microRNA information: hsa-miR-24-3p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-24-3p | miRbase |
| Accession: | MIMAT0000080 | miRbase |
| Precursor name: | hsa-mir-24-1 | miRbase |
| Precursor accession: | MI0000080 | miRbase |
| Symbol: | MIR24-1 | HGNC |
| RefSeq ID: | NR_029496 | GenBank |
| Sequence: | UGGCUCAGUUCAGCAGGAACAG |
Reported expression in cancers: hsa-miR-24-3p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-24-3p | acute myeloid leukemia | upregulation | "Increased expression of miR 24 is associated with ......" | 25550847 | qPCR |
| hsa-miR-24-3p | breast cancer | upregulation | "Among methods for profiling levels of miRNAs next- ......" | 22387599 | RNA-Seq; qPCR; Northern blot |
| hsa-miR-24-3p | breast cancer | upregulation | "Association between mir 24 and mir 378 in formalin ......" | 25120807 | qPCR |
| hsa-miR-24-3p | breast cancer | upregulation | "In the present study we found that the expression ......" | 26044523 | |
| hsa-miR-24-3p | colorectal cancer | deregulation | "Down regulation of miR 24 3p in colorectal cancer ......" | 25502080 | qPCR |
| hsa-miR-24-3p | gastric cancer | downregulation | "Tumor suppressor miR 24 restrains gastric cancer p ......" | 24886316 | qPCR |
| hsa-miR-24-3p | gastric cancer | upregulation | "The expression levels of miR-21 miR-24 miR-34a miR ......" | 26045155 | Reverse transcription PCR; qPCR |
| hsa-miR-24-3p | liver cancer | upregulation | "MicroRNA-24 miR-24 may be involved in neoplastic p ......" | 24800232 | |
| hsa-miR-24-3p | liver cancer | upregulation | "MiR-24-3p was previously reported to be significan ......" | 27650047 | Reverse transcription PCR |
| hsa-miR-24-3p | lung squamous cell cancer | upregulation | "Circulating miR 22 miR 24 and miR 34a as novel pre ......" | 23794259 | |
| hsa-miR-24-3p | lung squamous cell cancer | upregulation | "Recent studies have implied that aberration of miR ......" | 25725584 | |
| hsa-miR-24-3p | prostate cancer | downregulation | "miR 24 regulates CDKN1B/p27 expression in prostate ......" | 26847530 |
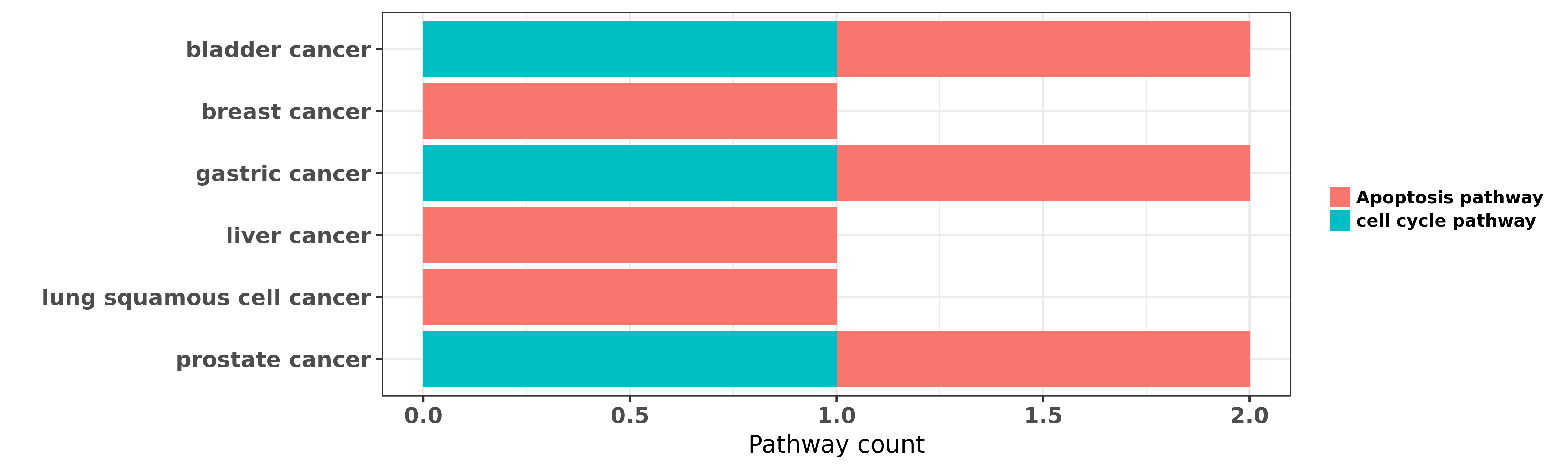
Reported cancer pathway affected by hsa-miR-24-3p
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-24-3p | bladder cancer | Apoptosis pathway | "Tumour suppressive microRNA 24 1 inhibits cancer c ......" | 24999187 | Luciferase |
| hsa-miR-24-3p | bladder cancer | cell cycle pathway; Apoptosis pathway | "Previous studied have shown dysregulation of miR-2 ......" | 26252200 | Luciferase |
| hsa-miR-24-3p | breast cancer | Apoptosis pathway | "In the present study we found that the expression ......" | 26044523 | |
| hsa-miR-24-3p | gastric cancer | cell cycle pathway; Apoptosis pathway | "Tumor suppressor miR 24 restrains gastric cancer p ......" | 24886316 | |
| hsa-miR-24-3p | gastric cancer | Apoptosis pathway | "Onco miR 24 regulates cell growth and apoptosis by ......" | 26758252 | |
| hsa-miR-24-3p | liver cancer | Apoptosis pathway | "MicroRNA-24 miR-24 may be involved in neoplastic p ......" | 24800232 | |
| hsa-miR-24-3p | lung squamous cell cancer | Apoptosis pathway | "Upregulation of miR 24 promotes cell proliferation ......" | 25725584 | |
| hsa-miR-24-3p | prostate cancer | cell cycle pathway; Apoptosis pathway | "miR 24 regulates CDKN1B/p27 expression in prostate ......" | 26847530 |
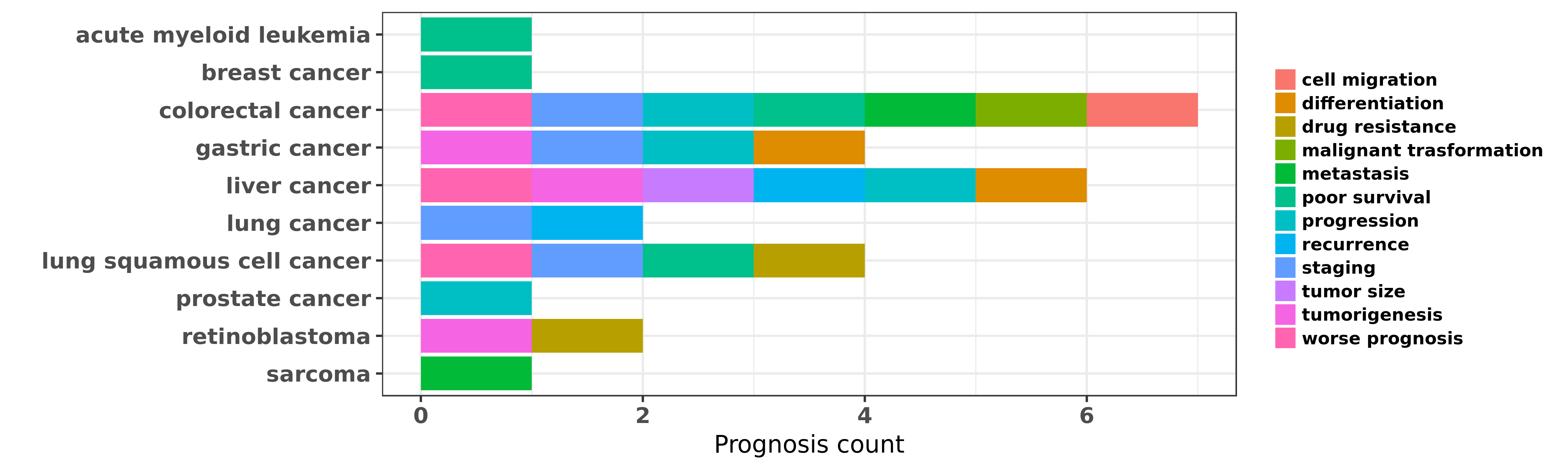
Reported cancer prognosis affected by hsa-miR-24-3p
| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-24-3p | acute myeloid leukemia | poor survival | "Increased expression of miR 24 is associated with ......" | 25550847 | |
| hsa-miR-24-3p | breast cancer | poor survival | "Preferential star strand biogenesis of pre miR 24 ......" | 22911661 | |
| hsa-miR-24-3p | colorectal cancer | malignant trasformation; staging; metastasis; worse prognosis; poor survival; cell migration; progression | "Down regulation of miR 24 3p in colorectal cancer ......" | 25502080 | |
| hsa-miR-24-3p | colorectal cancer | staging | "Analyses of online microRNA data base revealed tha ......" | 26297223 | |
| hsa-miR-24-3p | gastric cancer | progression; tumorigenesis; differentiation | "Tumor suppressor miR 24 restrains gastric cancer p ......" | 24886316 | |
| hsa-miR-24-3p | gastric cancer | staging | "Onco miR 24 regulates cell growth and apoptosis by ......" | 26758252 | |
| hsa-miR-24-3p | liver cancer | recurrence | "18 miRNAs including 6 up-regulated and 12 down-reg ......" | 22552153 | |
| hsa-miR-24-3p | liver cancer | worse prognosis | "Human hepatocellular carcinoma cell specific miRNA ......" | 23229173 | |
| hsa-miR-24-3p | liver cancer | differentiation; tumor size; worse prognosis; progression; tumorigenesis | "MicroRNA-24 miR-24 may be involved in neoplastic p ......" | 24800232 | |
| hsa-miR-24-3p | liver cancer | progression | "MiR 24 3p enhances cell growth in hepatocellular c ......" | 27650047 | Colony formation; Western blot; Luciferase |
| hsa-miR-24-3p | lung cancer | staging; recurrence | "Evaluation of dynamic change of serum miR 21 and m ......" | 22782668 | |
| hsa-miR-24-3p | lung squamous cell cancer | staging | "Circulating miR 22 miR 24 and miR 34a as novel pre ......" | 23794259 | |
| hsa-miR-24-3p | lung squamous cell cancer | drug resistance | "Mir 24 3p downregulation contributes to VP16 DDP r ......" | 25426560 | |
| hsa-miR-24-3p | lung squamous cell cancer | poor survival; worse prognosis | "Upregulation of miR 24 promotes cell proliferation ......" | 25725584 | |
| hsa-miR-24-3p | prostate cancer | progression | "miR 24 regulates CDKN1B/p27 expression in prostate ......" | 26847530 | |
| hsa-miR-24-3p | retinoblastoma | tumorigenesis; drug resistance | "In human fetal retinas adult retinas and retinobla ......" | 22336108 | |
| hsa-miR-24-3p | sarcoma | metastasis | "This potential path highlights the crucial role of ......" | 25984907 |
Reported gene related to hsa-miR-24-3p

| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-miR-24-3p | breast cancer | CDKN1B | "With the bioinformatic method we further identifie ......" | 26044523 |
| hsa-miR-24-3p | prostate cancer | CDKN1B | "The biological significance of miR-24 expression i ......" | 26847530 |
| hsa-miR-24-3p | prostate cancer | CDKN2A | "The biological significance of miR-24 expression i ......" | 26847530 |
| hsa-miR-24-3p | retinoblastoma | CDKN2A | "To study p14ARF biogenesis retinoblastoma cells we ......" | 22336108 |
| hsa-miR-24-3p | liver cancer | AFP | "Combined serum alpha-fetoprotein AFP and miR-24-3p ......" | 25129312 |
| hsa-miR-24-3p | lung squamous cell cancer | ATG4A | "Mir 24 3p downregulation contributes to VP16 DDP r ......" | 25426560 |
| hsa-miR-24-3p | gastric cancer | BCL2L11 | "Onco miR 24 regulates cell growth and apoptosis by ......" | 26758252 |
| hsa-miR-24-3p | bladder cancer | CARD10 | "Moreover the low level of miR-24 was associated wi ......" | 26252200 |
| hsa-miR-24-3p | prostate cancer | DUOXA1 | "We previously detected several miRNAs for example ......" | 22815235 |
| hsa-miR-24-3p | bladder cancer | FOXM1 | "Tumour suppressive microRNA 24 1 inhibits cancer c ......" | 24999187 |
| hsa-miR-24-3p | acute myeloid leukemia | FRTS1 | "Meanwhile among the group who obtained CR the case ......" | 25550847 |
| hsa-miR-24-3p | gastric cancer | ICOSLG | "Dual-luciferase reporter assays showed that miR-24 ......" | 23688438 |
| hsa-miR-24-3p | liver cancer | MT1M | "MiR 24 3p enhances cell growth in hepatocellular c ......" | 27650047 |
| hsa-miR-24-3p | lung squamous cell cancer | NAIF1 | "Upregulation of miR 24 promotes cell proliferation ......" | 25725584 |
| hsa-miR-24-3p | breast cancer | PRKCA | "Preferential star strand biogenesis of pre miR 24 ......" | 22911661 |
| hsa-miR-24-3p | lung squamous cell cancer | SCLC1 | "The present study found that miR-24-3p a recently ......" | 25426560 |
| hsa-miR-24-3p | breast cancer | STAR | "Preferential star strand biogenesis of pre miR 24 ......" | 22911661 |
| hsa-miR-24-3p | lung squamous cell cancer | TIMM8A | "Mir 24 3p downregulation contributes to VP16 DDP r ......" | 25426560 |
| hsa-miR-24-3p | retinoblastoma | TP53 | "p14ARF protein levels were restored without change ......" | 22336108 |
| hsa-miR-24-3p | gastric cancer | WWC2 | "Characteristic miR 24 Expression in Gastric Cancer ......" | 26045155 |
Expression profile in cancer corhorts:
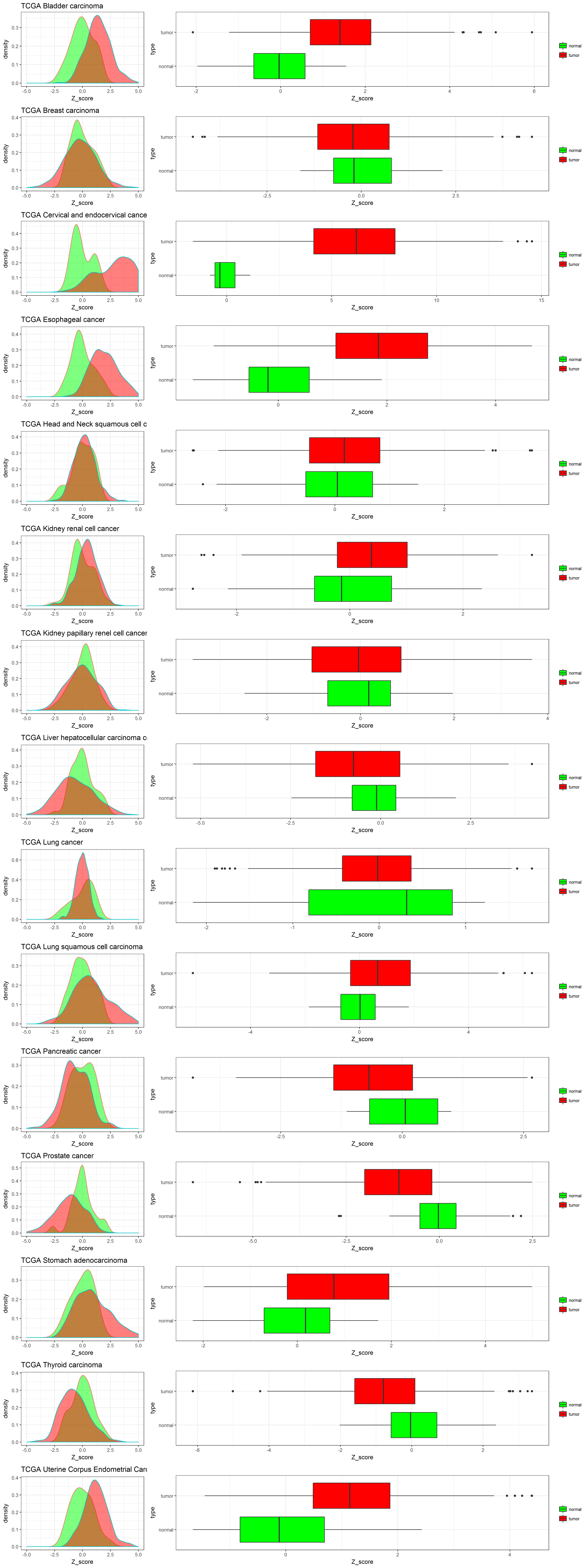
Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-24-3p | ALDH5A1 | 10 cancers: BLCA; CESC; ESCA; HNSC; KIRC; LGG; LIHC; LUAD; PAAD; PRAD | miRNAWalker2 validate | TCGA BLCA -0.18; TCGA CESC -0.248; TCGA ESCA -0.313; TCGA HNSC -0.249; TCGA KIRC -0.484; TCGA LGG -0.217; TCGA LIHC -0.433; TCGA LUAD -0.255; TCGA PAAD -0.298; TCGA PRAD -0.1 |
hsa-miR-24-3p | BEX1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LGG; LUSC; OV; PAAD; STAD | miRNAWalker2 validate | TCGA BLCA -0.721; TCGA BRCA -0.589; TCGA CESC -0.479; TCGA COAD -0.621; TCGA ESCA -0.822; TCGA HNSC -0.622; TCGA KIRC -0.7; TCGA LGG -0.348; TCGA LUSC -0.678; TCGA OV -1.182; TCGA PAAD -1.256; TCGA STAD -0.945 |
hsa-miR-24-3p | CCNG1 | 16 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PRAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.169; TCGA BRCA -0.136; TCGA CESC -0.168; TCGA ESCA -0.459; TCGA HNSC -0.163; TCGA KIRC -0.451; TCGA KIRP -0.204; TCGA LIHC -0.345; TCGA LUAD -0.138; TCGA LUSC -0.197; TCGA OV -0.21; TCGA PRAD -0.158; TCGA SARC -0.344; TCGA THCA -0.111; TCGA STAD -0.362; TCGA UCEC -0.232 |
hsa-miR-24-3p | CDKN1B | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LIHC; LUAD; LUSC; SARC; STAD | miRNAWalker2 validate; miRNATAP | TCGA BLCA -0.156; TCGA BRCA -0.13; TCGA CESC -0.228; TCGA ESCA -0.146; TCGA HNSC -0.096; TCGA LGG -0.114; TCGA LIHC -0.126; TCGA LUAD -0.11; TCGA LUSC -0.145; TCGA SARC -0.319; TCGA STAD -0.322 |
hsa-miR-24-3p | CIRBP | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LUSC; PAAD; PRAD; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.281; TCGA BRCA -0.444; TCGA CESC -0.178; TCGA ESCA -0.174; TCGA HNSC -0.351; TCGA LGG -0.147; TCGA LUSC -0.171; TCGA PAAD -0.369; TCGA PRAD -0.331; TCGA STAD -0.307; TCGA UCEC -0.208 |
hsa-miR-24-3p | DDX5 | 9 cancers: BLCA; ESCA; HNSC; LIHC; LUAD; LUSC; PAAD; PRAD; STAD | miRNAWalker2 validate | TCGA BLCA -0.072; TCGA ESCA -0.089; TCGA HNSC -0.086; TCGA LIHC -0.098; TCGA LUAD -0.107; TCGA LUSC -0.105; TCGA PAAD -0.121; TCGA PRAD -0.198; TCGA STAD -0.119 |
hsa-miR-24-3p | KLHDC3 | 10 cancers: BLCA; CESC; HNSC; LGG; LIHC; LUAD; OV; PAAD; PRAD; THCA | miRNAWalker2 validate; MirTarget | TCGA BLCA -0.074; TCGA CESC -0.087; TCGA HNSC -0.174; TCGA LGG -0.112; TCGA LIHC -0.243; TCGA LUAD -0.115; TCGA OV -0.191; TCGA PAAD -0.202; TCGA PRAD -0.256; TCGA THCA -0.101 |
hsa-miR-24-3p | KLHL23 | 9 cancers: BLCA; COAD; ESCA; LGG; LUAD; OV; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.321; TCGA COAD -0.198; TCGA ESCA -0.728; TCGA LGG -0.243; TCGA LUAD -0.179; TCGA OV -0.164; TCGA SARC -0.798; TCGA STAD -0.242; TCGA UCEC -0.339 |
hsa-miR-24-3p | MAGI1 | 9 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LGG; SARC; STAD; UCEC | miRNAWalker2 validate; MirTarget; miRNATAP | TCGA BLCA -0.433; TCGA BRCA -0.195; TCGA CESC -0.364; TCGA ESCA -0.748; TCGA KIRC -0.308; TCGA LGG -0.067; TCGA SARC -0.271; TCGA STAD -0.145; TCGA UCEC -0.32 |
hsa-miR-24-3p | PHOSPHO2 | 12 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LGG; OV; PRAD; SARC; STAD | miRNAWalker2 validate | TCGA BLCA -0.19; TCGA BRCA -0.416; TCGA CESC -0.177; TCGA ESCA -0.328; TCGA HNSC -0.137; TCGA KIRC -0.477; TCGA KIRP -0.213; TCGA LGG -0.09; TCGA OV -0.139; TCGA PRAD -0.513; TCGA SARC -0.279; TCGA STAD -0.24 |
hsa-miR-24-3p | SESN1 | 11 cancers: BLCA; BRCA; ESCA; HNSC; KIRC; LUSC; OV; PAAD; SARC; STAD; UCEC | miRNAWalker2 validate; MirTarget; miRNATAP | TCGA BLCA -0.15; TCGA BRCA -0.224; TCGA ESCA -0.441; TCGA HNSC -0.215; TCGA KIRC -0.189; TCGA LUSC -0.265; TCGA OV -0.187; TCGA PAAD -0.658; TCGA SARC -0.323; TCGA STAD -0.303; TCGA UCEC -0.327 |
hsa-miR-24-3p | STRADB | 14 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRP; LGG; LIHC; LUAD; LUSC; OV; PAAD; PRAD; STAD | miRNAWalker2 validate; MirTarget; miRNATAP | TCGA BLCA -0.101; TCGA BRCA -0.205; TCGA CESC -0.227; TCGA ESCA -0.174; TCGA HNSC -0.248; TCGA KIRP -0.214; TCGA LGG -0.105; TCGA LIHC -0.123; TCGA LUAD -0.204; TCGA LUSC -0.229; TCGA OV -0.254; TCGA PAAD -0.258; TCGA PRAD -0.34; TCGA STAD -0.249 |
hsa-miR-24-3p | TCEA3 | 9 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LIHC; PRAD; THCA; STAD | miRNAWalker2 validate | TCGA BLCA -0.429; TCGA BRCA -0.426; TCGA CESC -0.383; TCGA ESCA -0.604; TCGA HNSC -0.279; TCGA LIHC -0.344; TCGA PRAD -0.477; TCGA THCA -0.088; TCGA STAD -0.344 |
hsa-miR-24-3p | WRB | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.198; TCGA BRCA -0.082; TCGA CESC -0.15; TCGA ESCA -0.141; TCGA HNSC -0.164; TCGA OV -0.115; TCGA PRAD -0.119; TCGA SARC -0.161; TCGA STAD -0.087; TCGA UCEC -0.171 |
hsa-miR-24-3p | ANKRD44 | 9 cancers: BLCA; CESC; COAD; ESCA; LUAD; LUSC; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.461; TCGA CESC -0.259; TCGA COAD -0.82; TCGA ESCA -0.764; TCGA LUAD -0.258; TCGA LUSC -0.642; TCGA PAAD -0.447; TCGA STAD -0.49; TCGA UCEC -0.332 |
hsa-miR-24-3p | ATP1B2 | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; LUSC; OV; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.78; TCGA BRCA -0.167; TCGA CESC -0.505; TCGA COAD -0.565; TCGA ESCA -0.591; TCGA LUSC -0.374; TCGA OV -0.554; TCGA PAAD -0.641; TCGA STAD -0.989; TCGA UCEC -0.22 |
hsa-miR-24-3p | CPNE5 | 9 cancers: BLCA; CESC; COAD; ESCA; HNSC; LGG; LUSC; PAAD; THCA | MirTarget | TCGA BLCA -0.283; TCGA CESC -0.234; TCGA COAD -0.528; TCGA ESCA -0.667; TCGA HNSC -0.283; TCGA LGG -0.345; TCGA LUSC -0.609; TCGA PAAD -0.307; TCGA THCA -0.239 |
hsa-miR-24-3p | NUP210 | 9 cancers: BLCA; BRCA; CESC; HNSC; KIRC; LUAD; PRAD; SARC; THCA | MirTarget | TCGA BLCA -0.4; TCGA BRCA -0.173; TCGA CESC -0.147; TCGA HNSC -0.479; TCGA KIRC -0.205; TCGA LUAD -0.253; TCGA PRAD -0.334; TCGA SARC -0.349; TCGA THCA -0.124 |
hsa-miR-24-3p | TPSAB1 | 9 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LUSC; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.629; TCGA BRCA -0.607; TCGA COAD -0.912; TCGA ESCA -0.372; TCGA HNSC -0.232; TCGA LUSC -0.569; TCGA PAAD -0.51; TCGA STAD -0.492; TCGA UCEC -0.669 |
hsa-miR-24-3p | EXOG | 10 cancers: BLCA; CESC; ESCA; HNSC; LGG; LUAD; LUSC; PAAD; PRAD; STAD | MirTarget | TCGA BLCA -0.095; TCGA CESC -0.143; TCGA ESCA -0.205; TCGA HNSC -0.187; TCGA LGG -0.098; TCGA LUAD -0.105; TCGA LUSC -0.128; TCGA PAAD -0.251; TCGA PRAD -0.054; TCGA STAD -0.164 |
hsa-miR-24-3p | SCAMP5 | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; LGG; LUSC; PAAD; PRAD; THCA | MirTarget; miRNATAP | TCGA BLCA -0.283; TCGA BRCA -0.148; TCGA CESC -0.421; TCGA ESCA -0.667; TCGA HNSC -0.227; TCGA KIRC -0.351; TCGA LGG -0.218; TCGA LUSC -0.2; TCGA PAAD -0.752; TCGA PRAD -0.227; TCGA THCA -0.291 |
hsa-miR-24-3p | INSIG1 | 9 cancers: BLCA; CESC; ESCA; HNSC; KIRC; LGG; LIHC; LUAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.269; TCGA CESC -0.191; TCGA ESCA -0.269; TCGA HNSC -0.123; TCGA KIRC -0.214; TCGA LGG -0.134; TCGA LIHC -0.689; TCGA LUAD -0.147; TCGA UCEC -0.179 |
hsa-miR-24-3p | KLHDC7A | 9 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LUAD; LUSC; OV; THCA | MirTarget | TCGA BLCA -1.026; TCGA BRCA -1.207; TCGA CESC -1.64; TCGA ESCA -2.211; TCGA KIRC -0.459; TCGA LUAD -0.378; TCGA LUSC -0.849; TCGA OV -0.647; TCGA THCA -0.404 |
hsa-miR-24-3p | SFXN5 | 12 cancers: BLCA; BRCA; CESC; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; PAAD; PRAD; THCA | MirTarget; miRNATAP | TCGA BLCA -0.1; TCGA BRCA -0.399; TCGA CESC -0.137; TCGA HNSC -0.279; TCGA KIRC -0.262; TCGA KIRP -0.183; TCGA LIHC -0.301; TCGA LUAD -0.246; TCGA LUSC -0.226; TCGA PAAD -0.248; TCGA PRAD -0.16; TCGA THCA -0.147 |
hsa-miR-24-3p | EDA2R | 11 cancers: BLCA; BRCA; COAD; ESCA; LGG; LUSC; PAAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.427; TCGA BRCA -0.211; TCGA COAD -0.583; TCGA ESCA -0.41; TCGA LGG -0.226; TCGA LUSC -0.489; TCGA PAAD -0.391; TCGA SARC -0.533; TCGA THCA -0.158; TCGA STAD -0.276; TCGA UCEC -0.379 |
hsa-miR-24-3p | SLC19A2 | 10 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LGG; LIHC; LUSC; OV; PRAD | MirTarget; miRNATAP | TCGA BLCA -0.272; TCGA BRCA -0.363; TCGA CESC -0.119; TCGA ESCA -0.377; TCGA KIRC -0.773; TCGA LGG -0.162; TCGA LIHC -0.229; TCGA LUSC -0.169; TCGA OV -0.233; TCGA PRAD -0.195 |
hsa-miR-24-3p | STEAP2 | 10 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LGG; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.352; TCGA BRCA -0.386; TCGA CESC -0.337; TCGA ESCA -0.538; TCGA KIRC -0.53; TCGA LGG -0.139; TCGA PRAD -0.346; TCGA THCA -0.249; TCGA STAD -0.454; TCGA UCEC -0.432 |
hsa-miR-24-3p | CECR6 | 9 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LUAD; PAAD; PRAD | MirTarget | TCGA BLCA -0.159; TCGA BRCA -0.101; TCGA CESC -0.295; TCGA ESCA -0.266; TCGA HNSC -0.336; TCGA LGG -0.32; TCGA LUAD -0.317; TCGA PAAD -0.781; TCGA PRAD -0.575 |
hsa-miR-24-3p | SNN | 9 cancers: BLCA; COAD; KIRC; LUAD; LUSC; OV; SARC; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.218; TCGA COAD -0.247; TCGA KIRC -0.187; TCGA LUAD -0.137; TCGA LUSC -0.202; TCGA OV -0.254; TCGA SARC -0.336; TCGA THCA -0.169; TCGA STAD -0.14 |
hsa-miR-24-3p | TNS1 | 9 cancers: BLCA; CESC; COAD; ESCA; LUSC; PAAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.935; TCGA CESC -0.462; TCGA COAD -1.02; TCGA ESCA -0.71; TCGA LUSC -0.561; TCGA PAAD -0.328; TCGA SARC -0.729; TCGA STAD -0.932; TCGA UCEC -0.35 |
hsa-miR-24-3p | PATZ1 | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LUAD; PAAD; PRAD; SARC | MirTarget | TCGA BLCA -0.283; TCGA BRCA -0.32; TCGA CESC -0.357; TCGA ESCA -0.25; TCGA HNSC -0.173; TCGA LGG -0.2; TCGA LUAD -0.097; TCGA PAAD -0.194; TCGA PRAD -0.216; TCGA SARC -0.194 |
hsa-miR-24-3p | DLG2 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUSC; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -1.112; TCGA BRCA -0.252; TCGA CESC -0.546; TCGA COAD -1.09; TCGA ESCA -1.037; TCGA HNSC -0.37; TCGA KIRC -0.679; TCGA LUSC -0.84; TCGA PAAD -1.03; TCGA STAD -0.962; TCGA UCEC -0.742 |
hsa-miR-24-3p | GDPD1 | 10 cancers: BLCA; BRCA; CESC; ESCA; KIRC; KIRP; LGG; LUAD; PRAD; STAD | MirTarget | TCGA BLCA -0.174; TCGA BRCA -0.211; TCGA CESC -0.464; TCGA ESCA -0.838; TCGA KIRC -0.346; TCGA KIRP -0.174; TCGA LGG -0.226; TCGA LUAD -0.191; TCGA PRAD -0.34; TCGA STAD -0.182 |
hsa-miR-24-3p | GBA2 | 12 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; LIHC; LUAD; LUSC; PRAD; SARC; THCA | MirTarget | TCGA BLCA -0.112; TCGA BRCA -0.128; TCGA CESC -0.126; TCGA ESCA -0.47; TCGA HNSC -0.316; TCGA KIRC -0.266; TCGA LIHC -0.152; TCGA LUAD -0.212; TCGA LUSC -0.251; TCGA PRAD -0.245; TCGA SARC -0.106; TCGA THCA -0.17 |
hsa-miR-24-3p | ATP6V0E2 | 13 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; LGG; LIHC; LUAD; PAAD; PRAD; SARC; THCA | MirTarget | TCGA BLCA -0.465; TCGA BRCA -0.291; TCGA CESC -0.437; TCGA ESCA -0.475; TCGA HNSC -0.467; TCGA KIRC -0.492; TCGA LGG -0.139; TCGA LIHC -0.642; TCGA LUAD -0.193; TCGA PAAD -0.474; TCGA PRAD -0.431; TCGA SARC -0.299; TCGA THCA -0.251 |
hsa-miR-24-3p | RSPO3 | 9 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUSC; PAAD; SARC; STAD | MirTarget | TCGA BLCA -0.929; TCGA CESC -0.379; TCGA COAD -0.614; TCGA ESCA -0.66; TCGA HNSC -0.492; TCGA LUSC -0.83; TCGA PAAD -0.809; TCGA SARC -1.205; TCGA STAD -0.85 |
hsa-miR-24-3p | GPR155 | 10 cancers: BLCA; ESCA; HNSC; KIRC; LGG; LIHC; PAAD; SARC; THCA; STAD | MirTarget | TCGA BLCA -0.225; TCGA ESCA -0.648; TCGA HNSC -0.158; TCGA KIRC -0.511; TCGA LGG -0.159; TCGA LIHC -0.439; TCGA PAAD -0.575; TCGA SARC -0.259; TCGA THCA -0.285; TCGA STAD -0.652 |
hsa-miR-24-3p | PPM1K | 11 cancers: BLCA; CESC; COAD; ESCA; KIRC; LUSC; PAAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.172; TCGA CESC -0.349; TCGA COAD -0.441; TCGA ESCA -0.604; TCGA KIRC -0.483; TCGA LUSC -0.383; TCGA PAAD -0.46; TCGA PRAD -0.133; TCGA SARC -0.356; TCGA STAD -0.584; TCGA UCEC -0.299 |
hsa-miR-24-3p | SLC46A1 | 13 cancers: BLCA; BRCA; CESC; ESCA; KIRC; KIRP; LGG; LIHC; PAAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.198; TCGA BRCA -0.351; TCGA CESC -0.151; TCGA ESCA -0.622; TCGA KIRC -0.262; TCGA KIRP -0.124; TCGA LGG -0.058; TCGA LIHC -0.298; TCGA PAAD -0.421; TCGA PRAD -0.288; TCGA SARC -0.379; TCGA STAD -0.229; TCGA UCEC -0.131 |
hsa-miR-24-3p | BSN | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LGG; LIHC; LUAD; LUSC; PAAD; SARC; STAD | MirTarget; mirMAP; miRNATAP | TCGA BLCA -0.298; TCGA BRCA -0.411; TCGA CESC -0.268; TCGA COAD -0.407; TCGA ESCA -0.662; TCGA KIRC -0.63; TCGA LGG -0.177; TCGA LIHC -0.588; TCGA LUAD -0.233; TCGA LUSC -0.282; TCGA PAAD -0.754; TCGA SARC -0.313; TCGA STAD -0.44 |
hsa-miR-24-3p | TECPR1 | 9 cancers: BLCA; BRCA; CESC; ESCA; LIHC; LUAD; LUSC; PRAD; UCEC | MirTarget | TCGA BLCA -0.097; TCGA BRCA -0.163; TCGA CESC -0.183; TCGA ESCA -0.344; TCGA LIHC -0.194; TCGA LUAD -0.264; TCGA LUSC -0.255; TCGA PRAD -0.158; TCGA UCEC -0.197 |
hsa-miR-24-3p | APPL2 | 9 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LGG; OV; PRAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.076; TCGA BRCA -0.225; TCGA CESC -0.138; TCGA ESCA -0.15; TCGA KIRC -0.254; TCGA LGG -0.073; TCGA OV -0.158; TCGA PRAD -0.156; TCGA UCEC -0.186 |
hsa-miR-24-3p | ADCY6 | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PRAD; UCEC | MirTarget | TCGA BLCA -0.188; TCGA BRCA -0.296; TCGA CESC -0.364; TCGA ESCA -0.88; TCGA HNSC -0.536; TCGA KIRC -0.32; TCGA KIRP -0.124; TCGA LUAD -0.198; TCGA LUSC -0.296; TCGA PRAD -0.295; TCGA UCEC -0.201 |
hsa-miR-24-3p | PDGFRA | 9 cancers: BLCA; CESC; COAD; ESCA; LGG; LUSC; PAAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.446; TCGA CESC -0.558; TCGA COAD -0.473; TCGA ESCA -0.805; TCGA LGG -0.259; TCGA LUSC -0.645; TCGA PAAD -0.474; TCGA STAD -0.215; TCGA UCEC -0.37 |
hsa-miR-24-3p | TADA2B | 10 cancers: BLCA; BRCA; ESCA; KIRC; LGG; LIHC; LUSC; PAAD; SARC; UCEC | MirTarget | TCGA BLCA -0.053; TCGA BRCA -0.243; TCGA ESCA -0.308; TCGA KIRC -0.198; TCGA LGG -0.066; TCGA LIHC -0.078; TCGA LUSC -0.116; TCGA PAAD -0.095; TCGA SARC -0.148; TCGA UCEC -0.09 |
hsa-miR-24-3p | RSAD1 | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LIHC; LUAD; PAAD; PRAD; THCA | MirTarget | TCGA BLCA -0.088; TCGA BRCA -0.167; TCGA CESC -0.196; TCGA ESCA -0.226; TCGA HNSC -0.151; TCGA LGG -0.094; TCGA LIHC -0.292; TCGA LUAD -0.181; TCGA PAAD -0.298; TCGA PRAD -0.342; TCGA THCA -0.058 |
hsa-miR-24-3p | TPSB2 | 9 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LUSC; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.631; TCGA BRCA -0.594; TCGA COAD -0.922; TCGA ESCA -0.511; TCGA HNSC -0.314; TCGA LUSC -0.596; TCGA PAAD -0.633; TCGA STAD -0.417; TCGA UCEC -0.6 |
hsa-miR-24-3p | ICA1L | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; OV; PAAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.377; TCGA BRCA -0.146; TCGA CESC -0.538; TCGA ESCA -0.46; TCGA HNSC -0.174; TCGA LGG -0.119; TCGA OV -0.181; TCGA PAAD -0.48; TCGA SARC -0.299; TCGA STAD -0.738; TCGA UCEC -0.239 |
hsa-miR-24-3p | SYP | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LUAD; LUSC; PAAD; PRAD; SARC; STAD | MirTarget; miRNATAP | TCGA BLCA -0.28; TCGA BRCA -0.368; TCGA CESC -0.275; TCGA COAD -0.461; TCGA ESCA -0.838; TCGA HNSC -0.385; TCGA LGG -0.148; TCGA LUAD -0.347; TCGA LUSC -0.227; TCGA PAAD -0.89; TCGA PRAD -0.383; TCGA SARC -0.246; TCGA STAD -0.35 |
hsa-miR-24-3p | ACBD4 | 12 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LIHC; LUAD; LUSC; PRAD; SARC; STAD | MirTarget; miRNATAP | TCGA BLCA -0.333; TCGA BRCA -0.417; TCGA CESC -0.45; TCGA ESCA -0.729; TCGA HNSC -0.483; TCGA LGG -0.096; TCGA LIHC -0.383; TCGA LUAD -0.258; TCGA LUSC -0.406; TCGA PRAD -0.19; TCGA SARC -0.46; TCGA STAD -0.297 |
hsa-miR-24-3p | MGAT4A | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; LUAD; LUSC; PAAD; PRAD; STAD | MirTarget | TCGA BLCA -0.413; TCGA BRCA -0.281; TCGA CESC -0.548; TCGA ESCA -1.33; TCGA HNSC -0.366; TCGA KIRC -0.255; TCGA LUAD -0.16; TCGA LUSC -0.701; TCGA PAAD -0.41; TCGA PRAD -0.196; TCGA STAD -0.209 |
hsa-miR-24-3p | GNAO1 | 10 cancers: BLCA; BRCA; COAD; LGG; LUSC; PAAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.603; TCGA BRCA -0.245; TCGA COAD -1.19; TCGA LGG -0.099; TCGA LUSC -0.255; TCGA PAAD -0.756; TCGA SARC -1.228; TCGA THCA -0.25; TCGA STAD -1.093; TCGA UCEC -0.389 |
hsa-miR-24-3p | SYNGR1 | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; OV; PAAD; SARC; UCEC | mirMAP | TCGA BLCA -0.371; TCGA BRCA -0.239; TCGA CESC -0.417; TCGA COAD -0.621; TCGA HNSC -0.371; TCGA KIRC -0.467; TCGA OV -0.25; TCGA PAAD -0.596; TCGA SARC -0.472; TCGA UCEC -0.16 |
hsa-miR-24-3p | FAM189A1 | 9 cancers: BLCA; CESC; HNSC; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -1.086; TCGA CESC -0.779; TCGA HNSC -0.477; TCGA LUSC -0.737; TCGA OV -0.287; TCGA PAAD -0.47; TCGA PRAD -0.239; TCGA STAD -1.211; TCGA UCEC -0.614 |
hsa-miR-24-3p | AGFG2 | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LIHC; LUAD; PAAD; PRAD; SARC; UCEC | mirMAP | TCGA BLCA -0.127; TCGA BRCA -0.223; TCGA CESC -0.232; TCGA ESCA -0.295; TCGA HNSC -0.157; TCGA LIHC -0.365; TCGA LUAD -0.162; TCGA PAAD -0.228; TCGA PRAD -0.25; TCGA SARC -0.171; TCGA UCEC -0.095 |
hsa-miR-24-3p | MPP2 | 9 cancers: BLCA; BRCA; CESC; COAD; HNSC; OV; PAAD; SARC; STAD | mirMAP | TCGA BLCA -0.416; TCGA BRCA -0.412; TCGA CESC -0.232; TCGA COAD -0.606; TCGA HNSC -0.283; TCGA OV -0.201; TCGA PAAD -0.659; TCGA SARC -0.398; TCGA STAD -0.586 |
hsa-miR-24-3p | PACSIN1 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; PRAD; THCA | mirMAP | TCGA BLCA -0.482; TCGA BRCA -0.237; TCGA CESC -0.823; TCGA COAD -0.702; TCGA ESCA -1.455; TCGA HNSC -0.625; TCGA LUAD -0.528; TCGA LUSC -0.377; TCGA PAAD -0.987; TCGA PRAD -0.34; TCGA THCA -0.272 |
hsa-miR-24-3p | CLYBL | 11 cancers: BLCA; CESC; ESCA; HNSC; KIRC; LIHC; PAAD; PRAD; SARC; THCA; STAD | mirMAP | TCGA BLCA -0.415; TCGA CESC -0.184; TCGA ESCA -0.415; TCGA HNSC -0.27; TCGA KIRC -0.28; TCGA LIHC -0.335; TCGA PAAD -0.279; TCGA PRAD -0.144; TCGA SARC -0.335; TCGA THCA -0.214; TCGA STAD -0.194 |
hsa-miR-24-3p | SLC26A1 | 10 cancers: BLCA; BRCA; CESC; ESCA; LGG; LIHC; LUAD; LUSC; PRAD; UCEC | mirMAP | TCGA BLCA -0.2; TCGA BRCA -0.517; TCGA CESC -0.259; TCGA ESCA -0.692; TCGA LGG -0.227; TCGA LIHC -0.487; TCGA LUAD -0.242; TCGA LUSC -0.271; TCGA PRAD -0.3; TCGA UCEC -0.259 |
hsa-miR-24-3p | CSDC2 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; LGG; LUSC; PAAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.951; TCGA BRCA -0.277; TCGA CESC -0.789; TCGA COAD -0.684; TCGA ESCA -0.675; TCGA LGG -0.346; TCGA LUSC -0.735; TCGA PAAD -0.52; TCGA SARC -0.604; TCGA STAD -0.296; TCGA UCEC -0.335 |
hsa-miR-24-3p | DES | 9 cancers: BLCA; CESC; COAD; ESCA; LUSC; PAAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -1.805; TCGA CESC -0.585; TCGA COAD -2.06; TCGA ESCA -1.257; TCGA LUSC -0.857; TCGA PAAD -0.891; TCGA SARC -1.624; TCGA STAD -2.033; TCGA UCEC -0.759 |
hsa-miR-24-3p | WSCD1 | 9 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LUSC; OV; PAAD | mirMAP | TCGA BLCA -0.718; TCGA BRCA -0.228; TCGA CESC -0.65; TCGA ESCA -1.065; TCGA HNSC -0.249; TCGA LGG -0.166; TCGA LUSC -0.835; TCGA OV -0.513; TCGA PAAD -0.49 |
hsa-miR-24-3p | ZNF445 | 11 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LGG; LUAD; PAAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.095; TCGA BRCA -0.125; TCGA CESC -0.166; TCGA ESCA -0.358; TCGA KIRC -0.332; TCGA LGG -0.092; TCGA LUAD -0.116; TCGA PAAD -0.14; TCGA SARC -0.135; TCGA STAD -0.219; TCGA UCEC -0.081 |
hsa-miR-24-3p | MAPT | 10 cancers: BLCA; BRCA; COAD; LGG; LUAD; PAAD; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.839; TCGA BRCA -1.073; TCGA COAD -1.037; TCGA LGG -0.217; TCGA LUAD -0.273; TCGA PAAD -0.808; TCGA PRAD -0.209; TCGA SARC -0.803; TCGA STAD -1.068; TCGA UCEC -0.224 |
hsa-miR-24-3p | KLHL3 | 9 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LUSC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.181; TCGA BRCA -0.117; TCGA CESC -0.231; TCGA ESCA -0.474; TCGA KIRC -0.561; TCGA LUSC -0.266; TCGA THCA -0.305; TCGA STAD -0.28; TCGA UCEC -0.318 |
hsa-miR-24-3p | CREBL2 | 12 cancers: BLCA; BRCA; CESC; ESCA; KIRC; KIRP; LIHC; LUSC; PAAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.19; TCGA BRCA -0.133; TCGA CESC -0.181; TCGA ESCA -0.294; TCGA KIRC -0.299; TCGA KIRP -0.098; TCGA LIHC -0.168; TCGA LUSC -0.217; TCGA PAAD -0.262; TCGA SARC -0.116; TCGA STAD -0.347; TCGA UCEC -0.114 |
hsa-miR-24-3p | ROR1 | 9 cancers: BLCA; CESC; COAD; ESCA; KIRC; LUSC; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.437; TCGA CESC -0.311; TCGA COAD -1.22; TCGA ESCA -1.035; TCGA KIRC -0.298; TCGA LUSC -1.137; TCGA SARC -0.418; TCGA STAD -0.514; TCGA UCEC -0.673 |
hsa-miR-24-3p | REEP1 | 10 cancers: BLCA; BRCA; CESC; ESCA; LGG; LUSC; PAAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -1.135; TCGA BRCA -0.86; TCGA CESC -0.497; TCGA ESCA -0.575; TCGA LGG -0.216; TCGA LUSC -0.399; TCGA PAAD -0.466; TCGA SARC -0.843; TCGA STAD -1.046; TCGA UCEC -0.492 |
hsa-miR-24-3p | CRY2 | 12 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LGG; LIHC; LUSC; PAAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.275; TCGA BRCA -0.285; TCGA CESC -0.182; TCGA ESCA -0.497; TCGA KIRC -0.346; TCGA LGG -0.238; TCGA LIHC -0.293; TCGA LUSC -0.259; TCGA PAAD -0.286; TCGA SARC -0.201; TCGA STAD -0.323; TCGA UCEC -0.287 |
hsa-miR-24-3p | PRKCB | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; THCA; STAD | miRNATAP | TCGA BLCA -0.497; TCGA CESC -0.348; TCGA COAD -1.149; TCGA ESCA -0.723; TCGA HNSC -0.412; TCGA LUAD -0.201; TCGA LUSC -0.813; TCGA PAAD -0.802; TCGA THCA -0.326; TCGA STAD -0.999 |
hsa-miR-24-3p | NCOA1 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LGG; PAAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.233; TCGA BRCA -0.082; TCGA CESC -0.172; TCGA COAD -0.162; TCGA ESCA -0.268; TCGA KIRC -0.308; TCGA LGG -0.059; TCGA PAAD -0.208; TCGA SARC -0.175; TCGA STAD -0.297; TCGA UCEC -0.16 |
hsa-miR-24-3p | SPTBN4 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; PAAD; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.765; TCGA BRCA -0.842; TCGA CESC -0.351; TCGA COAD -0.623; TCGA ESCA -0.962; TCGA HNSC -0.22; TCGA KIRC -0.707; TCGA KIRP -0.268; TCGA LIHC -0.44; TCGA PAAD -0.846; TCGA PRAD -0.354; TCGA SARC -0.56; TCGA STAD -0.848; TCGA UCEC -0.261 |
hsa-miR-24-3p | MOCS1 | 9 cancers: BLCA; BRCA; CESC; HNSC; LGG; LIHC; LUSC; PAAD; STAD | miRNATAP | TCGA BLCA -0.388; TCGA BRCA -0.129; TCGA CESC -0.41; TCGA HNSC -0.182; TCGA LGG -0.064; TCGA LIHC -0.379; TCGA LUSC -0.31; TCGA PAAD -0.548; TCGA STAD -0.365 |
hsa-miR-24-3p | PLCL2 | 13 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; LGG; LIHC; LUAD; LUSC; PAAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.26; TCGA CESC -0.767; TCGA COAD -0.693; TCGA ESCA -1.113; TCGA HNSC -0.385; TCGA KIRC -0.376; TCGA LGG -0.159; TCGA LIHC -0.196; TCGA LUAD -0.201; TCGA LUSC -0.698; TCGA PAAD -0.422; TCGA STAD -0.368; TCGA UCEC -0.469 |
hsa-miR-24-3p | PPIL6 | 9 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; LUSC; OV; UCEC | miRNATAP | TCGA BLCA -0.292; TCGA BRCA -0.156; TCGA CESC -0.667; TCGA ESCA -0.458; TCGA HNSC -0.257; TCGA KIRC -0.331; TCGA LUSC -0.492; TCGA OV -0.217; TCGA UCEC -0.177 |
hsa-miR-24-3p | MRVI1 | 9 cancers: BLCA; CESC; COAD; ESCA; LGG; LUSC; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.75; TCGA CESC -0.443; TCGA COAD -0.739; TCGA ESCA -0.757; TCGA LGG -0.133; TCGA LUSC -0.598; TCGA SARC -1.353; TCGA STAD -0.903; TCGA UCEC -0.431 |
hsa-miR-24-3p | AHCYL1 | 10 cancers: BLCA; BRCA; CESC; ESCA; KIRC; KIRP; LIHC; PAAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.062; TCGA BRCA -0.089; TCGA CESC -0.122; TCGA ESCA -0.166; TCGA KIRC -0.308; TCGA KIRP -0.124; TCGA LIHC -0.081; TCGA PAAD -0.136; TCGA STAD -0.177; TCGA UCEC -0.112 |
hsa-miR-24-3p | RNF150 | 9 cancers: BLCA; CESC; COAD; ESCA; KIRC; LGG; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.944; TCGA CESC -0.897; TCGA COAD -1.24; TCGA ESCA -0.562; TCGA KIRC -0.886; TCGA LGG -0.138; TCGA SARC -0.645; TCGA STAD -1.182; TCGA UCEC -0.228 |
hsa-miR-24-3p | ESRRG | 11 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LGG; LUAD; LUSC; PAAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.776; TCGA BRCA -0.546; TCGA CESC -1.291; TCGA ESCA -1.967; TCGA KIRC -1.479; TCGA LGG -0.21; TCGA LUAD -0.404; TCGA LUSC -0.415; TCGA PAAD -0.804; TCGA STAD -0.647; TCGA UCEC -0.239 |
hsa-miR-24-3p | TSC22D3 | 9 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUSC; PAAD; STAD | miRNATAP | TCGA BLCA -0.343; TCGA BRCA -0.302; TCGA CESC -0.287; TCGA COAD -0.29; TCGA ESCA -0.642; TCGA HNSC -0.216; TCGA LUSC -0.479; TCGA PAAD -0.255; TCGA STAD -0.444 |
hsa-miR-24-3p | CELF2 | 9 cancers: BLCA; CESC; ESCA; HNSC; LUSC; PAAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.658; TCGA CESC -0.226; TCGA ESCA -0.76; TCGA HNSC -0.255; TCGA LUSC -0.826; TCGA PAAD -0.616; TCGA SARC -0.229; TCGA STAD -0.79; TCGA UCEC -0.387 |
hsa-miR-24-3p | FAM46C | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUSC; PAAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.422; TCGA BRCA -0.386; TCGA CESC -0.588; TCGA COAD -0.545; TCGA ESCA -1.412; TCGA HNSC -0.465; TCGA KIRC -0.461; TCGA LUSC -0.797; TCGA PAAD -0.939; TCGA SARC -0.528; TCGA THCA -0.151; TCGA STAD -0.362; TCGA UCEC -0.35 |
hsa-miR-24-3p | ZSCAN21 | 10 cancers: BLCA; BRCA; ESCA; KIRC; LGG; LIHC; PAAD; PRAD; SARC; STAD | miRNATAP | TCGA BLCA -0.076; TCGA BRCA -0.08; TCGA ESCA -0.132; TCGA KIRC -0.149; TCGA LGG -0.091; TCGA LIHC -0.236; TCGA PAAD -0.133; TCGA PRAD -0.075; TCGA SARC -0.098; TCGA STAD -0.128 |
hsa-miR-24-3p | UBR1 | 10 cancers: BLCA; BRCA; CESC; ESCA; KIRC; KIRP; LUSC; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.086; TCGA BRCA -0.16; TCGA CESC -0.161; TCGA ESCA -0.285; TCGA KIRC -0.411; TCGA KIRP -0.11; TCGA LUSC -0.152; TCGA SARC -0.29; TCGA STAD -0.246; TCGA UCEC -0.284 |
hsa-miR-24-3p | ARL1 | 9 cancers: BRCA; CESC; ESCA; KIRC; KIRP; OV; PRAD; STAD; UCEC | miRNAWalker2 validate | TCGA BRCA -0.132; TCGA CESC -0.164; TCGA ESCA -0.131; TCGA KIRC -0.223; TCGA KIRP -0.119; TCGA OV -0.128; TCGA PRAD -0.1; TCGA STAD -0.114; TCGA UCEC -0.125 |
hsa-miR-24-3p | BRD8 | 10 cancers: BRCA; HNSC; KIRC; LGG; LIHC; LUAD; LUSC; PRAD; SARC; STAD | miRNAWalker2 validate | TCGA BRCA -0.174; TCGA HNSC -0.136; TCGA KIRC -0.194; TCGA LGG -0.128; TCGA LIHC -0.164; TCGA LUAD -0.136; TCGA LUSC -0.107; TCGA PRAD -0.084; TCGA SARC -0.153; TCGA STAD -0.089 |
hsa-miR-24-3p | CYP20A1 | 10 cancers: BRCA; CESC; ESCA; KIRC; LGG; OV; PAAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BRCA -0.057; TCGA CESC -0.18; TCGA ESCA -0.348; TCGA KIRC -0.13; TCGA LGG -0.091; TCGA OV -0.077; TCGA PAAD -0.167; TCGA SARC -0.091; TCGA STAD -0.357; TCGA UCEC -0.104 |
hsa-miR-24-3p | MLEC | 9 cancers: BRCA; CESC; ESCA; KIRP; LIHC; OV; PAAD; PRAD; THCA | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | TCGA BRCA -0.1; TCGA CESC -0.118; TCGA ESCA -0.334; TCGA KIRP -0.116; TCGA LIHC -0.194; TCGA OV -0.116; TCGA PAAD -0.165; TCGA PRAD -0.197; TCGA THCA -0.181 |
hsa-miR-24-3p | ZXDB | 9 cancers: BRCA; CESC; ESCA; KIRC; KIRP; LGG; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.131; TCGA CESC -0.265; TCGA ESCA -0.249; TCGA KIRC -0.354; TCGA KIRP -0.093; TCGA LGG -0.139; TCGA SARC -0.288; TCGA STAD -0.131; TCGA UCEC -0.182 |
hsa-miR-24-3p | TMEM161B | 11 cancers: BRCA; ESCA; HNSC; KIRC; KIRP; LGG; LIHC; LUSC; PRAD; THCA; STAD | MirTarget; miRNATAP | TCGA BRCA -0.183; TCGA ESCA -0.283; TCGA HNSC -0.123; TCGA KIRC -0.547; TCGA KIRP -0.149; TCGA LGG -0.156; TCGA LIHC -0.23; TCGA LUSC -0.135; TCGA PRAD -0.139; TCGA THCA -0.079; TCGA STAD -0.183 |
hsa-miR-24-3p | CES3 | 9 cancers: BRCA; CESC; ESCA; HNSC; LGG; LIHC; LUAD; THCA; UCEC | mirMAP | TCGA BRCA -0.229; TCGA CESC -0.395; TCGA ESCA -0.571; TCGA HNSC -0.316; TCGA LGG -0.153; TCGA LIHC -0.518; TCGA LUAD -0.517; TCGA THCA -0.371; TCGA UCEC -0.376 |
hsa-miR-24-3p | GMPPB | 9 cancers: BRCA; CESC; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; THCA | mirMAP | TCGA BRCA -0.212; TCGA CESC -0.156; TCGA ESCA -0.188; TCGA HNSC -0.145; TCGA LIHC -0.144; TCGA LUAD -0.119; TCGA LUSC -0.151; TCGA PRAD -0.376; TCGA THCA -0.277 |
hsa-miR-24-3p | BTBD9 | 9 cancers: BRCA; CESC; ESCA; KIRC; LUSC; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BRCA -0.164; TCGA CESC -0.143; TCGA ESCA -0.269; TCGA KIRC -0.192; TCGA LUSC -0.244; TCGA PAAD -0.141; TCGA PRAD -0.114; TCGA STAD -0.107; TCGA UCEC -0.06 |
hsa-miR-24-3p | MIA3 | 9 cancers: BRCA; CESC; ESCA; KIRC; LIHC; PRAD; SARC; THCA; UCEC | miRNATAP | TCGA BRCA -0.177; TCGA CESC -0.122; TCGA ESCA -0.273; TCGA KIRC -0.138; TCGA LIHC -0.416; TCGA PRAD -0.274; TCGA SARC -0.087; TCGA THCA -0.074; TCGA UCEC -0.14 |
hsa-miR-24-3p | MAPK9 | 9 cancers: BRCA; ESCA; KIRC; KIRP; PRAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BRCA -0.09; TCGA ESCA -0.166; TCGA KIRC -0.189; TCGA KIRP -0.1; TCGA PRAD -0.064; TCGA SARC -0.128; TCGA THCA -0.058; TCGA STAD -0.099; TCGA UCEC -0.109 |
hsa-miR-24-3p | NLK | 10 cancers: BRCA; CESC; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; THCA | miRNATAP | TCGA BRCA -0.111; TCGA CESC -0.085; TCGA ESCA -0.308; TCGA KIRC -0.187; TCGA KIRP -0.121; TCGA LIHC -0.074; TCGA LUAD -0.122; TCGA PAAD -0.165; TCGA PRAD -0.082; TCGA THCA -0.067 |
hsa-miR-24-3p | OGT | 10 cancers: BRCA; CESC; ESCA; HNSC; KIRC; LGG; LUAD; LUSC; PRAD; STAD | miRNATAP | TCGA BRCA -0.075; TCGA CESC -0.106; TCGA ESCA -0.38; TCGA HNSC -0.145; TCGA KIRC -0.286; TCGA LGG -0.092; TCGA LUAD -0.306; TCGA LUSC -0.254; TCGA PRAD -0.297; TCGA STAD -0.17 |
hsa-miR-24-3p | GNE | 9 cancers: CESC; ESCA; HNSC; KIRC; LGG; LIHC; PRAD; SARC; UCEC | MirTarget | TCGA CESC -0.468; TCGA ESCA -0.56; TCGA HNSC -0.16; TCGA KIRC -0.329; TCGA LGG -0.081; TCGA LIHC -0.238; TCGA PRAD -0.275; TCGA SARC -0.145; TCGA UCEC -0.189 |
hsa-miR-24-3p | ACSL6 | 10 cancers: ESCA; HNSC; KIRC; LGG; LIHC; LUSC; PAAD; SARC; THCA; STAD | MirTarget | TCGA ESCA -0.845; TCGA HNSC -0.383; TCGA KIRC -0.753; TCGA LGG -0.36; TCGA LIHC -1.148; TCGA LUSC -0.374; TCGA PAAD -1.257; TCGA SARC -0.385; TCGA THCA -0.644; TCGA STAD -0.836 |
Enriched cancer pathways of putative targets