This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-10a-5p | ACAA1 | 0.79 | 0.00059 | -0.46 | 0.00059 | miRNAWalker2 validate | -0.13 | 0 | NA | |
| 2 | hsa-miR-590-3p | ACADL | 0.84 | 0.00129 | -4.45 | 0 | miRanda | -0.42 | 0.00074 | NA | |
| 3 | hsa-miR-148a-3p | ACOX1 | 2.31 | 0 | -0.01 | 0.90133 | mirMAP | -0.1 | 0 | NA | |
| 4 | hsa-miR-15a-5p | ACOX1 | 1.63 | 0 | -0.01 | 0.90133 | miRNATAP | -0.12 | 8.0E-5 | NA | |
| 5 | hsa-miR-199a-3p | ACOX1 | 1.68 | 0 | -0.01 | 0.90133 | PITA | -0.14 | 0 | NA | |
| 6 | hsa-miR-199b-3p | ACOX1 | 1.68 | 0 | -0.01 | 0.90133 | PITA | -0.14 | 0 | NA | |
| 7 | hsa-miR-497-5p | ACOX1 | -0.05 | 0.78621 | -0.01 | 0.90133 | miRNATAP | -0.11 | 5.0E-5 | NA | |
| 8 | hsa-miR-708-3p | ACOX1 | 3.44 | 0 | -0.01 | 0.90133 | mirMAP | -0.11 | 0 | NA | |
| 9 | hsa-miR-93-5p | ACOX1 | 1.51 | 0 | -0.01 | 0.90133 | mirMAP | -0.14 | 0 | NA | |
| 10 | hsa-miR-576-5p | ACOX2 | 1.03 | 0 | -0.57 | 0.05834 | MirTarget | -0.35 | 0 | NA | |
| 11 | hsa-miR-590-3p | ACOX2 | 0.84 | 0.00129 | -0.57 | 0.05834 | miRanda | -0.3 | 0 | NA | |
| 12 | hsa-miR-130b-3p | ACSL1 | 1.83 | 0 | -1.01 | 0 | MirTarget; miRNATAP | -0.12 | 0.00104 | NA | |
| 13 | hsa-miR-195-3p | ACSL3 | -1.33 | 0 | 0.2 | 0.18226 | mirMAP | -0.15 | 0 | NA | |
| 14 | hsa-miR-221-3p | ACSL3 | -0.1 | 0.65445 | 0.2 | 0.18226 | miRNAWalker2 validate | -0.14 | 1.0E-5 | NA | |
| 15 | hsa-miR-30b-5p | ACSL3 | 0.36 | 0.13803 | 0.2 | 0.18226 | mirMAP | -0.1 | 0.00043 | NA | |
| 16 | hsa-miR-30c-5p | ACSL3 | -0.33 | 0.1236 | 0.2 | 0.18226 | mirMAP | -0.19 | 0 | NA | |
| 17 | hsa-miR-30e-3p | ACSL3 | -0.1 | 0.52624 | 0.2 | 0.18226 | mirMAP | -0.12 | 0.00592 | NA | |
| 18 | hsa-miR-339-5p | ACSL3 | 0.54 | 0.04881 | 0.2 | 0.18226 | miRanda | -0.11 | 3.0E-5 | NA | |
| 19 | hsa-miR-342-3p | ACSL3 | -0.13 | 0.56103 | 0.2 | 0.18226 | miRanda | -0.18 | 0 | NA | |
| 20 | hsa-miR-106a-5p | ACSL4 | 1.39 | 6.0E-5 | -1.28 | 0 | MirTarget; miRNATAP | -0.14 | 1.0E-5 | NA | |
| 21 | hsa-miR-106b-5p | ACSL4 | 1.47 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.2 | 6.0E-5 | NA | |
| 22 | hsa-miR-130a-3p | ACSL4 | 0.88 | 0.00016 | -1.28 | 0 | MirTarget; miRNATAP | -0.14 | 0.00174 | NA | |
| 23 | hsa-miR-130b-3p | ACSL4 | 1.83 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.22 | 0 | NA | |
| 24 | hsa-miR-130b-5p | ACSL4 | 1.54 | 0 | -1.28 | 0 | MirTarget | -0.2 | 0 | NA | |
| 25 | hsa-miR-148a-3p | ACSL4 | 2.31 | 0 | -1.28 | 0 | miRNATAP | -0.12 | 0.00276 | NA | |
| 26 | hsa-miR-15a-5p | ACSL4 | 1.63 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.31 | 0 | NA | |
| 27 | hsa-miR-16-1-3p | ACSL4 | 1.5 | 0 | -1.28 | 0 | MirTarget | -0.22 | 2.0E-5 | NA | |
| 28 | hsa-miR-17-5p | ACSL4 | 2.07 | 0 | -1.28 | 0 | MirTarget; TargetScan; miRNATAP | -0.24 | 0 | NA | |
| 29 | hsa-miR-181b-5p | ACSL4 | 0.67 | 0.00024 | -1.28 | 0 | mirMAP | -0.21 | 0.00017 | NA | |
| 30 | hsa-miR-181c-5p | ACSL4 | 0.53 | 0.01259 | -1.28 | 0 | mirMAP | -0.13 | 0.00924 | NA | |
| 31 | hsa-miR-186-5p | ACSL4 | 0.85 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.24 | 0.00092 | NA | |
| 32 | hsa-miR-19a-3p | ACSL4 | 2.12 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.23 | 0 | NA | |
| 33 | hsa-miR-19b-3p | ACSL4 | 2.11 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.26 | 0 | NA | |
| 34 | hsa-miR-20a-5p | ACSL4 | 2.65 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.24 | 0 | NA | |
| 35 | hsa-miR-26b-3p | ACSL4 | 1.37 | 0 | -1.28 | 0 | MirTarget | -0.15 | 0.00024 | NA | |
| 36 | hsa-miR-26b-5p | ACSL4 | 0.72 | 5.0E-5 | -1.28 | 0 | mirMAP | -0.19 | 0.00082 | NA | |
| 37 | hsa-miR-301a-3p | ACSL4 | 2.7 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.23 | 0 | NA | |
| 38 | hsa-miR-335-3p | ACSL4 | 1.51 | 0 | -1.28 | 0 | MirTarget; mirMAP | -0.27 | 0 | NA | |
| 39 | hsa-miR-34a-5p | ACSL4 | 1.41 | 0 | -1.28 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.3 | 0 | NA | |
| 40 | hsa-miR-429 | ACSL4 | 2.38 | 0 | -1.28 | 0 | miRanda; miRNATAP | -0.24 | 0 | NA | |
| 41 | hsa-miR-450b-5p | ACSL4 | 1.69 | 0 | -1.28 | 0 | MirTarget; PITA; mirMAP | -0.11 | 0.00175 | NA | |
| 42 | hsa-miR-454-3p | ACSL4 | 1.49 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.29 | 0 | NA | |
| 43 | hsa-miR-664a-3p | ACSL4 | 0.44 | 0.02142 | -1.28 | 0 | mirMAP | -0.3 | 0 | NA | |
| 44 | hsa-miR-7-1-3p | ACSL4 | 2.61 | 0 | -1.28 | 0 | mirMAP | -0.18 | 9.0E-5 | NA | |
| 45 | hsa-miR-93-5p | ACSL4 | 1.51 | 0 | -1.28 | 0 | MirTarget; miRNATAP | -0.27 | 0 | NA | |
| 46 | hsa-miR-128-3p | ACSL5 | 1.04 | 0 | 0.21 | 0.49172 | miRNAWalker2 validate | -0.66 | 0 | NA | |
| 47 | hsa-miR-590-3p | ACSL5 | 0.84 | 0.00129 | 0.21 | 0.49172 | miRanda; mirMAP | -0.18 | 0.00436 | NA | |
| 48 | hsa-miR-106a-5p | ACSL6 | 1.39 | 6.0E-5 | 0.35 | 0.34401 | mirMAP | -0.16 | 0.00381 | NA | |
| 49 | hsa-miR-16-2-3p | ACSL6 | 0.5 | 0.02636 | 0.35 | 0.34401 | mirMAP | -0.22 | 0.00712 | NA | |
| 50 | hsa-miR-17-5p | ACSL6 | 2.07 | 0 | 0.35 | 0.34401 | mirMAP | -0.3 | 2.0E-5 | NA | |
| 51 | hsa-miR-20a-5p | ACSL6 | 2.65 | 0 | 0.35 | 0.34401 | mirMAP | -0.23 | 0.00057 | NA | |
| 52 | hsa-miR-582-5p | ACSL6 | 1.08 | 0.00149 | 0.35 | 0.34401 | mirMAP; miRNATAP | -0.17 | 0.00073 | NA | |
| 53 | hsa-miR-193a-3p | AQP7 | 0.55 | 0.0319 | -0.68 | 0.21739 | miRanda | -0.55 | 0 | NA | |
| 54 | hsa-miR-28-5p | CD36 | 1.2 | 0 | -3.79 | 0 | miRanda | -0.6 | 0 | NA | |
| 55 | hsa-miR-590-3p | CD36 | 0.84 | 0.00129 | -3.79 | 0 | miRanda | -0.35 | 3.0E-5 | NA | |
| 56 | hsa-miR-629-3p | CD36 | 1.32 | 0.00011 | -3.79 | 0 | MirTarget | -0.25 | 1.0E-5 | NA | |
| 57 | hsa-miR-96-5p | CD36 | 3.04 | 0 | -3.79 | 0 | MirTarget | -0.57 | 0 | NA | |
| 58 | hsa-miR-155-5p | CPT1A | 0.81 | 0.00061 | -0.73 | 1.0E-5 | miRNAWalker2 validate | -0.11 | 0.00071 | NA | |
| 59 | hsa-miR-17-5p | CPT1A | 2.07 | 0 | -0.73 | 1.0E-5 | mirMAP | -0.11 | 0.00045 | NA | |
| 60 | hsa-miR-199a-5p | CPT1A | 1.31 | 0 | -0.73 | 1.0E-5 | miRanda | -0.13 | 6.0E-5 | NA | |
| 61 | hsa-miR-20a-5p | CPT1A | 2.65 | 0 | -0.73 | 1.0E-5 | mirMAP | -0.13 | 0 | NA | |
| 62 | hsa-miR-3613-5p | CPT1A | 0.02 | 0.93473 | -0.73 | 1.0E-5 | mirMAP | -0.12 | 6.0E-5 | NA | |
| 63 | hsa-miR-455-5p | CPT1A | 1.37 | 0 | -0.73 | 1.0E-5 | miRanda | -0.12 | 0.00027 | NA | |
| 64 | hsa-miR-93-5p | CPT1A | 1.51 | 0 | -0.73 | 1.0E-5 | mirMAP | -0.11 | 0.00184 | NA | |
| 65 | hsa-miR-139-3p | CPT1B | -4 | 0 | 1.98 | 0 | PITA | -0.31 | 0 | NA | |
| 66 | hsa-miR-330-3p | CPT1B | -0.72 | 0.0008 | 1.98 | 0 | miRNATAP | -0.43 | 0 | NA | |
| 67 | hsa-miR-664a-3p | EHHADH | 0.44 | 0.02142 | 0.01 | 0.9632 | mirMAP | -0.19 | 1.0E-5 | NA | |
| 68 | hsa-miR-589-3p | FABP4 | 1.34 | 2.0E-5 | -6.28 | 0 | MirTarget | -0.32 | 0.00021 | NA | |
| 69 | hsa-miR-590-3p | FABP4 | 0.84 | 0.00129 | -6.28 | 0 | miRanda | -0.54 | 1.0E-5 | NA | |
| 70 | hsa-miR-429 | FABP5 | 2.38 | 0 | -2.4 | 0 | miRanda | -0.24 | 0 | NA | |
| 71 | hsa-let-7b-5p | FADS2 | -1.62 | 0 | -0.21 | 0.49738 | miRNAWalker2 validate | -0.24 | 0.00036 | NA | |
| 72 | hsa-miR-125b-5p | FADS2 | -0.55 | 0.01072 | -0.21 | 0.49738 | mirMAP | -0.25 | 0.00015 | NA | |
| 73 | hsa-miR-224-3p | FADS2 | 0.92 | 0.01001 | -0.21 | 0.49738 | mirMAP | -0.14 | 0.00053 | NA | |
| 74 | hsa-miR-26b-5p | FADS2 | 0.72 | 5.0E-5 | -0.21 | 0.49738 | miRNAWalker2 validate | -0.23 | 0.00394 | NA | |
| 75 | hsa-miR-335-5p | FADS2 | -0.47 | 0.0677 | -0.21 | 0.49738 | miRNAWalker2 validate | -0.17 | 0.00633 | NA | |
| 76 | hsa-miR-450b-5p | GK | 1.69 | 0 | -0.14 | 0.55876 | MirTarget | -0.17 | 1.0E-5 | NA | |
| 77 | hsa-miR-542-3p | GK | 1.62 | 0 | -0.14 | 0.55876 | miRanda | -0.19 | 2.0E-5 | NA | |
| 78 | hsa-miR-590-5p | HMGCS2 | 2.07 | 0 | -3.01 | 0.00014 | miRanda | -0.48 | 0.00259 | NA | |
| 79 | hsa-miR-335-3p | ILK | 1.51 | 0 | -0.61 | 0 | MirTarget | -0.16 | 0 | NA | |
| 80 | hsa-miR-155-5p | LPL | 0.81 | 0.00061 | -3.16 | 0 | miRNAWalker2 validate | -0.35 | 8.0E-5 | NA | |
| 81 | hsa-miR-29b-3p | LPL | 3.11 | 0 | -3.16 | 0 | miRNATAP | -0.27 | 0.00022 | NA | |
| 82 | hsa-miR-590-3p | LPL | 0.84 | 0.00129 | -3.16 | 0 | miRanda; mirMAP; miRNATAP | -0.52 | 0 | NA | |
| 83 | hsa-miR-146b-5p | ME1 | 1.09 | 1.0E-5 | 0.15 | 0.58684 | miRanda | -0.15 | 0.00474 | NA | |
| 84 | hsa-miR-3065-5p | ME1 | 0.65 | 0.09995 | 0.15 | 0.58684 | miRNATAP | -0.11 | 0.00128 | NA | |
| 85 | hsa-miR-30c-5p | ME1 | -0.33 | 0.1236 | 0.15 | 0.58684 | miRNATAP | -0.19 | 0.00164 | NA | |
| 86 | hsa-miR-34a-5p | ME1 | 1.41 | 0 | 0.15 | 0.58684 | miRNAWalker2 validate | -0.25 | 3.0E-5 | NA | |
| 87 | hsa-miR-326 | MMP1 | -0.99 | 0.00335 | 4.3 | 0 | miRanda | -0.27 | 0.00179 | NA | |
| 88 | hsa-miR-129-5p | OLR1 | -0.41 | 0.34149 | -2.94 | 0 | miRanda | -0.1 | 0.00791 | NA | |
| 89 | hsa-miR-21-5p | OLR1 | 4.38 | 0 | -2.94 | 0 | miRNAWalker2 validate | -0.29 | 0 | NA | |
| 90 | hsa-miR-320b | OLR1 | 0.23 | 0.37882 | -2.94 | 0 | miRanda | -0.32 | 0 | NA | |
| 91 | hsa-miR-335-3p | OLR1 | 1.51 | 0 | -2.94 | 0 | mirMAP | -0.39 | 0 | NA | |
| 92 | hsa-miR-590-3p | OLR1 | 0.84 | 0.00129 | -2.94 | 0 | miRanda; mirMAP | -0.25 | 0.00125 | NA | |
| 93 | hsa-miR-590-5p | OLR1 | 2.07 | 0 | -2.94 | 0 | miRanda | -0.37 | 0 | NA | |
| 94 | hsa-miR-96-5p | OLR1 | 3.04 | 0 | -2.94 | 0 | MirTarget | -0.46 | 0 | NA | |
| 95 | hsa-miR-24-3p | PCK2 | -0.04 | 0.78587 | -0.33 | 0.0252 | miRNAWalker2 validate | -0.16 | 0.00256 | NA | |
| 96 | hsa-miR-106b-5p | PDPK1 | 1.47 | 0 | -0.26 | 0.03408 | mirMAP | -0.13 | 0 | NA | |
| 97 | hsa-miR-128-3p | PDPK1 | 1.04 | 0 | -0.26 | 0.03408 | miRNAWalker2 validate | -0.16 | 0 | NA | |
| 98 | hsa-miR-16-5p | PDPK1 | 0.75 | 0 | -0.26 | 0.03408 | mirMAP | -0.12 | 0.00072 | NA | |
| 99 | hsa-miR-17-5p | PDPK1 | 2.07 | 0 | -0.26 | 0.03408 | miRNAWalker2 validate; mirMAP | -0.11 | 1.0E-5 | NA | |
| 100 | hsa-miR-185-5p | PDPK1 | 1.14 | 0 | -0.26 | 0.03408 | miRNAWalker2 validate | -0.11 | 0.0017 | NA | |
| 101 | hsa-miR-186-5p | PDPK1 | 0.85 | 0 | -0.26 | 0.03408 | mirMAP | -0.11 | 0.00858 | NA | |
| 102 | hsa-miR-22-5p | PDPK1 | 1.71 | 0 | -0.26 | 0.03408 | miRNATAP | -0.12 | 0 | NA | |
| 103 | hsa-miR-421 | PDPK1 | 0.17 | 0.53528 | -0.26 | 0.03408 | mirMAP | -0.13 | 0 | NA | |
| 104 | hsa-miR-424-5p | PDPK1 | 1.26 | 1.0E-5 | -0.26 | 0.03408 | mirMAP | -0.11 | 0 | NA | |
| 105 | hsa-miR-590-3p | PDPK1 | 0.84 | 0.00129 | -0.26 | 0.03408 | miRanda; miRNATAP | -0.15 | 0 | NA | |
| 106 | hsa-miR-590-5p | PDPK1 | 2.07 | 0 | -0.26 | 0.03408 | miRanda | -0.14 | 0 | NA | |
| 107 | hsa-miR-18a-5p | PPARA | 1.37 | 1.0E-5 | 0.11 | 0.39154 | mirMAP | -0.11 | 0 | NA | |
| 108 | hsa-miR-342-3p | PPARA | -0.13 | 0.56103 | 0.11 | 0.39154 | miRanda | -0.13 | 0 | NA | |
| 109 | hsa-miR-130b-3p | PPARG | 1.83 | 0 | -2.13 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.37 | 0 | NA | |
| 110 | hsa-miR-20a-5p | PPARG | 2.65 | 0 | -2.13 | 0 | miRTarBase | -0.27 | 2.0E-5 | NA | |
| 111 | hsa-miR-27b-3p | PPARG | 0.24 | 0.12264 | -2.13 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.39 | 0.00039 | NA | |
| 112 | hsa-miR-301a-3p | PPARG | 2.7 | 0 | -2.13 | 0 | MirTarget; miRNATAP | -0.23 | 7.0E-5 | NA | |
| 113 | hsa-miR-324-5p | PPARG | 1.07 | 5.0E-5 | -2.13 | 0 | miRanda | -0.28 | 1.0E-5 | NA | |
| 114 | hsa-miR-429 | PPARG | 2.38 | 0 | -2.13 | 0 | miRNATAP | -0.29 | 0 | NA | |
| 115 | hsa-miR-454-3p | PPARG | 1.49 | 0 | -2.13 | 0 | MirTarget; miRNATAP | -0.27 | 0.00035 | NA | |
| 116 | hsa-let-7a-2-3p | RXRG | 1.34 | 0.00037 | -4.18 | 0 | MirTarget | -0.28 | 7.0E-5 | NA | |
| 117 | hsa-let-7g-3p | RXRG | 2.38 | 0 | -4.18 | 0 | MirTarget; miRNATAP | -0.44 | 0 | NA | |
| 118 | hsa-let-7i-5p | SCD | 0.61 | 0 | -1.03 | 5.0E-5 | MirTarget; miRNATAP | -0.28 | 0.00359 | NA | |
| 119 | hsa-miR-10a-5p | SCD | 0.79 | 0.00059 | -1.03 | 5.0E-5 | miRNAWalker2 validate | -0.18 | 0.00055 | NA | |
| 120 | hsa-miR-140-5p | SCD | 0.67 | 0.00034 | -1.03 | 5.0E-5 | MirTarget; PITA | -0.19 | 0.00277 | NA | |
| 121 | hsa-miR-142-3p | SCD | 3.98 | 0 | -1.03 | 5.0E-5 | miRanda | -0.12 | 0.00043 | NA | |
| 122 | hsa-miR-145-3p | SCD | -0.69 | 0.00068 | -1.03 | 5.0E-5 | mirMAP | -0.15 | 0.00991 | NA | |
| 123 | hsa-miR-146a-5p | SCD | 0.68 | 0.01053 | -1.03 | 5.0E-5 | mirMAP | -0.19 | 2.0E-5 | 24561623 | miR-146a targets Numb to stabilize β-catenin which forms a feedback circuit to maintain Wnt activity and directs SCD |
| 124 | hsa-miR-181a-5p | SCD | -0.38 | 0.05621 | -1.03 | 5.0E-5 | miRNAWalker2 validate; MirTarget | -0.29 | 0 | NA | |
| 125 | hsa-miR-181b-5p | SCD | 0.67 | 0.00024 | -1.03 | 5.0E-5 | MirTarget | -0.3 | 0 | NA | |
| 126 | hsa-miR-181c-5p | SCD | 0.53 | 0.01259 | -1.03 | 5.0E-5 | MirTarget | -0.23 | 5.0E-5 | NA | |
| 127 | hsa-miR-181d-5p | SCD | 1.19 | 0 | -1.03 | 5.0E-5 | MirTarget | -0.2 | 0.00028 | NA | |
| 128 | hsa-miR-199a-3p | SCD | 1.68 | 0 | -1.03 | 5.0E-5 | MirTarget; PITA; miRNATAP | -0.28 | 0 | NA | |
| 129 | hsa-miR-199b-3p | SCD | 1.68 | 0 | -1.03 | 5.0E-5 | MirTarget; PITA; miRNATAP | -0.28 | 0 | NA | |
| 130 | hsa-miR-200b-3p | SCD | 1.55 | 0 | -1.03 | 5.0E-5 | MirTarget; TargetScan | -0.16 | 0.00033 | NA | |
| 131 | hsa-miR-34c-3p | SCD | -1.35 | 0.00619 | -1.03 | 5.0E-5 | mirMAP | -0.13 | 0 | NA | |
| 132 | hsa-miR-429 | SCD | 2.38 | 0 | -1.03 | 5.0E-5 | MirTarget; PITA; miRanda; miRNATAP | -0.16 | 4.0E-5 | NA | |
| 133 | hsa-miR-452-5p | SCD | 0.64 | 0.04582 | -1.03 | 5.0E-5 | mirMAP | -0.1 | 0.0068 | NA | |
| 134 | hsa-miR-664a-3p | SCD | 0.44 | 0.02142 | -1.03 | 5.0E-5 | mirMAP | -0.19 | 0.00158 | NA | |
| 135 | hsa-miR-664a-5p | SCD | -0.09 | 0.66227 | -1.03 | 5.0E-5 | MirTarget | -0.22 | 0.0001 | NA | |
| 136 | hsa-miR-141-3p | SCD5 | 3.37 | 0 | -2.12 | 0 | TargetScan | -0.34 | 0 | NA | |
| 137 | hsa-miR-16-2-3p | SCD5 | 0.5 | 0.02636 | -2.12 | 0 | mirMAP | -0.2 | 0.00789 | NA | |
| 138 | hsa-miR-17-5p | SCD5 | 2.07 | 0 | -2.12 | 0 | TargetScan | -0.3 | 0 | NA | |
| 139 | hsa-miR-324-5p | SCD5 | 1.07 | 5.0E-5 | -2.12 | 0 | miRNAWalker2 validate | -0.19 | 0.00133 | NA | |
| 140 | hsa-miR-590-5p | SCD5 | 2.07 | 0 | -2.12 | 0 | miRanda | -0.2 | 0.00297 | NA | |
| 141 | hsa-miR-629-3p | SCD5 | 1.32 | 0.00011 | -2.12 | 0 | MirTarget | -0.14 | 0.00358 | NA | |
| 142 | hsa-miR-193a-3p | SCP2 | 0.55 | 0.0319 | -0.17 | 0.16051 | MirTarget; miRanda | -0.1 | 1.0E-5 | NA | |
| 143 | hsa-miR-590-5p | SCP2 | 2.07 | 0 | -0.17 | 0.16051 | mirMAP | -0.13 | 0 | NA | |
| 144 | hsa-miR-185-3p | SLC27A1 | -0.38 | 0.09365 | -0.39 | 0.03469 | MirTarget | -0.21 | 0 | NA | |
| 145 | hsa-miR-18a-5p | SLC27A1 | 1.37 | 1.0E-5 | -0.39 | 0.03469 | mirMAP | -0.17 | 0 | NA | |
| 146 | hsa-miR-421 | SLC27A1 | 0.17 | 0.53528 | -0.39 | 0.03469 | miRanda; mirMAP | -0.31 | 0 | NA | |
| 147 | hsa-miR-940 | SLC27A1 | 0.01 | 0.97493 | -0.39 | 0.03469 | MirTarget | -0.11 | 0.0001 | NA | |
| 148 | hsa-let-7b-5p | SLC27A2 | -1.62 | 0 | 1.08 | 0.00231 | miRNAWalker2 validate | -0.31 | 6.0E-5 | NA | |
| 149 | hsa-let-7g-3p | SORBS1 | 2.38 | 0 | -1.8 | 0 | miRNATAP | -0.27 | 0 | NA | |
| 150 | hsa-miR-142-5p | SORBS1 | 1.3 | 0 | -1.8 | 0 | PITA; miRNATAP | -0.17 | 6.0E-5 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | FATTY ACID METABOLIC PROCESS | 21 | 296 | 8.396e-30 | 3.907e-26 |
| 2 | CELLULAR LIPID METABOLIC PROCESS | 25 | 913 | 8.291e-26 | 1.929e-22 |
| 3 | MONOCARBOXYLIC ACID METABOLIC PROCESS | 21 | 503 | 6.698e-25 | 8.731e-22 |
| 4 | LIPID METABOLIC PROCESS | 26 | 1158 | 7.505e-25 | 8.731e-22 |
| 5 | LONG CHAIN FATTY ACID METABOLIC PROCESS | 13 | 88 | 1.954e-22 | 1.818e-19 |
| 6 | ORGANIC ACID METABOLIC PROCESS | 22 | 953 | 1.314e-20 | 1.019e-17 |
| 7 | SMALL MOLECULE METABOLIC PROCESS | 26 | 1767 | 3.57e-20 | 2.373e-17 |
| 8 | FATTY ACID TRANSPORT | 10 | 56 | 3.044e-18 | 1.771e-15 |
| 9 | CELLULAR LIPID CATABOLIC PROCESS | 12 | 151 | 2.357e-17 | 1.218e-14 |
| 10 | LIPID OXIDATION | 10 | 70 | 3.336e-17 | 1.552e-14 |
| 11 | FATTY ACID BETA OXIDATION | 9 | 51 | 1.932e-16 | 8.173e-14 |
| 12 | LONG CHAIN FATTY ACID TRANSPORT | 8 | 42 | 5.399e-15 | 2.094e-12 |
| 13 | FATTY ACID CATABOLIC PROCESS | 9 | 73 | 6.003e-15 | 2.149e-12 |
| 14 | LIPID CATABOLIC PROCESS | 12 | 247 | 9.275e-15 | 3.083e-12 |
| 15 | MONOCARBOXYLIC ACID TRANSPORT | 10 | 124 | 1.287e-14 | 3.992e-12 |
| 16 | MONOCARBOXYLIC ACID CATABOLIC PROCESS | 9 | 96 | 7.795e-14 | 2.267e-11 |
| 17 | LIPID BIOSYNTHETIC PROCESS | 14 | 539 | 1.987e-13 | 5.439e-11 |
| 18 | LIPID LOCALIZATION | 11 | 264 | 7.725e-13 | 1.997e-10 |
| 19 | LIPID MODIFICATION | 10 | 210 | 2.633e-12 | 6.447e-10 |
| 20 | FATTY ACYL COA METABOLIC PROCESS | 7 | 51 | 3.602e-12 | 8.381e-10 |
| 21 | ACYL COA BIOSYNTHETIC PROCESS | 7 | 54 | 5.489e-12 | 1.161e-09 |
| 22 | THIOESTER BIOSYNTHETIC PROCESS | 7 | 54 | 5.489e-12 | 1.161e-09 |
| 23 | SMALL MOLECULE BIOSYNTHETIC PROCESS | 12 | 443 | 9.306e-12 | 1.883e-09 |
| 24 | OXIDATION REDUCTION PROCESS | 15 | 898 | 1.245e-11 | 2.414e-09 |
| 25 | ALPHA LINOLENIC ACID METABOLIC PROCESS | 5 | 13 | 1.802e-11 | 3.353e-09 |
| 26 | ORGANIC ACID TRANSPORT | 10 | 261 | 2.274e-11 | 4.07e-09 |
| 27 | SINGLE ORGANISM BIOSYNTHETIC PROCESS | 17 | 1340 | 2.537e-11 | 4.373e-09 |
| 28 | ORGANIC ANION TRANSPORT | 11 | 387 | 4.814e-11 | 7.999e-09 |
| 29 | COENZYME BIOSYNTHETIC PROCESS | 8 | 127 | 5.514e-11 | 8.848e-09 |
| 30 | CARBOXYLIC ACID CATABOLIC PROCESS | 9 | 205 | 7.76e-11 | 1.165e-08 |
| 31 | ORGANIC ACID CATABOLIC PROCESS | 9 | 205 | 7.76e-11 | 1.165e-08 |
| 32 | THIOESTER METABOLIC PROCESS | 7 | 83 | 1.24e-10 | 1.749e-08 |
| 33 | ACYL COA METABOLIC PROCESS | 7 | 83 | 1.24e-10 | 1.749e-08 |
| 34 | NEUTRAL LIPID METABOLIC PROCESS | 7 | 85 | 1.471e-10 | 2.013e-08 |
| 35 | SMALL MOLECULE CATABOLIC PROCESS | 10 | 328 | 2.138e-10 | 2.843e-08 |
| 36 | SINGLE ORGANISM CATABOLIC PROCESS | 14 | 957 | 4.253e-10 | 5.497e-08 |
| 37 | COFACTOR BIOSYNTHETIC PROCESS | 8 | 166 | 4.721e-10 | 5.937e-08 |
| 38 | MONOCARBOXYLIC ACID BIOSYNTHETIC PROCESS | 8 | 172 | 6.263e-10 | 7.669e-08 |
| 39 | ANION TRANSPORT | 11 | 507 | 8.449e-10 | 1.002e-07 |
| 40 | UNSATURATED FATTY ACID METABOLIC PROCESS | 7 | 109 | 8.612e-10 | 1.002e-07 |
| 41 | REGULATION OF LIPID METABOLIC PROCESS | 9 | 282 | 1.308e-09 | 1.485e-07 |
| 42 | CELLULAR CATABOLIC PROCESS | 15 | 1322 | 2.758e-09 | 3.056e-07 |
| 43 | FATTY ACID BETA OXIDATION USING ACYL COA OXIDASE | 4 | 13 | 6.247e-09 | 6.606e-07 |
| 44 | REGULATION OF CHOLESTEROL STORAGE | 4 | 13 | 6.247e-09 | 6.606e-07 |
| 45 | ORGANIC HYDROXY COMPOUND METABOLIC PROCESS | 10 | 482 | 8.728e-09 | 9.025e-07 |
| 46 | CATABOLIC PROCESS | 16 | 1773 | 1.785e-08 | 1.806e-06 |
| 47 | COENZYME METABOLIC PROCESS | 8 | 265 | 1.876e-08 | 1.858e-06 |
| 48 | CARBOXYLIC ACID BIOSYNTHETIC PROCESS | 8 | 270 | 2.17e-08 | 2.06e-06 |
| 49 | ORGANIC ACID BIOSYNTHETIC PROCESS | 8 | 270 | 2.17e-08 | 2.06e-06 |
| 50 | LIPID HOMEOSTASIS | 6 | 110 | 4.108e-08 | 3.823e-06 |
| 51 | SULFUR COMPOUND BIOSYNTHETIC PROCESS | 7 | 203 | 6.509e-08 | 5.938e-06 |
| 52 | POSITIVE REGULATION OF LIPID METABOLIC PROCESS | 6 | 128 | 1.016e-07 | 9.095e-06 |
| 53 | COFACTOR METABOLIC PROCESS | 8 | 334 | 1.12e-07 | 9.835e-06 |
| 54 | ION TRANSPORT | 13 | 1262 | 1.392e-07 | 1.2e-05 |
| 55 | REGULATION OF MACROPHAGE DERIVED FOAM CELL DIFFERENTIATION | 4 | 28 | 1.755e-07 | 1.458e-05 |
| 56 | REGULATION OF FATTY ACID OXIDATION | 4 | 28 | 1.755e-07 | 1.458e-05 |
| 57 | GLYCEROLIPID METABOLIC PROCESS | 8 | 356 | 1.825e-07 | 1.49e-05 |
| 58 | RESPONSE TO FATTY ACID | 5 | 83 | 3.713e-07 | 2.978e-05 |
| 59 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 13 | 1381 | 3.949e-07 | 3.114e-05 |
| 60 | BILE ACID METABOLIC PROCESS | 4 | 35 | 4.448e-07 | 3.449e-05 |
| 61 | REGULATION OF FATTY ACID METABOLIC PROCESS | 5 | 87 | 4.7e-07 | 3.585e-05 |
| 62 | ORGANIC HYDROXY COMPOUND BIOSYNTHETIC PROCESS | 6 | 175 | 6.452e-07 | 4.842e-05 |
| 63 | REGULATION OF LIPID STORAGE | 4 | 41 | 8.536e-07 | 6.305e-05 |
| 64 | RESPONSE TO COLD | 4 | 43 | 1.038e-06 | 7.544e-05 |
| 65 | REGULATION OF SEQUESTERING OF TRIGLYCERIDE | 3 | 12 | 1.165e-06 | 8.341e-05 |
| 66 | RESPONSE TO INSULIN | 6 | 205 | 1.625e-06 | 0.0001145 |
| 67 | FATTY ACID BIOSYNTHETIC PROCESS | 5 | 114 | 1.803e-06 | 0.0001252 |
| 68 | POSITIVE REGULATION OF FATTY ACID OXIDATION | 3 | 14 | 1.923e-06 | 0.0001316 |
| 69 | SULFUR COMPOUND METABOLIC PROCESS | 7 | 359 | 3.02e-06 | 0.0002036 |
| 70 | FATTY ACID BETA OXIDATION USING ACYL COA DEHYDROGENASE | 3 | 18 | 4.29e-06 | 0.0002852 |
| 71 | CELLULAR RESPONSE TO HORMONE STIMULUS | 8 | 552 | 4.898e-06 | 0.000321 |
| 72 | BILE ACID BIOSYNTHETIC PROCESS | 3 | 20 | 5.978e-06 | 0.0003864 |
| 73 | RESPONSE TO TEMPERATURE STIMULUS | 5 | 148 | 6.497e-06 | 0.0004141 |
| 74 | TRIGLYCERIDE CATABOLIC PROCESS | 3 | 21 | 6.966e-06 | 0.000438 |
| 75 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 10 | 1008 | 7.745e-06 | 0.0004805 |
| 76 | ORGANIC HYDROXY COMPOUND TRANSPORT | 5 | 155 | 8.138e-06 | 0.0004982 |
| 77 | HORMONE MEDIATED SIGNALING PATHWAY | 5 | 158 | 8.933e-06 | 0.0005398 |
| 78 | NEUTRAL LIPID CATABOLIC PROCESS | 3 | 26 | 1.353e-05 | 0.0007971 |
| 79 | ACYLGLYCEROL CATABOLIC PROCESS | 3 | 26 | 1.353e-05 | 0.0007971 |
| 80 | REGULATION OF CELLULAR KETONE METABOLIC PROCESS | 5 | 173 | 1.387e-05 | 0.0008065 |
| 81 | CHEMICAL HOMEOSTASIS | 9 | 874 | 1.81e-05 | 0.001039 |
| 82 | VERY LONG CHAIN FATTY ACID METABOLIC PROCESS | 3 | 29 | 1.895e-05 | 0.001075 |
| 83 | RESPONSE TO HORMONE | 9 | 893 | 2.146e-05 | 0.001203 |
| 84 | RESPONSE TO NUTRIENT | 5 | 191 | 2.234e-05 | 0.001238 |
| 85 | REGULATION OF LIPID TRANSPORT | 4 | 95 | 2.504e-05 | 0.001371 |
| 86 | POSITIVE REGULATION OF FATTY ACID METABOLIC PROCESS | 3 | 33 | 2.816e-05 | 0.001523 |
| 87 | RESPONSE TO ENDOGENOUS STIMULUS | 11 | 1450 | 3.093e-05 | 0.001654 |
| 88 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO CELL PERIPHERY | 3 | 37 | 3.99e-05 | 0.002063 |
| 89 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO PLASMA MEMBRANE | 3 | 37 | 3.99e-05 | 0.002063 |
| 90 | GLYCEROLIPID CATABOLIC PROCESS | 3 | 37 | 3.99e-05 | 0.002063 |
| 91 | ALCOHOL BIOSYNTHETIC PROCESS | 4 | 111 | 4.616e-05 | 0.00236 |
| 92 | STEROID BIOSYNTHETIC PROCESS | 4 | 114 | 5.123e-05 | 0.002591 |
| 93 | STEROID METABOLIC PROCESS | 5 | 237 | 6.257e-05 | 0.003131 |
| 94 | REGULATION OF PLASMA LIPOPROTEIN PARTICLE LEVELS | 3 | 45 | 7.215e-05 | 0.003571 |
| 95 | RESPONSE TO PEPTIDE | 6 | 404 | 7.632e-05 | 0.003738 |
| 96 | REGULATION OF LIPID BIOSYNTHETIC PROCESS | 4 | 128 | 8.045e-05 | 0.003899 |
| 97 | HOMEOSTATIC PROCESS | 10 | 1337 | 8.799e-05 | 0.004178 |
| 98 | REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION TO PLASMA MEMBRANE | 3 | 48 | 8.762e-05 | 0.004178 |
| 99 | CELLULAR RESPONSE TO FATTY ACID | 3 | 51 | 0.0001051 | 0.00494 |
| 100 | RESPONSE TO DRUG | 6 | 431 | 0.0001089 | 0.005067 |
| 101 | RESPONSE TO EXTRACELLULAR STIMULUS | 6 | 441 | 0.0001235 | 0.005688 |
| 102 | CELLULAR RESPONSE TO INSULIN STIMULUS | 4 | 146 | 0.0001339 | 0.006106 |
| 103 | CELLULAR RESPONSE TO LIPID | 6 | 457 | 0.0001499 | 0.006774 |
| 104 | UNSATURATED FATTY ACID BIOSYNTHETIC PROCESS | 3 | 58 | 0.0001544 | 0.006907 |
| 105 | POSITIVE REGULATION OF MEMBRANE INVAGINATION | 2 | 11 | 0.0001715 | 0.007528 |
| 106 | POSITIVE REGULATION OF PHAGOCYTOSIS ENGULFMENT | 2 | 11 | 0.0001715 | 0.007528 |
| 107 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 8 | 917 | 0.0001823 | 0.007927 |
| 108 | AMMONIUM ION METABOLIC PROCESS | 4 | 169 | 0.0002347 | 0.009569 |
| 109 | WHITE FAT CELL DIFFERENTIATION | 2 | 13 | 0.0002427 | 0.009569 |
| 110 | LIPOPROTEIN TRANSPORT | 2 | 13 | 0.0002427 | 0.009569 |
| 111 | REGULATION OF PHAGOCYTOSIS ENGULFMENT | 2 | 13 | 0.0002427 | 0.009569 |
| 112 | CARNITINE METABOLIC PROCESS | 2 | 13 | 0.0002427 | 0.009569 |
| 113 | LIPOPROTEIN LOCALIZATION | 2 | 13 | 0.0002427 | 0.009569 |
| 114 | NEGATIVE REGULATION OF MACROPHAGE DERIVED FOAM CELL DIFFERENTIATION | 2 | 13 | 0.0002427 | 0.009569 |
| 115 | INTRACELLULAR RECEPTOR SIGNALING PATHWAY | 4 | 168 | 0.0002294 | 0.009569 |
| 116 | REGULATION OF PHOSPHOLIPID BIOSYNTHETIC PROCESS | 2 | 13 | 0.0002427 | 0.009569 |
| 117 | NEGATIVE REGULATION OF CARDIAC MUSCLE CELL APOPTOTIC PROCESS | 2 | 13 | 0.0002427 | 0.009569 |
| 118 | REGULATION OF MEMBRANE INVAGINATION | 2 | 13 | 0.0002427 | 0.009569 |
| 119 | RESPONSE TO ACID CHEMICAL | 5 | 319 | 0.0002513 | 0.009827 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | LONG CHAIN FATTY ACID COA LIGASE ACTIVITY | 7 | 13 | 5.603e-17 | 5.206e-14 |
| 2 | FATTY ACID LIGASE ACTIVITY | 7 | 16 | 3.721e-16 | 1.729e-13 |
| 3 | LIGASE ACTIVITY FORMING CARBON SULFUR BONDS | 7 | 40 | 5.883e-13 | 1.822e-10 |
| 4 | FATTY ACID BINDING | 5 | 31 | 2.324e-09 | 5.397e-07 |
| 5 | MONOCARBOXYLIC ACID BINDING | 5 | 65 | 1.081e-07 | 2.009e-05 |
| 6 | FATTY ACYL COA BINDING | 4 | 31 | 2.686e-07 | 4.16e-05 |
| 7 | OXIDOREDUCTASE ACTIVITY ACTING ON THE CH CH GROUP OF DONORS | 4 | 57 | 3.262e-06 | 0.0003876 |
| 8 | OXIDOREDUCTASE ACTIVITY | 9 | 719 | 3.755e-06 | 0.0003876 |
| 9 | ACYL COA DEHYDROGENASE ACTIVITY | 3 | 17 | 3.579e-06 | 0.0003876 |
| 10 | LIGASE ACTIVITY | 7 | 406 | 6.773e-06 | 0.0006293 |
| 11 | LIPID BINDING | 8 | 657 | 1.741e-05 | 0.001348 |
| 12 | COENZYME BINDING | 5 | 179 | 1.635e-05 | 0.001348 |
| 13 | ORGANIC ACID BINDING | 5 | 209 | 3.44e-05 | 0.002459 |
| 14 | ELECTRON CARRIER ACTIVITY | 4 | 112 | 4.781e-05 | 0.003172 |
| 15 | SULFUR COMPOUND BINDING | 5 | 234 | 5.891e-05 | 0.003648 |
| 16 | TRANSFERASE ACTIVITY TRANSFERRING ACYL GROUPS | 5 | 249 | 7.903e-05 | 0.004588 |
| 17 | TRANSCRIPTION FACTOR ACTIVITY DIRECT LIGAND REGULATED SEQUENCE SPECIFIC DNA BINDING | 3 | 48 | 8.762e-05 | 0.004788 |
| 18 | COFACTOR BINDING | 5 | 263 | 0.0001022 | 0.005277 |
| 19 | STEROID HORMONE RECEPTOR ACTIVITY | 3 | 59 | 0.0001625 | 0.007943 |
| 20 | FATTY ACID TRANSPORTER ACTIVITY | 2 | 12 | 0.0002056 | 0.009094 |
| 21 | RECEPTOR BINDING | 10 | 1476 | 0.0001997 | 0.009094 |
| 22 | LOW DENSITY LIPOPROTEIN RECEPTOR ACTIVITY | 2 | 13 | 0.0002427 | 0.009928 |
| 23 | ADENYL NUCLEOTIDE BINDING | 10 | 1514 | 0.0002458 | 0.009928 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | MICROBODY PART | 10 | 93 | 6.611e-16 | 3.861e-13 |
| 2 | MICROBODY | 10 | 134 | 2.842e-14 | 8.298e-12 |
| 3 | MITOCHONDRION | 18 | 1633 | 5.175e-11 | 1.007e-08 |
| 4 | MICROBODY MEMBRANE | 6 | 58 | 8.301e-10 | 1.212e-07 |
| 5 | OUTER MEMBRANE | 8 | 190 | 1.379e-09 | 1.611e-07 |
| 6 | MICROBODY LUMEN | 5 | 45 | 1.641e-08 | 1.597e-06 |
| 7 | MITOCHONDRIAL ENVELOPE | 11 | 691 | 2.113e-08 | 1.763e-06 |
| 8 | MITOCHONDRIAL PART | 12 | 953 | 5.542e-08 | 4.046e-06 |
| 9 | ENVELOPE | 11 | 1090 | 2.04e-06 | 0.0001324 |
| 10 | NUCLEAR OUTER MEMBRANE ENDOPLASMIC RETICULUM MEMBRANE NETWORK | 9 | 1005 | 5.427e-05 | 0.003169 |
| 11 | ENDOPLASMIC RETICULUM PART | 9 | 1163 | 0.0001664 | 0.008832 |
| 12 | LIPID PARTICLE | 3 | 62 | 0.0001883 | 0.009164 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
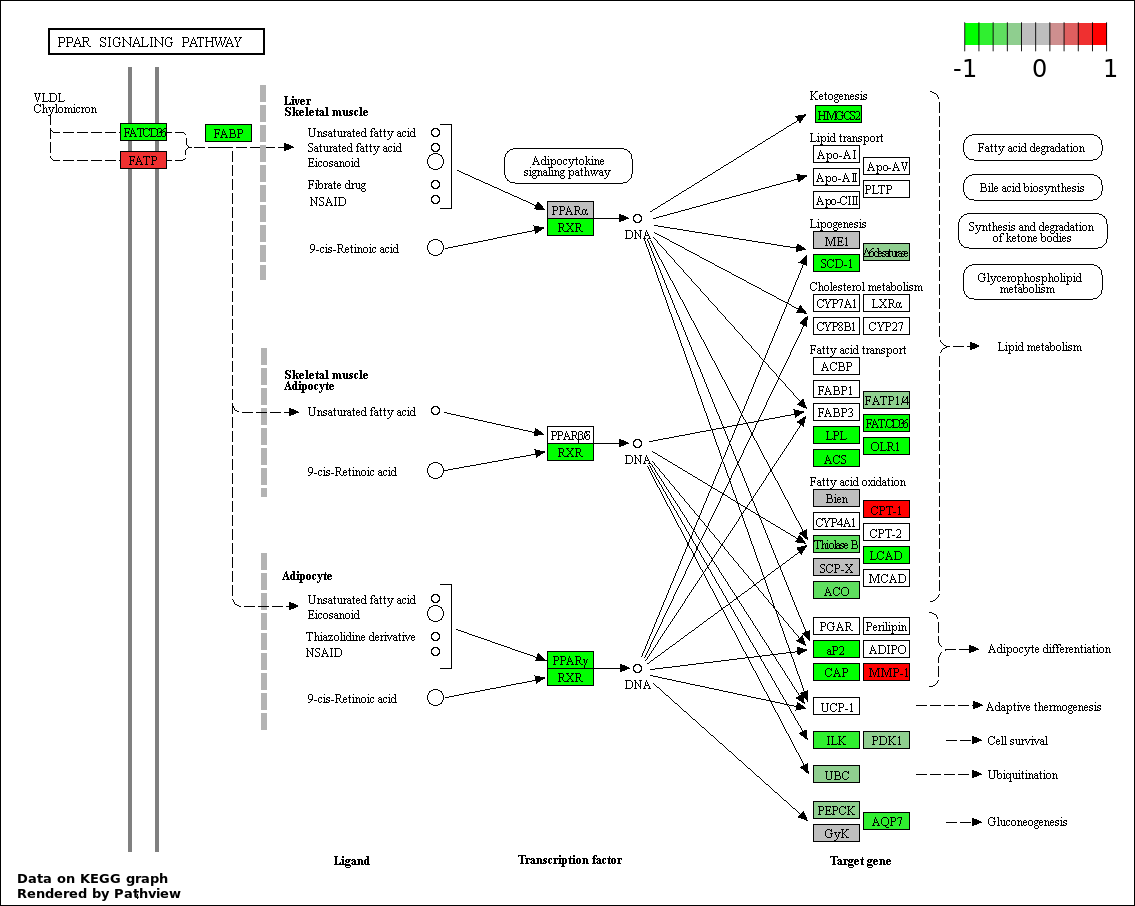
| 1 | hsa03320_PPAR_signaling_pathway | 36 | 70 | 6.093e-94 | 1.097e-91 | |
| 2 | hsa00071_Fatty_acid_metabolism | 11 | 43 | 6.511e-22 | 5.86e-20 | |
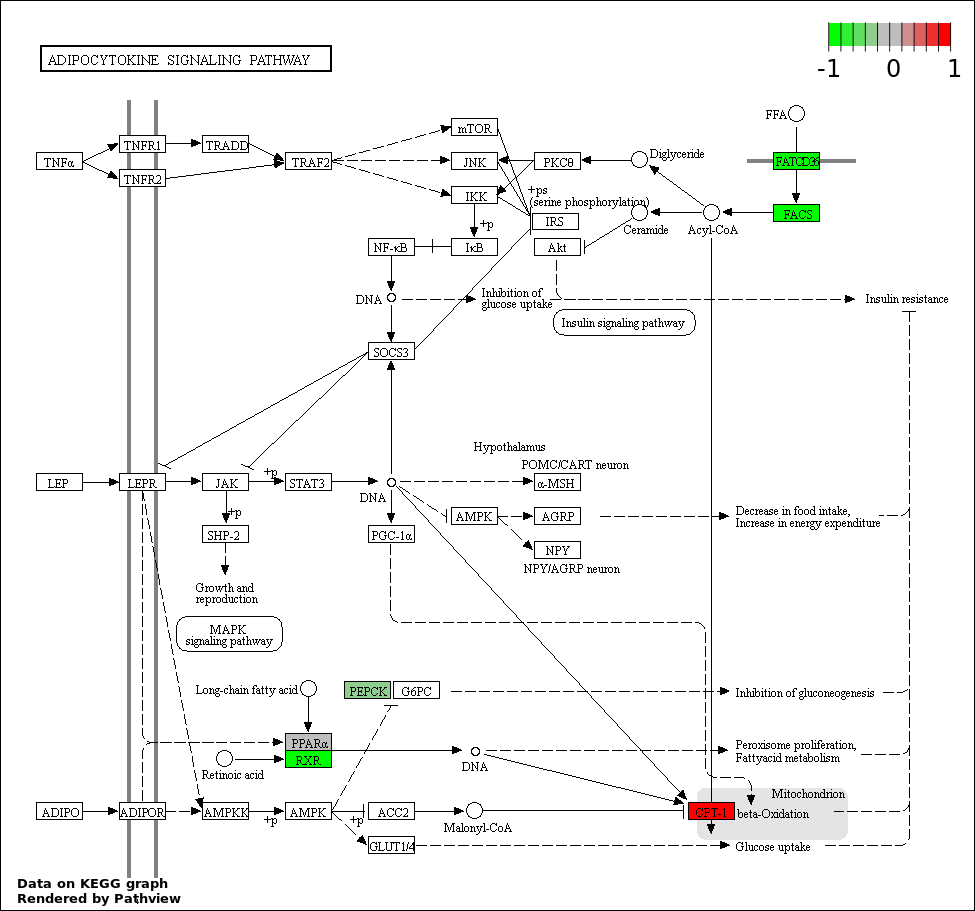
| 3 | hsa04920_Adipocytokine_signaling_pathway | 11 | 68 | 1.686e-19 | 1.012e-17 | |
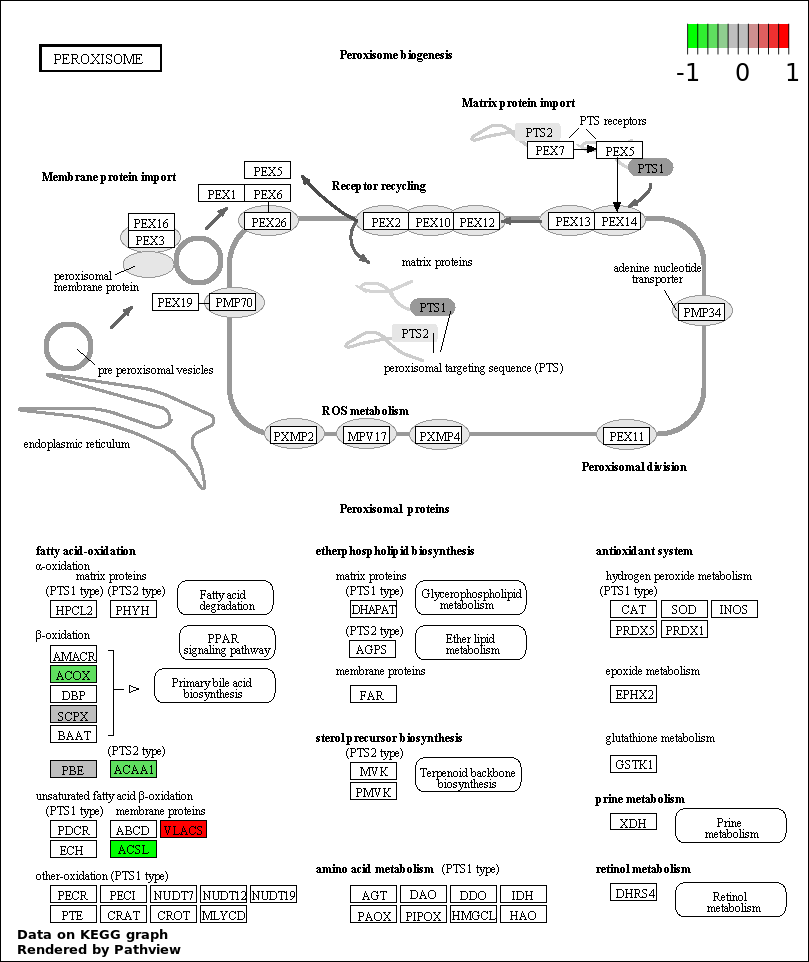
| 4 | hsa04146_Peroxisome | 11 | 79 | 9.814e-19 | 4.416e-17 | |
| 5 | hsa01040_Biosynthesis_of_unsaturated_fatty_acids | 5 | 21 | 2.819e-10 | 1.015e-08 | |
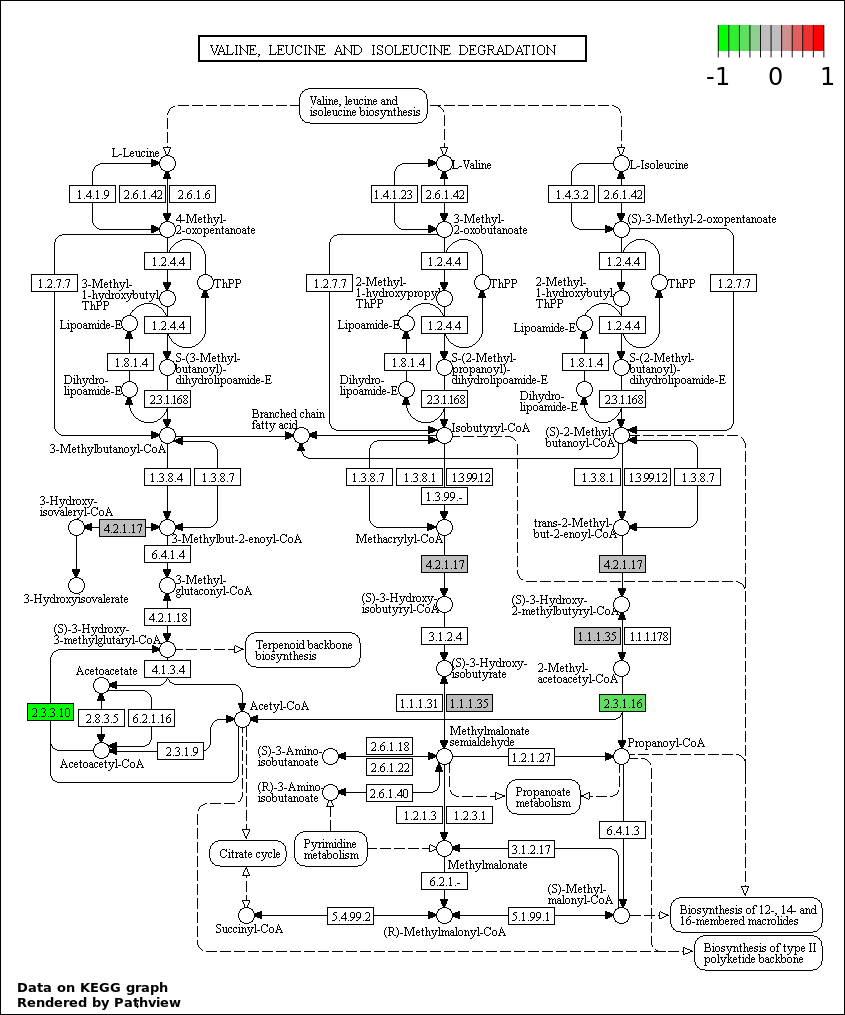
| 6 | hsa00280_Valine._leucine_and_isoleucine_degradation | 3 | 44 | 6.742e-05 | 0.002023 | |
| 7 | hsa00120_Primary_bile_acid_biosynthesis | 2 | 16 | 0.0003721 | 0.009567 | |
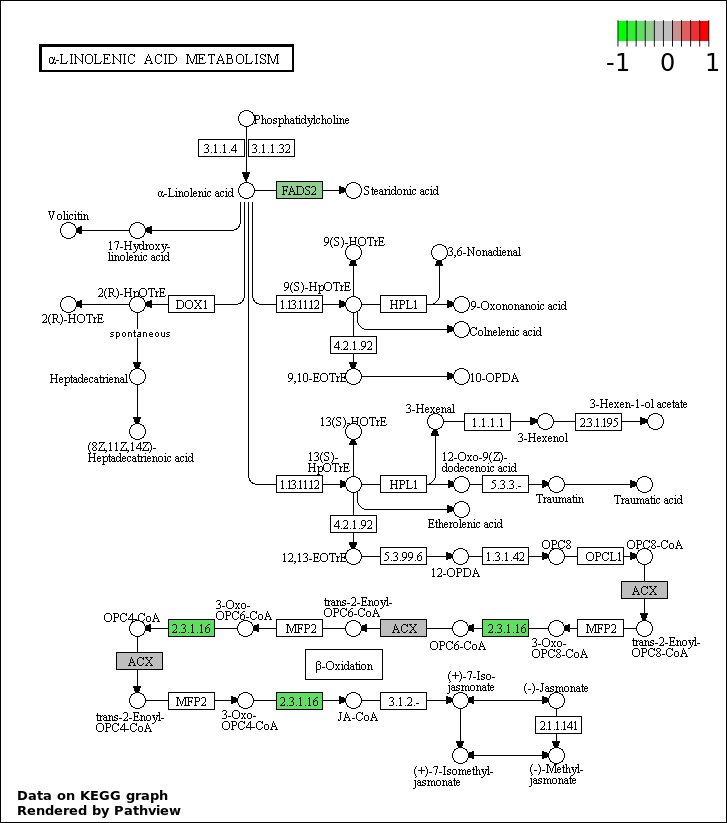
| 8 | hsa00592_alpha.Linolenic_acid_metabolism | 2 | 20 | 0.0005864 | 0.0132 | |
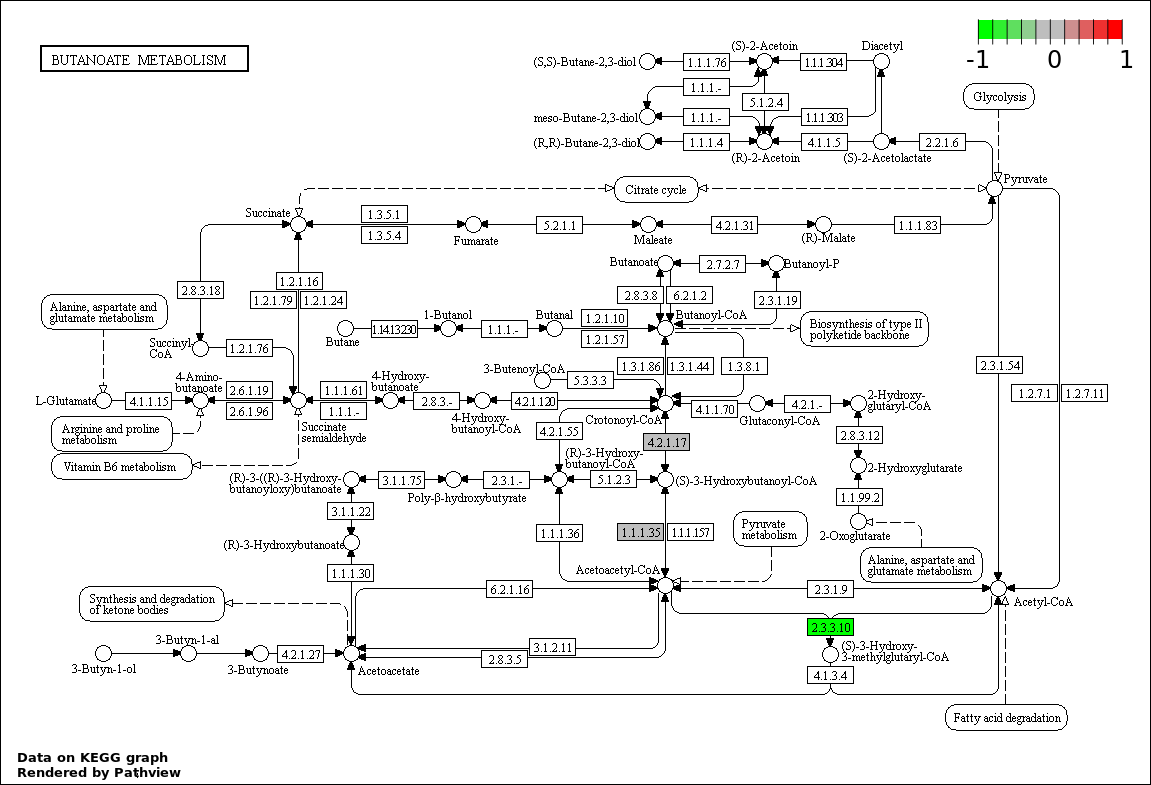
| 9 | hsa00650_Butanoate_metabolism | 2 | 30 | 0.001328 | 0.02655 | |
| 10 | hsa04910_Insulin_signaling_pathway | 3 | 138 | 0.001943 | 0.03497 | |
| 11 | hsa00620_Pyruvate_metabolism | 2 | 40 | 0.002354 | 0.03851 | |
| 12 | hsa00561_Glycerolipid_metabolism | 2 | 50 | 0.003655 | 0.05482 | |
| 13 | hsa04145_Phagosome | 2 | 156 | 0.03203 | 0.3843 | |
| 14 | hsa04510_Focal_adhesion | 2 | 200 | 0.0502 | 0.4988 | |
| 15 | hsa04151_PI3K_AKT_signaling_pathway | 2 | 351 | 0.1312 | 0.8437 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | MAGI2-AS3 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-26b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 18 | ACSL4 | Sponge network | -1.892 | 0 | -1.282 | 0 | 0.321 |
| 2 | RP11-166D19.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-26b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-7-1-3p | 15 | ACSL4 | Sponge network | -0.582 | 0.05253 | -1.282 | 0 | 0.307 |
| 3 | LINC00702 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-148a-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-26b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-34a-5p;hsa-miR-450b-5p;hsa-miR-454-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 20 | ACSL4 | Sponge network | -2.856 | 0 | -1.282 | 0 | 0.289 |
| 4 | RP11-532F6.3 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-26b-3p;hsa-miR-301a-3p | 10 | ACSL4 | Sponge network | -2.028 | 0 | -1.282 | 0 | 0.285 |
| 5 | CTD-2013N24.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-148a-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-26b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 19 | ACSL4 | Sponge network | -1.745 | 0 | -1.282 | 0 | 0.266 |
| 6 | CTD-2269F5.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 11 | ACSL4 | Sponge network | -1.576 | 0.00334 | -1.282 | 0 | 0.264 |
| 7 | LINC00968 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-148a-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-26b-3p;hsa-miR-301a-3p;hsa-miR-450b-5p;hsa-miR-93-5p | 16 | ACSL4 | Sponge network | -4.19 | 0 | -1.282 | 0 | 0.258 |