microRNA information: hsa-miR-381-3p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-381-3p | miRbase |
| Accession: | MIMAT0000736 | miRbase |
| Precursor name: | hsa-mir-381 | miRbase |
| Precursor accession: | MI0000789 | miRbase |
| Symbol: | MIR381 | HGNC |
| RefSeq ID: | NR_029873 | GenBank |
| Sequence: | UAUACAAGGGCAAGCUCUCUGU |
Reported expression in cancers: hsa-miR-381-3p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-381-3p | colon cancer | downregulation | "Down regulation of MicroRNA 381 promotes cell prol ......" | 26320367 | qPCR |
| hsa-miR-381-3p | colorectal cancer | downregulation | "MiR-381 has been reported to be dysregulated in se ......" | 27094913 | qPCR |
| hsa-miR-381-3p | glioblastoma | upregulation | "MicroRNA-381 miR-381 is a highly expressed onco-mi ......" | 25605243 | |
| hsa-miR-381-3p | lung cancer | downregulation | "MiR-381 expression was measured in 18 human lung a ......" | 22592211 | Reverse transcription PCR |
| hsa-miR-381-3p | ovarian cancer | deregulation | "A series of recent studies suggested that miR-381 ......" | 26768613 | |
| hsa-miR-381-3p | sarcoma | downregulation | "In this study we found that miR-381 is a positive ......" | 27612424 |
Reported cancer pathway affected by hsa-miR-381-3p

| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-381-3p | colorectal cancer | Epithelial mesenchymal transition pathway | "MiR 381 functions as a tumor suppressor in colorec ......" | 27094913 | Luciferase |
| hsa-miR-381-3p | liver cancer | cell cycle pathway; Wnt signaling pathway | "MicroRNA 381 suppresses cell growth and invasion b ......" | 26677080 | Colony formation; Western blot; Luciferase |
| hsa-miR-381-3p | ovarian cancer | Wnt signaling pathway | "MiR 381 inhibits epithelial ovarian cancer maligna ......" | 26768613 | |
| hsa-miR-381-3p | prostate cancer | Apoptosis pathway | "MicroRNAs miR 154 miR 299 5p miR 376a miR 376c miR ......" | 24166498 |
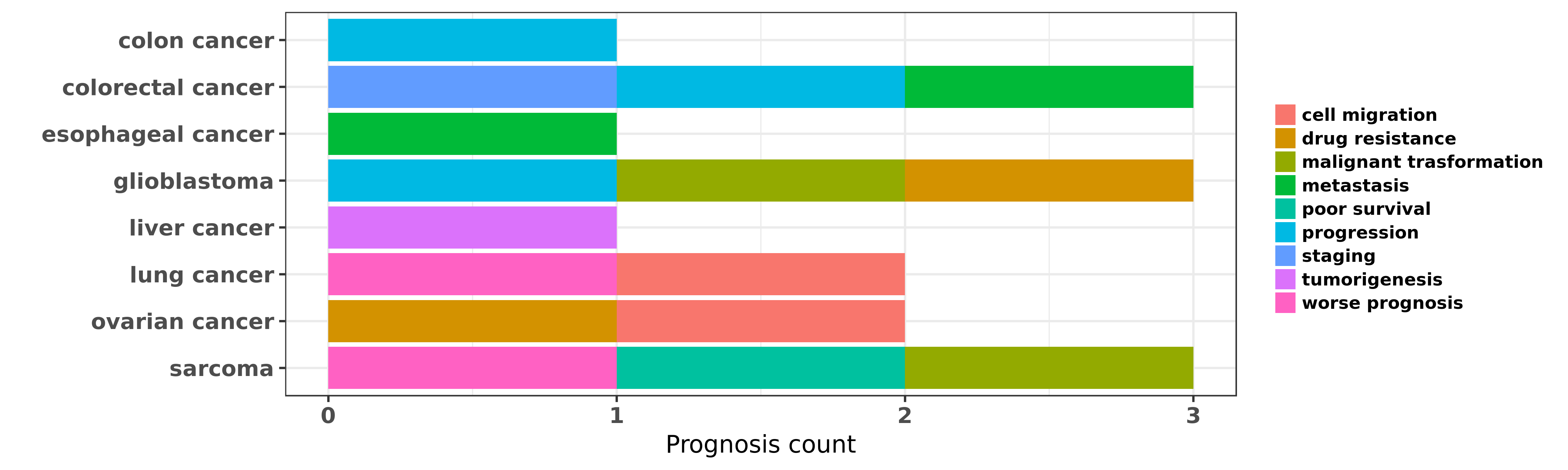
Reported cancer prognosis affected by hsa-miR-381-3p
| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-381-3p | colon cancer | progression | "Down regulation of MicroRNA 381 promotes cell prol ......" | 26320367 | Luciferase |
| hsa-miR-381-3p | colorectal cancer | staging; metastasis; progression | "MiR 381 functions as a tumor suppressor in colorec ......" | 27094913 | Luciferase |
| hsa-miR-381-3p | esophageal cancer | metastasis | "MicroRNA 381 increases radiosensitivity in esophag ......" | 25628936 | Colony formation; Transwell assay |
| hsa-miR-381-3p | glioblastoma | drug resistance; malignant trasformation; progression | "Targeting miR 381 NEFL axis sensitizes glioblastom ......" | 25605243 | |
| hsa-miR-381-3p | liver cancer | tumorigenesis | "MicroRNA 381 suppresses cell growth and invasion b ......" | 26677080 | Colony formation; Western blot; Luciferase |
| hsa-miR-381-3p | lung cancer | cell migration; worse prognosis | "MicroRNA 381 represses ID1 and is deregulated in l ......" | 22592211 | Luciferase |
| hsa-miR-381-3p | ovarian cancer | drug resistance | "Hybridization was carried out on miRNA microarray ......" | 24805828 | |
| hsa-miR-381-3p | ovarian cancer | cell migration | "MiR 381 inhibits epithelial ovarian cancer maligna ......" | 26768613 | |
| hsa-miR-381-3p | sarcoma | malignant trasformation | "SMARCB1 expression in epithelioid sarcoma is regul ......" | 24327545 | |
| hsa-miR-381-3p | sarcoma | worse prognosis; poor survival | "Low expression of miR 381 is a favorite prognosis ......" | 27612424 |
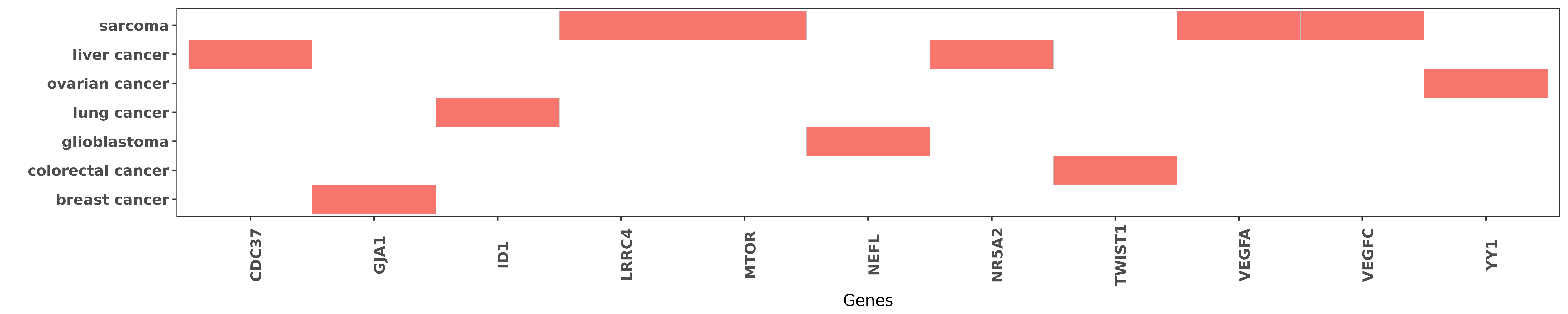
Reported gene related to hsa-miR-381-3p
| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-miR-381-3p | liver cancer | CDC37 | "MicroRNA 381 suppresses cell growth and invasion b ......" | 26677080 |
| hsa-miR-381-3p | breast cancer | GJA1 | "miR 381 suppresses C/EBPα dependent Cx43 expressi ......" | 26450928 |
| hsa-miR-381-3p | lung cancer | ID1 | "MicroRNA 381 represses ID1 and is deregulated in l ......" | 22592211 |
| hsa-miR-381-3p | sarcoma | LRRC4 | "Our results also showed a strong negative correlat ......" | 27612424 |
| hsa-miR-381-3p | sarcoma | MTOR | "This demonstrated that LRRC4 is a direct target ge ......" | 27612424 |
| hsa-miR-381-3p | glioblastoma | NEFL | "Targeting miR 381 NEFL axis sensitizes glioblastom ......" | 25605243 |
| hsa-miR-381-3p | liver cancer | NR5A2 | "Bioinformatics analysis and dual-luciferase report ......" | 26677080 |
| hsa-miR-381-3p | colorectal cancer | TWIST1 | "MiR 381 functions as a tumor suppressor in colorec ......" | 27094913 |
| hsa-miR-381-3p | sarcoma | VEGFA | "Basic fibroblast growth factor promotes VEGF C dep ......" | 27229532 |
| hsa-miR-381-3p | sarcoma | VEGFC | "Basic fibroblast growth factor promotes VEGF C dep ......" | 27229532 |
| hsa-miR-381-3p | ovarian cancer | YY1 | "MiR 381 inhibits epithelial ovarian cancer maligna ......" | 26768613 |
Expression profile in cancer corhorts:
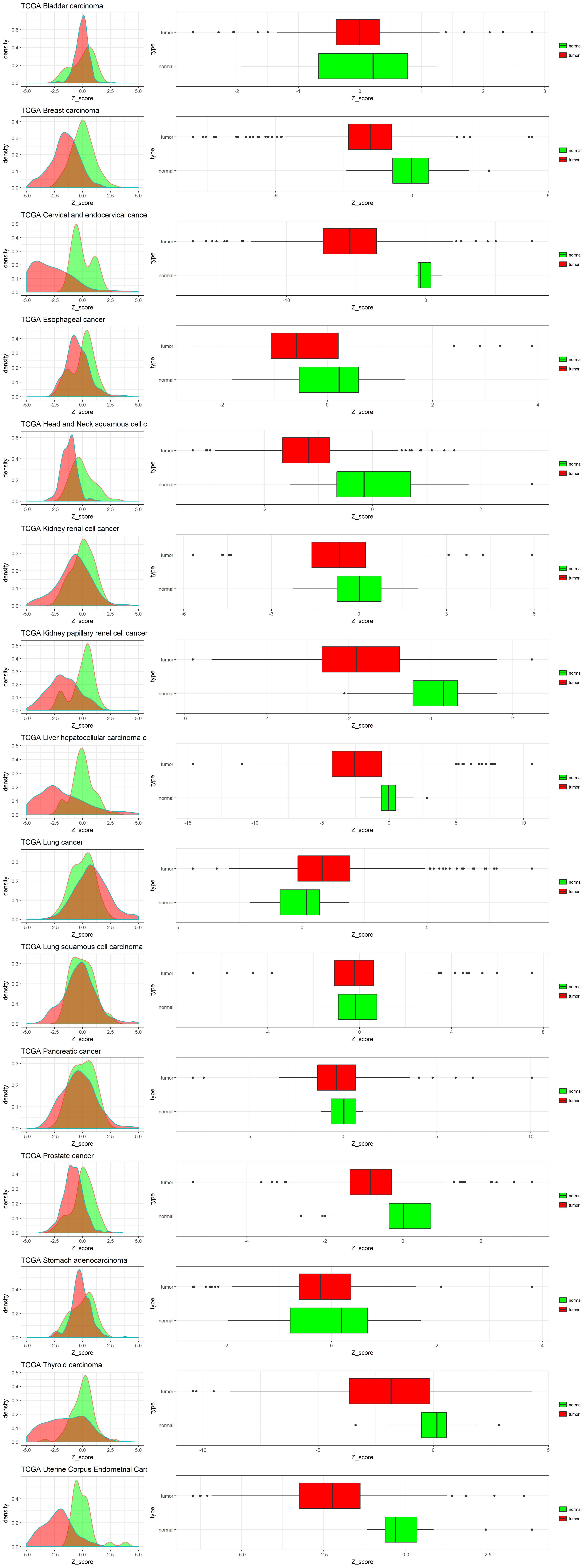
Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-381-3p | ACBD5 | 9 cancers: BLCA; BRCA; ESCA; KIRC; LGG; LIHC; LUAD; LUSC; THCA | MirTarget | TCGA BLCA -0.078; TCGA BRCA -0.159; TCGA ESCA -0.197; TCGA KIRC -0.052; TCGA LGG -0.054; TCGA LIHC -0.056; TCGA LUAD -0.082; TCGA LUSC -0.083; TCGA THCA -0.053 |
hsa-miR-381-3p | NDUFA5 | 9 cancers: BLCA; BRCA; COAD; ESCA; KIRC; KIRP; OV; THCA; STAD | mirMAP | TCGA BLCA -0.062; TCGA BRCA -0.142; TCGA COAD -0.119; TCGA ESCA -0.127; TCGA KIRC -0.081; TCGA KIRP -0.114; TCGA OV -0.055; TCGA THCA -0.066; TCGA STAD -0.104 |
hsa-miR-381-3p | VSIG10 | 12 cancers: BLCA; BRCA; ESCA; KIRC; KIRP; LGG; LIHC; LUAD; LUSC; PAAD; PRAD; SARC | mirMAP | TCGA BLCA -0.102; TCGA BRCA -0.118; TCGA ESCA -0.233; TCGA KIRC -0.094; TCGA KIRP -0.081; TCGA LGG -0.124; TCGA LIHC -0.075; TCGA LUAD -0.086; TCGA LUSC -0.241; TCGA PAAD -0.1; TCGA PRAD -0.129; TCGA SARC -0.092 |
hsa-miR-381-3p | PARD6B | 13 cancers: BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; PRAD; SARC; THCA; UCEC | MirTarget | TCGA BRCA -0.434; TCGA CESC -0.109; TCGA ESCA -0.21; TCGA HNSC -0.165; TCGA KIRC -0.154; TCGA KIRP -0.065; TCGA LUAD -0.093; TCGA LUSC -0.105; TCGA PAAD -0.211; TCGA PRAD -0.1; TCGA SARC -0.229; TCGA THCA -0.05; TCGA UCEC -0.129 |
Enriched cancer pathways of putative targets