microRNA information: hsa-miR-20b-5p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-20b-5p | miRbase |
| Accession: | MIMAT0001413 | miRbase |
| Precursor name: | hsa-mir-20b | miRbase |
| Precursor accession: | MI0001519 | miRbase |
| Symbol: | MIR20B | HGNC |
| RefSeq ID: | NR_029950 | GenBank |
| Sequence: | CAAAGUGCUCAUAGUGCAGGUAG |
Reported expression in cancers: hsa-miR-20b-5p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-20b-5p | cervical and endocervical cancer | upregulation | "Vote-counting analysis showed that up-regulation w ......" | 25920605 | |
| hsa-miR-20b-5p | cervical and endocervical cancer | upregulation | "Kaplan-Meier and log-rank analyses were used and i ......" | 26427662 | |
| hsa-miR-20b-5p | gastric cancer | upregulation | "Quantitative reverse transcriptase-polymerase chai ......" | 19148490 | qPCR |
| hsa-miR-20b-5p | gastric cancer | upregulation | "The most highly expressed miRNAs in gastric cancer ......" | 19175831 | |
| hsa-miR-20b-5p | gastric cancer | upregulation | "miR 20b overexpression is predictive of poor progn ......" | 26244024 | |
| hsa-miR-20b-5p | lung cancer | upregulation | "Taken together our data suggest that increased exp ......" | 26560875 | |
| hsa-miR-20b-5p | thyroid cancer | downregulation | "MiR 20b Displays Tumor Suppressor Functions in Pap ......" | 27717302 | qPCR |
Reported cancer pathway affected by hsa-miR-20b-5p
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-20b-5p | esophageal cancer | Apoptosis pathway | "Follow-up experimental verification of 7 miRNAs in ......" | 27462775 | |
| hsa-miR-20b-5p | esophageal cancer | Apoptosis pathway | "MicroRNA 20b miR 20b Promotes the Proliferation Mi ......" | 27701465 | Luciferase |
| hsa-miR-20b-5p | kidney renal cell cancer | Apoptosis pathway | "MicroRNA 20b 5p functions as a tumor suppressor in ......" | 26708577 | Flow cytometry |
Reported cancer prognosis affected by hsa-miR-20b-5p

| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-20b-5p | B cell lymphoma | poor survival | "Multivariate Cox analysis showed that higher miR-2 ......" | 25634356 | |
| hsa-miR-20b-5p | breast cancer | drug resistance; tumorigenesis | "Crucial role for early growth response 1 in the tr ......" | 23945289 | Luciferase; RNA pull-down |
| hsa-miR-20b-5p | breast cancer | recurrence | "Using microarray-based technology we have performe ......" | 24632820 | |
| hsa-miR-20b-5p | breast cancer | progression; tumorigenesis | "MicroRNA 20b promotes cell growth of breast cancer ......" | 25364498 | Luciferase; Colony formation |
| hsa-miR-20b-5p | breast cancer | metastasis | "miR 20b is up regulated in brain metastases from p ......" | 25893380 | Colony formation |
| hsa-miR-20b-5p | cervical and endocervical cancer | poor survival | "Kaplan-Meier and log-rank analyses were used and i ......" | 26427662 | |
| hsa-miR-20b-5p | colorectal cancer | staging | "MiR 20b 21 and 130b inhibit PTEN expression result ......" | 24468585 | Luciferase |
| hsa-miR-20b-5p | colorectal cancer | tumorigenesis | "Underexpression of miR 126 and miR 20b in heredita ......" | 24994098 | |
| hsa-miR-20b-5p | esophageal cancer | tumorigenesis | "MicroRNA 20b miR 20b Promotes the Proliferation Mi ......" | 27701465 | Luciferase |
| hsa-miR-20b-5p | gastric cancer | poor survival | "Quantitative reverse transcriptase-polymerase chai ......" | 19148490 | |
| hsa-miR-20b-5p | gastric cancer | worse prognosis; poor survival; staging; metastasis | "miR 20b overexpression is predictive of poor progn ......" | 26244024 | |
| hsa-miR-20b-5p | gastric cancer | drug resistance; progression | "Role of miR 27a miR 181a and miR 20b in gastric ca ......" | 26793992 | |
| hsa-miR-20b-5p | kidney renal cell cancer | cell migration | "MicroRNA 20b 5p functions as a tumor suppressor in ......" | 26708577 | Flow cytometry |
| hsa-miR-20b-5p | liver cancer | poor survival | "Clinicopathological Significance of MicroRNA 20b E ......" | 26612965 | |
| hsa-miR-20b-5p | lung squamous cell cancer | poor survival | "Five selected miRNAs let-7f miR-20b miR-30e-3p miR ......" | 20595154 | |
| hsa-miR-20b-5p | lymphoma | poor survival | "The expression of 12 selected miRNAs was studied b ......" | 20485376 | |
| hsa-miR-20b-5p | thyroid cancer | staging; metastasis; progression | "MiR 20b Displays Tumor Suppressor Functions in Pap ......" | 27717302 |
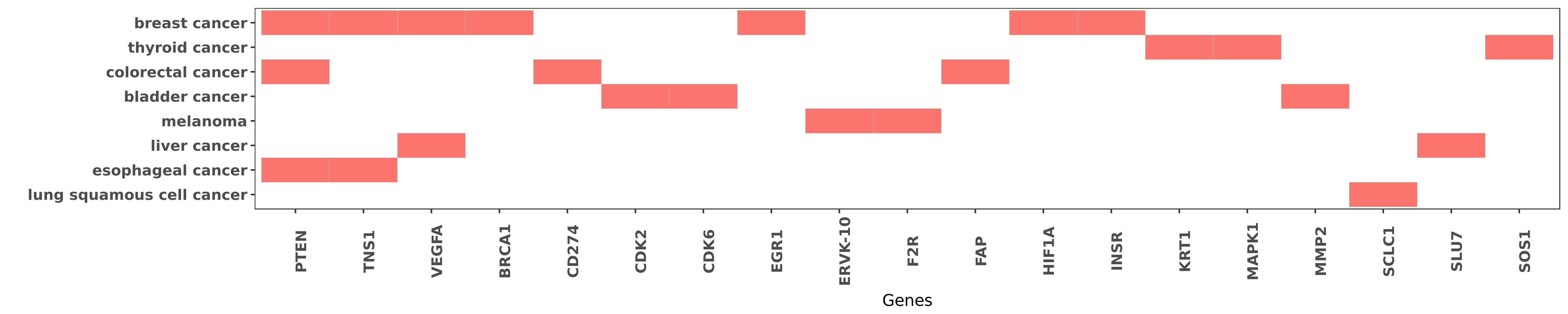
Reported gene related to hsa-miR-20b-5p
| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-miR-20b-5p | breast cancer | PTEN | "In vitro RNA-pull down assays indicated that miR-2 ......" | 23945289 |
| hsa-miR-20b-5p | breast cancer | PTEN | "MicroRNA 20b promotes cell growth of breast cancer ......" | 25364498 |
| hsa-miR-20b-5p | colorectal cancer | PTEN | "Finally the impact of these up-regulated miRNAs on ......" | 24468585 |
| hsa-miR-20b-5p | esophageal cancer | PTEN | "Additionally the 3'-untranslated region 3'-UTR of ......" | 27701465 |
| hsa-miR-20b-5p | breast cancer | VEGFA | "Correlation analysis showed that the key miRNAs mi ......" | 22901144 |
| hsa-miR-20b-5p | breast cancer | VEGFA | "We investigated whether and how miR-20b can regula ......" | 20232316 |
| hsa-miR-20b-5p | liver cancer | VEGFA | "Clinicopathological Significance of MicroRNA 20b E ......" | 26612965 |
| hsa-miR-20b-5p | breast cancer | TNS1 | "MicroRNA 20b promotes cell growth of breast cancer ......" | 25364498 |
| hsa-miR-20b-5p | esophageal cancer | TNS1 | "MicroRNA 20b miR 20b Promotes the Proliferation Mi ......" | 27701465 |
| hsa-miR-20b-5p | breast cancer | BRCA1 | "In vitro RNA-pull down assays indicated that miR-2 ......" | 23945289 |
| hsa-miR-20b-5p | colorectal cancer | CD274 | "These findings suggest that miR-20b -21 and -130b ......" | 24468585 |
| hsa-miR-20b-5p | bladder cancer | CDK2 | "The transfection of miR-20b into EJ cells induced ......" | 26166554 |
| hsa-miR-20b-5p | bladder cancer | CDK6 | "The transfection of miR-20b into EJ cells induced ......" | 26166554 |
| hsa-miR-20b-5p | breast cancer | EGR1 | "We identified several GC-rich consensus binding mo ......" | 23945289 |
| hsa-miR-20b-5p | melanoma | ERVK-10 | "miR 20b regulates expression of proteinase activat ......" | 24405508 |
| hsa-miR-20b-5p | melanoma | F2R | "miR 20b regulates expression of proteinase activat ......" | 24405508 |
| hsa-miR-20b-5p | colorectal cancer | FAP | "We analyzed the expressions of miR-126 and miR-20b ......" | 24994098 |
| hsa-miR-20b-5p | breast cancer | HIF1A | "Correlation analysis showed that the key miRNAs mi ......" | 22901144 |
| hsa-miR-20b-5p | breast cancer | INSR | "miR-20b was upregulated by IR and its upregulation ......" | 23945289 |
| hsa-miR-20b-5p | thyroid cancer | KRT1 | "MiR-20b was over-expressed in PTC cell lines K1 an ......" | 27717302 |
| hsa-miR-20b-5p | thyroid cancer | MAPK1 | "By targeting SOS1 and ERK2 miR-20b inhibits the ac ......" | 27717302 |
| hsa-miR-20b-5p | bladder cancer | MMP2 | "MicroRNA 20b inhibits the proliferation migration ......" | 26166554 |
| hsa-miR-20b-5p | lung squamous cell cancer | SCLC1 | "Among them five downregulated miRNAs let-7 miR-20 ......" | 23117485 |
| hsa-miR-20b-5p | liver cancer | SLU7 | "Interestingly altered splicing of miR-17-92 and do ......" | 26804174 |
| hsa-miR-20b-5p | thyroid cancer | SOS1 | "By targeting SOS1 and ERK2 miR-20b inhibits the ac ......" | 27717302 |
Expression profile in cancer corhorts:

Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-20b-5p | CDKN1A | 9 cancers: BLCA; BRCA; CESC; HNSC; KIRC; KIRP; LUSC; PRAD; THCA | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | TCGA BLCA -0.112; TCGA BRCA -0.056; TCGA CESC -0.077; TCGA HNSC -0.105; TCGA KIRC -0.138; TCGA KIRP -0.108; TCGA LUSC -0.167; TCGA PRAD -0.076; TCGA THCA -0.117 |
hsa-miR-20b-5p | CRIM1 | 9 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LUAD; LUSC; PRAD; UCEC | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | TCGA BLCA -0.077; TCGA BRCA -0.078; TCGA CESC -0.106; TCGA ESCA -0.081; TCGA HNSC -0.194; TCGA LUAD -0.07; TCGA LUSC -0.09; TCGA PRAD -0.236; TCGA UCEC -0.105 |
hsa-miR-20b-5p | CYBRD1 | 9 cancers: BLCA; BRCA; CESC; HNSC; LUAD; LUSC; PRAD; SARC; UCEC | MirTarget | TCGA BLCA -0.267; TCGA BRCA -0.091; TCGA CESC -0.095; TCGA HNSC -0.142; TCGA LUAD -0.229; TCGA LUSC -0.117; TCGA PRAD -0.277; TCGA SARC -0.116; TCGA UCEC -0.17 |
hsa-miR-20b-5p | JAK1 | 10 cancers: BLCA; CESC; ESCA; HNSC; KIRP; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.091; TCGA CESC -0.058; TCGA ESCA -0.05; TCGA HNSC -0.083; TCGA KIRP -0.071; TCGA LUAD -0.054; TCGA LUSC -0.102; TCGA PRAD -0.147; TCGA THCA -0.071; TCGA UCEC -0.075 |
hsa-miR-20b-5p | IL1RAP | 9 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LUSC; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.178; TCGA BRCA -0.067; TCGA CESC -0.178; TCGA HNSC -0.162; TCGA KIRC -0.213; TCGA LUSC -0.179; TCGA PRAD -0.162; TCGA THCA -0.495; TCGA UCEC -0.061 |
hsa-miR-20b-5p | PRRX1 | 9 cancers: BLCA; CESC; KIRC; LGG; LUAD; LUSC; PRAD; SARC; STAD | MirTarget; miRNATAP | TCGA BLCA -0.374; TCGA CESC -0.22; TCGA KIRC -0.201; TCGA LGG -0.133; TCGA LUAD -0.072; TCGA LUSC -0.178; TCGA PRAD -0.257; TCGA SARC -0.15; TCGA STAD -0.124 |
hsa-miR-20b-5p | TGM2 | 9 cancers: BLCA; CESC; HNSC; KIRC; KIRP; LIHC; LUSC; THCA; UCEC | MirTarget | TCGA BLCA -0.332; TCGA CESC -0.184; TCGA HNSC -0.098; TCGA KIRC -0.309; TCGA KIRP -0.319; TCGA LIHC -0.087; TCGA LUSC -0.249; TCGA THCA -0.344; TCGA UCEC -0.197 |
hsa-miR-20b-5p | ABCA1 | 9 cancers: BLCA; CESC; HNSC; KIRC; KIRP; LGG; LUAD; LUSC; PRAD | MirTarget; miRNATAP | TCGA BLCA -0.086; TCGA CESC -0.138; TCGA HNSC -0.111; TCGA KIRC -0.136; TCGA KIRP -0.08; TCGA LGG -0.169; TCGA LUAD -0.084; TCGA LUSC -0.089; TCGA PRAD -0.137 |
hsa-miR-20b-5p | ELK3 | 9 cancers: BLCA; CESC; HNSC; KIRC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.175; TCGA CESC -0.188; TCGA HNSC -0.206; TCGA KIRC -0.113; TCGA LUAD -0.093; TCGA LUSC -0.154; TCGA PRAD -0.257; TCGA THCA -0.101; TCGA UCEC -0.123 |
hsa-miR-20b-5p | ANKH | 9 cancers: BLCA; BRCA; CESC; COAD; HNSC; LIHC; LUAD; LUSC; THCA | MirTarget | TCGA BLCA -0.136; TCGA BRCA -0.068; TCGA CESC -0.121; TCGA COAD -0.069; TCGA HNSC -0.066; TCGA LIHC -0.052; TCGA LUAD -0.059; TCGA LUSC -0.2; TCGA THCA -0.164 |
hsa-miR-20b-5p | ZBTB4 | 9 cancers: BLCA; BRCA; CESC; HNSC; LUAD; LUSC; PRAD; SARC; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.082; TCGA BRCA -0.058; TCGA CESC -0.077; TCGA HNSC -0.095; TCGA LUAD -0.11; TCGA LUSC -0.094; TCGA PRAD -0.111; TCGA SARC -0.062; TCGA UCEC -0.105 |
hsa-miR-20b-5p | EDA2R | 10 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.119; TCGA BRCA -0.177; TCGA CESC -0.218; TCGA HNSC -0.162; TCGA KIRC -0.32; TCGA LUAD -0.154; TCGA LUSC -0.164; TCGA PRAD -0.279; TCGA THCA -0.143; TCGA UCEC -0.223 |
hsa-miR-20b-5p | ARHGAP1 | 9 cancers: BLCA; CESC; ESCA; HNSC; KIRC; KIRP; LUSC; PRAD; THCA | MirTarget; miRNATAP | TCGA BLCA -0.118; TCGA CESC -0.058; TCGA ESCA -0.087; TCGA HNSC -0.109; TCGA KIRC -0.087; TCGA KIRP -0.058; TCGA LUSC -0.128; TCGA PRAD -0.067; TCGA THCA -0.088 |
hsa-miR-20b-5p | FIBIN | 9 cancers: BLCA; BRCA; CESC; HNSC; LUAD; LUSC; PRAD; SARC; UCEC | MirTarget | TCGA BLCA -0.416; TCGA BRCA -0.136; TCGA CESC -0.25; TCGA HNSC -0.35; TCGA LUAD -0.109; TCGA LUSC -0.26; TCGA PRAD -0.163; TCGA SARC -0.34; TCGA UCEC -0.113 |
hsa-miR-20b-5p | APBB2 | 10 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LGG; LIHC; LUAD; LUSC; PRAD | MirTarget; miRNATAP | TCGA BLCA -0.112; TCGA BRCA -0.057; TCGA CESC -0.072; TCGA HNSC -0.197; TCGA KIRC -0.053; TCGA LGG -0.066; TCGA LIHC -0.079; TCGA LUAD -0.06; TCGA LUSC -0.135; TCGA PRAD -0.061 |
hsa-miR-20b-5p | CCND1 | 11 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LIHC; LUAD; LUSC; PAAD; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.251; TCGA BRCA -0.099; TCGA CESC -0.307; TCGA HNSC -0.25; TCGA KIRC -0.399; TCGA LIHC -0.23; TCGA LUAD -0.179; TCGA LUSC -0.134; TCGA PAAD -0.153; TCGA THCA -0.255; TCGA STAD -0.181 |
hsa-miR-20b-5p | AKAP13 | 12 cancers: BLCA; BRCA; HNSC; KIRC; KIRP; LGG; LIHC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.13; TCGA BRCA -0.059; TCGA HNSC -0.078; TCGA KIRC -0.08; TCGA KIRP -0.073; TCGA LGG -0.057; TCGA LIHC -0.061; TCGA LUAD -0.089; TCGA LUSC -0.087; TCGA PRAD -0.138; TCGA THCA -0.068; TCGA UCEC -0.068 |
hsa-miR-20b-5p | DPYSL2 | 9 cancers: BLCA; BRCA; KIRC; KIRP; LGG; LUAD; LUSC; PRAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.152; TCGA BRCA -0.057; TCGA KIRC -0.109; TCGA KIRP -0.061; TCGA LGG -0.066; TCGA LUAD -0.156; TCGA LUSC -0.14; TCGA PRAD -0.179; TCGA UCEC -0.057 |
hsa-miR-20b-5p | FNDC3A | 9 cancers: BLCA; BRCA; HNSC; KIRP; LIHC; LUAD; LUSC; PRAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.088; TCGA BRCA -0.054; TCGA HNSC -0.089; TCGA KIRP -0.06; TCGA LIHC -0.141; TCGA LUAD -0.056; TCGA LUSC -0.064; TCGA PRAD -0.21; TCGA UCEC -0.063 |
hsa-miR-20b-5p | RAB11FIP5 | 9 cancers: BLCA; CESC; HNSC; LUAD; LUSC; PAAD; SARC; THCA; UCEC | MirTarget | TCGA BLCA -0.103; TCGA CESC -0.091; TCGA HNSC -0.18; TCGA LUAD -0.073; TCGA LUSC -0.071; TCGA PAAD -0.132; TCGA SARC -0.139; TCGA THCA -0.078; TCGA UCEC -0.111 |
hsa-miR-20b-5p | PDCD1LG2 | 9 cancers: BLCA; CESC; HNSC; KIRC; LUSC; PRAD; SARC; THCA; UCEC | MirTarget | TCGA BLCA -0.327; TCGA CESC -0.279; TCGA HNSC -0.102; TCGA KIRC -0.124; TCGA LUSC -0.097; TCGA PRAD -0.191; TCGA SARC -0.125; TCGA THCA -0.255; TCGA UCEC -0.107 |
hsa-miR-20b-5p | CTSK | 11 cancers: BLCA; BRCA; CESC; HNSC; LUAD; LUSC; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.308; TCGA BRCA -0.078; TCGA CESC -0.166; TCGA HNSC -0.204; TCGA LUAD -0.06; TCGA LUSC -0.219; TCGA PRAD -0.111; TCGA SARC -0.291; TCGA THCA -0.128; TCGA STAD -0.091; TCGA UCEC -0.115 |
hsa-miR-20b-5p | ZBTB47 | 9 cancers: BLCA; BRCA; CESC; LGG; LUAD; LUSC; PRAD; SARC; UCEC | MirTarget | TCGA BLCA -0.076; TCGA BRCA -0.067; TCGA CESC -0.074; TCGA LGG -0.098; TCGA LUAD -0.062; TCGA LUSC -0.084; TCGA PRAD -0.097; TCGA SARC -0.059; TCGA UCEC -0.068 |
hsa-miR-20b-5p | PDGFRA | 10 cancers: BLCA; BRCA; CESC; HNSC; LGG; LUAD; LUSC; PRAD; SARC; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.2; TCGA BRCA -0.121; TCGA CESC -0.131; TCGA HNSC -0.093; TCGA LGG -0.281; TCGA LUAD -0.126; TCGA LUSC -0.165; TCGA PRAD -0.246; TCGA SARC -0.24; TCGA UCEC -0.105 |
hsa-miR-20b-5p | THBS2 | 12 cancers: BLCA; CESC; ESCA; HNSC; KIRC; KIRP; LUSC; PAAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.345; TCGA CESC -0.2; TCGA ESCA -0.17; TCGA HNSC -0.387; TCGA KIRC -0.271; TCGA KIRP -0.223; TCGA LUSC -0.286; TCGA PAAD -0.286; TCGA SARC -0.188; TCGA THCA -0.253; TCGA STAD -0.152; TCGA UCEC -0.072 |
hsa-miR-20b-5p | APCDD1 | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; PRAD; SARC; THCA | MirTarget | TCGA BLCA -0.08; TCGA BRCA -0.124; TCGA CESC -0.101; TCGA COAD -0.169; TCGA HNSC -0.22; TCGA LUAD -0.071; TCGA LUSC -0.158; TCGA PRAD -0.199; TCGA SARC -0.125; TCGA THCA -0.06 |
hsa-miR-20b-5p | SCN2B | 9 cancers: BLCA; BRCA; CESC; HNSC; LUAD; LUSC; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.345; TCGA BRCA -0.29; TCGA CESC -0.319; TCGA HNSC -0.391; TCGA LUAD -0.252; TCGA LUSC -0.277; TCGA PRAD -0.347; TCGA THCA -0.147; TCGA UCEC -0.297 |
hsa-miR-20b-5p | TBC1D2 | 12 cancers: BLCA; CESC; HNSC; KIRC; KIRP; LGG; LIHC; LUAD; LUSC; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.144; TCGA CESC -0.154; TCGA HNSC -0.113; TCGA KIRC -0.127; TCGA KIRP -0.14; TCGA LGG -0.058; TCGA LIHC -0.067; TCGA LUAD -0.067; TCGA LUSC -0.081; TCGA PRAD -0.119; TCGA THCA -0.407; TCGA STAD -0.153 |
hsa-miR-20b-5p | S1PR1 | 9 cancers: BLCA; BRCA; CESC; KIRC; LIHC; LUAD; LUSC; PRAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.135; TCGA BRCA -0.075; TCGA CESC -0.125; TCGA KIRC -0.172; TCGA LIHC -0.105; TCGA LUAD -0.103; TCGA LUSC -0.13; TCGA PRAD -0.137; TCGA UCEC -0.096 |
hsa-miR-20b-5p | PLEKHO2 | 9 cancers: BLCA; CESC; ESCA; HNSC; KIRC; LGG; LUSC; PRAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.124; TCGA CESC -0.093; TCGA ESCA -0.065; TCGA HNSC -0.052; TCGA KIRC -0.187; TCGA LGG -0.059; TCGA LUSC -0.128; TCGA PRAD -0.123; TCGA UCEC -0.064 |
hsa-miR-20b-5p | REEP3 | 9 cancers: BLCA; CESC; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; THCA | MirTarget; miRNATAP | TCGA BLCA -0.052; TCGA CESC -0.059; TCGA ESCA -0.1; TCGA HNSC -0.159; TCGA LIHC -0.05; TCGA LUAD -0.106; TCGA LUSC -0.091; TCGA PRAD -0.057; TCGA THCA -0.078 |
hsa-miR-20b-5p | TRAF3IP2 | 12 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; SARC; STAD | mirMAP | TCGA BLCA -0.118; TCGA CESC -0.098; TCGA COAD -0.069; TCGA ESCA -0.086; TCGA HNSC -0.125; TCGA KIRC -0.192; TCGA KIRP -0.139; TCGA LIHC -0.103; TCGA LUAD -0.056; TCGA LUSC -0.099; TCGA SARC -0.064; TCGA STAD -0.09 |
hsa-miR-20b-5p | FHL2 | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRP; LUSC; PAAD; PRAD; STAD | mirMAP | TCGA BLCA -0.107; TCGA BRCA -0.09; TCGA CESC -0.157; TCGA ESCA -0.18; TCGA HNSC -0.198; TCGA KIRP -0.226; TCGA LUSC -0.257; TCGA PAAD -0.177; TCGA PRAD -0.214; TCGA STAD -0.146 |
hsa-miR-20b-5p | ANXA11 | 10 cancers: BLCA; ESCA; KIRP; LIHC; LUAD; LUSC; PAAD; SARC; THCA; STAD | mirMAP | TCGA BLCA -0.058; TCGA ESCA -0.1; TCGA KIRP -0.054; TCGA LIHC -0.079; TCGA LUAD -0.097; TCGA LUSC -0.066; TCGA PAAD -0.139; TCGA SARC -0.066; TCGA THCA -0.092; TCGA STAD -0.08 |
hsa-miR-20b-5p | CXorf36 | 9 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LUAD; LUSC; PRAD; UCEC | mirMAP | TCGA BLCA -0.084; TCGA BRCA -0.099; TCGA CESC -0.103; TCGA HNSC -0.076; TCGA KIRC -0.391; TCGA LUAD -0.089; TCGA LUSC -0.149; TCGA PRAD -0.142; TCGA UCEC -0.123 |
hsa-miR-20b-5p | MMP16 | 10 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LGG; LUAD; LUSC; PRAD; THCA | mirMAP | TCGA BLCA -0.223; TCGA BRCA -0.123; TCGA CESC -0.224; TCGA HNSC -0.306; TCGA KIRC -0.484; TCGA LGG -0.119; TCGA LUAD -0.132; TCGA LUSC -0.165; TCGA PRAD -0.388; TCGA THCA -0.615 |
hsa-miR-20b-5p | BACH1 | 9 cancers: BLCA; CESC; ESCA; HNSC; LGG; LUSC; PRAD; THCA; UCEC | mirMAP | TCGA BLCA -0.084; TCGA CESC -0.12; TCGA ESCA -0.052; TCGA HNSC -0.098; TCGA LGG -0.061; TCGA LUSC -0.098; TCGA PRAD -0.057; TCGA THCA -0.11; TCGA UCEC -0.078 |
hsa-miR-20b-5p | KLF13 | 9 cancers: BLCA; BRCA; ESCA; HNSC; LUAD; LUSC; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.13; TCGA BRCA -0.056; TCGA ESCA -0.061; TCGA HNSC -0.09; TCGA LUAD -0.087; TCGA LUSC -0.179; TCGA PRAD -0.197; TCGA STAD -0.057; TCGA UCEC -0.089 |
hsa-miR-20b-5p | COL8A2 | 13 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LGG; LUAD; LUSC; PAAD; PRAD; SARC; THCA; UCEC | mirMAP | TCGA BLCA -0.294; TCGA BRCA -0.069; TCGA CESC -0.139; TCGA HNSC -0.117; TCGA KIRC -0.125; TCGA LGG -0.14; TCGA LUAD -0.091; TCGA LUSC -0.216; TCGA PAAD -0.236; TCGA PRAD -0.207; TCGA SARC -0.137; TCGA THCA -0.551; TCGA UCEC -0.109 |
hsa-miR-20b-5p | PTRF | 13 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PRAD; SARC; THCA; UCEC | mirMAP | TCGA BLCA -0.37; TCGA BRCA -0.102; TCGA CESC -0.256; TCGA ESCA -0.138; TCGA HNSC -0.238; TCGA KIRC -0.261; TCGA KIRP -0.096; TCGA LUAD -0.13; TCGA LUSC -0.197; TCGA PRAD -0.217; TCGA SARC -0.05; TCGA THCA -0.127; TCGA UCEC -0.143 |
hsa-miR-20b-5p | MXRA7 | 9 cancers: BLCA; BRCA; CESC; HNSC; LUAD; LUSC; PRAD; SARC; UCEC | mirMAP | TCGA BLCA -0.185; TCGA BRCA -0.083; TCGA CESC -0.113; TCGA HNSC -0.101; TCGA LUAD -0.135; TCGA LUSC -0.051; TCGA PRAD -0.183; TCGA SARC -0.06; TCGA UCEC -0.105 |
hsa-miR-20b-5p | THSD4 | 9 cancers: BLCA; BRCA; CESC; HNSC; LIHC; LUAD; LUSC; PRAD; THCA | mirMAP | TCGA BLCA -0.202; TCGA BRCA -0.159; TCGA CESC -0.182; TCGA HNSC -0.229; TCGA LIHC -0.13; TCGA LUAD -0.101; TCGA LUSC -0.188; TCGA PRAD -0.226; TCGA THCA -0.122 |
hsa-miR-20b-5p | MYO1C | 9 cancers: BLCA; HNSC; KIRC; LIHC; LUAD; LUSC; PAAD; THCA; UCEC | mirMAP | TCGA BLCA -0.068; TCGA HNSC -0.142; TCGA KIRC -0.098; TCGA LIHC -0.058; TCGA LUAD -0.087; TCGA LUSC -0.118; TCGA PAAD -0.064; TCGA THCA -0.113; TCGA UCEC -0.063 |
hsa-miR-20b-5p | SPRED1 | 9 cancers: BLCA; CESC; HNSC; KIRC; LUAD; LUSC; PAAD; PRAD; UCEC | miRNATAP | TCGA BLCA -0.153; TCGA CESC -0.062; TCGA HNSC -0.056; TCGA KIRC -0.119; TCGA LUAD -0.153; TCGA LUSC -0.121; TCGA PAAD -0.102; TCGA PRAD -0.172; TCGA UCEC -0.063 |
hsa-miR-20b-5p | BNC2 | 9 cancers: BLCA; BRCA; CESC; HNSC; LUAD; LUSC; PRAD; SARC; UCEC | miRNATAP | TCGA BLCA -0.379; TCGA BRCA -0.07; TCGA CESC -0.141; TCGA HNSC -0.128; TCGA LUAD -0.087; TCGA LUSC -0.226; TCGA PRAD -0.306; TCGA SARC -0.078; TCGA UCEC -0.278 |
hsa-miR-20b-5p | PTEN | 9 cancers: BLCA; CESC; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; UCEC | miRNATAP | TCGA BLCA -0.088; TCGA CESC -0.079; TCGA ESCA -0.054; TCGA HNSC -0.1; TCGA LIHC -0.06; TCGA LUAD -0.096; TCGA LUSC -0.094; TCGA PRAD -0.094; TCGA UCEC -0.08 |
hsa-miR-20b-5p | TGFBR2 | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; UCEC | miRNATAP | TCGA BLCA -0.106; TCGA BRCA -0.101; TCGA CESC -0.08; TCGA ESCA -0.121; TCGA HNSC -0.203; TCGA LIHC -0.08; TCGA LUAD -0.192; TCGA LUSC -0.226; TCGA PRAD -0.217; TCGA UCEC -0.126 |
hsa-miR-20b-5p | RUNX1 | 10 cancers: BLCA; KIRC; KIRP; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD | miRNATAP | TCGA BLCA -0.053; TCGA KIRC -0.35; TCGA KIRP -0.214; TCGA LUAD -0.078; TCGA LUSC -0.094; TCGA PAAD -0.118; TCGA PRAD -0.102; TCGA SARC -0.09; TCGA THCA -0.544; TCGA STAD -0.079 |
hsa-miR-20b-5p | CCND2 | 11 cancers: BLCA; BRCA; CESC; HNSC; KIRC; KIRP; LGG; LUAD; PRAD; THCA; UCEC | miRNATAP | TCGA BLCA -0.296; TCGA BRCA -0.105; TCGA CESC -0.295; TCGA HNSC -0.096; TCGA KIRC -0.26; TCGA KIRP -0.325; TCGA LGG -0.103; TCGA LUAD -0.097; TCGA PRAD -0.229; TCGA THCA -0.158; TCGA UCEC -0.186 |
hsa-miR-20b-5p | SMAD7 | 9 cancers: BLCA; CESC; HNSC; LGG; LUAD; LUSC; PRAD; SARC; UCEC | miRNATAP | TCGA BLCA -0.111; TCGA CESC -0.097; TCGA HNSC -0.053; TCGA LGG -0.086; TCGA LUAD -0.098; TCGA LUSC -0.171; TCGA PRAD -0.138; TCGA SARC -0.08; TCGA UCEC -0.073 |
hsa-miR-20b-5p | PTGER3 | 9 cancers: BLCA; BRCA; CESC; HNSC; LGG; LUAD; LUSC; PRAD; UCEC | miRNATAP | TCGA BLCA -0.234; TCGA BRCA -0.194; TCGA CESC -0.264; TCGA HNSC -0.183; TCGA LGG -0.201; TCGA LUAD -0.193; TCGA LUSC -0.18; TCGA PRAD -0.326; TCGA UCEC -0.313 |
hsa-miR-20b-5p | HEG1 | 9 cancers: BLCA; BRCA; CESC; HNSC; LUAD; LUSC; PRAD; THCA; UCEC | miRNATAP | TCGA BLCA -0.239; TCGA BRCA -0.056; TCGA CESC -0.251; TCGA HNSC -0.149; TCGA LUAD -0.12; TCGA LUSC -0.199; TCGA PRAD -0.24; TCGA THCA -0.058; TCGA UCEC -0.073 |
hsa-miR-20b-5p | OTUD1 | 9 cancers: BLCA; CESC; HNSC; LGG; LUAD; PRAD; SARC; THCA; UCEC | miRNATAP | TCGA BLCA -0.092; TCGA CESC -0.159; TCGA HNSC -0.053; TCGA LGG -0.143; TCGA LUAD -0.05; TCGA PRAD -0.073; TCGA SARC -0.069; TCGA THCA -0.061; TCGA UCEC -0.148 |
hsa-miR-20b-5p | CSF1 | 9 cancers: BLCA; CESC; KIRC; LGG; LUAD; LUSC; PRAD; THCA; UCEC | miRNATAP | TCGA BLCA -0.212; TCGA CESC -0.126; TCGA KIRC -0.144; TCGA LGG -0.134; TCGA LUAD -0.065; TCGA LUSC -0.148; TCGA PRAD -0.192; TCGA THCA -0.134; TCGA UCEC -0.075 |
hsa-miR-20b-5p | SKI | 10 cancers: BLCA; CESC; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PRAD; UCEC | miRNATAP | TCGA BLCA -0.065; TCGA CESC -0.062; TCGA ESCA -0.064; TCGA HNSC -0.061; TCGA KIRC -0.158; TCGA KIRP -0.066; TCGA LUAD -0.073; TCGA LUSC -0.123; TCGA PRAD -0.051; TCGA UCEC -0.051 |
hsa-miR-20b-5p | TIMP2 | 10 cancers: BLCA; BRCA; CESC; HNSC; LUAD; LUSC; PRAD; SARC; THCA; UCEC | miRNATAP | TCGA BLCA -0.275; TCGA BRCA -0.062; TCGA CESC -0.107; TCGA HNSC -0.219; TCGA LUAD -0.076; TCGA LUSC -0.212; TCGA PRAD -0.193; TCGA SARC -0.057; TCGA THCA -0.06; TCGA UCEC -0.131 |
hsa-miR-20b-5p | CALD1 | 10 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LIHC; LUAD; LUSC; PRAD; UCEC | miRNATAP | TCGA BLCA -0.337; TCGA BRCA -0.066; TCGA CESC -0.242; TCGA HNSC -0.215; TCGA KIRC -0.092; TCGA LIHC -0.102; TCGA LUAD -0.075; TCGA LUSC -0.205; TCGA PRAD -0.257; TCGA UCEC -0.135 |
hsa-miR-20b-5p | MYO1D | 9 cancers: BLCA; COAD; ESCA; HNSC; LUSC; PAAD; SARC; THCA; STAD | miRNATAP | TCGA BLCA -0.072; TCGA COAD -0.067; TCGA ESCA -0.098; TCGA HNSC -0.122; TCGA LUSC -0.078; TCGA PAAD -0.119; TCGA SARC -0.129; TCGA THCA -0.181; TCGA STAD -0.115 |
hsa-miR-20b-5p | NBL1 | 14 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.202; TCGA BRCA -0.108; TCGA CESC -0.051; TCGA ESCA -0.182; TCGA HNSC -0.119; TCGA KIRC -0.247; TCGA KIRP -0.403; TCGA LUAD -0.063; TCGA LUSC -0.189; TCGA PAAD -0.226; TCGA SARC -0.226; TCGA THCA -0.215; TCGA STAD -0.142; TCGA UCEC -0.135 |
hsa-miR-20b-5p | TRIP10 | 10 cancers: BLCA; CESC; ESCA; KIRC; KIRP; LUSC; PAAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.051; TCGA CESC -0.066; TCGA ESCA -0.145; TCGA KIRC -0.16; TCGA KIRP -0.116; TCGA LUSC -0.064; TCGA PAAD -0.225; TCGA THCA -0.102; TCGA STAD -0.082; TCGA UCEC -0.054 |
hsa-miR-20b-5p | FYN | 9 cancers: BLCA; CESC; KIRC; LGG; LUAD; LUSC; PRAD; THCA; UCEC | miRNATAP | TCGA BLCA -0.083; TCGA CESC -0.129; TCGA KIRC -0.201; TCGA LGG -0.082; TCGA LUAD -0.072; TCGA LUSC -0.058; TCGA PRAD -0.123; TCGA THCA -0.215; TCGA UCEC -0.065 |
hsa-miR-20b-5p | AHNAK | 12 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LUAD; LUSC; PRAD; SARC; THCA; UCEC | miRNATAP | TCGA BLCA -0.263; TCGA BRCA -0.109; TCGA CESC -0.146; TCGA ESCA -0.117; TCGA HNSC -0.197; TCGA LGG -0.118; TCGA LUAD -0.172; TCGA LUSC -0.194; TCGA PRAD -0.144; TCGA SARC -0.07; TCGA THCA -0.087; TCGA UCEC -0.147 |
hsa-miR-20b-5p | RNF128 | 9 cancers: BRCA; COAD; HNSC; LIHC; LUAD; LUSC; PAAD; THCA; STAD | MirTarget; miRNATAP | TCGA BRCA -0.215; TCGA COAD -0.131; TCGA HNSC -0.289; TCGA LIHC -0.118; TCGA LUAD -0.195; TCGA LUSC -0.164; TCGA PAAD -0.25; TCGA THCA -0.061; TCGA STAD -0.247 |
hsa-miR-20b-5p | PRRG1 | 9 cancers: BRCA; CESC; ESCA; HNSC; LIHC; LUSC; PRAD; THCA; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.066; TCGA CESC -0.29; TCGA ESCA -0.131; TCGA HNSC -0.276; TCGA LIHC -0.082; TCGA LUSC -0.146; TCGA PRAD -0.132; TCGA THCA -0.142; TCGA UCEC -0.126 |
hsa-miR-20b-5p | ARHGAP23 | 9 cancers: BRCA; CESC; HNSC; KIRC; LUAD; LUSC; PRAD; THCA; UCEC | mirMAP | TCGA BRCA -0.063; TCGA CESC -0.257; TCGA HNSC -0.065; TCGA KIRC -0.132; TCGA LUAD -0.087; TCGA LUSC -0.093; TCGA PRAD -0.304; TCGA THCA -0.299; TCGA UCEC -0.155 |
hsa-miR-20b-5p | CAMK2N1 | 9 cancers: BRCA; CESC; ESCA; HNSC; LIHC; LUSC; PAAD; PRAD; THCA | miRNATAP | TCGA BRCA -0.104; TCGA CESC -0.195; TCGA ESCA -0.224; TCGA HNSC -0.295; TCGA LIHC -0.094; TCGA LUSC -0.25; TCGA PAAD -0.243; TCGA PRAD -0.127; TCGA THCA -0.601 |
hsa-miR-20b-5p | BHLHE41 | 9 cancers: BRCA; HNSC; KIRC; KIRP; LGG; LUAD; LUSC; PRAD; THCA | miRNATAP | TCGA BRCA -0.082; TCGA HNSC -0.07; TCGA KIRC -0.49; TCGA KIRP -0.134; TCGA LGG -0.158; TCGA LUAD -0.085; TCGA LUSC -0.234; TCGA PRAD -0.269; TCGA THCA -0.221 |
hsa-miR-20b-5p | EFNB1 | 10 cancers: BRCA; CESC; ESCA; HNSC; KIRP; LUSC; PAAD; PRAD; THCA; STAD | miRNATAP | TCGA BRCA -0.059; TCGA CESC -0.184; TCGA ESCA -0.145; TCGA HNSC -0.142; TCGA KIRP -0.106; TCGA LUSC -0.158; TCGA PAAD -0.227; TCGA PRAD -0.156; TCGA THCA -0.087; TCGA STAD -0.165 |
hsa-miR-20b-5p | YPEL4 | 9 cancers: BRCA; CESC; KIRC; KIRP; LGG; LUSC; SARC; THCA; UCEC | miRNATAP | TCGA BRCA -0.132; TCGA CESC -0.178; TCGA KIRC -0.492; TCGA KIRP -0.213; TCGA LGG -0.229; TCGA LUSC -0.094; TCGA SARC -0.103; TCGA THCA -0.369; TCGA UCEC -0.175 |
hsa-miR-20b-5p | LAPTM4A | 10 cancers: CESC; COAD; HNSC; KIRP; LIHC; LUAD; PAAD; PRAD; SARC; UCEC | MirTarget; miRNATAP | TCGA CESC -0.067; TCGA COAD -0.057; TCGA HNSC -0.054; TCGA KIRP -0.075; TCGA LIHC -0.073; TCGA LUAD -0.056; TCGA PAAD -0.085; TCGA PRAD -0.079; TCGA SARC -0.068; TCGA UCEC -0.053 |
hsa-miR-20b-5p | TAOK3 | 9 cancers: CESC; ESCA; HNSC; LGG; LUAD; LUSC; PRAD; SARC; UCEC | MirTarget | TCGA CESC -0.118; TCGA ESCA -0.103; TCGA HNSC -0.096; TCGA LGG -0.094; TCGA LUAD -0.072; TCGA LUSC -0.106; TCGA PRAD -0.065; TCGA SARC -0.062; TCGA UCEC -0.089 |
hsa-miR-20b-5p | QSOX1 | 9 cancers: ESCA; HNSC; KIRC; LUAD; LUSC; PAAD; PRAD; SARC; THCA | mirMAP | TCGA ESCA -0.129; TCGA HNSC -0.146; TCGA KIRC -0.217; TCGA LUAD -0.074; TCGA LUSC -0.129; TCGA PAAD -0.186; TCGA PRAD -0.191; TCGA SARC -0.108; TCGA THCA -0.304 |
Enriched cancer pathways of putative targets