microRNA information: hsa-miR-500b-5p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-500b-5p | miRbase |
| Accession: | MIMAT0016925 | miRbase |
| Precursor name: | hsa-mir-500b | miRbase |
| Precursor accession: | MI0015903 | miRbase |
| Symbol: | MIR500B | HGNC |
| RefSeq ID: | NR_036257 | GenBank |
| Sequence: | AAUCCUUGCUACCUGGGU |
Reported expression in cancers: hsa-miR-500b-5p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-500b-5p | gastric cancer | upregulation | "Furthermore we found that miR-500 promoted gastric ......" | 25595906 |
Reported cancer pathway affected by hsa-miR-500b-5p
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|
Reported cancer prognosis affected by hsa-miR-500b-5p

| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-500b-5p | gastric cancer | poor survival; malignant trasformation; progression | "Here we report that the miR-500 directly repressed ......" | 25595906 | |
| hsa-miR-500b-5p | lung cancer | progression | "We succeeded in establishing the MTA1 knockdown NS ......" | 22576802 | RNAi |
Reported gene related to hsa-miR-500b-5p
| miRNA | cancer | gene | reporting | PUBMED |
|---|
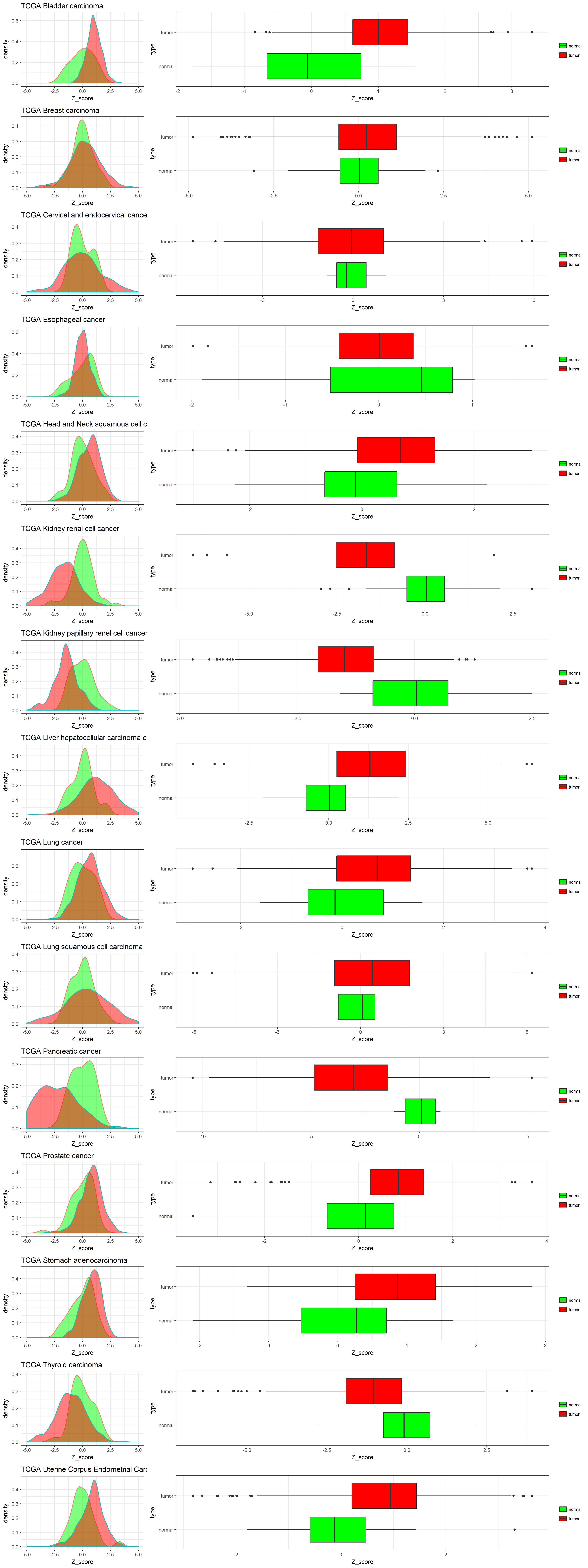
Expression profile in cancer corhorts:
Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-500b-5p | PLEKHM3 | 11 cancers: BLCA; ESCA; HNSC; LGG; LIHC; LUAD; LUSC; PAAD; PRAD; SARC; STAD | MirTarget | TCGA BLCA -0.119; TCGA ESCA -0.108; TCGA HNSC -0.239; TCGA LGG -0.252; TCGA LIHC -0.125; TCGA LUAD -0.092; TCGA LUSC -0.078; TCGA PAAD -0.147; TCGA PRAD -0.32; TCGA SARC -0.138; TCGA STAD -0.218 |
hsa-miR-500b-5p | STRN3 | 11 cancers: BLCA; BRCA; ESCA; KIRP; LGG; LIHC; LUAD; PAAD; PRAD; SARC; STAD | MirTarget | TCGA BLCA -0.142; TCGA BRCA -0.169; TCGA ESCA -0.124; TCGA KIRP -0.069; TCGA LGG -0.075; TCGA LIHC -0.117; TCGA LUAD -0.07; TCGA PAAD -0.094; TCGA PRAD -0.124; TCGA SARC -0.108; TCGA STAD -0.121 |
hsa-miR-500b-5p | CCDC91 | 9 cancers: BLCA; BRCA; CESC; ESCA; LUAD; PAAD; PRAD; SARC; STAD | MirTarget | TCGA BLCA -0.09; TCGA BRCA -0.096; TCGA CESC -0.122; TCGA ESCA -0.174; TCGA LUAD -0.163; TCGA PAAD -0.149; TCGA PRAD -0.098; TCGA SARC -0.096; TCGA STAD -0.098 |
hsa-miR-500b-5p | SLITRK6 | 10 cancers: BLCA; BRCA; ESCA; HNSC; LGG; LIHC; PAAD; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.476; TCGA BRCA -0.67; TCGA ESCA -0.586; TCGA HNSC -0.221; TCGA LGG -0.288; TCGA LIHC -0.365; TCGA PAAD -0.5; TCGA PRAD -0.246; TCGA THCA -0.327; TCGA STAD -0.685 |
hsa-miR-500b-5p | TGFB2 | 9 cancers: BLCA; CESC; HNSC; LUSC; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.231; TCGA CESC -0.327; TCGA HNSC -0.16; TCGA LUSC -0.213; TCGA PAAD -0.297; TCGA PRAD -0.098; TCGA THCA -0.27; TCGA STAD -0.19; TCGA UCEC -0.208 |
hsa-miR-500b-5p | FAM102B | 9 cancers: BLCA; BRCA; HNSC; KIRC; KIRP; LGG; LUAD; PRAD; THCA | MirTarget | TCGA BLCA -0.09; TCGA BRCA -0.115; TCGA HNSC -0.148; TCGA KIRC -0.184; TCGA KIRP -0.081; TCGA LGG -0.081; TCGA LUAD -0.097; TCGA PRAD -0.099; TCGA THCA -0.148 |
hsa-miR-500b-5p | DPP8 | 11 cancers: BLCA; BRCA; CESC; HNSC; LGG; LIHC; LUAD; PAAD; PRAD; SARC; STAD | MirTarget | TCGA BLCA -0.142; TCGA BRCA -0.203; TCGA CESC -0.148; TCGA HNSC -0.301; TCGA LGG -0.264; TCGA LIHC -0.108; TCGA LUAD -0.135; TCGA PAAD -0.167; TCGA PRAD -0.381; TCGA SARC -0.107; TCGA STAD -0.075 |
hsa-miR-500b-5p | RAPGEF6 | 10 cancers: BLCA; BRCA; HNSC; KIRP; LGG; LUAD; LUSC; PRAD; SARC; STAD | MirTarget | TCGA BLCA -0.161; TCGA BRCA -0.325; TCGA HNSC -0.513; TCGA KIRP -0.162; TCGA LGG -0.267; TCGA LUAD -0.172; TCGA LUSC -0.145; TCGA PRAD -0.464; TCGA SARC -0.191; TCGA STAD -0.05 |
hsa-miR-500b-5p | HMCN1 | 10 cancers: BLCA; BRCA; ESCA; HNSC; KIRC; LGG; LUSC; PAAD; PRAD; STAD | MirTarget | TCGA BLCA -0.211; TCGA BRCA -0.322; TCGA ESCA -0.419; TCGA HNSC -0.387; TCGA KIRC -0.349; TCGA LGG -0.202; TCGA LUSC -0.417; TCGA PAAD -0.515; TCGA PRAD -0.176; TCGA STAD -0.479 |
hsa-miR-500b-5p | RPS6KB1 | 10 cancers: BRCA; CESC; HNSC; KIRC; KIRP; LGG; LIHC; LUAD; PAAD; PRAD | MirTarget | TCGA BRCA -0.075; TCGA CESC -0.069; TCGA HNSC -0.064; TCGA KIRC -0.053; TCGA KIRP -0.128; TCGA LGG -0.125; TCGA LIHC -0.065; TCGA LUAD -0.088; TCGA PAAD -0.095; TCGA PRAD -0.123 |
hsa-miR-500b-5p | CHSY1 | 10 cancers: BRCA; CESC; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; PRAD; THCA | MirTarget | TCGA BRCA -0.069; TCGA CESC -0.109; TCGA HNSC -0.122; TCGA KIRC -0.238; TCGA KIRP -0.105; TCGA LUAD -0.152; TCGA LUSC -0.065; TCGA PAAD -0.178; TCGA PRAD -0.096; TCGA THCA -0.074 |
hsa-miR-500b-5p | CYP1B1 | 9 cancers: ESCA; HNSC; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD | MirTarget | TCGA ESCA -0.345; TCGA HNSC -0.342; TCGA LUAD -0.196; TCGA LUSC -0.269; TCGA PAAD -0.32; TCGA PRAD -0.22; TCGA SARC -0.259; TCGA THCA -0.668; TCGA STAD -0.668 |
Enriched cancer pathways of putative targets