microRNA information: hsa-miR-181a-2-3p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-181a-2-3p | miRbase |
| Accession: | MIMAT0004558 | miRbase |
| Precursor name: | hsa-mir-181a-2 | miRbase |
| Precursor accession: | MI0000269 | miRbase |
| Symbol: | NA | HGNC |
| RefSeq ID: | NA | GenBank |
| Sequence: | ACCACUGACCGUUGACUGUACC |
Reported expression in cancers: hsa-miR-181a-2-3p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-181a-2-3p | glioblastoma | deregulation | "Hypoxic signature of microRNAs in glioblastoma: in ......" | 25129238 | RNA-Seq |
| hsa-miR-181a-2-3p | head and neck cancer | deregulation | "A global miRNA profiling was performed on 12 sampl ......" | 21637912 | qPCR; Microarray |
| hsa-miR-181a-2-3p | thyroid cancer | upregulation | "The aim of this study was to assess miRNA expressi ......" | 23427895 | Microarray |
Reported cancer pathway affected by hsa-miR-181a-2-3p
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|
Reported cancer prognosis affected by hsa-miR-181a-2-3p

| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-181a-2-3p | head and neck cancer | malignant trasformation | "A global miRNA profiling was performed on 12 sampl ......" | 21637912 | |
| hsa-miR-181a-2-3p | thyroid cancer | poor survival | "The aim of this study was to assess miRNA expressi ......" | 23427895 |
Reported gene related to hsa-miR-181a-2-3p
| miRNA | cancer | gene | reporting | PUBMED |
|---|
Expression profile in cancer corhorts:
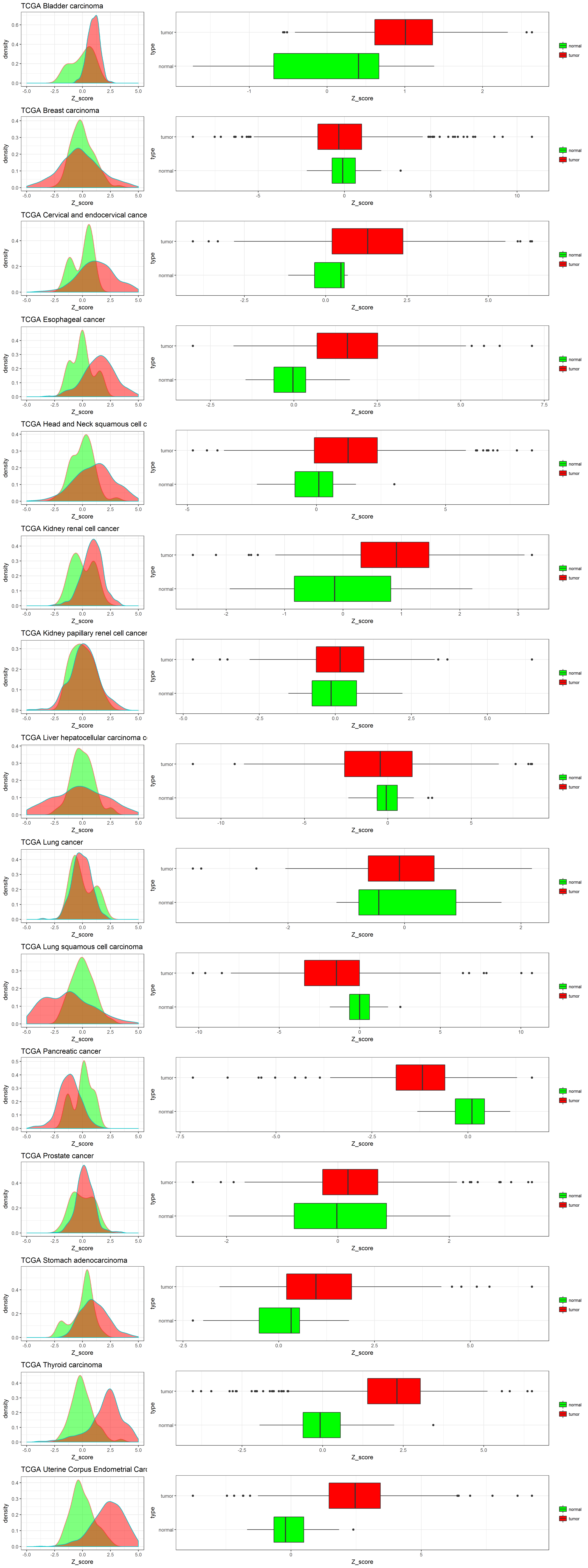
Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-181a-2-3p | SYNJ1 | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; PAAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.065; TCGA BRCA -0.07; TCGA CESC -0.17; TCGA ESCA -0.115; TCGA HNSC -0.109; TCGA PAAD -0.126; TCGA SARC -0.069; TCGA THCA -0.052; TCGA STAD -0.075; TCGA UCEC -0.164 |
hsa-miR-181a-2-3p | EEF2K | 9 cancers: BLCA; BRCA; CESC; HNSC; OV; PRAD; SARC; THCA; UCEC | MirTarget | TCGA BLCA -0.108; TCGA BRCA -0.146; TCGA CESC -0.074; TCGA HNSC -0.063; TCGA OV -0.11; TCGA PRAD -0.129; TCGA SARC -0.09; TCGA THCA -0.067; TCGA UCEC -0.056 |
hsa-miR-181a-2-3p | ATP8A1 | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; OV; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.327; TCGA BRCA -0.297; TCGA CESC -0.218; TCGA ESCA -0.779; TCGA HNSC -0.238; TCGA OV -0.202; TCGA PAAD -0.522; TCGA PRAD -0.185; TCGA STAD -0.444; TCGA UCEC -0.126 |
hsa-miR-181a-2-3p | SEL1L | 11 cancers: BLCA; BRCA; ESCA; KIRC; LIHC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.089; TCGA BRCA -0.125; TCGA ESCA -0.141; TCGA KIRC -0.099; TCGA LIHC -0.06; TCGA LUAD -0.086; TCGA LUSC -0.067; TCGA PRAD -0.184; TCGA THCA -0.104; TCGA STAD -0.123; TCGA UCEC -0.113 |
hsa-miR-181a-2-3p | PPP2R2C | 9 cancers: BLCA; BRCA; CESC; KIRC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.448; TCGA BRCA -0.448; TCGA CESC -0.32; TCGA KIRC -0.727; TCGA LUAD -0.629; TCGA LUSC -0.818; TCGA PRAD -0.308; TCGA THCA -0.541; TCGA UCEC -0.223 |
hsa-miR-181a-2-3p | USP12 | 9 cancers: BLCA; CESC; COAD; ESCA; HNSC; OV; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.222; TCGA CESC -0.158; TCGA COAD -0.16; TCGA ESCA -0.122; TCGA HNSC -0.125; TCGA OV -0.099; TCGA SARC -0.125; TCGA STAD -0.131; TCGA UCEC -0.125 |
hsa-miR-181a-2-3p | MAGI2 | 9 cancers: BLCA; BRCA; HNSC; KIRC; LGG; PAAD; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.201; TCGA BRCA -0.163; TCGA HNSC -0.157; TCGA KIRC -0.415; TCGA LGG -0.08; TCGA PAAD -0.434; TCGA PRAD -0.112; TCGA THCA -0.14; TCGA UCEC -0.139 |
hsa-miR-181a-2-3p | CBX7 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LIHC; OV; PAAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.399; TCGA BRCA -0.293; TCGA CESC -0.35; TCGA COAD -0.235; TCGA ESCA -0.301; TCGA HNSC -0.268; TCGA LGG -0.177; TCGA LIHC -0.231; TCGA OV -0.323; TCGA PAAD -0.415; TCGA SARC -0.397; TCGA STAD -0.351; TCGA UCEC -0.524 |
hsa-miR-181a-2-3p | SNPH | 9 cancers: BLCA; BRCA; COAD; ESCA; KIRC; LGG; PAAD; THCA; UCEC | mirMAP | TCGA BLCA -0.107; TCGA BRCA -0.223; TCGA COAD -0.232; TCGA ESCA -0.256; TCGA KIRC -0.208; TCGA LGG -0.267; TCGA PAAD -0.636; TCGA THCA -0.234; TCGA UCEC -0.299 |
hsa-miR-181a-2-3p | NFIC | 10 cancers: BLCA; BRCA; COAD; HNSC; LIHC; OV; PRAD; SARC; THCA; STAD | mirMAP | TCGA BLCA -0.308; TCGA BRCA -0.143; TCGA COAD -0.141; TCGA HNSC -0.21; TCGA LIHC -0.123; TCGA OV -0.148; TCGA PRAD -0.128; TCGA SARC -0.172; TCGA THCA -0.091; TCGA STAD -0.135 |
hsa-miR-181a-2-3p | GNG7 | 11 cancers: BLCA; ESCA; HNSC; KIRC; LGG; LIHC; OV; PAAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.336; TCGA ESCA -0.358; TCGA HNSC -0.202; TCGA KIRC -0.181; TCGA LGG -0.263; TCGA LIHC -0.167; TCGA OV -0.37; TCGA PAAD -0.318; TCGA THCA -0.2; TCGA STAD -0.249; TCGA UCEC -0.601 |
hsa-miR-181a-2-3p | LYNX1 | 10 cancers: BLCA; CESC; COAD; HNSC; KIRC; LGG; LIHC; OV; SARC; UCEC | mirMAP | TCGA BLCA -0.362; TCGA CESC -0.579; TCGA COAD -0.293; TCGA HNSC -0.371; TCGA KIRC -0.609; TCGA LGG -0.37; TCGA LIHC -0.19; TCGA OV -0.44; TCGA SARC -0.336; TCGA UCEC -0.328 |
hsa-miR-181a-2-3p | PPARA | 12 cancers: BLCA; CESC; ESCA; HNSC; KIRC; LIHC; OV; PRAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.191; TCGA CESC -0.179; TCGA ESCA -0.2; TCGA HNSC -0.131; TCGA KIRC -0.355; TCGA LIHC -0.285; TCGA OV -0.14; TCGA PRAD -0.082; TCGA SARC -0.097; TCGA THCA -0.068; TCGA STAD -0.286; TCGA UCEC -0.121 |
hsa-miR-181a-2-3p | ABCA5 | 9 cancers: BRCA; CESC; ESCA; HNSC; KIRC; LGG; LIHC; PAAD; STAD | MirTarget | TCGA BRCA -0.12; TCGA CESC -0.111; TCGA ESCA -0.166; TCGA HNSC -0.13; TCGA KIRC -0.192; TCGA LGG -0.469; TCGA LIHC -0.136; TCGA PAAD -0.513; TCGA STAD -0.234 |
hsa-miR-181a-2-3p | ATP1A3 | 11 cancers: BRCA; CESC; COAD; ESCA; HNSC; LUAD; OV; PAAD; THCA; STAD; UCEC | MirTarget | TCGA BRCA -0.503; TCGA CESC -0.482; TCGA COAD -0.685; TCGA ESCA -0.358; TCGA HNSC -0.216; TCGA LUAD -0.238; TCGA OV -0.763; TCGA PAAD -1.073; TCGA THCA -0.713; TCGA STAD -0.269; TCGA UCEC -0.418 |
hsa-miR-181a-2-3p | RMND5B | 9 cancers: BRCA; CESC; ESCA; HNSC; KIRC; LIHC; PAAD; SARC; STAD | MirTarget | TCGA BRCA -0.221; TCGA CESC -0.154; TCGA ESCA -0.124; TCGA HNSC -0.252; TCGA KIRC -0.087; TCGA LIHC -0.071; TCGA PAAD -0.151; TCGA SARC -0.113; TCGA STAD -0.115 |
hsa-miR-181a-2-3p | CA12 | 9 cancers: BRCA; KIRC; LGG; LUAD; LUSC; OV; PRAD; SARC; UCEC | MirTarget | TCGA BRCA -0.869; TCGA KIRC -0.525; TCGA LGG -0.316; TCGA LUAD -0.378; TCGA LUSC -0.716; TCGA OV -0.264; TCGA PRAD -0.166; TCGA SARC -0.231; TCGA UCEC -0.167 |
hsa-miR-181a-2-3p | HEMK1 | 10 cancers: BRCA; ESCA; KIRC; KIRP; LIHC; OV; PAAD; SARC; THCA; STAD | mirMAP | TCGA BRCA -0.305; TCGA ESCA -0.233; TCGA KIRC -0.182; TCGA KIRP -0.155; TCGA LIHC -0.158; TCGA OV -0.091; TCGA PAAD -0.101; TCGA SARC -0.114; TCGA THCA -0.088; TCGA STAD -0.089 |
hsa-miR-181a-2-3p | MTFR1 | 11 cancers: CESC; COAD; KIRC; KIRP; LIHC; LUAD; LUSC; PRAD; SARC; THCA; UCEC | MirTarget | TCGA CESC -0.083; TCGA COAD -0.105; TCGA KIRC -0.202; TCGA KIRP -0.138; TCGA LIHC -0.098; TCGA LUAD -0.196; TCGA LUSC -0.235; TCGA PRAD -0.072; TCGA SARC -0.061; TCGA THCA -0.195; TCGA UCEC -0.061 |
hsa-miR-181a-2-3p | SLC6A17 | 10 cancers: CESC; COAD; KIRC; KIRP; LUAD; PAAD; PRAD; SARC; THCA; UCEC | mirMAP | TCGA CESC -0.27; TCGA COAD -0.544; TCGA KIRC -1.282; TCGA KIRP -0.389; TCGA LUAD -0.35; TCGA PAAD -0.822; TCGA PRAD -0.459; TCGA SARC -0.419; TCGA THCA -0.746; TCGA UCEC -0.324 |
Enriched cancer pathways of putative targets