microRNA information: hsa-miR-450a-5p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-450a-5p | miRbase |
| Accession: | MIMAT0001545 | miRbase |
| Precursor name: | hsa-mir-450a-1 | miRbase |
| Precursor accession: | MI0001652 | miRbase |
| Symbol: | MIR450A1 | HGNC |
| RefSeq ID: | NR_029962 | GenBank |
| Sequence: | UUUUGCGAUGUGUUCCUAAUAU |
Reported expression in cancers: hsa-miR-450a-5p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-450a-5p | NA | NA | "NA ......" | NA | NA |
| hsa-miR-450a-5p | " ......" |
Reported cancer pathway affected by hsa-miR-450a-5p
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|
Reported cancer prognosis affected by hsa-miR-450a-5p
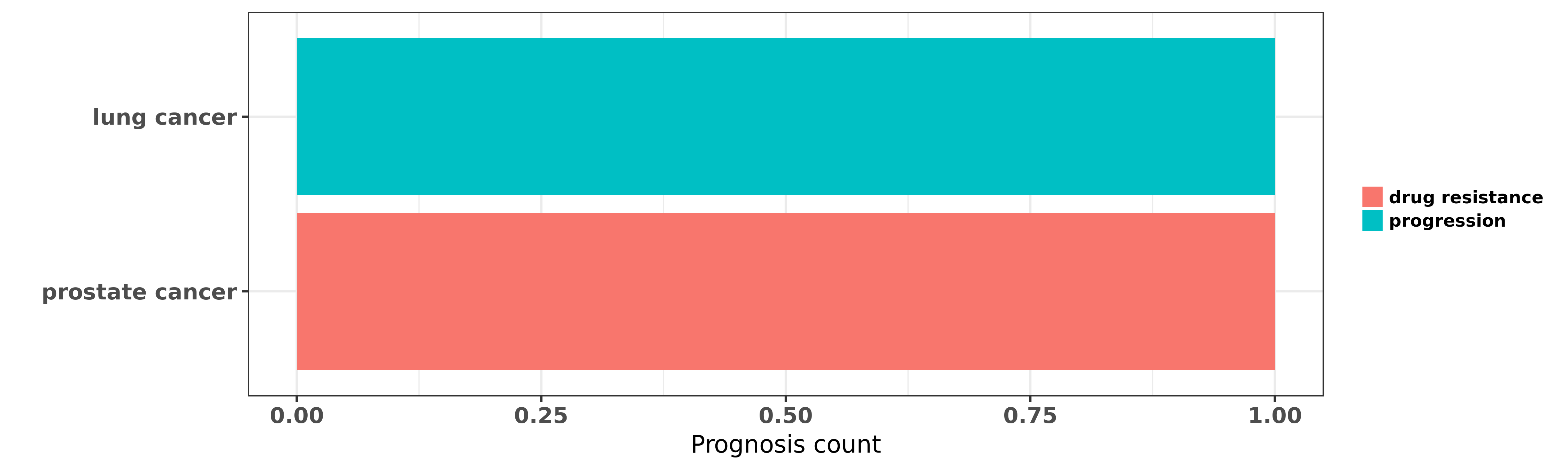
| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-450a-5p | lung cancer | progression | "Upregulation of microRNA 450 inhibits the progress ......" | 27246609 | |
| hsa-miR-450a-5p | prostate cancer | drug resistance | "To screen for epigenetically silenced miRNAs in pr ......" | 22310291 |
Reported gene related to hsa-miR-450a-5p
| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-miR-450a-5p | lung cancer | IRF2 | "miR-450 targets IRF2 and thus supresses lung cance ......" | 27246609 |
Expression profile in cancer corhorts:
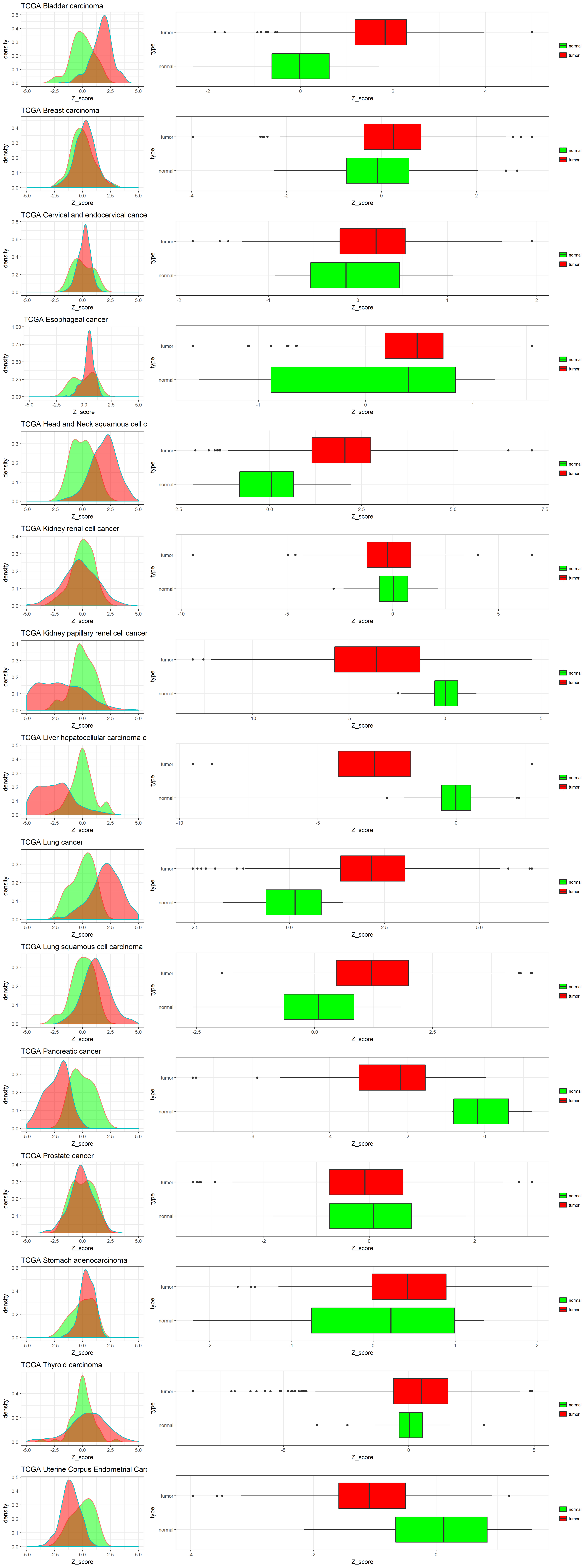
Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-450a-5p | FBXW11 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; LGG; LUAD; LUSC; PAAD; PRAD; STAD | miRNAWalker2 validate | TCGA BLCA -0.107; TCGA BRCA -0.062; TCGA CESC -0.074; TCGA COAD -0.06; TCGA HNSC -0.141; TCGA KIRC -0.057; TCGA LGG -0.085; TCGA LUAD -0.085; TCGA LUSC -0.094; TCGA PAAD -0.173; TCGA PRAD -0.057; TCGA STAD -0.088 |
hsa-miR-450a-5p | SMG1 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; PRAD; SARC; STAD | MirTarget | TCGA BLCA -0.087; TCGA CESC -0.076; TCGA COAD -0.056; TCGA ESCA -0.077; TCGA HNSC -0.066; TCGA KIRC -0.088; TCGA KIRP -0.094; TCGA PRAD -0.118; TCGA SARC -0.074; TCGA STAD -0.096 |
hsa-miR-450a-5p | NFIX | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LGG; LIHC; LUAD; LUSC; SARC; STAD | mirMAP | TCGA BLCA -0.165; TCGA BRCA -0.097; TCGA CESC -0.104; TCGA ESCA -0.171; TCGA HNSC -0.314; TCGA LGG -0.12; TCGA LIHC -0.101; TCGA LUAD -0.162; TCGA LUSC -0.2; TCGA SARC -0.077; TCGA STAD -0.135 |
hsa-miR-450a-5p | PBX1 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LGG; LUAD; LUSC; PAAD; PRAD; SARC; STAD | miRNATAP | TCGA BLCA -0.281; TCGA BRCA -0.153; TCGA CESC -0.167; TCGA COAD -0.186; TCGA HNSC -0.494; TCGA LGG -0.114; TCGA LUAD -0.113; TCGA LUSC -0.208; TCGA PAAD -0.171; TCGA PRAD -0.083; TCGA SARC -0.265; TCGA STAD -0.32 |
hsa-miR-450a-5p | CEP350 | 12 cancers: BLCA; CESC; COAD; HNSC; KIRC; LIHC; LUAD; PAAD; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.089; TCGA CESC -0.082; TCGA COAD -0.057; TCGA HNSC -0.162; TCGA KIRC -0.066; TCGA LIHC -0.134; TCGA LUAD -0.05; TCGA PAAD -0.125; TCGA PRAD -0.104; TCGA THCA -0.055; TCGA STAD -0.073; TCGA UCEC -0.063 |
Enriched cancer pathways of putative targets