This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-106b-5p | AADAC | 1.22 | 0 | -4.47 | 0 | MirTarget | -0.47 | 0 | NA | |
| 2 | hsa-miR-93-5p | AADAC | 1.02 | 0 | -4.47 | 0 | MirTarget | -0.44 | 1.0E-5 | NA | |
| 3 | hsa-miR-106b-5p | ABCA1 | 1.22 | 0 | -1.41 | 0 | MirTarget; miRNATAP | -0.35 | 0 | NA | |
| 4 | hsa-miR-93-5p | ABCA1 | 1.02 | 0 | -1.41 | 0 | MirTarget; miRNATAP | -0.36 | 0 | NA | |
| 5 | hsa-miR-106b-5p | ABCD2 | 1.22 | 0 | -4.13 | 0 | MirTarget | -0.54 | 0 | NA | |
| 6 | hsa-miR-93-5p | ABCD2 | 1.02 | 0 | -4.13 | 0 | MirTarget | -0.62 | 0 | NA | |
| 7 | hsa-miR-106b-5p | ABHD2 | 1.22 | 0 | 0.45 | 0.00053 | miRNATAP | -0.3 | 0 | NA | |
| 8 | hsa-miR-93-5p | ABHD2 | 1.02 | 0 | 0.45 | 0.00053 | miRNATAP | -0.29 | 0 | NA | |
| 9 | hsa-miR-106b-5p | ABHD5 | 1.22 | 0 | -0.76 | 0 | MirTarget | -0.11 | 0 | NA | |
| 10 | hsa-miR-93-5p | ACER3 | 1.02 | 0 | -0.64 | 0 | mirMAP | -0.12 | 0.00102 | NA | |
| 11 | hsa-miR-106b-5p | ACOX1 | 1.22 | 0 | -0.61 | 0 | mirMAP | -0.12 | 0 | NA | |
| 12 | hsa-miR-106b-5p | ACSL4 | 1.22 | 0 | -1.48 | 0 | MirTarget; miRNATAP | -0.18 | 0 | NA | |
| 13 | hsa-miR-93-5p | ACSL4 | 1.02 | 0 | -1.48 | 0 | MirTarget; miRNATAP | -0.23 | 0 | NA | |
| 14 | hsa-miR-106b-5p | ACVR1B | 1.22 | 0 | 0.15 | 0.02139 | miRNATAP | -0.1 | 0 | NA | |
| 15 | hsa-miR-106b-5p | ADAM22 | 1.22 | 0 | -1.22 | 0 | MirTarget; mirMAP | -0.51 | 0 | NA | |
| 16 | hsa-miR-93-5p | ADAM22 | 1.02 | 0 | -1.22 | 0 | MirTarget; mirMAP | -0.54 | 0 | NA | |
| 17 | hsa-miR-106b-5p | ADAMTS5 | 1.22 | 0 | -3.55 | 0 | miRNATAP | -0.71 | 0 | NA | |
| 18 | hsa-miR-20a-3p | ADAMTS6 | -0.22 | 0.17987 | 1.1 | 0 | miRNATAP | -0.13 | 0 | NA | |
| 19 | hsa-miR-106a-5p | ADCY1 | 0.57 | 0.00015 | -0.05 | 0.84149 | mirMAP | -0.21 | 0.00014 | NA | |
| 20 | hsa-miR-106b-5p | ADCY1 | 1.22 | 0 | -0.05 | 0.84149 | mirMAP | -0.46 | 0 | NA | |
| 21 | hsa-miR-20a-3p | ADCY1 | -0.22 | 0.17987 | -0.05 | 0.84149 | mirMAP | -0.27 | 0 | NA | |
| 22 | hsa-miR-93-5p | ADCY1 | 1.02 | 0 | -0.05 | 0.84149 | mirMAP | -0.48 | 0 | NA | |
| 23 | hsa-miR-106b-5p | ADCY9 | 1.22 | 0 | 0.12 | 0.23839 | mirMAP | -0.3 | 0 | NA | |
| 24 | hsa-miR-93-5p | ADCY9 | 1.02 | 0 | 0.12 | 0.23839 | mirMAP | -0.26 | 0 | NA | |
| 25 | hsa-miR-106b-5p | ADRA2A | 1.22 | 0 | -1.76 | 0 | miRNATAP | -0.78 | 0 | NA | |
| 26 | hsa-miR-106b-5p | AFF4 | 1.22 | 0 | -0.28 | 0.00153 | miRNATAP | -0.31 | 0 | NA | |
| 27 | hsa-miR-93-5p | AFF4 | 1.02 | 0 | -0.28 | 0.00153 | miRNATAP | -0.28 | 0 | NA | |
| 28 | hsa-miR-106b-5p | AFP | 1.22 | 0 | -3.14 | 0 | miRNAWalker2 validate | -0.93 | 0 | NA | |
| 29 | hsa-miR-20a-3p | AGFG2 | -0.22 | 0.17987 | -0.32 | 2.0E-5 | mirMAP | -0.12 | 0 | NA | |
| 30 | hsa-miR-106b-5p | AGPS | 1.22 | 0 | -0.4 | 0.00019 | mirMAP | -0.18 | 0 | NA | |
| 31 | hsa-miR-93-5p | AGPS | 1.02 | 0 | -0.4 | 0.00019 | mirMAP | -0.19 | 0 | NA | |
| 32 | hsa-miR-106a-5p | AHNAK | 0.57 | 0.00015 | -1.77 | 0 | miRNATAP | -0.18 | 0 | NA | |
| 33 | hsa-miR-106b-5p | AHNAK | 1.22 | 0 | -1.77 | 0 | miRNATAP | -0.67 | 0 | NA | |
| 34 | hsa-miR-93-5p | AHNAK | 1.02 | 0 | -1.77 | 0 | miRNATAP | -0.58 | 0 | NA | |
| 35 | hsa-miR-106b-5p | AKAP11 | 1.22 | 0 | -0.85 | 0 | miRNAWalker2 validate | -0.35 | 0 | NA | |
| 36 | hsa-miR-93-5p | AKAP11 | 1.02 | 0 | -0.85 | 0 | miRNAWalker2 validate | -0.33 | 0 | NA | |
| 37 | hsa-miR-106b-5p | AKAP13 | 1.22 | 0 | -0.61 | 0 | MirTarget; mirMAP; miRNATAP | -0.23 | 0 | NA | |
| 38 | hsa-miR-93-5p | AKAP13 | 1.02 | 0 | -0.61 | 0 | MirTarget; miRNATAP | -0.26 | 0 | NA | |
| 39 | hsa-miR-106b-5p | AKT3 | 1.22 | 0 | -1.39 | 0 | miRNATAP | -0.33 | 0 | NA | |
| 40 | hsa-miR-93-5p | AKT3 | 1.02 | 0 | -1.39 | 0 | miRNATAP | -0.36 | 0 | NA | |
| 41 | hsa-miR-93-5p | ALDH1A3 | 1.02 | 0 | -2.22 | 0 | mirMAP | -0.4 | 0 | NA | |
| 42 | hsa-miR-106b-5p | ALDH3A2 | 1.22 | 0 | -0.86 | 0 | mirMAP | -0.3 | 0 | NA | |
| 43 | hsa-miR-106b-5p | ALPK3 | 1.22 | 0 | -1.48 | 0 | mirMAP | -0.31 | 0 | NA | |
| 44 | hsa-miR-106a-5p | ALX4 | 0.57 | 0.00015 | -3.22 | 0 | miRNATAP | -0.26 | 3.0E-5 | NA | |
| 45 | hsa-miR-106b-5p | ALX4 | 1.22 | 0 | -3.22 | 0 | miRNATAP | -0.65 | 0 | NA | |
| 46 | hsa-miR-93-5p | ALX4 | 1.02 | 0 | -3.22 | 0 | miRNATAP | -0.62 | 0 | NA | |
| 47 | hsa-miR-20a-3p | ANGPTL1 | -0.22 | 0.17987 | -3.72 | 0 | MirTarget | -0.26 | 0 | NA | |
| 48 | hsa-miR-106a-5p | ANK2 | 0.57 | 0.00015 | -2.51 | 0 | MirTarget; miRNATAP | -0.17 | 1.0E-5 | NA | |
| 49 | hsa-miR-106b-5p | ANK2 | 1.22 | 0 | -2.51 | 0 | MirTarget; miRNATAP | -0.57 | 0 | NA | |
| 50 | hsa-miR-93-5p | ANK2 | 1.02 | 0 | -2.51 | 0 | MirTarget; miRNATAP | -0.6 | 0 | NA | |
| 51 | hsa-miR-106b-5p | ANKFY1 | 1.22 | 0 | -0.49 | 0 | miRNAWalker2 validate; MirTarget | -0.21 | 0 | NA | |
| 52 | hsa-miR-93-5p | ANKFY1 | 1.02 | 0 | -0.49 | 0 | miRNAWalker2 validate | -0.17 | 0 | NA | |
| 53 | hsa-miR-106b-5p | ANKH | 1.22 | 0 | -0.31 | 0.00327 | MirTarget | -0.14 | 2.0E-5 | NA | |
| 54 | hsa-miR-93-5p | ANKH | 1.02 | 0 | -0.31 | 0.00327 | MirTarget | -0.16 | 0 | NA | |
| 55 | hsa-miR-106b-5p | ANKRD12 | 1.22 | 0 | -0.64 | 0 | MirTarget | -0.24 | 0 | NA | |
| 56 | hsa-miR-93-5p | ANKRD12 | 1.02 | 0 | -0.64 | 0 | MirTarget | -0.3 | 0 | NA | |
| 57 | hsa-miR-106a-5p | ANKRD29 | 0.57 | 0.00015 | -2.95 | 0 | MirTarget | -0.25 | 0 | NA | |
| 58 | hsa-miR-106b-5p | ANKRD29 | 1.22 | 0 | -2.95 | 0 | MirTarget | -0.7 | 0 | NA | |
| 59 | hsa-miR-93-5p | ANKRD29 | 1.02 | 0 | -2.95 | 0 | miRNAWalker2 validate; MirTarget | -0.66 | 0 | NA | |
| 60 | hsa-miR-106b-5p | ANKRD50 | 1.22 | 0 | 0.25 | 0.02231 | MirTarget; miRNATAP | -0.2 | 0 | NA | |
| 61 | hsa-miR-93-5p | ANKRD50 | 1.02 | 0 | 0.25 | 0.02231 | MirTarget; miRNATAP | -0.2 | 0 | NA | |
| 62 | hsa-miR-106b-5p | ANO6 | 1.22 | 0 | -1.5 | 0 | MirTarget | -0.42 | 0 | NA | |
| 63 | hsa-miR-93-5p | ANO6 | 1.02 | 0 | -1.5 | 0 | MirTarget; miRNATAP | -0.37 | 0 | NA | |
| 64 | hsa-miR-106b-5p | ANXA11 | 1.22 | 0 | -0.27 | 1.0E-5 | mirMAP | -0.11 | 0 | NA | |
| 65 | hsa-miR-106b-5p | AP1G1 | 1.22 | 0 | -0.11 | 0.12378 | miRNAWalker2 validate | -0.1 | 0 | NA | |
| 66 | hsa-miR-106b-5p | AP2B1 | 1.22 | 0 | 0.26 | 0.00183 | miRNAWalker2 validate | -0.15 | 0 | NA | |
| 67 | hsa-miR-93-5p | AP3S2 | 1.02 | 0 | -0.11 | 0.06712 | mirMAP | -0.1 | 0 | NA | |
| 68 | hsa-miR-106a-5p | APBB2 | 0.57 | 0.00015 | -0.39 | 0.0002 | MirTarget; miRNATAP | -0.15 | 0 | NA | |
| 69 | hsa-miR-106b-5p | APBB2 | 1.22 | 0 | -0.39 | 0.0002 | MirTarget; miRNATAP | -0.42 | 0 | NA | |
| 70 | hsa-miR-93-5p | APBB2 | 1.02 | 0 | -0.39 | 0.0002 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.4 | 0 | NA | |
| 71 | hsa-miR-106b-5p | APC | 1.22 | 0 | -0.69 | 0 | miRNAWalker2 validate; miRTarBase | -0.31 | 0 | 23087084 | miR 106b downregulates adenomatous polyposis coli and promotes cell proliferation in human hepatocellular carcinoma; Moreover we demonstrated that miR-106b downregulates APC expression by directly targeting the 3'-untranslated region of APC messenger RNA; Taken together our results suggest that miR-106b plays an important role in promoting the proliferation of human hepatoma cells and presents a novel mechanism of micro RNA-mediated direct suppression of APC expression in cancer cells |
| 72 | hsa-miR-20a-3p | APC2 | -0.22 | 0.17987 | 0.94 | 0 | mirMAP | -0.14 | 1.0E-5 | NA | |
| 73 | hsa-miR-106a-5p | APCDD1 | 0.57 | 0.00015 | -2.4 | 0 | MirTarget | -0.12 | 0.00041 | NA | |
| 74 | hsa-miR-106b-5p | APCDD1 | 1.22 | 0 | -2.4 | 0 | MirTarget | -0.33 | 0 | NA | |
| 75 | hsa-miR-93-5p | APCDD1 | 1.02 | 0 | -2.4 | 0 | MirTarget | -0.32 | 0 | NA | |
| 76 | hsa-miR-106b-5p | APLP2 | 1.22 | 0 | -0.34 | 1.0E-5 | miRNAWalker2 validate | -0.17 | 0 | NA | |
| 77 | hsa-miR-93-5p | APLP2 | 1.02 | 0 | -0.34 | 1.0E-5 | miRNAWalker2 validate | -0.12 | 0 | NA | |
| 78 | hsa-miR-106a-5p | AR | 0.57 | 0.00015 | -1 | 0.00094 | mirMAP | -0.45 | 0 | NA | |
| 79 | hsa-miR-106b-5p | AR | 1.22 | 0 | -1 | 0.00094 | mirMAP | -1.35 | 0 | NA | |
| 80 | hsa-miR-93-5p | AR | 1.02 | 0 | -1 | 0.00094 | mirMAP | -1.31 | 0 | NA | |
| 81 | hsa-miR-93-5p | ARAP2 | 1.02 | 0 | -0.25 | 0.02981 | MirTarget | -0.13 | 0.00033 | NA | |
| 82 | hsa-miR-106b-5p | ARHGAP1 | 1.22 | 0 | 0.21 | 1.0E-5 | MirTarget; miRNATAP | -0.11 | 0 | NA | |
| 83 | hsa-miR-106b-5p | ARHGAP12 | 1.22 | 0 | -0.58 | 0 | miRNATAP | -0.24 | 0 | NA | |
| 84 | hsa-miR-93-5p | ARHGAP12 | 1.02 | 0 | -0.58 | 0 | miRNAWalker2 validate; miRNATAP | -0.26 | 0 | NA | |
| 85 | hsa-miR-106b-5p | ARHGAP19 | 1.22 | 0 | -0.97 | 0 | mirMAP | -0.17 | 0 | NA | |
| 86 | hsa-miR-93-5p | ARHGAP19 | 1.02 | 0 | -0.97 | 0 | mirMAP | -0.12 | 0 | NA | |
| 87 | hsa-miR-106b-5p | ARHGAP23 | 1.22 | 0 | -0.93 | 0 | mirMAP | -0.22 | 0 | NA | |
| 88 | hsa-miR-93-5p | ARHGAP23 | 1.02 | 0 | -0.93 | 0 | mirMAP | -0.22 | 0 | NA | |
| 89 | hsa-miR-106a-5p | ARHGAP24 | 0.57 | 0.00015 | -1.41 | 0 | MirTarget | -0.11 | 0 | NA | |
| 90 | hsa-miR-106b-5p | ARHGAP24 | 1.22 | 0 | -1.41 | 0 | MirTarget | -0.32 | 0 | NA | |
| 91 | hsa-miR-93-5p | ARHGAP24 | 1.02 | 0 | -1.41 | 0 | MirTarget | -0.35 | 0 | NA | |
| 92 | hsa-miR-106b-5p | ARHGAP26 | 1.22 | 0 | -1.51 | 0 | MirTarget; mirMAP; miRNATAP | -0.29 | 0 | NA | |
| 93 | hsa-miR-93-5p | ARHGAP26 | 1.02 | 0 | -1.51 | 0 | MirTarget; mirMAP; miRNATAP | -0.32 | 0 | NA | |
| 94 | hsa-miR-93-5p | ARHGAP32 | 1.02 | 0 | 0.43 | 9.0E-5 | miRNAWalker2 validate | -0.25 | 0 | NA | |
| 95 | hsa-miR-106b-5p | ARHGEF3 | 1.22 | 0 | -0.04 | 0.65559 | MirTarget; miRNATAP | -0.18 | 0 | NA | |
| 96 | hsa-miR-93-5p | ARHGEF3 | 1.02 | 0 | -0.04 | 0.65559 | MirTarget; miRNATAP | -0.21 | 0 | NA | |
| 97 | hsa-miR-106a-5p | ARHGEF38 | 0.57 | 0.00015 | 0.07 | 0.77513 | mirMAP | -0.16 | 0.00387 | NA | |
| 98 | hsa-miR-106b-5p | ARHGEF38 | 1.22 | 0 | 0.07 | 0.77513 | mirMAP | -0.75 | 0 | NA | |
| 99 | hsa-miR-106b-5p | ARID4A | 1.22 | 0 | -0.86 | 0 | MirTarget; miRNATAP | -0.3 | 0 | NA | |
| 100 | hsa-miR-93-5p | ARID4A | 1.02 | 0 | -0.86 | 0 | MirTarget; miRNATAP | -0.32 | 0 | NA | |
| 101 | hsa-miR-106b-5p | ARL10 | 1.22 | 0 | -1.27 | 0 | mirMAP | -0.33 | 0 | NA | |
| 102 | hsa-miR-93-5p | ARL10 | 1.02 | 0 | -1.27 | 0 | mirMAP | -0.3 | 0 | NA | |
| 103 | hsa-miR-106b-5p | ARL13B | 1.22 | 0 | -0.46 | 0 | miRNAWalker2 validate; MirTarget | -0.16 | 0 | NA | |
| 104 | hsa-miR-93-5p | ARL13B | 1.02 | 0 | -0.46 | 0 | MirTarget | -0.19 | 0 | NA | |
| 105 | hsa-miR-106b-5p | ARL4A | 1.22 | 0 | -1.62 | 0 | miRNATAP | -0.21 | 0 | NA | |
| 106 | hsa-miR-93-5p | ARL4A | 1.02 | 0 | -1.62 | 0 | miRNATAP | -0.26 | 0 | NA | |
| 107 | hsa-miR-93-5p | ARNT2 | 1.02 | 0 | 1.54 | 0 | mirMAP | -0.18 | 0.00214 | NA | |
| 108 | hsa-miR-106b-5p | ARSD | 1.22 | 0 | 0.28 | 0.00266 | mirMAP | -0.23 | 0 | NA | |
| 109 | hsa-miR-93-5p | ARSJ | 1.02 | 0 | -0.61 | 7.0E-5 | miRNAWalker2 validate | -0.29 | 0 | NA | |
| 110 | hsa-miR-93-5p | ASB1 | 1.02 | 0 | -0.93 | 0 | miRNAWalker2 validate | -0.13 | 0 | NA | |
| 111 | hsa-miR-106b-5p | ASH1L | 1.22 | 0 | -0.4 | 1.0E-5 | miRNATAP | -0.26 | 0 | NA | |
| 112 | hsa-miR-93-5p | ASH1L | 1.02 | 0 | -0.4 | 1.0E-5 | miRNATAP | -0.23 | 0 | NA | |
| 113 | hsa-miR-106a-5p | ASPA | 0.57 | 0.00015 | -3.96 | 0 | mirMAP | -0.31 | 0 | NA | |
| 114 | hsa-miR-106b-5p | ASPA | 1.22 | 0 | -3.96 | 0 | mirMAP | -0.9 | 0 | NA | |
| 115 | hsa-miR-20a-3p | ASPA | -0.22 | 0.17987 | -3.96 | 0 | mirMAP | -0.24 | 0 | NA | |
| 116 | hsa-miR-93-5p | ASPA | 1.02 | 0 | -3.96 | 0 | mirMAP | -0.87 | 0 | NA | |
| 117 | hsa-miR-106b-5p | ATE1 | 1.22 | 0 | -0.72 | 0 | MirTarget | -0.35 | 0 | NA | |
| 118 | hsa-miR-93-5p | ATE1 | 1.02 | 0 | -0.72 | 0 | MirTarget | -0.35 | 0 | NA | |
| 119 | hsa-miR-106b-5p | ATF6 | 1.22 | 0 | 0.17 | 0.01386 | mirMAP | -0.13 | 0 | NA | |
| 120 | hsa-miR-106b-5p | ATL3 | 1.22 | 0 | -0.22 | 0.01349 | MirTarget; miRNATAP | -0.1 | 0.00027 | NA | |
| 121 | hsa-miR-106a-5p | ATP1A2 | 0.57 | 0.00015 | -5.48 | 0 | MirTarget | -0.6 | 0 | NA | |
| 122 | hsa-miR-106b-5p | ATP1A2 | 1.22 | 0 | -5.48 | 0 | MirTarget; miRNATAP | -1.47 | 0 | NA | |
| 123 | hsa-miR-93-5p | ATP1A2 | 1.02 | 0 | -5.48 | 0 | MirTarget | -1.33 | 0 | NA | |
| 124 | hsa-miR-20a-3p | ATRNL1 | -0.22 | 0.17987 | -0.74 | 0.02081 | MirTarget; mirMAP | -0.21 | 0.00197 | NA | |
| 125 | hsa-miR-106b-5p | ATRX | 1.22 | 0 | -0.59 | 0 | mirMAP | -0.26 | 0 | NA | |
| 126 | hsa-miR-106b-5p | ATXN1 | 1.22 | 0 | -0.11 | 0.17245 | miRNATAP | -0.25 | 0 | NA | |
| 127 | hsa-miR-93-5p | ATXN1 | 1.02 | 0 | -0.11 | 0.17245 | miRNAWalker2 validate; miRNATAP | -0.26 | 0 | NA | |
| 128 | hsa-miR-106b-5p | ATXN1L | 1.22 | 0 | -0.97 | 0 | miRNATAP | -0.18 | 0 | NA | |
| 129 | hsa-miR-93-5p | ATXN1L | 1.02 | 0 | -0.97 | 0 | miRNATAP | -0.19 | 0 | NA | |
| 130 | hsa-miR-106b-5p | B3GALT2 | 1.22 | 0 | -1.54 | 0 | MirTarget | -0.32 | 0 | NA | |
| 131 | hsa-miR-93-5p | B3GALT2 | 1.02 | 0 | -1.54 | 0 | MirTarget | -0.46 | 0 | NA | |
| 132 | hsa-miR-106b-5p | BACH1 | 1.22 | 0 | -0.49 | 0 | mirMAP | -0.15 | 0 | NA | |
| 133 | hsa-miR-93-5p | BACH1 | 1.02 | 0 | -0.49 | 0 | mirMAP | -0.18 | 0 | NA | |
| 134 | hsa-miR-93-5p | BAZ2A | 1.02 | 0 | -0.1 | 0.10091 | miRNAWalker2 validate | -0.14 | 0 | NA | |
| 135 | hsa-miR-106b-5p | BBX | 1.22 | 0 | -0.61 | 0 | MirTarget; miRNATAP | -0.2 | 0 | NA | |
| 136 | hsa-miR-93-5p | BBX | 1.02 | 0 | -0.61 | 0 | MirTarget; miRNATAP | -0.21 | 0 | NA | |
| 137 | hsa-miR-106a-5p | BCAS1 | 0.57 | 0.00015 | 1.43 | 1.0E-5 | mirMAP | -0.5 | 0 | NA | |
| 138 | hsa-miR-106b-5p | BCAS1 | 1.22 | 0 | 1.43 | 1.0E-5 | mirMAP | -0.92 | 0 | NA | |
| 139 | hsa-miR-93-5p | BCAS1 | 1.02 | 0 | 1.43 | 1.0E-5 | mirMAP | -0.87 | 0 | NA | |
| 140 | hsa-miR-20a-3p | BCL2 | -0.22 | 0.17987 | -0.9 | 0 | mirMAP | -0.14 | 0.0001 | NA | |
| 141 | hsa-miR-106b-5p | BCL2L2 | 1.22 | 0 | -0.92 | 0 | miRNATAP | -0.21 | 0 | NA | |
| 142 | hsa-miR-93-5p | BCL2L2 | 1.02 | 0 | -0.92 | 0 | miRNATAP | -0.18 | 0 | NA | |
| 143 | hsa-miR-106b-5p | BCL6 | 1.22 | 0 | -1.26 | 0 | miRNATAP | -0.29 | 0 | NA | |
| 144 | hsa-miR-106b-5p | BHLHE41 | 1.22 | 0 | -1.53 | 0 | miRNATAP | -0.31 | 0 | NA | |
| 145 | hsa-miR-93-5p | BHLHE41 | 1.02 | 0 | -1.53 | 0 | miRNATAP | -0.35 | 0 | NA | |
| 146 | hsa-miR-106b-5p | BMP2 | 1.22 | 0 | -2.66 | 0 | miRNATAP | -0.27 | 1.0E-5 | NA | |
| 147 | hsa-miR-106b-5p | BMPR2 | 1.22 | 0 | -0.62 | 0 | MirTarget; miRNATAP | -0.28 | 0 | NA | |
| 148 | hsa-miR-93-5p | BMPR2 | 1.02 | 0 | -0.62 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.3 | 0 | NA | |
| 149 | hsa-miR-106a-5p | BNC2 | 0.57 | 0.00015 | -0.78 | 0 | miRNATAP | -0.17 | 0 | NA | |
| 150 | hsa-miR-106b-5p | BNC2 | 1.22 | 0 | -0.78 | 0 | miRNATAP | -0.57 | 0 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | POSITIVE REGULATION OF GENE EXPRESSION | 148 | 1733 | 1.988e-14 | 9.248e-11 |
| 2 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 150 | 1805 | 1.197e-13 | 2.784e-10 |
| 3 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 148 | 1784 | 2.074e-13 | 3.217e-10 |
| 4 | NEUROGENESIS | 120 | 1402 | 7.589e-12 | 8.827e-09 |
| 5 | RESPONSE TO ENDOGENOUS STIMULUS | 122 | 1450 | 1.539e-11 | 1.194e-08 |
| 6 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 94 | 1004 | 1.521e-11 | 1.194e-08 |
| 7 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 78 | 788 | 6.814e-11 | 3.963e-08 |
| 8 | CIRCULATORY SYSTEM DEVELOPMENT | 78 | 788 | 6.814e-11 | 3.963e-08 |
| 9 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 71 | 689 | 8.577e-11 | 4.435e-08 |
| 10 | INTRACELLULAR SIGNAL TRANSDUCTION | 126 | 1572 | 1.68e-10 | 7.815e-08 |
| 11 | CELL DEVELOPMENT | 117 | 1426 | 2.036e-10 | 8.614e-08 |
| 12 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 130 | 1656 | 3.211e-10 | 1.245e-07 |
| 13 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 136 | 1791 | 1.063e-09 | 3.694e-07 |
| 14 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 89 | 1008 | 1.111e-09 | 3.694e-07 |
| 15 | HEART DEVELOPMENT | 52 | 466 | 2.077e-09 | 6.038e-07 |
| 16 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 125 | 1618 | 1.998e-09 | 6.038e-07 |
| 17 | TISSUE DEVELOPMENT | 119 | 1518 | 2.206e-09 | 6.038e-07 |
| 18 | REGULATION OF PROTEIN MODIFICATION PROCESS | 130 | 1710 | 2.469e-09 | 6.382e-07 |
| 19 | REGULATION OF CELL DIFFERENTIATION | 116 | 1492 | 5.802e-09 | 1.421e-06 |
| 20 | REGULATION OF GTPASE ACTIVITY | 65 | 673 | 7.514e-09 | 1.748e-06 |
| 21 | REGULATION OF SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 36 | 278 | 1.47e-08 | 3.256e-06 |
| 22 | NEURON DIFFERENTIATION | 77 | 874 | 1.745e-08 | 3.691e-06 |
| 23 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 134 | 1848 | 2.543e-08 | 5.144e-06 |
| 24 | REGULATION OF HYDROLASE ACTIVITY | 102 | 1327 | 9.43e-08 | 1.828e-05 |
| 25 | EPITHELIUM DEVELOPMENT | 79 | 945 | 1.058e-07 | 1.969e-05 |
| 26 | BONE DEVELOPMENT | 24 | 156 | 1.523e-07 | 2.726e-05 |
| 27 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 112 | 1518 | 1.754e-07 | 3.022e-05 |
| 28 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 64 | 724 | 2.614e-07 | 4.343e-05 |
| 29 | REGULATION OF KINASE ACTIVITY | 67 | 776 | 3.23e-07 | 5.182e-05 |
| 30 | SINGLE ORGANISM BEHAVIOR | 41 | 384 | 3.511e-07 | 5.446e-05 |
| 31 | BEHAVIOR | 50 | 516 | 3.803e-07 | 5.709e-05 |
| 32 | NEURON DEVELOPMENT | 61 | 687 | 4.317e-07 | 6.277e-05 |
| 33 | RESPONSE TO GROWTH FACTOR | 47 | 475 | 4.682e-07 | 6.601e-05 |
| 34 | MUSCLE STRUCTURE DEVELOPMENT | 44 | 432 | 4.934e-07 | 6.617e-05 |
| 35 | CELLULAR RESPONSE TO OXYGEN LEVELS | 22 | 143 | 4.977e-07 | 6.617e-05 |
| 36 | NEGATIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 64 | 740 | 5.634e-07 | 7.282e-05 |
| 37 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 72 | 872 | 6.348e-07 | 7.984e-05 |
| 38 | RESPONSE TO OXYGEN LEVELS | 35 | 311 | 7.398e-07 | 9.059e-05 |
| 39 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 118 | 1672 | 7.939e-07 | 9.472e-05 |
| 40 | REGULATION OF RAS PROTEIN SIGNAL TRANSDUCTION | 25 | 184 | 9.455e-07 | 0.00011 |
| 41 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 101 | 1381 | 1.124e-06 | 0.0001276 |
| 42 | ORGAN MORPHOGENESIS | 69 | 841 | 1.374e-06 | 0.0001523 |
| 43 | POSITIVE REGULATION OF CELL COMMUNICATION | 109 | 1532 | 1.508e-06 | 0.000161 |
| 44 | RESPONSE TO HORMONE | 72 | 893 | 1.523e-06 | 0.000161 |
| 45 | PROTEIN PHOSPHORYLATION | 75 | 944 | 1.586e-06 | 0.000164 |
| 46 | POSITIVE REGULATION OF CATABOLIC PROCESS | 40 | 395 | 1.877e-06 | 0.0001878 |
| 47 | VASCULATURE DEVELOPMENT | 45 | 469 | 1.896e-06 | 0.0001878 |
| 48 | SKELETAL SYSTEM DEVELOPMENT | 44 | 455 | 2.004e-06 | 0.0001943 |
| 49 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 80 | 1036 | 2.122e-06 | 0.0001975 |
| 50 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 80 | 1036 | 2.122e-06 | 0.0001975 |
| 51 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 86 | 1142 | 2.382e-06 | 0.0002174 |
| 52 | POSITIVE REGULATION OF HYDROLASE ACTIVITY | 72 | 905 | 2.461e-06 | 0.0002202 |
| 53 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 49 | 541 | 3.445e-06 | 0.0002968 |
| 54 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 49 | 541 | 3.445e-06 | 0.0002968 |
| 55 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 62 | 750 | 3.75e-06 | 0.0003061 |
| 56 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 42 | 437 | 3.986e-06 | 0.0003061 |
| 57 | REGULATION OF CELL PROJECTION ORGANIZATION | 50 | 558 | 3.71e-06 | 0.0003061 |
| 58 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 47 | 513 | 4.002e-06 | 0.0003061 |
| 59 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 105 | 1492 | 3.909e-06 | 0.0003061 |
| 60 | NEGATIVE REGULATION OF PHOSPHORYLATION | 41 | 422 | 3.969e-06 | 0.0003061 |
| 61 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 46 | 498 | 4.078e-06 | 0.0003061 |
| 62 | NEGATIVE REGULATION OF CELL COMMUNICATION | 88 | 1192 | 4.015e-06 | 0.0003061 |
| 63 | REGULATION OF CELL DEVELOPMENT | 67 | 836 | 4.35e-06 | 0.0003213 |
| 64 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 66 | 823 | 5.029e-06 | 0.0003656 |
| 65 | POSITIVE REGULATION OF CELL DEVELOPMENT | 44 | 472 | 5.211e-06 | 0.000373 |
| 66 | RESPONSE TO NITROGEN COMPOUND | 68 | 859 | 5.582e-06 | 0.0003935 |
| 67 | TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 24 | 190 | 5.669e-06 | 0.0003937 |
| 68 | CELLULAR RESPONSE TO HORMONE STIMULUS | 49 | 552 | 5.997e-06 | 0.0004103 |
| 69 | UROGENITAL SYSTEM DEVELOPMENT | 32 | 299 | 6.359e-06 | 0.0004288 |
| 70 | REGULATION OF SYSTEM PROCESS | 46 | 507 | 6.539e-06 | 0.0004347 |
| 71 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 64 | 799 | 7.279e-06 | 0.0004704 |
| 72 | PROTEIN LOCALIZATION | 121 | 1805 | 7.192e-06 | 0.0004704 |
| 73 | DEVELOPMENTAL MATURATION | 24 | 193 | 7.438e-06 | 0.0004741 |
| 74 | REGULATION OF RHO PROTEIN SIGNAL TRANSDUCTION | 17 | 109 | 8.028e-06 | 0.0005048 |
| 75 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 62 | 771 | 8.968e-06 | 0.0005564 |
| 76 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 98 | 1395 | 9.092e-06 | 0.0005567 |
| 77 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 127 | 1929 | 9.98e-06 | 0.0006031 |
| 78 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 32 | 306 | 1.026e-05 | 0.0006121 |
| 79 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 91 | 1275 | 1.054e-05 | 0.0006206 |
| 80 | MUSCLE CELL DIFFERENTIATION | 27 | 237 | 1.068e-05 | 0.0006212 |
| 81 | VESICLE ORGANIZATION | 30 | 280 | 1.201e-05 | 0.0006897 |
| 82 | REGULATION OF CELL MORPHOGENESIS | 48 | 552 | 1.286e-05 | 0.0007298 |
| 83 | REGULATION OF NEURON DIFFERENTIATION | 48 | 554 | 1.414e-05 | 0.0007854 |
| 84 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 76 | 1021 | 1.418e-05 | 0.0007854 |
| 85 | REGULATION OF ACTIN FILAMENT BASED PROCESS | 32 | 312 | 1.524e-05 | 0.0008341 |
| 86 | RHYTHMIC PROCESS | 31 | 298 | 1.559e-05 | 0.0008434 |
| 87 | FOREBRAIN DEVELOPMENT | 35 | 357 | 1.642e-05 | 0.000878 |
| 88 | ACTIVATION OF MAPKK ACTIVITY | 11 | 52 | 1.719e-05 | 0.000909 |
| 89 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER INVOLVED IN CELLULAR RESPONSE TO CHEMICAL STIMULUS | 8 | 27 | 1.783e-05 | 0.000932 |
| 90 | NEGATIVE REGULATION OF GENE EXPRESSION | 102 | 1493 | 1.89e-05 | 0.0009773 |
| 91 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 82 | 1135 | 1.913e-05 | 0.0009781 |
| 92 | NEURON PROJECTION DEVELOPMENT | 47 | 545 | 1.953e-05 | 0.0009879 |
| 93 | REGULATION OF TRANSFERASE ACTIVITY | 71 | 946 | 2.093e-05 | 0.001047 |
| 94 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 31 | 303 | 2.163e-05 | 0.001071 |
| 95 | REGULATION OF DEVELOPMENTAL GROWTH | 30 | 289 | 2.215e-05 | 0.001074 |
| 96 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 103 | 1517 | 2.212e-05 | 0.001074 |
| 97 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 42 | 470 | 2.346e-05 | 0.001126 |
| 98 | REGULATION OF CATABOLIC PROCESS | 58 | 731 | 2.573e-05 | 0.001222 |
| 99 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 51 | 616 | 2.621e-05 | 0.001232 |
| 100 | TUBE DEVELOPMENT | 47 | 552 | 2.696e-05 | 0.001255 |
| 101 | COGNITION | 27 | 251 | 3.008e-05 | 0.001386 |
| 102 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 71 | 957 | 3.043e-05 | 0.001388 |
| 103 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 82 | 1152 | 3.237e-05 | 0.001462 |
| 104 | MEMORY | 15 | 98 | 3.396e-05 | 0.001505 |
| 105 | NEGATIVE REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 15 | 98 | 3.396e-05 | 0.001505 |
| 106 | EPITHELIAL CELL DIFFERENTIATION | 43 | 495 | 3.688e-05 | 0.001619 |
| 107 | HOMEOSTATIC PROCESS | 92 | 1337 | 3.821e-05 | 0.001662 |
| 108 | STRIATED MUSCLE CELL DIFFERENTIATION | 21 | 173 | 3.905e-05 | 0.001667 |
| 109 | REGULATION OF SODIUM ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 9 | 38 | 3.874e-05 | 0.001667 |
| 110 | REGULATION OF MAPK CASCADE | 53 | 660 | 4.163e-05 | 0.001745 |
| 111 | REGULATION OF ORGANELLE ORGANIZATION | 83 | 1178 | 4.139e-05 | 0.001745 |
| 112 | POSITIVE REGULATION OF KINASE ACTIVITY | 42 | 482 | 4.219e-05 | 0.001753 |
| 113 | TELENCEPHALON DEVELOPMENT | 25 | 228 | 4.265e-05 | 0.001756 |
| 114 | REGULATION OF TRANSPORT | 117 | 1804 | 4.489e-05 | 0.001832 |
| 115 | REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 25 | 229 | 4.587e-05 | 0.001856 |
| 116 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 39 | 437 | 4.679e-05 | 0.001877 |
| 117 | ORGAN GROWTH | 12 | 68 | 4.911e-05 | 0.001953 |
| 118 | KIDNEY EPITHELIUM DEVELOPMENT | 17 | 125 | 4.985e-05 | 0.001966 |
| 119 | SODIUM ION TRANSMEMBRANE TRANSPORT | 14 | 90 | 5.126e-05 | 0.001971 |
| 120 | POSITIVE REGULATION OF MAPK CASCADE | 41 | 470 | 5.06e-05 | 0.001971 |
| 121 | MESONEPHROS DEVELOPMENT | 14 | 90 | 5.126e-05 | 0.001971 |
| 122 | BONE MORPHOGENESIS | 13 | 79 | 5.169e-05 | 0.001971 |
| 123 | CELLULAR RESPONSE TO PEPTIDE | 28 | 274 | 5.437e-05 | 0.002057 |
| 124 | MUSCLE TISSUE DEVELOPMENT | 28 | 275 | 5.798e-05 | 0.002128 |
| 125 | ENSHEATHMENT OF NEURONS | 14 | 91 | 5.809e-05 | 0.002128 |
| 126 | AXON ENSHEATHMENT | 14 | 91 | 5.809e-05 | 0.002128 |
| 127 | BLOOD VESSEL MORPHOGENESIS | 34 | 364 | 5.772e-05 | 0.002128 |
| 128 | CELLULAR RESPONSE TO NITROGEN COMPOUND | 43 | 505 | 5.868e-05 | 0.002133 |
| 129 | CHEMICAL HOMEOSTASIS | 65 | 874 | 6.149e-05 | 0.002218 |
| 130 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 77 | 1087 | 6.642e-05 | 0.002377 |
| 131 | SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 33 | 352 | 6.873e-05 | 0.002441 |
| 132 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 92 | 1360 | 7.146e-05 | 0.002507 |
| 133 | CELL MORPHOGENESIS INVOLVED IN NEURON DIFFERENTIATION | 34 | 368 | 7.167e-05 | 0.002507 |
| 134 | REGULATION OF CELL DEATH | 98 | 1472 | 7.52e-05 | 0.002611 |
| 135 | HEAD DEVELOPMENT | 55 | 709 | 7.596e-05 | 0.002618 |
| 136 | NEURON PROJECTION MORPHOGENESIS | 36 | 402 | 8.439e-05 | 0.002832 |
| 137 | RESPONSE TO ABIOTIC STIMULUS | 73 | 1024 | 8.452e-05 | 0.002832 |
| 138 | REGULATION OF CYTOPLASMIC TRANSPORT | 41 | 481 | 8.459e-05 | 0.002832 |
| 139 | ACTION POTENTIAL | 14 | 94 | 8.357e-05 | 0.002832 |
| 140 | ACTIN FILAMENT BASED PROCESS | 39 | 450 | 8.804e-05 | 0.002926 |
| 141 | SODIUM ION TRANSPORT | 18 | 144 | 9.321e-05 | 0.003076 |
| 142 | POSITIVE REGULATION OF LOCOMOTION | 37 | 420 | 9.49e-05 | 0.00311 |
| 143 | CARDIAC SEPTUM DEVELOPMENT | 13 | 85 | 0.0001126 | 0.003614 |
| 144 | NEGATIVE REGULATION OF CELL CYCLE PHASE TRANSITION | 18 | 146 | 0.0001115 | 0.003614 |
| 145 | HEART PROCESS | 13 | 85 | 0.0001126 | 0.003614 |
| 146 | REPRODUCTIVE SYSTEM DEVELOPMENT | 36 | 408 | 0.0001137 | 0.003624 |
| 147 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 64 | 876 | 0.0001166 | 0.003692 |
| 148 | ADULT BEHAVIOR | 17 | 135 | 0.0001319 | 0.004148 |
| 149 | MULTICELLULAR ORGANISMAL SIGNALING | 16 | 123 | 0.0001404 | 0.004386 |
| 150 | CELLULAR COMPONENT MORPHOGENESIS | 65 | 900 | 0.0001434 | 0.004449 |
| 151 | POSITIVE REGULATION OF NEURON PROJECTION DEVELOPMENT | 24 | 232 | 0.0001499 | 0.004619 |
| 152 | SECRETORY GRANULE ORGANIZATION | 7 | 27 | 0.0001542 | 0.004671 |
| 153 | CELL PROJECTION ORGANIZATION | 65 | 902 | 0.0001527 | 0.004671 |
| 154 | INOSITOL LIPID MEDIATED SIGNALING | 16 | 124 | 0.0001546 | 0.004671 |
| 155 | ENDOCHONDRAL BONE MORPHOGENESIS | 9 | 45 | 0.0001584 | 0.004755 |
| 156 | PROTEIN UBIQUITINATION | 49 | 629 | 0.0001675 | 0.004944 |
| 157 | REGULATION OF CELL ADHESION | 49 | 629 | 0.0001675 | 0.004944 |
| 158 | PHOSPHORYLATION | 83 | 1228 | 0.0001679 | 0.004944 |
| 159 | RESPONSE TO STEROID HORMONE | 41 | 497 | 0.0001714 | 0.005017 |
| 160 | NEGATIVE REGULATION OF CELL CYCLE | 37 | 433 | 0.0001757 | 0.005109 |
| 161 | REGULATION OF GROWTH | 49 | 633 | 0.0001945 | 0.005586 |
| 162 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 48 | 616 | 0.0001934 | 0.005586 |
| 163 | RESPONSE TO PEPTIDE | 35 | 404 | 0.0002004 | 0.005722 |
| 164 | CONNECTIVE TISSUE DEVELOPMENT | 21 | 194 | 0.0002037 | 0.00578 |
| 165 | CARDIAC MUSCLE CELL ACTION POTENTIAL | 8 | 37 | 0.0002072 | 0.005844 |
| 166 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 44 | 552 | 0.0002161 | 0.006058 |
| 167 | MUSCLE CELL DEVELOPMENT | 16 | 128 | 0.0002243 | 0.006251 |
| 168 | REGULATION OF CIRCADIAN RHYTHM | 14 | 103 | 0.0002263 | 0.006269 |
| 169 | REGULATION OF BINDING | 27 | 283 | 0.0002298 | 0.006327 |
| 170 | RESPONSE TO MECHANICAL STIMULUS | 22 | 210 | 0.0002326 | 0.006365 |
| 171 | LOCOMOTION | 76 | 1114 | 0.0002395 | 0.006517 |
| 172 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 35 | 408 | 0.0002417 | 0.006539 |
| 173 | ADULT LOCOMOTORY BEHAVIOR | 12 | 80 | 0.0002469 | 0.006581 |
| 174 | APPENDAGE DEVELOPMENT | 19 | 169 | 0.0002476 | 0.006581 |
| 175 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 122 | 1977 | 0.0002503 | 0.006581 |
| 176 | CARDIAC MUSCLE CELL CONTRACTION | 7 | 29 | 0.0002503 | 0.006581 |
| 177 | LIMB DEVELOPMENT | 19 | 169 | 0.0002476 | 0.006581 |
| 178 | PERIPHERAL NERVOUS SYSTEM DEVELOPMENT | 11 | 69 | 0.0002571 | 0.006721 |
| 179 | POSITIVE REGULATION OF CELL PROLIFERATION | 59 | 814 | 0.0002633 | 0.006845 |
| 180 | REGULATION OF SODIUM ION TRANSMEMBRANE TRANSPORT | 9 | 48 | 0.0002654 | 0.006861 |
| 181 | MEMBRANE DEPOLARIZATION DURING CARDIAC MUSCLE CELL ACTION POTENTIAL | 5 | 14 | 0.0002696 | 0.00693 |
| 182 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 47 | 609 | 0.0002794 | 0.007142 |
| 183 | SENSORY ORGAN DEVELOPMENT | 40 | 493 | 0.000288 | 0.007324 |
| 184 | VASCULOGENESIS | 10 | 59 | 0.0002914 | 0.007369 |
| 185 | VESICLE CYTOSKELETAL TRAFFICKING | 8 | 39 | 0.0003047 | 0.007665 |
| 186 | EPITHELIAL CELL DEVELOPMENT | 20 | 186 | 0.0003123 | 0.007728 |
| 187 | REGULATION OF VESICLE MEDIATED TRANSPORT | 38 | 462 | 0.00031 | 0.007728 |
| 188 | POSITIVE REGULATION OF SMOOTH MUSCLE CELL MIGRATION | 7 | 30 | 0.0003138 | 0.007728 |
| 189 | RESPONSE TO BMP | 13 | 94 | 0.0003156 | 0.007728 |
| 190 | CELLULAR RESPONSE TO BMP STIMULUS | 13 | 94 | 0.0003156 | 0.007728 |
| 191 | SKELETAL SYSTEM MORPHOGENESIS | 21 | 201 | 0.0003315 | 0.008075 |
| 192 | REGULATION OF CELL SUBSTRATE ADHESION | 19 | 173 | 0.0003342 | 0.008099 |
| 193 | CELLULAR RESPONSE TO INSULIN STIMULUS | 17 | 146 | 0.0003409 | 0.008219 |
| 194 | REGULATION OF CELLULAR LOCALIZATION | 84 | 1277 | 0.0003665 | 0.00879 |
| 195 | REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 17 | 147 | 0.0003696 | 0.008819 |
| 196 | CELLULAR RESPONSE TO GLUCOSE STARVATION | 7 | 31 | 0.0003896 | 0.009249 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | ENZYME BINDING | 143 | 1737 | 1.034e-12 | 9.609e-10 |
| 2 | NUCLEIC ACID BINDING TRANSCRIPTION FACTOR ACTIVITY | 102 | 1199 | 4.808e-10 | 2.233e-07 |
| 3 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 42 | 328 | 1.307e-09 | 4.047e-07 |
| 4 | RNA POLYMERASE II TRANSCRIPTION FACTOR ACTIVITY SEQUENCE SPECIFIC DNA BINDING | 62 | 629 | 7.844e-09 | 1.822e-06 |
| 5 | MOLECULAR FUNCTION REGULATOR | 105 | 1353 | 3.676e-08 | 6.831e-06 |
| 6 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION REGULATORY REGION SEQUENCE SPECIFIC BINDING | 36 | 315 | 3.541e-07 | 5.142e-05 |
| 7 | ENZYME REGULATOR ACTIVITY | 78 | 959 | 3.874e-07 | 5.142e-05 |
| 8 | GTPASE BINDING | 34 | 295 | 6.127e-07 | 7.115e-05 |
| 9 | SEQUENCE SPECIFIC DNA BINDING | 81 | 1037 | 1.168e-06 | 0.0001206 |
| 10 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 28 | 226 | 1.429e-06 | 0.0001207 |
| 11 | TRANSCRIPTION FACTOR BINDING | 49 | 524 | 1.4e-06 | 0.0001207 |
| 12 | MACROMOLECULAR COMPLEX BINDING | 101 | 1399 | 2.007e-06 | 0.0001554 |
| 13 | NUCLEOSIDE TRIPHOSPHATASE REGULATOR ACTIVITY | 35 | 329 | 2.719e-06 | 0.0001943 |
| 14 | SMAD BINDING | 14 | 72 | 3.614e-06 | 0.0002298 |
| 15 | BINDING BRIDGING | 23 | 173 | 3.71e-06 | 0.0002298 |
| 16 | DOUBLE STRANDED DNA BINDING | 62 | 764 | 6.745e-06 | 0.0003916 |
| 17 | REGULATORY REGION NUCLEIC ACID BINDING | 65 | 818 | 7.986e-06 | 0.0004122 |
| 18 | RAB GTPASE BINDING | 18 | 120 | 7.688e-06 | 0.0004122 |
| 19 | PROTEIN HOMODIMERIZATION ACTIVITY | 59 | 722 | 9.159e-06 | 0.0004254 |
| 20 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 19 | 133 | 8.965e-06 | 0.0004254 |
| 21 | CALCIUM ION BINDING | 57 | 697 | 1.271e-05 | 0.0005621 |
| 22 | RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 16 | 104 | 1.757e-05 | 0.0007421 |
| 23 | ENZYME ACTIVATOR ACTIVITY | 42 | 471 | 2.467e-05 | 0.0009963 |
| 24 | PROTEIN DIMERIZATION ACTIVITY | 82 | 1149 | 2.954e-05 | 0.001144 |
| 25 | MAP KINASE KINASE KINASE ACTIVITY | 7 | 22 | 3.611e-05 | 0.001342 |
| 26 | CYTOSKELETAL PROTEIN BINDING | 62 | 819 | 5.476e-05 | 0.001956 |
| 27 | CORE PROMOTER PROXIMAL REGION DNA BINDING | 34 | 371 | 8.407e-05 | 0.002893 |
| 28 | UBIQUITIN LIKE PROTEIN TRANSFERASE ACTIVITY | 37 | 420 | 9.49e-05 | 0.003149 |
| 29 | ACTIN BINDING | 35 | 393 | 0.0001174 | 0.003592 |
| 30 | KINASE ACTIVITY | 62 | 842 | 0.0001199 | 0.003592 |
| 31 | TRANSCRIPTIONAL REPRESSOR ACTIVITY RNA POLYMERASE II ACTIVATING TRANSCRIPTION FACTOR BINDING | 10 | 53 | 0.0001156 | 0.003592 |
| 32 | TRANSCRIPTION FACTOR ACTIVITY PROTEIN BINDING | 47 | 588 | 0.0001249 | 0.003626 |
| 33 | PROTEIN KINASE ACTIVITY | 50 | 640 | 0.0001345 | 0.003786 |
| 34 | X1 PHOSPHATIDYLINOSITOL BINDING | 6 | 19 | 0.0001396 | 0.003814 |
| 35 | RNA POLYMERASE II ACTIVATING TRANSCRIPTION FACTOR BINDING | 8 | 36 | 0.0001691 | 0.004488 |
| 36 | VOLTAGE GATED SODIUM CHANNEL ACTIVITY | 6 | 20 | 0.0001918 | 0.004949 |
| 37 | ACTIVATING TRANSCRIPTION FACTOR BINDING | 10 | 57 | 0.0002174 | 0.005458 |
| 38 | RECEPTOR SIGNALING PROTEIN SERINE THREONINE KINASE ACTIVITY | 13 | 92 | 0.0002542 | 0.006215 |
| 39 | RAS GUANYL NUCLEOTIDE EXCHANGE FACTOR ACTIVITY | 23 | 228 | 0.0002946 | 0.007018 |
| 40 | RECEPTOR SIGNALING PROTEIN ACTIVITY | 19 | 172 | 0.0003104 | 0.007209 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | NEURON PART | 100 | 1265 | 3.252e-08 | 1.899e-05 |
| 2 | NEURON PROJECTION | 79 | 942 | 9.269e-08 | 2.706e-05 |
| 3 | CELL PROJECTION | 123 | 1786 | 1.58e-06 | 0.0003076 |
| 4 | MEMBRANE REGION | 85 | 1134 | 3.287e-06 | 0.0003839 |
| 5 | EARLY ENDOSOME | 33 | 301 | 2.715e-06 | 0.0003839 |
| 6 | ENDOSOME | 64 | 793 | 5.715e-06 | 0.0005563 |
| 7 | PLASMA MEMBRANE RAFT | 15 | 86 | 6.72e-06 | 0.0005606 |
| 8 | ACTIN CYTOSKELETON | 41 | 444 | 1.36e-05 | 0.0008845 |
| 9 | GOLGI APPARATUS | 100 | 1445 | 1.363e-05 | 0.0008845 |
| 10 | AXON | 39 | 418 | 1.735e-05 | 0.001013 |
| 11 | PML BODY | 15 | 97 | 3e-05 | 0.001365 |
| 12 | CELL JUNCTION | 82 | 1151 | 3.14e-05 | 0.001365 |
| 13 | SYNAPSE | 59 | 754 | 3.273e-05 | 0.001365 |
| 14 | CUL3 RING UBIQUITIN LIGASE COMPLEX | 12 | 64 | 2.616e-05 | 0.001365 |
| 15 | CELL PROJECTION PART | 70 | 946 | 3.763e-05 | 0.001465 |
| 16 | VACUOLE | 83 | 1180 | 4.39e-05 | 0.001602 |
| 17 | MEMBRANE MICRODOMAIN | 29 | 288 | 5.269e-05 | 0.00181 |
| 18 | CELL LEADING EDGE | 33 | 350 | 6.152e-05 | 0.001996 |
| 19 | PLASMA MEMBRANE REGION | 68 | 929 | 6.764e-05 | 0.002079 |
| 20 | INTRACELLULAR VESICLE | 85 | 1259 | 0.0001452 | 0.00424 |
| 21 | PROTEIN KINASE COMPLEX | 13 | 90 | 0.0002034 | 0.005399 |
| 22 | SOMATODENDRITIC COMPARTMENT | 50 | 650 | 0.0001951 | 0.005399 |
| 23 | PERINUCLEAR REGION OF CYTOPLASM | 49 | 642 | 0.0002701 | 0.006573 |
| 24 | VOLTAGE GATED SODIUM CHANNEL COMPLEX | 5 | 14 | 0.0002696 | 0.006573 |
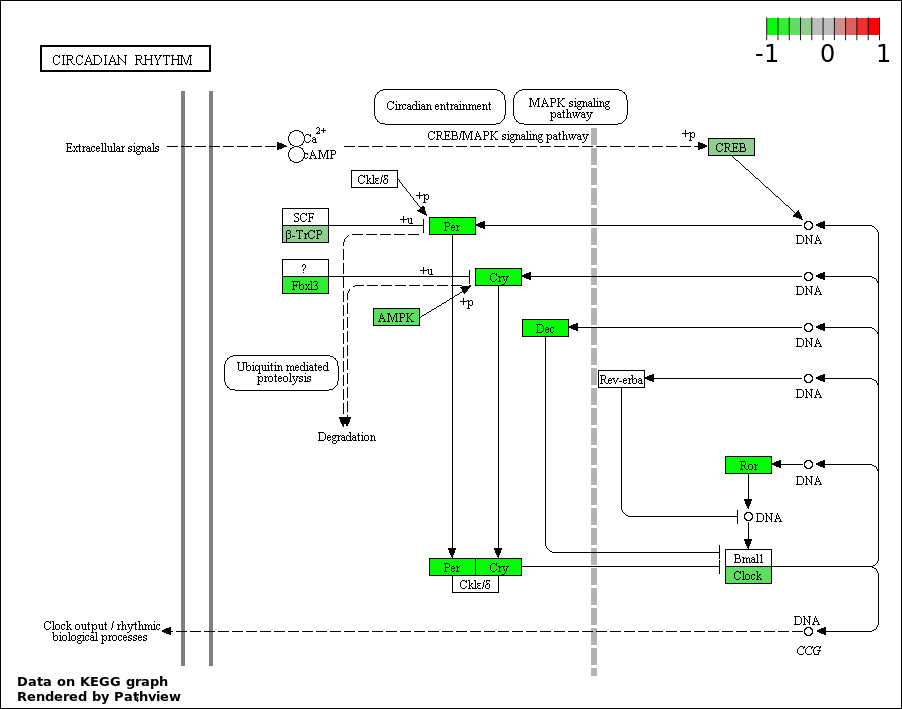
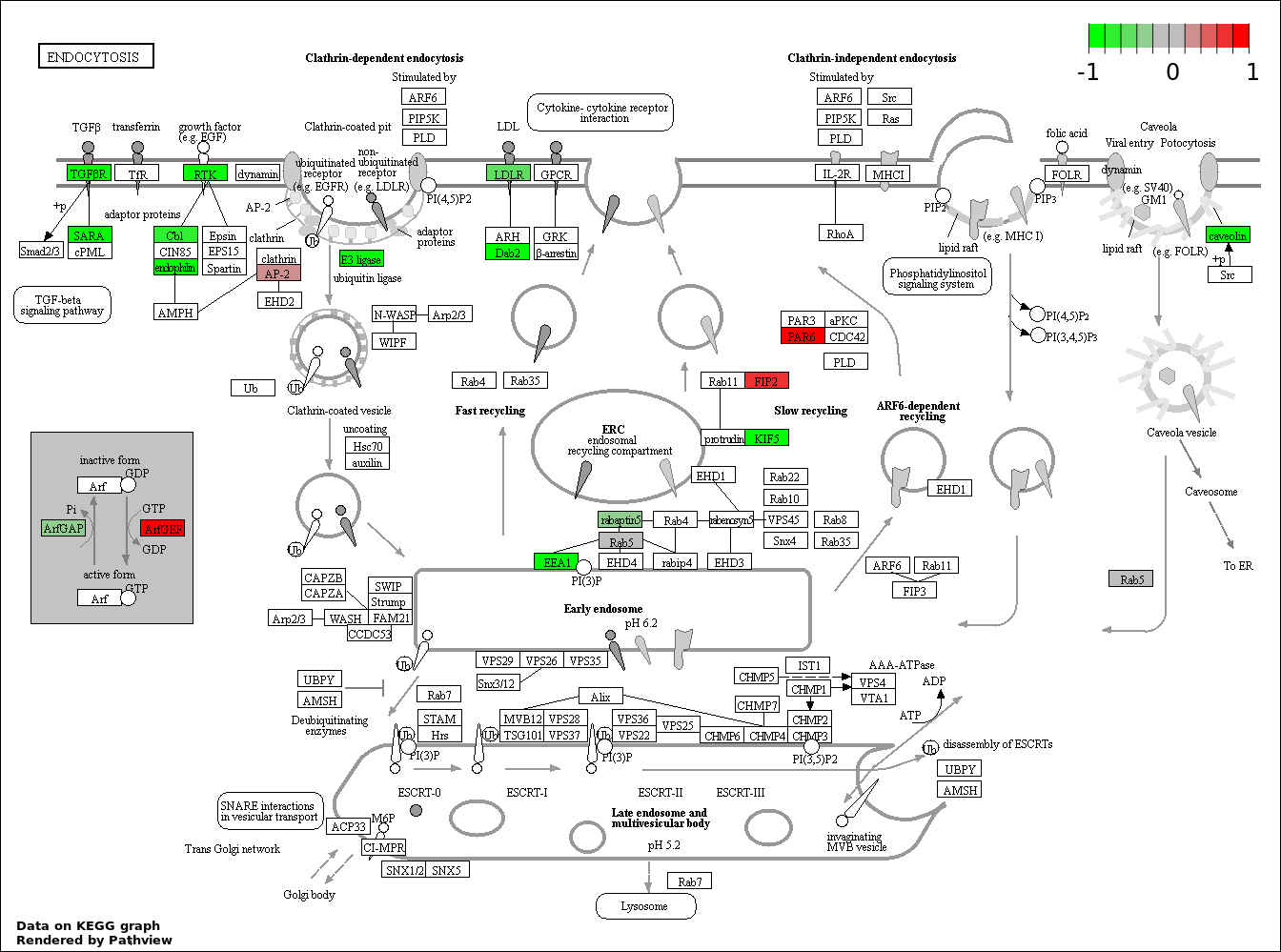
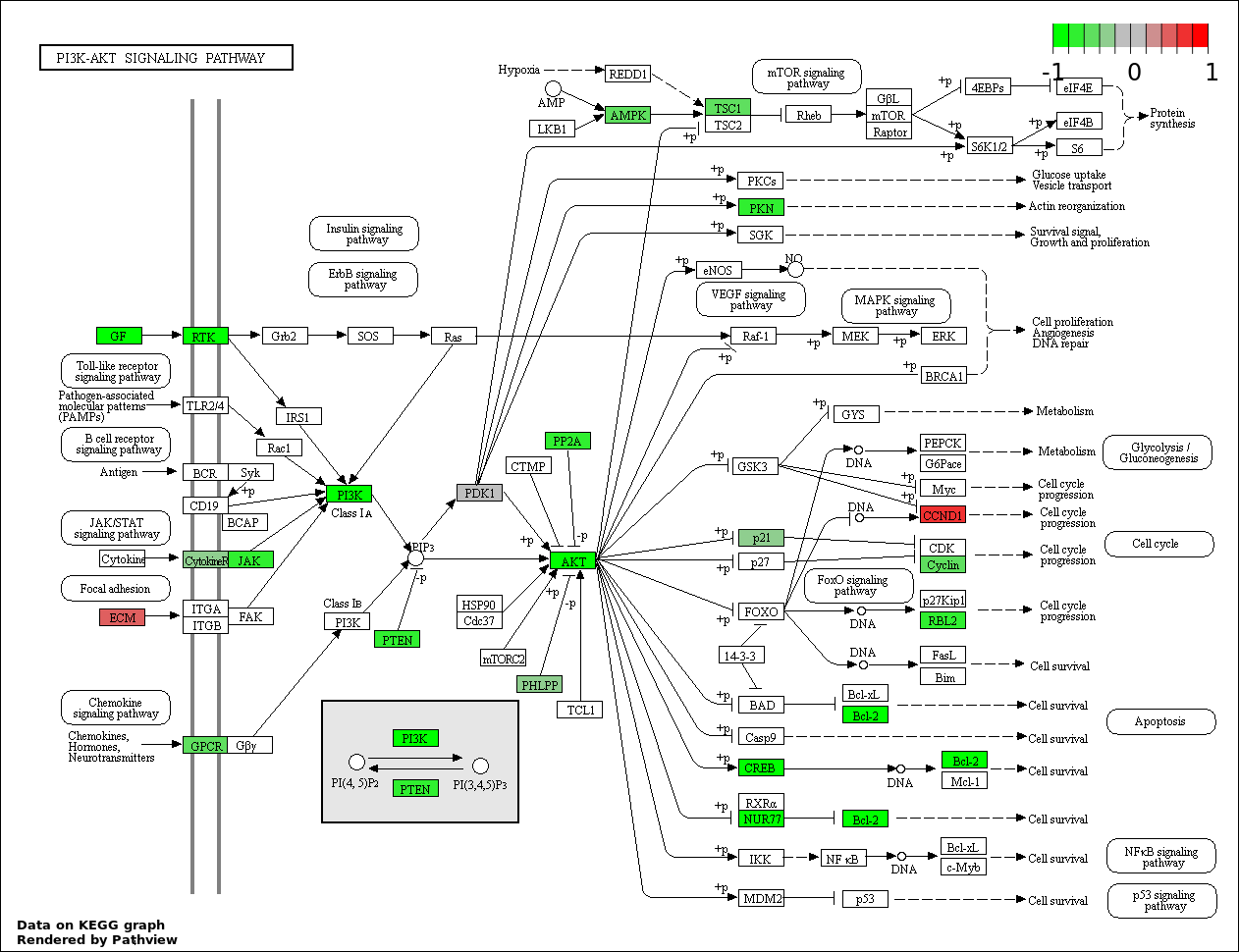
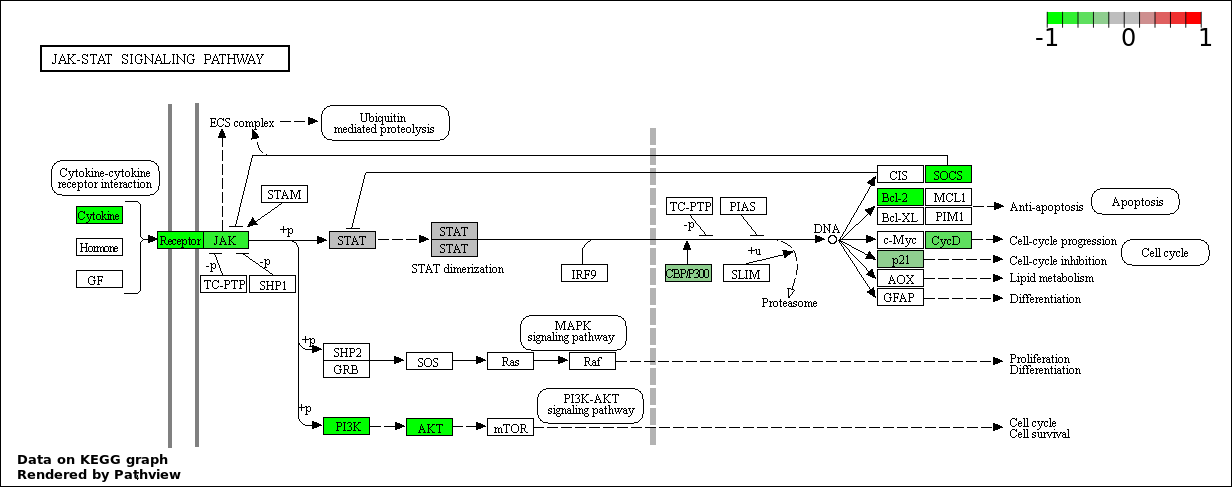
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04710_Circadian_rhythm_._mammal | 7 | 23 | 4.988e-05 | 0.004318 | |
| 2 | hsa04144_Endocytosis | 23 | 203 | 5.127e-05 | 0.004318 | |
| 3 | hsa04390_Hippo_signaling_pathway | 19 | 154 | 7.197e-05 | 0.004318 | |
| 4 | hsa04151_PI3K_AKT_signaling_pathway | 31 | 351 | 0.0003257 | 0.01466 | |
| 5 | hsa04630_Jak.STAT_signaling_pathway | 17 | 155 | 0.0006837 | 0.02461 | |
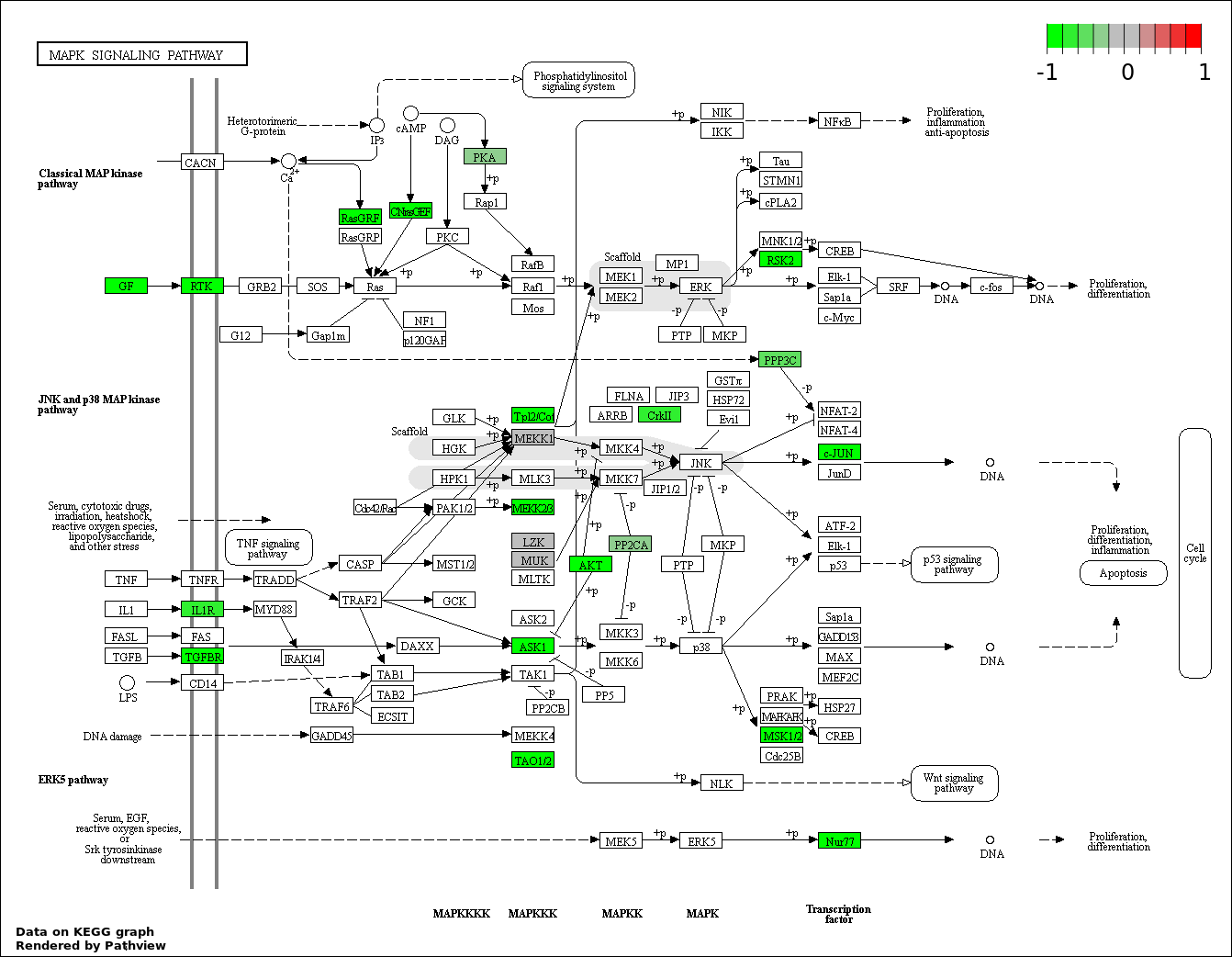
| 6 | hsa04010_MAPK_signaling_pathway | 24 | 268 | 0.001217 | 0.03651 | |
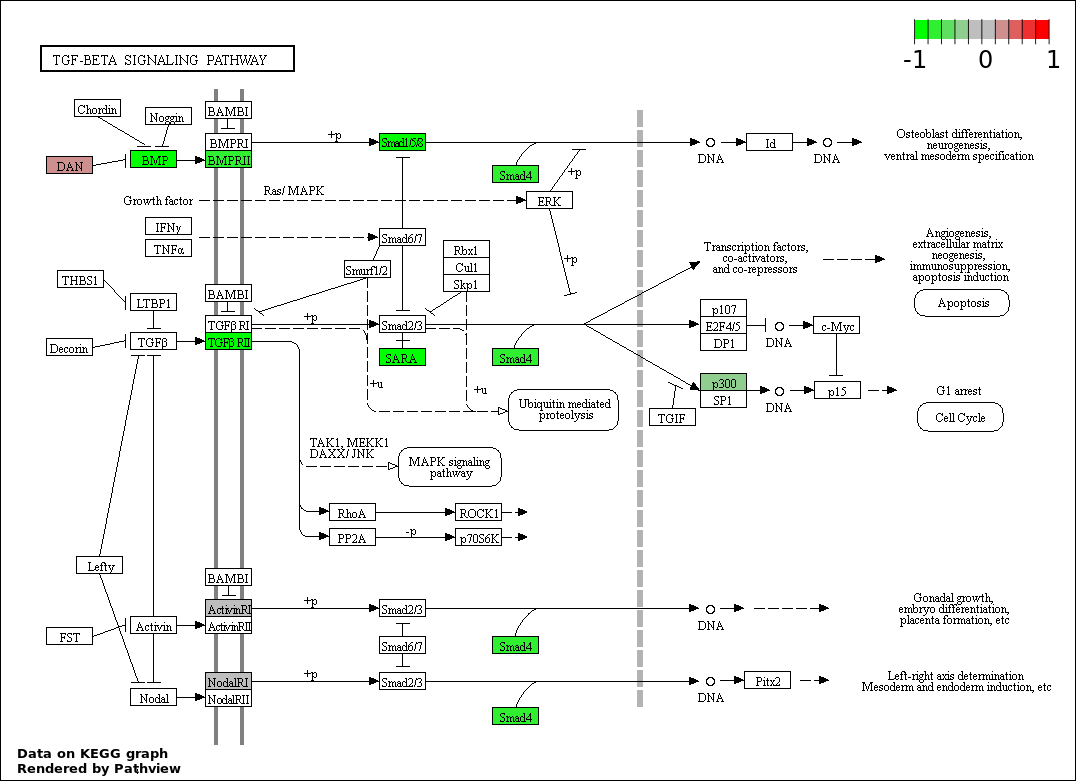
| 7 | hsa04350_TGF.beta_signaling_pathway | 11 | 85 | 0.001559 | 0.04009 | |
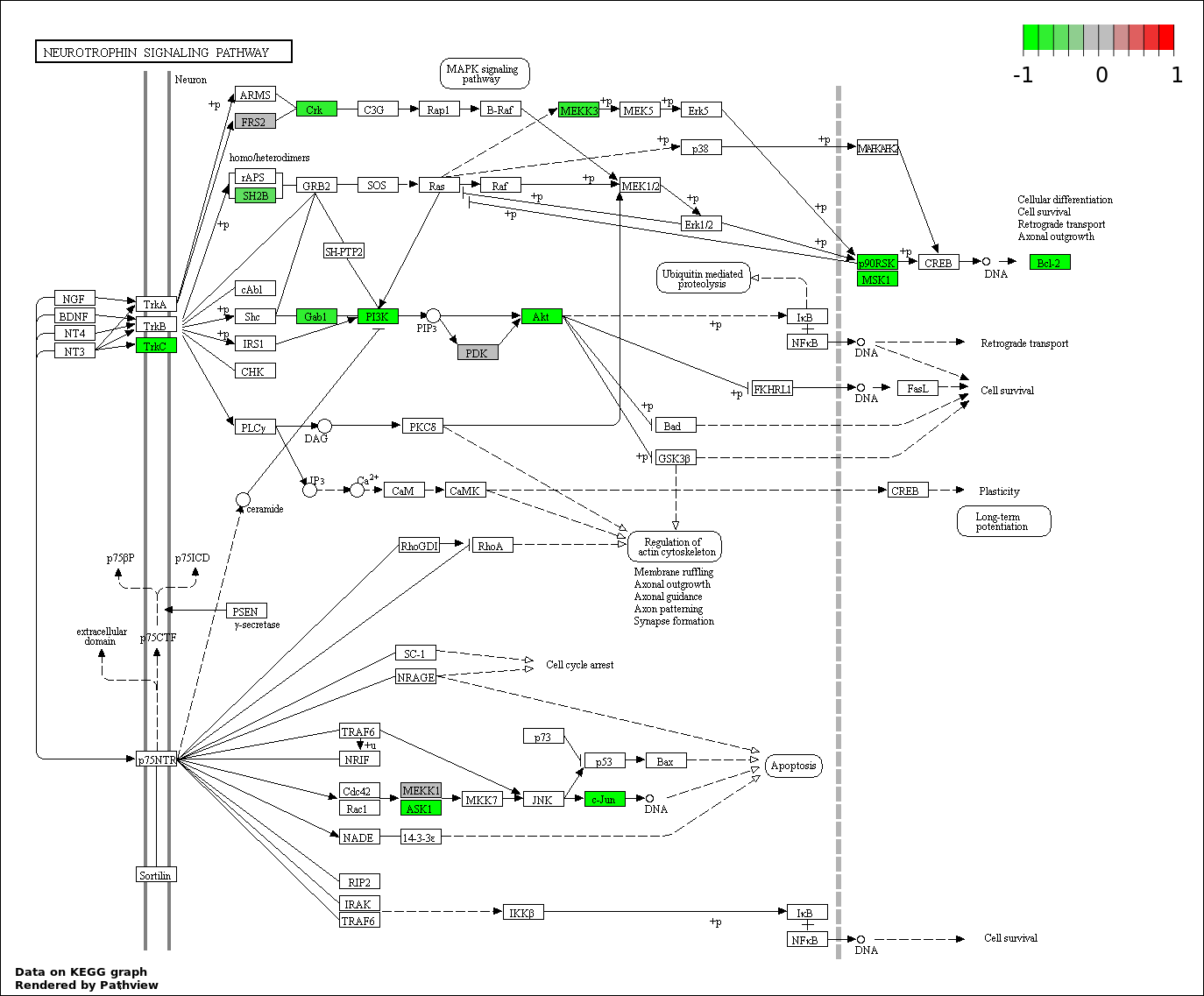
| 8 | hsa04722_Neurotrophin_signaling_pathway | 14 | 127 | 0.001873 | 0.04215 | |
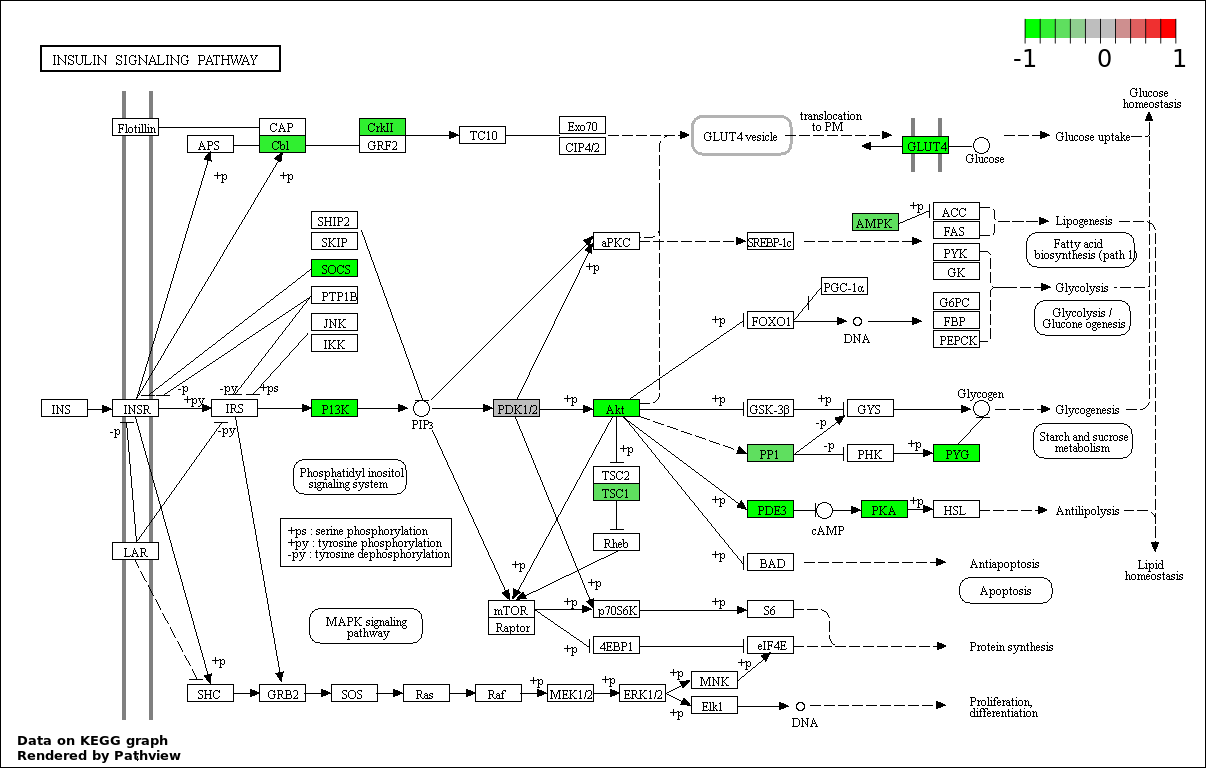
| 9 | hsa04910_Insulin_signaling_pathway | 14 | 138 | 0.004038 | 0.08077 | |
| 10 | hsa04912_GnRH_signaling_pathway | 11 | 101 | 0.006049 | 0.1089 | |
| 11 | hsa04540_Gap_junction | 10 | 90 | 0.00754 | 0.1234 | |
| 12 | hsa04150_mTOR_signaling_pathway | 7 | 52 | 0.008698 | 0.1305 | |
| 13 | hsa04510_Focal_adhesion | 17 | 200 | 0.009766 | 0.1352 | |
| 14 | hsa04920_Adipocytokine_signaling_pathway | 8 | 68 | 0.01157 | 0.1487 | |
| 15 | hsa04115_p53_signaling_pathway | 8 | 69 | 0.01258 | 0.151 | |
| 16 | hsa04962_Vasopressin.regulated_water_reabsorption | 6 | 44 | 0.01387 | 0.156 | |
| 17 | hsa04914_Progesterone.mediated_oocyte_maturation | 9 | 87 | 0.01698 | 0.1798 | |
| 18 | hsa04210_Apoptosis | 9 | 89 | 0.01945 | 0.1899 | |
| 19 | hsa04310_Wnt_signaling_pathway | 13 | 151 | 0.02004 | 0.1899 | |
| 20 | hsa00770_Pantothenate_and_CoA_biosynthesis | 3 | 16 | 0.03347 | 0.2772 | |
| 21 | hsa04114_Oocyte_meiosis | 10 | 114 | 0.03437 | 0.2772 | |
| 22 | hsa04270_Vascular_smooth_muscle_contraction | 10 | 116 | 0.03806 | 0.2772 | |
| 23 | hsa04720_Long.term_potentiation | 7 | 70 | 0.03894 | 0.2772 | |
| 24 | hsa04972_Pancreatic_secretion | 9 | 101 | 0.03983 | 0.2772 | |
| 25 | hsa04960_Aldosterone.regulated_sodium_reabsorption | 5 | 42 | 0.04034 | 0.2772 | |
| 26 | hsa00340_Histidine_metabolism | 4 | 29 | 0.04052 | 0.2772 | |
| 27 | hsa04976_Bile_secretion | 7 | 71 | 0.04159 | 0.2772 | |
| 28 | hsa04970_Salivary_secretion | 8 | 89 | 0.04875 | 0.3116 | |
| 29 | hsa04971_Gastric_acid_secretion | 7 | 74 | 0.05019 | 0.3116 | |
| 30 | hsa04380_Osteoclast_differentiation | 10 | 128 | 0.06592 | 0.3883 | |
| 31 | hsa04146_Peroxisome | 7 | 79 | 0.06688 | 0.3883 | |
| 32 | hsa04360_Axon_guidance | 10 | 130 | 0.07158 | 0.3909 | |
| 33 | hsa04014_Ras_signaling_pathway | 16 | 236 | 0.07167 | 0.3909 | |
| 34 | hsa00565_Ether_lipid_metabolism | 4 | 36 | 0.07864 | 0.4146 | |
| 35 | hsa04530_Tight_junction | 10 | 133 | 0.08061 | 0.4146 | |
| 36 | hsa04916_Melanogenesis | 8 | 101 | 0.08798 | 0.4399 | |
| 37 | hsa04012_ErbB_signaling_pathway | 7 | 87 | 0.09975 | 0.4769 | |
| 38 | hsa04120_Ubiquitin_mediated_proteolysis | 10 | 139 | 0.1007 | 0.4769 | |
| 39 | hsa04020_Calcium_signaling_pathway | 12 | 177 | 0.1079 | 0.4914 | |
| 40 | hsa04340_Hedgehog_signaling_pathway | 5 | 56 | 0.1092 | 0.4914 | |
| 41 | hsa00071_Fatty_acid_metabolism | 4 | 43 | 0.1294 | 0.5681 | |
| 42 | hsa00310_Lysine_degradation | 4 | 44 | 0.1376 | 0.5896 | |
| 43 | hsa04141_Protein_processing_in_endoplasmic_reticulum | 11 | 168 | 0.1423 | 0.5956 | |
| 44 | hsa00604_Glycosphingolipid_biosynthesis_._ganglio_series | 2 | 15 | 0.1462 | 0.5983 | |
| 45 | hsa04514_Cell_adhesion_molecules_.CAMs. | 9 | 136 | 0.1652 | 0.6606 | |
| 46 | hsa00561_Glycerolipid_metabolism | 4 | 50 | 0.1904 | 0.745 | |
| 47 | hsa00670_One_carbon_pool_by_folate | 2 | 18 | 0.1954 | 0.7484 | |
| 48 | hsa03320_PPAR_signaling_pathway | 5 | 70 | 0.2111 | 0.7756 | |
| 49 | hsa04730_Long.term_depression | 5 | 70 | 0.2111 | 0.7756 | |
| 50 | hsa04660_T_cell_receptor_signaling_pathway | 7 | 108 | 0.2189 | 0.7879 | |
| 51 | hsa00592_alpha.Linolenic_acid_metabolism | 2 | 20 | 0.2292 | 0.8089 | |
| 52 | hsa04520_Adherens_junction | 5 | 73 | 0.2361 | 0.8172 | |
| 53 | hsa04662_B_cell_receptor_signaling_pathway | 5 | 75 | 0.2532 | 0.8463 | |
| 54 | hsa04370_VEGF_signaling_pathway | 5 | 76 | 0.2618 | 0.8463 | |
| 55 | hsa00410_beta.Alanine_metabolism | 2 | 22 | 0.2633 | 0.8463 | |
| 56 | hsa00532_Glycosaminoglycan_biosynthesis_._chondroitin_sulfate | 2 | 22 | 0.2633 | 0.8463 | |
| 57 | hsa00600_Sphingolipid_metabolism | 3 | 40 | 0.2718 | 0.8585 | |
| 58 | hsa04964_Proximal_tubule_bicarbonate_reclamation | 2 | 23 | 0.2804 | 0.8674 | |
| 59 | hsa00350_Tyrosine_metabolism | 3 | 41 | 0.2843 | 0.8674 | |
| 60 | hsa00564_Glycerophospholipid_metabolism | 5 | 80 | 0.297 | 0.8777 | |
| 61 | hsa00760_Nicotinate_and_nicotinamide_metabolism | 2 | 24 | 0.2974 | 0.8777 | |
| 62 | hsa04973_Carbohydrate_digestion_and_absorption | 3 | 44 | 0.322 | 0.9256 | |
| 63 | hsa03015_mRNA_surveillance_pathway | 5 | 83 | 0.3239 | 0.9256 | |
| 64 | hsa04110_Cell_cycle | 7 | 128 | 0.3615 | 1 | |
| 65 | hsa04810_Regulation_of_actin_cytoskeleton | 11 | 214 | 0.379 | 1 | |
| 66 | hsa00512_Mucin_type_O.Glycan_biosynthesis | 2 | 30 | 0.3974 | 1 | |
| 67 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 5 | 95 | 0.4326 | 1 | |
| 68 | hsa00230_Purine_metabolism | 8 | 162 | 0.4542 | 1 | |
| 69 | hsa04142_Lysosome | 6 | 121 | 0.4711 | 1 | |
| 70 | hsa04070_Phosphatidylinositol_signaling_system | 4 | 78 | 0.4743 | 1 | |
| 71 | hsa04062_Chemokine_signaling_pathway | 9 | 189 | 0.4888 | 1 | |
| 72 | hsa00270_Cysteine_and_methionine_metabolism | 2 | 36 | 0.4904 | 1 | |
| 73 | hsa04620_Toll.like_receptor_signaling_pathway | 5 | 102 | 0.4946 | 1 | |
| 74 | hsa00280_Valine._leucine_and_isoleucine_degradation | 2 | 44 | 0.5993 | 1 | |
| 75 | hsa02010_ABC_transporters | 2 | 44 | 0.5993 | 1 | |
| 76 | hsa04610_Complement_and_coagulation_cascades | 3 | 69 | 0.6111 | 1 | |
| 77 | hsa04975_Fat_digestion_and_absorption | 2 | 46 | 0.6236 | 1 | |
| 78 | hsa03018_RNA_degradation | 3 | 71 | 0.6305 | 1 | |
| 79 | hsa04330_Notch_signaling_pathway | 2 | 47 | 0.6354 | 1 | |
| 80 | hsa00520_Amino_sugar_and_nucleotide_sugar_metabolism | 2 | 48 | 0.6468 | 1 | |
| 81 | hsa04260_Cardiac_muscle_contraction | 3 | 77 | 0.6846 | 1 | |
| 82 | hsa00983_Drug_metabolism_._other_enzymes | 2 | 52 | 0.6898 | 1 | |
| 83 | hsa04664_Fc_epsilon_RI_signaling_pathway | 3 | 79 | 0.7013 | 1 | |
| 84 | hsa00500_Starch_and_sucrose_metabolism | 2 | 54 | 0.7096 | 1 | |
| 85 | hsa04974_Protein_digestion_and_absorption | 3 | 81 | 0.7172 | 1 | |
| 86 | hsa00562_Inositol_phosphate_metabolism | 2 | 57 | 0.7373 | 1 | |
| 87 | hsa04512_ECM.receptor_interaction | 3 | 85 | 0.7472 | 1 | |
| 88 | hsa00590_Arachidonic_acid_metabolism | 2 | 59 | 0.7545 | 1 | |
| 89 | hsa04640_Hematopoietic_cell_lineage | 3 | 88 | 0.768 | 1 | |
| 90 | hsa04670_Leukocyte_transendothelial_migration | 4 | 117 | 0.7827 | 1 | |
| 91 | hsa00010_Glycolysis_._Gluconeogenesis | 2 | 65 | 0.8003 | 1 | |
| 92 | hsa00240_Pyrimidine_metabolism | 3 | 99 | 0.8324 | 1 | |
| 93 | hsa00980_Metabolism_of_xenobiotics_by_cytochrome_P450 | 2 | 71 | 0.8384 | 1 | |
| 94 | hsa04145_Phagosome | 5 | 156 | 0.841 | 1 | |
| 95 | hsa00982_Drug_metabolism_._cytochrome_P450 | 2 | 73 | 0.8496 | 1 | |
| 96 | hsa04612_Antigen_processing_and_presentation | 2 | 78 | 0.8745 | 1 | |
| 97 | hsa03008_Ribosome_biogenesis_in_eukaryotes | 2 | 81 | 0.8875 | 1 | |
| 98 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 3 | 136 | 0.9494 | 1 | |
| 99 | hsa03013_RNA_transport | 2 | 152 | 0.993 | 1 | |
| 100 | hsa04740_Olfactory_transduction | 2 | 388 | 1 | 1 |