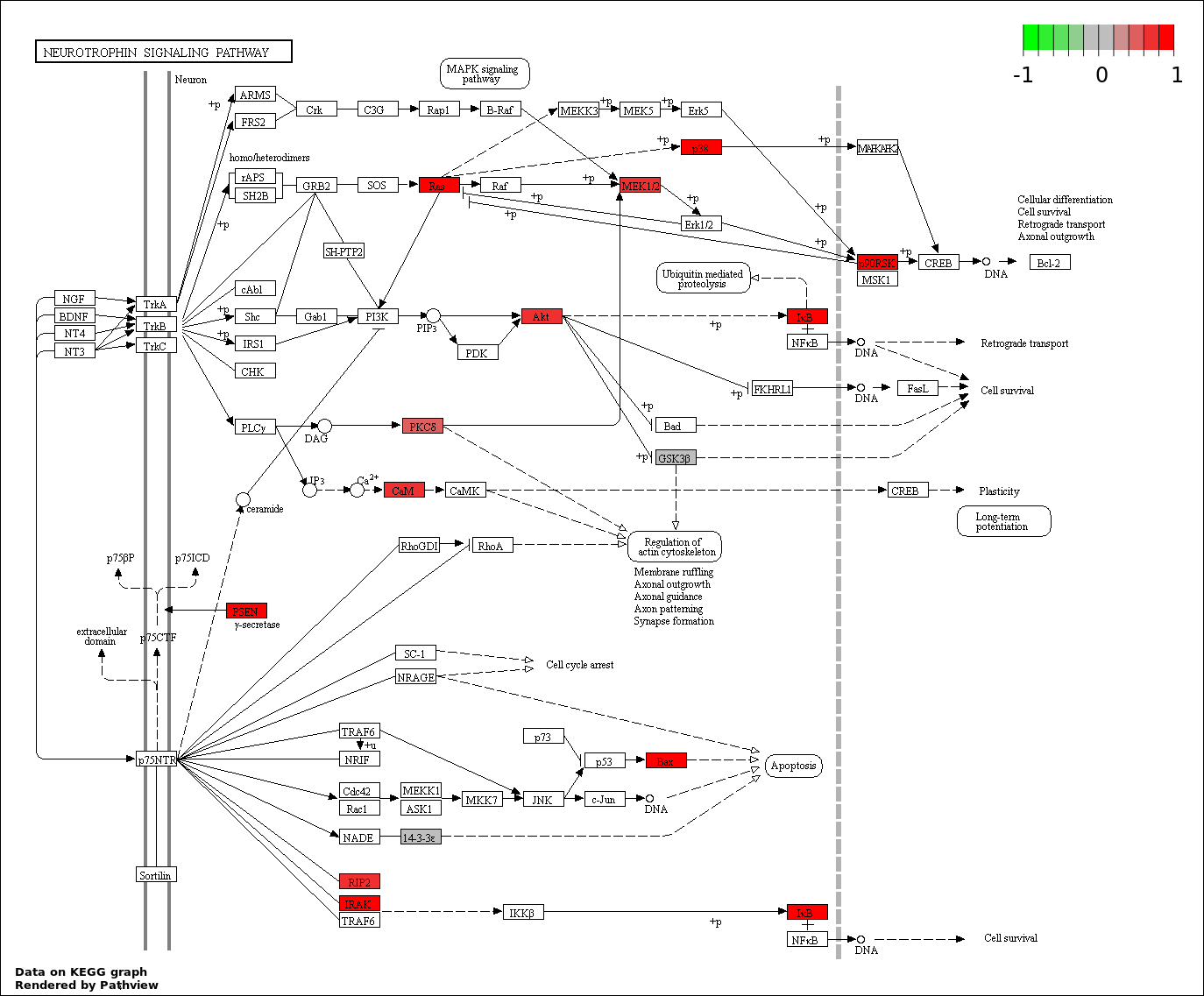
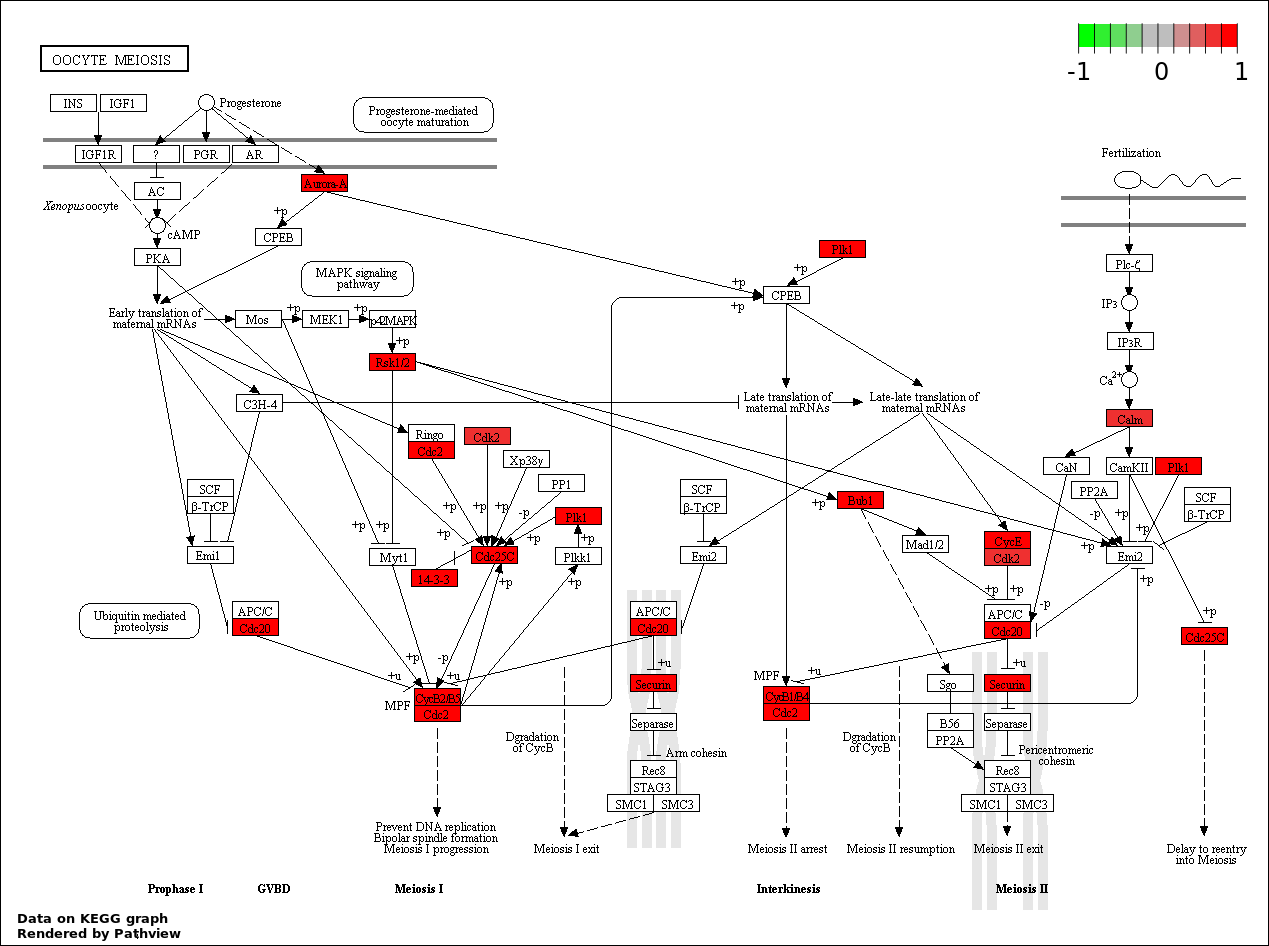
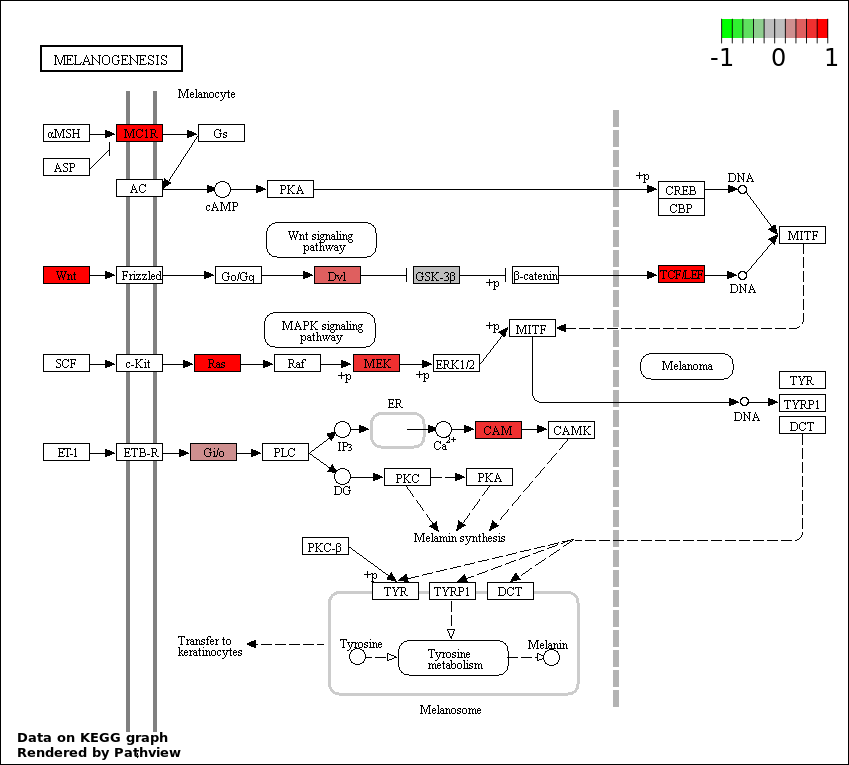
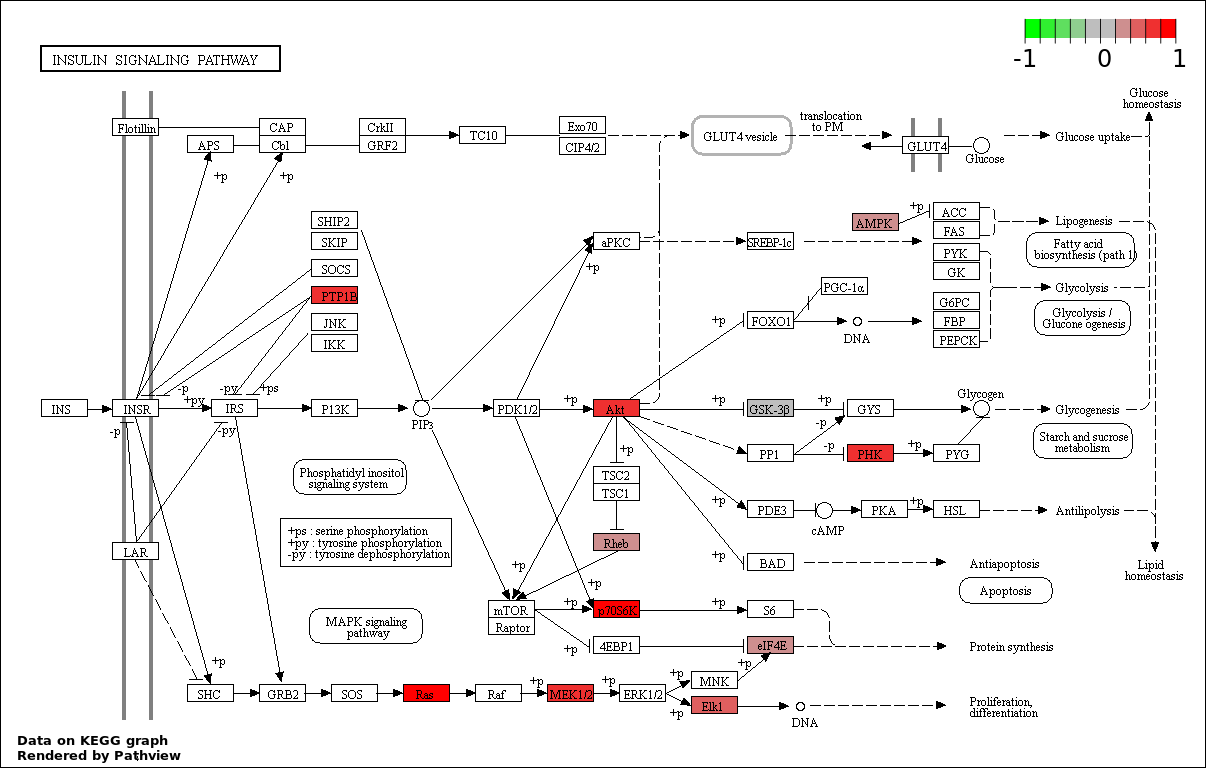
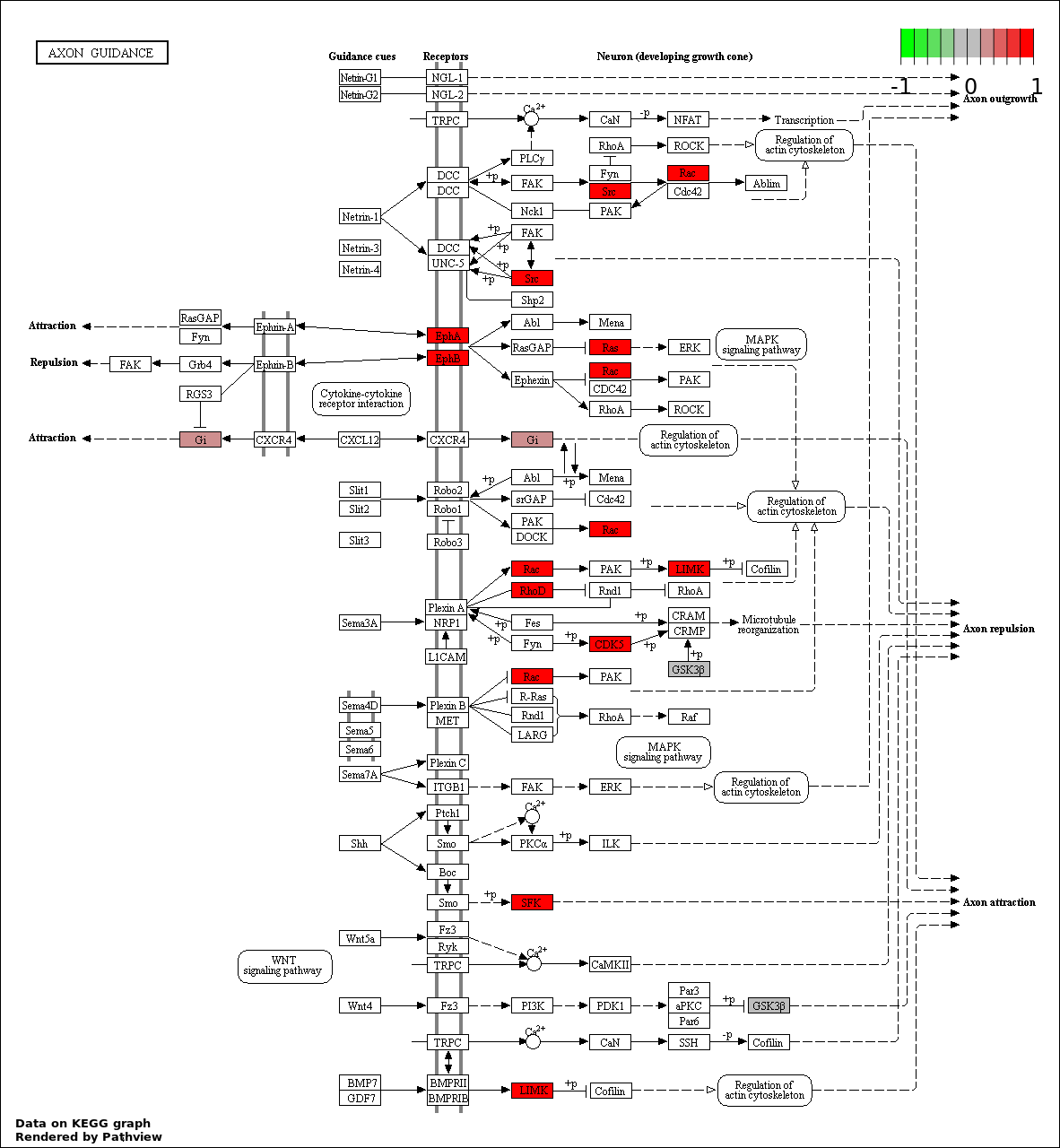
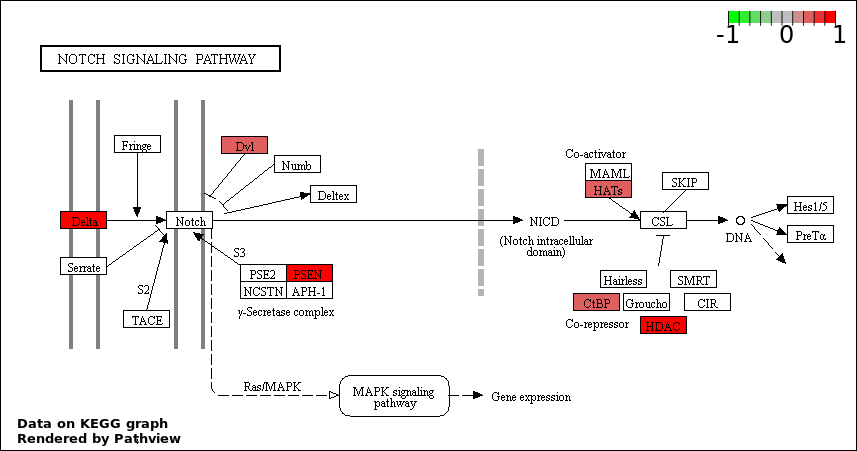
This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-let-7a-3p | KLF4 | 0.47 | 0 | -2.57 | 0 | MirTarget; miRNATAP | -0.4 | 0 | NA | |
| 2 | hsa-let-7f-1-3p | KLF4 | 0.38 | 0.00156 | -2.57 | 0 | MirTarget | -0.14 | 0.00245 | NA | |
| 3 | hsa-miR-103a-3p | KLF4 | 0.61 | 0 | -2.57 | 0 | miRNAWalker2 validate; miRTarBase | -0.37 | 0 | NA | |
| 4 | hsa-miR-107 | KLF4 | 0.68 | 0 | -2.57 | 0 | miRNAWalker2 validate; miRTarBase; miRanda; miRNATAP | -0.46 | 0 | NA | |
| 5 | hsa-miR-128-3p | KLF4 | 0.9 | 0 | -2.57 | 0 | miRNAWalker2 validate; MirTarget | -0.53 | 0 | NA | |
| 6 | hsa-miR-142-3p | KLF4 | 1.54 | 0 | -2.57 | 0 | miRNATAP | -0.18 | 0 | NA | |
| 7 | hsa-miR-148a-3p | KLF4 | 0.71 | 0 | -2.57 | 0 | MirTarget; miRNATAP | -0.2 | 1.0E-5 | NA | |
| 8 | hsa-miR-148b-3p | KLF4 | 1.06 | 0 | -2.57 | 0 | MirTarget | -0.57 | 0 | NA | |
| 9 | hsa-miR-200b-3p | KLF4 | 2.12 | 0 | -2.57 | 0 | MirTarget; TargetScan | -0.32 | 0 | NA | |
| 10 | hsa-miR-200c-3p | KLF4 | 2.07 | 0 | -2.57 | 0 | MirTarget; miRNATAP | -0.48 | 0 | NA | |
| 11 | hsa-miR-25-3p | KLF4 | -0.01 | 0.94406 | -2.57 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.24 | 0.00069 | NA | |
| 12 | hsa-miR-29b-3p | KLF4 | 0.44 | 6.0E-5 | -2.57 | 0 | MirTarget; miRNATAP | -0.2 | 9.0E-5 | 22751119 | Progestin suppression of miR 29 potentiates dedifferentiation of breast cancer cells via KLF4; We demonstrate that miR-29 directly targets Krüppel-like factor 4 KLF4 a transcription factor required for the reprogramming of differentiated cells to pluripotent stem cells and for the maintenance of breast cancer stem cells; These results reveal a novel mechanism whereby progestins increase the stem cell-like population in hormone-responsive breast cancers by decreasing miR-29 to augment PR-mediated upregulation of KLF4 |
| 13 | hsa-miR-301a-3p | KLF4 | 1.32 | 0 | -2.57 | 0 | miRNATAP | -0.36 | 0 | NA | |
| 14 | hsa-miR-30b-5p | KLF4 | -0.47 | 2.0E-5 | -2.57 | 0 | miRNAWalker2 validate | -0.1 | 0.03994 | NA | |
| 15 | hsa-miR-32-5p | KLF4 | 1.39 | 0 | -2.57 | 0 | MirTarget; miRNATAP | -0.41 | 0 | 26471298 | We also identified Kruppel-like factor 4 KLF4 as a direct target of miR-32; We conclude that miR-32 promotes GC cell proliferation migration and invasion by targeting KLF4 suggesting that the miR-32-KLF4 pathway may be useful in clinical diagnosis and therapeutics |
| 16 | hsa-miR-34a-5p | KLF4 | 0.13 | 0.13326 | -2.57 | 0 | MirTarget; miRNATAP | -0.16 | 0.01412 | NA | |
| 17 | hsa-miR-375 | KLF4 | 2.1 | 0 | -2.57 | 0 | miRanda; miRNATAP | -0.11 | 0 | 26224477 | MiR 375 targets KLF4 and impacts the proliferation of colorectal carcinoma; Subsequently using the luciferase reporter assays we found that the KLF4 untranslated region 3'UTR carries the direct binding site of miR-375; Inhibition of KLF4 performed similar effects with miR-375 overexpression on CRC cells and overexpression of KLF4 could significantly reverse the tumor suppressive effects of miR-375 on CRC cells; Furthermore we found overexpressed miR-375 effectively repressed tumor growth via KLF4 in xenograft animal experiment; Taken together these results illustrated that miR-375 depresses proliferation of CRC through regulating 3'UTR of KLF4 mRNA which might be a promising therapeutic target for treating colorectal cancers |
| 18 | hsa-miR-429 | KLF4 | 2.84 | 0 | -2.57 | 0 | MirTarget; PITA; miRanda; miRNATAP | -0.35 | 0 | NA | |
| 19 | hsa-miR-493-5p | KLF4 | 1.25 | 0 | -2.57 | 0 | miRNATAP | -0.15 | 9.0E-5 | NA | |
| 20 | hsa-miR-508-3p | KLF4 | 0.26 | 0.18993 | -2.57 | 0 | PITA; miRNATAP | -0.07 | 0.01438 | NA | |
| 21 | hsa-miR-590-3p | KLF4 | 1.27 | 0 | -2.57 | 0 | miRanda | -0.41 | 0 | NA | |
| 22 | hsa-miR-92a-3p | KLF4 | -0.34 | 0.00073 | -2.57 | 0 | MirTarget; miRNATAP | -0.26 | 0 | 27131314 | Additionally KLF4 was identified as a direct target of miR-92a in CRC cells through bioinformatics and luciferase reporter analysis; KLF4 overexpression attenuated the effects of miR-92a on the regulation of CRC cell motility; In conclusion our data demonstrated that miR-92a may play a positive role in the colorectal carcinogenesis by promoting the proliferation and migration of CRC cells through targeting KLF4 as well as downstream p21 |
| 23 | hsa-miR-92b-3p | KLF4 | 1.3 | 0 | -2.57 | 0 | MirTarget; miRNATAP | -0.26 | 0 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CELL CYCLE | 114 | 1316 | 3.551e-42 | 1.652e-38 |
| 2 | CHROMOSOME ORGANIZATION | 99 | 1009 | 6.517e-41 | 1.516e-37 |
| 3 | CELL CYCLE PROCESS | 95 | 1081 | 3.417e-35 | 5.3e-32 |
| 4 | MITOTIC CELL CYCLE | 80 | 766 | 1.134e-34 | 1.319e-31 |
| 5 | REGULATION OF CELL CYCLE | 87 | 949 | 1.806e-33 | 1.681e-30 |
| 6 | POSITIVE REGULATION OF GENE EXPRESSION | 115 | 1733 | 1.705e-31 | 1.322e-28 |
| 7 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 115 | 1805 | 6.885e-30 | 4.577e-27 |
| 8 | CHROMATIN MODIFICATION | 62 | 539 | 3.984e-29 | 2.317e-26 |
| 9 | CHROMATIN ORGANIZATION | 68 | 663 | 6.549e-29 | 3.386e-26 |
| 10 | PEPTIDYL AMINO ACID MODIFICATION | 75 | 841 | 5.898e-28 | 2.744e-25 |
| 11 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 111 | 1784 | 7.285e-28 | 3.081e-25 |
| 12 | COVALENT CHROMATIN MODIFICATION | 49 | 345 | 2.869e-27 | 1.113e-24 |
| 13 | CELLULAR RESPONSE TO STRESS | 100 | 1565 | 8.568e-26 | 3.067e-23 |
| 14 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 98 | 1517 | 1.315e-25 | 4.371e-23 |
| 15 | PROTEIN PHOSPHORYLATION | 75 | 944 | 8.58e-25 | 2.662e-22 |
| 16 | CELL CYCLE CHECKPOINT | 36 | 194 | 2.428e-24 | 7.06e-22 |
| 17 | NEGATIVE REGULATION OF GENE EXPRESSION | 95 | 1493 | 2.693e-24 | 7.37e-22 |
| 18 | CELL CYCLE PHASE TRANSITION | 40 | 255 | 4.659e-24 | 1.204e-21 |
| 19 | DNA METABOLIC PROCESS | 66 | 758 | 5.409e-24 | 1.325e-21 |
| 20 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 72 | 917 | 1.737e-23 | 4.041e-21 |
| 21 | REGULATION OF CELL CYCLE PROCESS | 56 | 558 | 2.278e-23 | 5.046e-21 |
| 22 | REGULATION OF ORGANELLE ORGANIZATION | 82 | 1178 | 2.41e-23 | 5.097e-21 |
| 23 | REGULATION OF PROTEIN MODIFICATION PROCESS | 100 | 1710 | 7.294e-23 | 1.476e-20 |
| 24 | CELLULAR RESPONSE TO DNA DAMAGE STIMULUS | 61 | 720 | 1.401e-21 | 2.716e-19 |
| 25 | REGULATION OF MITOTIC CELL CYCLE | 49 | 468 | 2.595e-21 | 4.831e-19 |
| 26 | NEGATIVE REGULATION OF CELL CYCLE | 47 | 433 | 3.993e-21 | 7.146e-19 |
| 27 | REGULATION OF DNA METABOLIC PROCESS | 42 | 340 | 4.224e-21 | 7.28e-19 |
| 28 | PHOSPHORYLATION | 80 | 1228 | 5.771e-21 | 9.59e-19 |
| 29 | NEGATIVE REGULATION OF CELL CYCLE PROCESS | 34 | 214 | 8.799e-21 | 1.412e-18 |
| 30 | MITOTIC CELL CYCLE CHECKPOINT | 28 | 139 | 3.451e-20 | 5.352e-18 |
| 31 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 73 | 1087 | 7.401e-20 | 1.111e-17 |
| 32 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 87 | 1492 | 1.104e-19 | 1.606e-17 |
| 33 | REGULATION OF CELL PROLIFERATION | 87 | 1496 | 1.311e-19 | 1.849e-17 |
| 34 | REGULATION OF CELLULAR RESPONSE TO STRESS | 56 | 691 | 5.964e-19 | 8.162e-17 |
| 35 | RESPONSE TO ABIOTIC STIMULUS | 69 | 1024 | 7.537e-19 | 1.002e-16 |
| 36 | NEGATIVE REGULATION OF MITOTIC CELL CYCLE | 31 | 199 | 8.645e-19 | 1.117e-16 |
| 37 | INTRACELLULAR SIGNAL TRANSDUCTION | 88 | 1572 | 8.916e-19 | 1.121e-16 |
| 38 | REGULATION OF BINDING | 36 | 283 | 1.196e-18 | 1.464e-16 |
| 39 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 52 | 616 | 2.11e-18 | 2.517e-16 |
| 40 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 80 | 1381 | 6.692e-18 | 7.785e-16 |
| 41 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 99 | 1977 | 7.901e-18 | 8.966e-16 |
| 42 | NEGATIVE REGULATION OF CELL CYCLE PHASE TRANSITION | 26 | 146 | 1.971e-17 | 2.133e-15 |
| 43 | DNA INTEGRITY CHECKPOINT | 26 | 146 | 1.971e-17 | 2.133e-15 |
| 44 | REGULATION OF CELL DEATH | 82 | 1472 | 2.414e-17 | 2.553e-15 |
| 45 | RESPONSE TO LIPID | 61 | 888 | 4.277e-17 | 4.423e-15 |
| 46 | CELL DIVISION | 43 | 460 | 5.804e-17 | 5.871e-15 |
| 47 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 65 | 1004 | 6.644e-17 | 6.578e-15 |
| 48 | NEGATIVE REGULATION OF CELL DEATH | 60 | 872 | 7.416e-17 | 7.189e-15 |
| 49 | RESPONSE TO ENDOGENOUS STIMULUS | 80 | 1450 | 1.122e-16 | 1.065e-14 |
| 50 | NEGATIVE REGULATION OF ORGANELLE ORGANIZATION | 39 | 387 | 1.421e-16 | 1.322e-14 |
| 51 | ORGANELLE FISSION | 44 | 496 | 1.733e-16 | 1.581e-14 |
| 52 | REGULATION OF TRANSFERASE ACTIVITY | 62 | 946 | 2.152e-16 | 1.926e-14 |
| 53 | REGULATION OF SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 26 | 162 | 2.806e-16 | 2.418e-14 |
| 54 | REGULATION OF CHROMOSOME ORGANIZATION | 33 | 278 | 2.807e-16 | 2.418e-14 |
| 55 | REGULATION OF CELL CYCLE ARREST | 22 | 108 | 3.088e-16 | 2.612e-14 |
| 56 | RESPONSE TO ALCOHOL | 37 | 362 | 5.633e-16 | 4.68e-14 |
| 57 | NEGATIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 51 | 684 | 8e-16 | 6.53e-14 |
| 58 | RESPONSE TO STEROID HORMONE | 43 | 497 | 9.632e-16 | 7.728e-14 |
| 59 | NEGATIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 53 | 740 | 1.107e-15 | 8.733e-14 |
| 60 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 68 | 1152 | 1.238e-15 | 9.599e-14 |
| 61 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 67 | 1135 | 2.072e-15 | 1.581e-13 |
| 62 | REGULATION OF CELL CYCLE PHASE TRANSITION | 34 | 321 | 3.126e-15 | 2.346e-13 |
| 63 | RESPONSE TO HORMONE | 58 | 893 | 3.381e-15 | 2.497e-13 |
| 64 | REGULATION OF NUCLEASE ACTIVITY | 12 | 23 | 4.473e-15 | 3.154e-13 |
| 65 | SIGNAL TRANSDUCTION IN RESPONSE TO DNA DAMAGE | 20 | 96 | 4.456e-15 | 3.154e-13 |
| 66 | G1 DNA DAMAGE CHECKPOINT | 18 | 73 | 4.472e-15 | 3.154e-13 |
| 67 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 89 | 1848 | 6.025e-15 | 4.184e-13 |
| 68 | CELL CYCLE G2 M PHASE TRANSITION | 23 | 138 | 6.512e-15 | 4.456e-13 |
| 69 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 83 | 1656 | 6.796e-15 | 4.583e-13 |
| 70 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 51 | 724 | 7.862e-15 | 5.152e-13 |
| 71 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 64 | 1079 | 7.776e-15 | 5.152e-13 |
| 72 | POSITIVE REGULATION OF CELL CYCLE | 34 | 332 | 8.544e-15 | 5.521e-13 |
| 73 | MITOTIC DNA INTEGRITY CHECKPOINT | 20 | 100 | 1.026e-14 | 6.45e-13 |
| 74 | REGULATION OF PROTEIN STABILITY | 28 | 221 | 1.026e-14 | 6.45e-13 |
| 75 | MITOTIC NUCLEAR DIVISION | 35 | 361 | 1.784e-14 | 1.107e-12 |
| 76 | PROTEIN AUTOPHOSPHORYLATION | 26 | 192 | 1.89e-14 | 1.157e-12 |
| 77 | POSITIVE REGULATION OF CELL CYCLE ARREST | 18 | 85 | 7.857e-14 | 4.748e-12 |
| 78 | NEGATIVE REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 19 | 98 | 8.744e-14 | 5.216e-12 |
| 79 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 45 | 616 | 9.552e-14 | 5.626e-12 |
| 80 | REGULATION OF KINASE ACTIVITY | 51 | 776 | 1.195e-13 | 6.951e-12 |
| 81 | DNA REPAIR | 39 | 480 | 1.709e-13 | 9.819e-12 |
| 82 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 42 | 552 | 1.746e-13 | 9.905e-12 |
| 83 | EMBRYO DEVELOPMENT | 55 | 894 | 1.776e-13 | 9.955e-12 |
| 84 | HEAD DEVELOPMENT | 48 | 709 | 2.28e-13 | 1.263e-11 |
| 85 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 54 | 872 | 2.327e-13 | 1.274e-11 |
| 86 | REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 22 | 147 | 2.515e-13 | 1.361e-11 |
| 87 | INTERSPECIES INTERACTION BETWEEN ORGANISMS | 46 | 662 | 2.995e-13 | 1.583e-11 |
| 88 | SYMBIOSIS ENCOMPASSING MUTUALISM THROUGH PARASITISM | 46 | 662 | 2.995e-13 | 1.583e-11 |
| 89 | REGULATION OF CELL DIVISION | 29 | 272 | 3.08e-13 | 1.61e-11 |
| 90 | REGULATION OF CELL DIFFERENTIATION | 74 | 1492 | 4.588e-13 | 2.372e-11 |
| 91 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 42 | 573 | 6.06e-13 | 3.099e-11 |
| 92 | REGULATION OF PROTEIN MODIFICATION BY SMALL PROTEIN CONJUGATION OR REMOVAL | 29 | 280 | 6.447e-13 | 3.261e-11 |
| 93 | REGULATION OF CATABOLIC PROCESS | 48 | 731 | 6.97e-13 | 3.487e-11 |
| 94 | RESPONSE TO INORGANIC SUBSTANCE | 38 | 479 | 7.443e-13 | 3.65e-11 |
| 95 | POSITIVE REGULATION OF CELL PROLIFERATION | 51 | 814 | 7.453e-13 | 3.65e-11 |
| 96 | CELL CYCLE G1 S PHASE TRANSITION | 19 | 111 | 9.245e-13 | 4.435e-11 |
| 97 | G1 S TRANSITION OF MITOTIC CELL CYCLE | 19 | 111 | 9.245e-13 | 4.435e-11 |
| 98 | SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 20 | 127 | 1.172e-12 | 5.564e-11 |
| 99 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 37 | 465 | 1.387e-12 | 6.519e-11 |
| 100 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 86 | 1929 | 1.445e-12 | 6.722e-11 |
| 101 | NEUROGENESIS | 70 | 1402 | 1.631e-12 | 7.515e-11 |
| 102 | RESPONSE TO IONIZING RADIATION | 21 | 145 | 1.707e-12 | 7.788e-11 |
| 103 | PEPTIDYL LYSINE MODIFICATION | 30 | 312 | 1.757e-12 | 7.938e-11 |
| 104 | POSITIVE REGULATION OF NUCLEASE ACTIVITY | 9 | 15 | 2.357e-12 | 1.054e-10 |
| 105 | REGULATION OF PROTEASOMAL PROTEIN CATABOLIC PROCESS | 23 | 181 | 2.436e-12 | 1.079e-10 |
| 106 | REGULATION OF PROTEASOMAL UBIQUITIN DEPENDENT PROTEIN CATABOLIC PROCESS | 21 | 148 | 2.565e-12 | 1.126e-10 |
| 107 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 76 | 1618 | 3.158e-12 | 1.373e-10 |
| 108 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 69 | 1395 | 3.736e-12 | 1.61e-10 |
| 109 | RESPONSE TO RADIATION | 34 | 413 | 4.582e-12 | 1.956e-10 |
| 110 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 49 | 799 | 4.77e-12 | 2.018e-10 |
| 111 | REGULATION OF DNA BINDING | 17 | 93 | 4.903e-12 | 2.039e-10 |
| 112 | REGULATION OF RESPONSE TO STRESS | 71 | 1468 | 4.908e-12 | 2.039e-10 |
| 113 | REGULATION OF CELL DEVELOPMENT | 50 | 836 | 7.061e-12 | 2.907e-10 |
| 114 | POSITIVE REGULATION OF CELL CYCLE PROCESS | 26 | 247 | 7.187e-12 | 2.933e-10 |
| 115 | NEGATIVE REGULATION OF CHROMOSOME ORGANIZATION | 17 | 96 | 8.382e-12 | 3.391e-10 |
| 116 | REPRODUCTION | 65 | 1297 | 9.968e-12 | 3.998e-10 |
| 117 | FOREBRAIN DEVELOPMENT | 31 | 357 | 1.068e-11 | 4.247e-10 |
| 118 | REGULATION OF CELLULAR PROTEIN CATABOLIC PROCESS | 27 | 274 | 1.325e-11 | 5.225e-10 |
| 119 | REGULATION OF CELLULAR LOCALIZATION | 64 | 1277 | 1.476e-11 | 5.77e-10 |
| 120 | CELL DEATH | 55 | 1001 | 1.552e-11 | 6.019e-10 |
| 121 | REGULATION OF NUCLEAR DIVISION | 21 | 163 | 1.702e-11 | 6.546e-10 |
| 122 | PEPTIDYL SERINE MODIFICATION | 20 | 148 | 2.143e-11 | 8.175e-10 |
| 123 | REGULATION OF PROTEIN CATABOLIC PROCESS | 32 | 393 | 2.667e-11 | 1.009e-09 |
| 124 | CHROMATIN REMODELING | 20 | 150 | 2.753e-11 | 1.033e-09 |
| 125 | CELL PROLIFERATION | 43 | 672 | 2.795e-11 | 1.04e-09 |
| 126 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 59 | 1142 | 3.122e-11 | 1.153e-09 |
| 127 | REGULATION OF GENE EXPRESSION EPIGENETIC | 24 | 229 | 5.147e-11 | 1.886e-09 |
| 128 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 30 | 363 | 7.914e-11 | 2.877e-09 |
| 129 | RESPONSE TO ESTROGEN | 23 | 218 | 1.144e-10 | 4.126e-09 |
| 130 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 47 | 823 | 1.52e-10 | 5.442e-09 |
| 131 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 34 | 470 | 1.545e-10 | 5.489e-09 |
| 132 | DNA TEMPLATED TRANSCRIPTION INITIATION | 22 | 202 | 1.577e-10 | 5.56e-09 |
| 133 | REGULATION OF PROTEOLYSIS | 43 | 711 | 1.671e-10 | 5.847e-09 |
| 134 | POSITIVE REGULATION OF DNA METABOLIC PROCESS | 21 | 185 | 1.91e-10 | 6.582e-09 |
| 135 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 53 | 1008 | 1.898e-10 | 6.582e-09 |
| 136 | REGULATION OF CENTROSOME CYCLE | 11 | 39 | 2.066e-10 | 7.07e-09 |
| 137 | REGULATION OF PROTEIN LOCALIZATION | 51 | 950 | 2.104e-10 | 7.145e-09 |
| 138 | REGULATION OF PROTEIN BINDING | 20 | 168 | 2.206e-10 | 7.438e-09 |
| 139 | RESPONSE TO GAMMA RADIATION | 12 | 50 | 2.451e-10 | 8.206e-09 |
| 140 | INTRINSIC APOPTOTIC SIGNALING PATHWAY | 19 | 152 | 2.692e-10 | 8.948e-09 |
| 141 | DNA REPLICATION | 22 | 208 | 2.784e-10 | 9.187e-09 |
| 142 | RESPONSE TO DRUG | 32 | 431 | 2.85e-10 | 9.34e-09 |
| 143 | PROTEIN COMPLEX BIOGENESIS | 56 | 1132 | 5.363e-10 | 1.733e-08 |
| 144 | PROTEIN COMPLEX ASSEMBLY | 56 | 1132 | 5.363e-10 | 1.733e-08 |
| 145 | PROTEIN COMPLEX SUBUNIT ORGANIZATION | 68 | 1527 | 5.418e-10 | 1.739e-08 |
| 146 | GLAND DEVELOPMENT | 30 | 395 | 6.117e-10 | 1.95e-08 |
| 147 | REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 11 | 43 | 6.594e-10 | 2.087e-08 |
| 148 | POSITIVE REGULATION OF BINDING | 17 | 127 | 8.088e-10 | 2.526e-08 |
| 149 | HISTONE PHOSPHORYLATION | 9 | 25 | 8.038e-10 | 2.526e-08 |
| 150 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 36 | 554 | 8.363e-10 | 2.594e-08 |
| 151 | NEGATIVE REGULATION OF GENE EXPRESSION EPIGENETIC | 16 | 112 | 9.626e-10 | 2.947e-08 |
| 152 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 75 | 1791 | 9.573e-10 | 2.947e-08 |
| 153 | AGING | 24 | 264 | 9.694e-10 | 2.948e-08 |
| 154 | POSITIVE REGULATION OF KINASE ACTIVITY | 33 | 482 | 1.187e-09 | 3.585e-08 |
| 155 | CELLULAR RESPONSE TO LIPID | 32 | 457 | 1.231e-09 | 3.694e-08 |
| 156 | NEGATIVE REGULATION OF BINDING | 17 | 131 | 1.319e-09 | 3.932e-08 |
| 157 | REGULATION OF MEMBRANE PERMEABILITY | 13 | 70 | 1.327e-09 | 3.932e-08 |
| 158 | PEPTIDYL THREONINE MODIFICATION | 11 | 46 | 1.448e-09 | 4.265e-08 |
| 159 | REGULATION OF INTRACELLULAR TRANSPORT | 38 | 621 | 1.497e-09 | 4.382e-08 |
| 160 | POSITIVE REGULATION OF CHROMOSOME ORGANIZATION | 18 | 150 | 1.561e-09 | 4.541e-08 |
| 161 | POSITIVE REGULATION OF CHROMATIN MODIFICATION | 14 | 85 | 1.605e-09 | 4.589e-08 |
| 162 | TELENCEPHALON DEVELOPMENT | 22 | 228 | 1.608e-09 | 4.589e-08 |
| 163 | NUCLEAR CHROMOSOME SEGREGATION | 22 | 228 | 1.608e-09 | 4.589e-08 |
| 164 | POSITIVE REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 10 | 36 | 1.649e-09 | 4.68e-08 |
| 165 | CHROMOSOME SEGREGATION | 24 | 272 | 1.768e-09 | 4.986e-08 |
| 166 | RESPONSE TO NITROGEN COMPOUND | 46 | 859 | 1.909e-09 | 5.35e-08 |
| 167 | REGULATION OF CHROMATIN ORGANIZATION | 18 | 152 | 1.938e-09 | 5.399e-08 |
| 168 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 49 | 957 | 2.383e-09 | 6.56e-08 |
| 169 | POSITIVE REGULATION OF CELLULAR PROTEIN CATABOLIC PROCESS | 20 | 192 | 2.38e-09 | 6.56e-08 |
| 170 | POSITIVE REGULATION OF CATABOLIC PROCESS | 29 | 395 | 2.594e-09 | 7.099e-08 |
| 171 | ATP DEPENDENT CHROMATIN REMODELING | 13 | 74 | 2.719e-09 | 7.397e-08 |
| 172 | SISTER CHROMATID SEGREGATION | 19 | 176 | 3.304e-09 | 8.939e-08 |
| 173 | NEGATIVE REGULATION OF TRANSFERASE ACTIVITY | 27 | 351 | 3.439e-09 | 9.251e-08 |
| 174 | MACROMOLECULAR COMPLEX ASSEMBLY | 62 | 1398 | 4.171e-09 | 1.115e-07 |
| 175 | EPITHELIUM DEVELOPMENT | 48 | 945 | 4.556e-09 | 1.211e-07 |
| 176 | POSITIVE REGULATION OF PROTEIN CATABOLIC PROCESS | 23 | 263 | 4.6e-09 | 1.216e-07 |
| 177 | MITOCHONDRIAL MEMBRANE ORGANIZATION | 14 | 92 | 4.681e-09 | 1.231e-07 |
| 178 | REGULATION OF MICROTUBULE BASED PROCESS | 22 | 243 | 5.287e-09 | 1.382e-07 |
| 179 | POSITIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 27 | 360 | 5.921e-09 | 1.53e-07 |
| 180 | APOPTOTIC SIGNALING PATHWAY | 24 | 289 | 5.888e-09 | 1.53e-07 |
| 181 | REGULATION OF RESPONSE TO DNA DAMAGE STIMULUS | 17 | 145 | 6.382e-09 | 1.641e-07 |
| 182 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 41 | 750 | 8.66e-09 | 2.214e-07 |
| 183 | ORGAN MORPHOGENESIS | 44 | 841 | 8.87e-09 | 2.255e-07 |
| 184 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 69 | 1672 | 9.338e-09 | 2.361e-07 |
| 185 | REGULATION OF NEURON DIFFERENTIATION | 34 | 554 | 1.061e-08 | 2.667e-07 |
| 186 | RESPONSE TO VITAMIN | 14 | 98 | 1.086e-08 | 2.716e-07 |
| 187 | MICROTUBULE CYTOSKELETON ORGANIZATION | 26 | 348 | 1.233e-08 | 3.069e-07 |
| 188 | RESPONSE TO NUTRIENT | 19 | 191 | 1.285e-08 | 3.164e-07 |
| 189 | RESPONSE TO REACTIVE OXYGEN SPECIES | 19 | 191 | 1.285e-08 | 3.164e-07 |
| 190 | HISTONE METHYLATION | 13 | 84 | 1.354e-08 | 3.315e-07 |
| 191 | TRANSCRIPTION INITIATION FROM RNA POLYMERASE II PROMOTER | 17 | 153 | 1.447e-08 | 3.524e-07 |
| 192 | TISSUE DEVELOPMENT | 64 | 1518 | 1.587e-08 | 3.846e-07 |
| 193 | EMBRYONIC MORPHOGENESIS | 33 | 539 | 1.867e-08 | 4.477e-07 |
| 194 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 42 | 801 | 1.862e-08 | 4.477e-07 |
| 195 | POSITIVE REGULATION OF PROTEIN MODIFICATION BY SMALL PROTEIN CONJUGATION OR REMOVAL | 19 | 196 | 1.962e-08 | 4.681e-07 |
| 196 | REGULATION OF HISTONE H3 K9 METHYLATION | 7 | 17 | 2.165e-08 | 5.139e-07 |
| 197 | POSITIVE REGULATION OF CELL COMMUNICATION | 64 | 1532 | 2.244e-08 | 5.301e-07 |
| 198 | PROTEIN UBIQUITINATION | 36 | 629 | 2.331e-08 | 5.478e-07 |
| 199 | POSITIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 23 | 289 | 2.722e-08 | 6.364e-07 |
| 200 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 49 | 1036 | 3.013e-08 | 6.975e-07 |
| 201 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 49 | 1036 | 3.013e-08 | 6.975e-07 |
| 202 | NEGATIVE REGULATION OF HISTONE MODIFICATION | 9 | 36 | 3.041e-08 | 7.004e-07 |
| 203 | REGULATION OF DNA REPAIR | 12 | 75 | 3.342e-08 | 7.624e-07 |
| 204 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 35 | 609 | 3.328e-08 | 7.624e-07 |
| 205 | RESPONSE TO KETONE | 18 | 182 | 3.406e-08 | 7.73e-07 |
| 206 | POSITIVE REGULATION OF CELL DEVELOPMENT | 30 | 472 | 3.712e-08 | 8.384e-07 |
| 207 | PROTEIN LOCALIZATION | 71 | 1805 | 3.84e-08 | 8.605e-07 |
| 208 | PROTEASOMAL PROTEIN CATABOLIC PROCESS | 22 | 271 | 3.847e-08 | 8.605e-07 |
| 209 | RESPONSE TO UV | 15 | 126 | 4.061e-08 | 8.997e-07 |
| 210 | MACROMOLECULE METHYLATION | 19 | 205 | 4.06e-08 | 8.997e-07 |
| 211 | REGULATION OF CYTOSKELETON ORGANIZATION | 31 | 502 | 4.214e-08 | 9.293e-07 |
| 212 | DNA CONFORMATION CHANGE | 22 | 273 | 4.387e-08 | 9.628e-07 |
| 213 | ANAPHASE PROMOTING COMPLEX DEPENDENT CATABOLIC PROCESS | 12 | 77 | 4.534e-08 | 9.904e-07 |
| 214 | RESPONSE TO ESTRADIOL | 16 | 146 | 4.693e-08 | 1.02e-06 |
| 215 | CELL DEVELOPMENT | 60 | 1426 | 5.151e-08 | 1.115e-06 |
| 216 | NEGATIVE REGULATION OF CELL COMMUNICATION | 53 | 1192 | 5.961e-08 | 1.284e-06 |
| 217 | NEGATIVE REGULATION OF PROTEIN BINDING | 12 | 79 | 6.091e-08 | 1.306e-06 |
| 218 | CYTOSKELETON ORGANIZATION | 42 | 838 | 6.591e-08 | 1.407e-06 |
| 219 | PROTEIN MODIFICATION BY SMALL PROTEIN CONJUGATION OR REMOVAL | 43 | 873 | 7.409e-08 | 1.574e-06 |
| 220 | POSITIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 17 | 171 | 7.601e-08 | 1.608e-06 |
| 221 | REPRODUCTIVE SYSTEM DEVELOPMENT | 27 | 408 | 8.052e-08 | 1.695e-06 |
| 222 | POSITIVE REGULATION OF CELL DEATH | 34 | 605 | 9.011e-08 | 1.889e-06 |
| 223 | PALLIUM DEVELOPMENT | 16 | 153 | 9.087e-08 | 1.896e-06 |
| 224 | PROTEIN UBIQUITINATION INVOLVED IN UBIQUITIN DEPENDENT PROTEIN CATABOLIC PROCESS | 15 | 134 | 9.313e-08 | 1.934e-06 |
| 225 | SENSORY ORGAN DEVELOPMENT | 30 | 493 | 9.673e-08 | 2e-06 |
| 226 | PROTEIN CATABOLIC PROCESS | 33 | 579 | 1.014e-07 | 2.07e-06 |
| 227 | MICROTUBULE BASED PROCESS | 31 | 522 | 1.014e-07 | 2.07e-06 |
| 228 | REGULATION OF IRE1 MEDIATED UNFOLDED PROTEIN RESPONSE | 6 | 13 | 1.011e-07 | 2.07e-06 |
| 229 | IN UTERO EMBRYONIC DEVELOPMENT | 23 | 311 | 1.045e-07 | 2.123e-06 |
| 230 | CELLULAR RESPONSE TO HORMONE STIMULUS | 32 | 552 | 1.087e-07 | 2.198e-06 |
| 231 | RESPONSE TO EXTRACELLULAR STIMULUS | 28 | 441 | 1.094e-07 | 2.205e-06 |
| 232 | POSITIVE REGULATION OF PROTEOLYSIS | 25 | 363 | 1.152e-07 | 2.31e-06 |
| 233 | CELLULAR RESPONSE TO RADIATION | 15 | 137 | 1.251e-07 | 2.493e-06 |
| 234 | NEGATIVE REGULATION OF CELL PROLIFERATION | 35 | 643 | 1.254e-07 | 2.493e-06 |
| 235 | PROTEIN LOCALIZATION TO ORGANELLE | 32 | 556 | 1.28e-07 | 2.534e-06 |
| 236 | POSITIVE REGULATION OF DNA BINDING | 9 | 42 | 1.293e-07 | 2.55e-06 |
| 237 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 57 | 1360 | 1.329e-07 | 2.609e-06 |
| 238 | NEGATIVE REGULATION OF CATALYTIC ACTIVITY | 41 | 829 | 1.366e-07 | 2.67e-06 |
| 239 | REGULATION OF HOMEOSTATIC PROCESS | 28 | 447 | 1.447e-07 | 2.817e-06 |
| 240 | NEGATIVE REGULATION OF PROTEIN MODIFICATION BY SMALL PROTEIN CONJUGATION OR REMOVAL | 15 | 139 | 1.517e-07 | 2.929e-06 |
| 241 | POSTTRANSCRIPTIONAL REGULATION OF GENE EXPRESSION | 28 | 448 | 1.515e-07 | 2.929e-06 |
| 242 | NEGATIVE REGULATION OF PHOSPHORYLATION | 27 | 422 | 1.591e-07 | 3.06e-06 |
| 243 | TISSUE MORPHOGENESIS | 31 | 533 | 1.609e-07 | 3.08e-06 |
| 244 | CELLULAR RESPONSE TO NITROGEN COMPOUND | 30 | 505 | 1.629e-07 | 3.106e-06 |
| 245 | POSITIVE REGULATION OF INTRACELLULAR TRANSPORT | 25 | 370 | 1.658e-07 | 3.149e-06 |
| 246 | CELLULAR MACROMOLECULE LOCALIZATION | 53 | 1234 | 1.803e-07 | 3.411e-06 |
| 247 | INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO ENDOPLASMIC RETICULUM STRESS | 8 | 32 | 1.813e-07 | 3.416e-06 |
| 248 | CELLULAR RESPONSE TO REACTIVE OXYGEN SPECIES | 13 | 104 | 1.835e-07 | 3.443e-06 |
| 249 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 61 | 1518 | 1.983e-07 | 3.706e-06 |
| 250 | NEGATIVE REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 12 | 88 | 2.073e-07 | 3.859e-06 |
| 251 | CELLULAR RESPONSE TO OXYGEN LEVELS | 15 | 143 | 2.207e-07 | 4.091e-06 |
| 252 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 31 | 541 | 2.231e-07 | 4.103e-06 |
| 253 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 31 | 541 | 2.231e-07 | 4.103e-06 |
| 254 | PROTEIN DNA COMPLEX SUBUNIT ORGANIZATION | 19 | 229 | 2.342e-07 | 4.291e-06 |
| 255 | EYE DEVELOPMENT | 23 | 326 | 2.431e-07 | 4.428e-06 |
| 256 | NEGATIVE REGULATION OF CHROMATIN MODIFICATION | 9 | 45 | 2.436e-07 | 4.428e-06 |
| 257 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 33 | 602 | 2.484e-07 | 4.497e-06 |
| 258 | PEPTIDYL TYROSINE MODIFICATION | 17 | 186 | 2.578e-07 | 4.648e-06 |
| 259 | ESTABLISHMENT OF PROTEIN LOCALIZATION | 58 | 1423 | 2.591e-07 | 4.655e-06 |
| 260 | REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY | 15 | 145 | 2.648e-07 | 4.74e-06 |
| 261 | NEGATIVE REGULATION OF CELL DIVISION | 10 | 60 | 3.173e-07 | 5.656e-06 |
| 262 | RESPONSE TO HYDROGEN PEROXIDE | 13 | 109 | 3.201e-07 | 5.686e-06 |
| 263 | RESPONSE TO OXYGEN LEVELS | 22 | 311 | 4.254e-07 | 7.513e-06 |
| 264 | DNA RECOMBINATION | 18 | 215 | 4.263e-07 | 7.513e-06 |
| 265 | NEGATIVE REGULATION OF HISTONE METHYLATION | 6 | 16 | 4.481e-07 | 7.869e-06 |
| 266 | SPINDLE CHECKPOINT | 7 | 25 | 4.651e-07 | 8.106e-06 |
| 267 | POSITIVE REGULATION OF PEPTIDYL THREONINE PHOSPHORYLATION | 7 | 25 | 4.651e-07 | 8.106e-06 |
| 268 | REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 24 | 365 | 4.814e-07 | 8.358e-06 |
| 269 | MAINTENANCE OF CELL NUMBER | 14 | 132 | 4.884e-07 | 8.422e-06 |
| 270 | REGULATION OF PROTEIN COMPLEX DISASSEMBLY | 18 | 217 | 4.887e-07 | 8.422e-06 |
| 271 | REGULATION OF MITOCHONDRION ORGANIZATION | 18 | 218 | 5.229e-07 | 8.913e-06 |
| 272 | CELLULAR RESPONSE TO STEROID HORMONE STIMULUS | 18 | 218 | 5.229e-07 | 8.913e-06 |
| 273 | REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 18 | 218 | 5.229e-07 | 8.913e-06 |
| 274 | RESPONSE TO EXTERNAL STIMULUS | 68 | 1821 | 5.442e-07 | 9.241e-06 |
| 275 | REGULATION OF PROTEIN ACETYLATION | 10 | 64 | 5.931e-07 | 1.003e-05 |
| 276 | REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 18 | 220 | 5.981e-07 | 1.007e-05 |
| 277 | CELLULAR RESPONSE TO ALCOHOL | 13 | 115 | 5.995e-07 | 1.007e-05 |
| 278 | REGULATION OF CYTOPLASMIC TRANSPORT | 28 | 481 | 6.385e-07 | 1.069e-05 |
| 279 | REGULATION OF DNA METHYLATION | 6 | 17 | 6.808e-07 | 1.135e-05 |
| 280 | POSITIVE REGULATION OF PROTEASOMAL PROTEIN CATABOLIC PROCESS | 12 | 98 | 6.836e-07 | 1.136e-05 |
| 281 | DNA GEOMETRIC CHANGE | 11 | 81 | 6.974e-07 | 1.155e-05 |
| 282 | MORPHOGENESIS OF AN EPITHELIUM | 25 | 400 | 7.113e-07 | 1.174e-05 |
| 283 | PROTEIN METHYLATION | 13 | 117 | 7.323e-07 | 1.191e-05 |
| 284 | MAMMARY GLAND DEVELOPMENT | 13 | 117 | 7.323e-07 | 1.191e-05 |
| 285 | REGULATION OF MICROTUBULE POLYMERIZATION OR DEPOLYMERIZATION | 16 | 178 | 7.266e-07 | 1.191e-05 |
| 286 | PROTEIN ALKYLATION | 13 | 117 | 7.323e-07 | 1.191e-05 |
| 287 | CELLULAR RESPONSE TO UV | 10 | 66 | 7.969e-07 | 1.292e-05 |
| 288 | ORGAN REGENERATION | 11 | 83 | 8.954e-07 | 1.447e-05 |
| 289 | MACROMOLECULE DEACYLATION | 10 | 67 | 9.199e-07 | 1.471e-05 |
| 290 | CELL AGING | 10 | 67 | 9.199e-07 | 1.471e-05 |
| 291 | REGULATION OF SISTER CHROMATID SEGREGATION | 10 | 67 | 9.199e-07 | 1.471e-05 |
| 292 | RESPONSE TO OXIDATIVE STRESS | 23 | 352 | 9.315e-07 | 1.484e-05 |
| 293 | ERBB2 SIGNALING PATHWAY | 8 | 39 | 9.37e-07 | 1.488e-05 |
| 294 | NUCLEOBASE BIOSYNTHETIC PROCESS | 6 | 18 | 1.004e-06 | 1.589e-05 |
| 295 | REGULATION OF NEURON DEATH | 19 | 252 | 1.014e-06 | 1.599e-05 |
| 296 | REGULATION OF DNA REPLICATION | 15 | 161 | 1.025e-06 | 1.611e-05 |
| 297 | REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 24 | 381 | 1.036e-06 | 1.623e-05 |
| 298 | TUBE DEVELOPMENT | 30 | 552 | 1.062e-06 | 1.658e-05 |
| 299 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 26 | 437 | 1.078e-06 | 1.674e-05 |
| 300 | DNA BIOSYNTHETIC PROCESS | 13 | 121 | 1.079e-06 | 1.674e-05 |
| 301 | CELLULAR RESPONSE TO OXIDATIVE STRESS | 16 | 184 | 1.132e-06 | 1.75e-05 |
| 302 | REGULATION OF CHROMOSOME SEGREGATION | 11 | 85 | 1.141e-06 | 1.753e-05 |
| 303 | EPHRIN RECEPTOR SIGNALING PATHWAY | 11 | 85 | 1.141e-06 | 1.753e-05 |
| 304 | VENTRICULAR SEPTUM DEVELOPMENT | 9 | 54 | 1.245e-06 | 1.905e-05 |
| 305 | MEIOTIC CELL CYCLE | 16 | 186 | 1.307e-06 | 1.994e-05 |
| 306 | RESPONSE TO METAL ION | 22 | 333 | 1.34e-06 | 2.038e-05 |
| 307 | DOUBLE STRAND BREAK REPAIR | 15 | 165 | 1.399e-06 | 2.121e-05 |
| 308 | METHYLATION | 20 | 284 | 1.532e-06 | 2.315e-05 |
| 309 | MACROMOLECULE CATABOLIC PROCESS | 41 | 926 | 2.366e-06 | 3.563e-05 |
| 310 | REGULATION OF LIGASE ACTIVITY | 13 | 130 | 2.441e-06 | 3.664e-05 |
| 311 | SISTER CHROMATID COHESION | 12 | 111 | 2.62e-06 | 3.908e-05 |
| 312 | NEGATIVE REGULATION OF DNA METABOLIC PROCESS | 12 | 111 | 2.62e-06 | 3.908e-05 |
| 313 | PROTEIN STABILIZATION | 13 | 131 | 2.661e-06 | 3.956e-05 |
| 314 | PROTEOLYSIS | 49 | 1208 | 2.8e-06 | 4.149e-05 |
| 315 | MEMBRANE ORGANIZATION | 40 | 899 | 2.811e-06 | 4.152e-05 |
| 316 | REGULATION OF CENTROSOME DUPLICATION | 7 | 32 | 2.882e-06 | 4.243e-05 |
| 317 | DNA METHYLATION | 8 | 45 | 2.952e-06 | 4.293e-05 |
| 318 | CENTROSOME CYCLE | 8 | 45 | 2.952e-06 | 4.293e-05 |
| 319 | PROTEIN LOCALIZATION TO CHROMOSOME | 8 | 45 | 2.952e-06 | 4.293e-05 |
| 320 | DNA ALKYLATION | 8 | 45 | 2.952e-06 | 4.293e-05 |
| 321 | TUBE MORPHOGENESIS | 21 | 323 | 3.003e-06 | 4.354e-05 |
| 322 | RHYTHMIC PROCESS | 20 | 298 | 3.2e-06 | 4.625e-05 |
| 323 | MITOCHONDRIAL TRANSPORT | 15 | 177 | 3.369e-06 | 4.838e-05 |
| 324 | CHROMATIN ASSEMBLY OR DISASSEMBLY | 15 | 177 | 3.369e-06 | 4.838e-05 |
| 325 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 24 | 408 | 3.414e-06 | 4.888e-05 |
| 326 | CANONICAL WNT SIGNALING PATHWAY | 11 | 95 | 3.486e-06 | 4.976e-05 |
| 327 | RESPONSE TO COCAINE | 8 | 46 | 3.511e-06 | 4.996e-05 |
| 328 | CHROMOSOME LOCALIZATION | 9 | 61 | 3.582e-06 | 5.066e-05 |
| 329 | CELLULAR RESPONSE TO HYDROGEN PEROXIDE | 9 | 61 | 3.582e-06 | 5.066e-05 |
| 330 | POSITIVE REGULATION OF ENDOPLASMIC RETICULUM UNFOLDED PROTEIN RESPONSE | 5 | 13 | 3.74e-06 | 5.273e-05 |
| 331 | REGULATION OF CELL PROJECTION ORGANIZATION | 29 | 558 | 3.881e-06 | 5.456e-05 |
| 332 | NEGATIVE REGULATION OF PROTEOLYSIS | 21 | 329 | 4.005e-06 | 5.614e-05 |
| 333 | NEGATIVE REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 13 | 136 | 4.046e-06 | 5.637e-05 |
| 334 | NUCLEUS ORGANIZATION | 13 | 136 | 4.046e-06 | 5.637e-05 |
| 335 | ERBB SIGNALING PATHWAY | 10 | 79 | 4.307e-06 | 5.982e-05 |
| 336 | RESPONSE TO ALKALOID | 13 | 137 | 4.39e-06 | 6.079e-05 |
| 337 | NEGATIVE REGULATION OF MITOTIC NUCLEAR DIVISION | 7 | 34 | 4.448e-06 | 6.141e-05 |
| 338 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 36 | 788 | 5.06e-06 | 6.945e-05 |
| 339 | CIRCULATORY SYSTEM DEVELOPMENT | 36 | 788 | 5.06e-06 | 6.945e-05 |
| 340 | REGULATION OF MAPK CASCADE | 32 | 660 | 5.204e-06 | 7.122e-05 |
| 341 | DNA DEPENDENT DNA REPLICATION | 11 | 99 | 5.235e-06 | 7.143e-05 |
| 342 | REGULATION OF CYTOKINESIS | 9 | 64 | 5.392e-06 | 7.336e-05 |
| 343 | REGULATION OF TRANSCRIPTION REGULATORY REGION DNA BINDING | 7 | 36 | 6.665e-06 | 9.042e-05 |
| 344 | MICROTUBULE ORGANIZING CENTER ORGANIZATION | 10 | 84 | 7.544e-06 | 0.000102 |
| 345 | CELLULAR RESPONSE TO ABIOTIC STIMULUS | 18 | 263 | 7.582e-06 | 0.0001023 |
| 346 | POSITIVE REGULATION OF MITOTIC CELL CYCLE | 12 | 123 | 7.685e-06 | 0.0001033 |
| 347 | REGULATION OF PEPTIDYL THREONINE PHOSPHORYLATION | 7 | 37 | 8.078e-06 | 0.0001083 |
| 348 | CARDIAC SEPTUM DEVELOPMENT | 10 | 85 | 8.398e-06 | 0.0001123 |
| 349 | INTRACELLULAR RECEPTOR SIGNALING PATHWAY | 14 | 168 | 8.656e-06 | 0.0001154 |
| 350 | RESPONSE TO ACID CHEMICAL | 20 | 319 | 8.855e-06 | 0.0001174 |
| 351 | REGULATION OF MAP KINASE ACTIVITY | 20 | 319 | 8.855e-06 | 0.0001174 |
| 352 | STEROID HORMONE MEDIATED SIGNALING PATHWAY | 12 | 125 | 9.076e-06 | 0.00012 |
| 353 | REGULATION OF GENE SILENCING | 8 | 52 | 9.108e-06 | 0.0001201 |
| 354 | CEREBRAL CORTEX DEVELOPMENT | 11 | 105 | 9.284e-06 | 0.000122 |
| 355 | POSITIVE REGULATION OF DNA REPAIR | 7 | 38 | 9.731e-06 | 0.0001275 |
| 356 | EMBRYONIC ORGAN DEVELOPMENT | 23 | 406 | 1.008e-05 | 0.0001318 |
| 357 | CARDIAC VENTRICLE DEVELOPMENT | 11 | 106 | 1.017e-05 | 0.0001326 |
| 358 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 24 | 437 | 1.086e-05 | 0.0001412 |
| 359 | SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 21 | 352 | 1.13e-05 | 0.0001465 |
| 360 | REGULATION OF MAMMARY GLAND EPITHELIAL CELL PROLIFERATION | 5 | 16 | 1.207e-05 | 0.0001552 |
| 361 | NEGATIVE REGULATION OF CYTOSKELETON ORGANIZATION | 16 | 221 | 1.207e-05 | 0.0001552 |
| 362 | V D J RECOMBINATION | 5 | 16 | 1.207e-05 | 0.0001552 |
| 363 | REGULATION OF CELLULAR AMIDE METABOLIC PROCESS | 21 | 354 | 1.231e-05 | 0.0001578 |
| 364 | REGULATION OF CYTOKINE PRODUCTION | 28 | 563 | 1.29e-05 | 0.0001644 |
| 365 | INTRACELLULAR STEROID HORMONE RECEPTOR SIGNALING PATHWAY | 9 | 71 | 1.287e-05 | 0.0001644 |
| 366 | MULTI ORGANISM REPRODUCTIVE PROCESS | 38 | 891 | 1.319e-05 | 0.0001677 |
| 367 | REGULATION OF HISTONE H3 K4 METHYLATION | 6 | 27 | 1.371e-05 | 0.0001734 |
| 368 | MEIOTIC CELL CYCLE PROCESS | 13 | 152 | 1.369e-05 | 0.0001734 |
| 369 | CELLULAR RESPONSE TO ACID CHEMICAL | 14 | 175 | 1.38e-05 | 0.000174 |
| 370 | NEGATIVE REGULATION OF DNA REPLICATION | 8 | 55 | 1.398e-05 | 0.0001758 |
| 371 | NEGATIVE REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY | 10 | 90 | 1.404e-05 | 0.000176 |
| 372 | POSITIVE REGULATION OF LIGASE ACTIVITY | 11 | 110 | 1.452e-05 | 0.0001816 |
| 373 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 15 | 200 | 1.484e-05 | 0.0001851 |
| 374 | CELLULAR RESPONSE TO LIGHT STIMULUS | 10 | 91 | 1.549e-05 | 0.0001922 |
| 375 | MITOTIC SISTER CHROMATID SEGREGATION | 10 | 91 | 1.549e-05 | 0.0001922 |
| 376 | POSITIVE REGULATION OF CELLULAR AMIDE METABOLIC PROCESS | 11 | 111 | 1.582e-05 | 0.0001958 |
| 377 | REGULATION OF HISTONE METHYLATION | 8 | 56 | 1.602e-05 | 0.0001977 |
| 378 | NEGATIVE REGULATION OF CHROMOSOME SEGREGATION | 6 | 28 | 1.716e-05 | 0.0002096 |
| 379 | REGULATION OF INTRACELLULAR ESTROGEN RECEPTOR SIGNALING PATHWAY | 6 | 28 | 1.716e-05 | 0.0002096 |
| 380 | REGULATION OF MEGAKARYOCYTE DIFFERENTIATION | 6 | 28 | 1.716e-05 | 0.0002096 |
| 381 | REGULATION OF ENDOPLASMIC RETICULUM UNFOLDED PROTEIN RESPONSE | 6 | 28 | 1.716e-05 | 0.0002096 |
| 382 | RESPONSE TO LIGHT STIMULUS | 18 | 280 | 1.776e-05 | 0.0002158 |
| 383 | REGULATION OF PROTEIN TARGETING | 19 | 307 | 1.776e-05 | 0.0002158 |
| 384 | MULTICELLULAR ORGANISM REPRODUCTION | 34 | 768 | 1.795e-05 | 0.0002176 |
| 385 | NEGATIVE REGULATION OF CELLULAR CATABOLIC PROCESS | 13 | 156 | 1.809e-05 | 0.0002183 |
| 386 | PEPTIDYL LYSINE METHYLATION | 9 | 74 | 1.811e-05 | 0.0002183 |
| 387 | MORPHOGENESIS OF EMBRYONIC EPITHELIUM | 12 | 134 | 1.844e-05 | 0.0002217 |
| 388 | METAPHASE PLATE CONGRESSION | 7 | 42 | 1.94e-05 | 0.0002327 |
| 389 | REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 6 | 29 | 2.127e-05 | 0.0002531 |
| 390 | POSITIVE REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 6 | 29 | 2.127e-05 | 0.0002531 |
| 391 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 37 | 876 | 2.123e-05 | 0.0002531 |
| 392 | RESPONSE TO ETHANOL | 12 | 136 | 2.141e-05 | 0.0002541 |
| 393 | PROTEIN SUMOYLATION | 11 | 115 | 2.214e-05 | 0.0002621 |
| 394 | POSITIVE REGULATION OF MAP KINASE ACTIVITY | 15 | 207 | 2.229e-05 | 0.0002632 |
| 395 | REGULATION OF PROTEIN IMPORT | 14 | 183 | 2.284e-05 | 0.0002683 |
| 396 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 40 | 983 | 2.282e-05 | 0.0002683 |
| 397 | IMMUNE SYSTEM DEVELOPMENT | 28 | 582 | 2.349e-05 | 0.0002753 |
| 398 | REGULATION OF INTRACELLULAR STEROID HORMONE RECEPTOR SIGNALING PATHWAY | 8 | 59 | 2.372e-05 | 0.0002766 |
| 399 | DNA METHYLATION OR DEMETHYLATION | 8 | 59 | 2.372e-05 | 0.0002766 |
| 400 | PLACENTA DEVELOPMENT | 12 | 138 | 2.478e-05 | 0.0002883 |
| 401 | RESPONSE TO CYTOKINE | 32 | 714 | 2.51e-05 | 0.0002913 |
| 402 | REGENERATION | 13 | 161 | 2.53e-05 | 0.0002928 |
| 403 | REGULATION OF RELEASE OF CYTOCHROME C FROM MITOCHONDRIA | 7 | 44 | 2.662e-05 | 0.0003073 |
| 404 | REGULATION OF MONOOXYGENASE ACTIVITY | 8 | 60 | 2.689e-05 | 0.0003097 |
| 405 | REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 10 | 97 | 2.721e-05 | 0.0003127 |
| 406 | MUSCLE CELL DIFFERENTIATION | 16 | 237 | 2.861e-05 | 0.0003279 |
| 407 | GENE SILENCING | 15 | 212 | 2.946e-05 | 0.0003368 |
| 408 | ENDOMEMBRANE SYSTEM ORGANIZATION | 24 | 465 | 2.986e-05 | 0.0003406 |
| 409 | CELLULAR RESPONSE TO GAMMA RADIATION | 5 | 19 | 3.056e-05 | 0.0003468 |
| 410 | REGULATION OF DOUBLE STRAND BREAK REPAIR VIA HOMOLOGOUS RECOMBINATION | 5 | 19 | 3.056e-05 | 0.0003468 |
| 411 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 15 | 213 | 3.112e-05 | 0.0003523 |
| 412 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 25 | 498 | 3.229e-05 | 0.0003647 |
| 413 | RECOMBINATIONAL REPAIR | 9 | 80 | 3.414e-05 | 0.0003847 |
| 414 | RESPONSE TO TOXIC SUBSTANCE | 16 | 241 | 3.506e-05 | 0.000394 |
| 415 | NEGATIVE REGULATION OF DNA BINDING | 7 | 46 | 3.59e-05 | 0.0004006 |
| 416 | GROWTH | 22 | 410 | 3.587e-05 | 0.0004006 |
| 417 | NEGATIVE REGULATION OF NUCLEAR DIVISION | 7 | 46 | 3.59e-05 | 0.0004006 |
| 418 | POSITIVE REGULATION OF HOMEOSTATIC PROCESS | 15 | 216 | 3.659e-05 | 0.0004073 |
| 419 | POSITIVE REGULATION OF MITOCHONDRION ORGANIZATION | 13 | 167 | 3.717e-05 | 0.0004128 |
| 420 | REGULATION OF FIBROBLAST PROLIFERATION | 9 | 81 | 3.773e-05 | 0.000417 |
| 421 | NUCLEAR ENVELOPE ORGANIZATION | 9 | 81 | 3.773e-05 | 0.000417 |
| 422 | CARDIAC CHAMBER DEVELOPMENT | 12 | 144 | 3.783e-05 | 0.0004171 |
| 423 | REGULATION OF NEURON APOPTOTIC PROCESS | 14 | 192 | 3.889e-05 | 0.0004278 |
| 424 | ESTABLISHMENT OF LOCALIZATION IN CELL | 58 | 1676 | 3.962e-05 | 0.0004348 |
| 425 | RESPONSE TO VITAMIN A | 5 | 20 | 4.007e-05 | 0.0004387 |
| 426 | MEMBRANE DISASSEMBLY | 7 | 47 | 4.146e-05 | 0.0004518 |
| 427 | NUCLEAR ENVELOPE DISASSEMBLY | 7 | 47 | 4.146e-05 | 0.0004518 |
| 428 | APPENDAGE DEVELOPMENT | 13 | 169 | 4.209e-05 | 0.0004565 |
| 429 | LIMB DEVELOPMENT | 13 | 169 | 4.209e-05 | 0.0004565 |
| 430 | NEGATIVE REGULATION OF CELLULAR PROTEIN CATABOLIC PROCESS | 8 | 64 | 4.335e-05 | 0.000468 |
| 431 | POSITIVE REGULATION OF RESPONSE TO DNA DAMAGE STIMULUS | 8 | 64 | 4.335e-05 | 0.000468 |
| 432 | ANTERIOR POSTERIOR PATTERN SPECIFICATION | 14 | 194 | 4.357e-05 | 0.0004693 |
| 433 | REGULATION OF PROTEIN EXPORT FROM NUCLEUS | 6 | 33 | 4.631e-05 | 0.0004965 |
| 434 | G2 DNA DAMAGE CHECKPOINT | 6 | 33 | 4.631e-05 | 0.0004965 |
| 435 | CELLULAR RESPONSE TO PEPTIDE | 17 | 274 | 4.732e-05 | 0.0005062 |
| 436 | RESPONSE TO AMINE | 7 | 48 | 4.77e-05 | 0.0005079 |
| 437 | CENTROMERE COMPLEX ASSEMBLY | 7 | 48 | 4.77e-05 | 0.0005079 |
| 438 | IMP BIOSYNTHETIC PROCESS | 4 | 11 | 4.884e-05 | 0.0005189 |
| 439 | RESPONSE TO TRANSITION METAL NANOPARTICLE | 12 | 148 | 4.953e-05 | 0.000525 |
| 440 | TELOMERE ORGANIZATION | 10 | 104 | 4.982e-05 | 0.0005269 |
| 441 | NEGATIVE REGULATION OF CELL DEVELOPMENT | 18 | 303 | 5.029e-05 | 0.0005306 |
| 442 | REGULATION OF PROTEIN IMPORT INTO NUCLEUS TRANSLOCATION | 5 | 21 | 5.173e-05 | 0.0005433 |
| 443 | NEGATIVE REGULATION OF PROTEIN ACETYLATION | 5 | 21 | 5.173e-05 | 0.0005433 |
| 444 | STRIATED MUSCLE CELL DIFFERENTIATION | 13 | 173 | 5.364e-05 | 0.0005621 |
| 445 | POSITIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 25 | 514 | 5.397e-05 | 0.000563 |
| 446 | DEVELOPMENTAL GROWTH | 19 | 333 | 5.391e-05 | 0.000563 |
| 447 | LENS DEVELOPMENT IN CAMERA TYPE EYE | 8 | 66 | 5.431e-05 | 0.0005653 |
| 448 | NEGATIVE REGULATION OF KINASE ACTIVITY | 16 | 250 | 5.445e-05 | 0.0005656 |
| 449 | REGULATION OF NITRIC OXIDE SYNTHASE ACTIVITY | 7 | 49 | 5.469e-05 | 0.0005668 |
| 450 | GLIOGENESIS | 13 | 175 | 6.039e-05 | 0.0006244 |
| 451 | LYMPHOCYTE HOMEOSTASIS | 7 | 50 | 6.25e-05 | 0.0006448 |
| 452 | RESPONSE TO RETINOIC ACID | 10 | 107 | 6.356e-05 | 0.0006544 |
| 453 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 11 | 129 | 6.43e-05 | 0.0006576 |
| 454 | PROTEIN ACETYLATION | 11 | 129 | 6.43e-05 | 0.0006576 |
| 455 | TUBE FORMATION | 11 | 129 | 6.43e-05 | 0.0006576 |
| 456 | RESPONSE TO MINERALOCORTICOID | 6 | 35 | 6.558e-05 | 0.0006692 |
| 457 | HISTONE H3 DEACETYLATION | 5 | 22 | 6.583e-05 | 0.0006703 |
| 458 | POSITIVE REGULATION OF CYTOPLASMIC TRANSPORT | 17 | 282 | 6.763e-05 | 0.0006856 |
| 459 | REGULATION OF WNT SIGNALING PATHWAY | 18 | 310 | 6.749e-05 | 0.0006856 |
| 460 | REGIONALIZATION | 18 | 311 | 7.033e-05 | 0.0007114 |
| 461 | PURINE NUCLEOBASE BIOSYNTHETIC PROCESS | 4 | 12 | 7.21e-05 | 0.0007277 |
| 462 | MITOTIC SPINDLE ORGANIZATION | 8 | 69 | 7.5e-05 | 0.0007553 |
| 463 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 30 | 689 | 7.605e-05 | 0.0007643 |
| 464 | CELLULAR RESPONSE TO HEAT | 6 | 36 | 7.737e-05 | 0.0007742 |
| 465 | POSITIVE REGULATION OF PROTEIN ACETYLATION | 6 | 36 | 7.737e-05 | 0.0007742 |
| 466 | RESPONSE TO HEAT | 9 | 89 | 7.975e-05 | 0.000793 |
| 467 | ORGANELLE ASSEMBLY | 24 | 495 | 7.976e-05 | 0.000793 |
| 468 | EPITHELIAL CELL DIFFERENTIATION | 24 | 495 | 7.976e-05 | 0.000793 |
| 469 | POSITIVE REGULATION OF ERYTHROCYTE DIFFERENTIATION | 5 | 23 | 8.273e-05 | 0.0008207 |
| 470 | NON RECOMBINATIONAL REPAIR | 8 | 70 | 8.32e-05 | 0.0008219 |
| 471 | SPINDLE ASSEMBLY | 8 | 70 | 8.32e-05 | 0.0008219 |
| 472 | CELLULAR COMPONENT MORPHOGENESIS | 36 | 900 | 8.403e-05 | 0.0008283 |
| 473 | MIDBRAIN DEVELOPMENT | 9 | 90 | 8.705e-05 | 0.0008546 |
| 474 | REGULATION OF GLIOGENESIS | 9 | 90 | 8.705e-05 | 0.0008546 |
| 475 | GAMETE GENERATION | 27 | 595 | 8.834e-05 | 0.0008654 |
| 476 | REGULATION OF TRANSPORT | 60 | 1804 | 8.865e-05 | 0.0008666 |
| 477 | POSITIVE REGULATION OF CELLULAR COMPONENT BIOGENESIS | 21 | 406 | 9.062e-05 | 0.000884 |
| 478 | SEXUAL REPRODUCTION | 31 | 730 | 9.122e-05 | 0.0008879 |
| 479 | MAMMARY GLAND EPITHELIUM DEVELOPMENT | 7 | 53 | 9.155e-05 | 0.0008893 |
| 480 | HORMONE MEDIATED SIGNALING PATHWAY | 12 | 158 | 9.331e-05 | 0.0009045 |
| 481 | NEGATIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 16 | 262 | 9.468e-05 | 0.0009159 |
| 482 | REGULATION OF GROWTH | 28 | 633 | 0.0001012 | 0.0009767 |
| 483 | POSITIVE REGULATION OF PROTEIN IMPORT INTO NUCLEUS TRANSLOCATION | 4 | 13 | 0.0001025 | 0.0009791 |
| 484 | PROTEIN LOCALIZATION TO CHROMOSOME CENTROMERIC REGION | 4 | 13 | 0.0001025 | 0.0009791 |
| 485 | EYELID DEVELOPMENT IN CAMERA TYPE EYE | 4 | 13 | 0.0001025 | 0.0009791 |
| 486 | HEPATOCYTE APOPTOTIC PROCESS | 4 | 13 | 0.0001025 | 0.0009791 |
| 487 | S ADENOSYLHOMOCYSTEINE METABOLIC PROCESS | 4 | 13 | 0.0001025 | 0.0009791 |
| 488 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 46 | 1275 | 0.0001039 | 0.0009886 |
| 489 | GLIAL CELL DIFFERENTIATION | 11 | 136 | 0.0001038 | 0.0009886 |
| 490 | REGULATION OF DOUBLE STRAND BREAK REPAIR | 6 | 38 | 0.000106 | 0.001006 |
| 491 | WNT SIGNALING PATHWAY | 19 | 351 | 0.0001081 | 0.001025 |
| 492 | HINDBRAIN DEVELOPMENT | 11 | 137 | 0.0001108 | 0.001048 |
| 493 | HIPPOCAMPUS DEVELOPMENT | 8 | 73 | 0.0001124 | 0.001061 |
| 494 | RESPONSE TO GROWTH FACTOR | 23 | 475 | 0.0001142 | 0.001076 |
| 495 | CELLULAR RESPONSE TO CYTOKINE STIMULUS | 27 | 606 | 0.0001195 | 0.001123 |
| 496 | NEURAL TUBE FORMATION | 9 | 94 | 0.0001221 | 0.001146 |
| 497 | NUCLEOBASE METABOLIC PROCESS | 6 | 39 | 0.0001231 | 0.001153 |
| 498 | DNA SYNTHESIS INVOLVED IN DNA REPAIR | 8 | 74 | 0.0001238 | 0.001157 |
| 499 | NUCLEAR TRANSPORT | 19 | 355 | 0.0001253 | 0.001168 |
| 500 | REGULATION OF RNA STABILITY | 11 | 139 | 0.0001261 | 0.001173 |
| 501 | EPITHELIAL CELL APOPTOTIC PROCESS | 5 | 25 | 0.0001263 | 0.001173 |
| 502 | CHROMATIN SILENCING | 9 | 95 | 0.0001325 | 0.001228 |
| 503 | PATTERN SPECIFICATION PROCESS | 21 | 418 | 0.0001359 | 0.001258 |
| 504 | ATTACHMENT OF SPINDLE MICROTUBULES TO KINETOCHORE | 4 | 14 | 0.0001412 | 0.001298 |
| 505 | IMP METABOLIC PROCESS | 4 | 14 | 0.0001412 | 0.001298 |
| 506 | REGULATION OF HISTONE H3 K9 ACETYLATION | 4 | 14 | 0.0001412 | 0.001298 |
| 507 | MESODERM DEVELOPMENT | 10 | 118 | 0.0001448 | 0.001329 |
| 508 | CELLULAR COMPONENT DISASSEMBLY | 24 | 515 | 0.0001457 | 0.001335 |
| 509 | NEGATIVE REGULATION OF NEURON DIFFERENTIATION | 13 | 191 | 0.0001465 | 0.001337 |
| 510 | APOPTOTIC MITOCHONDRIAL CHANGES | 7 | 57 | 0.0001465 | 0.001337 |
| 511 | GLIAL CELL DEVELOPMENT | 8 | 76 | 0.0001495 | 0.001362 |
| 512 | CELLULAR RESPONSE TO INTERLEUKIN 4 | 5 | 26 | 0.0001538 | 0.001394 |
| 513 | REGULATION OF CELL GROWTH | 20 | 391 | 0.000154 | 0.001394 |
| 514 | RESPONSE TO CORTICOSTERONE | 5 | 26 | 0.0001538 | 0.001394 |
| 515 | REGULATION OF DNA RECOMBINATION | 7 | 58 | 0.0001637 | 0.001476 |
| 516 | REGULATION OF HEMATOPOIETIC PROGENITOR CELL DIFFERENTIATION | 6 | 41 | 0.000164 | 0.001476 |
| 517 | MITOTIC RECOMBINATION | 6 | 41 | 0.000164 | 0.001476 |
| 518 | DNA PACKAGING | 13 | 194 | 0.000171 | 0.001536 |
| 519 | NEGATIVE REGULATION OF CELL PROJECTION ORGANIZATION | 11 | 144 | 0.0001725 | 0.001544 |
| 520 | REGULATION OF RESPONSE TO CYTOKINE STIMULUS | 11 | 144 | 0.0001725 | 0.001544 |
| 521 | SINGLE ORGANISM CELLULAR LOCALIZATION | 35 | 898 | 0.000174 | 0.001554 |
| 522 | POSITIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 10 | 121 | 0.0001782 | 0.001588 |
| 523 | SOMITE DEVELOPMENT | 8 | 78 | 0.0001795 | 0.001597 |
| 524 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 17 | 306 | 0.0001815 | 0.001612 |
| 525 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 9 | 99 | 0.0001819 | 0.001612 |
| 526 | NEGATIVE REGULATION OF PROTEIN COMPLEX DISASSEMBLY | 12 | 170 | 0.0001865 | 0.00165 |
| 527 | SOMATIC DIVERSIFICATION OF IMMUNE RECEPTORS | 6 | 42 | 0.0001881 | 0.001661 |
| 528 | NEGATIVE REGULATION OF HISTONE ACETYLATION | 4 | 15 | 0.0001895 | 0.001666 |
| 529 | REGULATION OF SPINDLE ASSEMBLY | 4 | 15 | 0.0001895 | 0.001666 |
| 530 | REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 10 | 122 | 0.0001907 | 0.001674 |
| 531 | REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 18 | 337 | 0.0001924 | 0.001686 |
| 532 | METENCEPHALON DEVELOPMENT | 9 | 100 | 0.0001964 | 0.001711 |
| 533 | DNA MODIFICATION | 8 | 79 | 0.0001963 | 0.001711 |
| 534 | LIMBIC SYSTEM DEVELOPMENT | 9 | 100 | 0.0001964 | 0.001711 |
| 535 | NEGATIVE REGULATION OF NEURON DEATH | 12 | 171 | 0.000197 | 0.001713 |
| 536 | OLIGODENDROCYTE DIFFERENTIATION | 7 | 60 | 0.0002032 | 0.00176 |
| 537 | LEUKOCYTE HOMEOSTASIS | 7 | 60 | 0.0002032 | 0.00176 |
| 538 | ENDOCRINE SYSTEM DEVELOPMENT | 10 | 123 | 0.0002039 | 0.001763 |
| 539 | CELLULAR PROCESS INVOLVED IN REPRODUCTION IN MULTICELLULAR ORGANISM | 15 | 252 | 0.0002058 | 0.001776 |
| 540 | MUSCLE STRUCTURE DEVELOPMENT | 21 | 432 | 0.0002133 | 0.001838 |
| 541 | IMMUNE SYSTEM PROCESS | 63 | 1984 | 0.000214 | 0.00184 |
| 542 | BETA CATENIN TCF COMPLEX ASSEMBLY | 6 | 43 | 0.0002149 | 0.001845 |
| 543 | HEART DEVELOPMENT | 22 | 466 | 0.0002286 | 0.001959 |
| 544 | CELL FATE COMMITMENT | 14 | 227 | 0.0002313 | 0.001979 |
| 545 | CELLULAR CATABOLIC PROCESS | 46 | 1322 | 0.0002367 | 0.002021 |
| 546 | POSITIVE REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 14 | 228 | 0.000242 | 0.002063 |
| 547 | HOMEOSTASIS OF NUMBER OF CELLS | 12 | 175 | 0.0002441 | 0.002077 |
| 548 | REGULATION OF PROTEIN UBIQUITINATION INVOLVED IN UBIQUITIN DEPENDENT PROTEIN CATABOLIC PROCESS | 9 | 103 | 0.0002457 | 0.002086 |
| 549 | POSITIVE REGULATION OF HISTONE H3 K4 METHYLATION | 4 | 16 | 0.0002486 | 0.002099 |
| 550 | REGULATION OF GENE EXPRESSION BY GENETIC IMPRINTING | 4 | 16 | 0.0002486 | 0.002099 |
| 551 | SOMITOGENESIS | 7 | 62 | 0.0002499 | 0.002099 |
| 552 | CARDIAC RIGHT VENTRICLE MORPHOGENESIS | 4 | 16 | 0.0002486 | 0.002099 |
| 553 | REGULATION OF TISSUE REMODELING | 7 | 62 | 0.0002499 | 0.002099 |
| 554 | PIGMENT METABOLIC PROCESS | 7 | 62 | 0.0002499 | 0.002099 |
| 555 | JNK CASCADE | 8 | 82 | 0.0002544 | 0.002133 |
| 556 | PROTEIN DEALKYLATION | 5 | 29 | 0.0002642 | 0.002203 |
| 557 | DNA REPLICATION INITIATION | 5 | 29 | 0.0002642 | 0.002203 |
| 558 | PROTEIN DEMETHYLATION | 5 | 29 | 0.0002642 | 0.002203 |
| 559 | POSITIVE REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 15 | 258 | 0.0002651 | 0.002207 |
| 560 | NEGATIVE REGULATION OF CATABOLIC PROCESS | 13 | 203 | 0.0002668 | 0.002217 |
| 561 | POSITIVE REGULATION OF WNT SIGNALING PATHWAY | 11 | 152 | 0.0002767 | 0.002295 |
| 562 | SUBSTANTIA NIGRA DEVELOPMENT | 6 | 45 | 0.0002775 | 0.002298 |
| 563 | REGULATION OF SYMBIOSIS ENCOMPASSING MUTUALISM THROUGH PARASITISM | 13 | 205 | 0.0002934 | 0.002425 |
| 564 | CYTOKINESIS | 8 | 84 | 0.0003005 | 0.002479 |
| 565 | RESPONSE TO ENDOPLASMIC RETICULUM STRESS | 14 | 233 | 0.0003021 | 0.002488 |
| 566 | HOMEOSTATIC PROCESS | 46 | 1337 | 0.0003039 | 0.002499 |
| 567 | PROTEIN EXPORT FROM NUCLEUS | 5 | 30 | 0.0003118 | 0.00255 |
| 568 | RESPONSE TO AMPHETAMINE | 5 | 30 | 0.0003118 | 0.00255 |
| 569 | RESPONSE TO X RAY | 5 | 30 | 0.0003118 | 0.00255 |
| 570 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 6 | 46 | 0.0003137 | 0.002557 |
| 571 | HISTONE H3 ACETYLATION | 6 | 46 | 0.0003137 | 0.002557 |
| 572 | RESPONSE TO ANGIOTENSIN | 4 | 17 | 0.0003199 | 0.002593 |
| 573 | NEGATIVE REGULATION OF CELL AGING | 4 | 17 | 0.0003199 | 0.002593 |
| 574 | CELL SEPARATION AFTER CYTOKINESIS | 4 | 17 | 0.0003199 | 0.002593 |
| 575 | CELLULAR RESPONSE TO RETINOIC ACID | 7 | 65 | 0.0003358 | 0.002713 |
| 576 | ORGANELLE LOCALIZATION | 20 | 415 | 0.0003353 | 0.002713 |
| 577 | CELLULAR RESPONSE TO EXTERNAL STIMULUS | 15 | 264 | 0.0003388 | 0.002732 |
| 578 | IMMUNE RESPONSE REGULATING CELL SURFACE RECEPTOR SIGNALING PATHWAY | 17 | 323 | 0.0003415 | 0.002749 |
| 579 | REGULATION OF CANONICAL WNT SIGNALING PATHWAY | 14 | 236 | 0.0003439 | 0.002764 |
| 580 | CELLULAR RESPONSE TO INORGANIC SUBSTANCE | 11 | 156 | 0.000346 | 0.002776 |
| 581 | THYMUS DEVELOPMENT | 6 | 47 | 0.0003536 | 0.002822 |
| 582 | PIGMENT BIOSYNTHETIC PROCESS | 6 | 47 | 0.0003536 | 0.002822 |
| 583 | POSITIVE REGULATION OF NEURON APOPTOTIC PROCESS | 6 | 47 | 0.0003536 | 0.002822 |
| 584 | RESPONSE TO INTERLEUKIN 4 | 5 | 31 | 0.0003656 | 0.002908 |
| 585 | MULTIVESICULAR BODY ORGANIZATION | 5 | 31 | 0.0003656 | 0.002908 |
| 586 | NEURAL NUCLEUS DEVELOPMENT | 7 | 66 | 0.0003692 | 0.002932 |
| 587 | POSITIVE REGULATION OF GROWTH | 14 | 238 | 0.0003745 | 0.002969 |
| 588 | POSITIVE REGULATION OF TRANSPORT | 35 | 936 | 0.0003768 | 0.002982 |
| 589 | SENSORY ORGAN MORPHOGENESIS | 14 | 239 | 0.0003907 | 0.003086 |
| 590 | POSITIVE REGULATION OF NEURON DEATH | 7 | 67 | 0.0004053 | 0.003167 |
| 591 | REGULATION OF TELOMERE MAINTENANCE | 7 | 67 | 0.0004053 | 0.003167 |
| 592 | CELL JUNCTION ORGANIZATION | 12 | 185 | 0.0004056 | 0.003167 |
| 593 | RESPONSE TO PLATELET DERIVED GROWTH FACTOR | 4 | 18 | 0.0004047 | 0.003167 |
| 594 | RESPONSE TO CAFFEINE | 4 | 18 | 0.0004047 | 0.003167 |
| 595 | NEGATIVE REGULATION OF MEGAKARYOCYTE DIFFERENTIATION | 4 | 18 | 0.0004047 | 0.003167 |
| 596 | LYMPHOCYTE APOPTOTIC PROCESS | 4 | 18 | 0.0004047 | 0.003167 |
| 597 | PROTEIN ACYLATION | 11 | 159 | 0.0004071 | 0.003173 |
| 598 | MEIOSIS I | 8 | 88 | 0.000413 | 0.003214 |
| 599 | POSITIVE REGULATION OF CELL CYCLE PHASE TRANSITION | 7 | 68 | 0.000444 | 0.003449 |
| 600 | SEGMENTATION | 8 | 89 | 0.000446 | 0.003458 |
| 601 | NEGATIVE REGULATION OF TRANSPORT | 21 | 458 | 0.0004636 | 0.003589 |
| 602 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 37 | 1021 | 0.0004656 | 0.003599 |
| 603 | NEURON DIFFERENTIATION | 33 | 874 | 0.0004675 | 0.003608 |
| 604 | REGULATION OF CELLULAR COMPONENT BIOGENESIS | 30 | 767 | 0.0004773 | 0.003671 |
| 605 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 11 | 162 | 0.0004771 | 0.003671 |
| 606 | REGULATION OF OXIDOREDUCTASE ACTIVITY | 8 | 90 | 0.000481 | 0.003693 |
| 607 | ACTIVATION OF MAPK ACTIVITY | 10 | 137 | 0.0004855 | 0.003717 |
| 608 | RESPONSE TO ACTIVITY | 7 | 69 | 0.0004857 | 0.003717 |
| 609 | SOMATIC CELL DNA RECOMBINATION | 5 | 33 | 0.0004941 | 0.003744 |
| 610 | SIGNAL TRANSDUCTION IN ABSENCE OF LIGAND | 5 | 33 | 0.0004941 | 0.003744 |
| 611 | SOMATIC DIVERSIFICATION OF IMMUNE RECEPTORS VIA GERMLINE RECOMBINATION WITHIN A SINGLE LOCUS | 5 | 33 | 0.0004941 | 0.003744 |
| 612 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 5 | 33 | 0.0004941 | 0.003744 |
| 613 | REGULATION OF BONE RESORPTION | 5 | 33 | 0.0004941 | 0.003744 |
| 614 | POSITIVE REGULATION OF HISTONE METHYLATION | 5 | 33 | 0.0004941 | 0.003744 |
| 615 | POSITIVE REGULATION OF STEM CELL DIFFERENTIATION | 6 | 50 | 0.0004973 | 0.003762 |
| 616 | POSITIVE REGULATION OF HEMOPOIESIS | 11 | 163 | 0.0005025 | 0.00379 |
| 617 | RESPONSE TO TOPOLOGICALLY INCORRECT PROTEIN | 11 | 163 | 0.0005025 | 0.00379 |
| 618 | REGULATION OF INTERLEUKIN 5 PRODUCTION | 4 | 19 | 0.0005045 | 0.003799 |
| 619 | RESPONSE TO VIRUS | 14 | 247 | 0.0005426 | 0.004078 |
| 620 | T CELL HOMEOSTASIS | 5 | 34 | 0.0005697 | 0.004255 |
| 621 | PROTEIN DESTABILIZATION | 5 | 34 | 0.0005697 | 0.004255 |
| 622 | OLIGODENDROCYTE DEVELOPMENT | 5 | 34 | 0.0005697 | 0.004255 |
| 623 | CATABOLIC PROCESS | 56 | 1773 | 0.0005677 | 0.004255 |
| 624 | INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO DNA DAMAGE | 7 | 71 | 0.0005783 | 0.004313 |
| 625 | POSITIVE REGULATION OF CYTOKINE PRODUCTION | 18 | 370 | 0.0005885 | 0.004382 |
| 626 | PROTEIN OLIGOMERIZATION | 20 | 434 | 0.0005907 | 0.004391 |
| 627 | NEGATIVE REGULATION OF RESPONSE TO DNA DAMAGE STIMULUS | 6 | 52 | 0.0006158 | 0.004555 |
| 628 | REGULATION OF SPINDLE ORGANIZATION | 4 | 20 | 0.0006206 | 0.004555 |
| 629 | NEGATIVE REGULATION OF CELL CYCLE ARREST | 4 | 20 | 0.0006206 | 0.004555 |
| 630 | GENETIC IMPRINTING | 4 | 20 | 0.0006206 | 0.004555 |
| 631 | TRACHEA DEVELOPMENT | 4 | 20 | 0.0006206 | 0.004555 |
| 632 | CELLULAR RESPONSE TO IONIZING RADIATION | 6 | 52 | 0.0006158 | 0.004555 |
| 633 | ENDOPLASMIC RETICULUM CALCIUM ION HOMEOSTASIS | 4 | 20 | 0.0006206 | 0.004555 |
| 634 | RESPONSE TO LEAD ION | 4 | 20 | 0.0006206 | 0.004555 |
| 635 | RESPONSE TO PEPTIDE | 19 | 404 | 0.0006306 | 0.004617 |
| 636 | NEGATIVE REGULATION OF CYTOPLASMIC TRANSPORT | 9 | 117 | 0.0006311 | 0.004617 |
| 637 | REGULATION OF AXONOGENESIS | 11 | 168 | 0.0006474 | 0.004729 |
| 638 | REGULATION OF MUSCLE SYSTEM PROCESS | 12 | 195 | 0.0006495 | 0.004737 |
| 639 | ACTIVATION OF JUN KINASE ACTIVITY | 5 | 35 | 0.0006537 | 0.004753 |
| 640 | RESPONSE TO IRON ION | 5 | 35 | 0.0006537 | 0.004753 |
| 641 | POSITIVE REGULATION OF FIBROBLAST PROLIFERATION | 6 | 53 | 0.0006827 | 0.004917 |
| 642 | HISTONE EXCHANGE | 6 | 53 | 0.0006827 | 0.004917 |
| 643 | RAS PROTEIN SIGNAL TRANSDUCTION | 10 | 143 | 0.0006796 | 0.004917 |
| 644 | DNA REPLICATION INDEPENDENT NUCLEOSOME ASSEMBLY | 6 | 53 | 0.0006827 | 0.004917 |
| 645 | DNA REPLICATION INDEPENDENT NUCLEOSOME ORGANIZATION | 6 | 53 | 0.0006827 | 0.004917 |
| 646 | NEGATIVE REGULATION OF INTRACELLULAR TRANSPORT | 10 | 143 | 0.0006796 | 0.004917 |
| 647 | POSITIVE REGULATION OF PROTEIN BINDING | 7 | 73 | 0.0006845 | 0.004923 |
| 648 | REGULATION OF PROTEIN COMPLEX ASSEMBLY | 18 | 375 | 0.0006877 | 0.004938 |
| 649 | REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 12 | 197 | 0.0007106 | 0.005087 |
| 650 | POSITIVE REGULATION OF PROTEIN COMPLEX ASSEMBLY | 12 | 197 | 0.0007106 | 0.005087 |
| 651 | REGULATION OF HEMOPOIESIS | 16 | 314 | 0.0007172 | 0.005126 |
| 652 | REGULATION OF ERYTHROCYTE DIFFERENTIATION | 5 | 36 | 0.0007467 | 0.005329 |
| 653 | REGULATION OF GENE SILENCING BY RNA | 4 | 21 | 0.0007545 | 0.005332 |
| 654 | B CELL HOMEOSTASIS | 4 | 21 | 0.0007545 | 0.005332 |
| 655 | SYMPATHETIC NERVOUS SYSTEM DEVELOPMENT | 4 | 21 | 0.0007545 | 0.005332 |
| 656 | PURINE NUCLEOBASE METABOLIC PROCESS | 4 | 21 | 0.0007545 | 0.005332 |
| 657 | REGULATION OF MITOCHONDRIAL MEMBRANE POTENTIAL | 6 | 54 | 0.0007551 | 0.005332 |
| 658 | REGULATION OF POSTTRANSCRIPTIONAL GENE SILENCING | 4 | 21 | 0.0007545 | 0.005332 |
| 659 | ANATOMICAL STRUCTURE HOMEOSTASIS | 15 | 285 | 0.0007509 | 0.005332 |
| 660 | REGULATION OF CELL SIZE | 11 | 172 | 0.0007869 | 0.005548 |
| 661 | REGULATION OF INTERFERON GAMMA PRODUCTION | 8 | 97 | 0.0007934 | 0.005582 |
| 662 | NEURON PROJECTION DEVELOPMENT | 23 | 545 | 0.0007942 | 0.005582 |
| 663 | RESPONSE TO NERVE GROWTH FACTOR | 5 | 37 | 0.0008492 | 0.005933 |
| 664 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 8 | 98 | 0.000849 | 0.005933 |
| 665 | VIRION ASSEMBLY | 5 | 37 | 0.0008492 | 0.005933 |
| 666 | NEGATIVE REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 5 | 37 | 0.0008492 | 0.005933 |
| 667 | REGULATION OF CELLULAR RESPONSE TO HEAT | 7 | 76 | 0.0008721 | 0.006084 |
| 668 | POSITIVE REGULATION OF CELL GROWTH | 10 | 148 | 0.0008865 | 0.006175 |
| 669 | REGULATION OF PROTEIN HOMODIMERIZATION ACTIVITY | 4 | 22 | 0.0009075 | 0.006293 |
| 670 | RELEASE OF CYTOCHROME C FROM MITOCHONDRIA | 4 | 22 | 0.0009075 | 0.006293 |
| 671 | POSITIVE REGULATION OF NITRIC OXIDE SYNTHASE ACTIVITY | 4 | 22 | 0.0009075 | 0.006293 |
| 672 | CELL MOTILITY | 31 | 835 | 0.0009103 | 0.006293 |
| 673 | LOCALIZATION OF CELL | 31 | 835 | 0.0009103 | 0.006293 |
| 674 | CELLULAR MACROMOLECULAR COMPLEX ASSEMBLY | 28 | 727 | 0.0009262 | 0.006394 |
| 675 | RESPONSE TO CORTICOSTEROID | 11 | 176 | 0.0009505 | 0.006552 |
| 676 | RESPONSE TO TESTOSTERONE | 5 | 38 | 0.0009618 | 0.00661 |
| 677 | REGULATION OF CD4 POSITIVE ALPHA BETA T CELL ACTIVATION | 5 | 38 | 0.0009618 | 0.00661 |
| 678 | MALE GAMETE GENERATION | 21 | 486 | 0.0009891 | 0.006788 |
| 679 | FC RECEPTOR SIGNALING PATHWAY | 12 | 206 | 0.001049 | 0.007188 |
| 680 | LEUKOCYTE APOPTOTIC PROCESS | 4 | 23 | 0.001081 | 0.007398 |
| 681 | CELLULAR RESPONSE TO NUTRIENT | 5 | 39 | 0.001085 | 0.007403 |
| 682 | NEGATIVE REGULATION OF MITOCHONDRION ORGANIZATION | 5 | 39 | 0.001085 | 0.007403 |
| 683 | DENDRITE DEVELOPMENT | 7 | 79 | 0.001098 | 0.00747 |
| 684 | REGULATION OF STRIATED MUSCLE CONTRACTION | 7 | 79 | 0.001098 | 0.00747 |
| 685 | CELLULAR GLUCAN METABOLIC PROCESS | 6 | 58 | 0.001106 | 0.007501 |
| 686 | GLUCAN METABOLIC PROCESS | 6 | 58 | 0.001106 | 0.007501 |
| 687 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 10 | 153 | 0.001143 | 0.007739 |
| 688 | STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 8 | 103 | 0.001176 | 0.00795 |
| 689 | NEGATIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 12 | 209 | 0.001188 | 0.007999 |
| 690 | RESPONSE TO BIOTIC STIMULUS | 32 | 886 | 0.001185 | 0.007999 |
| 691 | GERM CELL DEVELOPMENT | 12 | 209 | 0.001188 | 0.007999 |
| 692 | RESPONSE TO UV C | 3 | 11 | 0.001205 | 0.00801 |
| 693 | PROTEIN LOCALIZATION TO KINETOCHORE | 3 | 11 | 0.001205 | 0.00801 |
| 694 | LOCOMOTION | 38 | 1114 | 0.001202 | 0.00801 |
| 695 | REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION TO MITOCHONDRION | 9 | 128 | 0.0012 | 0.00801 |
| 696 | ORGANELLE INHERITANCE | 3 | 11 | 0.001205 | 0.00801 |
| 697 | REGULATION OF HISTONE H4 ACETYLATION | 3 | 11 | 0.001205 | 0.00801 |
| 698 | LYMPHOID PROGENITOR CELL DIFFERENTIATION | 3 | 11 | 0.001205 | 0.00801 |
| 699 | MUSCLE CELL DEVELOPMENT | 9 | 128 | 0.0012 | 0.00801 |
| 700 | CORPUS CALLOSUM DEVELOPMENT | 3 | 11 | 0.001205 | 0.00801 |
| 701 | REGULATION OF CELL CYCLE G2 M PHASE TRANSITION | 6 | 59 | 0.00121 | 0.008034 |
| 702 | MAMMARY GLAND MORPHOGENESIS | 5 | 40 | 0.00122 | 0.008074 |
| 703 | CYTOPLASMIC SEQUESTERING OF PROTEIN | 5 | 40 | 0.00122 | 0.008074 |
| 704 | CARDIAC CHAMBER MORPHOGENESIS | 8 | 104 | 0.001251 | 0.008248 |
| 705 | POSITIVE REGULATION OF PROTEIN IMPORT | 8 | 104 | 0.001251 | 0.008248 |
| 706 | DEVELOPMENTAL GROWTH INVOLVED IN MORPHOGENESIS | 8 | 104 | 0.001251 | 0.008248 |
| 707 | VIRAL BUDDING | 4 | 24 | 0.001277 | 0.008356 |
| 708 | REGULATION OF HISTONE DEACETYLATION | 4 | 24 | 0.001277 | 0.008356 |
| 709 | NEGATIVE REGULATION OF HEMATOPOIETIC PROGENITOR CELL DIFFERENTIATION | 4 | 24 | 0.001277 | 0.008356 |
| 710 | MULTI ORGANISM ORGANELLE ORGANIZATION | 4 | 24 | 0.001277 | 0.008356 |
| 711 | MULTI ORGANISM MEMBRANE BUDDING | 4 | 24 | 0.001277 | 0.008356 |
| 712 | NEGATIVE REGULATION OF CYTOKINE PRODUCTION | 12 | 211 | 0.001289 | 0.008422 |
| 713 | REGULATION OF MYELOID CELL DIFFERENTIATION | 11 | 183 | 0.001304 | 0.008511 |
| 714 | INTRACELLULAR PROTEIN TRANSPORT | 29 | 781 | 0.00132 | 0.008604 |
| 715 | MITOTIC SPINDLE ASSEMBLY | 5 | 41 | 0.001366 | 0.008855 |
| 716 | MICROTUBULE CYTOSKELETON ORGANIZATION INVOLVED IN MITOSIS | 5 | 41 | 0.001366 | 0.008855 |
| 717 | CELLULAR RESPONSE TO ESTROGEN STIMULUS | 5 | 41 | 0.001366 | 0.008855 |
| 718 | ANDROGEN RECEPTOR SIGNALING PATHWAY | 5 | 41 | 0.001366 | 0.008855 |
| 719 | POSITIVE REGULATION OF MULTI ORGANISM PROCESS | 10 | 157 | 0.001389 | 0.008986 |
| 720 | REGULATION OF GLUCOSE METABOLIC PROCESS | 8 | 106 | 0.001415 | 0.009145 |
| 721 | EMBRYONIC PLACENTA DEVELOPMENT | 7 | 83 | 0.001468 | 0.009462 |
| 722 | NUCLEOSIDE MONOPHOSPHATE BIOSYNTHETIC PROCESS | 7 | 83 | 0.001468 | 0.009462 |
| 723 | LENS FIBER CELL DIFFERENTIATION | 4 | 25 | 0.001496 | 0.009615 |
| 724 | SIGNAL PEPTIDE PROCESSING | 4 | 25 | 0.001496 | 0.009615 |
| 725 | VASCULATURE DEVELOPMENT | 20 | 469 | 0.001515 | 0.00972 |
| 726 | DENDRITE MORPHOGENESIS | 5 | 42 | 0.001526 | 0.009765 |
| 727 | REGULATION OF BONE REMODELING | 5 | 42 | 0.001526 | 0.009765 |
| 728 | NEGATIVE REGULATION OF PEPTIDASE ACTIVITY | 13 | 245 | 0.001541 | 0.009852 |
| 729 | POSITIVE REGULATION OF MAPK CASCADE | 20 | 470 | 0.001553 | 0.009913 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | RIBONUCLEOTIDE BINDING | 114 | 1860 | 4.114e-28 | 3.822e-25 |
| 2 | PROTEIN KINASE ACTIVITY | 62 | 640 | 5.208e-25 | 2.419e-22 |
| 3 | ENZYME BINDING | 104 | 1737 | 1.213e-24 | 3.756e-22 |
| 4 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 76 | 992 | 3.824e-24 | 8.882e-22 |
| 5 | KINASE ACTIVITY | 69 | 842 | 1.463e-23 | 2.717e-21 |
| 6 | ADENYL NUCLEOTIDE BINDING | 94 | 1514 | 2.936e-23 | 4.545e-21 |
| 7 | KINASE BINDING | 57 | 606 | 2.195e-22 | 2.913e-20 |
| 8 | MACROMOLECULAR COMPLEX BINDING | 88 | 1399 | 4.307e-22 | 5.001e-20 |
| 9 | TRANSCRIPTION FACTOR ACTIVITY PROTEIN BINDING | 55 | 588 | 1.679e-21 | 1.733e-19 |
| 10 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 47 | 445 | 1.25e-20 | 1.161e-18 |
| 11 | CHROMATIN BINDING | 45 | 435 | 1.989e-19 | 1.68e-17 |
| 12 | TRANSCRIPTION FACTOR BINDING | 48 | 524 | 1.782e-18 | 1.38e-16 |
| 13 | SEQUENCE SPECIFIC DNA BINDING | 67 | 1037 | 2.274e-17 | 1.625e-15 |
| 14 | REGULATORY REGION NUCLEIC ACID BINDING | 58 | 818 | 6.668e-17 | 4.425e-15 |
| 15 | HISTONE DEACETYLASE BINDING | 22 | 105 | 1.636e-16 | 1.013e-14 |
| 16 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 22 | 133 | 2.998e-14 | 1.741e-12 |
| 17 | DOUBLE STRANDED DNA BINDING | 51 | 764 | 6.527e-14 | 3.567e-12 |
| 18 | TRANSCRIPTION COACTIVATOR ACTIVITY | 31 | 296 | 7.419e-14 | 3.829e-12 |
| 19 | IDENTICAL PROTEIN BINDING | 64 | 1209 | 1.37e-12 | 6.7e-11 |
| 20 | PROTEIN DOMAIN SPECIFIC BINDING | 43 | 624 | 2.493e-12 | 1.158e-10 |
| 21 | CORE PROMOTER BINDING | 20 | 152 | 3.521e-11 | 1.558e-09 |
| 22 | RNA POLYMERASE II TRANSCRIPTION COFACTOR ACTIVITY | 16 | 91 | 3.881e-11 | 1.639e-09 |
| 23 | ENHANCER BINDING | 16 | 93 | 5.469e-11 | 2.209e-09 |
| 24 | HISTONE METHYLTRANSFERASE ACTIVITY | 13 | 58 | 1.099e-10 | 4.254e-09 |
| 25 | PROTEIN COMPLEX BINDING | 50 | 935 | 3.686e-10 | 1.37e-08 |
| 26 | HISTONE KINASE ACTIVITY | 8 | 19 | 1.641e-09 | 5.865e-08 |
| 27 | STRUCTURE SPECIFIC DNA BINDING | 16 | 118 | 2.109e-09 | 7.256e-08 |
| 28 | NUCLEOSOMAL DNA BINDING | 9 | 30 | 5.146e-09 | 1.707e-07 |
| 29 | PROTEIN METHYLTRANSFERASE ACTIVITY | 13 | 82 | 1.001e-08 | 3.206e-07 |
| 30 | PROTEIN DEACETYLASE ACTIVITY | 10 | 43 | 1.096e-08 | 3.362e-07 |
| 31 | CORE PROMOTER PROXIMAL REGION DNA BINDING | 27 | 371 | 1.122e-08 | 3.362e-07 |
| 32 | PROTEIN DIMERIZATION ACTIVITY | 53 | 1149 | 1.785e-08 | 5.025e-07 |
| 33 | NUCLEOSOME BINDING | 10 | 45 | 1.76e-08 | 5.025e-07 |
| 34 | RNA POLYMERASE II TRANSCRIPTION FACTOR ACTIVITY SEQUENCE SPECIFIC DNA BINDING | 36 | 629 | 2.331e-08 | 6.188e-07 |
| 35 | S ADENOSYLMETHIONINE DEPENDENT METHYLTRANSFERASE ACTIVITY | 16 | 139 | 2.326e-08 | 6.188e-07 |
| 36 | HEAT SHOCK PROTEIN BINDING | 13 | 89 | 2.775e-08 | 7.162e-07 |
| 37 | NUCLEIC ACID BINDING TRANSCRIPTION FACTOR ACTIVITY | 54 | 1199 | 2.876e-08 | 7.22e-07 |
| 38 | TRANSFERASE ACTIVITY TRANSFERRING ONE CARBON GROUPS | 20 | 225 | 3.6e-08 | 8.8e-07 |
| 39 | HORMONE RECEPTOR BINDING | 17 | 168 | 5.856e-08 | 1.395e-06 |
| 40 | CHROMATIN DNA BINDING | 12 | 80 | 7.037e-08 | 1.634e-06 |
| 41 | PROTEIN TYROSINE KINASE ACTIVITY | 17 | 176 | 1.159e-07 | 2.626e-06 |
| 42 | BETA CATENIN BINDING | 12 | 84 | 1.227e-07 | 2.714e-06 |
| 43 | DEACETYLASE ACTIVITY | 10 | 55 | 1.347e-07 | 2.91e-06 |
| 44 | RECEPTOR BINDING | 60 | 1476 | 1.711e-07 | 3.612e-06 |
| 45 | N METHYLTRANSFERASE ACTIVITY | 12 | 88 | 2.073e-07 | 4.281e-06 |
| 46 | HISTONE LYSINE N METHYLTRANSFERASE ACTIVITY | 9 | 45 | 2.436e-07 | 4.92e-06 |
| 47 | CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 8 | 34 | 3.021e-07 | 5.972e-06 |
| 48 | PROTEIN HETERODIMERIZATION ACTIVITY | 28 | 468 | 3.688e-07 | 7.138e-06 |
| 49 | PROTEIN HOMODIMERIZATION ACTIVITY | 36 | 722 | 6.834e-07 | 1.296e-05 |
| 50 | STEROID HORMONE RECEPTOR BINDING | 11 | 81 | 6.974e-07 | 1.296e-05 |
| 51 | CORE PROMOTER SEQUENCE SPECIFIC DNA BINDING | 12 | 101 | 9.499e-07 | 1.73e-05 |
| 52 | RNA BINDING | 61 | 1598 | 1.124e-06 | 2.009e-05 |
| 53 | RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 12 | 104 | 1.304e-06 | 2.286e-05 |
| 54 | DNA BINDING BENDING | 6 | 20 | 2.025e-06 | 3.31e-05 |
| 55 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION REGULATORY REGION SEQUENCE SPECIFIC BINDING | 21 | 315 | 2.021e-06 | 3.31e-05 |
| 56 | UBIQUITIN LIKE PROTEIN LIGASE BINDING | 19 | 264 | 2.031e-06 | 3.31e-05 |
| 57 | LYSINE N METHYLTRANSFERASE ACTIVITY | 9 | 57 | 1.996e-06 | 3.31e-05 |
| 58 | HISTONE DEACETYLASE ACTIVITY H3 K14 SPECIFIC | 5 | 12 | 2.34e-06 | 3.748e-05 |
| 59 | HISTONE BINDING | 15 | 177 | 3.369e-06 | 5.305e-05 |
| 60 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 17 | 226 | 3.878e-06 | 6.005e-05 |
| 61 | RNA POLYMERASE II DISTAL ENHANCER SEQUENCE SPECIFIC DNA BINDING | 9 | 65 | 6.148e-06 | 9.363e-05 |
| 62 | HYDROLASE ACTIVITY ACTING ON CARBON NITROGEN BUT NOT PEPTIDE BONDS IN LINEAR AMIDES | 10 | 84 | 7.544e-06 | 0.000113 |
| 63 | LIGAND DEPENDENT NUCLEAR RECEPTOR TRANSCRIPTION COACTIVATOR ACTIVITY | 8 | 53 | 1.054e-05 | 0.0001554 |
| 64 | HISTONE DEMETHYLASE ACTIVITY | 6 | 26 | 1.085e-05 | 0.0001575 |
| 65 | PROTEIN SERINE THREONINE TYROSINE KINASE ACTIVITY | 7 | 39 | 1.165e-05 | 0.0001666 |
| 66 | HYDROLASE ACTIVITY ACTING ON ACID ANHYDRIDES | 36 | 820 | 1.21e-05 | 0.0001704 |
| 67 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 20 | 328 | 1.331e-05 | 0.0001845 |
| 68 | ION CHANNEL BINDING | 11 | 111 | 1.582e-05 | 0.0002162 |
| 69 | GUANYL NUCLEOTIDE BINDING | 22 | 390 | 1.677e-05 | 0.0002232 |
| 70 | NAD DEPENDENT PROTEIN DEACETYLASE ACTIVITY | 5 | 17 | 1.682e-05 | 0.0002232 |
| 71 | BHLH TRANSCRIPTION FACTOR BINDING | 6 | 28 | 1.716e-05 | 0.0002245 |
| 72 | KINASE REGULATOR ACTIVITY | 14 | 186 | 2.738e-05 | 0.0003532 |
| 73 | DNA DEPENDENT ATPASE ACTIVITY | 9 | 79 | 3.085e-05 | 0.0003872 |
| 74 | EPHRIN RECEPTOR ACTIVITY | 5 | 19 | 3.056e-05 | 0.0003872 |
| 75 | HYDROLASE ACTIVITY ACTING ON CARBON NITROGEN BUT NOT PEPTIDE BONDS | 12 | 143 | 3.531e-05 | 0.0004374 |
| 76 | NON MEMBRANE SPANNING PROTEIN TYROSINE KINASE ACTIVITY | 7 | 46 | 3.59e-05 | 0.0004389 |
| 77 | GTPASE ACTIVITY | 16 | 246 | 4.49e-05 | 0.0005417 |
| 78 | PROTEIN N TERMINUS BINDING | 10 | 103 | 4.585e-05 | 0.000546 |
| 79 | DEMETHYLASE ACTIVITY | 6 | 34 | 5.528e-05 | 0.00065 |
| 80 | PEPTIDE N ACETYLTRANSFERASE ACTIVITY | 8 | 67 | 6.06e-05 | 0.0006895 |
| 81 | TRANSCRIPTIONAL REPRESSOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 9 | 86 | 6.086e-05 | 0.0006895 |
| 82 | P53 BINDING | 8 | 67 | 6.06e-05 | 0.0006895 |
| 83 | N ACETYLTRANSFERASE ACTIVITY | 9 | 87 | 6.668e-05 | 0.0007464 |
| 84 | TAU PROTEIN KINASE ACTIVITY | 4 | 12 | 7.21e-05 | 0.0007974 |
| 85 | TRANSCRIPTIONAL REPRESSOR ACTIVITY RNA POLYMERASE II ACTIVATING TRANSCRIPTION FACTOR BINDING | 7 | 53 | 9.155e-05 | 0.000989 |
| 86 | RNA POLYMERASE II CORE PROMOTER SEQUENCE SPECIFIC DNA BINDING | 7 | 53 | 9.155e-05 | 0.000989 |
| 87 | PROTEIN C TERMINUS BINDING | 13 | 186 | 0.0001123 | 0.001199 |
| 88 | LIGASE REGULATOR ACTIVITY | 4 | 14 | 0.0001412 | 0.00149 |
| 89 | REPRESSING TRANSCRIPTION FACTOR BINDING | 7 | 57 | 0.0001465 | 0.001529 |
| 90 | TRANSCRIPTION COREPRESSOR ACTIVITY | 14 | 221 | 0.0001753 | 0.00181 |
| 91 | ZINC ION BINDING | 42 | 1155 | 0.0001783 | 0.00182 |
| 92 | FOUR WAY JUNCTION DNA BINDING | 4 | 15 | 0.0001895 | 0.001913 |
| 93 | POLY A RNA BINDING | 42 | 1170 | 0.0002343 | 0.002341 |
| 94 | ACETYLTRANSFERASE ACTIVITY | 9 | 103 | 0.0002457 | 0.002428 |
| 95 | N ACYLTRANSFERASE ACTIVITY | 9 | 104 | 0.0002642 | 0.002584 |
| 96 | NF KAPPAB BINDING | 5 | 30 | 0.0003118 | 0.003017 |
| 97 | RAN GTPASE BINDING | 5 | 31 | 0.0003656 | 0.003502 |
| 98 | NITRIC OXIDE SYNTHASE BINDING | 4 | 19 | 0.0005045 | 0.004783 |
| 99 | UBIQUITIN LIKE PROTEIN LIGASE ACTIVITY | 12 | 199 | 0.0007766 | 0.007287 |
| 100 | TRANSITION METAL ION BINDING | 46 | 1400 | 0.0008136 | 0.007558 |
| 101 | UBIQUITIN LIKE PROTEIN TRANSFERASE ACTIVITY | 19 | 420 | 0.0009998 | 0.009197 |
| 102 | ANDROGEN RECEPTOR BINDING | 5 | 39 | 0.001085 | 0.009601 |
| 103 | ZINC ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 4 | 23 | 0.001081 | 0.009601 |
| 104 | CYSTEINE TYPE ENDOPEPTIDASE INHIBITOR ACTIVITY INVOLVED IN APOPTOTIC PROCESS | 4 | 23 | 0.001081 | 0.009601 |
| 105 | TRANSITION METAL ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 5 | 39 | 0.001085 | 0.009601 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CHROMOSOME | 76 | 880 | 1.963e-27 | 1.146e-24 |
| 2 | NUCLEAR CHROMOSOME | 56 | 523 | 9.228e-25 | 2.694e-22 |
| 3 | TRANSCRIPTION FACTOR COMPLEX | 37 | 298 | 8.861e-19 | 1.725e-16 |
| 4 | CHROMATIN | 42 | 441 | 6.999e-17 | 1.022e-14 |
| 5 | NUCLEAR TRANSCRIPTION FACTOR COMPLEX | 23 | 127 | 9.874e-16 | 1.153e-13 |
| 6 | CHROMOSOMAL REGION | 34 | 330 | 7.139e-15 | 6.948e-13 |
| 7 | TRANSFERASE COMPLEX | 47 | 703 | 6.508e-13 | 5.429e-11 |
| 8 | RNA POLYMERASE II TRANSCRIPTION FACTOR COMPLEX | 18 | 101 | 1.807e-12 | 1.173e-10 |
| 9 | NUCLEAR CHROMATIN | 29 | 291 | 1.706e-12 | 1.173e-10 |
| 10 | CATALYTIC COMPLEX | 58 | 1038 | 2.023e-12 | 1.181e-10 |
| 11 | NUCLEOPLASM PART | 46 | 708 | 3.168e-12 | 1.682e-10 |
| 12 | MICROTUBULE CYTOSKELETON | 58 | 1068 | 6.445e-12 | 2.895e-10 |
| 13 | PERINUCLEAR REGION OF CYTOPLASM | 43 | 642 | 6.359e-12 | 2.895e-10 |
| 14 | CYTOSKELETAL PART | 69 | 1436 | 1.395e-11 | 5.821e-10 |
| 15 | MEDIATOR COMPLEX | 11 | 34 | 3.863e-11 | 1.504e-09 |
| 16 | MICROTUBULE ORGANIZING CENTER | 40 | 623 | 1.303e-10 | 4.756e-09 |
| 17 | CENTROSOME | 34 | 487 | 3.949e-10 | 1.356e-08 |
| 18 | CONDENSED CHROMOSOME | 21 | 195 | 5.104e-10 | 1.656e-08 |
| 19 | CYTOSKELETON | 80 | 1967 | 9.125e-10 | 2.805e-08 |
| 20 | MIDBODY | 16 | 132 | 1.102e-08 | 3.217e-07 |
| 21 | CONDENSED NUCLEAR CHROMOSOME | 13 | 85 | 1.569e-08 | 4.363e-07 |
| 22 | CHD TYPE COMPLEX | 7 | 17 | 2.165e-08 | 5.747e-07 |
| 23 | CONDENSED NUCLEAR CHROMOSOME CENTROMERIC REGION | 7 | 18 | 3.481e-08 | 8.47e-07 |
| 24 | CHROMOSOME TELOMERIC REGION | 17 | 162 | 3.414e-08 | 8.47e-07 |
| 25 | SEX CHROMOSOME | 8 | 27 | 4.183e-08 | 9.771e-07 |
| 26 | CHROMOSOME CENTROMERIC REGION | 17 | 174 | 9.808e-08 | 2.203e-06 |
| 27 | SPINDLE | 22 | 289 | 1.2e-07 | 2.595e-06 |
| 28 | PCG PROTEIN COMPLEX | 9 | 43 | 1.607e-07 | 3.351e-06 |
| 29 | NUCLEAR TRANSCRIPTIONAL REPRESSOR COMPLEX | 7 | 22 | 1.739e-07 | 3.437e-06 |
| 30 | SWI SNF SUPERFAMILY TYPE COMPLEX | 11 | 71 | 1.765e-07 | 3.437e-06 |
| 31 | SPINDLE MICROTUBULE | 10 | 58 | 2.276e-07 | 4.288e-06 |
| 32 | HISTONE DEACETYLASE COMPLEX | 10 | 61 | 3.727e-07 | 6.802e-06 |
| 33 | ESC E Z COMPLEX | 6 | 16 | 4.481e-07 | 7.931e-06 |
| 34 | PROTEIN DNA COMPLEX | 16 | 175 | 5.778e-07 | 9.925e-06 |
| 35 | HISTONE METHYLTRANSFERASE COMPLEX | 10 | 71 | 1.594e-06 | 2.659e-05 |
| 36 | METHYLTRANSFERASE COMPLEX | 11 | 90 | 2.033e-06 | 3.298e-05 |
| 37 | TRANSCRIPTIONAL REPRESSOR COMPLEX | 10 | 74 | 2.35e-06 | 3.611e-05 |
| 38 | CONDENSED CHROMOSOME OUTER KINETOCHORE | 5 | 12 | 2.34e-06 | 3.611e-05 |
| 39 | NUCLEAR CHROMOSOME TELOMERIC REGION | 13 | 132 | 2.898e-06 | 4.34e-05 |
| 40 | TRANSCRIPTION FACTOR TFTC COMPLEX | 5 | 14 | 5.721e-06 | 8.352e-05 |
| 41 | KINETOCHORE | 12 | 120 | 5.948e-06 | 8.472e-05 |
| 42 | UBIQUITIN LIGASE COMPLEX | 18 | 262 | 7.195e-06 | 9.694e-05 |
| 43 | CELL SUBSTRATE JUNCTION | 23 | 398 | 7.304e-06 | 9.694e-05 |
| 44 | CONDENSED CHROMOSOME CENTROMERIC REGION | 11 | 102 | 7.008e-06 | 9.694e-05 |
| 45 | HETEROCHROMATIN | 9 | 67 | 7.935e-06 | 0.000103 |
| 46 | PRONUCLEUS | 5 | 15 | 8.439e-06 | 0.0001049 |
| 47 | XY BODY | 5 | 15 | 8.439e-06 | 0.0001049 |
| 48 | SPINDLE POLE | 12 | 126 | 9.851e-06 | 0.0001199 |
| 49 | NUCLEOLUS | 37 | 848 | 1.045e-05 | 0.0001246 |
| 50 | SPINDLE MIDZONE | 6 | 27 | 1.371e-05 | 0.0001602 |
| 51 | ANCHORING JUNCTION | 25 | 489 | 2.391e-05 | 0.0002737 |
| 52 | TRANSFERASE COMPLEX TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 16 | 237 | 2.861e-05 | 0.0003213 |
| 53 | EUCHROMATIN | 6 | 31 | 3.186e-05 | 0.000351 |
| 54 | PIGMENT GRANULE | 10 | 103 | 4.585e-05 | 0.0004958 |
| 55 | NUCLEAR EUCHROMATIN | 5 | 24 | 0.0001028 | 0.001072 |
| 56 | SIN3 COMPLEX | 4 | 13 | 0.0001025 | 0.001072 |
| 57 | CONTRACTILE FIBER | 14 | 211 | 0.0001078 | 0.001104 |
| 58 | CELL JUNCTION | 42 | 1151 | 0.0001655 | 0.001667 |
| 59 | SIN3 TYPE COMPLEX | 4 | 16 | 0.0002486 | 0.002461 |
| 60 | CELL CELL JUNCTION | 19 | 383 | 0.0003285 | 0.003197 |
| 61 | LATERAL PLASMA MEMBRANE | 6 | 50 | 0.0004973 | 0.004761 |
| 62 | SAGA TYPE COMPLEX | 5 | 34 | 0.0005697 | 0.005366 |
| 63 | CYTOPLASMIC SIDE OF MEMBRANE | 11 | 170 | 0.0007143 | 0.006621 |
| 64 | DNA DIRECTED RNA POLYMERASE II HOLOENZYME | 8 | 97 | 0.0007934 | 0.00724 |
| 65 | RNA POLYMERASE COMPLEX | 9 | 122 | 0.0008533 | 0.007667 |
| 66 | SITE OF POLARIZED GROWTH | 10 | 149 | 0.0009336 | 0.008261 |
| 67 | MCM COMPLEX | 3 | 11 | 0.001205 | 0.009912 |
| 68 | SET1C COMPASS COMPLEX | 3 | 11 | 0.001205 | 0.009912 |
| 69 | ARP2 3 PROTEIN COMPLEX | 3 | 11 | 0.001205 | 0.009912 |
| 70 | WNT SIGNALOSOME | 3 | 11 | 0.001205 | 0.009912 |
| 71 | ESCRT III COMPLEX | 3 | 11 | 0.001205 | 0.009912 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04110_Cell_cycle | 37 | 128 | 1.641e-32 | 2.904e-30 | |
| 2 | hsa04722_Neurotrophin_signaling_pathway | 21 | 127 | 1.172e-13 | 1.037e-11 | |
| 3 | hsa04114_Oocyte_meiosis | 18 | 114 | 1.526e-11 | 9.001e-10 | |
| 4 | hsa04914_Progesterone.mediated_oocyte_maturation | 15 | 87 | 2.129e-10 | 9.423e-09 | |
| 5 | hsa04115_p53_signaling_pathway | 11 | 69 | 1.304e-07 | 4.616e-06 | |
| 6 | hsa04916_Melanogenesis | 12 | 101 | 9.499e-07 | 2.802e-05 | |
| 7 | hsa04910_Insulin_signaling_pathway | 13 | 138 | 4.759e-06 | 0.0001203 | |
| 8 | hsa04360_Axon_guidance | 12 | 130 | 1.356e-05 | 3e-04 | |
| 9 | hsa04330_Notch_signaling_pathway | 7 | 47 | 4.146e-05 | 0.0008154 | |
| 10 | hsa04530_Tight_junction | 11 | 133 | 8.487e-05 | 0.001502 | |
| 11 | hsa04370_VEGF_signaling_pathway | 8 | 76 | 0.0001495 | 0.002406 | |
| 12 | hsa04912_GnRH_signaling_pathway | 9 | 101 | 0.0002118 | 0.003027 | |
| 13 | hsa03440_Homologous_recombination | 5 | 28 | 0.0002223 | 0.003027 | |
| 14 | hsa00310_Lysine_degradation | 6 | 44 | 0.0002446 | 0.003093 | |
| 15 | hsa04012_ErbB_signaling_pathway | 8 | 87 | 0.0003821 | 0.004509 | |
| 16 | hsa04520_Adherens_junction | 7 | 73 | 0.0006845 | 0.007572 | |
| 17 | hsa04662_B_cell_receptor_signaling_pathway | 7 | 75 | 0.0008056 | 0.007979 | |
| 18 | hsa04510_Focal_adhesion | 12 | 200 | 0.0008114 | 0.007979 | |
| 19 | hsa04310_Wnt_signaling_pathway | 10 | 151 | 0.001034 | 0.009256 | |
| 20 | hsa03060_Protein_export | 4 | 23 | 0.001081 | 0.009256 | |
| 21 | hsa04664_Fc_epsilon_RI_signaling_pathway | 7 | 79 | 0.001098 | 0.009256 | |
| 22 | hsa04660_T_cell_receptor_signaling_pathway | 8 | 108 | 0.001595 | 0.01283 | |
| 23 | hsa04120_Ubiquitin_mediated_proteolysis | 9 | 139 | 0.002125 | 0.01636 | |
| 24 | hsa04141_Protein_processing_in_endoplasmic_reticulum | 10 | 168 | 0.002293 | 0.01659 | |
| 25 | hsa04540_Gap_junction | 7 | 90 | 0.002343 | 0.01659 | |
| 26 | hsa04720_Long.term_potentiation | 6 | 70 | 0.002925 | 0.01931 | |
| 27 | hsa04144_Endocytosis | 11 | 203 | 0.002946 | 0.01931 | |
| 28 | hsa04010_MAPK_signaling_pathway | 13 | 268 | 0.003365 | 0.02127 | |
| 29 | hsa04150_mTOR_signaling_pathway | 5 | 52 | 0.003953 | 0.02413 | |
| 30 | hsa04612_Antigen_processing_and_presentation | 6 | 78 | 0.005008 | 0.02955 | |
| 31 | hsa04062_Chemokine_signaling_pathway | 10 | 189 | 0.0053 | 0.0301 | |
| 32 | hsa04340_Hedgehog_signaling_pathway | 5 | 56 | 0.005441 | 0.0301 | |
| 33 | hsa03030_DNA_replication | 4 | 36 | 0.005849 | 0.03137 | |
| 34 | hsa04810_Regulation_of_actin_cytoskeleton | 10 | 214 | 0.01217 | 0.06336 | |
| 35 | hsa04620_Toll.like_receptor_signaling_pathway | 6 | 102 | 0.01758 | 0.08891 | |
| 36 | hsa03008_Ribosome_biogenesis_in_eukaryotes | 5 | 81 | 0.0243 | 0.1195 | |
| 37 | hsa04621_NOD.like_receptor_signaling_pathway | 4 | 59 | 0.0315 | 0.1484 | |
| 38 | hsa04670_Leukocyte_transendothelial_migration | 6 | 117 | 0.03186 | 0.1484 | |
| 39 | hsa04210_Apoptosis | 5 | 89 | 0.0346 | 0.157 | |
| 40 | hsa00270_Cysteine_and_methionine_metabolism | 3 | 36 | 0.03595 | 0.1591 | |
| 41 | hsa03022_Basal_transcription_factors | 3 | 37 | 0.03855 | 0.1664 | |
| 42 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 5 | 95 | 0.04386 | 0.1848 | |
| 43 | hsa04920_Adipocytokine_signaling_pathway | 4 | 68 | 0.04904 | 0.2019 | |
| 44 | hsa00670_One_carbon_pool_by_folate | 2 | 18 | 0.05055 | 0.2033 | |
| 45 | hsa04730_Long.term_depression | 4 | 70 | 0.05354 | 0.2106 | |
| 46 | hsa04971_Gastric_acid_secretion | 4 | 74 | 0.06317 | 0.2431 | |
| 47 | hsa04270_Vascular_smooth_muscle_contraction | 5 | 116 | 0.08695 | 0.3219 | |
| 48 | hsa00983_Drug_metabolism_._other_enzymes | 3 | 52 | 0.08817 | 0.3219 | |
| 49 | hsa03013_RNA_transport | 6 | 152 | 0.08912 | 0.3219 | |
| 50 | hsa00230_Purine_metabolism | 6 | 162 | 0.1118 | 0.3959 | |
| 51 | hsa04744_Phototransduction | 2 | 29 | 0.1162 | 0.4033 | |
| 52 | hsa00020_Citrate_cycle_.TCA_cycle. | 2 | 30 | 0.1229 | 0.4184 | |
| 53 | hsa00010_Glycolysis_._Gluconeogenesis | 3 | 65 | 0.1447 | 0.4744 | |
| 54 | hsa03410_Base_excision_repair | 2 | 34 | 0.1506 | 0.4846 | |
| 55 | hsa00620_Pyruvate_metabolism | 2 | 40 | 0.194 | 0.6025 | |
| 56 | hsa03420_Nucleotide_excision_repair | 2 | 45 | 0.2312 | 0.7057 | |
| 57 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 4 | 136 | 0.2971 | 0.8764 | |
| 58 | hsa04630_Jak.STAT_signaling_pathway | 4 | 155 | 0.3842 | 1 | |
| 59 | hsa04145_Phagosome | 4 | 156 | 0.3888 | 1 | |
| 60 | hsa04622_RIG.I.like_receptor_signaling_pathway | 2 | 71 | 0.4229 | 1 | |
| 61 | hsa04070_Phosphatidylinositol_signaling_system | 2 | 78 | 0.4707 | 1 | |
| 62 | hsa04380_Osteoclast_differentiation | 3 | 128 | 0.4815 | 1 | |
| 63 | hsa04020_Calcium_signaling_pathway | 4 | 177 | 0.4832 | 1 | |
| 64 | hsa03015_mRNA_surveillance_pathway | 2 | 83 | 0.5034 | 1 | |
| 65 | hsa04350_TGF.beta_signaling_pathway | 2 | 85 | 0.5161 | 1 | |
| 66 | hsa04512_ECM.receptor_interaction | 2 | 85 | 0.5161 | 1 | |
| 67 | hsa04514_Cell_adhesion_molecules_.CAMs. | 3 | 136 | 0.522 | 1 | |
| 68 | hsa04640_Hematopoietic_cell_lineage | 2 | 88 | 0.5347 | 1 | |
| 69 | hsa04970_Salivary_secretion | 2 | 89 | 0.5408 | 1 | |
| 70 | hsa00240_Pyrimidine_metabolism | 2 | 99 | 0.5987 | 1 | |
| 71 | hsa03040_Spliceosome | 2 | 128 | 0.7352 | 1 | |
| 72 | hsa04740_Olfactory_transduction | 2 | 388 | 0.997 | 1 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | RP11-175K6.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p | 11 | KLF4 | Sponge network | -3.312 | 0 | -2.566 | 0 | 0.659 |
| 2 | RP11-1024P17.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 14 | KLF4 | Sponge network | -2.541 | 0 | -2.566 | 0 | 0.641 |
| 3 | MIR22HG | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-375;hsa-miR-92a-3p;hsa-miR-92b-3p | 16 | KLF4 | Sponge network | -1.551 | 0 | -2.566 | 0 | 0.632 |
| 4 | AC011526.1 | hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-142-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-32-5p;hsa-miR-375;hsa-miR-493-5p;hsa-miR-590-3p;hsa-miR-92a-3p | 11 | KLF4 | Sponge network | -1.879 | 0 | -2.566 | 0 | 0.63 |
| 5 | RP11-736K20.5 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-375;hsa-miR-508-3p;hsa-miR-590-3p | 16 | KLF4 | Sponge network | -3.119 | 0 | -2.566 | 0 | 0.623 |
| 6 | MAGI2-AS3 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-375;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 19 | KLF4 | Sponge network | -2.333 | 0 | -2.566 | 0 | 0.622 |
| 7 | MIR143HG | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 20 | KLF4 | Sponge network | -4.451 | 0 | -2.566 | 0 | 0.601 |
| 8 | ADAMTS9-AS2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-375;hsa-miR-493-5p;hsa-miR-590-3p;hsa-miR-92b-3p | 18 | KLF4 | Sponge network | -4.039 | 0 | -2.566 | 0 | 0.577 |
| 9 | RP11-389C8.2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-429;hsa-miR-590-3p | 15 | KLF4 | Sponge network | -1.579 | 0 | -2.566 | 0 | 0.563 |
| 10 | RP11-1101K5.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-508-3p | 13 | KLF4 | Sponge network | -8.619 | 0 | -2.566 | 0 | 0.54 |
| 11 | TRHDE-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-375;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p;hsa-miR-92b-3p | 18 | KLF4 | Sponge network | -7.569 | 0 | -2.566 | 0 | 0.539 |
| 12 | LIPE-AS1 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p | 12 | KLF4 | Sponge network | -0.907 | 0 | -2.566 | 0 | 0.537 |
| 13 | LINC00968 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-493-5p;hsa-miR-508-3p;hsa-miR-590-3p | 14 | KLF4 | Sponge network | -4.245 | 0 | -2.566 | 0 | 0.52 |
| 14 | RP11-166D19.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 18 | KLF4 | Sponge network | -1.912 | 0 | -2.566 | 0 | 0.516 |
| 15 | AL035610.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p | 17 | KLF4 | Sponge network | -6.728 | 0 | -2.566 | 0 | 0.513 |
| 16 | ALDH1L1-AS2 | hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 11 | KLF4 | Sponge network | -7.428 | 0 | -2.566 | 0 | 0.512 |
| 17 | LINC00924 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -3.023 | 0 | -2.566 | 0 | 0.497 |
| 18 | RP11-276H19.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 15 | KLF4 | Sponge network | -2.749 | 0 | -2.566 | 0 | 0.494 |
| 19 | RP5-1052I5.1 | hsa-miR-107;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-29b-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-493-5p;hsa-miR-590-3p;hsa-miR-92b-3p | 10 | KLF4 | Sponge network | -2.903 | 0 | -2.566 | 0 | 0.48 |
| 20 | EMX2OS | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-375;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 19 | KLF4 | Sponge network | -2.548 | 0 | -2.566 | 0 | 0.478 |
| 21 | MEG3 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-429 | 14 | KLF4 | Sponge network | -2.353 | 0 | -2.566 | 0 | 0.475 |
| 22 | CTD-2013N24.2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p | 15 | KLF4 | Sponge network | -1.842 | 0 | -2.566 | 0 | 0.473 |
| 23 | ADAMTS9-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-375;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 21 | KLF4 | Sponge network | -4.696 | 0 | -2.566 | 0 | 0.472 |
| 24 | RP11-384P7.7 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -4.533 | 0 | -2.566 | 0 | 0.464 |
| 25 | PGM5-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p | 10 | KLF4 | Sponge network | -7.694 | 0 | -2.566 | 0 | 0.464 |
| 26 | LINC00641 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -1.721 | 0 | -2.566 | 0 | 0.462 |
| 27 | ADIPOQ-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-590-3p | 14 | KLF4 | Sponge network | -8 | 0 | -2.566 | 0 | 0.46 |
| 28 | HOXB-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-508-3p | 11 | KLF4 | Sponge network | -2.063 | 0 | -2.566 | 0 | 0.452 |
| 29 | WDR86-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-429 | 10 | KLF4 | Sponge network | -3.263 | 0 | -2.566 | 0 | 0.45 |
| 30 | AC003991.3 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 17 | KLF4 | Sponge network | -2.021 | 0 | -2.566 | 0 | 0.44 |
| 31 | RP11-180N14.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-375 | 11 | KLF4 | Sponge network | -2.382 | 0 | -2.566 | 0 | 0.428 |
| 32 | DIO3OS | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p | 12 | KLF4 | Sponge network | -3.064 | 0 | -2.566 | 0 | 0.425 |
| 33 | FZD10-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 18 | KLF4 | Sponge network | -1.884 | 0 | -2.566 | 0 | 0.424 |
| 34 | RP13-270P17.2 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -1.378 | 0 | -2.566 | 0 | 0.421 |
| 35 | RP11-1008C21.2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-375;hsa-miR-92a-3p;hsa-miR-92b-3p | 17 | KLF4 | Sponge network | -1.055 | 0 | -2.566 | 0 | 0.42 |
| 36 | ACTA2-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-590-3p | 15 | KLF4 | Sponge network | -3.065 | 0 | -2.566 | 0 | 0.418 |
| 37 | AGAP11 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-34a-5p | 13 | KLF4 | Sponge network | -2.72 | 0 | -2.566 | 0 | 0.417 |
| 38 | MIR497HG | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375 | 14 | KLF4 | Sponge network | -2.527 | 0 | -2.566 | 0 | 0.415 |
| 39 | LINC00639 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-493-5p | 13 | KLF4 | Sponge network | -2.449 | 0 | -2.566 | 0 | 0.412 |
| 40 | AC144831.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p | 12 | KLF4 | Sponge network | -1.421 | 0 | -2.566 | 0 | 0.411 |
| 41 | WDFY3-AS2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p | 16 | KLF4 | Sponge network | -1.618 | 0 | -2.566 | 0 | 0.407 |
| 42 | LINC00702 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-30b-5p;hsa-miR-32-5p;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 18 | KLF4 | Sponge network | -1.308 | 0 | -2.566 | 0 | 0.403 |
| 43 | AC005682.5 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -3.076 | 0 | -2.566 | 0 | 0.4 |
| 44 | HOXA-AS2 | hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -2.242 | 0 | -2.566 | 0 | 0.399 |
| 45 | PDZRN3-AS1 | hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -2.387 | 0 | -2.566 | 0 | 0.398 |
| 46 | NR2F1-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 18 | KLF4 | Sponge network | -0.993 | 0 | -2.566 | 0 | 0.39 |
| 47 | RP11-57H14.4 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-493-5p;hsa-miR-590-3p | 14 | KLF4 | Sponge network | -1.436 | 0 | -2.566 | 0 | 0.385 |
| 48 | HOTAIRM1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 11 | KLF4 | Sponge network | -2.146 | 0 | -2.566 | 0 | 0.384 |
| 49 | CTD-3247F14.2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p | 12 | KLF4 | Sponge network | -2.833 | 0 | -2.566 | 0 | 0.381 |
| 50 | AC097724.3 | hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 10 | KLF4 | Sponge network | -2.411 | 0 | -2.566 | 0 | 0.377 |
| 51 | AC108142.1 | hsa-let-7a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -2.247 | 0 | -2.566 | 0 | 0.377 |
| 52 | TPTEP1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-375;hsa-miR-590-3p;hsa-miR-92b-3p | 16 | KLF4 | Sponge network | -2.33 | 0 | -2.566 | 0 | 0.377 |
| 53 | RP11-400K9.4 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-148b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-30b-5p | 10 | KLF4 | Sponge network | -1.058 | 0 | -2.566 | 0 | 0.374 |
| 54 | RP11-736K20.6 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -1.407 | 0 | -2.566 | 0 | 0.371 |
| 55 | RP11-276H19.2 | hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 10 | KLF4 | Sponge network | -3.719 | 0 | -2.566 | 0 | 0.37 |
| 56 | RP11-356J5.12 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-375;hsa-miR-493-5p;hsa-miR-590-3p;hsa-miR-92b-3p | 16 | KLF4 | Sponge network | -1.682 | 0 | -2.566 | 0 | 0.367 |
| 57 | LINC01088 | hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-375;hsa-miR-429;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -1.675 | 0 | -2.566 | 0 | 0.366 |
| 58 | RP11-2B6.2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 10 | KLF4 | Sponge network | -1.878 | 0 | -2.566 | 0 | 0.365 |
| 59 | RP1-193H18.2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-493-5p;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -1.975 | 0 | -2.566 | 0 | 0.358 |
| 60 | RP11-286B14.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 11 | KLF4 | Sponge network | -5.866 | 0 | -2.566 | 0 | 0.356 |
| 61 | RP11-434B12.1 | hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -1.324 | 0 | -2.566 | 0 | 0.352 |
| 62 | RP11-448G15.3 | hsa-let-7a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -1.586 | 0 | -2.566 | 0 | 0.35 |
| 63 | LL22NC03-86G7.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 17 | KLF4 | Sponge network | -1.159 | 0 | -2.566 | 0 | 0.349 |
| 64 | HLA-F-AS1 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-590-3p;hsa-miR-92b-3p | 11 | KLF4 | Sponge network | -1.168 | 0 | -2.566 | 0 | 0.348 |
| 65 | RP11-645C24.5 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-128-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-590-3p | 10 | KLF4 | Sponge network | -2.365 | 0 | -2.566 | 0 | 0.348 |
| 66 | FAM13A-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 13 | KLF4 | Sponge network | -1.451 | 0 | -2.566 | 0 | 0.348 |
| 67 | AF131215.2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-30b-5p;hsa-miR-429 | 14 | KLF4 | Sponge network | -2.008 | 0 | -2.566 | 0 | 0.348 |
| 68 | TBX5-AS1 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-92a-3p | 13 | KLF4 | Sponge network | -0.804 | 0 | -2.566 | 0 | 0.344 |
| 69 | GS1-124K5.11 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 11 | KLF4 | Sponge network | -0.939 | 0 | -2.566 | 0 | 0.341 |
| 70 | RP11-822E23.8 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-375 | 14 | KLF4 | Sponge network | -2.406 | 0 | -2.566 | 0 | 0.336 |
| 71 | AC004947.2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -3.692 | 0 | -2.566 | 0 | 0.336 |
| 72 | HCG11 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -1.369 | 0 | -2.566 | 0 | 0.336 |
| 73 | RP11-401P9.4 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p | 13 | KLF4 | Sponge network | -0.963 | 0 | -2.566 | 0 | 0.333 |
| 74 | RP11-978I15.10 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-429 | 14 | KLF4 | Sponge network | -3.591 | 0 | -2.566 | 0 | 0.332 |
| 75 | CTD-2647L4.4 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-429 | 11 | KLF4 | Sponge network | -1.068 | 0 | -2.566 | 0 | 0.331 |
| 76 | RP11-981G7.6 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-493-5p;hsa-miR-590-3p | 14 | KLF4 | Sponge network | -2.292 | 0 | -2.566 | 0 | 0.331 |
| 77 | LINC00987 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-590-3p;hsa-miR-92b-3p | 14 | KLF4 | Sponge network | -1.247 | 0 | -2.566 | 0 | 0.327 |
| 78 | AC007743.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-375;hsa-miR-429;hsa-miR-92b-3p | 15 | KLF4 | Sponge network | -2.327 | 0 | -2.566 | 0 | 0.325 |
| 79 | RP11-244O19.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-590-3p;hsa-miR-92b-3p | 16 | KLF4 | Sponge network | -1.068 | 0 | -2.566 | 0 | 0.325 |
| 80 | AF131215.9 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-590-3p;hsa-miR-92b-3p | 14 | KLF4 | Sponge network | -1.776 | 0 | -2.566 | 0 | 0.325 |
| 81 | RP11-116O18.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-493-5p;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -3.1 | 0 | -2.566 | 0 | 0.325 |
| 82 | LINC00667 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p | 15 | KLF4 | Sponge network | -1.129 | 0 | -2.566 | 0 | 0.322 |
| 83 | SH3BP5-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p | 10 | KLF4 | Sponge network | -1.175 | 0 | -2.566 | 0 | 0.321 |
| 84 | CTB-107G13.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 11 | KLF4 | Sponge network | -3.353 | 0 | -2.566 | 0 | 0.311 |
| 85 | RP11-545I5.3 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 16 | KLF4 | Sponge network | -1.007 | 0 | -2.566 | 0 | 0.311 |
| 86 | RP4-798P15.3 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-30b-5p;hsa-miR-34a-5p;hsa-miR-375;hsa-miR-493-5p;hsa-miR-590-3p | 17 | KLF4 | Sponge network | -1.167 | 0 | -2.566 | 0 | 0.309 |
| 87 | RP11-244H3.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 11 | KLF4 | Sponge network | -0.877 | 0 | -2.566 | 0 | 0.308 |
| 88 | RP11-5C23.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 11 | KLF4 | Sponge network | -0.816 | 0 | -2.566 | 0 | 0.307 |
| 89 | SNHG18 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -0.741 | 0 | -2.566 | 0 | 0.305 |
| 90 | RP11-573G6.6 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-429 | 10 | KLF4 | Sponge network | -1.242 | 0 | -2.566 | 0 | 0.301 |
| 91 | CTD-3092A11.2 | hsa-let-7a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -1.102 | 0 | -2.566 | 0 | 0.301 |
| 92 | RP11-890B15.3 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -0.935 | 0 | -2.566 | 0 | 0.3 |
| 93 | AC007246.3 | hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 13 | KLF4 | Sponge network | -0.496 | 0 | -2.566 | 0 | 0.291 |
| 94 | TPT1-AS1 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-375;hsa-miR-493-5p;hsa-miR-590-3p;hsa-miR-92b-3p | 14 | KLF4 | Sponge network | -1.187 | 0 | -2.566 | 0 | 0.29 |
| 95 | APCDD1L-AS1 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-493-5p | 13 | KLF4 | Sponge network | -3.451 | 0 | -2.566 | 0 | 0.288 |
| 96 | RP11-403B2.7 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -1.946 | 0 | -2.566 | 0 | 0.287 |
| 97 | LINC01085 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-429;hsa-miR-590-3p | 15 | KLF4 | Sponge network | -3.703 | 0 | -2.566 | 0 | 0.281 |
| 98 | CKMT2-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 11 | KLF4 | Sponge network | -0.724 | 0 | -2.566 | 0 | 0.28 |
| 99 | RP11-274B21.10 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 10 | KLF4 | Sponge network | -1.456 | 0 | -2.566 | 0 | 0.278 |
| 100 | RP11-67L2.2 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-92b-3p | 17 | KLF4 | Sponge network | -0.677 | 0 | -2.566 | 0 | 0.278 |
| 101 | RP11-411K7.1 | hsa-let-7a-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-92b-3p | 11 | KLF4 | Sponge network | -2.478 | 0 | -2.566 | 0 | 0.274 |
| 102 | RP11-631N16.2 | hsa-let-7a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -1.389 | 0 | -2.566 | 0 | 0.274 |
| 103 | SOCS2-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -1.697 | 0 | -2.566 | 0 | 0.271 |
| 104 | RP11-398E10.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -3.93 | 0 | -2.566 | 0 | 0.267 |
| 105 | RP11-6O2.3 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p | 10 | KLF4 | Sponge network | -3.108 | 0 | -2.566 | 0 | 0.265 |
| 106 | RP11-293M10.6 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-429 | 12 | KLF4 | Sponge network | -0.601 | 0.00056 | -2.566 | 0 | 0.264 |
| 107 | OR2A1-AS1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-493-5p;hsa-miR-590-3p | 12 | KLF4 | Sponge network | -1 | 0 | -2.566 | 0 | 0.262 |
| 108 | LINC00648 | hsa-let-7a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p | 14 | KLF4 | Sponge network | -5.143 | 0 | -2.566 | 0 | 0.261 |
| 109 | RP11-378A13.1 | hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 11 | KLF4 | Sponge network | -0.486 | 0 | -2.566 | 0 | 0.26 |
| 110 | DNM3OS | hsa-let-7a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-590-3p | 13 | KLF4 | Sponge network | -0.276 | 0.07533 | -2.566 | 0 | 0.258 |
| 111 | CTD-2619J13.17 | hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-590-3p | 10 | KLF4 | Sponge network | -0.985 | 0 | -2.566 | 0 | 0.258 |
| 112 | AC083843.1 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92b-3p | 12 | KLF4 | Sponge network | -1.426 | 0 | -2.566 | 0 | 0.254 |
| 113 | ZNF833P | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-128-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-32-5p;hsa-miR-34a-5p;hsa-miR-92b-3p | 10 | KLF4 | Sponge network | -0.772 | 0 | -2.566 | 0 | 0.253 |
| 114 | RP11-299J3.8 | hsa-let-7f-1-3p;hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200c-3p;hsa-miR-301a-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92b-3p | 13 | KLF4 | Sponge network | -1.007 | 0 | -2.566 | 0 | 0.251 |
| 115 | PWAR6 | hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-25-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-34a-5p;hsa-miR-429;hsa-miR-493-5p;hsa-miR-590-3p | 16 | KLF4 | Sponge network | -1.696 | 0 | -2.566 | 0 | 0.251 |