This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-708-3p | APAF1 | 0.14 | 0.89698 | -0.17 | 0.88665 | mirMAP | -0.11 | 0.0011 | NA | |
| 2 | hsa-miR-324-5p | ATM | -0.5 | 0.53742 | 0.19 | 0.86463 | miRanda | -0.12 | 0.00155 | NA | |
| 3 | hsa-miR-455-5p | ATM | -0.14 | 0.86574 | 0.19 | 0.86463 | miRanda | -0.12 | 0.00128 | NA | |
| 4 | hsa-miR-590-5p | ATM | -0.55 | 0.47274 | 0.19 | 0.86463 | mirMAP | -0.1 | 0.00142 | NA | |
| 5 | hsa-miR-30a-5p | CASP3 | 0.22 | 0.93395 | -0.16 | 0.90316 | miRNATAP | -0.2 | 6.0E-5 | NA | |
| 6 | hsa-miR-143-3p | CASP8 | 0.37 | 0.92734 | -0.27 | 0.82859 | MirTarget | -0.11 | 0.00055 | NA | |
| 7 | hsa-miR-133b | CCNB1 | 0.39 | 0.56076 | -0.28 | 0.85013 | miRanda | -0.13 | 0 | NA | |
| 8 | hsa-miR-139-5p | CCNB1 | -0.08 | 0.92869 | -0.28 | 0.85013 | miRanda | -0.18 | 0.00084 | NA | |
| 9 | hsa-let-7c-5p | CCNB2 | -0.06 | 0.9749 | -0.2 | 0.87078 | miRNAWalker2 validate | -0.19 | 0 | NA | |
| 10 | hsa-miR-23b-3p | CCNB2 | -0.11 | 0.96135 | -0.2 | 0.87078 | miRNAWalker2 validate | -0.33 | 0.00011 | NA | |
| 11 | hsa-miR-16-5p | CCND1 | 0.01 | 0.99448 | -0.25 | 0.89067 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.13 | 0.00823 | 23991964; 22922827; 18483394 | At the molecular level our results further revealed that cyclin D1 expression was negatively regulated by miR-16;CCND1 has been found to be a target of miR-15a and miR-16-1 through analysis of complementary sequences between microRNAs and CCND1 mRNA; Moreover the transcription of CCND1 is suppressed by miR-15a and miR-16-1 via direct binding to the CCND1 3'-untranslated region 3'-UTR;Truncation in CCND1 mRNA alters miR 16 1 regulation in mantle cell lymphoma; Furthermore we demonstrated that this truncation alters miR-16-1 binding sites and through the use of reporter constructs we were able to show that miR-16-1 regulates CCND1 mRNA expression; This study introduces the role of miR-16-1 in the regulation of CCND1 in MCL |
| 12 | hsa-miR-195-5p | CCND1 | 0.34 | 0.74962 | -0.25 | 0.89067 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.14 | 0.00272 | 21350001; 26631043; 25823925 | Raf-1 and Ccnd1 were identified as novel direct targets of miR-195 and miR-497 miR-195/497 expression levels in clinical specimens were found to be correlated inversely with malignancy of breast cancer;MiR 195 inhibits the proliferation of human cervical cancer cells by directly targeting cyclin D1; The present study was to evaluate the level of miR-195 and cyclin D1 in CC tissues and cells; We further investigated the molecular mechanisms of miR-195 and cyclin D1 in CC cell lines HeLa and SiHa; Furthermore the expression of miR-195 was inversely proportional to that of cyclin D1 mRNA or protein p = 0.013 p = 0.015 respectively; However the inhibitor of miR-195 promoted the expression of cyclin D1 and cell proliferation; In conclusion our data suggest that miR-195 may have the potential role in treatment of CC patients as well as miR-195 is a novel regulator of invasiveness and tumorigenicity in CC cells by targeting cyclin D1;MicroRNA profiling identifies MiR 195 suppresses osteosarcoma cell metastasis by targeting CCND1; Meanwhile CCND1 was identified as the target gene of miR-195 and further studied; More importantly using real-time PCR we evaluated the expression of miR-195 and CCND1 in osteosarcoma samples from 107 frozen biopsy tissues and 99 formalin- or paraformalin-fixed paraffin-embedded FFPE tissues; Results indicated lowly expressed miR-195 or highly CCND1 correlated with positive overall survival and their expression inversely related to each other; In summary our study suggests miR-195 functions as a tumor metastasis suppressor gene by down-regulating CCND1 and can be used as a potential target in the treatment of osteosarcoma |
| 13 | hsa-miR-29c-3p | CCND1 | 0.16 | 0.94272 | -0.25 | 0.89067 | mirMAP | -0.14 | 0.00181 | NA | |
| 14 | hsa-miR-125b-5p | CCNE1 | 0.04 | 0.98059 | -0.28 | 0.73651 | miRNAWalker2 validate | -0.15 | 0.00074 | NA | |
| 15 | hsa-miR-195-5p | CCNE1 | 0.34 | 0.74962 | -0.28 | 0.73651 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.18 | 0.00302 | 24402230 | Furthermore through qPCR and western blot assays we showed that overexpression of miR-195-5p reduced CCNE1 mRNA and protein levels respectively |
| 16 | hsa-miR-26a-5p | CCNE2 | 0.01 | 0.99772 | -0.15 | 0.8473 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.36 | 0.00084 | 24116110; 21901171 | The loss of miR 26a mediated post transcriptional regulation of cyclin E2 in pancreatic cancer cell proliferation and decreased patient survival; The in vitro and in vivo assays showed that overexpression of miR-26a resulted in cell cycle arrest inhibited cell proliferation and decreased tumor growth which was associated with cyclin E2 downregulation;We also show that enforced expression of miR-26a in AML cells is able to inhibit cell cycle progression by downregulating cyclin E2 expression |
| 17 | hsa-miR-30a-5p | CCNE2 | 0.22 | 0.93395 | -0.15 | 0.8473 | miRNATAP | -0.44 | 0 | NA | |
| 18 | hsa-miR-34a-5p | CCNE2 | -0.5 | 0.74203 | -0.15 | 0.8473 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.22 | 0.00018 | NA | |
| 19 | hsa-miR-34c-5p | CCNE2 | -0.02 | 0.95279 | -0.15 | 0.8473 | miRNAWalker2 validate; miRTarBase; PITA; miRanda; miRNATAP | -0.28 | 0 | NA | |
| 20 | hsa-let-7c-5p | CCNG1 | -0.06 | 0.9749 | -0.02 | 0.9911 | miRNAWalker2 validate | -0.11 | 0.00149 | NA | |
| 21 | hsa-let-7e-5p | CCNG1 | 0.21 | 0.92234 | -0.02 | 0.9911 | miRNAWalker2 validate | -0.23 | 0.00082 | NA | |
| 22 | hsa-miR-100-5p | CCNG1 | 0.02 | 0.99282 | -0.02 | 0.9911 | miRNAWalker2 validate | -0.12 | 0.00075 | NA | |
| 23 | hsa-miR-23b-3p | CCNG1 | -0.11 | 0.96135 | -0.02 | 0.9911 | MirTarget; miRNATAP | -0.22 | 0.0086 | 26872615 | MiR 23b targets cyclin G1 and suppresses ovarian cancer tumorigenesis and progression; Dual-luciferase reporter assay and a xenograft mouse model were used to examine the expression of miR-23b and its target gene CCNG1; Dual-luciferase reporter assay showed that miR-23b bound with the 3' untranslated region of CCNG1; Furthermore miR-23b inhibited tumor growth and suppressed CCNG1 expression in vitro; Our findings show that miR-23b may inhibit ovarian cancer tumorigenesis and progression by downregulating CCNG1 and the expression of the relevant genes |
| 24 | hsa-miR-106a-5p | CCNG2 | -0.2 | 0.80221 | -0.11 | 0.92934 | MirTarget; miRNATAP | -0.13 | 0.00134 | NA | |
| 25 | hsa-miR-106b-5p | CCNG2 | -0.3 | 0.86929 | -0.11 | 0.92934 | MirTarget; miRNATAP | -0.12 | 0.00599 | NA | |
| 26 | hsa-miR-17-5p | CCNG2 | -0.18 | 0.93454 | -0.11 | 0.92934 | MirTarget; TargetScan; miRNATAP | -0.15 | 2.0E-5 | NA | |
| 27 | hsa-miR-20a-5p | CCNG2 | -0.18 | 0.92812 | -0.11 | 0.92934 | MirTarget; miRNATAP | -0.12 | 0.00028 | NA | |
| 28 | hsa-miR-224-5p | CCNG2 | -0.24 | 0.85735 | -0.11 | 0.92934 | mirMAP | -0.13 | 0 | NA | |
| 29 | hsa-miR-29a-5p | CCNG2 | -0.32 | 0.60044 | -0.11 | 0.92934 | mirMAP | -0.11 | 0.00322 | NA | |
| 30 | hsa-miR-331-5p | CCNG2 | -0.45 | 0.25036 | -0.11 | 0.92934 | miRNATAP | -0.12 | 0.00824 | NA | |
| 31 | hsa-miR-335-3p | CCNG2 | -0.24 | 0.8845 | -0.11 | 0.92934 | MirTarget; mirMAP | -0.16 | 1.0E-5 | NA | |
| 32 | hsa-miR-503-5p | CCNG2 | 0.18 | 0.7664 | -0.11 | 0.92934 | miRNAWalker2 validate | -0.1 | 0.00027 | NA | |
| 33 | hsa-miR-93-5p | CCNG2 | -0.61 | 0.8253 | -0.11 | 0.92934 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.14 | 0.00129 | NA | |
| 34 | hsa-miR-23b-3p | CDK2 | -0.11 | 0.96135 | -0.01 | 0.99411 | miRNAWalker2 validate | -0.17 | 0.00275 | NA | |
| 35 | hsa-miR-195-5p | CDK4 | 0.34 | 0.74962 | -0.06 | 0.97429 | miRNAWalker2 validate; miRTarBase | -0.18 | 0 | NA | |
| 36 | hsa-miR-34c-5p | CDK4 | -0.02 | 0.95279 | -0.06 | 0.97429 | miRNAWalker2 validate; miRTarBase | -0.11 | 0.00308 | NA | |
| 37 | hsa-miR-21-5p | CDK6 | -0.15 | 0.97024 | 0.26 | 0.8515 | miRNAWalker2 validate; mirMAP | -0.32 | 5.0E-5 | NA | |
| 38 | hsa-miR-22-3p | CDK6 | 0.06 | 0.98656 | 0.26 | 0.8515 | miRNAWalker2 validate | -0.27 | 0.00121 | NA | |
| 39 | hsa-let-7c-5p | CDKN1A | -0.06 | 0.9749 | 0.1 | 0.95314 | MirTarget | -0.15 | 0.0033 | NA | |
| 40 | hsa-miR-101-3p | CDKN1A | 0.01 | 0.99704 | 0.1 | 0.95314 | MirTarget | -0.4 | 4.0E-5 | NA | |
| 41 | hsa-miR-708-5p | CDKN1A | -0.03 | 0.97493 | 0.1 | 0.95314 | MirTarget | -0.14 | 0.00964 | NA | |
| 42 | hsa-let-7g-5p | CDKN2A | -0.2 | 0.92299 | 1.38 | 0.01803 | miRNAWalker2 validate; miRTarBase | -0.54 | 0.00297 | NA | |
| 43 | hsa-miR-139-5p | CHEK1 | -0.08 | 0.92869 | -0.28 | 0.78554 | miRanda | -0.13 | 0.00213 | NA | |
| 44 | hsa-miR-195-5p | CHEK1 | 0.34 | 0.74962 | -0.28 | 0.78554 | MirTarget; miRNATAP | -0.16 | 0.00025 | 25840419 | MiR 195 suppresses non small cell lung cancer by targeting CHEK1; We discovered that CHEK1 was a direct target of miR-195 which decreased CHEK1 expression in lung cancer cells |
| 45 | hsa-miR-497-5p | CHEK1 | -0.01 | 0.98915 | -0.28 | 0.78554 | MirTarget; miRNATAP | -0.21 | 5.0E-5 | 24464213 | Checkpoint kinase 1 is negatively regulated by miR 497 in hepatocellular carcinoma; In silico analysis showed that CHEK1 was a candidate target of miR-497 which was previously found to be downregulated in HCC by us; To test whether miR-497 could bind to 3'untranslated region 3'UTR of CHEK1 luciferase reporter assay was conducted; The result revealed that miR-497 could bind to the 3'untranslated region 3'UTR of CHEK1 mRNA; Western blot showed that ectopic expression of miR-497 suppressed the CHEK1 expression and inhibition of miR-497 led to significant upregulation of CHEK1; Finally miR-497 expression was measured in the same 30 HCC samples and the correlation between miR-497 and CHEK1 was analyzed; The results indicated that miR-497 was downregulated in HCC and had a significant negative correlation with CHEK1; Taken together these results demonstrated that CHEK1 was negatively regulated by miR-497 and the overexpressed CHEK1 was resulted from the downregulated miR-497 in HCC which provided a potential molecular target for HCC therapy |
| 46 | hsa-miR-139-5p | CYCS | -0.08 | 0.92869 | -0.08 | 0.96574 | miRanda | -0.18 | 7.0E-5 | NA | |
| 47 | hsa-miR-34c-3p | CYCS | -0.07 | 0.82545 | -0.08 | 0.96574 | mirMAP | -0.1 | 0.008 | NA | |
| 48 | hsa-miR-34c-5p | CYCS | -0.02 | 0.95279 | -0.08 | 0.96574 | miRanda | -0.16 | 0.00033 | NA | |
| 49 | hsa-miR-106a-5p | FAS | -0.2 | 0.80221 | -0.35 | 0.70281 | miRNAWalker2 validate; miRTarBase | -0.31 | 0 | 22431000; 27142596 | miR 106a is frequently upregulated in gastric cancer and inhibits the extrinsic apoptotic pathway by targeting FAS; Bioinformatic analysis combining with validation experiments identified FAS as a direct target of miR-106a; Moreover a significant inverse correlation was found between miR-106a and FAS expression not only in gastric cancer cell lines but also in gastric cancer specimens; Taken together these findings suggest that ectopicly overexpressed miR-106a may play an oncogenic role in gastric carcinogenesis and impair extrinsic apoptotic pathway through targeting FAS;Functional experiment ascertained that miR-106a interacted with FAS and mediated caspase3 pathway |
| 50 | hsa-miR-361-5p | FAS | -0.14 | 0.9415 | -0.35 | 0.70281 | miRanda | -0.42 | 0.00016 | NA | |
| 51 | hsa-miR-590-5p | FAS | -0.55 | 0.47274 | -0.35 | 0.70281 | miRanda | -0.17 | 0.00403 | NA | |
| 52 | hsa-miR-98-5p | FAS | -0.17 | 0.8988 | -0.35 | 0.70281 | miRNAWalker2 validate | -0.38 | 2.0E-5 | NA | |
| 53 | hsa-miR-324-3p | GADD45B | -0.42 | 0.66153 | 0.52 | 0.61353 | MirTarget; miRNATAP | -0.22 | 0.00285 | NA | |
| 54 | hsa-miR-590-3p | GADD45B | -0.28 | 0.59127 | 0.52 | 0.61353 | miRanda | -0.23 | 0 | NA | |
| 55 | hsa-miR-181c-5p | GTSE1 | 0.04 | 0.96916 | -0.25 | 0.80831 | MirTarget | -0.2 | 4.0E-5 | NA | |
| 56 | hsa-let-7a-3p | IGF1 | -0.22 | 0.85543 | 0.01 | 0.98747 | mirMAP | -1.08 | 0 | NA | |
| 57 | hsa-let-7f-1-3p | IGF1 | -0.31 | 0.69341 | 0.01 | 0.98747 | mirMAP | -1.34 | 0 | NA | |
| 58 | hsa-miR-1275 | IGF1 | -0.83 | 0.08038 | 0.01 | 0.98747 | MirTarget; PITA | -0.38 | 5.0E-5 | NA | |
| 59 | hsa-miR-130b-3p | IGF1 | -0.22 | 0.82466 | 0.01 | 0.98747 | MirTarget | -1.33 | 0 | NA | |
| 60 | hsa-miR-148a-5p | IGF1 | -0.45 | 0.74842 | 0.01 | 0.98747 | mirMAP | -0.73 | 0 | NA | |
| 61 | hsa-miR-15b-3p | IGF1 | -0.56 | 0.60918 | 0.01 | 0.98747 | mirMAP | -1.37 | 0 | NA | |
| 62 | hsa-miR-16-1-3p | IGF1 | 0.15 | 0.73374 | 0.01 | 0.98747 | mirMAP | -0.87 | 0 | NA | |
| 63 | hsa-miR-186-5p | IGF1 | -0.32 | 0.85413 | 0.01 | 0.98747 | mirMAP | -1.35 | 0 | NA | |
| 64 | hsa-miR-19a-3p | IGF1 | -0.21 | 0.84464 | 0.01 | 0.98747 | MirTarget | -0.99 | 0 | NA | |
| 65 | hsa-miR-19b-1-5p | IGF1 | -0.01 | 0.98851 | 0.01 | 0.98747 | mirMAP | -0.96 | 0 | NA | |
| 66 | hsa-miR-19b-3p | IGF1 | -0.03 | 0.98666 | 0.01 | 0.98747 | MirTarget | -1.19 | 0 | NA | |
| 67 | hsa-miR-20a-3p | IGF1 | 0.46 | 0.57674 | 0.01 | 0.98747 | mirMAP | -0.83 | 0 | NA | |
| 68 | hsa-miR-224-3p | IGF1 | 0.11 | 0.74909 | 0.01 | 0.98747 | mirMAP | -0.32 | 0.00753 | NA | |
| 69 | hsa-miR-26b-5p | IGF1 | -0.02 | 0.99038 | 0.01 | 0.98747 | mirMAP | -1.33 | 0 | NA | |
| 70 | hsa-miR-27a-3p | IGF1 | -0.12 | 0.9588 | 0.01 | 0.98747 | miRNAWalker2 validate; miRTarBase | -1.33 | 0 | NA | |
| 71 | hsa-miR-29a-3p | IGF1 | 0.01 | 0.99698 | 0.01 | 0.98747 | MirTarget | -1.22 | 0 | NA | |
| 72 | hsa-miR-29b-3p | IGF1 | -0.1 | 0.95899 | 0.01 | 0.98747 | MirTarget | -1.13 | 0 | 25592039 | Luciferase reporter assays were conducted to determine the association between miR-29b and the insulin-like growth factor 1 IGF1 3' untranslated region 3'UTR; IGF1 an activator of PI3K/Akt signaling was confirmed as a novel target of miR-29b |
| 73 | hsa-miR-301a-3p | IGF1 | -0.11 | 0.83169 | 0.01 | 0.98747 | MirTarget | -0.72 | 0 | NA | |
| 74 | hsa-miR-32-3p | IGF1 | -0.57 | 0.13133 | 0.01 | 0.98747 | mirMAP | -0.85 | 0 | NA | |
| 75 | hsa-miR-320a | IGF1 | -0.42 | 0.8402 | 0.01 | 0.98747 | miRNATAP | -0.53 | 0.00309 | NA | |
| 76 | hsa-miR-33a-3p | IGF1 | -0.79 | 0.01052 | 0.01 | 0.98747 | MirTarget | -0.71 | 0 | NA | |
| 77 | hsa-miR-361-5p | IGF1 | -0.14 | 0.9415 | 0.01 | 0.98747 | PITA; mirMAP | -0.76 | 0.00313 | NA | |
| 78 | hsa-miR-362-5p | IGF1 | -0.27 | 0.75202 | 0.01 | 0.98747 | mirMAP | -0.69 | 0 | NA | |
| 79 | hsa-miR-369-3p | IGF1 | 0.12 | 0.8323 | 0.01 | 0.98747 | mirMAP | -0.42 | 0.00732 | NA | |
| 80 | hsa-miR-374b-3p | IGF1 | -0.05 | 0.86075 | 0.01 | 0.98747 | mirMAP | -0.9 | 0 | NA | |
| 81 | hsa-miR-376b-3p | IGF1 | -0.01 | 0.98243 | 0.01 | 0.98747 | mirMAP | -0.4 | 0.00219 | NA | |
| 82 | hsa-miR-377-3p | IGF1 | -0.04 | 0.89351 | 0.01 | 0.98747 | mirMAP | -0.35 | 0.00945 | NA | |
| 83 | hsa-miR-421 | IGF1 | -0.18 | 0.7347 | 0.01 | 0.98747 | PITA | -0.68 | 0 | NA | |
| 84 | hsa-miR-450b-5p | IGF1 | -0.01 | 0.98315 | 0.01 | 0.98747 | MirTarget; PITA; mirMAP; miRNATAP | -0.44 | 0.00527 | NA | |
| 85 | hsa-miR-452-5p | IGF1 | -0.05 | 0.97387 | 0.01 | 0.98747 | MirTarget; mirMAP | -0.39 | 0.00722 | NA | |
| 86 | hsa-miR-454-3p | IGF1 | -0.1 | 0.84355 | 0.01 | 0.98747 | MirTarget | -1.21 | 0 | NA | |
| 87 | hsa-miR-486-5p | IGF1 | -0.55 | 0.6964 | 0.01 | 0.98747 | PITA; miRNATAP | -0.28 | 0.00602 | NA | |
| 88 | hsa-miR-576-5p | IGF1 | -0.51 | 0.41719 | 0.01 | 0.98747 | PITA; mirMAP; miRNATAP | -1.08 | 0 | NA | |
| 89 | hsa-miR-577 | IGF1 | -1.06 | 0.32606 | 0.01 | 0.98747 | PITA | -0.61 | 0 | NA | |
| 90 | hsa-miR-590-3p | IGF1 | -0.28 | 0.59127 | 0.01 | 0.98747 | MirTarget; miRanda; mirMAP; miRNATAP | -0.9 | 0 | NA | |
| 91 | hsa-miR-592 | IGF1 | -0.18 | 0.85623 | 0.01 | 0.98747 | mirMAP | -0.18 | 0.00559 | NA | |
| 92 | hsa-miR-629-5p | IGF1 | -0.39 | 0.79854 | 0.01 | 0.98747 | mirMAP | -0.58 | 3.0E-5 | NA | |
| 93 | hsa-miR-940 | IGF1 | -0.23 | 0.68006 | 0.01 | 0.98747 | MirTarget; PITA; miRNATAP | -0.57 | 0 | NA | |
| 94 | hsa-let-7a-3p | IGFBP3 | -0.22 | 0.85543 | 0.57 | 0.72672 | miRNATAP | -0.43 | 0 | NA | |
| 95 | hsa-miR-197-3p | IGFBP3 | -0.46 | 0.79492 | 0.57 | 0.72672 | MirTarget; miRNATAP | -0.35 | 2.0E-5 | NA | |
| 96 | hsa-miR-19a-3p | IGFBP3 | -0.21 | 0.84464 | 0.57 | 0.72672 | MirTarget; miRNATAP | -0.41 | 0 | NA | |
| 97 | hsa-miR-19b-3p | IGFBP3 | -0.03 | 0.98666 | 0.57 | 0.72672 | MirTarget; miRNATAP | -0.48 | 0 | NA | |
| 98 | hsa-miR-224-5p | IGFBP3 | -0.24 | 0.85735 | 0.57 | 0.72672 | MirTarget | -0.3 | 0 | NA | |
| 99 | hsa-miR-339-5p | IGFBP3 | -0.3 | 0.71291 | 0.57 | 0.72672 | miRanda | -0.32 | 0 | NA | |
| 100 | hsa-miR-374a-5p | IGFBP3 | -0.34 | 0.76692 | 0.57 | 0.72672 | MirTarget; mirMAP | -0.5 | 0 | NA | |
| 101 | hsa-miR-374b-5p | IGFBP3 | -0.29 | 0.8357 | 0.57 | 0.72672 | MirTarget | -0.54 | 0 | NA | |
| 102 | hsa-miR-375 | IGFBP3 | -0.69 | 0.83172 | 0.57 | 0.72672 | miRNAWalker2 validate | -0.17 | 0.00046 | NA | |
| 103 | hsa-miR-590-3p | IGFBP3 | -0.28 | 0.59127 | 0.57 | 0.72672 | miRanda | -0.33 | 0 | NA | |
| 104 | hsa-miR-590-5p | IGFBP3 | -0.55 | 0.47274 | 0.57 | 0.72672 | miRanda | -0.42 | 0 | NA | |
| 105 | hsa-miR-143-3p | MDM2 | 0.37 | 0.92734 | -0.49 | 0.72349 | miRNAWalker2 validate | -0.16 | 0.00124 | NA | |
| 106 | hsa-let-7b-5p | MDM4 | -0.23 | 0.93895 | 0.25 | 0.82318 | miRNAWalker2 validate; MirTarget | -0.14 | 0.00444 | NA | |
| 107 | hsa-miR-150-5p | PERP | -0.38 | 0.83879 | 0.23 | 0.91127 | MirTarget | -0.17 | 0 | NA | |
| 108 | hsa-miR-629-3p | PERP | -0.5 | 0.33397 | 0.23 | 0.91127 | MirTarget | -0.12 | 0.00543 | NA | |
| 109 | hsa-miR-200b-3p | PMAIP1 | -0.43 | 0.86396 | 0.02 | 0.98032 | MirTarget; TargetScan | -0.15 | 0.00941 | NA | |
| 110 | hsa-miR-195-5p | PPM1D | 0.34 | 0.74962 | -0.19 | 0.83465 | MirTarget; miRNATAP | -0.1 | 0.00097 | NA | |
| 111 | hsa-miR-26b-5p | PPM1D | -0.02 | 0.99038 | -0.19 | 0.83465 | miRNAWalker2 validate | -0.12 | 0.00263 | NA | |
| 112 | hsa-miR-29a-3p | PPM1D | 0.01 | 0.99698 | -0.19 | 0.83465 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.16 | 5.0E-5 | NA | |
| 113 | hsa-miR-29b-3p | PPM1D | -0.1 | 0.95899 | -0.19 | 0.83465 | MirTarget; miRNATAP | -0.11 | 0.00016 | NA | |
| 114 | hsa-miR-106a-5p | PTEN | -0.2 | 0.80221 | -0.27 | 0.8446 | miRNATAP | -0.14 | 0.00145 | 26097565; 26318586 | miR 106a promotes growth and metastasis of non small cell lung cancer by targeting PTEN; Furthermore the presence of miR-106a was inversely correlated with PTEN in NSCLC tissues; Overall this study suggested that miR-106a inhibited the growth and metastasis of NSCLC cells by decreasing PTEN expression;Further pterostilbene through downregulation of miR-17-5p and miR-106a-5p expression both in tumors and systemic circulation rescued PTEN mRNA and protein levels leading to reduced tumor growth in vivo |
| 115 | hsa-miR-188-5p | PTEN | -0.57 | 0.32482 | -0.27 | 0.8446 | MirTarget; PITA; miRNATAP | -0.1 | 0.00127 | NA | |
| 116 | hsa-miR-29b-3p | PTEN | -0.1 | 0.95899 | -0.27 | 0.8446 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.12 | 0.0055 | 26471361; 23179556; 26063204; 21359530 | Mechanistically ATDC exerted its oncogenic effects by suppressing miR-29 and subsequent upregulation of DNMT3A leading to DNA methylation and silencing of the tumor suppressor PTEN;Anticancer role of MUC1 aptamer miR 29b chimera in epithelial ovarian carcinoma cells through regulation of PTEN methylation; Our study indicated that Chi-29b chimera can effectively exert antitumor effect through specific delivery of miR-29b into OVCAR-3 tumor cells subsequently reexpressing PTEN gene and inducing cell apoptosis;Furthermore the dual-luciferase reporter assay demonstrated that miR-29b inhibited the expression of the luciferase gene containing the 3'-UTRs of MMP2 and PTEN mRNA;In contrast enhanced miR-29b expression by transfection with pre-miR-29b decreased the expression of PTEN and impaired apoptosis increasing tumor cell migration and invasion; Moreover PTEN was shown to be a direct target of miR-29b and was also shown to contribute to the miR-29b-mediated effects on cell invasion; Modulation of miR-29b altered the role of PTEN involved in cell migration and invasion; Aberrant expression of miR-29b which modulates PTEN expression can contribute to migration invasion and anti-apoptosis |
| 117 | hsa-miR-146b-5p | RFWD2 | -0.4 | 0.83751 | 0.13 | 0.92354 | miRNATAP | -0.12 | 4.0E-5 | NA | |
| 118 | hsa-miR-338-3p | RPRM | 0.49 | 0.78848 | 0.53 | 0.54344 | miRanda | -0.36 | 0.00289 | NA | |
| 119 | hsa-miR-375 | RPRM | -0.69 | 0.83172 | 0.53 | 0.54344 | miRanda | -0.5 | 0 | NA | |
| 120 | hsa-miR-501-3p | RPRM | -0.75 | 0.55276 | 0.53 | 0.54344 | miRNATAP | -0.62 | 4.0E-5 | NA | |
| 121 | hsa-miR-100-5p | RRM2 | 0.02 | 0.99282 | -0.25 | 0.87456 | miRNAWalker2 validate | -0.25 | 0 | NA | |
| 122 | hsa-miR-125a-5p | RRM2 | -0 | 0.99916 | -0.25 | 0.87456 | miRanda | -0.27 | 9.0E-5 | NA | |
| 123 | hsa-miR-199a-5p | RRM2 | 0.16 | 0.9358 | -0.25 | 0.87456 | miRanda | -0.24 | 4.0E-5 | NA | |
| 124 | hsa-miR-199b-5p | RRM2 | -0.04 | 0.97717 | -0.25 | 0.87456 | miRanda | -0.15 | 0.00321 | NA | |
| 125 | hsa-miR-217 | RRM2 | 0.34 | 0.761 | -0.25 | 0.87456 | miRanda | -0.2 | 0 | NA | |
| 126 | hsa-miR-26a-5p | RRM2 | 0.01 | 0.99772 | -0.25 | 0.87456 | miRNAWalker2 validate | -0.42 | 3.0E-5 | NA | |
| 127 | hsa-miR-30a-5p | RRM2 | 0.22 | 0.93395 | -0.25 | 0.87456 | miRNAWalker2 validate | -0.5 | 0 | NA | |
| 128 | hsa-miR-140-5p | RRM2B | -0.23 | 0.84733 | -0.42 | 0.72245 | miRanda | -0.18 | 0.00211 | NA | |
| 129 | hsa-miR-142-3p | RRM2B | -0.15 | 0.9461 | -0.42 | 0.72245 | miRNAWalker2 validate; miRanda | -0.15 | 0 | NA | |
| 130 | hsa-miR-30e-5p | RRM2B | -0.07 | 0.97968 | -0.42 | 0.72245 | mirMAP | -0.19 | 0.00223 | NA | |
| 131 | hsa-miR-369-3p | RRM2B | 0.12 | 0.8323 | -0.42 | 0.72245 | mirMAP | -0.11 | 0.00769 | NA | |
| 132 | hsa-miR-495-3p | RRM2B | 0 | 0.99753 | -0.42 | 0.72245 | mirMAP | -0.17 | 0.0005 | NA | |
| 133 | hsa-miR-7-1-3p | RRM2B | -0.46 | 0.6659 | -0.42 | 0.72245 | mirMAP | -0.16 | 1.0E-5 | NA | |
| 134 | hsa-miR-335-5p | SERPINB5 | -0.03 | 0.97338 | 1 | 0.44591 | miRNAWalker2 validate | -0.58 | 0.00012 | NA | |
| 135 | hsa-miR-148a-3p | SERPINE1 | -0.1 | 0.97698 | 0.97 | 0.43183 | miRNATAP | -0.63 | 0 | NA | |
| 136 | hsa-miR-148a-5p | SERPINE1 | -0.45 | 0.74842 | 0.97 | 0.43183 | miRNATAP | -0.49 | 0 | NA | |
| 137 | hsa-miR-2110 | SERPINE1 | -0.32 | 0.25098 | 0.97 | 0.43183 | miRNATAP | -0.26 | 0.00814 | NA | |
| 138 | hsa-miR-301a-3p | SERPINE1 | -0.11 | 0.83169 | 0.97 | 0.43183 | miRNAWalker2 validate; miRTarBase | -0.4 | 1.0E-5 | NA | |
| 139 | hsa-miR-30b-5p | SERPINE1 | -0.01 | 0.99462 | 0.97 | 0.43183 | MirTarget | -0.51 | 0 | 25170877 | miR-30b may function as a novel tumor suppressor gene in gastric cancer by targeting PAI-1 and regulating the apoptosis of cancer cells |
| 140 | hsa-miR-30c-5p | SERPINE1 | -0.3 | 0.86581 | 0.97 | 0.43183 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.66 | 0 | NA | |
| 141 | hsa-miR-30d-5p | SERPINE1 | -0.17 | 0.95173 | 0.97 | 0.43183 | miRNATAP | -0.62 | 0 | NA | |
| 142 | hsa-miR-378a-5p | SERPINE1 | -0.56 | 0.72488 | 0.97 | 0.43183 | MirTarget | -0.55 | 0 | NA | |
| 143 | hsa-miR-421 | SERPINE1 | -0.18 | 0.7347 | 0.97 | 0.43183 | miRanda | -0.42 | 2.0E-5 | NA | |
| 144 | hsa-miR-425-5p | SERPINE1 | -0.35 | 0.84598 | 0.97 | 0.43183 | MirTarget; miRNATAP | -0.54 | 0 | NA | |
| 145 | hsa-miR-452-3p | SERPINE1 | -0.17 | 0.79901 | 0.97 | 0.43183 | mirMAP | -0.39 | 0 | NA | |
| 146 | hsa-miR-514a-3p | SERPINE1 | 0.23 | 0.69823 | 0.97 | 0.43183 | MirTarget | -0.14 | 0.00373 | NA | |
| 147 | hsa-miR-590-3p | SERPINE1 | -0.28 | 0.59127 | 0.97 | 0.43183 | miRanda | -0.43 | 0 | NA | |
| 148 | hsa-miR-629-3p | SERPINE1 | -0.5 | 0.33397 | 0.97 | 0.43183 | MirTarget | -0.28 | 0.00066 | NA | |
| 149 | hsa-miR-769-5p | SERPINE1 | -0.31 | 0.7357 | 0.97 | 0.43183 | MirTarget; miRNATAP | -0.6 | 1.0E-5 | NA | |
| 150 | hsa-miR-942-5p | SERPINE1 | -0.51 | 0.50778 | 0.97 | 0.43183 | MirTarget | -0.45 | 0 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | REGULATION OF CELL CYCLE | 25 | 949 | 6.983e-23 | 3.249e-19 |
| 2 | CELL CYCLE PROCESS | 24 | 1081 | 4.194e-20 | 9.757e-17 |
| 3 | CELL CYCLE | 25 | 1316 | 1.973e-19 | 2.295e-16 |
| 4 | NEGATIVE REGULATION OF CELL CYCLE | 18 | 433 | 1.689e-19 | 2.295e-16 |
| 5 | MITOTIC CELL CYCLE | 21 | 766 | 3.405e-19 | 3.169e-16 |
| 6 | REGULATION OF CELL DEATH | 24 | 1472 | 5.18e-17 | 4.017e-14 |
| 7 | POSITIVE REGULATION OF CELL DEATH | 18 | 605 | 6.31e-17 | 4.194e-14 |
| 8 | RESPONSE TO ABIOTIC STIMULUS | 21 | 1024 | 1.239e-16 | 7.205e-14 |
| 9 | REGULATION OF CELL CYCLE ARREST | 11 | 108 | 2.514e-16 | 1.3e-13 |
| 10 | REGULATION OF CELL PROLIFERATION | 23 | 1496 | 1.195e-15 | 5.559e-13 |
| 11 | SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 11 | 127 | 1.577e-15 | 6.672e-13 |
| 12 | POSITIVE REGULATION OF CELL CYCLE PROCESS | 13 | 247 | 2.104e-15 | 8.158e-13 |
| 13 | CELL CYCLE PHASE TRANSITION | 13 | 255 | 3.182e-15 | 1.139e-12 |
| 14 | CELL CYCLE CHECKPOINT | 12 | 194 | 4.223e-15 | 1.404e-12 |
| 15 | NEGATIVE REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 10 | 98 | 6.438e-15 | 1.997e-12 |
| 16 | NEGATIVE REGULATION OF CELL CYCLE PROCESS | 12 | 214 | 1.378e-14 | 4.007e-12 |
| 17 | REGULATION OF MITOTIC CELL CYCLE | 15 | 468 | 1.529e-14 | 4.186e-12 |
| 18 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 22 | 1492 | 1.641e-14 | 4.243e-12 |
| 19 | REGULATION OF PROTEOLYSIS | 17 | 711 | 2.126e-14 | 5.207e-12 |
| 20 | CELL DEATH | 19 | 1001 | 2.458e-14 | 5.719e-12 |
| 21 | G1 DNA DAMAGE CHECKPOINT | 9 | 73 | 2.794e-14 | 6.191e-12 |
| 22 | RESPONSE TO OXYGEN LEVELS | 13 | 311 | 4.122e-14 | 8.718e-12 |
| 23 | CELLULAR RESPONSE TO STRESS | 22 | 1565 | 4.387e-14 | 8.875e-12 |
| 24 | POSITIVE REGULATION OF CELL CYCLE | 13 | 332 | 9.52e-14 | 1.846e-11 |
| 25 | POSITIVE REGULATION OF CELL CYCLE ARREST | 9 | 85 | 1.164e-13 | 2.167e-11 |
| 26 | NEGATIVE REGULATION OF MITOTIC CELL CYCLE | 11 | 199 | 2.344e-13 | 4.195e-11 |
| 27 | POSITIVE REGULATION OF PROTEOLYSIS | 13 | 363 | 2.973e-13 | 5.123e-11 |
| 28 | ACTIVATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 9 | 95 | 3.273e-13 | 5.439e-11 |
| 29 | NEGATIVE REGULATION OF CELL CYCLE PHASE TRANSITION | 10 | 146 | 3.792e-13 | 5.882e-11 |
| 30 | DNA INTEGRITY CHECKPOINT | 10 | 146 | 3.792e-13 | 5.882e-11 |
| 31 | REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 10 | 147 | 4.063e-13 | 6.098e-11 |
| 32 | APOPTOTIC SIGNALING PATHWAY | 12 | 289 | 4.958e-13 | 6.991e-11 |
| 33 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 11 | 213 | 4.946e-13 | 6.991e-11 |
| 34 | MITOTIC DNA INTEGRITY CHECKPOINT | 9 | 100 | 5.259e-13 | 7.198e-11 |
| 35 | REGULATION OF PEPTIDASE ACTIVITY | 13 | 392 | 7.888e-13 | 1.049e-10 |
| 36 | CELL CYCLE G1 S PHASE TRANSITION | 9 | 111 | 1.374e-12 | 1.728e-10 |
| 37 | G1 S TRANSITION OF MITOTIC CELL CYCLE | 9 | 111 | 1.374e-12 | 1.728e-10 |
| 38 | ZYMOGEN ACTIVATION | 9 | 112 | 1.492e-12 | 1.827e-10 |
| 39 | REGULATION OF CELL CYCLE PHASE TRANSITION | 12 | 321 | 1.711e-12 | 2.042e-10 |
| 40 | REGULATION OF CELL CYCLE PROCESS | 14 | 558 | 3.798e-12 | 4.418e-10 |
| 41 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 12 | 363 | 7.229e-12 | 8.204e-10 |
| 42 | CELLULAR RESPONSE TO DNA DAMAGE STIMULUS | 15 | 720 | 7.645e-12 | 8.469e-10 |
| 43 | MITOTIC CELL CYCLE CHECKPOINT | 9 | 139 | 1.069e-11 | 1.157e-09 |
| 44 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 7 | 55 | 2.005e-11 | 2.12e-09 |
| 45 | SIGNAL TRANSDUCTION IN RESPONSE TO DNA DAMAGE | 8 | 96 | 2.16e-11 | 2.234e-09 |
| 46 | INTRINSIC APOPTOTIC SIGNALING PATHWAY | 9 | 152 | 2.399e-11 | 2.426e-09 |
| 47 | POSITIVE REGULATION OF PEPTIDASE ACTIVITY | 9 | 154 | 2.699e-11 | 2.672e-09 |
| 48 | REGULATION OF TRANSFERASE ACTIVITY | 16 | 946 | 2.894e-11 | 2.805e-09 |
| 49 | REGULATION OF PROTEIN MODIFICATION PROCESS | 20 | 1710 | 3.197e-11 | 3.036e-09 |
| 50 | REGULATION OF SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 9 | 162 | 4.257e-11 | 3.962e-09 |
| 51 | RESPONSE TO DRUG | 12 | 431 | 5.31e-11 | 4.844e-09 |
| 52 | POSITIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 9 | 171 | 6.916e-11 | 6.189e-09 |
| 53 | INTRACELLULAR SIGNAL TRANSDUCTION | 19 | 1572 | 7.155e-11 | 6.282e-09 |
| 54 | RESPONSE TO LIPID | 15 | 888 | 1.466e-10 | 1.263e-08 |
| 55 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 16 | 1087 | 2.268e-10 | 1.919e-08 |
| 56 | RESPONSE TO STEROID HORMONE | 12 | 497 | 2.73e-10 | 2.268e-08 |
| 57 | NEGATIVE REGULATION OF CELL PROLIFERATION | 13 | 643 | 3.761e-10 | 3.07e-08 |
| 58 | RESPONSE TO ESTROGEN | 9 | 218 | 6.009e-10 | 4.821e-08 |
| 59 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 8 | 153 | 9.239e-10 | 7.286e-08 |
| 60 | CELL CYCLE ARREST | 8 | 154 | 9.731e-10 | 7.546e-08 |
| 61 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 18 | 1618 | 1.08e-09 | 8.239e-08 |
| 62 | REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 7 | 97 | 1.19e-09 | 8.93e-08 |
| 63 | RESPONSE TO METAL ION | 10 | 333 | 1.313e-09 | 9.696e-08 |
| 64 | REGENERATION | 8 | 161 | 1.385e-09 | 1.007e-07 |
| 65 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 18 | 1656 | 1.569e-09 | 1.123e-07 |
| 66 | CELL DIVISION | 11 | 460 | 1.904e-09 | 1.342e-07 |
| 67 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 11 | 470 | 2.384e-09 | 1.656e-07 |
| 68 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 14 | 917 | 2.617e-09 | 1.791e-07 |
| 69 | RESPONSE TO ALCOHOL | 10 | 362 | 2.932e-09 | 1.977e-07 |
| 70 | CELLULAR RESPONSE TO ABIOTIC STIMULUS | 9 | 263 | 3.13e-09 | 2.081e-07 |
| 71 | CELLULAR RESPONSE TO EXTERNAL STIMULUS | 9 | 264 | 3.236e-09 | 2.091e-07 |
| 72 | AGING | 9 | 264 | 3.236e-09 | 2.091e-07 |
| 73 | PROTEIN MATURATION | 9 | 265 | 3.345e-09 | 2.132e-07 |
| 74 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 17 | 1518 | 3.444e-09 | 2.166e-07 |
| 75 | REGULATION OF KINASE ACTIVITY | 13 | 776 | 3.68e-09 | 2.283e-07 |
| 76 | RESPONSE TO X RAY | 5 | 30 | 4.376e-09 | 2.679e-07 |
| 77 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 18 | 1791 | 5.485e-09 | 3.315e-07 |
| 78 | RESPONSE TO EXTERNAL STIMULUS | 18 | 1821 | 7.138e-09 | 4.258e-07 |
| 79 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 16 | 1381 | 7.32e-09 | 4.311e-07 |
| 80 | RESPONSE TO UV | 7 | 126 | 7.48e-09 | 4.35e-07 |
| 81 | NEGATIVE REGULATION OF CATALYTIC ACTIVITY | 13 | 829 | 8.123e-09 | 4.66e-07 |
| 82 | REPLICATIVE SENESCENCE | 4 | 12 | 8.213e-09 | 4.66e-07 |
| 83 | NEURON APOPTOTIC PROCESS | 5 | 35 | 9.892e-09 | 5.546e-07 |
| 84 | RESPONSE TO RADIATION | 10 | 413 | 1.032e-08 | 5.705e-07 |
| 85 | DNA REPLICATION | 8 | 208 | 1.042e-08 | 5.705e-07 |
| 86 | NEGATIVE REGULATION OF CELL DEATH | 13 | 872 | 1.483e-08 | 8.025e-07 |
| 87 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 13 | 876 | 1.566e-08 | 8.376e-07 |
| 88 | REGULATION OF PROTEIN STABILITY | 8 | 221 | 1.673e-08 | 8.846e-07 |
| 89 | REGULATION OF FIBROBLAST PROLIFERATION | 6 | 81 | 1.707e-08 | 8.924e-07 |
| 90 | RESPONSE TO IONIZING RADIATION | 7 | 145 | 1.986e-08 | 1.015e-06 |
| 91 | ORGAN REGENERATION | 6 | 83 | 1.979e-08 | 1.015e-06 |
| 92 | RESPONSE TO ESTRADIOL | 7 | 146 | 2.083e-08 | 1.042e-06 |
| 93 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 14 | 1079 | 2.078e-08 | 1.042e-06 |
| 94 | RESPONSE TO TOXIC SUBSTANCE | 8 | 241 | 3.281e-08 | 1.624e-06 |
| 95 | RESPONSE TO INORGANIC SUBSTANCE | 10 | 479 | 4.185e-08 | 2.05e-06 |
| 96 | NEURON DEATH | 5 | 47 | 4.588e-08 | 2.224e-06 |
| 97 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 6 | 98 | 5.393e-08 | 2.587e-06 |
| 98 | NEGATIVE REGULATION OF CELL CYCLE ARREST | 4 | 20 | 7.941e-08 | 3.77e-06 |
| 99 | POSITIVE REGULATION OF FIBROBLAST PROLIFERATION | 5 | 53 | 8.505e-08 | 3.957e-06 |
| 100 | INTRINSIC APOPTOTIC SIGNALING PATHWAY BY P53 CLASS MEDIATOR | 5 | 53 | 8.505e-08 | 3.957e-06 |
| 101 | RESPONSE TO ENDOGENOUS STIMULUS | 15 | 1450 | 1.137e-07 | 5.237e-06 |
| 102 | RESPONSE TO NITROGEN COMPOUND | 12 | 859 | 1.227e-07 | 5.598e-06 |
| 103 | REGULATION OF RESPONSE TO STRESS | 15 | 1468 | 1.336e-07 | 6.036e-06 |
| 104 | REGULATION OF CELL CYCLE G2 M PHASE TRANSITION | 5 | 59 | 1.47e-07 | 6.577e-06 |
| 105 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 7 | 200 | 1.806e-07 | 8.002e-06 |
| 106 | RESPONSE TO HORMONE | 12 | 893 | 1.868e-07 | 8.199e-06 |
| 107 | HISTONE PHOSPHORYLATION | 4 | 25 | 2.058e-07 | 8.948e-06 |
| 108 | POSITIVE REGULATION OF MITOTIC CELL CYCLE | 6 | 123 | 2.095e-07 | 9.024e-06 |
| 109 | RESPONSE TO EXTRACELLULAR STIMULUS | 9 | 441 | 2.656e-07 | 1.134e-05 |
| 110 | CELL AGING | 5 | 67 | 2.801e-07 | 1.185e-05 |
| 111 | PROTEIN STABILIZATION | 6 | 131 | 3.043e-07 | 1.275e-05 |
| 112 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 13 | 1135 | 3.224e-07 | 1.339e-05 |
| 113 | REGULATION OF DNA DAMAGE RESPONSE SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 4 | 28 | 3.315e-07 | 1.365e-05 |
| 114 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 14 | 1360 | 3.67e-07 | 1.498e-05 |
| 115 | CELL CYCLE G2 M PHASE TRANSITION | 6 | 138 | 4.138e-07 | 1.674e-05 |
| 116 | NEGATIVE REGULATION OF B CELL ACTIVATION | 4 | 30 | 4.424e-07 | 1.759e-05 |
| 117 | NEGATIVE REGULATION OF CELL MATRIX ADHESION | 4 | 30 | 4.424e-07 | 1.759e-05 |
| 118 | CELLULAR RESPONSE TO OXYGEN LEVELS | 6 | 143 | 5.103e-07 | 2.012e-05 |
| 119 | NEGATIVE REGULATION OF CELL COMMUNICATION | 13 | 1192 | 5.643e-07 | 2.207e-05 |
| 120 | NEGATIVE REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 4 | 32 | 5.788e-07 | 2.241e-05 |
| 121 | NEGATIVE REGULATION OF TRANSFERASE ACTIVITY | 8 | 351 | 5.829e-07 | 2.241e-05 |
| 122 | PROTEOLYSIS | 13 | 1208 | 6.567e-07 | 2.505e-05 |
| 123 | POSITIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 8 | 360 | 7.054e-07 | 2.669e-05 |
| 124 | T CELL HOMEOSTASIS | 4 | 34 | 7.441e-07 | 2.77e-05 |
| 125 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 4 | 34 | 7.441e-07 | 2.77e-05 |
| 126 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 12 | 1036 | 9.133e-07 | 3.346e-05 |
| 127 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 12 | 1036 | 9.133e-07 | 3.346e-05 |
| 128 | REGULATION OF CELLULAR RESPONSE TO STRESS | 10 | 691 | 1.228e-06 | 4.438e-05 |
| 129 | REGULATION OF CELL MATRIX ADHESION | 5 | 90 | 1.23e-06 | 4.438e-05 |
| 130 | POSITIVE REGULATION OF CELL COMMUNICATION | 14 | 1532 | 1.543e-06 | 5.522e-05 |
| 131 | RESPONSE TO CORTICOSTEROID | 6 | 176 | 1.72e-06 | 6.07e-05 |
| 132 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 9 | 552 | 1.723e-06 | 6.07e-05 |
| 133 | RESPONSE TO LIGHT STIMULUS | 7 | 280 | 1.735e-06 | 6.07e-05 |
| 134 | REGULATION OF HYDROLASE ACTIVITY | 13 | 1327 | 1.893e-06 | 6.573e-05 |
| 135 | REGULATION OF CATABOLIC PROCESS | 10 | 731 | 2.038e-06 | 7.025e-05 |
| 136 | RESPONSE TO KETONE | 6 | 182 | 2.09e-06 | 7.149e-05 |
| 137 | NEGATIVE REGULATION OF PHOSPHORYLATION | 8 | 422 | 2.314e-06 | 7.861e-05 |
| 138 | POSITIVE REGULATION OF DNA DAMAGE RESPONSE SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 3 | 13 | 2.427e-06 | 8.066e-05 |
| 139 | ACTIVATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 13 | 2.427e-06 | 8.066e-05 |
| 140 | RESPONSE TO COBALT ION | 3 | 13 | 2.427e-06 | 8.066e-05 |
| 141 | RESPONSE TO NUTRIENT | 6 | 191 | 2.764e-06 | 9.12e-05 |
| 142 | DNA METABOLIC PROCESS | 10 | 758 | 2.82e-06 | 9.24e-05 |
| 143 | LYMPHOCYTE HOMEOSTASIS | 4 | 50 | 3.606e-06 | 0.0001173 |
| 144 | NEGATIVE REGULATION OF B CELL PROLIFERATION | 3 | 15 | 3.85e-06 | 0.0001244 |
| 145 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 9 | 616 | 4.227e-06 | 0.0001356 |
| 146 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 15 | 1929 | 4.364e-06 | 0.0001391 |
| 147 | NEGATIVE REGULATION OF CELL SUBSTRATE ADHESION | 4 | 53 | 4.565e-06 | 0.0001435 |
| 148 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 4 | 53 | 4.565e-06 | 0.0001435 |
| 149 | RESPONSE TO MECHANICAL STIMULUS | 6 | 210 | 4.775e-06 | 0.0001491 |
| 150 | NEGATIVE REGULATION OF PROTEOLYSIS | 7 | 329 | 5.031e-06 | 0.0001561 |
| 151 | REGULATION OF B CELL ACTIVATION | 5 | 121 | 5.305e-06 | 0.0001624 |
| 152 | REGULATION OF B CELL PROLIFERATION | 4 | 55 | 5.3e-06 | 0.0001624 |
| 153 | POSITIVE REGULATION OF SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 3 | 17 | 5.737e-06 | 0.0001722 |
| 154 | POSITIVE REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 17 | 5.737e-06 | 0.0001722 |
| 155 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 3 | 17 | 5.737e-06 | 0.0001722 |
| 156 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 15 | 1977 | 5.914e-06 | 0.0001764 |
| 157 | REGULATION OF DNA METABOLIC PROCESS | 7 | 340 | 6.24e-06 | 0.0001849 |
| 158 | NEGATIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 5 | 126 | 6.467e-06 | 0.0001905 |
| 159 | LEUKOCYTE HOMEOSTASIS | 4 | 60 | 7.521e-06 | 0.0002201 |
| 160 | RESPONSE TO ETHANOL | 5 | 136 | 9.385e-06 | 0.0002729 |
| 161 | CELLULAR RESPONSE TO RADIATION | 5 | 137 | 9.726e-06 | 0.0002811 |
| 162 | CELLULAR RESPONSE TO UV | 4 | 66 | 1.102e-05 | 0.0003164 |
| 163 | RESPONSE TO BIOTIC STIMULUS | 10 | 886 | 1.118e-05 | 0.0003192 |
| 164 | POSITIVE REGULATION OF CELL CYCLE PHASE TRANSITION | 4 | 68 | 1.241e-05 | 0.0003521 |
| 165 | NEGATIVE REGULATION OF KINASE ACTIVITY | 6 | 250 | 1.293e-05 | 0.0003602 |
| 166 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 22 | 1.29e-05 | 0.0003602 |
| 167 | REGULATION OF RESPONSE TO DNA DAMAGE STIMULUS | 5 | 145 | 1.281e-05 | 0.0003602 |
| 168 | REGULATION OF PROTEASOMAL UBIQUITIN DEPENDENT PROTEIN CATABOLIC PROCESS | 5 | 148 | 1.415e-05 | 0.0003887 |
| 169 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 8 | 541 | 1.428e-05 | 0.0003887 |
| 170 | RESPONSE TO TRANSITION METAL NANOPARTICLE | 5 | 148 | 1.415e-05 | 0.0003887 |
| 171 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 8 | 541 | 1.428e-05 | 0.0003887 |
| 172 | INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO DNA DAMAGE | 4 | 71 | 1.474e-05 | 0.0003938 |
| 173 | RESPONSE TO MAGNESIUM ION | 3 | 23 | 1.481e-05 | 0.0003938 |
| 174 | RESPONSE TO INCREASED OXYGEN LEVELS | 3 | 23 | 1.481e-05 | 0.0003938 |
| 175 | RESPONSE TO HYPEROXIA | 3 | 23 | 1.481e-05 | 0.0003938 |
| 176 | REGULATION OF PROTEIN CATABOLIC PROCESS | 7 | 393 | 1.599e-05 | 0.0004226 |
| 177 | PROTEIN PHOSPHORYLATION | 10 | 944 | 1.939e-05 | 0.0005096 |
| 178 | RESPONSE TO CORTICOSTERONE | 3 | 26 | 2.165e-05 | 0.0005641 |
| 179 | REGULATION OF CELLULAR PROTEIN CATABOLIC PROCESS | 6 | 274 | 2.17e-05 | 0.0005641 |
| 180 | REGULATION OF NUCLEAR DIVISION | 5 | 163 | 2.254e-05 | 0.0005827 |
| 181 | CELLULAR RESPONSE TO MECHANICAL STIMULUS | 4 | 80 | 2.366e-05 | 0.0006083 |
| 182 | RESPONSE TO CARBOHYDRATE | 5 | 168 | 2.607e-05 | 0.0006664 |
| 183 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 6 | 285 | 2.708e-05 | 0.0006885 |
| 184 | REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 29 | 3.029e-05 | 0.0007578 |
| 185 | POSITIVE REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 29 | 3.029e-05 | 0.0007578 |
| 186 | REGULATION OF CELL SUBSTRATE ADHESION | 5 | 173 | 3e-05 | 0.0007578 |
| 187 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 8 | 602 | 3.074e-05 | 0.0007649 |
| 188 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 7 | 437 | 3.157e-05 | 0.0007813 |
| 189 | PHOSPHORYLATION | 11 | 1228 | 3.271e-05 | 0.0008053 |
| 190 | INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO DNA DAMAGE BY P53 CLASS MEDIATOR | 3 | 30 | 3.361e-05 | 0.000823 |
| 191 | POSITIVE REGULATION OF GENE EXPRESSION | 13 | 1733 | 3.45e-05 | 0.0008405 |
| 192 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 8 | 616 | 3.62e-05 | 0.0008772 |
| 193 | REGULATION OF PROTEASOMAL PROTEIN CATABOLIC PROCESS | 5 | 181 | 3.725e-05 | 0.000898 |
| 194 | REGULATION OF INTRACELLULAR TRANSPORT | 8 | 621 | 3.834e-05 | 0.0009195 |
| 195 | POSITIVE REGULATION OF CELL PROLIFERATION | 9 | 814 | 3.901e-05 | 0.0009307 |
| 196 | CELLULAR RESPONSE TO LIGHT STIMULUS | 4 | 91 | 3.933e-05 | 0.0009336 |
| 197 | INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO ENDOPLASMIC RETICULUM STRESS | 3 | 32 | 4.094e-05 | 0.0009669 |
| 198 | REGULATION OF GROWTH | 8 | 633 | 4.39e-05 | 0.001032 |
| 199 | CELLULAR RESPONSE TO EXTRACELLULAR STIMULUS | 5 | 188 | 4.464e-05 | 0.001041 |
| 200 | REGULATION OF PROTEIN EXPORT FROM NUCLEUS | 3 | 33 | 4.497e-05 | 0.001041 |
| 201 | REGULATION OF CELL AGING | 3 | 33 | 4.497e-05 | 0.001041 |
| 202 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 7 | 465 | 4.683e-05 | 0.001079 |
| 203 | RESPONSE TO REACTIVE OXYGEN SPECIES | 5 | 191 | 4.813e-05 | 0.001103 |
| 204 | PROTEIN DESTABILIZATION | 3 | 34 | 4.925e-05 | 0.001123 |
| 205 | POSITIVE REGULATION OF MAPK CASCADE | 7 | 470 | 5.01e-05 | 0.001137 |
| 206 | RESPONSE TO MOLECULE OF BACTERIAL ORIGIN | 6 | 321 | 5.26e-05 | 0.001188 |
| 207 | NEGATIVE REGULATION OF PROTEIN PROCESSING | 3 | 35 | 5.378e-05 | 0.001191 |
| 208 | RESPONSE TO MINERALOCORTICOID | 3 | 35 | 5.378e-05 | 0.001191 |
| 209 | REPRODUCTION | 11 | 1297 | 5.399e-05 | 0.001191 |
| 210 | NEGATIVE REGULATION OF PROTEIN MATURATION | 3 | 35 | 5.378e-05 | 0.001191 |
| 211 | RESPONSE TO IRON ION | 3 | 35 | 5.378e-05 | 0.001191 |
| 212 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 4 | 99 | 5.472e-05 | 0.001201 |
| 213 | POSITIVE REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 36 | 5.859e-05 | 0.00128 |
| 214 | REGULATION OF CELL ACTIVATION | 7 | 484 | 6.029e-05 | 0.001311 |
| 215 | REGULATION OF LEUKOCYTE PROLIFERATION | 5 | 206 | 6.89e-05 | 0.001491 |
| 216 | ORGANELLE FISSION | 7 | 496 | 7.033e-05 | 0.001515 |
| 217 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 3 | 39 | 7.466e-05 | 0.001601 |
| 218 | CELLULAR RESPONSE TO ESTROGEN STIMULUS | 3 | 41 | 8.683e-05 | 0.001853 |
| 219 | RESPONSE TO OXIDATIVE STRESS | 6 | 352 | 8.752e-05 | 0.00186 |
| 220 | POSITIVE REGULATION OF HYDROLASE ACTIVITY | 9 | 905 | 8.853e-05 | 0.001872 |
| 221 | REGULATION OF MITOCHONDRION ORGANIZATION | 5 | 218 | 9.002e-05 | 0.001895 |
| 222 | REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 5 | 220 | 9.397e-05 | 0.00197 |
| 223 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 10 | 1142 | 9.741e-05 | 0.002032 |
| 224 | MITOTIC NUCLEAR DIVISION | 6 | 361 | 0.0001005 | 0.00207 |
| 225 | REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 43 | 0.0001002 | 0.00207 |
| 226 | NEGATIVE REGULATION OF CELL ADHESION | 5 | 223 | 0.0001002 | 0.00207 |
| 227 | RESPONSE TO BACTERIUM | 7 | 528 | 0.0001039 | 0.002131 |
| 228 | REGULATION OF IMMUNE SYSTEM PROCESS | 11 | 1403 | 0.0001098 | 0.00224 |
| 229 | POSITIVE REGULATION OF INTRACELLULAR TRANSPORT | 6 | 370 | 0.000115 | 0.002338 |
| 230 | NEGATIVE REGULATION OF IMMUNE SYSTEM PROCESS | 6 | 372 | 0.0001185 | 0.002397 |
| 231 | RESPONSE TO ENDOPLASMIC RETICULUM STRESS | 5 | 233 | 0.0001231 | 0.002479 |
| 232 | REGULATION OF PROTEIN LOCALIZATION | 9 | 950 | 0.0001282 | 0.00257 |
| 233 | RESPONSE TO ANTIBIOTIC | 3 | 47 | 0.0001309 | 0.002604 |
| 234 | POSITIVE REGULATION OF NEURON APOPTOTIC PROCESS | 3 | 47 | 0.0001309 | 0.002604 |
| 235 | REGULATION OF SMOOTH MUSCLE CELL MIGRATION | 3 | 49 | 0.0001483 | 0.002926 |
| 236 | REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION TO MITOCHONDRION | 4 | 128 | 0.0001484 | 0.002926 |
| 237 | NEGATIVE REGULATION OF PEPTIDASE ACTIVITY | 5 | 245 | 0.0001556 | 0.003055 |
| 238 | REGULATION OF LIGASE ACTIVITY | 4 | 130 | 0.0001575 | 0.00308 |
| 239 | GLAND DEVELOPMENT | 6 | 395 | 0.0001643 | 0.003198 |
| 240 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 7 | 573 | 0.0001724 | 0.003343 |
| 241 | REGULATION OF NEURON DEATH | 5 | 252 | 0.0001774 | 0.003425 |
| 242 | PROTEIN CATABOLIC PROCESS | 7 | 579 | 0.0001838 | 0.003534 |
| 243 | CIRCADIAN RHYTHM | 4 | 137 | 0.0001927 | 0.00369 |
| 244 | REPRODUCTIVE SYSTEM DEVELOPMENT | 6 | 408 | 0.0001957 | 0.003733 |
| 245 | TISSUE DEVELOPMENT | 11 | 1518 | 0.0002205 | 0.004187 |
| 246 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 8 | 801 | 0.000225 | 0.004255 |
| 247 | RAS PROTEIN SIGNAL TRANSDUCTION | 4 | 143 | 0.0002271 | 0.004278 |
| 248 | REGULATION OF CELLULAR LOCALIZATION | 10 | 1277 | 0.0002438 | 0.004573 |
| 249 | REGULATION OF CELL DIVISION | 5 | 272 | 0.0002528 | 0.004724 |
| 250 | REGULATION OF EPITHELIAL CELL APOPTOTIC PROCESS | 3 | 59 | 0.0002579 | 0.004801 |
| 251 | RESPONSE TO TEMPERATURE STIMULUS | 4 | 148 | 0.000259 | 0.004801 |
| 252 | DEOXYRIBONUCLEOTIDE BIOSYNTHETIC PROCESS | 2 | 12 | 0.0002804 | 0.005157 |
| 253 | POSITIVE REGULATION OF INSULIN LIKE GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 2 | 12 | 0.0002804 | 0.005157 |
| 254 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 12 | 1848 | 0.0002973 | 0.005447 |
| 255 | REGULATION OF CELL ADHESION | 7 | 629 | 0.0003047 | 0.005559 |
| 256 | POSITIVE REGULATION OF RESPONSE TO DNA DAMAGE STIMULUS | 3 | 64 | 0.0003282 | 0.005965 |
| 257 | REGULATION OF HISTONE PHOSPHORYLATION | 2 | 13 | 0.0003309 | 0.005968 |
| 258 | NEGATIVE REGULATION OF CELL ACTIVATION | 4 | 158 | 0.0003322 | 0.005968 |
| 259 | RESPONSE TO PURINE CONTAINING COMPOUND | 4 | 158 | 0.0003322 | 0.005968 |
| 260 | POSITIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 5 | 289 | 0.0003342 | 0.00598 |
| 261 | CELLULAR RESPONSE TO LIPID | 6 | 457 | 0.0003597 | 0.006413 |
| 262 | POSITIVE REGULATION OF NEURON DEATH | 3 | 67 | 0.0003757 | 0.006647 |
| 263 | REGULATION OF SISTER CHROMATID SEGREGATION | 3 | 67 | 0.0003757 | 0.006647 |
| 264 | REGULATION OF FIBRINOLYSIS | 2 | 14 | 0.0003856 | 0.006744 |
| 265 | RHYTHMIC PROCESS | 5 | 298 | 0.0003846 | 0.006744 |
| 266 | REGULATION OF SMOOTH MUSCLE CELL APOPTOTIC PROCESS | 2 | 14 | 0.0003856 | 0.006744 |
| 267 | REGULATION OF MAPK CASCADE | 7 | 660 | 0.0004073 | 0.007098 |
| 268 | NEGATIVE REGULATION OF LEUKOCYTE PROLIFERATION | 3 | 69 | 0.0004097 | 0.007113 |
| 269 | REGULATION OF MEMBRANE PERMEABILITY | 3 | 70 | 0.0004274 | 0.007393 |
| 270 | DENTATE GYRUS DEVELOPMENT | 2 | 15 | 0.0004443 | 0.007628 |
| 271 | RESPONSE TO VITAMIN E | 2 | 15 | 0.0004443 | 0.007628 |
| 272 | POSITIVE REGULATION OF KINASE ACTIVITY | 6 | 482 | 0.0004772 | 0.008163 |
| 273 | HIPPOCAMPUS DEVELOPMENT | 3 | 73 | 0.0004835 | 0.008211 |
| 274 | REGULATION OF ORGAN GROWTH | 3 | 73 | 0.0004835 | 0.008211 |
| 275 | HOMEOSTASIS OF NUMBER OF CELLS | 4 | 175 | 0.0004891 | 0.008275 |
| 276 | NEGATIVE REGULATION OF ADHERENS JUNCTION ORGANIZATION | 2 | 16 | 0.0005071 | 0.008504 |
| 277 | NEGATIVE REGULATION OF SMOOTH MUSCLE CELL MIGRATION | 2 | 16 | 0.0005071 | 0.008504 |
| 278 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 11 | 1672 | 0.0005081 | 0.008504 |
| 279 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 9 | 1152 | 0.000537 | 0.008955 |
| 280 | NEGATIVE REGULATION OF CELL AGING | 2 | 17 | 0.0005739 | 0.009537 |
| 281 | MACROMOLECULE CATABOLIC PROCESS | 8 | 926 | 0.0005946 | 0.009846 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CYCLIN DEPENDENT PROTEIN SERINE THREONINE KINASE REGULATOR ACTIVITY | 7 | 28 | 1.219e-13 | 1.133e-10 |
| 2 | CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 5 | 34 | 8.492e-09 | 3.945e-06 |
| 3 | CYCLIN BINDING | 4 | 19 | 6.363e-08 | 1.97e-05 |
| 4 | KINASE REGULATOR ACTIVITY | 7 | 186 | 1.101e-07 | 2.557e-05 |
| 5 | PROTEIN COMPLEX BINDING | 12 | 935 | 3.063e-07 | 5.691e-05 |
| 6 | ENZYME BINDING | 15 | 1737 | 1.171e-06 | 0.0001812 |
| 7 | CYCLIN DEPENDENT PROTEIN SERINE THREONINE KINASE INHIBITOR ACTIVITY | 3 | 12 | 1.87e-06 | 0.0002481 |
| 8 | KINASE BINDING | 9 | 606 | 3.7e-06 | 0.0003437 |
| 9 | ENZYME REGULATOR ACTIVITY | 11 | 959 | 3.167e-06 | 0.0003437 |
| 10 | MACROMOLECULAR COMPLEX BINDING | 13 | 1399 | 3.406e-06 | 0.0003437 |
| 11 | PROTEIN KINASE ACTIVITY | 9 | 640 | 5.762e-06 | 0.0004866 |
| 12 | KINASE ACTIVITY | 10 | 842 | 7.157e-06 | 0.0005541 |
| 13 | HISTONE KINASE ACTIVITY | 3 | 19 | 8.151e-06 | 0.0005825 |
| 14 | P53 BINDING | 4 | 67 | 1.17e-05 | 0.0007762 |
| 15 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 10 | 992 | 2.969e-05 | 0.001839 |
| 16 | PROTEIN SERINE THREONINE KINASE INHIBITOR ACTIVITY | 3 | 30 | 3.361e-05 | 0.001936 |
| 17 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 7 | 445 | 3.543e-05 | 0.001936 |
| 18 | MOLECULAR FUNCTION REGULATOR | 11 | 1353 | 7.922e-05 | 0.004088 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CYCLIN DEPENDENT PROTEIN KINASE HOLOENZYME COMPLEX | 6 | 31 | 4.184e-11 | 2.443e-08 |
| 2 | PROTEIN KINASE COMPLEX | 6 | 90 | 3.23e-08 | 9.43e-06 |
| 3 | TRANSFERASE COMPLEX TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 6 | 237 | 9.541e-06 | 0.001857 |
| 4 | CATALYTIC COMPLEX | 10 | 1038 | 4.368e-05 | 0.005102 |
| 5 | MICROTUBULE ORGANIZING CENTER | 8 | 623 | 3.922e-05 | 0.005102 |
| 6 | MICROTUBULE CYTOSKELETON | 10 | 1068 | 5.558e-05 | 0.00523 |
| 7 | CENTROSOME | 7 | 487 | 6.268e-05 | 0.00523 |
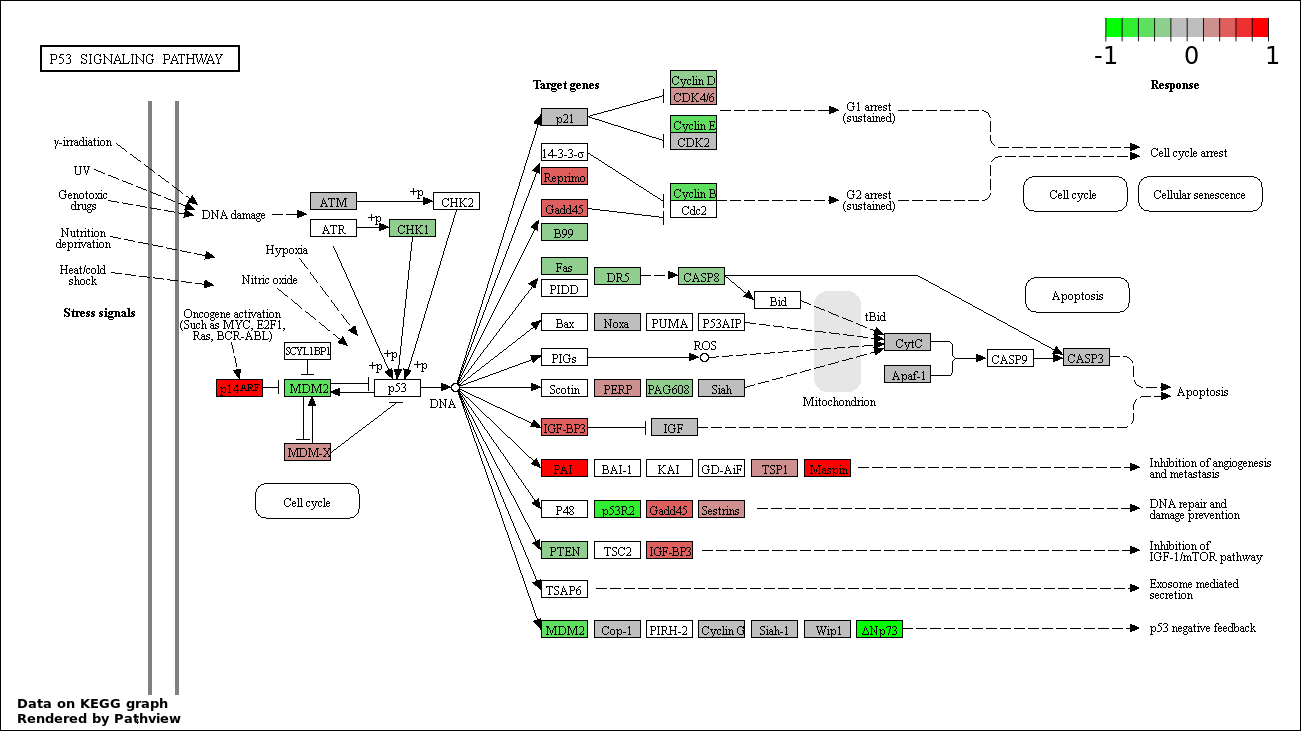
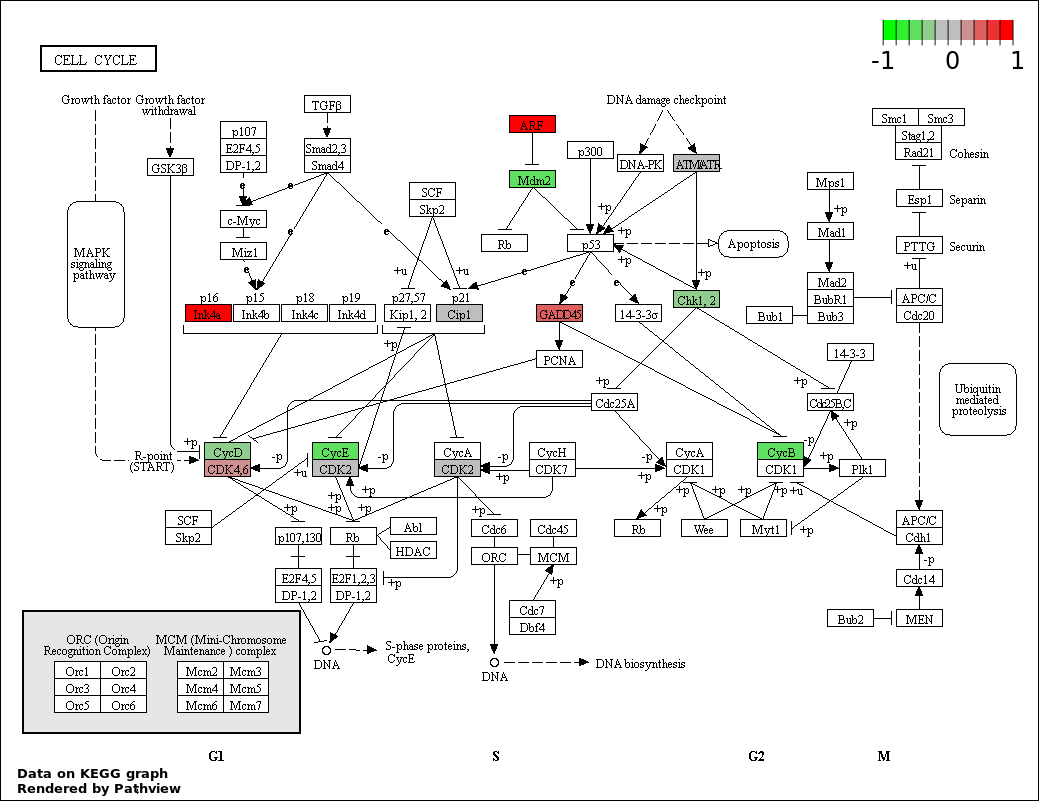
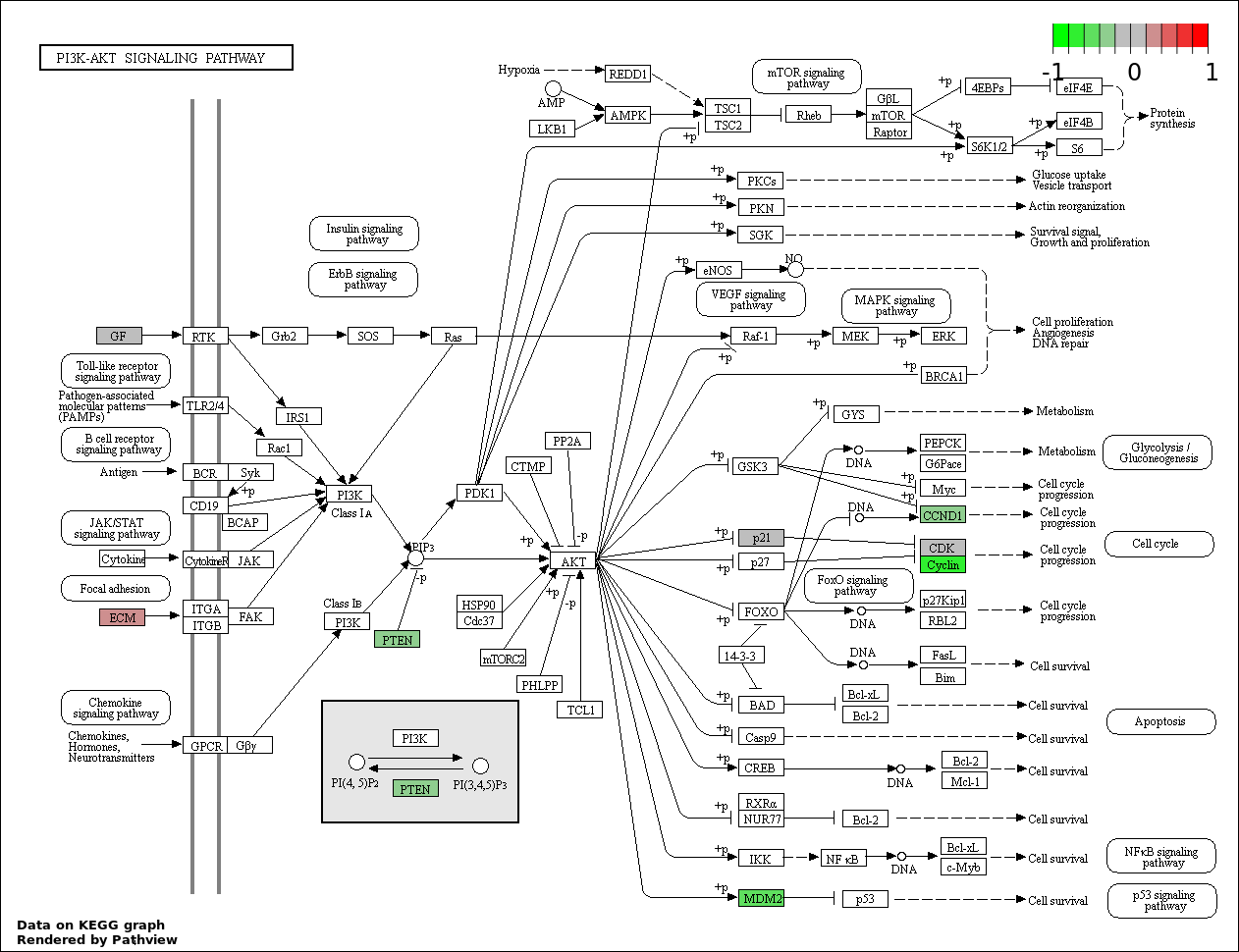
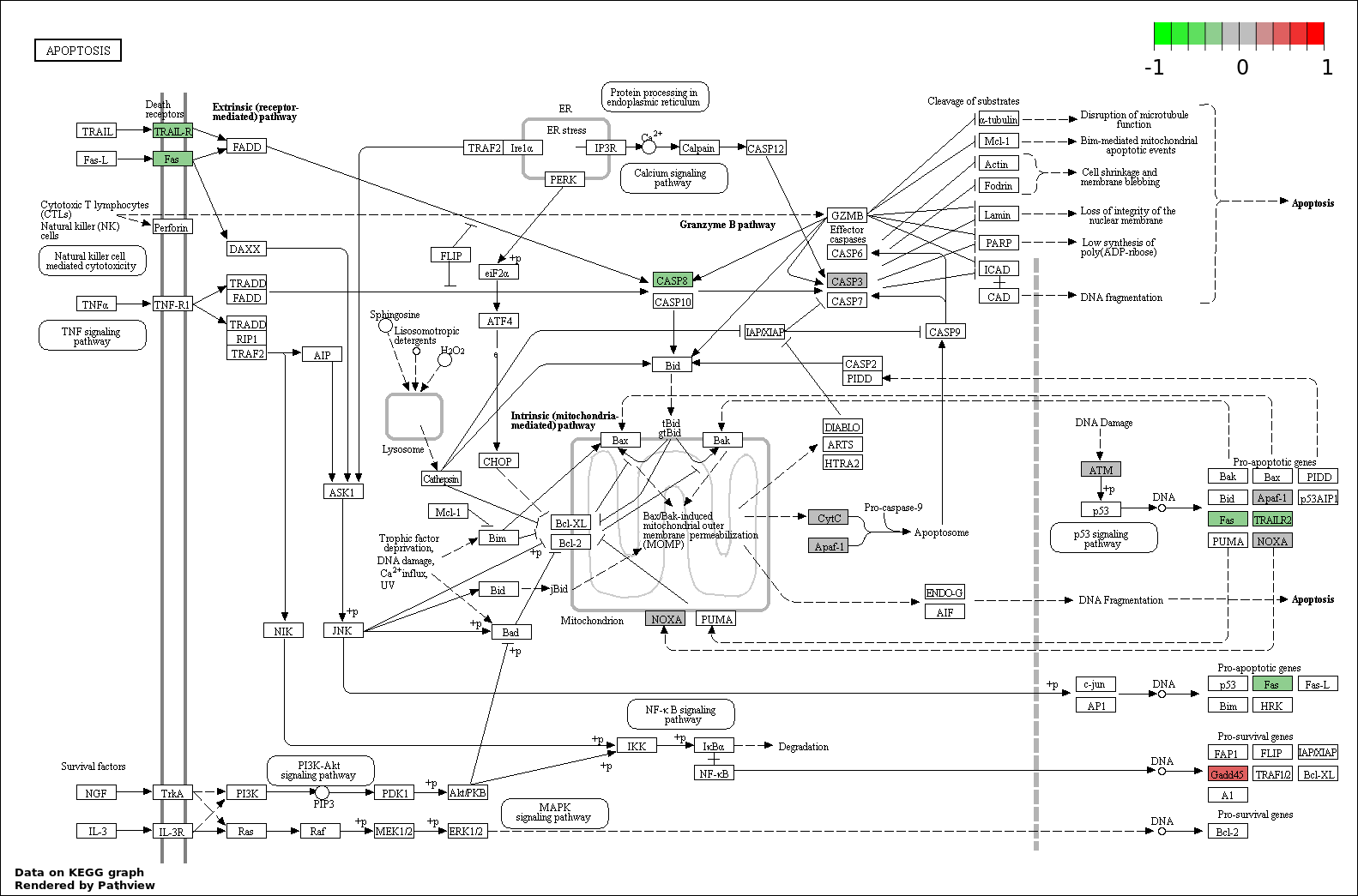
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04115_p53_signaling_pathway | 42 | 69 | 3.731e-111 | 6.715e-109 | |
| 2 | hsa04110_Cell_cycle | 14 | 128 | 4.231e-21 | 3.808e-19 | |
| 3 | hsa04151_PI3K_AKT_signaling_pathway | 11 | 351 | 1.096e-10 | 6.574e-09 | |
| 4 | hsa04210_Apoptosis | 7 | 89 | 6.461e-10 | 2.907e-08 | |
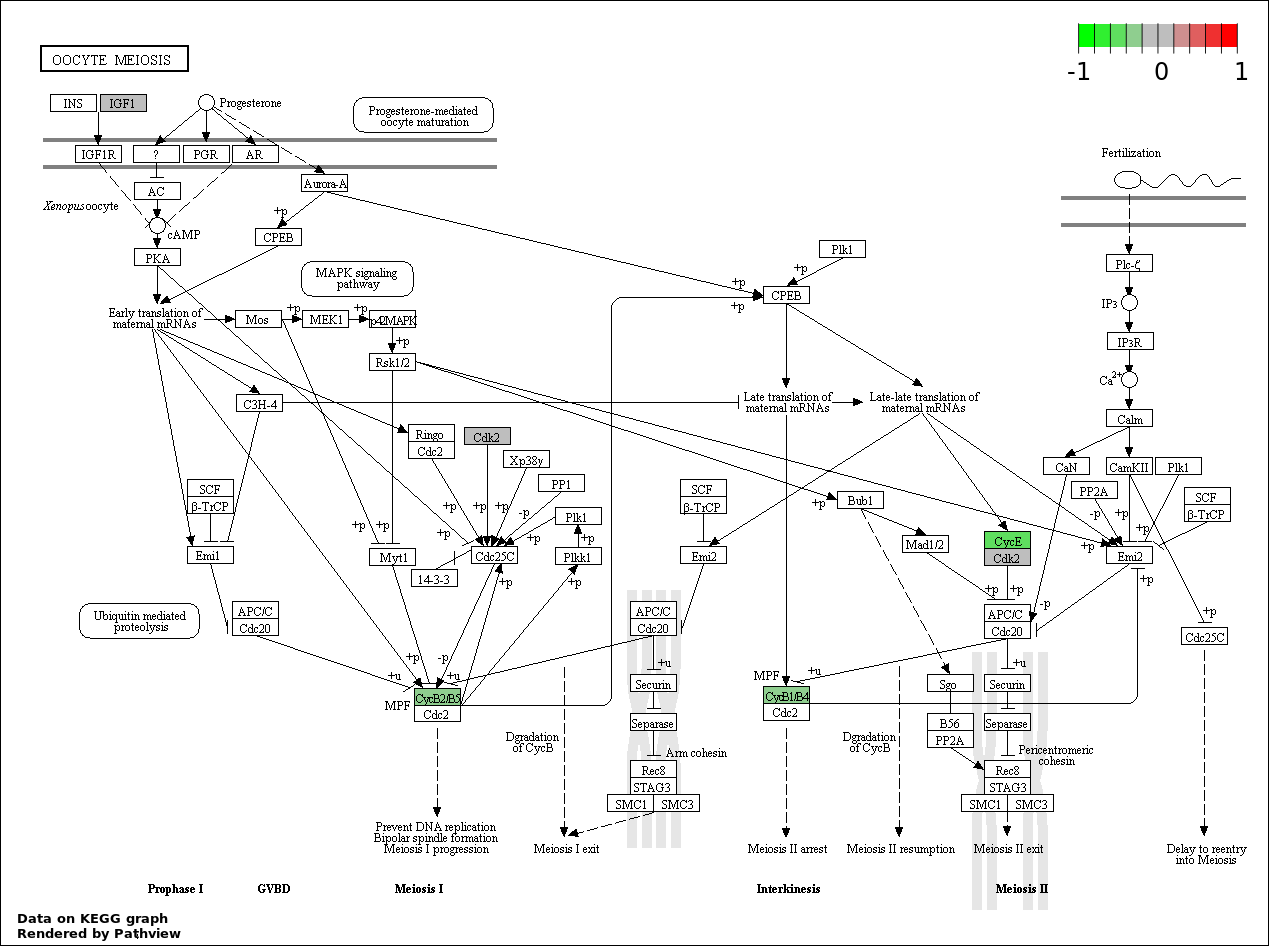
| 5 | hsa04114_Oocyte_meiosis | 6 | 114 | 1.333e-07 | 4.799e-06 | |
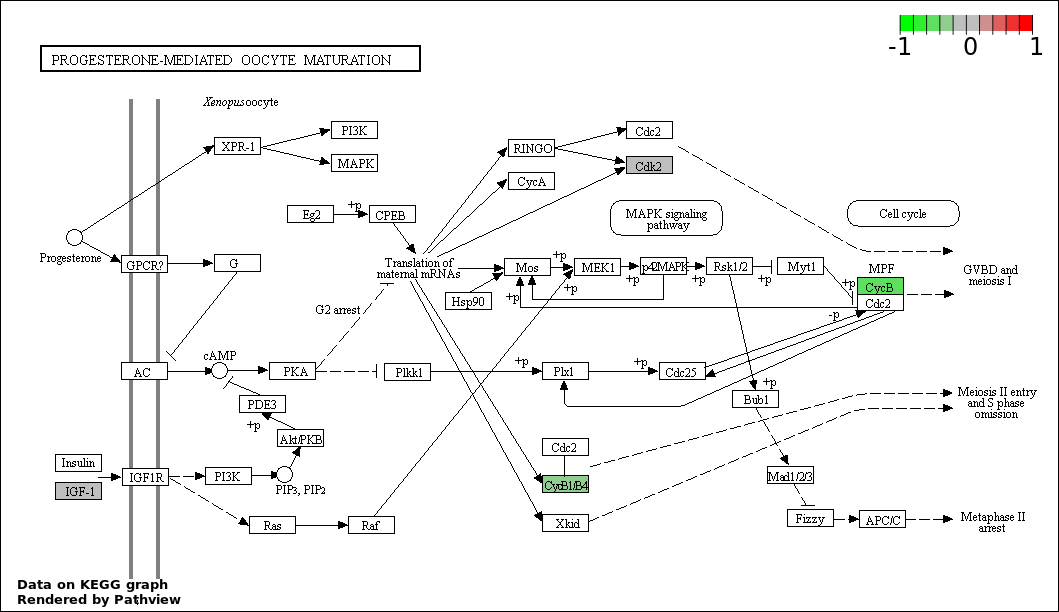
| 6 | hsa04914_Progesterone.mediated_oocyte_maturation | 4 | 87 | 3.295e-05 | 0.0009886 | |
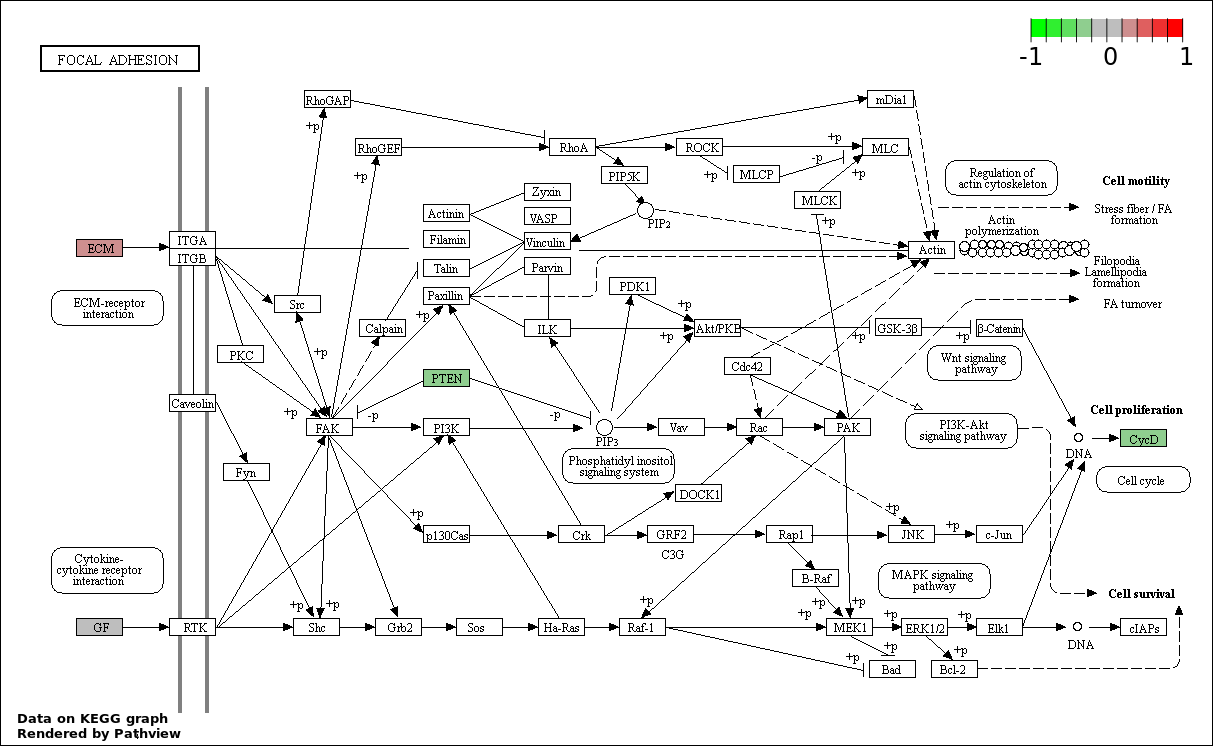
| 7 | hsa04510_Focal_adhesion | 4 | 200 | 0.0008068 | 0.02075 | |
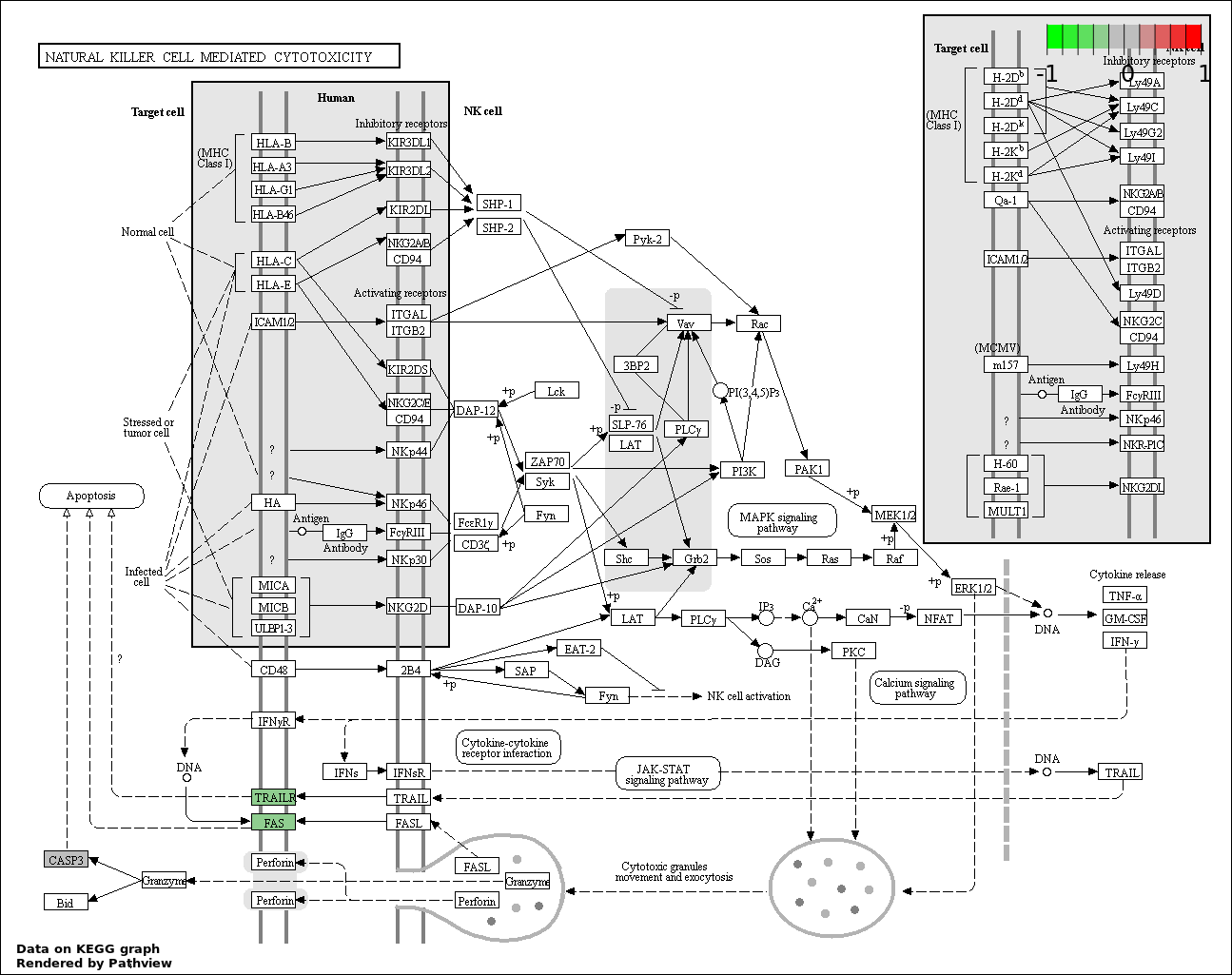
| 8 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 3 | 136 | 0.002909 | 0.06187 | |
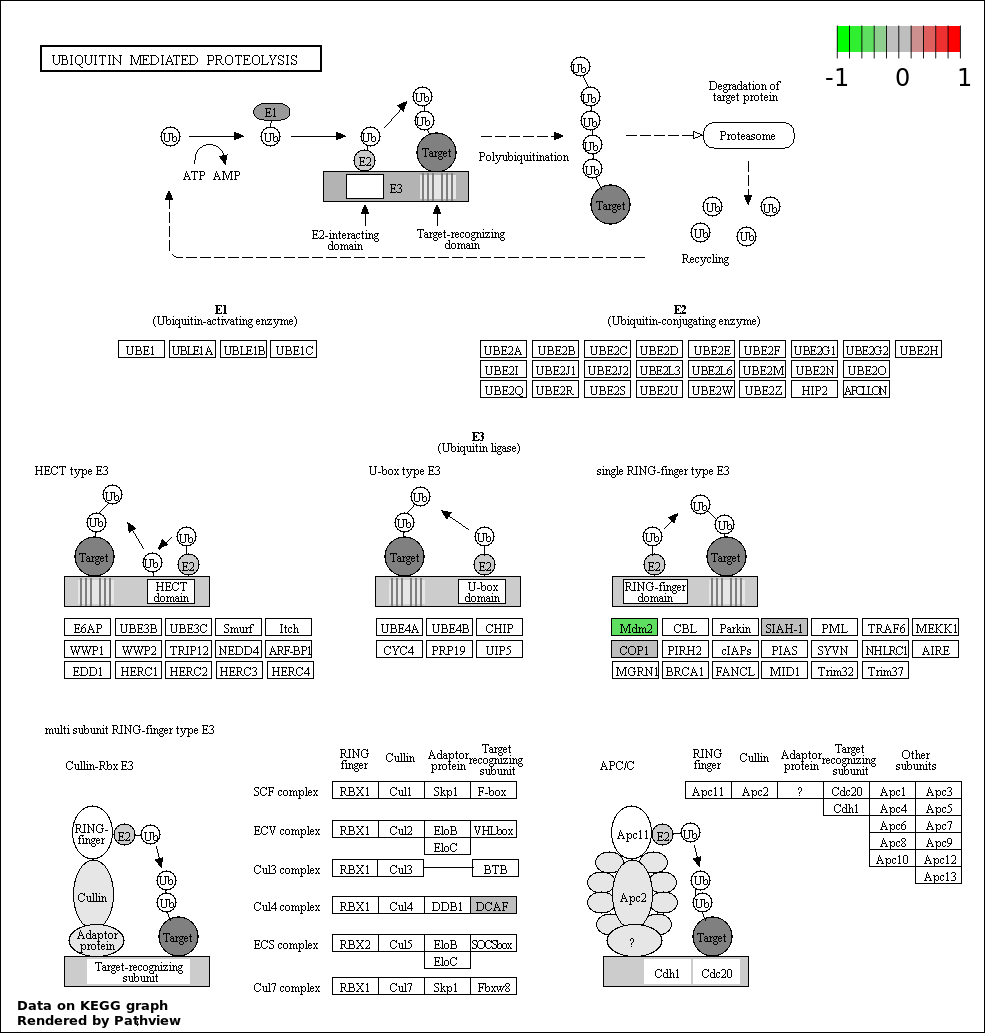
| 9 | hsa04120_Ubiquitin_mediated_proteolysis | 3 | 139 | 0.003093 | 0.06187 | |
| 10 | hsa04390_Hippo_signaling_pathway | 3 | 154 | 0.004125 | 0.07425 | |
| 11 | hsa00480_Glutathione_metabolism | 2 | 50 | 0.004948 | 0.08096 | |
| 12 | hsa00240_Pyrimidine_metabolism | 2 | 99 | 0.01836 | 0.2574 | |
| 13 | hsa04010_MAPK_signaling_pathway | 3 | 268 | 0.01859 | 0.2574 | |
| 14 | hsa04530_Tight_junction | 2 | 133 | 0.03178 | 0.4086 | |
| 15 | hsa04310_Wnt_signaling_pathway | 2 | 151 | 0.04004 | 0.4805 | |
| 16 | hsa00230_Purine_metabolism | 2 | 162 | 0.04545 | 0.5114 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | EMX2OS |
hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-130b-3p;hsa-miR-148a-5p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-27a-3p;hsa-miR-29a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-33a-3p;hsa-miR-374b-3p;hsa-miR-421;hsa-miR-452-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-577;hsa-miR-590-3p;hsa-miR-592;hsa-miR-629-5p;hsa-miR-940 | 29 | IGF1 | Sponge network | 1.057 | 0.31716 | 0.009 | 0.98747 | 0.587 |
| 2 | CECR7 |
hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-1275;hsa-miR-130b-3p;hsa-miR-148a-5p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-26b-5p;hsa-miR-29a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-362-5p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-592;hsa-miR-629-5p | 24 | IGF1 | Sponge network | 0.551 | 0.56177 | 0.009 | 0.98747 | 0.513 |
| 3 | EMX2OS |
hsa-let-7a-3p;hsa-let-7d-5p;hsa-let-7f-1-3p;hsa-miR-1304-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-375;hsa-miR-421;hsa-miR-576-5p;hsa-miR-577;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-940 | 31 | THBS1 | Sponge network | 1.057 | 0.31716 | 0.376 | 0.83609 | 0.511 |
| 4 | CECR7 |
hsa-let-7a-3p;hsa-let-7d-5p;hsa-let-7f-1-3p;hsa-miR-1304-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-375;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-671-5p;hsa-miR-7-1-3p | 26 | THBS1 | Sponge network | 0.551 | 0.56177 | 0.376 | 0.83609 | 0.491 |
| 5 | CECR7 |
hsa-miR-148a-3p;hsa-miR-148a-5p;hsa-miR-301a-3p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-425-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-942-5p | 10 | SERPINE1 | Sponge network | 0.551 | 0.56177 | 0.967 | 0.43183 | 0.452 |
| 6 | MEG3 |
hsa-miR-1304-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-576-5p;hsa-miR-577;hsa-miR-590-5p;hsa-miR-671-5p;hsa-miR-7-1-3p | 15 | THBS1 | Sponge network | 0.433 | 0.33816 | 0.376 | 0.83609 | 0.414 |
| 7 | ZNF883 |
hsa-let-7a-3p;hsa-let-7d-5p;hsa-let-7f-1-3p;hsa-miR-1304-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-576-5p;hsa-miR-590-5p | 18 | THBS1 | Sponge network | 0.913 | 0.16772 | 0.376 | 0.83609 | 0.396 |
| 8 | EMX2OS |
hsa-miR-148a-3p;hsa-miR-148a-5p;hsa-miR-301a-3p;hsa-miR-378a-5p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-452-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-942-5p | 10 | SERPINE1 | Sponge network | 1.057 | 0.31716 | 0.967 | 0.43183 | 0.395 |
| 9 | MEG3 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-186-5p;hsa-miR-20a-3p;hsa-miR-26b-5p;hsa-miR-29a-3p;hsa-miR-29b-3p;hsa-miR-374b-3p;hsa-miR-576-5p;hsa-miR-577 | 10 | IGF1 | Sponge network | 0.433 | 0.33816 | 0.009 | 0.98747 | 0.37 |
| 10 | CECR7 |
hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-148a-3p;hsa-miR-148b-5p;hsa-miR-15a-5p;hsa-miR-15b-3p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-192-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-33a-3p;hsa-miR-362-3p;hsa-miR-362-5p;hsa-miR-425-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-671-5p;hsa-miR-93-5p;hsa-miR-942-5p | 32 | ZMAT3 | Sponge network | 0.551 | 0.56177 | -0.325 | 0.50394 | 0.314 |
| 11 | ZNF883 |
hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-130b-3p;hsa-miR-148a-5p;hsa-miR-15b-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-26b-5p;hsa-miR-29a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-33a-3p;hsa-miR-576-5p | 15 | IGF1 | Sponge network | 0.913 | 0.16772 | 0.009 | 0.98747 | 0.269 |
| 12 | EMX2OS |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-144-3p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-192-3p;hsa-miR-19b-1-5p;hsa-miR-200b-5p;hsa-miR-20a-3p;hsa-miR-222-5p;hsa-miR-25-3p;hsa-miR-26b-3p;hsa-miR-29a-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-30d-3p;hsa-miR-32-3p;hsa-miR-32-5p;hsa-miR-33a-3p;hsa-miR-33a-5p;hsa-miR-33b-5p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-577;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-625-5p;hsa-miR-7-1-3p;hsa-miR-877-5p | 30 | SESN3 | Sponge network | 1.057 | 0.31716 | -0.016 | 0.97504 | 0.252 |
| 13 | HCG11 | hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-222-5p;hsa-miR-26b-3p;hsa-miR-33a-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-625-3p;hsa-miR-7-1-3p;hsa-miR-877-5p | 11 | SESN3 | Sponge network | 0.419 | 0.63279 | -0.016 | 0.97504 | 0.251 |