microRNA information: hsa-miR-200b-5p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-200b-5p | miRbase |
| Accession: | MIMAT0004571 | miRbase |
| Precursor name: | hsa-mir-200b | miRbase |
| Precursor accession: | MI0000342 | miRbase |
| Symbol: | MIR200B | HGNC |
| RefSeq ID: | NR_029639 | GenBank |
| Sequence: | CAUCUUACUGGGCAGCAUUGGA |
Reported expression in cancers: hsa-miR-200b-5p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-200b-5p | bladder cancer | upregulation | "MicroRNA expression signatures of bladder cancer r ......" | 21464941 | RNA-Seq |
| hsa-miR-200b-5p | bladder cancer | upregulation | "Samples were analyzed with a miRNA array containin ......" | 22863868 | Microarray |
| hsa-miR-200b-5p | breast cancer | downregulation | "miR 200b as a prognostic factor in breast cancer t ......" | 24447584 | qPCR; Microarray; in situ hybridization |
| hsa-miR-200b-5p | breast cancer | downregulation | "MicroRNA 200b targets protein kinase Cα and suppr ......" | 24925028 | |
| hsa-miR-200b-5p | breast cancer | downregulation | "In the present study we investigated the roles and ......" | 25639535 | |
| hsa-miR-200b-5p | breast cancer | downregulation | "Expression levels of miR-200a miR-200b miR-200c mi ......" | 26201425 | qPCR |
| hsa-miR-200b-5p | breast cancer | downregulation | "Loss of expression of miR-200 family members has b ......" | 27717206 | |
| hsa-miR-200b-5p | cervical and endocervical cancer | upregulation | "Previous studies have revealed the important role ......" | 27044840 | |
| hsa-miR-200b-5p | colorectal cancer | upregulation | "In this study we revealed a role for miR-200b in c ......" | 24151081 | |
| hsa-miR-200b-5p | colorectal cancer | upregulation | "The processes of EMT and metastasis are highly reg ......" | 25422078 | |
| hsa-miR-200b-5p | endometrial cancer | upregulation | "We evaluated the differential expressions of miRNA ......" | 21035172 | Microarray |
| hsa-miR-200b-5p | endometrial cancer | upregulation | "In this study we investigated miR-200b expression ......" | 23205572 | |
| hsa-miR-200b-5p | endometrial cancer | upregulation | "Using integrated statistical analyses we identifie ......" | 25750291 | |
| hsa-miR-200b-5p | esophageal cancer | downregulation | "To further our understanding of the pathobiology o ......" | 24064224 | qPCR |
| hsa-miR-200b-5p | gastric cancer | downregulation | "miR 200b and miR 200c as prognostic factors and me ......" | 23995857 | qPCR; Microarray |
| hsa-miR-200b-5p | gastric cancer | downregulation | "Although microRNA-200 miR-200 family members are t ......" | 25502084 | |
| hsa-miR-200b-5p | gastric cancer | downregulation | "To find a potentially useful prognostic predictor ......" | 26137064 | |
| hsa-miR-200b-5p | head and neck cancer | downregulation | "The miR-200 family and miR-203 were downregulated ......" | 26882562 | |
| hsa-miR-200b-5p | head and neck cancer | downregulation | "Expression profiles of miR 29c miR 200b and miR 37 ......" | 27440205 | qPCR |
| hsa-miR-200b-5p | kidney renal cell cancer | downregulation | "The set of miRNAs with significantly decreased exp ......" | 23074016 | |
| hsa-miR-200b-5p | liver cancer | downregulation | "We investigated the diagnostic impact of microRNA- ......" | 24895326 | qPCR |
| hsa-miR-200b-5p | liver cancer | downregulation | "MiR-200 family is an important regulator of epithe ......" | 25909223 | |
| hsa-miR-200b-5p | liver cancer | downregulation | "Significant downregulation of microRNA-200b was ob ......" | 26919246 | |
| hsa-miR-200b-5p | lung cancer | upregulation | "However the molecular mechanisms involved in the s ......" | 24830600 | |
| hsa-miR-200b-5p | lung cancer | upregulation | "Among these 28 miRNAs were up-regulated in both th ......" | 27189341 | qPCR |
| hsa-miR-200b-5p | melanoma | deregulation | "A functional role of microRNAs miRNAs or miRs in n ......" | 20957176 | |
| hsa-miR-200b-5p | ovarian cancer | upregulation | "The prognostic value of the miR 200 family in ovar ......" | 26910180 | |
| hsa-miR-200b-5p | pancreatic cancer | upregulation | "Profiling of 95 microRNAs in pancreatic cancer cel ......" | 19030927 | qPCR |
| hsa-miR-200b-5p | prostate cancer | downregulation | "miR 200b suppresses cell proliferation migration a ......" | 24317363 | |
| hsa-miR-200b-5p | prostate cancer | deregulation | "Comparative microRNA profiling of prostate carcino ......" | 24337069 | RNA-Seq; qPCR |
| hsa-miR-200b-5p | prostate cancer | upregulation | "miR 200b inhibits prostate cancer EMT growth and m ......" | 24391862 | |
| hsa-miR-200b-5p | sarcoma | deregulation | "The most significantly downregulated miRNAs were m ......" | 24027049 | |
| hsa-miR-200b-5p | thyroid cancer | upregulation | "The miR-200 family was recently identified as a su ......" | 22797360 | |
| hsa-miR-200b-5p | thyroid cancer | downregulation | "MiR 200 Regulates Epithelial Mesenchymal Transitio ......" | 25542369 |
Reported cancer pathway affected by hsa-miR-200b-5p
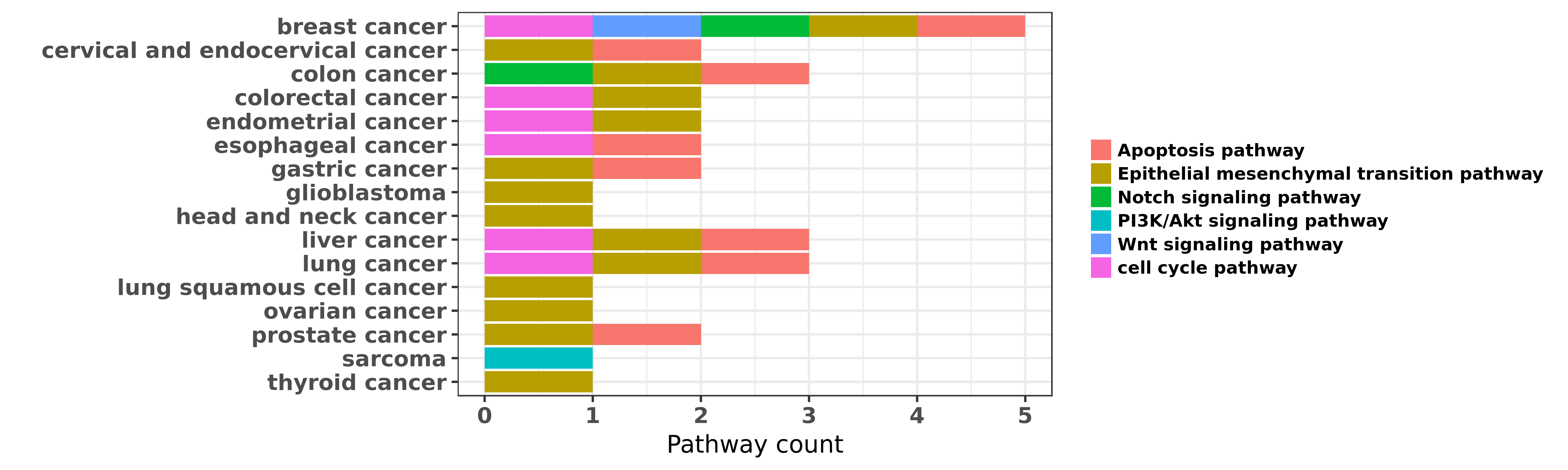
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-200b-5p | breast cancer | cell cycle pathway; Apoptosis pathway | "The genes encoding microRNAs of the human miR-200 ......" | 20514023 | Luciferase |
| hsa-miR-200b-5p | breast cancer | Epithelial mesenchymal transition pathway | "Phosphoglucose isomerase/autocrine motility factor ......" | 21389093 | |
| hsa-miR-200b-5p | breast cancer | Epithelial mesenchymal transition pathway | "Global miRNA expression profiling was performed on ......" | 22294488 | |
| hsa-miR-200b-5p | breast cancer | Epithelial mesenchymal transition pathway | "Epigenetic modulation of the miR 200 family is ass ......" | 23525011 | |
| hsa-miR-200b-5p | breast cancer | Epithelial mesenchymal transition pathway | "MiR 200 can repress breast cancer metastasis throu ......" | 24037528 | |
| hsa-miR-200b-5p | breast cancer | Epithelial mesenchymal transition pathway | "miR 200b as a prognostic factor in breast cancer t ......" | 24447584 | Western blot; Luciferase; MTT assay |
| hsa-miR-200b-5p | breast cancer | Epithelial mesenchymal transition pathway | "Some miRNAs especially the miR-200 family miR-9 an ......" | 25086633 | |
| hsa-miR-200b-5p | breast cancer | Apoptosis pathway; cell cycle pathway | "MiR 200b expression in breast cancer: a prognostic ......" | 25639535 | Western blot; Luciferase |
| hsa-miR-200b-5p | breast cancer | Wnt signaling pathway; Apoptosis pathway; Notch signaling pathway | "The miRNAs and their clusters such as the miR-200 ......" | 26712794 | |
| hsa-miR-200b-5p | cervical and endocervical cancer | Epithelial mesenchymal transition pathway | "MicroRNA 200b suppresses cell invasion and metasta ......" | 26935156 | Cell migration assay; Western blot |
| hsa-miR-200b-5p | cervical and endocervical cancer | Epithelial mesenchymal transition pathway | "MicroRNA 200b inhibits epithelial mesenchymal tran ......" | 26935796 | Transwell assay; Luciferase |
| hsa-miR-200b-5p | cervical and endocervical cancer | Apoptosis pathway | "MiR 200b promotes the cell proliferation and metas ......" | 27044840 | Western blot; Luciferase |
| hsa-miR-200b-5p | colon cancer | Epithelial mesenchymal transition pathway | "MicroRNA 200 miR 200 cluster regulation by achaete ......" | 25371200 | |
| hsa-miR-200b-5p | colon cancer | Apoptosis pathway; Notch signaling pathway | "Niclosamide inhibits colon cancer progression thro ......" | 27460529 | Flow cytometry; Cell migration assay |
| hsa-miR-200b-5p | colorectal cancer | cell cycle pathway | "MicroRNA 200b stimulates tumour growth in TGFBR2 n ......" | 24151081 | |
| hsa-miR-200b-5p | colorectal cancer | Epithelial mesenchymal transition pathway | "Epithelial-mesenchymal transition EMT and changes ......" | 25757925 | |
| hsa-miR-200b-5p | colorectal cancer | Epithelial mesenchymal transition pathway | "The miR-200 family presented as the most powerful ......" | 27157610 | |
| hsa-miR-200b-5p | endometrial cancer | Epithelial mesenchymal transition pathway | "Tamoxifen represses miR 200 microRNAs and promotes ......" | 23295740 | |
| hsa-miR-200b-5p | endometrial cancer | cell cycle pathway | "The expression of miRNAs and related genes were de ......" | 23680357 | Western blot; Flow cytometry; Colony formation |
| hsa-miR-200b-5p | esophageal cancer | cell cycle pathway; Apoptosis pathway | "miR 200b induces cell cycle arrest and represses c ......" | 27496804 | |
| hsa-miR-200b-5p | gastric cancer | Epithelial mesenchymal transition pathway | "MicroRNA 200b regulates cell proliferation invasio ......" | 22311119 | |
| hsa-miR-200b-5p | gastric cancer | Apoptosis pathway | "Diallyl disulfide suppresses proliferation and ind ......" | 23851184 | |
| hsa-miR-200b-5p | gastric cancer | Epithelial mesenchymal transition pathway | "Epigenetic modulation and repression of miR 200b b ......" | 25411357 | |
| hsa-miR-200b-5p | glioblastoma | Epithelial mesenchymal transition pathway | "NPV LDE 225 Erismodegib inhibits epithelial mesenc ......" | 23482671 | Luciferase; Western blot |
| hsa-miR-200b-5p | head and neck cancer | Epithelial mesenchymal transition pathway | "Epithelial-mesenchymal-transition EMT is a critica ......" | 24424572 | |
| hsa-miR-200b-5p | liver cancer | cell cycle pathway | "Expression of the microRNA-200 miR-200 family has ......" | 23760980 | Flow cytometry; MTT assay |
| hsa-miR-200b-5p | liver cancer | Epithelial mesenchymal transition pathway | "We investigated the diagnostic impact of microRNA- ......" | 24895326 | |
| hsa-miR-200b-5p | liver cancer | Epithelial mesenchymal transition pathway | "MiR-200 family is an important regulator of epithe ......" | 25909223 | |
| hsa-miR-200b-5p | liver cancer | cell cycle pathway; Apoptosis pathway | "Methylation associated silencing of miR 200b facil ......" | 26919246 | Luciferase; Colony formation |
| hsa-miR-200b-5p | lung cancer | cell cycle pathway; Apoptosis pathway | "MicroRNA 200b reverses chemoresistance of docetaxe ......" | 22139708 | Luciferase |
| hsa-miR-200b-5p | lung cancer | Epithelial mesenchymal transition pathway | "Nobiletin inhibited hypoxia induced epithelial mes ......" | 26012256 | |
| hsa-miR-200b-5p | lung cancer | Epithelial mesenchymal transition pathway | "BMP4 depletion by miR 200 inhibits tumorigenesis a ......" | 26395571 | Western blot; Luciferase |
| hsa-miR-200b-5p | lung squamous cell cancer | Epithelial mesenchymal transition pathway | "The microRNA‑200 miR-200 family is a powerful re ......" | 23708087 | Western blot |
| hsa-miR-200b-5p | lung squamous cell cancer | Epithelial mesenchymal transition pathway | "The difference in miRNA expression profiles betwee ......" | 26783187 | |
| hsa-miR-200b-5p | ovarian cancer | Epithelial mesenchymal transition pathway | "Differences in miRNA expression between HGSC CCC a ......" | 24512620 | |
| hsa-miR-200b-5p | ovarian cancer | Epithelial mesenchymal transition pathway | "Epithelial mesenchymal transition associated miRNA ......" | 24952258 | |
| hsa-miR-200b-5p | ovarian cancer | Epithelial mesenchymal transition pathway | "The miR 200 family differentially regulates sensit ......" | 26025631 | |
| hsa-miR-200b-5p | prostate cancer | Epithelial mesenchymal transition pathway | "miR 200 regulates PDGF D mediated epithelial mesen ......" | 19544444 | |
| hsa-miR-200b-5p | prostate cancer | Apoptosis pathway | "Nevertheless decreased expressions of tumor suppre ......" | 26843836 | |
| hsa-miR-200b-5p | sarcoma | PI3K/Akt signaling pathway | "CCL5 promotes VEGF dependent angiogenesis by down ......" | 25301739 | Western blot |
| hsa-miR-200b-5p | thyroid cancer | Epithelial mesenchymal transition pathway | "We identified two significantly decreased microRNA ......" | 20498632 | |
| hsa-miR-200b-5p | thyroid cancer | Epithelial mesenchymal transition pathway | "The miR 200 family regulates the epithelial mesenc ......" | 22797360 | Western blot |
| hsa-miR-200b-5p | thyroid cancer | Epithelial mesenchymal transition pathway | "MiR 200 Regulates Epithelial Mesenchymal Transitio ......" | 25542369 | Western blot |
Reported cancer prognosis affected by hsa-miR-200b-5p

| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-200b-5p | B cell lymphoma | worse prognosis | "Inhibition of ZEB1 by miR 200 characterizes Helico ......" | 24390222 | |
| hsa-miR-200b-5p | bladder cancer | drug resistance; cell migration | "miR 200 expression regulates epithelial to mesench ......" | 19671845 | Western blot; Luciferase |
| hsa-miR-200b-5p | bladder cancer | staging; progression | "Coordinated epigenetic repression of the miR 200 f ......" | 20473948 | |
| hsa-miR-200b-5p | bladder cancer | staging | "Samples were analyzed with a miRNA array containin ......" | 22863868 | |
| hsa-miR-200b-5p | bladder cancer | staging; poor survival; recurrence | "Expression levels of miRNAs were assessed by quant ......" | 23169479 | |
| hsa-miR-200b-5p | bladder cancer | malignant trasformation | "Analyses in human urothelial cells identify methyl ......" | 23867826 | |
| hsa-miR-200b-5p | breast cancer | drug resistance | "The drug resistance of MCF-7/ADR cells was evaluat ......" | 18971180 | Flow cytometry; MTT assay |
| hsa-miR-200b-5p | breast cancer | metastasis | "miR 200 enhances mouse breast cancer cell coloniza ......" | 19787069 | |
| hsa-miR-200b-5p | breast cancer | progression | "Recent evidence demonstrates that up-regulation of ......" | 19839049 | |
| hsa-miR-200b-5p | breast cancer | progression | "The genes encoding microRNAs of the human miR-200 ......" | 20514023 | Luciferase |
| hsa-miR-200b-5p | breast cancer | motility; metastasis | "Phosphoglucose isomerase/autocrine motility factor ......" | 21389093 | |
| hsa-miR-200b-5p | breast cancer | metastasis | "Decreased expression of miRNAs of the miR-200 fami ......" | 21682933 | |
| hsa-miR-200b-5p | breast cancer | staging; metastasis | "Mapping the regulatory sequences controlling 93 br ......" | 22231446 | |
| hsa-miR-200b-5p | breast cancer | metastasis; staging; progression | "Global miRNA expression profiling was performed on ......" | 22294488 | |
| hsa-miR-200b-5p | breast cancer | differentiation | "In order to identify which miRNAs are involved in ......" | 22562546 | |
| hsa-miR-200b-5p | breast cancer | progression; poor survival | "Circulating miRNAs from plasma of CTC-positive and ......" | 22952344 | |
| hsa-miR-200b-5p | breast cancer | differentiation | "As the family of miR-200 microRNAs has been shown ......" | 23112837 | |
| hsa-miR-200b-5p | breast cancer | progression | "Epigenetic modulation of the miR 200 family is ass ......" | 23525011 | |
| hsa-miR-200b-5p | breast cancer | drug resistance; progression; cell migration | "Reduced expression of miR 200 family members contr ......" | 23626803 | |
| hsa-miR-200b-5p | breast cancer | metastasis; staging; poor survival | "MiR 200 can repress breast cancer metastasis throu ......" | 24037528 | |
| hsa-miR-200b-5p | breast cancer | drug resistance | "miR 200b as a prognostic factor in breast cancer t ......" | 24447584 | Western blot; Luciferase; MTT assay |
| hsa-miR-200b-5p | breast cancer | metastasis | "The expression of miRNAs in patients with primary ......" | 24846313 | |
| hsa-miR-200b-5p | breast cancer | metastasis; cell migration | "MicroRNA 200b targets protein kinase Cα and suppr ......" | 24925028 | |
| hsa-miR-200b-5p | breast cancer | metastasis; progression | "Members of three miRNA families i.e miR-17 miR-200 ......" | 25001613 | |
| hsa-miR-200b-5p | breast cancer | worse prognosis | "Some miRNAs especially the miR-200 family miR-9 an ......" | 25086633 | |
| hsa-miR-200b-5p | breast cancer | staging | "MiR 200b expression in breast cancer: a prognostic ......" | 25639535 | Western blot; Luciferase |
| hsa-miR-200b-5p | breast cancer | drug resistance | "Therefore we investigated the role of known EMT re ......" | 25746005 | |
| hsa-miR-200b-5p | breast cancer | progression | "ADAM12 L is a direct target of the miR 29 and miR ......" | 25886595 | Western blot; Luciferase |
| hsa-miR-200b-5p | breast cancer | metastasis | "The upregulation of fibronectin and lysyl oxidase ......" | 26068592 | |
| hsa-miR-200b-5p | breast cancer | metastasis | "Expression levels of miR-200a miR-200b miR-200c mi ......" | 26201425 | |
| hsa-miR-200b-5p | breast cancer | metastasis | "These included miR-141 miR-144 miR-193b miR-200a m ......" | 26785733 | |
| hsa-miR-200b-5p | breast cancer | cell migration; metastasis | "MicroRNA 200b Impacts Breast Cancer Cell Migration ......" | 27276064 | Western blot; Transwell assay; Wound Healing Assay |
| hsa-miR-200b-5p | breast cancer | differentiation | "miR 9 and miR 200 Regulate PDGFRβ Mediated Endoth ......" | 27402080 | |
| hsa-miR-200b-5p | breast cancer | drug resistance | "Loss of expression of miR-200 family members has b ......" | 27717206 | |
| hsa-miR-200b-5p | breast cancer | drug resistance | "Specifically we will discuss key miRNAs involved i ......" | 27721895 | |
| hsa-miR-200b-5p | breast cancer | worse prognosis; drug resistance | "The aim of this study was to investigate the role ......" | 27746365 | |
| hsa-miR-200b-5p | cervical and endocervical cancer | metastasis | "MicroRNA 200b suppresses cell invasion and metasta ......" | 26935156 | Cell migration assay; Western blot |
| hsa-miR-200b-5p | cervical and endocervical cancer | metastasis; progression; cell migration | "MiR 200b promotes the cell proliferation and metas ......" | 27044840 | Western blot; Luciferase |
| hsa-miR-200b-5p | cervical and endocervical cancer | progression | "Long noncoding RNA PVT1 promotes cervical cancer p ......" | 27272214 | |
| hsa-miR-200b-5p | colon cancer | progression | "Niclosamide inhibits colon cancer progression thro ......" | 27460529 | Flow cytometry; Cell migration assay |
| hsa-miR-200b-5p | colorectal cancer | metastasis | "Applying real-time PCR we detected the expression ......" | 21827717 | |
| hsa-miR-200b-5p | colorectal cancer | metastasis | "Although the microRNA-200 miR-200 family is a cruc ......" | 22735571 | |
| hsa-miR-200b-5p | colorectal cancer | metastasis; staging | "Here we used in situ hybridization and immunohisto ......" | 23441132 | |
| hsa-miR-200b-5p | colorectal cancer | staging; metastasis | "In the first phase we selected candidate miRNAs as ......" | 23982750 | |
| hsa-miR-200b-5p | colorectal cancer | progression | "MicroRNA 200b stimulates tumour growth in TGFBR2 n ......" | 24151081 | |
| hsa-miR-200b-5p | colorectal cancer | poor survival; staging | "Role of miR 200 family members in survival of colo ......" | 24510588 | |
| hsa-miR-200b-5p | colorectal cancer | metastasis | "The processes of EMT and metastasis are highly reg ......" | 25422078 | |
| hsa-miR-200b-5p | colorectal cancer | drug resistance | "Epithelial-mesenchymal transition EMT and changes ......" | 25757925 | |
| hsa-miR-200b-5p | colorectal cancer | metastasis | "MiR 200 suppresses metastases of colorectal cancer ......" | 26242262 | Luciferase |
| hsa-miR-200b-5p | colorectal cancer | poor survival; worse prognosis | "Survival analyses showed that plasma miR-96 and mi ......" | 26863633 | |
| hsa-miR-200b-5p | colorectal cancer | worse prognosis; poor survival | "The miR-200 family presented as the most powerful ......" | 27157610 | |
| hsa-miR-200b-5p | colorectal cancer | worse prognosis | "Plasma miR 122 and miR 200 family are prognostic m ......" | 27632639 | |
| hsa-miR-200b-5p | endometrial cancer | tumorigenesis | "We compared the expression profiles of 723 human m ......" | 21897839 | |
| hsa-miR-200b-5p | endometrial cancer | metastasis | "MicroRNA 200b is overexpressed in endometrial aden ......" | 23205572 | Western blot; Luciferase |
| hsa-miR-200b-5p | endometrial cancer | motility; drug resistance | "Tamoxifen represses miR 200 microRNAs and promotes ......" | 23295740 | |
| hsa-miR-200b-5p | endometrial cancer | progression; metastasis; tumorigenesis | "The expression of miRNAs and related genes were de ......" | 23680357 | Western blot; Flow cytometry; Colony formation |
| hsa-miR-200b-5p | endometrial cancer | metastasis | "The abnormal expression of the long noncoding RNA ......" | 27693631 | |
| hsa-miR-200b-5p | esophageal cancer | staging; metastasis; poor survival | "miR 200b suppresses invasiveness and modulates the ......" | 24064224 | |
| hsa-miR-200b-5p | esophageal cancer | progression; poor survival | "miR 200b induces cell cycle arrest and represses c ......" | 27496804 | |
| hsa-miR-200b-5p | gastric cancer | tumorigenesis | "The effect of microRNA abnormalities in carcinogen ......" | 20484038 | |
| hsa-miR-200b-5p | gastric cancer | differentiation; metastasis | "MicroRNA 200b regulates cell proliferation invasio ......" | 22311119 | |
| hsa-miR-200b-5p | gastric cancer | progression; worse prognosis; staging; metastasis; poor survival | "miR 200b and miR 200c as prognostic factors and me ......" | 23995857 | Luciferase; Cell proliferation assay; Cell migration assay; Cell Proliferation Assay |
| hsa-miR-200b-5p | gastric cancer | worse prognosis; poor survival; cell migration | "Integrated microRNA network analyses identify a po ......" | 24352645 | |
| hsa-miR-200b-5p | gastric cancer | metastasis; worse prognosis | "Epigenetic modulation and repression of miR 200b b ......" | 25411357 | |
| hsa-miR-200b-5p | gastric cancer | tumorigenesis; progression | "Although microRNA-200 miR-200 family members are t ......" | 25502084 | Wound Healing Assay |
| hsa-miR-200b-5p | gastric cancer | staging; metastasis | "In the first step of this study preliminary experi ......" | 26233325 | |
| hsa-miR-200b-5p | glioblastoma | metastasis | "We determined CSF levels of several cancer-associa ......" | 22492962 | |
| hsa-miR-200b-5p | head and neck cancer | metastasis; tumorigenesis | "Epithelial-mesenchymal-transition EMT is a critica ......" | 24424572 | |
| hsa-miR-200b-5p | head and neck cancer | progression; worse prognosis | "Expression profiles of miR 29c miR 200b and miR 37 ......" | 27440205 | |
| hsa-miR-200b-5p | kidney renal cell cancer | malignant trasformation | "We performed genome-wide expression profiling of m ......" | 19228262 | |
| hsa-miR-200b-5p | kidney renal cell cancer | progression; tumorigenesis | "MicroRNA-429 miR-429 a short noncoding RNA belongi ......" | 25953723 | |
| hsa-miR-200b-5p | kidney renal cell cancer | staging | "Comparisons of RCC and BC expression signatures re ......" | 27357429 | |
| hsa-miR-200b-5p | liver cancer | malignant trasformation | "Our study identified and validated miR-224 overexp ......" | 18433021 | |
| hsa-miR-200b-5p | liver cancer | metastasis | "Epigenetic activation of the MiR 200 family contri ......" | 23222811 | |
| hsa-miR-200b-5p | liver cancer | metastasis; cell migration; differentiation | "miR 200b restoration and DNA methyltransferase inh ......" | 23552699 | |
| hsa-miR-200b-5p | liver cancer | drug resistance | "Expression of the microRNA-200 miR-200 family has ......" | 23760980 | Flow cytometry; MTT assay |
| hsa-miR-200b-5p | liver cancer | metastasis; progression | "The aim of this study was to assess the role of th ......" | 24135722 | |
| hsa-miR-200b-5p | liver cancer | cell migration | "Transfection of Huh7 cell with an activated mutant ......" | 25065598 | |
| hsa-miR-200b-5p | liver cancer | tumorigenesis; cell migration; metastasis | "MiR-200 family is an important regulator of epithe ......" | 25909223 | |
| hsa-miR-200b-5p | liver cancer | progression | "Methylation associated silencing of miR 200b facil ......" | 26919246 | Luciferase; Colony formation |
| hsa-miR-200b-5p | lung cancer | metastasis; worse prognosis; tumorigenesis | "miR 200 Inhibits lung adenocarcinoma cell invasion ......" | 21115742 | |
| hsa-miR-200b-5p | lung cancer | metastasis | "The Notch ligand Jagged2 promotes lung adenocarcin ......" | 21403400 | |
| hsa-miR-200b-5p | lung cancer | drug resistance; worse prognosis | "MicroRNA 200b reverses chemoresistance of docetaxe ......" | 22139708 | Luciferase |
| hsa-miR-200b-5p | lung cancer | metastasis | "Loss of miR-200s has been shown to enhance cancer ......" | 24791940 | |
| hsa-miR-200b-5p | lung cancer | drug resistance | "HDAC 1/4 mediated silencing of microRNA 200b promo ......" | 24830600 | |
| hsa-miR-200b-5p | lung cancer | drug resistance | "Recently we have identified microRNA miR-200b as a ......" | 25279705 | |
| hsa-miR-200b-5p | lung cancer | metastasis; cell migration | "The miR 200 family and the miR 183~96~182 cluster ......" | 25798833 | |
| hsa-miR-200b-5p | lung cancer | staging; tumorigenesis | "MicroRNA-200 miR-200 has emerged as a regulator of ......" | 26314828 | |
| hsa-miR-200b-5p | lung cancer | metastasis; tumorigenesis | "BMP4 depletion by miR 200 inhibits tumorigenesis a ......" | 26395571 | Western blot; Luciferase |
| hsa-miR-200b-5p | lung cancer | drug resistance | "MiR 200b regulates autophagy associated with chemo ......" | 26416454 | |
| hsa-miR-200b-5p | lung cancer | progression | "Interestingly mir-200 family mir-200a mir-200b and ......" | 27695346 | |
| hsa-miR-200b-5p | lung squamous cell cancer | metastasis; immune resistance | "The microRNA‑200 miR-200 family is a powerful re ......" | 23708087 | Western blot |
| hsa-miR-200b-5p | lung squamous cell cancer | drug resistance | "Zinc finger E box binding homeobox 2 ZEB2 regulate ......" | 24769353 | Luciferase |
| hsa-miR-200b-5p | lung squamous cell cancer | worse prognosis; staging | "The miRNA family miR-200 has been associated with ......" | 25003366 | |
| hsa-miR-200b-5p | lung squamous cell cancer | drug resistance | "The difference in miRNA expression profiles betwee ......" | 26783187 | |
| hsa-miR-200b-5p | lung squamous cell cancer | metastasis | "miR 200b inhibits migration and invasion in non sm ......" | 27356635 | Luciferase |
| hsa-miR-200b-5p | lung squamous cell cancer | progression; tumorigenesis | "The microRNA miR-200 family has been demonstrated ......" | 27602157 | |
| hsa-miR-200b-5p | lymphoma | progression | "The profiles of miRNAs in conjunctival MALT lympho ......" | 22183793 | Luciferase |
| hsa-miR-200b-5p | melanoma | metastasis; progression; cell migration | "A functional role of microRNAs miRNAs or miRs in n ......" | 20957176 | |
| hsa-miR-200b-5p | melanoma | progression | "MiR-200 in situ hybridization and E-cadherin immun ......" | 22956368 | |
| hsa-miR-200b-5p | melanoma | metastasis | "From a discovery analysis using 40 thick primary m ......" | 26743475 | |
| hsa-miR-200b-5p | ovarian cancer | worse prognosis | "The microRNA expression profiles were examined usi ......" | 18451233 | |
| hsa-miR-200b-5p | ovarian cancer | staging | "Levels of 8 microRNAs miR-21 miR-141 miR-200a miR- ......" | 18589210 | |
| hsa-miR-200b-5p | ovarian cancer | staging; poor survival; recurrence; cell migration | "A miR 200 microRNA cluster as prognostic marker in ......" | 19501389 | |
| hsa-miR-200b-5p | ovarian cancer | tumorigenesis | "Regulation of miR 200 family microRNAs and ZEB tra ......" | 19854497 | Luciferase |
| hsa-miR-200b-5p | ovarian cancer | worse prognosis | "We highlight the role of the let-7 and miR-200 fam ......" | 20083225 | |
| hsa-miR-200b-5p | ovarian cancer | progression; poor survival; drug resistance | "The miR 200 family controls beta tubulin III expre ......" | 21051560 | |
| hsa-miR-200b-5p | ovarian cancer | poor survival | "The Cox proportional hazards model and the log-ran ......" | 21345725 | |
| hsa-miR-200b-5p | ovarian cancer | worse prognosis; staging | "Two families of miRNAs miR-200 and let-7 are frequ ......" | 23237306 | |
| hsa-miR-200b-5p | ovarian cancer | drug resistance | "Sequence variation among members of the miR 200 mi ......" | 24447705 | |
| hsa-miR-200b-5p | ovarian cancer | progression; poor survival | "Differences in miRNA expression between HGSC CCC a ......" | 24512620 | |
| hsa-miR-200b-5p | ovarian cancer | metastasis; recurrence | "We recently determined that the ectopic over-expre ......" | 24802724 | |
| hsa-miR-200b-5p | ovarian cancer | metastasis; drug resistance | "For example deficiencies of enzymes including Dice ......" | 24822185 | |
| hsa-miR-200b-5p | ovarian cancer | progression; metastasis | "Epithelial mesenchymal transition associated miRNA ......" | 24952258 | |
| hsa-miR-200b-5p | ovarian cancer | poor survival; staging; progression | "Clinicopathological and prognostic implications of ......" | 24966949 | |
| hsa-miR-200b-5p | ovarian cancer | progression; worse prognosis; poor survival | "Expression of serum miR 200a miR 200b and miR 200c ......" | 26063644 | |
| hsa-miR-200b-5p | ovarian cancer | progression; poor survival; malignant trasformation | "Plasma miR 200b in ovarian carcinoma patients: dis ......" | 26416421 | |
| hsa-miR-200b-5p | ovarian cancer | progression; poor survival | "The prognostic value of the miR 200 family in ovar ......" | 26910180 | |
| hsa-miR-200b-5p | ovarian cancer | malignant trasformation; staging; metastasis; poor survival; progression | "Diagnostic and prognostic relevance of circulating ......" | 26943577 | |
| hsa-miR-200b-5p | ovarian cancer | metastasis | "The miR-200 family especially miR-200c has been sh ......" | 27601996 | |
| hsa-miR-200b-5p | ovarian cancer | poor survival | "Furthermore some of the identified DNA-methylated ......" | 27746113 | |
| hsa-miR-200b-5p | ovarian cancer | malignant trasformation | "Circulating Cell Free miR 373 miR 200a miR 200b an ......" | 27753009 | |
| hsa-miR-200b-5p | pancreatic cancer | metastasis | "Additionally the miR-200 family regulates several ......" | 24040120 | Luciferase |
| hsa-miR-200b-5p | prostate cancer | cell migration | "miR 200 regulates PDGF D mediated epithelial mesen ......" | 19544444 | |
| hsa-miR-200b-5p | prostate cancer | metastasis | "Regulation of epithelial plasticity by miR 424 and ......" | 24193225 | |
| hsa-miR-200b-5p | prostate cancer | staging | "miR 200b suppresses cell proliferation migration a ......" | 24317363 | |
| hsa-miR-200b-5p | prostate cancer | staging; malignant trasformation | "Comparative microRNA profiling of prostate carcino ......" | 24337069 | |
| hsa-miR-200b-5p | prostate cancer | metastasis; progression | "miR 200b inhibits prostate cancer EMT growth and m ......" | 24391862 | |
| hsa-miR-200b-5p | prostate cancer | poor survival; metastasis | "Non-responders to docetaxel and patients with shor ......" | 24714754 | |
| hsa-miR-200b-5p | prostate cancer | poor survival; recurrence | "The expression of 10 genes and 18 miRNAs were asse ......" | 25409297 | |
| hsa-miR-200b-5p | prostate cancer | drug resistance | "Nevertheless decreased expressions of tumor suppre ......" | 26843836 | |
| hsa-miR-200b-5p | sarcoma | tumorigenesis | "MiR-141 which belong to miR-200 family take a part ......" | 24307282 | |
| hsa-miR-200b-5p | sarcoma | progression; staging; metastasis | "MicroRNA 200b acts as a tumor suppressor in osteos ......" | 27307751 | |
| hsa-miR-200b-5p | thyroid cancer | cell migration | "However specific miRNAs are downregulated in ATC s ......" | 25202329 |
Reported gene related to hsa-miR-200b-5p

| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-miR-200b-5p | B cell lymphoma | ZEB1 | "Inhibition of ZEB1 by miR 200 characterizes Helico ......" | 24390222 |
| hsa-miR-200b-5p | bladder cancer | ZEB1 | "ZEB1 is also known to repress miR-200c-141 transcr ......" | 20473948 |
| hsa-miR-200b-5p | breast cancer | ZEB1 | "Conversely ectopic expression of miR-200 inhibited ......" | 27402080 |
| hsa-miR-200b-5p | breast cancer | ZEB1 | "Here we show that 53BP1 negatively regulated EMT b ......" | 26011542 |
| hsa-miR-200b-5p | breast cancer | ZEB1 | "Concomitant with miR-200 decrease there was an inc ......" | 23626803 |
| hsa-miR-200b-5p | breast cancer | ZEB1 | "MiR 200 can repress breast cancer metastasis throu ......" | 24037528 |
| hsa-miR-200b-5p | colorectal cancer | ZEB1 | "MiR 200 suppresses metastases of colorectal cancer ......" | 26242262 |
| hsa-miR-200b-5p | endometrial cancer | ZEB1 | "This finding was accompanied by a sharp downregula ......" | 23743934 |
| hsa-miR-200b-5p | gastric cancer | ZEB1 | "Functional experiments demonstrated that miR-200b ......" | 24352645 |
| hsa-miR-200b-5p | head and neck cancer | ZEB1 | "Furthermore our data suggest that the promoter hyp ......" | 24424572 |
| hsa-miR-200b-5p | lung squamous cell cancer | ZEB1 | "Nintedanib was able to reverse TGF-β1-induced EMT ......" | 26783187 |
| hsa-miR-200b-5p | ovarian cancer | ZEB1 | "We used qRT-PCR to examine expression of the miR-2 ......" | 19854497 |
| hsa-miR-200b-5p | ovarian cancer | ZEB1 | "GRHL2 miR 200 ZEB1 maintains the epithelial status ......" | 26887977 |
| hsa-miR-200b-5p | prostate cancer | ZEB1 | "Recent studies have shown that the miR-200 family ......" | 19544444 |
| hsa-miR-200b-5p | sarcoma | ZEB1 | "This circuit comprises the microRNA 200 miR-200 fa ......" | 27402864 |
| hsa-miR-200b-5p | sarcoma | ZEB1 | "Moreover a series of loss-of-function and gain-of- ......" | 24815002 |
| hsa-miR-200b-5p | sarcoma | ZEB1 | "MicroRNA 200b acts as a tumor suppressor in osteos ......" | 27307751 |
| hsa-miR-200b-5p | breast cancer | CDH1 | "β1 integrin inhibition elicits a prometastatic sw ......" | 24518294 |
| hsa-miR-200b-5p | breast cancer | CDH1 | "These findings are surprising since the miR-200 fa ......" | 19787069 |
| hsa-miR-200b-5p | breast cancer | CDH1 | "Here we demonstrate that the tumor suppressive miR ......" | 26062653 |
| hsa-miR-200b-5p | esophageal cancer | CDH1 | "Loss of miR 200b promotes invasion via activating ......" | 26334393 |
| hsa-miR-200b-5p | gastric cancer | CDH1 | "Functional experiments demonstrated that miR-200b ......" | 24352645 |
| hsa-miR-200b-5p | gastric cancer | CDH1 | "E-cadherin expression was partially restored by tr ......" | 20484038 |
| hsa-miR-200b-5p | gastric cancer | CDH1 | "There was a strong correlation between the levels ......" | 22311119 |
| hsa-miR-200b-5p | liver cancer | CDH1 | "Spearman's Rho analysis revealed a significant neg ......" | 24895326 |
| hsa-miR-200b-5p | liver cancer | CDH1 | "We found that overexpression of miR-200 family mem ......" | 22868917 |
| hsa-miR-200b-5p | liver cancer | CDH1 | "In mesenchymal cells forced expression of miR-200b ......" | 23552699 |
| hsa-miR-200b-5p | melanoma | CDH1 | "MiR-200 in situ hybridization and E-cadherin immun ......" | 22956368 |
| hsa-miR-200b-5p | ovarian cancer | CDH1 | "Our findings showed that miR-101 represents a redu ......" | 24677166 |
| hsa-miR-200b-5p | pancreatic cancer | CDH1 | "In a panel of 23 pancreatic cell lines we observed ......" | 20551052 |
| hsa-miR-200b-5p | thyroid cancer | CDH1 | "Restoration of miR-200 expression by pre-miR-200a/ ......" | 25542369 |
| hsa-miR-200b-5p | breast cancer | ZEB2 | "This switch involved activation of the transformin ......" | 24518294 |
| hsa-miR-200b-5p | breast cancer | ZEB2 | "These findings are surprising since the miR-200 fa ......" | 19787069 |
| hsa-miR-200b-5p | gastric cancer | ZEB2 | "MicroRNA 200b regulates cell proliferation invasio ......" | 22311119 |
| hsa-miR-200b-5p | head and neck cancer | ZEB2 | "Furthermore our data suggest that the promoter hyp ......" | 24424572 |
| hsa-miR-200b-5p | lung squamous cell cancer | ZEB2 | "Zinc finger E box binding homeobox 2 ZEB2 regulate ......" | 24769353 |
| hsa-miR-200b-5p | ovarian cancer | ZEB2 | "We used qRT-PCR to examine expression of the miR-2 ......" | 19854497 |
| hsa-miR-200b-5p | prostate cancer | ZEB2 | "Recent studies have shown that the miR-200 family ......" | 19544444 |
| hsa-miR-200b-5p | sarcoma | ZEB2 | "Moreover a series of loss-of-function and gain-of- ......" | 24815002 |
| hsa-miR-200b-5p | esophageal cancer | VIM | "Here we show that miR-200b represses ESCC cell inv ......" | 26334393 |
| hsa-miR-200b-5p | liver cancer | VIM | "Spearman's Rho analysis revealed a significant neg ......" | 24895326 |
| hsa-miR-200b-5p | prostate cancer | VIM | "In contrast mesenchymal markers fibronectin and vi ......" | 24391862 |
| hsa-miR-200b-5p | thyroid cancer | VIM | "Restoration of miR-200 expression by pre-miR-200a/ ......" | 25542369 |
| hsa-miR-200b-5p | breast cancer | FERMT2 | "Here we report that Kindlin 2 markedly downregulat ......" | 23483548 |
| hsa-miR-200b-5p | esophageal cancer | FERMT2 | "miR 200b suppresses invasiveness and modulates the ......" | 24064224 |
| hsa-miR-200b-5p | esophageal cancer | FERMT2 | "We revealed that miR-200b suppresses the integrin ......" | 26334393 |
| hsa-miR-200b-5p | endometrial cancer | VEGFA | "In contrast a positive correlation was observed be ......" | 22904162 |
| hsa-miR-200b-5p | kidney renal cell cancer | VEGFA | "We also found strong anti-correlation between VEGF ......" | 20420713 |
| hsa-miR-200b-5p | sarcoma | VEGFA | "CCL5 promotes VEGF dependent angiogenesis by down ......" | 25301739 |
| hsa-miR-200b-5p | thyroid cancer | ATM | "However specific miRNAs are downregulated in ATC s ......" | 25202329 |
| hsa-miR-200b-5p | thyroid cancer | ATM | "This study was set to study the molecular mechanis ......" | 25542369 |
| hsa-miR-200b-5p | lung cancer | E2F3 | "MicroRNA 200b reverses chemoresistance of docetaxe ......" | 22139708 |
| hsa-miR-200b-5p | lung cancer | E2F3 | "Previously we have shown that restoration of micro ......" | 24830600 |
| hsa-miR-200b-5p | ovarian cancer | GRHL2 | "GRHL2 miR 200 ZEB1 maintains the epithelial status ......" | 26887977 |
| hsa-miR-200b-5p | sarcoma | GRHL2 | "This circuit comprises the microRNA 200 miR-200 fa ......" | 27402864 |
| hsa-miR-200b-5p | lung cancer | HDAC9 | "HDAC 1/4 mediated silencing of microRNA 200b promo ......" | 24830600 |
| hsa-miR-200b-5p | lung cancer | HDAC9 | "Additionally overexpression of histone deacetylase ......" | 25279705 |
| hsa-miR-200b-5p | breast cancer | ITGA9 | "β1 integrin inhibition elicits a prometastatic sw ......" | 24518294 |
| hsa-miR-200b-5p | esophageal cancer | ITGA9 | "We revealed that miR-200b suppresses the integrin ......" | 26334393 |
| hsa-miR-200b-5p | endometrial cancer | MMP2 | "Using reverse gelatin zymography we showed that mi ......" | 23205572 |
| hsa-miR-200b-5p | ovarian cancer | MMP2 | "Interestingly our findings show that catalpol trea ......" | 25347277 |
| hsa-miR-200b-5p | ovarian cancer | MUC16 | "Plasma miR 200b in ovarian carcinoma patients: dis ......" | 26416421 |
| hsa-miR-200b-5p | ovarian cancer | MUC16 | "The increased levels of miR-200b and miR-200c were ......" | 26943577 |
| hsa-miR-200b-5p | lung cancer | NOTCH1 | "Nobiletin inhibited hypoxia induced epithelial mes ......" | 26012256 |
| hsa-miR-200b-5p | sarcoma | NOTCH1 | "We also found that reexpression of miR-34a and miR ......" | 23430952 |
| hsa-miR-200b-5p | pancreatic cancer | PTEN | "Re expression of miR 200 by novel approaches regul ......" | 22637745 |
| hsa-miR-200b-5p | pancreatic cancer | PTEN | "Anti tumor activity of a novel compound CDF is med ......" | 21408027 |
| hsa-miR-200b-5p | lung cancer | QRSL1 | "It was reported that miR-200 can activate PI3K/AKT ......" | 26314828 |
| hsa-miR-200b-5p | lung cancer | QRSL1 | "Mechanistically Jagged2 was found to promote metas ......" | 21403400 |
| hsa-miR-200b-5p | breast cancer | TAM | "miR-200 family expression was progressively reduce ......" | 23626803 |
| hsa-miR-200b-5p | endometrial cancer | TAM | "When treated with TAM ECC-1 and Ishikawa cells wer ......" | 23295740 |
| hsa-miR-200b-5p | breast cancer | ADAM12 | "ADAM12 L is a direct target of the miR 29 and miR ......" | 25886595 |
| hsa-miR-200b-5p | prostate cancer | AR | "We identified miR-200 b as a downstream target of ......" | 24391862 |
| hsa-miR-200b-5p | lung cancer | ATG12 | "In the present study an inverse correlation betwee ......" | 26416454 |
| hsa-miR-200b-5p | lung squamous cell cancer | ATRX | "Additionally miR-200 family downregulates HNRNPR3 ......" | 23708087 |
| hsa-miR-200b-5p | B cell lymphoma | BCL6 | "In 30 H pylori-positive and 30 H pylori-negative g ......" | 24390222 |
| hsa-miR-200b-5p | liver cancer | BMI1 | "Methylation associated silencing of miR 200b facil ......" | 26919246 |
| hsa-miR-200b-5p | lung cancer | BMP4 | "BMP4 depletion by miR 200 inhibits tumorigenesis a ......" | 26395571 |
| hsa-miR-200b-5p | colon cancer | CCL16 | "This observation indicates a common molecular axis ......" | 25832648 |
| hsa-miR-200b-5p | sarcoma | CCL5 | "CCL5 promotes VEGF dependent angiogenesis by down ......" | 25301739 |
| hsa-miR-200b-5p | colorectal cancer | CD2AP | "Importantly epigenetic silencing of the miR-200 fa ......" | 27157610 |
| hsa-miR-200b-5p | pancreatic cancer | CD44 | "In the current study we showed for the first time ......" | 21408027 |
| hsa-miR-200b-5p | esophageal cancer | CDK2 | "Correlating with the frequent loss of miR-200b in ......" | 27496804 |
| hsa-miR-200b-5p | colorectal cancer | CDKN1B | "This study suggests that miR-200b plays a tumour-p ......" | 24151081 |
| hsa-miR-200b-5p | melanoma | CHL1 | "Overall our findings call into question the genera ......" | 20957176 |
| hsa-miR-200b-5p | thyroid cancer | CHST14 | "We identified two significantly decreased microRNA ......" | 20498632 |
| hsa-miR-200b-5p | liver cancer | DNMT1 | "After the mesenchymal cells were treated with comb ......" | 23552699 |
| hsa-miR-200b-5p | bladder cancer | EGF | "miR 200 expression regulates epithelial to mesench ......" | 19671845 |
| hsa-miR-200b-5p | bladder cancer | EGFR | "Protein expression and signaling pathway modulatio ......" | 19671845 |
| hsa-miR-200b-5p | pancreatic cancer | EPCAM | "In the current study we showed for the first time ......" | 21408027 |
| hsa-miR-200b-5p | prostate cancer | ERG | "The tumor suppressive miR 200b subfamily is an ERG ......" | 27191272 |
| hsa-miR-200b-5p | breast cancer | ESR1 | "Further miR-200b and miR-200c overexpression sensi ......" | 23626803 |
| hsa-miR-200b-5p | breast cancer | ETV5 | "This study aimed to investigate the effect of miR- ......" | 27276064 |
| hsa-miR-200b-5p | breast cancer | EXOSC10 | "The novel promoter of the Hsa-mir-200b cluster den ......" | 22231446 |
| hsa-miR-200b-5p | lung cancer | FLT1 | "Forced miR-200 expression suppressed Flt1 levels i ......" | 21115742 |
| hsa-miR-200b-5p | prostate cancer | FN1 | "In contrast mesenchymal markers fibronectin and vi ......" | 24391862 |
| hsa-miR-200b-5p | lung cancer | FOXF2 | "The miR 200 family and the miR 183~96~182 cluster ......" | 25798833 |
| hsa-miR-200b-5p | cervical and endocervical cancer | FOXG1 | "MiR 200b promotes the cell proliferation and metas ......" | 27044840 |
| hsa-miR-200b-5p | breast cancer | FOXM1 | "SUMOylation of FOXM1B alters its transcriptional a ......" | 24918286 |
| hsa-miR-200b-5p | lung squamous cell cancer | FSCN1 | "miR 200b inhibits migration and invasion in non sm ......" | 27356635 |
| hsa-miR-200b-5p | lung cancer | GATA3 | "Reciprocally miR-200 inhibited expression of Gata3 ......" | 21403400 |
| hsa-miR-200b-5p | prostate cancer | GNA13 | "These data provide strong evidence that GNA13 is a ......" | 23329838 |
| hsa-miR-200b-5p | liver cancer | H19 | "Epigenetic activation of the MiR 200 family contri ......" | 23222811 |
| hsa-miR-200b-5p | lung cancer | HDAC1 | "Additionally overexpression of histone deacetylase ......" | 25279705 |
| hsa-miR-200b-5p | lung squamous cell cancer | HFE | "Additionally miR-200 family downregulates HNRNPR3 ......" | 23708087 |
| hsa-miR-200b-5p | lung squamous cell cancer | HMI | "The miR-200 family and these potential targets are ......" | 23708087 |
| hsa-miR-200b-5p | liver cancer | IBSP | "Methylation specific PCR MSP and bisulfite sequenc ......" | 26919246 |
| hsa-miR-200b-5p | breast cancer | IKBKB | "Importantly IKBKB overexpression attenuates the in ......" | 26433127 |
| hsa-miR-200b-5p | lung cancer | IRS1 | "Taken together our results suggest that miR-200 ma ......" | 26314828 |
| hsa-miR-200b-5p | lung cancer | JAG2 | "The Notch ligand Jagged2 promotes lung adenocarcin ......" | 21403400 |
| hsa-miR-200b-5p | endometrial cancer | KCNE1 | "Furthermore it was observed that the expression pa ......" | 26788150 |
| hsa-miR-200b-5p | esophageal cancer | KIAA0101 | "Correlating with the frequent loss of miR-200b in ......" | 27496804 |
| hsa-miR-200b-5p | colorectal cancer | KRAS | "In KRAS mutated tumours increased miR-200b and dec ......" | 22804917 |
| hsa-miR-200b-5p | endometrial cancer | MALAT1 | "The abnormal expression of the long noncoding RNA ......" | 27693631 |
| hsa-miR-200b-5p | breast cancer | MAPK8 | "SUMOylation of FOXM1B alters its transcriptional a ......" | 24918286 |
| hsa-miR-200b-5p | pancreatic cancer | MMP14 | "Re expression of miR 200 by novel approaches regul ......" | 22637745 |
| hsa-miR-200b-5p | breast cancer | MSN | "MiR 200 can repress breast cancer metastasis throu ......" | 24037528 |
| hsa-miR-200b-5p | endometrial cancer | MYC | "Tamoxifen represses miR 200 microRNAs and promotes ......" | 23295740 |
| hsa-miR-200b-5p | breast cancer | NR4A1 | "Mapping the regulatory sequences controlling 93 br ......" | 22231446 |
| hsa-miR-200b-5p | breast cancer | P1 | "P2 has comparable promoter activity to the previou ......" | 22231446 |
| hsa-miR-200b-5p | prostate cancer | PDGFD | "miR 200 regulates PDGF D mediated epithelial mesen ......" | 19544444 |
| hsa-miR-200b-5p | breast cancer | PELP1 | "Whole genome miR array analysis using PELP1-overex ......" | 23975430 |
| hsa-miR-200b-5p | breast cancer | PHLDA3 | "Our data indicate that Kindlin 2 plays a novel rol ......" | 23483548 |
| hsa-miR-200b-5p | breast cancer | PRDX2 | "Demethylating agent 5-aza-2'-deoxycytidine 5-aza-d ......" | 23626803 |
| hsa-miR-200b-5p | pancreatic cancer | PTGS2 | "In a xenograft mouse model of human PC CDF treatme ......" | 21408027 |
| hsa-miR-200b-5p | cervical and endocervical cancer | PVT1 | "Long noncoding RNA PVT1 promotes cervical cancer p ......" | 27272214 |
| hsa-miR-200b-5p | breast cancer | RAC1 | "Further mechanistic studies revealed that PKCα do ......" | 24925028 |
| hsa-miR-200b-5p | cervical and endocervical cancer | RND3 | "MicroRNA 200b inhibits epithelial mesenchymal tran ......" | 26935796 |
| hsa-miR-200b-5p | breast cancer | SGSM3 | "The genes encoding microRNAs of the human miR-200 ......" | 20514023 |
| hsa-miR-200b-5p | breast cancer | SLC10A4 | "Here we investigated P4 downregulation of miR-141 ......" | 25241899 |
| hsa-miR-200b-5p | colorectal cancer | SNAI1 | "EMT-TFs and microRNAs such as ZEB1/2 and miR-200 o ......" | 27573895 |
| hsa-miR-200b-5p | breast cancer | SP1 | "MiR 200b expression in breast cancer: a prognostic ......" | 25639535 |
| hsa-miR-200b-5p | endometrial cancer | SPEN | "This finding was accompanied by a sharp downregula ......" | 23743934 |
| hsa-miR-200b-5p | thyroid cancer | TGFB1 | "Inhibition of TGFbeta receptor 1 TGFBR1 in these c ......" | 20498632 |
| hsa-miR-200b-5p | thyroid cancer | TGFBR1 | "Inhibition of TGFbeta receptor 1 TGFBR1 in these c ......" | 20498632 |
| hsa-miR-200b-5p | colorectal cancer | TGFBR2 | "MicroRNA 200b stimulates tumour growth in TGFBR2 n ......" | 24151081 |
| hsa-miR-200b-5p | endometrial cancer | TIMP2 | "A novel target of miR-200b tissue inhibitor of met ......" | 23205572 |
| hsa-miR-200b-5p | breast cancer | TP53BP1 | "Here we show that 53BP1 negatively regulated EMT b ......" | 26011542 |
| hsa-miR-200b-5p | prostate cancer | TP73 | "Down regulation of miR 200b 3p by low p73 contribu ......" | 23389960 |
| hsa-miR-200b-5p | bladder cancer | TWIST1 | "In addition we observe that the mesoderm transcrip ......" | 20473948 |
| hsa-miR-200b-5p | lung cancer | ZFPM2 | "It was reported that miR-200 can activate PI3K/AKT ......" | 26314828 |
Expression profile in cancer corhorts:

Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-200b-5p | AFF3 | 12 cancers: BLCA; CESC; COAD; ESCA; KIRC; KIRP; LUSC; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.463; TCGA CESC -0.415; TCGA COAD -0.68; TCGA ESCA -0.383; TCGA KIRC -0.348; TCGA KIRP -0.296; TCGA LUSC -0.703; TCGA PAAD -0.583; TCGA PRAD -0.196; TCGA THCA -0.125; TCGA STAD -0.553; TCGA UCEC -0.278 |
hsa-miR-200b-5p | THBS1 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.404; TCGA CESC -0.501; TCGA COAD -0.585; TCGA ESCA -0.429; TCGA HNSC -0.429; TCGA LUAD -0.19; TCGA LUSC -0.493; TCGA PAAD -0.552; TCGA STAD -0.342; TCGA UCEC -0.37 |
hsa-miR-200b-5p | TXLNB | 13 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.367; TCGA CESC -0.389; TCGA COAD -0.633; TCGA ESCA -0.549; TCGA HNSC -0.532; TCGA KIRC -0.447; TCGA KIRP -0.216; TCGA LUAD -0.196; TCGA LUSC -0.158; TCGA PAAD -0.668; TCGA THCA -0.136; TCGA STAD -0.507; TCGA UCEC -0.427 |
hsa-miR-200b-5p | RFTN2 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.157; TCGA BRCA -0.277; TCGA CESC -0.217; TCGA COAD -0.43; TCGA ESCA -0.089; TCGA HNSC -0.153; TCGA KIRC -0.213; TCGA LUAD -0.135; TCGA LUSC -0.301; TCGA OV -0.133; TCGA PAAD -0.29; TCGA PRAD -0.073; TCGA STAD -0.232; TCGA UCEC -0.36 |
hsa-miR-200b-5p | ST6GALNAC3 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.228; TCGA BRCA -0.257; TCGA CESC -0.387; TCGA COAD -0.244; TCGA HNSC -0.26; TCGA LUSC -0.44; TCGA OV -0.233; TCGA PAAD -0.248; TCGA PRAD -0.172; TCGA THCA -0.054; TCGA STAD -0.197; TCGA UCEC -0.401 |
hsa-miR-200b-5p | EID1 | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.052; TCGA BRCA -0.069; TCGA CESC -0.1; TCGA COAD -0.173; TCGA ESCA -0.094; TCGA HNSC -0.145; TCGA KIRP -0.101; TCGA PAAD -0.126; TCGA STAD -0.258; TCGA UCEC -0.173 |
hsa-miR-200b-5p | STX2 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.362; TCGA BRCA -0.085; TCGA CESC -0.175; TCGA COAD -0.222; TCGA ESCA -0.168; TCGA HNSC -0.315; TCGA KIRC -0.122; TCGA LUAD -0.101; TCGA LUSC -0.367; TCGA OV -0.126; TCGA PAAD -0.215; TCGA PRAD -0.15; TCGA STAD -0.284; TCGA UCEC -0.263 |
hsa-miR-200b-5p | HCFC2 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.091; TCGA BRCA -0.076; TCGA CESC -0.13; TCGA COAD -0.233; TCGA ESCA -0.141; TCGA HNSC -0.117; TCGA LUAD -0.092; TCGA LUSC -0.154; TCGA OV -0.073; TCGA PAAD -0.204; TCGA STAD -0.234; TCGA UCEC -0.205 |
hsa-miR-200b-5p | RIMKLB | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LUSC; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.185; TCGA BRCA -0.062; TCGA CESC -0.12; TCGA COAD -0.643; TCGA ESCA -0.455; TCGA KIRC -0.131; TCGA KIRP -0.103; TCGA LUSC -0.088; TCGA PAAD -0.21; TCGA STAD -0.388; TCGA UCEC -0.089 |
hsa-miR-200b-5p | NCF2 | 11 cancers: BLCA; BRCA; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; PRAD | MirTarget | TCGA BLCA -0.429; TCGA BRCA -0.055; TCGA COAD -0.439; TCGA ESCA -0.299; TCGA HNSC -0.129; TCGA KIRC -0.386; TCGA KIRP -0.303; TCGA LUAD -0.257; TCGA LUSC -0.47; TCGA PAAD -0.321; TCGA PRAD -0.111 |
hsa-miR-200b-5p | MBNL1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LUAD; LUSC; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.111; TCGA BRCA -0.084; TCGA CESC -0.087; TCGA COAD -0.076; TCGA ESCA -0.161; TCGA KIRC -0.161; TCGA KIRP -0.06; TCGA LUAD -0.069; TCGA LUSC -0.056; TCGA PAAD -0.158; TCGA STAD -0.201; TCGA UCEC -0.16 |
hsa-miR-200b-5p | RORA | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LUSC; OV; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.102; TCGA BRCA -0.114; TCGA CESC -0.148; TCGA COAD -0.499; TCGA ESCA -0.263; TCGA KIRC -0.156; TCGA LUSC -0.114; TCGA OV -0.095; TCGA PAAD -0.274; TCGA STAD -0.304; TCGA UCEC -0.227 |
hsa-miR-200b-5p | SIGLEC8 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LUAD; LUSC; PAAD; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.296; TCGA BRCA -0.083; TCGA CESC -0.347; TCGA COAD -0.427; TCGA ESCA -0.255; TCGA KIRC -1.147; TCGA KIRP -0.416; TCGA LUAD -0.155; TCGA LUSC -0.68; TCGA PAAD -0.562; TCGA PRAD -0.137; TCGA THCA -0.2; TCGA UCEC -0.139 |
hsa-miR-200b-5p | KLF9 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.382; TCGA BRCA -0.265; TCGA CESC -0.225; TCGA COAD -0.34; TCGA ESCA -0.25; TCGA HNSC -0.199; TCGA LUAD -0.113; TCGA LUSC -0.426; TCGA OV -0.109; TCGA PAAD -0.306; TCGA STAD -0.394; TCGA UCEC -0.33 |
hsa-miR-200b-5p | ATP8B4 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.377; TCGA BRCA -0.184; TCGA CESC -0.366; TCGA COAD -0.347; TCGA ESCA -0.307; TCGA HNSC -0.229; TCGA KIRC -0.118; TCGA LUAD -0.179; TCGA LUSC -0.345; TCGA PAAD -0.468; TCGA PRAD -0.121; TCGA STAD -0.205; TCGA UCEC -0.363 |
hsa-miR-200b-5p | COPS8 | 10 cancers: BLCA; COAD; ESCA; HNSC; KIRP; OV; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.101; TCGA COAD -0.079; TCGA ESCA -0.079; TCGA HNSC -0.135; TCGA KIRP -0.182; TCGA OV -0.065; TCGA PAAD -0.138; TCGA PRAD -0.054; TCGA STAD -0.084; TCGA UCEC -0.072 |
hsa-miR-200b-5p | HIVEP3 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; STAD; UCEC | MirTarget | TCGA BLCA -0.229; TCGA CESC -0.262; TCGA COAD -0.176; TCGA ESCA -0.261; TCGA HNSC -0.124; TCGA LUAD -0.135; TCGA LUSC -0.265; TCGA OV -0.122; TCGA STAD -0.089; TCGA UCEC -0.067 |
hsa-miR-200b-5p | GAS7 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.484; TCGA BRCA -0.312; TCGA CESC -0.445; TCGA COAD -0.729; TCGA ESCA -0.395; TCGA HNSC -0.307; TCGA KIRP -0.17; TCGA LUAD -0.21; TCGA LUSC -0.582; TCGA PAAD -0.49; TCGA PRAD -0.098; TCGA STAD -0.415; TCGA UCEC -0.263 |
hsa-miR-200b-5p | BCAT1 | 12 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; PRAD; STAD | mirMAP | TCGA BLCA -0.283; TCGA CESC -0.236; TCGA COAD -0.644; TCGA ESCA -0.331; TCGA HNSC -0.232; TCGA KIRC -0.379; TCGA KIRP -0.384; TCGA LUAD -0.216; TCGA LUSC -0.245; TCGA PAAD -0.326; TCGA PRAD -0.159; TCGA STAD -0.132 |
hsa-miR-200b-5p | ADD2 | 9 cancers: BLCA; CESC; COAD; ESCA; KIRC; KIRP; PAAD; THCA; STAD | mirMAP | TCGA BLCA -0.262; TCGA CESC -0.315; TCGA COAD -1.21; TCGA ESCA -0.725; TCGA KIRC -0.478; TCGA KIRP -0.227; TCGA PAAD -0.691; TCGA THCA -0.165; TCGA STAD -0.528 |
hsa-miR-200b-5p | FGFR1 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUAD; LUSC; PAAD; PRAD; THCA; STAD | mirMAP | TCGA BLCA -0.423; TCGA BRCA -0.166; TCGA CESC -0.205; TCGA COAD -0.636; TCGA ESCA -0.38; TCGA HNSC -0.32; TCGA KIRP -0.146; TCGA LUAD -0.138; TCGA LUSC -0.104; TCGA PAAD -0.343; TCGA PRAD -0.128; TCGA THCA -0.077; TCGA STAD -0.423 |
hsa-miR-200b-5p | CYLD | 12 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.148; TCGA CESC -0.125; TCGA COAD -0.148; TCGA ESCA -0.206; TCGA HNSC -0.132; TCGA KIRC -0.052; TCGA KIRP -0.053; TCGA LUAD -0.089; TCGA LUSC -0.249; TCGA PAAD -0.151; TCGA STAD -0.138; TCGA UCEC -0.1 |
hsa-miR-200b-5p | HEPH | 10 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.435; TCGA BRCA -0.102; TCGA CESC -0.375; TCGA HNSC -0.375; TCGA KIRC -0.143; TCGA LUAD -0.098; TCGA LUSC -0.393; TCGA PRAD -0.189; TCGA STAD -0.118; TCGA UCEC -0.324 |
hsa-miR-200b-5p | PTGS1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; PRAD; THCA; STAD | mirMAP | TCGA BLCA -0.701; TCGA BRCA -0.073; TCGA CESC -0.366; TCGA COAD -0.433; TCGA ESCA -0.26; TCGA HNSC -0.206; TCGA LUAD -0.179; TCGA LUSC -0.385; TCGA PAAD -0.365; TCGA PRAD -0.219; TCGA THCA -0.061; TCGA STAD -0.284 |
hsa-miR-200b-5p | FKBP5 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; LUAD; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.416; TCGA BRCA -0.135; TCGA CESC -0.167; TCGA COAD -0.106; TCGA ESCA -0.394; TCGA KIRP -0.319; TCGA LUAD -0.306; TCGA LUSC -0.281; TCGA OV -0.13; TCGA PAAD -0.492; TCGA STAD -0.343; TCGA UCEC -0.237 |
hsa-miR-200b-5p | PPARGC1A | 10 cancers: BLCA; BRCA; CESC; HNSC; KIRP; LUSC; OV; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.419; TCGA BRCA -0.223; TCGA CESC -0.399; TCGA HNSC -0.314; TCGA KIRP -0.236; TCGA LUSC -0.228; TCGA OV -0.305; TCGA THCA -0.253; TCGA STAD -0.171; TCGA UCEC -0.317 |
hsa-miR-200b-5p | PRRX1 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.599; TCGA BRCA -0.244; TCGA CESC -0.428; TCGA COAD -0.832; TCGA ESCA -0.576; TCGA HNSC -0.482; TCGA KIRC -0.448; TCGA LUAD -0.206; TCGA LUSC -0.149; TCGA OV -0.264; TCGA PAAD -0.527; TCGA PRAD -0.102; TCGA STAD -0.352; TCGA UCEC -0.325 |
hsa-miR-200b-5p | KLF12 | 9 cancers: BLCA; COAD; ESCA; HNSC; LUSC; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.249; TCGA COAD -0.348; TCGA ESCA -0.274; TCGA HNSC -0.095; TCGA LUSC -0.184; TCGA PAAD -0.27; TCGA PRAD -0.1; TCGA STAD -0.333; TCGA UCEC -0.194 |
hsa-miR-200b-5p | SEPT7 | 11 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; OV; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.067; TCGA CESC -0.096; TCGA COAD -0.111; TCGA ESCA -0.099; TCGA HNSC -0.086; TCGA KIRC -0.13; TCGA LUAD -0.056; TCGA OV -0.077; TCGA PAAD -0.126; TCGA STAD -0.145; TCGA UCEC -0.15 |
hsa-miR-200b-5p | TMOD2 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.119; TCGA BRCA -0.081; TCGA CESC -0.289; TCGA COAD -0.348; TCGA ESCA -0.124; TCGA HNSC -0.211; TCGA KIRC -0.241; TCGA LUAD -0.186; TCGA LUSC -0.322; TCGA OV -0.104; TCGA PAAD -0.29; TCGA STAD -0.326; TCGA UCEC -0.417 |
hsa-miR-200b-5p | NOTCH2 | 9 cancers: BLCA; CESC; COAD; ESCA; LUAD; LUSC; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.127; TCGA CESC -0.076; TCGA COAD -0.182; TCGA ESCA -0.16; TCGA LUAD -0.08; TCGA LUSC -0.143; TCGA PAAD -0.187; TCGA STAD -0.202; TCGA UCEC -0.11 |
hsa-miR-200b-5p | DZIP1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.225; TCGA BRCA -0.132; TCGA CESC -0.293; TCGA COAD -0.73; TCGA ESCA -0.586; TCGA HNSC -0.17; TCGA LUAD -0.116; TCGA OV -0.149; TCGA PAAD -0.385; TCGA PRAD -0.07; TCGA STAD -0.481; TCGA UCEC -0.373 |
hsa-miR-200b-5p | TTLL7 | 9 cancers: BLCA; COAD; ESCA; HNSC; KIRC; LUSC; PAAD; THCA; STAD | mirMAP | TCGA BLCA -0.197; TCGA COAD -0.528; TCGA ESCA -0.208; TCGA HNSC -0.18; TCGA KIRC -0.365; TCGA LUSC -0.245; TCGA PAAD -0.186; TCGA THCA -0.139; TCGA STAD -0.547 |
hsa-miR-200b-5p | ENTPD1 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.192; TCGA BRCA -0.052; TCGA CESC -0.187; TCGA COAD -0.345; TCGA ESCA -0.118; TCGA HNSC -0.133; TCGA KIRC -0.261; TCGA LUAD -0.131; TCGA LUSC -0.253; TCGA OV -0.077; TCGA PAAD -0.281; TCGA PRAD -0.108; TCGA STAD -0.2; TCGA UCEC -0.185 |
hsa-miR-200b-5p | NTRK3 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.383; TCGA BRCA -0.286; TCGA CESC -0.569; TCGA COAD -0.975; TCGA ESCA -0.525; TCGA LUAD -0.17; TCGA LUSC -0.598; TCGA OV -0.358; TCGA PAAD -0.415; TCGA PRAD -0.099; TCGA STAD -0.629; TCGA UCEC -0.485 |
hsa-miR-200b-5p | ABL2 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; PAAD; STAD | mirMAP | TCGA BLCA -0.133; TCGA CESC -0.126; TCGA COAD -0.163; TCGA ESCA -0.156; TCGA HNSC -0.181; TCGA KIRC -0.243; TCGA LUAD -0.097; TCGA LUSC -0.151; TCGA PAAD -0.264; TCGA STAD -0.077 |
hsa-miR-200b-5p | RNF217 | 9 cancers: BLCA; BRCA; COAD; ESCA; KIRC; KIRP; LUAD; PAAD; STAD | mirMAP | TCGA BLCA -0.304; TCGA BRCA -0.14; TCGA COAD -0.557; TCGA ESCA -0.65; TCGA KIRC -0.183; TCGA KIRP -0.116; TCGA LUAD -0.124; TCGA PAAD -0.319; TCGA STAD -0.423 |
hsa-miR-200b-5p | GFRA1 | 12 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.305; TCGA CESC -0.736; TCGA COAD -0.86; TCGA ESCA -0.442; TCGA HNSC -0.185; TCGA LUSC -0.453; TCGA OV -0.429; TCGA PAAD -0.459; TCGA PRAD -0.156; TCGA THCA -0.275; TCGA STAD -0.682; TCGA UCEC -0.502 |
hsa-miR-200b-5p | CAMK4 | 11 cancers: BLCA; CESC; COAD; ESCA; KIRC; LUAD; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.263; TCGA CESC -0.209; TCGA COAD -0.323; TCGA ESCA -0.27; TCGA KIRC -0.2; TCGA LUAD -0.174; TCGA LUSC -0.178; TCGA PRAD -0.117; TCGA THCA -0.097; TCGA STAD -0.187; TCGA UCEC -0.212 |
hsa-miR-200b-5p | LGI2 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUSC; OV; PAAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.439; TCGA BRCA -0.179; TCGA CESC -0.399; TCGA COAD -0.678; TCGA ESCA -0.543; TCGA HNSC -0.351; TCGA LUSC -0.188; TCGA OV -0.161; TCGA PAAD -0.472; TCGA THCA -0.102; TCGA STAD -0.528; TCGA UCEC -0.516 |
hsa-miR-200b-5p | MMP16 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.304; TCGA BRCA -0.22; TCGA CESC -0.383; TCGA COAD -0.671; TCGA ESCA -0.301; TCGA HNSC -0.58; TCGA KIRC -0.873; TCGA LUSC -0.26; TCGA OV -0.343; TCGA PAAD -0.472; TCGA PRAD -0.142; TCGA STAD -0.399; TCGA UCEC -0.293 |
hsa-miR-200b-5p | FBXO32 | 11 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRP; LUAD; PAAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.307; TCGA CESC -0.253; TCGA COAD -0.458; TCGA ESCA -0.34; TCGA HNSC -0.283; TCGA KIRP -0.278; TCGA LUAD -0.147; TCGA PAAD -0.238; TCGA THCA -0.095; TCGA STAD -0.546; TCGA UCEC -0.288 |
hsa-miR-200b-5p | PALM2-AKAP2 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.392; TCGA BRCA -0.258; TCGA CESC -0.385; TCGA COAD -0.404; TCGA ESCA -0.221; TCGA HNSC -0.388; TCGA KIRC -0.159; TCGA LUAD -0.163; TCGA LUSC -0.651; TCGA OV -0.081; TCGA PAAD -0.368; TCGA STAD -0.294; TCGA UCEC -0.332 |
hsa-miR-200b-5p | APBB2 | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LUAD; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.153; TCGA BRCA -0.061; TCGA CESC -0.182; TCGA ESCA -0.079; TCGA HNSC -0.282; TCGA LUAD -0.062; TCGA LUSC -0.308; TCGA OV -0.08; TCGA PAAD -0.152; TCGA STAD -0.149; TCGA UCEC -0.091 |
hsa-miR-200b-5p | AMOTL1 | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.242; TCGA BRCA -0.194; TCGA CESC -0.262; TCGA COAD -0.829; TCGA ESCA -0.355; TCGA HNSC -0.074; TCGA KIRC -0.194; TCGA PAAD -0.33; TCGA STAD -0.46; TCGA UCEC -0.181 |
hsa-miR-200b-5p | PYGO1 | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRP; LUSC; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.232; TCGA BRCA -0.211; TCGA CESC -0.286; TCGA ESCA -0.373; TCGA HNSC -0.166; TCGA KIRP -0.34; TCGA LUSC -0.154; TCGA PAAD -0.414; TCGA STAD -0.541; TCGA UCEC -0.37 |
hsa-miR-200b-5p | SYNPO2 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.528; TCGA BRCA -0.399; TCGA CESC -0.438; TCGA COAD -1.229; TCGA ESCA -0.65; TCGA HNSC -0.409; TCGA LUAD -0.2; TCGA LUSC -0.308; TCGA OV -0.155; TCGA PAAD -0.484; TCGA PRAD -0.107; TCGA THCA -0.075; TCGA STAD -0.988; TCGA UCEC -0.597 |
hsa-miR-200b-5p | FAM26E | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.403; TCGA BRCA -0.25; TCGA CESC -0.41; TCGA COAD -0.775; TCGA ESCA -0.305; TCGA HNSC -0.6; TCGA KIRC -0.128; TCGA LUAD -0.267; TCGA LUSC -0.424; TCGA PAAD -0.396; TCGA PRAD -0.138; TCGA THCA -0.071; TCGA STAD -0.244; TCGA UCEC -0.335 |
hsa-miR-200b-5p | SLC36A4 | 9 cancers: BLCA; ESCA; HNSC; KIRP; LUAD; LUSC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.153; TCGA ESCA -0.091; TCGA HNSC -0.087; TCGA KIRP -0.074; TCGA LUAD -0.06; TCGA LUSC -0.126; TCGA THCA -0.067; TCGA STAD -0.163; TCGA UCEC -0.172 |
hsa-miR-200b-5p | PLCXD3 | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; LUSC; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.162; TCGA BRCA -0.592; TCGA CESC -0.357; TCGA COAD -1.163; TCGA ESCA -0.419; TCGA LUSC -0.716; TCGA PAAD -0.394; TCGA PRAD -0.108; TCGA STAD -0.642; TCGA UCEC -0.621 |
hsa-miR-200b-5p | PRELP | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.507; TCGA BRCA -0.419; TCGA CESC -0.429; TCGA COAD -1.31; TCGA ESCA -0.671; TCGA HNSC -0.239; TCGA LUAD -0.131; TCGA LUSC -0.592; TCGA PAAD -0.288; TCGA PRAD -0.076; TCGA STAD -0.807; TCGA UCEC -0.581 |
hsa-miR-200b-5p | BEND4 | 9 cancers: BLCA; COAD; ESCA; LUAD; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.191; TCGA COAD -0.49; TCGA ESCA -0.323; TCGA LUAD -0.279; TCGA LUSC -0.384; TCGA OV -0.335; TCGA PAAD -0.598; TCGA STAD -0.181; TCGA UCEC -0.373 |
hsa-miR-200b-5p | ADH1B | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.5; TCGA BRCA -1.003; TCGA CESC -0.506; TCGA COAD -1.219; TCGA ESCA -0.517; TCGA LUSC -1.023; TCGA OV -0.448; TCGA PAAD -0.752; TCGA PRAD -0.356; TCGA STAD -0.722; TCGA UCEC -0.816 |
hsa-miR-200b-5p | TCF4 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.244; TCGA BRCA -0.242; TCGA CESC -0.142; TCGA COAD -0.566; TCGA ESCA -0.263; TCGA HNSC -0.156; TCGA KIRC -0.36; TCGA LUAD -0.125; TCGA LUSC -0.1; TCGA OV -0.144; TCGA PAAD -0.377; TCGA PRAD -0.058; TCGA STAD -0.264; TCGA UCEC -0.25 |
hsa-miR-200b-5p | ITGA1 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.269; TCGA BRCA -0.178; TCGA CESC -0.271; TCGA COAD -0.277; TCGA ESCA -0.13; TCGA HNSC -0.325; TCGA KIRC -0.348; TCGA KIRP -0.222; TCGA LUAD -0.148; TCGA LUSC -0.476; TCGA OV -0.118; TCGA PAAD -0.195; TCGA PRAD -0.102; TCGA STAD -0.353; TCGA UCEC -0.15 |
hsa-miR-200b-5p | ZNF788 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.196; TCGA CESC -0.315; TCGA COAD -0.432; TCGA ESCA -0.364; TCGA HNSC -0.243; TCGA LUAD -0.137; TCGA LUSC -0.279; TCGA PAAD -0.277; TCGA STAD -0.268; TCGA UCEC -0.119 |
hsa-miR-200b-5p | AKAP2 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PAAD; UCEC | mirMAP | TCGA BLCA -0.478; TCGA BRCA -0.215; TCGA CESC -0.54; TCGA COAD -0.479; TCGA ESCA -0.205; TCGA HNSC -0.386; TCGA LUAD -0.221; TCGA LUSC -0.801; TCGA OV -0.104; TCGA PAAD -0.352; TCGA UCEC -0.534 |
hsa-miR-200b-5p | KCTD7 | 10 cancers: BLCA; CESC; COAD; ESCA; KIRC; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.081; TCGA CESC -0.118; TCGA COAD -0.147; TCGA ESCA -0.088; TCGA KIRC -0.057; TCGA LUSC -0.109; TCGA OV -0.121; TCGA PAAD -0.146; TCGA STAD -0.268; TCGA UCEC -0.238 |
hsa-miR-200b-5p | CRIM1 | 9 cancers: BRCA; CESC; COAD; ESCA; HNSC; LUSC; OV; STAD; UCEC | MirTarget | TCGA BRCA -0.134; TCGA CESC -0.228; TCGA COAD -0.105; TCGA ESCA -0.105; TCGA HNSC -0.14; TCGA LUSC -0.242; TCGA OV -0.072; TCGA STAD -0.209; TCGA UCEC -0.253 |
hsa-miR-200b-5p | KIAA0355 | 9 cancers: BRCA; CESC; COAD; ESCA; KIRC; LUSC; PAAD; STAD; UCEC | MirTarget | TCGA BRCA -0.102; TCGA CESC -0.104; TCGA COAD -0.11; TCGA ESCA -0.081; TCGA KIRC -0.117; TCGA LUSC -0.205; TCGA PAAD -0.122; TCGA STAD -0.136; TCGA UCEC -0.131 |
hsa-miR-200b-5p | NPHP3 | 10 cancers: BRCA; CESC; ESCA; HNSC; KIRC; LUAD; OV; PAAD; STAD; UCEC | MirTarget | TCGA BRCA -0.125; TCGA CESC -0.086; TCGA ESCA -0.153; TCGA HNSC -0.056; TCGA KIRC -0.231; TCGA LUAD -0.055; TCGA OV -0.088; TCGA PAAD -0.108; TCGA STAD -0.204; TCGA UCEC -0.145 |
hsa-miR-200b-5p | ZC3H12C | 9 cancers: BRCA; CESC; ESCA; HNSC; KIRP; LUSC; PAAD; STAD; UCEC | MirTarget | TCGA BRCA -0.125; TCGA CESC -0.163; TCGA ESCA -0.156; TCGA HNSC -0.101; TCGA KIRP -0.107; TCGA LUSC -0.129; TCGA PAAD -0.181; TCGA STAD -0.192; TCGA UCEC -0.231 |
hsa-miR-200b-5p | MCC | 9 cancers: BRCA; CESC; COAD; ESCA; LUAD; OV; PAAD; STAD; UCEC | MirTarget | TCGA BRCA -0.151; TCGA CESC -0.415; TCGA COAD -0.613; TCGA ESCA -0.51; TCGA LUAD -0.101; TCGA OV -0.197; TCGA PAAD -0.287; TCGA STAD -0.382; TCGA UCEC -0.257 |
hsa-miR-200b-5p | LIN7A | 12 cancers: BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUAD; LUSC; PAAD; THCA; STAD; UCEC | mirMAP | TCGA BRCA -0.268; TCGA CESC -0.277; TCGA COAD -0.5; TCGA ESCA -0.43; TCGA HNSC -0.295; TCGA KIRP -0.219; TCGA LUAD -0.193; TCGA LUSC -0.476; TCGA PAAD -0.333; TCGA THCA -0.253; TCGA STAD -0.205; TCGA UCEC -0.363 |
hsa-miR-200b-5p | ZNF677 | 10 cancers: BRCA; CESC; COAD; ESCA; HNSC; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BRCA -0.201; TCGA CESC -0.368; TCGA COAD -0.693; TCGA ESCA -0.457; TCGA HNSC -0.153; TCGA LUSC -0.479; TCGA OV -0.098; TCGA PAAD -0.324; TCGA STAD -0.487; TCGA UCEC -0.315 |
hsa-miR-200b-5p | ITSN1 | 9 cancers: BRCA; COAD; KIRC; LUAD; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BRCA -0.213; TCGA COAD -0.081; TCGA KIRC -0.059; TCGA LUAD -0.079; TCGA LUSC -0.159; TCGA OV -0.054; TCGA PAAD -0.135; TCGA STAD -0.127; TCGA UCEC -0.134 |
hsa-miR-200b-5p | IDS | 9 cancers: CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; STAD; UCEC | mirMAP | TCGA CESC -0.097; TCGA COAD -0.191; TCGA ESCA -0.106; TCGA HNSC -0.091; TCGA KIRC -0.097; TCGA LUAD -0.149; TCGA LUSC -0.166; TCGA STAD -0.208; TCGA UCEC -0.133 |
hsa-miR-200b-5p | PHTF2 | 9 cancers: COAD; ESCA; HNSC; KIRC; LUAD; OV; PAAD; STAD; UCEC | MirTarget | TCGA COAD -0.175; TCGA ESCA -0.202; TCGA HNSC -0.08; TCGA KIRC -0.153; TCGA LUAD -0.104; TCGA OV -0.052; TCGA PAAD -0.239; TCGA STAD -0.1; TCGA UCEC -0.075 |
Enriched cancer pathways of putative targets