miR-106a-5p | IL6R | 9 cancers: BLCA; COAD; HNSC; KIRP; LGG; LUAD; LUSC; PRAD; UCEC | mirMAP | TCGA BLCA -0.189; TCGA COAD -0.214; TCGA HNSC -0.207; TCGA KIRP -0.138; TCGA LGG -0.132; TCGA LUAD -0.218; TCGA LUSC -0.285; TCGA PRAD -0.193; TCGA UCEC -0.176 |
hsa-miR-106a-5p | ADCY9 | 11 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.206; TCGA BRCA -0.084; TCGA COAD -0.154; TCGA ESCA -0.17; TCGA HNSC -0.088; TCGA LIHC -0.081; TCGA LUAD -0.17; TCGA LUSC -0.201; TCGA PRAD -0.191; TCGA STAD -0.159; TCGA UCEC -0.132 |
hsa-miR-106a-5p | KLHL21 | 9 cancers: BLCA; CESC; HNSC; LIHC; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.145; TCGA CESC -0.142; TCGA HNSC -0.057; TCGA LIHC -0.16; TCGA LUSC -0.087; TCGA PRAD -0.107; TCGA THCA -0.083; TCGA STAD -0.169; TCGA UCEC -0.116 |
hsa-miR-106a-5p | NFIA | 9 cancers: BLCA; ESCA; KIRC; KIRP; LGG; LIHC; LUSC; PRAD; STAD | mirMAP | TCGA BLCA -0.195; TCGA ESCA -0.12; TCGA KIRC -0.147; TCGA KIRP -0.202; TCGA LGG -0.148; TCGA LIHC -0.143; TCGA LUSC -0.147; TCGA PRAD -0.081; TCGA STAD -0.219 |
hsa-miR-106a-5p | NFASC | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.343; TCGA BRCA -0.109; TCGA CESC -0.311; TCGA COAD -0.343; TCGA ESCA -0.331; TCGA HNSC -0.228; TCGA LUAD -0.134; TCGA LUSC -0.17; TCGA PRAD -0.292; TCGA STAD -0.486; TCGA UCEC -0.154 |
hsa-miR-106a-5p | SLMAP | 9 cancers: BLCA; CESC; COAD; ESCA; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.205; TCGA CESC -0.108; TCGA COAD -0.125; TCGA ESCA -0.122; TCGA LUAD -0.06; TCGA LUSC -0.068; TCGA PRAD -0.218; TCGA STAD -0.105; TCGA UCEC -0.18 |
hsa-miR-106a-5p | SCN3B | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.351; TCGA BRCA -0.312; TCGA CESC -0.279; TCGA COAD -0.718; TCGA HNSC -0.345; TCGA LUAD -0.205; TCGA LUSC -0.215; TCGA PRAD -0.219; TCGA STAD -0.257; TCGA UCEC -0.322 |
hsa-miR-106a-5p | SLFN5 | 13 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRP; LIHC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.105; TCGA CESC -0.097; TCGA COAD -0.42; TCGA ESCA -0.124; TCGA HNSC -0.077; TCGA KIRP -0.22; TCGA LIHC -0.207; TCGA LUAD -0.118; TCGA LUSC -0.162; TCGA PRAD -0.315; TCGA THCA -0.194; TCGA STAD -0.095; TCGA UCEC -0.073 |
hsa-miR-106a-5p | LUZP1 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP; miRNATAP | TCGA BLCA -0.068; TCGA CESC -0.198; TCGA COAD -0.092; TCGA ESCA -0.081; TCGA HNSC -0.146; TCGA LUAD -0.077; TCGA LUSC -0.171; TCGA PRAD -0.195; TCGA STAD -0.075; TCGA UCEC -0.105 |
hsa-miR-106a-5p | KLF13 | 10 cancers: BLCA; BRCA; ESCA; HNSC; KIRP; LIHC; LUAD; LUSC; PRAD; UCEC | mirMAP | TCGA BLCA -0.126; TCGA BRCA -0.054; TCGA ESCA -0.092; TCGA HNSC -0.095; TCGA KIRP -0.119; TCGA LIHC -0.095; TCGA LUAD -0.149; TCGA LUSC -0.273; TCGA PRAD -0.234; TCGA UCEC -0.13 |
hsa-miR-106a-5p | SGCD | 10 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LGG; LUAD; LUSC; SARC; STAD | mirMAP | TCGA BLCA -0.71; TCGA BRCA -0.231; TCGA COAD -0.827; TCGA ESCA -0.385; TCGA HNSC -0.545; TCGA LGG -0.137; TCGA LUAD -0.35; TCGA LUSC -0.538; TCGA SARC -0.194; TCGA STAD -0.408 |
hsa-miR-106a-5p | KCND3 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LGG; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.601; TCGA BRCA -0.348; TCGA CESC -0.215; TCGA COAD -0.779; TCGA ESCA -0.523; TCGA HNSC -0.243; TCGA KIRP -0.418; TCGA LGG -0.198; TCGA LIHC -0.299; TCGA LUAD -0.476; TCGA LUSC -0.452; TCGA PRAD -0.351; TCGA STAD -0.409; TCGA UCEC -0.335 |
hsa-miR-106a-5p | PDE7B | 9 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.188; TCGA BRCA -0.193; TCGA CESC -0.113; TCGA COAD -0.461; TCGA HNSC -0.169; TCGA LUSC -0.336; TCGA PRAD -0.302; TCGA STAD -0.386; TCGA UCEC -0.265 |
hsa-miR-106a-5p | COL8A2 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LUAD; LUSC; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.442; TCGA BRCA -0.138; TCGA CESC -0.207; TCGA COAD -0.714; TCGA ESCA -0.23; TCGA HNSC -0.191; TCGA LGG -0.197; TCGA LUAD -0.194; TCGA LUSC -0.335; TCGA PAAD -0.411; TCGA PRAD -0.233; TCGA THCA -0.637; TCGA STAD -0.39; TCGA UCEC -0.149 |
hsa-miR-106a-5p | ZMAT3 | 9 cancers: BLCA; BRCA; COAD; HNSC; LIHC; LUAD; PRAD; THCA; UCEC | mirMAP | TCGA BLCA -0.142; TCGA BRCA -0.079; TCGA COAD -0.269; TCGA HNSC -0.085; TCGA LIHC -0.108; TCGA LUAD -0.175; TCGA PRAD -0.311; TCGA THCA -0.353; TCGA UCEC -0.109 |
hsa-miR-106a-5p | MSRB3 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.581; TCGA BRCA -0.208; TCGA CESC -0.35; TCGA COAD -0.767; TCGA ESCA -0.315; TCGA HNSC -0.276; TCGA LUAD -0.272; TCGA LUSC -0.432; TCGA PRAD -0.354; TCGA STAD -0.458; TCGA UCEC -0.296 |
hsa-miR-106a-5p | ZHX3 | 13 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LIHC; LUAD; LUSC; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.134; TCGA BRCA -0.071; TCGA CESC -0.061; TCGA ESCA -0.095; TCGA HNSC -0.143; TCGA LGG -0.137; TCGA LIHC -0.138; TCGA LUAD -0.084; TCGA LUSC -0.131; TCGA PRAD -0.077; TCGA SARC -0.142; TCGA STAD -0.128; TCGA UCEC -0.157 |
hsa-miR-106a-5p | LRRN4CL | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.462; TCGA BRCA -0.377; TCGA CESC -0.189; TCGA COAD -0.559; TCGA ESCA -0.315; TCGA HNSC -0.263; TCGA LUAD -0.271; TCGA LUSC -0.402; TCGA OV -0.235; TCGA PRAD -0.106; TCGA STAD -0.445; TCGA UCEC -0.36 |
hsa-miR-106a-5p | PTRF | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PRAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.479; TCGA BRCA -0.162; TCGA CESC -0.334; TCGA COAD -0.464; TCGA ESCA -0.29; TCGA HNSC -0.243; TCGA KIRC -0.212; TCGA KIRP -0.129; TCGA LUAD -0.218; TCGA LUSC -0.312; TCGA PRAD -0.228; TCGA SARC -0.06; TCGA THCA -0.158; TCGA STAD -0.371; TCGA UCEC -0.246 |
hsa-miR-106a-5p | MAF | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; STAD; UCEC | mirMAP | TCGA BLCA -0.305; TCGA BRCA -0.107; TCGA CESC -0.292; TCGA COAD -0.431; TCGA HNSC -0.186; TCGA LGG -0.12; TCGA LUAD -0.103; TCGA LUSC -0.168; TCGA STAD -0.208; TCGA UCEC -0.235 |
hsa-miR-106a-5p | GREM2 | 9 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUSC; OV; STAD | mirMAP | TCGA BLCA -0.444; TCGA BRCA -0.37; TCGA CESC -0.239; TCGA COAD -1.07; TCGA ESCA -0.482; TCGA HNSC -0.356; TCGA LUSC -0.524; TCGA OV -0.24; TCGA STAD -0.903 |
hsa-miR-106a-5p | MXRA7 | 11 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; OV; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.284; TCGA BRCA -0.105; TCGA CESC -0.144; TCGA COAD -0.357; TCGA HNSC -0.108; TCGA LUAD -0.197; TCGA LUSC -0.087; TCGA OV -0.127; TCGA PRAD -0.2; TCGA STAD -0.324; TCGA UCEC -0.175 |
hsa-miR-106a-5p | CCBE1 | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.191; TCGA BRCA -0.142; TCGA CESC -0.575; TCGA COAD -0.944; TCGA HNSC -0.366; TCGA LUAD -0.23; TCGA LUSC -0.633; TCGA PRAD -0.5; TCGA STAD -0.205; TCGA UCEC -0.314 |
hsa-miR-106a-5p | RBM43 | 13 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRP; LGG; LUAD; LUSC; OV; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.099; TCGA BRCA -0.113; TCGA CESC -0.078; TCGA COAD -0.196; TCGA HNSC -0.09; TCGA KIRP -0.094; TCGA LGG -0.117; TCGA LUAD -0.183; TCGA LUSC -0.156; TCGA OV -0.058; TCGA PRAD -0.243; TCGA STAD -0.16; TCGA UCEC -0.151 |
hsa-miR-106a-5p | SRL | 11 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.408; TCGA BRCA -0.189; TCGA COAD -0.434; TCGA ESCA -0.229; TCGA HNSC -0.407; TCGA LUAD -0.239; TCGA LUSC -0.284; TCGA PRAD -0.289; TCGA THCA -0.332; TCGA STAD -0.321; TCGA UCEC -0.39 |
hsa-miR-106a-5p | BCL9L | 12 cancers: BLCA; CESC; COAD; HNSC; KIRC; KIRP; LGG; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.092; TCGA CESC -0.174; TCGA COAD -0.158; TCGA HNSC -0.174; TCGA KIRC -0.113; TCGA KIRP -0.082; TCGA LGG -0.133; TCGA LUSC -0.158; TCGA PRAD -0.075; TCGA THCA -0.121; TCGA STAD -0.075; TCGA UCEC -0.064 |
hsa-miR-106a-5p | TEAD1 | 9 cancers: BLCA; COAD; HNSC; LGG; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.121; TCGA COAD -0.115; TCGA HNSC -0.052; TCGA LGG -0.126; TCGA LUSC -0.104; TCGA PRAD -0.261; TCGA THCA -0.163; TCGA STAD -0.175; TCGA UCEC -0.115 |
hsa-miR-106a-5p | THSD4 | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; THCA; STAD | mirMAP | TCGA BLCA -0.226; TCGA BRCA -0.397; TCGA CESC -0.253; TCGA COAD -0.284; TCGA HNSC -0.249; TCGA LUAD -0.182; TCGA LUSC -0.343; TCGA PRAD -0.26; TCGA THCA -0.135; TCGA STAD -0.381 |
hsa-miR-106a-5p | PRELP | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.751; TCGA BRCA -0.146; TCGA CESC -0.302; TCGA COAD -1.155; TCGA ESCA -0.522; TCGA HNSC -0.173; TCGA LUAD -0.387; TCGA LUSC -0.533; TCGA PAAD -0.265; TCGA PRAD -0.274; TCGA STAD -0.85; TCGA UCEC -0.358 |
hsa-miR-106a-5p | GM2A | 11 cancers: BLCA; CESC; COAD; HNSC; KIRC; LUAD; LUSC; PRAD; SARC; THCA; STAD | mirMAP | TCGA BLCA -0.117; TCGA CESC -0.181; TCGA COAD -0.118; TCGA HNSC -0.107; TCGA KIRC -0.084; TCGA LUAD -0.056; TCGA LUSC -0.101; TCGA PRAD -0.068; TCGA SARC -0.065; TCGA THCA -0.08; TCGA STAD -0.11 |
hsa-miR-106a-5p | MYO5A | 9 cancers: BLCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.156; TCGA CESC -0.254; TCGA COAD -0.45; TCGA HNSC -0.25; TCGA LUAD -0.094; TCGA LUSC -0.147; TCGA PRAD -0.128; TCGA STAD -0.135; TCGA UCEC -0.097 |
hsa-miR-106a-5p | ENPP1 | 9 cancers: BLCA; BRCA; HNSC; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.343; TCGA BRCA -0.122; TCGA HNSC -0.144; TCGA LUSC -0.167; TCGA OV -0.154; TCGA PAAD -0.327; TCGA PRAD -0.094; TCGA STAD -0.164; TCGA UCEC -0.195 |
hsa-miR-106a-5p | MYO1C | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.103; TCGA BRCA -0.068; TCGA CESC -0.062; TCGA COAD -0.181; TCGA ESCA -0.1; TCGA HNSC -0.135; TCGA KIRC -0.118; TCGA KIRP -0.07; TCGA LIHC -0.059; TCGA LUAD -0.101; TCGA LUSC -0.222; TCGA OV -0.111; TCGA THCA -0.181; TCGA STAD -0.139; TCGA UCEC -0.102 |
hsa-miR-106a-5p | DMD | 9 cancers: BLCA; CESC; COAD; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.449; TCGA CESC -0.299; TCGA COAD -0.269; TCGA LUAD -0.084; TCGA LUSC -0.202; TCGA PRAD -0.358; TCGA THCA -0.255; TCGA STAD -0.359; TCGA UCEC -0.198 |
hsa-miR-106a-5p | GLIS3 | 9 cancers: BLCA; COAD; HNSC; KIRP; LGG; LUAD; LUSC; PRAD; SARC | miRNATAP | TCGA BLCA -0.202; TCGA COAD -0.777; TCGA HNSC -0.229; TCGA KIRP -0.283; TCGA LGG -0.235; TCGA LUAD -0.134; TCGA LUSC -0.343; TCGA PRAD -0.344; TCGA SARC -0.227 |
hsa-miR-106a-5p | CELF2 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LGG; LUAD; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.476; TCGA BRCA -0.067; TCGA CESC -0.231; TCGA COAD -0.13; TCGA ESCA -0.197; TCGA KIRC -0.105; TCGA KIRP -0.162; TCGA LGG -0.088; TCGA LUAD -0.203; TCGA LUSC -0.344; TCGA PRAD -0.29; TCGA STAD -0.318; TCGA UCEC -0.121 |
hsa-miR-106a-5p | SLITRK3 | 9 cancers: BLCA; COAD; ESCA; LGG; OV; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.536; TCGA COAD -1.201; TCGA ESCA -0.832; TCGA LGG -0.375; TCGA OV -0.234; TCGA PAAD -0.413; TCGA PRAD -0.543; TCGA STAD -1.055; TCGA UCEC -0.503 |
hsa-miR-106a-5p | SLC40A1 | 9 cancers: BLCA; BRCA; HNSC; LIHC; LUAD; LUSC; OV; PRAD; THCA | miRNATAP | TCGA BLCA -0.152; TCGA BRCA -0.184; TCGA HNSC -0.158; TCGA LIHC -0.081; TCGA LUAD -0.292; TCGA LUSC -0.254; TCGA OV -0.138; TCGA PRAD -0.222; TCGA THCA -0.124 |
hsa-miR-106a-5p | PTPRD | 10 cancers: BLCA; CESC; HNSC; KIRP; LGG; LUAD; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.485; TCGA CESC -0.356; TCGA HNSC -0.402; TCGA KIRP -0.243; TCGA LGG -0.217; TCGA LUAD -0.194; TCGA LUSC -0.275; TCGA PRAD -0.297; TCGA STAD -0.314; TCGA UCEC -0.186 |
hsa-miR-106a-5p | ATXN1 | 10 cancers: BLCA; COAD; HNSC; KIRC; KIRP; LUAD; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.276; TCGA COAD -0.238; TCGA HNSC -0.169; TCGA KIRC -0.091; TCGA KIRP -0.104; TCGA LUAD -0.095; TCGA LUSC -0.162; TCGA PRAD -0.179; TCGA STAD -0.186; TCGA UCEC -0.096 |
hsa-miR-106a-5p | BNC2 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUAD; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.601; TCGA BRCA -0.171; TCGA CESC -0.269; TCGA COAD -0.85; TCGA ESCA -0.409; TCGA HNSC -0.269; TCGA KIRP -0.13; TCGA LUAD -0.192; TCGA LUSC -0.404; TCGA PRAD -0.336; TCGA STAD -0.489; TCGA UCEC -0.467 |
hsa-miR-106a-5p | ATXN1L | 11 cancers: BLCA; CESC; COAD; HNSC; LGG; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.058; TCGA CESC -0.062; TCGA COAD -0.065; TCGA HNSC -0.077; TCGA LGG -0.065; TCGA LIHC -0.107; TCGA LUAD -0.104; TCGA LUSC -0.073; TCGA PRAD -0.189; TCGA STAD -0.064; TCGA UCEC -0.117 |
hsa-miR-106a-5p | PTEN | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.119; TCGA BRCA -0.054; TCGA CESC -0.121; TCGA COAD -0.135; TCGA ESCA -0.083; TCGA HNSC -0.106; TCGA LIHC -0.064; TCGA LUAD -0.131; TCGA LUSC -0.159; TCGA OV -0.109; TCGA PRAD -0.091; TCGA SARC -0.066; TCGA STAD -0.07; TCGA UCEC -0.139 |
hsa-miR-106a-5p | FCHO2 | 9 cancers: BLCA; BRCA; COAD; LUAD; LUSC; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.085; TCGA BRCA -0.062; TCGA COAD -0.103; TCGA LUAD -0.185; TCGA LUSC -0.199; TCGA PRAD -0.149; TCGA THCA -0.117; TCGA STAD -0.078; TCGA UCEC -0.146 |
hsa-miR-106a-5p | PTGER3 | 13 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; OV; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.441; TCGA BRCA -0.38; TCGA CESC -0.462; TCGA COAD -0.954; TCGA HNSC -0.324; TCGA LGG -0.255; TCGA LUAD -0.246; TCGA LUSC -0.385; TCGA OV -0.34; TCGA PRAD -0.37; TCGA THCA -0.115; TCGA STAD -0.432; TCGA UCEC -0.547 |
hsa-miR-106a-5p | SPRED1 | 10 cancers: BLCA; CESC; COAD; HNSC; KIRC; LUAD; LUSC; PAAD; PRAD; UCEC | miRNATAP | TCGA BLCA -0.183; TCGA CESC -0.1; TCGA COAD -0.124; TCGA HNSC -0.073; TCGA KIRC -0.089; TCGA LUAD -0.22; TCGA LUSC -0.207; TCGA PAAD -0.123; TCGA PRAD -0.186; TCGA UCEC -0.115 |
hsa-miR-106a-5p | KCNMA1 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LUAD; LUSC; PRAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.517; TCGA BRCA -0.287; TCGA CESC -0.219; TCGA COAD -0.913; TCGA ESCA -0.552; TCGA KIRC -0.25; TCGA LUAD -0.175; TCGA LUSC -0.28; TCGA PRAD -0.308; TCGA SARC -0.187; TCGA THCA -0.217; TCGA STAD -0.751; TCGA UCEC -0.341 |
hsa-miR-106a-5p | FOXN3 | 10 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LGG; LUAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.108; TCGA BRCA -0.065; TCGA COAD -0.196; TCGA ESCA -0.094; TCGA HNSC -0.111; TCGA LGG -0.091; TCGA LUAD -0.09; TCGA PRAD -0.139; TCGA STAD -0.199; TCGA UCEC -0.135 |
hsa-miR-106a-5p | YPEL2 | 12 cancers: BLCA; BRCA; COAD; HNSC; KIRC; KIRP; LUAD; LUSC; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.056; TCGA BRCA -0.059; TCGA COAD -0.146; TCGA HNSC -0.112; TCGA KIRC -0.11; TCGA KIRP -0.195; TCGA LUAD -0.099; TCGA LUSC -0.116; TCGA PRAD -0.095; TCGA THCA -0.059; TCGA STAD -0.077; TCGA UCEC -0.184 |
hsa-miR-106a-5p | NFIC | 11 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LGG; LIHC; LUAD; LUSC; PRAD; STAD | miRNATAP | TCGA BLCA -0.142; TCGA BRCA -0.051; TCGA COAD -0.255; TCGA ESCA -0.11; TCGA HNSC -0.099; TCGA LGG -0.138; TCGA LIHC -0.172; TCGA LUAD -0.197; TCGA LUSC -0.149; TCGA PRAD -0.249; TCGA STAD -0.155 |
hsa-miR-106a-5p | AFF4 | 9 cancers: BLCA; COAD; HNSC; LGG; LIHC; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.064; TCGA COAD -0.1; TCGA HNSC -0.092; TCGA LGG -0.08; TCGA LIHC -0.063; TCGA LUSC -0.1; TCGA PRAD -0.149; TCGA STAD -0.071; TCGA UCEC -0.087 |
hsa-miR-106a-5p | RBMS2 | 13 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.118; TCGA CESC -0.101; TCGA COAD -0.177; TCGA ESCA -0.139; TCGA HNSC -0.148; TCGA KIRC -0.153; TCGA KIRP -0.119; TCGA LUAD -0.115; TCGA LUSC -0.265; TCGA PRAD -0.158; TCGA THCA -0.126; TCGA STAD -0.074; TCGA UCEC -0.15 |
hsa-miR-106a-5p | HEG1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.31; TCGA BRCA -0.098; TCGA CESC -0.365; TCGA COAD -0.493; TCGA ESCA -0.136; TCGA HNSC -0.206; TCGA LUAD -0.218; TCGA LUSC -0.426; TCGA PRAD -0.282; TCGA THCA -0.125; TCGA STAD -0.228; TCGA UCEC -0.15 |
hsa-miR-106a-5p | ZFPM2 | 10 cancers: BLCA; BRCA; COAD; ESCA; LUAD; LUSC; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.627; TCGA BRCA -0.159; TCGA COAD -0.71; TCGA ESCA -0.278; TCGA LUAD -0.285; TCGA LUSC -0.467; TCGA PRAD -0.346; TCGA SARC -0.295; TCGA STAD -0.398; TCGA UCEC -0.418 |
hsa-miR-106a-5p | SKI | 12 cancers: BLCA; CESC; COAD; HNSC; KIRC; KIRP; LUAD; LUSC; OV; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.083; TCGA CESC -0.106; TCGA COAD -0.082; TCGA HNSC -0.054; TCGA KIRC -0.15; TCGA KIRP -0.078; TCGA LUAD -0.091; TCGA LUSC -0.175; TCGA OV -0.064; TCGA PRAD -0.067; TCGA STAD -0.084; TCGA UCEC -0.081 |
hsa-miR-106a-5p | TCF4 | 11 cancers: BLCA; BRCA; COAD; HNSC; KIRC; LGG; LUAD; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.234; TCGA BRCA -0.119; TCGA COAD -0.5; TCGA HNSC -0.094; TCGA KIRC -0.284; TCGA LGG -0.191; TCGA LUAD -0.172; TCGA LUSC -0.092; TCGA PRAD -0.211; TCGA STAD -0.156; TCGA UCEC -0.1 |
hsa-miR-106a-5p | TIMP3 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LIHC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.355; TCGA BRCA -0.211; TCGA CESC -0.458; TCGA COAD -0.532; TCGA ESCA -0.338; TCGA HNSC -0.429; TCGA LGG -0.152; TCGA LIHC -0.134; TCGA LUAD -0.288; TCGA LUSC -0.6; TCGA OV -0.188; TCGA PAAD -0.212; TCGA PRAD -0.263; TCGA STAD -0.32; TCGA UCEC -0.32 |
hsa-miR-106a-5p | OSBPL5 | 10 cancers: BLCA; BRCA; ESCA; HNSC; KIRC; LUAD; LUSC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.172; TCGA BRCA -0.065; TCGA ESCA -0.267; TCGA HNSC -0.13; TCGA KIRC -0.119; TCGA LUAD -0.095; TCGA LUSC -0.187; TCGA THCA -0.124; TCGA STAD -0.207; TCGA UCEC -0.154 |
hsa-miR-106a-5p | EPAS1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.179; TCGA BRCA -0.104; TCGA CESC -0.175; TCGA COAD -0.199; TCGA ESCA -0.165; TCGA HNSC -0.101; TCGA KIRC -0.198; TCGA LUAD -0.133; TCGA LUSC -0.474; TCGA PRAD -0.172; TCGA STAD -0.188; TCGA UCEC -0.191 |
hsa-miR-106a-5p | DNAJB9 | 11 cancers: BLCA; BRCA; COAD; KIRP; LIHC; LUAD; LUSC; OV; PRAD; SARC; UCEC | miRNATAP | TCGA BLCA -0.11; TCGA BRCA -0.072; TCGA COAD -0.092; TCGA KIRP -0.071; TCGA LIHC -0.115; TCGA LUAD -0.15; TCGA LUSC -0.157; TCGA OV -0.098; TCGA PRAD -0.116; TCGA SARC -0.06; TCGA UCEC -0.103 |
hsa-miR-106a-5p | CSF1 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LGG; LUAD; LUSC; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.292; TCGA BRCA -0.113; TCGA CESC -0.207; TCGA COAD -0.323; TCGA ESCA -0.17; TCGA HNSC -0.098; TCGA KIRC -0.105; TCGA LGG -0.241; TCGA LUAD -0.158; TCGA LUSC -0.367; TCGA PRAD -0.227; TCGA THCA -0.23; TCGA STAD -0.147; TCGA UCEC -0.109 |
hsa-miR-106a-5p | NBL1 | 15 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; OV; PAAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.31; TCGA BRCA -0.202; TCGA CESC -0.086; TCGA ESCA -0.267; TCGA HNSC -0.168; TCGA KIRC -0.157; TCGA KIRP -0.487; TCGA LUAD -0.131; TCGA LUSC -0.322; TCGA OV -0.124; TCGA PAAD -0.269; TCGA SARC -0.269; TCGA THCA -0.193; TCGA STAD -0.214; TCGA UCEC -0.233 |
hsa-miR-106a-5p | OTUD1 | 13 cancers: BLCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; OV; PRAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.133; TCGA CESC -0.202; TCGA COAD -0.132; TCGA HNSC -0.067; TCGA LGG -0.151; TCGA LUAD -0.087; TCGA LUSC -0.124; TCGA OV -0.134; TCGA PRAD -0.07; TCGA SARC -0.074; TCGA THCA -0.087; TCGA STAD -0.086; TCGA UCEC -0.219 |
hsa-miR-106a-5p | CCND2 | 11 cancers: BLCA; BRCA; CESC; ESCA; KIRC; KIRP; LUAD; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.46; TCGA BRCA -0.117; TCGA CESC -0.445; TCGA ESCA -0.196; TCGA KIRC -0.225; TCGA KIRP -0.447; TCGA LUAD -0.149; TCGA PRAD -0.251; TCGA THCA -0.233; TCGA STAD -0.174; TCGA UCEC -0.314 |
hsa-miR-106a-5p | RBMS1 | 9 cancers: BLCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.073; TCGA CESC -0.137; TCGA COAD -0.407; TCGA HNSC -0.094; TCGA LUAD -0.074; TCGA LUSC -0.141; TCGA PRAD -0.285; TCGA STAD -0.113; TCGA UCEC -0.161 |
hsa-miR-106a-5p | MYO1D | 11 cancers: BLCA; ESCA; HNSC; KIRP; LUAD; LUSC; PAAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.08; TCGA ESCA -0.129; TCGA HNSC -0.126; TCGA KIRP -0.13; TCGA LUAD -0.074; TCGA LUSC -0.131; TCGA PAAD -0.176; TCGA SARC -0.144; TCGA THCA -0.286; TCGA STAD -0.106; TCGA UCEC -0.062 |
hsa-miR-106a-5p | ISM1 | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; PAAD; PRAD; STAD | miRNATAP | TCGA BLCA -0.277; TCGA BRCA -0.215; TCGA CESC -0.219; TCGA COAD -1.026; TCGA HNSC -0.332; TCGA LUAD -0.337; TCGA LUSC -0.501; TCGA PAAD -0.45; TCGA PRAD -0.229; TCGA STAD -0.581 |
hsa-miR-106a-5p | CALD1 | 13 cancers: BLCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.494; TCGA CESC -0.356; TCGA COAD -0.633; TCGA ESCA -0.242; TCGA HNSC -0.253; TCGA LIHC -0.089; TCGA LUAD -0.167; TCGA LUSC -0.363; TCGA OV -0.078; TCGA PRAD -0.305; TCGA THCA -0.102; TCGA STAD -0.339; TCGA UCEC -0.251 |
hsa-miR-106a-5p | UBR1 | 9 cancers: BLCA; BRCA; COAD; HNSC; LGG; LUAD; LUSC; PRAD; UCEC | miRNATAP | TCGA BLCA -0.083; TCGA BRCA -0.075; TCGA COAD -0.079; TCGA HNSC -0.091; TCGA LGG -0.059; TCGA LUAD -0.092; TCGA LUSC -0.116; TCGA PRAD -0.182; TCGA UCEC -0.118 |
hsa-miR-106a-5p | BMP2 | 9 cancers: BLCA; CESC; COAD; KIRC; LGG; LUAD; LUSC; PRAD; UCEC | miRNATAP | TCGA BLCA -0.137; TCGA CESC -0.285; TCGA COAD -0.332; TCGA KIRC -0.123; TCGA LGG -0.379; TCGA LUAD -0.25; TCGA LUSC -0.555; TCGA PRAD -0.247; TCGA UCEC -0.193 |
hsa-miR-106a-5p | TRIP10 | 10 cancers: BLCA; CESC; ESCA; KIRC; KIRP; LUAD; LUSC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.061; TCGA CESC -0.089; TCGA ESCA -0.188; TCGA KIRC -0.121; TCGA KIRP -0.084; TCGA LUAD -0.051; TCGA LUSC -0.083; TCGA THCA -0.07; TCGA STAD -0.104; TCGA UCEC -0.07 |
hsa-miR-106a-5p | SMAD7 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.125; TCGA BRCA -0.053; TCGA CESC -0.15; TCGA COAD -0.164; TCGA HNSC -0.055; TCGA LGG -0.086; TCGA LUAD -0.126; TCGA LUSC -0.312; TCGA PAAD -0.126; TCGA PRAD -0.141; TCGA STAD -0.1; TCGA UCEC -0.117 |
hsa-miR-106a-5p | AHNAK | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LGG; LUAD; LUSC; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.313; TCGA BRCA -0.185; TCGA CESC -0.22; TCGA COAD -0.355; TCGA ESCA -0.234; TCGA HNSC -0.23; TCGA KIRP -0.098; TCGA LGG -0.245; TCGA LUAD -0.31; TCGA LUSC -0.313; TCGA PRAD -0.187; TCGA SARC -0.086; TCGA STAD -0.338; TCGA UCEC -0.245 |
hsa-miR-106a-5p | TIMP2 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PRAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.411; TCGA BRCA -0.104; TCGA CESC -0.197; TCGA COAD -0.694; TCGA ESCA -0.157; TCGA HNSC -0.323; TCGA LUAD -0.171; TCGA LUSC -0.428; TCGA OV -0.08; TCGA PRAD -0.222; TCGA SARC -0.078; TCGA THCA -0.122; TCGA STAD -0.311; TCGA UCEC -0.231 |
hsa-miR-106a-5p | JAM2 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LGG; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.435; TCGA BRCA -0.216; TCGA CESC -0.236; TCGA COAD -0.79; TCGA ESCA -0.325; TCGA HNSC -0.202; TCGA KIRC -0.125; TCGA LGG -0.131; TCGA LIHC -0.123; TCGA LUAD -0.221; TCGA LUSC -0.401; TCGA OV -0.143; TCGA PRAD -0.189; TCGA STAD -0.505; TCGA UCEC -0.214 |
hsa-miR-106a-5p | ZNF25 | 13 cancers: BLCA; BRCA; CESC; COAD; KIRC; LIHC; LUAD; LUSC; OV; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.144; TCGA BRCA -0.063; TCGA CESC -0.083; TCGA COAD -0.226; TCGA KIRC -0.067; TCGA LIHC -0.118; TCGA LUAD -0.105; TCGA LUSC -0.098; TCGA OV -0.052; TCGA PRAD -0.103; TCGA SARC -0.07; TCGA STAD -0.218; TCGA UCEC -0.128 |
hsa-miR-106a-5p | UBC | 11 cancers: BLCA; CESC; ESCA; HNSC; KIRC; LIHC; LUAD; LUSC; OV; SARC; STAD | miRNATAP | TCGA BLCA -0.082; TCGA CESC -0.072; TCGA ESCA -0.104; TCGA HNSC -0.074; TCGA KIRC -0.109; TCGA LIHC -0.089; TCGA LUAD -0.069; TCGA LUSC -0.058; TCGA OV -0.055; TCGA SARC -0.059; TCGA STAD -0.114 |
hsa-miR-106a-5p | KIF13A | 12 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LGG; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.15; TCGA BRCA -0.089; TCGA COAD -0.102; TCGA ESCA -0.15; TCGA HNSC -0.179; TCGA LGG -0.143; TCGA LUAD -0.188; TCGA LUSC -0.057; TCGA PAAD -0.168; TCGA PRAD -0.198; TCGA STAD -0.081; TCGA UCEC -0.104 |
hsa-miR-106a-5p | GPR137B | 10 cancers: BLCA; CESC; COAD; HNSC; LGG; OV; PAAD; PRAD; THCA; UCEC | miRNATAP | TCGA BLCA -0.067; TCGA CESC -0.088; TCGA COAD -0.281; TCGA HNSC -0.084; TCGA LGG -0.13; TCGA OV -0.1; TCGA PAAD -0.155; TCGA PRAD -0.162; TCGA THCA -0.106; TCGA UCEC -0.154 |
hsa-miR-106a-5p | AKT3 | 10 cancers: BLCA; CESC; COAD; KIRC; LUAD; OV; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.236; TCGA CESC -0.123; TCGA COAD -0.549; TCGA KIRC -0.083; TCGA LUAD -0.088; TCGA OV -0.16; TCGA PRAD -0.098; TCGA THCA -0.117; TCGA STAD -0.239; TCGA UCEC -0.317 |
hsa-miR-106a-5p | FBXL5 | 13 cancers: BLCA; BRCA; COAD; ESCA; HNSC; KIRC; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.075; TCGA BRCA -0.122; TCGA COAD -0.138; TCGA ESCA -0.069; TCGA HNSC -0.054; TCGA KIRC -0.08; TCGA LIHC -0.135; TCGA LUAD -0.104; TCGA LUSC -0.109; TCGA OV -0.134; TCGA PRAD -0.112; TCGA STAD -0.099; TCGA UCEC -0.09 |
hsa-miR-106a-5p | LRRK1 | 11 cancers: BLCA; CESC; ESCA; KIRC; KIRP; LUAD; LUSC; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.105; TCGA CESC -0.283; TCGA ESCA -0.105; TCGA KIRC -0.316; TCGA KIRP -0.13; TCGA LUAD -0.124; TCGA LUSC -0.087; TCGA PRAD -0.209; TCGA THCA -0.104; TCGA STAD -0.093; TCGA UCEC -0.133 |
hsa-miR-106a-5p | FYN | 10 cancers: BLCA; CESC; COAD; KIRC; LGG; LUAD; LUSC; PRAD; THCA; UCEC | miRNATAP | TCGA BLCA -0.136; TCGA CESC -0.21; TCGA COAD -0.282; TCGA KIRC -0.168; TCGA LGG -0.087; TCGA LUAD -0.1; TCGA LUSC -0.166; TCGA PRAD -0.139; TCGA THCA -0.249; TCGA UCEC -0.125 |
hsa-miR-106a-5p | CAMTA2 | 10 cancers: BRCA; COAD; HNSC; KIRC; KIRP; LIHC; LUSC; SARC; STAD; UCEC | MirTarget | TCGA BRCA -0.08; TCGA COAD -0.105; TCGA HNSC -0.054; TCGA KIRC -0.076; TCGA KIRP -0.102; TCGA LIHC -0.06; TCGA LUSC -0.065; TCGA SARC -0.056; TCGA STAD -0.07; TCGA UCEC -0.069 |
hsa-miR-106a-5p | MINK1 | 12 cancers: BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LUSC; OV; PRAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.057; TCGA CESC -0.134; TCGA ESCA -0.1; TCGA HNSC -0.067; TCGA KIRC -0.069; TCGA KIRP -0.15; TCGA LUSC -0.111; TCGA OV -0.054; TCGA PRAD -0.075; TCGA THCA -0.074; TCGA STAD -0.069; TCGA UCEC -0.064 |
hsa-miR-106a-5p | PLEKHM1 | 11 cancers: BRCA; ESCA; HNSC; LIHC; LUAD; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.077; TCGA ESCA -0.093; TCGA HNSC -0.113; TCGA LIHC -0.061; TCGA LUAD -0.087; TCGA PAAD -0.094; TCGA PRAD -0.055; TCGA SARC -0.064; TCGA THCA -0.081; TCGA STAD -0.117; TCGA UCEC -0.052 |
hsa-miR-106a-5p | CASC4 | 12 cancers: BRCA; COAD; HNSC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; PRAD; SARC; UCEC | MirTarget | TCGA BRCA -0.085; TCGA COAD -0.089; TCGA HNSC -0.059; TCGA KIRP -0.061; TCGA LIHC -0.061; TCGA LUAD -0.118; TCGA LUSC -0.117; TCGA OV -0.103; TCGA PAAD -0.111; TCGA PRAD -0.131; TCGA SARC -0.074; TCGA UCEC -0.134 |
hsa-miR-106a-5p | GAB1 | 9 cancers: BRCA; CESC; HNSC; LGG; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.075; TCGA CESC -0.077; TCGA HNSC -0.122; TCGA LGG -0.158; TCGA LUAD -0.107; TCGA LUSC -0.116; TCGA PRAD -0.228; TCGA STAD -0.191; TCGA UCEC -0.168 |
hsa-miR-106a-5p | WDFY3 | 10 cancers: BRCA; COAD; HNSC; KIRP; LGG; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.055; TCGA COAD -0.093; TCGA HNSC -0.089; TCGA KIRP -0.099; TCGA LGG -0.163; TCGA LUAD -0.083; TCGA LUSC -0.051; TCGA PRAD -0.138; TCGA STAD -0.123; TCGA UCEC -0.06 |
hsa-miR-106a-5p | GCNT4 | 11 cancers: BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BRCA -0.237; TCGA CESC -0.238; TCGA COAD -0.636; TCGA HNSC -0.214; TCGA LGG -0.182; TCGA LUAD -0.285; TCGA LUSC -0.265; TCGA PRAD -0.575; TCGA THCA -0.42; TCGA STAD -0.405; TCGA UCEC -0.378 |
hsa-miR-106a-5p | APCDD1 | 9 cancers: BRCA; CESC; HNSC; KIRC; LGG; LUAD; LUSC; PRAD; SARC | MirTarget | TCGA BRCA -0.124; TCGA CESC -0.151; TCGA HNSC -0.234; TCGA KIRC -0.135; TCGA LGG -0.116; TCGA LUAD -0.12; TCGA LUSC -0.227; TCGA PRAD -0.215; TCGA SARC -0.215 |
hsa-miR-106a-5p | SUMF1 | 9 cancers: BRCA; CESC; KIRP; LIHC; LUAD; LUSC; OV; SARC; STAD | MirTarget | TCGA BRCA -0.068; TCGA CESC -0.072; TCGA KIRP -0.067; TCGA LIHC -0.072; TCGA LUAD -0.069; TCGA LUSC -0.103; TCGA OV -0.12; TCGA SARC -0.07; TCGA STAD -0.078 |
hsa-miR-106a-5p | MARCH8 | 10 cancers: BRCA; COAD; HNSC; LGG; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.139; TCGA COAD -0.113; TCGA HNSC -0.077; TCGA LGG -0.342; TCGA LIHC -0.151; TCGA LUAD -0.143; TCGA LUSC -0.131; TCGA PRAD -0.274; TCGA STAD -0.111; TCGA UCEC -0.182 |
hsa-miR-106a-5p | NTN4 | 12 cancers: BRCA; CESC; COAD; HNSC; LGG; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.249; TCGA CESC -0.262; TCGA COAD -0.215; TCGA HNSC -0.087; TCGA LGG -0.17; TCGA LIHC -0.165; TCGA LUAD -0.24; TCGA LUSC -0.388; TCGA OV -0.148; TCGA PRAD -0.322; TCGA STAD -0.309; TCGA UCEC -0.251 |
hsa-miR-106a-5p | QSOX1 | 10 cancers: BRCA; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; PRAD; SARC; THCA | mirMAP | TCGA BRCA -0.066; TCGA COAD -0.203; TCGA ESCA -0.207; TCGA HNSC -0.173; TCGA KIRC -0.168; TCGA LUAD -0.082; TCGA LUSC -0.271; TCGA PRAD -0.227; TCGA SARC -0.096; TCGA THCA -0.424 |
hsa-miR-106a-5p | AR | 10 cancers: BRCA; COAD; HNSC; KIRP; LGG; LIHC; LUAD; LUSC; PRAD; STAD | mirMAP | TCGA BRCA -0.445; TCGA COAD -0.824; TCGA HNSC -0.218; TCGA KIRP -0.265; TCGA LGG -0.469; TCGA LIHC -0.483; TCGA LUAD -0.319; TCGA LUSC -0.524; TCGA PRAD -0.298; TCGA STAD -0.547 |
hsa-miR-106a-5p | ARHGAP23 | 10 cancers: BRCA; CESC; COAD; KIRC; LUAD; OV; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BRCA -0.079; TCGA CESC -0.364; TCGA COAD -0.231; TCGA KIRC -0.104; TCGA LUAD -0.128; TCGA OV -0.088; TCGA PRAD -0.335; TCGA THCA -0.402; TCGA STAD -0.255; TCGA UCEC -0.22 |
hsa-miR-106a-5p | CAMK2N1 | 11 cancers: BRCA; CESC; ESCA; HNSC; LUAD; LUSC; OV; PAAD; PRAD; THCA; UCEC | miRNATAP | TCGA BRCA -0.184; TCGA CESC -0.305; TCGA ESCA -0.256; TCGA HNSC -0.318; TCGA LUAD -0.18; TCGA LUSC -0.506; TCGA OV -0.159; TCGA PAAD -0.222; TCGA PRAD -0.125; TCGA THCA -0.777; TCGA UCEC -0.167 |
hsa-miR-106a-5p | TBC1D9 | 9 cancers: BRCA; COAD; LIHC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BRCA -0.372; TCGA COAD -0.205; TCGA LIHC -0.131; TCGA LUAD -0.078; TCGA LUSC -0.195; TCGA PRAD -0.105; TCGA THCA -0.192; TCGA STAD -0.179; TCGA UCEC -0.057 |
hsa-miR-106a-5p | BHLHE41 | 13 cancers: BRCA; COAD; ESCA; HNSC; KIRC; KIRP; LGG; LUAD; LUSC; OV; PRAD; THCA; STAD | miRNATAP | TCGA BRCA -0.069; TCGA COAD -0.571; TCGA ESCA -0.22; TCGA HNSC -0.163; TCGA KIRC -0.498; TCGA KIRP -0.177; TCGA LGG -0.235; TCGA LUAD -0.223; TCGA LUSC -0.427; TCGA OV -0.242; TCGA PRAD -0.291; TCGA THCA -0.318; TCGA STAD -0.238 |
hsa-miR-106a-5p | ARAP2 | 9 cancers: CESC; HNSC; KIRP; LGG; LUAD; LUSC; OV; PRAD; UCEC | MirTarget | TCGA CESC -0.112; TCGA HNSC -0.079; TCGA KIRP -0.084; TCGA LGG -0.183; TCGA LUAD -0.159; TCGA LUSC -0.177; TCGA OV -0.083; TCGA PRAD -0.104; TCGA UCEC -0.135 |
hsa-miR-106a-5p | HBP1 | 12 cancers: CESC; COAD; KIRP; LGG; LIHC; LUAD; LUSC; OV; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA CESC -0.06; TCGA COAD -0.107; TCGA KIRP -0.07; TCGA LGG -0.089; TCGA LIHC -0.102; TCGA LUAD -0.113; TCGA LUSC -0.068; TCGA OV -0.115; TCGA PRAD -0.07; TCGA SARC -0.066; TCGA STAD -0.091; TCGA UCEC -0.1 |
hsa-miR-106a-5p | PRRG1 | 9 cancers: CESC; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget; miRNATAP | TCGA CESC -0.41; TCGA ESCA -0.149; TCGA HNSC -0.316; TCGA LIHC -0.114; TCGA LUAD -0.122; TCGA LUSC -0.293; TCGA PRAD -0.179; TCGA THCA -0.2; TCGA UCEC -0.188 |
hsa-miR-106a-5p | RNF213 | 9 cancers: CESC; KIRC; KIRP; LGG; LIHC; LUSC; PRAD; THCA; UCEC | mirMAP | TCGA CESC -0.075; TCGA KIRC -0.085; TCGA KIRP -0.174; TCGA LGG -0.114; TCGA LIHC -0.107; TCGA LUSC -0.09; TCGA PRAD -0.061; TCGA THCA -0.109; TCGA UCEC -0.06 |
hsa-miR-106a-5p | TANC2 | 9 cancers: COAD; HNSC; LGG; LUAD; LUSC; PAAD; PRAD; THCA; STAD | mirMAP; miRNATAP | TCGA COAD -0.499; TCGA HNSC -0.135; TCGA LGG -0.125; TCGA LUAD -0.065; TCGA LUSC -0.187; TCGA PAAD -0.155; TCGA PRAD -0.072; TCGA THCA -0.273; TCGA STAD -0.139 |
hsa-miR-106a-5p | RAPGEF2 | 9 cancers: HNSC; KIRC; KIRP; LGG; LIHC; LUAD; LUSC; PRAD; UCEC | MirTarget; miRNATAP | TCGA HNSC -0.075; TCGA KIRC -0.137; TCGA KIRP -0.114; TCGA LGG -0.063; TCGA LIHC -0.069; TCGA LUAD -0.113; TCGA LUSC -0.182; TCGA PRAD -0.112; TCGA UCEC -0.06 |
Enriched cancer pathways of putative targets