microRNA information: hsa-miR-24-1-5p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-24-1-5p | miRbase |
| Accession: | MIMAT0000079 | miRbase |
| Precursor name: | hsa-mir-24-1 | miRbase |
| Precursor accession: | MI0000080 | miRbase |
| Symbol: | MIR24-1 | HGNC |
| RefSeq ID: | NR_029496 | GenBank |
| Sequence: | UGCCUACUGAGCUGAUAUCAGU |
Reported expression in cancers: hsa-miR-24-1-5p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-24-1-5p | melanoma | deregulation | "Comparative microarray analysis of microRNA expres ......" | 23111773 | Microarray; RNA-Seq; qPCR |
Reported cancer pathway affected by hsa-miR-24-1-5p

| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-24-1-5p | bladder cancer | Apoptosis pathway | "Tumour suppressive microRNA 24 1 inhibits cancer c ......" | 24999187 | Luciferase |
Reported cancer prognosis affected by hsa-miR-24-1-5p
| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-24-1-5p | melanoma | metastasis; malignant trasformation | "Comparative microarray analysis of microRNA expres ......" | 23111773 |
Reported gene related to hsa-miR-24-1-5p
| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-miR-24-1-5p | bladder cancer | FOXM1 | "Tumour suppressive microRNA 24 1 inhibits cancer c ......" | 24999187 |
Expression profile in cancer corhorts:
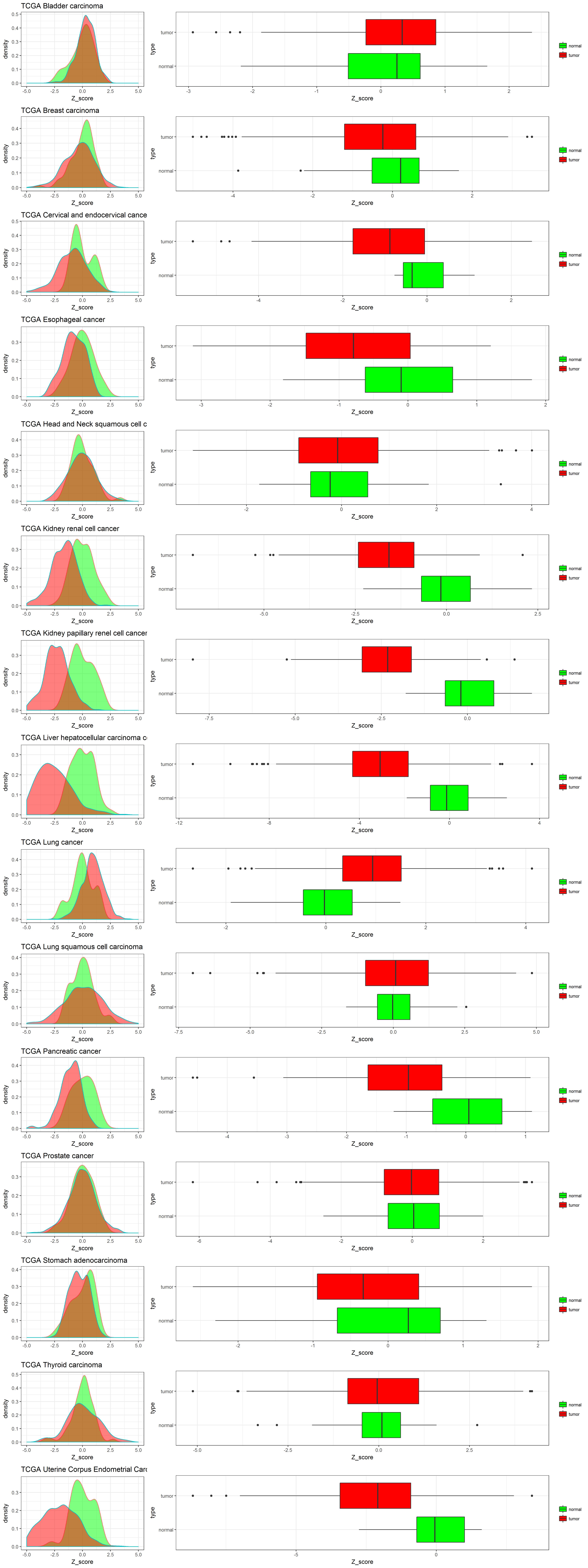
Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-24-1-5p | SRPK2 | 10 cancers: BLCA; HNSC; KIRP; LIHC; LUAD; PAAD; PRAD; SARC; THCA; UCEC | MirTarget | TCGA BLCA -0.073; TCGA HNSC -0.146; TCGA KIRP -0.107; TCGA LIHC -0.073; TCGA LUAD -0.121; TCGA PAAD -0.147; TCGA PRAD -0.235; TCGA SARC -0.075; TCGA THCA -0.101; TCGA UCEC -0.145 |
hsa-miR-24-1-5p | ANKRD28 | 13 cancers: BLCA; CESC; HNSC; KIRP; LGG; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.078; TCGA CESC -0.084; TCGA HNSC -0.068; TCGA KIRP -0.09; TCGA LGG -0.12; TCGA LUAD -0.07; TCGA LUSC -0.101; TCGA PAAD -0.111; TCGA PRAD -0.21; TCGA SARC -0.073; TCGA THCA -0.103; TCGA STAD -0.098; TCGA UCEC -0.073 |
hsa-miR-24-1-5p | HNRNPU | 9 cancers: BLCA; ESCA; LGG; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.071; TCGA ESCA -0.079; TCGA LGG -0.06; TCGA LIHC -0.128; TCGA LUAD -0.084; TCGA PAAD -0.073; TCGA PRAD -0.093; TCGA STAD -0.135; TCGA UCEC -0.146 |
hsa-miR-24-1-5p | WDR26 | 9 cancers: BLCA; HNSC; LGG; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.065; TCGA HNSC -0.06; TCGA LGG -0.115; TCGA LIHC -0.056; TCGA LUAD -0.052; TCGA PAAD -0.081; TCGA PRAD -0.141; TCGA STAD -0.084; TCGA UCEC -0.079 |
hsa-miR-24-1-5p | IL6ST | 10 cancers: BLCA; CESC; COAD; HNSC; LUAD; LUSC; PAAD; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.306; TCGA CESC -0.365; TCGA COAD -0.274; TCGA HNSC -0.592; TCGA LUAD -0.284; TCGA LUSC -0.266; TCGA PAAD -0.527; TCGA PRAD -0.539; TCGA THCA -0.271; TCGA STAD -0.089 |
hsa-miR-24-1-5p | ACTR2 | 11 cancers: BLCA; CESC; ESCA; HNSC; LUAD; LUSC; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.054; TCGA CESC -0.112; TCGA ESCA -0.078; TCGA HNSC -0.184; TCGA LUAD -0.191; TCGA LUSC -0.089; TCGA PAAD -0.15; TCGA PRAD -0.215; TCGA THCA -0.179; TCGA STAD -0.131; TCGA UCEC -0.172 |
hsa-miR-24-1-5p | DNAJC10 | 10 cancers: BLCA; CESC; HNSC; KIRP; LIHC; LUAD; LUSC; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.084; TCGA CESC -0.258; TCGA HNSC -0.098; TCGA KIRP -0.11; TCGA LIHC -0.152; TCGA LUAD -0.083; TCGA LUSC -0.119; TCGA PRAD -0.308; TCGA THCA -0.089; TCGA STAD -0.059 |
hsa-miR-24-1-5p | RFX7 | 9 cancers: BLCA; COAD; HNSC; KIRP; LGG; LUAD; PAAD; PRAD; STAD | MirTarget | TCGA BLCA -0.107; TCGA COAD -0.084; TCGA HNSC -0.127; TCGA KIRP -0.104; TCGA LGG -0.259; TCGA LUAD -0.152; TCGA PAAD -0.133; TCGA PRAD -0.184; TCGA STAD -0.15 |
hsa-miR-24-1-5p | BDP1 | 10 cancers: BLCA; HNSC; KIRC; KIRP; LGG; LUAD; LUSC; PAAD; PRAD; STAD | MirTarget | TCGA BLCA -0.106; TCGA HNSC -0.131; TCGA KIRC -0.078; TCGA KIRP -0.083; TCGA LGG -0.108; TCGA LUAD -0.122; TCGA LUSC -0.148; TCGA PAAD -0.133; TCGA PRAD -0.248; TCGA STAD -0.18 |
hsa-miR-24-1-5p | CYP27B1 | 9 cancers: CESC; HNSC; LGG; LIHC; LUSC; PRAD; SARC; THCA; UCEC | MirTarget | TCGA CESC -0.184; TCGA HNSC -0.241; TCGA LGG -0.146; TCGA LIHC -0.44; TCGA LUSC -0.215; TCGA PRAD -0.243; TCGA SARC -0.393; TCGA THCA -0.204; TCGA UCEC -0.133 |
hsa-miR-24-1-5p | TM9SF3 | 9 cancers: CESC; ESCA; HNSC; LGG; LUAD; LUSC; PRAD; SARC; UCEC | MirTarget | TCGA CESC -0.079; TCGA ESCA -0.139; TCGA HNSC -0.067; TCGA LGG -0.067; TCGA LUAD -0.061; TCGA LUSC -0.091; TCGA PRAD -0.216; TCGA SARC -0.1; TCGA UCEC -0.101 |
hsa-miR-24-1-5p | C1GALT1 | 10 cancers: CESC; HNSC; KIRP; LUAD; OV; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA CESC -0.204; TCGA HNSC -0.102; TCGA KIRP -0.088; TCGA LUAD -0.128; TCGA OV -0.121; TCGA PRAD -0.134; TCGA SARC -0.084; TCGA THCA -0.142; TCGA STAD -0.089; TCGA UCEC -0.207 |
hsa-miR-24-1-5p | HELLS | 9 cancers: COAD; KIRP; LGG; LIHC; LUAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA COAD -0.219; TCGA KIRP -0.217; TCGA LGG -0.27; TCGA LIHC -0.773; TCGA LUAD -0.281; TCGA PRAD -0.208; TCGA THCA -0.267; TCGA STAD -0.178; TCGA UCEC -0.329 |
Enriched cancer pathways of putative targets