This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-301a-3p | ABHD2 | -0.76 | 0.09613 | 0.63 | 0.15716 | mirMAP | -0.32 | 0 | NA | |
| 2 | hsa-miR-301a-3p | ABTB2 | -0.76 | 0.09613 | 0.09 | 0.87066 | mirMAP | -0.41 | 0 | NA | |
| 3 | hsa-miR-301a-3p | ACBD3 | -0.76 | 0.09613 | 0.56 | 0.02656 | MirTarget; miRNATAP | -0.14 | 0.00051 | NA | |
| 4 | hsa-miR-301a-3p | ACSL4 | -0.76 | 0.09613 | -0.57 | 0.06388 | MirTarget; miRNATAP | -0.17 | 0.00063 | NA | |
| 5 | hsa-miR-301a-3p | ACVR1 | -0.76 | 0.09613 | 0.3 | 0.22609 | MirTarget; miRNATAP | -0.18 | 1.0E-5 | NA | |
| 6 | hsa-miR-301a-3p | ADAM12 | -0.76 | 0.09613 | -1.12 | 0.25198 | miRNATAP | -0.65 | 4.0E-5 | NA | |
| 7 | hsa-miR-301a-3p | AGFG1 | -0.76 | 0.09613 | 0.01 | 0.95401 | MirTarget; miRNATAP | -0.13 | 3.0E-5 | NA | |
| 8 | hsa-miR-301a-3p | ANKRD28 | -0.76 | 0.09613 | -0.03 | 0.87325 | MirTarget; miRNATAP | -0.11 | 0.00066 | NA | |
| 9 | hsa-miR-301a-3p | ANXA11 | -0.76 | 0.09613 | 0.21 | 0.52584 | mirMAP | -0.29 | 0 | NA | |
| 10 | hsa-miR-301a-3p | APBB2 | -0.76 | 0.09613 | -0.11 | 0.68147 | mirMAP | -0.13 | 0.00265 | NA | |
| 11 | hsa-miR-301a-3p | APCDD1 | -0.76 | 0.09613 | 0.26 | 0.63295 | MirTarget | -0.33 | 0.00017 | NA | |
| 12 | hsa-miR-301a-3p | AR | -0.76 | 0.09613 | -0.52 | 0.55352 | mirMAP; miRNATAP | -0.4 | 0.00488 | 25940439 | Mechanism dissection revealed infiltrating pre-adipocytes might function through down-regulation of the androgen receptor AR via modulation of miR-301a and then increase PCa cell invasion via induction of TGF-β1/Smad/MMP9 signals |
| 13 | hsa-miR-301a-3p | ARAP2 | -0.76 | 0.09613 | -0.25 | 0.55742 | MirTarget | -0.34 | 0 | NA | |
| 14 | hsa-miR-301a-3p | ARHGAP12 | -0.76 | 0.09613 | 0.37 | 0.2122 | miRNATAP | -0.15 | 0.00123 | NA | |
| 15 | hsa-miR-301a-3p | ARHGAP21 | -0.76 | 0.09613 | 0.3 | 0.28692 | MirTarget; miRNATAP | -0.24 | 0 | NA | |
| 16 | hsa-miR-301a-3p | ARHGAP26 | -0.76 | 0.09613 | 0.27 | 0.55575 | mirMAP | -0.35 | 0 | NA | |
| 17 | hsa-miR-301a-3p | ARL6IP1 | -0.76 | 0.09613 | 0.51 | 0.12034 | MirTarget | -0.16 | 0.00307 | NA | |
| 18 | hsa-miR-301a-3p | ATF6 | -0.76 | 0.09613 | 0.23 | 0.30334 | mirMAP | -0.11 | 0.00271 | NA | |
| 19 | hsa-miR-301a-3p | ATG16L1 | -0.76 | 0.09613 | 0.32 | 0.16633 | MirTarget; miRNATAP | -0.14 | 0.00012 | NA | |
| 20 | hsa-miR-301a-3p | ATP11A | -0.76 | 0.09613 | 0.11 | 0.75717 | MirTarget; miRNATAP | -0.16 | 0.00587 | NA | |
| 21 | hsa-miR-301a-3p | ATR | -0.76 | 0.09613 | -0.04 | 0.82153 | mirMAP | -0.1 | 0.00132 | NA | |
| 22 | hsa-miR-301a-3p | ATXN1 | -0.76 | 0.09613 | -0.51 | 0.09797 | miRNATAP | -0.17 | 0.00057 | NA | |
| 23 | hsa-miR-301a-3p | B4GALT5 | -0.76 | 0.09613 | 0.16 | 0.60085 | miRNATAP | -0.15 | 0.00206 | NA | |
| 24 | hsa-miR-301a-3p | BCL9L | -0.76 | 0.09613 | 0.28 | 0.39221 | mirMAP | -0.16 | 0.00185 | NA | |
| 25 | hsa-miR-301a-3p | BMPR1B | -0.76 | 0.09613 | 0.22 | 0.768 | miRNATAP | -0.75 | 0 | NA | |
| 26 | hsa-miR-301a-3p | BMPR2 | -0.76 | 0.09613 | -0.03 | 0.9248 | MirTarget; miRNATAP | -0.12 | 0.00892 | NA | |
| 27 | hsa-miR-301a-3p | BTBD10 | -0.76 | 0.09613 | 0.37 | 0.08561 | MirTarget | -0.11 | 0.00144 | NA | |
| 28 | hsa-miR-301a-3p | BTN2A2 | -0.76 | 0.09613 | -1.12 | 0.00207 | mirMAP | -0.18 | 0.00333 | NA | |
| 29 | hsa-miR-301a-3p | C15orf52 | -0.76 | 0.09613 | 0.11 | 0.85147 | mirMAP | -0.25 | 0.00658 | NA | |
| 30 | hsa-miR-301a-3p | C1S | -0.76 | 0.09613 | -0.53 | 0.36559 | MirTarget | -0.5 | 0 | NA | |
| 31 | hsa-miR-301a-3p | C2orf49 | -0.76 | 0.09613 | 0.24 | 0.24355 | mirMAP | -0.22 | 0 | NA | |
| 32 | hsa-miR-301a-3p | CA13 | -0.76 | 0.09613 | 1.34 | 0.0047 | MirTarget | -0.22 | 0.00456 | NA | |
| 33 | hsa-miR-301a-3p | CAMK4 | -0.76 | 0.09613 | -0.61 | 0.35184 | mirMAP | -0.31 | 0.00432 | NA | |
| 34 | hsa-miR-301a-3p | CCDC50 | -0.76 | 0.09613 | -0.36 | 0.09894 | mirMAP | -0.12 | 0.00086 | NA | |
| 35 | hsa-miR-301a-3p | CCDC6 | -0.76 | 0.09613 | -0.08 | 0.72287 | MirTarget | -0.15 | 4.0E-5 | NA | |
| 36 | hsa-miR-301a-3p | CCNA2 | -0.76 | 0.09613 | 0.67 | 0.16545 | miRNATAP | -0.2 | 0.00997 | NA | |
| 37 | hsa-miR-301a-3p | CD109 | -0.76 | 0.09613 | 0.78 | 0.26548 | mirMAP | -0.44 | 0.0001 | NA | |
| 38 | hsa-miR-301a-3p | CDC73 | -0.76 | 0.09613 | 0.22 | 0.34941 | miRNATAP | -0.13 | 0.00064 | NA | |
| 39 | hsa-miR-301a-3p | CDH11 | -0.76 | 0.09613 | -0.6 | 0.41785 | mirMAP | -0.66 | 0 | NA | |
| 40 | hsa-miR-301a-3p | CEP55 | -0.76 | 0.09613 | 0.99 | 0.14629 | MirTarget | -0.58 | 0 | NA | |
| 41 | hsa-miR-301a-3p | CFLAR | -0.76 | 0.09613 | -0.19 | 0.52443 | mirMAP | -0.2 | 2.0E-5 | NA | |
| 42 | hsa-miR-301a-3p | CHRNA7 | -0.76 | 0.09613 | -0.37 | 0.67746 | mirMAP | -0.66 | 0 | NA | |
| 43 | hsa-miR-301a-3p | CMPK1 | -0.76 | 0.09613 | 0.21 | 0.42634 | MirTarget; miRNATAP | -0.15 | 0.00022 | NA | |
| 44 | hsa-miR-301a-3p | COL4A4 | -0.76 | 0.09613 | -0.35 | 0.65939 | mirMAP | -0.42 | 0.00112 | NA | |
| 45 | hsa-miR-301a-3p | CTSK | -0.76 | 0.09613 | -0.05 | 0.94011 | MirTarget | -0.49 | 0 | NA | |
| 46 | hsa-miR-301a-3p | CYBRD1 | -0.76 | 0.09613 | 0.13 | 0.82152 | miRNAWalker2 validate | -0.35 | 9.0E-5 | NA | |
| 47 | hsa-miR-301a-3p | DCBLD2 | -0.76 | 0.09613 | 1.11 | 0.10472 | MirTarget; miRNATAP | -0.37 | 0.00096 | NA | |
| 48 | hsa-miR-301a-3p | DEPDC1 | -0.76 | 0.09613 | 0.87 | 0.19097 | MirTarget | -0.38 | 0.00038 | NA | |
| 49 | hsa-miR-301a-3p | DIAPH2 | -0.76 | 0.09613 | 0.37 | 0.26798 | mirMAP | -0.37 | 0 | NA | |
| 50 | hsa-miR-301a-3p | DIAPH3 | -0.76 | 0.09613 | 1.01 | 0.0985 | MirTarget; miRNATAP | -0.53 | 0 | NA | |
| 51 | hsa-miR-301a-3p | DSG2 | -0.76 | 0.09613 | 1.16 | 0.02127 | mirMAP | -0.28 | 0.00054 | NA | |
| 52 | hsa-miR-301a-3p | E2F2 | -0.76 | 0.09613 | -0.19 | 0.7383 | miRNATAP | -0.29 | 0.00119 | NA | |
| 53 | hsa-miR-301a-3p | E2F7 | -0.76 | 0.09613 | 0.87 | 0.2158 | miRNATAP | -0.51 | 0 | NA | |
| 54 | hsa-miR-301a-3p | EDN1 | -0.76 | 0.09613 | -0.52 | 0.36358 | MirTarget | -0.39 | 2.0E-5 | NA | |
| 55 | hsa-miR-301a-3p | EFNB2 | -0.76 | 0.09613 | 0.86 | 0.03481 | miRNATAP | -0.37 | 0 | NA | |
| 56 | hsa-miR-301a-3p | EIF4EBP2 | -0.76 | 0.09613 | -0.05 | 0.76069 | mirMAP | -0.18 | 0 | NA | |
| 57 | hsa-miR-301a-3p | ELK3 | -0.76 | 0.09613 | -0.28 | 0.5575 | MirTarget; miRNATAP | -0.22 | 0.00519 | NA | |
| 58 | hsa-miR-301a-3p | ENDOD1 | -0.76 | 0.09613 | 0.39 | 0.28326 | MirTarget | -0.17 | 0.00315 | NA | |
| 59 | hsa-miR-301a-3p | EPHB4 | -0.76 | 0.09613 | 0.6 | 0.15515 | miRNATAP | -0.41 | 0 | NA | |
| 60 | hsa-miR-301a-3p | EREG | -0.76 | 0.09613 | 4.41 | 0.00138 | miRNATAP | -1.27 | 0 | NA | |
| 61 | hsa-miR-301a-3p | ERLIN2 | -0.76 | 0.09613 | 0.2 | 0.43102 | mirMAP | -0.14 | 0.00071 | NA | |
| 62 | hsa-miR-301a-3p | EXT1 | -0.76 | 0.09613 | 0.16 | 0.54187 | mirMAP | -0.22 | 0 | NA | |
| 63 | hsa-miR-301a-3p | FBXO28 | -0.76 | 0.09613 | 0.35 | 0.09605 | MirTarget; miRNATAP | -0.11 | 0.00087 | NA | |
| 64 | hsa-miR-301a-3p | FGD4 | -0.76 | 0.09613 | 0.52 | 0.17379 | mirMAP | -0.16 | 0.00759 | NA | |
| 65 | hsa-miR-301a-3p | FIBIN | -0.76 | 0.09613 | -0.38 | 0.63537 | MirTarget | -0.59 | 0 | NA | |
| 66 | hsa-miR-301a-3p | FMN1 | -0.76 | 0.09613 | 1.42 | 0.04177 | mirMAP | -0.52 | 0 | NA | |
| 67 | hsa-miR-301a-3p | FNDC4 | -0.76 | 0.09613 | -0.45 | 0.36303 | miRNATAP | -0.23 | 0.0042 | NA | |
| 68 | hsa-miR-301a-3p | FOSL1 | -0.76 | 0.09613 | 2.38 | 0.00428 | MirTarget | -0.6 | 1.0E-5 | NA | |
| 69 | hsa-miR-301a-3p | FOXF2 | -0.76 | 0.09613 | 1.14 | 0.0316 | miRNATAP | -0.45 | 0 | NA | |
| 70 | hsa-miR-301a-3p | FOXP1 | -0.76 | 0.09613 | -0.02 | 0.93749 | mirMAP | -0.18 | 0 | NA | |
| 71 | hsa-miR-301a-3p | FRMD6 | -0.76 | 0.09613 | -0.32 | 0.5949 | MirTarget | -0.5 | 0 | NA | |
| 72 | hsa-miR-301a-3p | G0S2 | -0.76 | 0.09613 | 0.15 | 0.84254 | MirTarget | -0.37 | 0.00224 | NA | |
| 73 | hsa-miR-301a-3p | GDA | -0.76 | 0.09613 | 2.64 | 0.01236 | miRNATAP | -0.51 | 0.00299 | NA | |
| 74 | hsa-miR-301a-3p | GJA1 | -0.76 | 0.09613 | -0.29 | 0.55396 | MirTarget; miRNATAP | -0.3 | 0.00012 | NA | |
| 75 | hsa-miR-301a-3p | GPR157 | -0.76 | 0.09613 | 0.08 | 0.88757 | mirMAP | -0.27 | 0.00223 | NA | |
| 76 | hsa-miR-301a-3p | GPRC5A | -0.76 | 0.09613 | 3.96 | 0.00162 | mirMAP | -1.51 | 0 | NA | |
| 77 | hsa-miR-301a-3p | GSPT1 | -0.76 | 0.09613 | 0.25 | 0.20157 | mirMAP | -0.17 | 0 | NA | |
| 78 | hsa-miR-301a-3p | H6PD | -0.76 | 0.09613 | -0.58 | 0.04995 | mirMAP | -0.23 | 0 | NA | |
| 79 | hsa-miR-301a-3p | HECA | -0.76 | 0.09613 | -0.49 | 0.04604 | MirTarget; miRNATAP | -0.16 | 4.0E-5 | NA | |
| 80 | hsa-miR-301a-3p | HES2 | -0.76 | 0.09613 | 0.16 | 0.86833 | mirMAP | -0.55 | 0.00049 | NA | |
| 81 | hsa-miR-301a-3p | HOXA3 | -0.76 | 0.09613 | 1.12 | 0.05646 | miRNATAP | -0.52 | 0 | NA | |
| 82 | hsa-miR-301a-3p | HOXA5 | -0.76 | 0.09613 | 1.18 | 0.02773 | miRNATAP | -0.41 | 0 | NA | |
| 83 | hsa-miR-301a-3p | HRH1 | -0.76 | 0.09613 | 1.23 | 0.01267 | mirMAP | -0.47 | 0 | NA | |
| 84 | hsa-miR-301a-3p | IGF1 | -0.76 | 0.09613 | -1.41 | 0.20856 | MirTarget | -0.51 | 0.00543 | NA | |
| 85 | hsa-miR-301a-3p | IGF2 | -0.76 | 0.09613 | 0.45 | 0.58211 | mirMAP | -0.42 | 0.00151 | NA | |
| 86 | hsa-miR-301a-3p | IGF2BP1 | -0.76 | 0.09613 | 0.61 | 0.61732 | mirMAP; miRNATAP | -0.67 | 0.00073 | NA | |
| 87 | hsa-miR-301a-3p | IGSF3 | -0.76 | 0.09613 | 0.78 | 0.14083 | MirTarget | -0.43 | 0 | NA | |
| 88 | hsa-miR-301a-3p | IL15 | -0.76 | 0.09613 | -0.95 | 0.00887 | MirTarget | -0.17 | 0.00339 | NA | |
| 89 | hsa-miR-301a-3p | IL17RA | -0.76 | 0.09613 | -0.28 | 0.27875 | mirMAP | -0.13 | 0.00101 | NA | |
| 90 | hsa-miR-301a-3p | IL17RD | -0.76 | 0.09613 | -0.42 | 0.32296 | mirMAP | -0.39 | 0 | NA | |
| 91 | hsa-miR-301a-3p | IL1RAP | -0.76 | 0.09613 | 0.24 | 0.6772 | MirTarget | -0.47 | 0 | NA | |
| 92 | hsa-miR-301a-3p | ITGA2 | -0.76 | 0.09613 | 0.8 | 0.18125 | mirMAP | -0.46 | 0 | NA | |
| 93 | hsa-miR-301a-3p | ITPRIPL2 | -0.76 | 0.09613 | 0.18 | 0.53284 | MirTarget; miRNATAP | -0.32 | 0 | NA | |
| 94 | hsa-miR-301a-3p | IYD | -0.76 | 0.09613 | 3.23 | 0.01005 | mirMAP | -1.29 | 0 | NA | |
| 95 | hsa-miR-301a-3p | KCNE3 | -0.76 | 0.09613 | 1.28 | 0.07886 | mirMAP | -0.59 | 0 | NA | |
| 96 | hsa-miR-301a-3p | KDELR2 | -0.76 | 0.09613 | 0.88 | 0.00046 | miRNAWalker2 validate | -0.18 | 1.0E-5 | NA | |
| 97 | hsa-miR-301a-3p | KDSR | -0.76 | 0.09613 | 0.09 | 0.62561 | mirMAP | -0.13 | 3.0E-5 | NA | |
| 98 | hsa-miR-301a-3p | KIAA1211 | -0.76 | 0.09613 | 1.02 | 0.07908 | MirTarget | -0.25 | 0.00753 | NA | |
| 99 | hsa-miR-301a-3p | KIAA1217 | -0.76 | 0.09613 | 0.15 | 0.72848 | MirTarget | -0.52 | 0 | NA | |
| 100 | hsa-miR-301a-3p | KIF18B | -0.76 | 0.09613 | 0.91 | 0.15141 | mirMAP | -0.4 | 8.0E-5 | NA | |
| 101 | hsa-miR-301a-3p | KIF1C | -0.76 | 0.09613 | -0.08 | 0.7809 | mirMAP | -0.24 | 0 | NA | |
| 102 | hsa-miR-301a-3p | KIF26B | -0.76 | 0.09613 | 0.51 | 0.44044 | mirMAP | -0.55 | 0 | NA | |
| 103 | hsa-miR-301a-3p | KIRREL | -0.76 | 0.09613 | -0.45 | 0.56708 | mirMAP | -0.35 | 0.00696 | NA | |
| 104 | hsa-miR-301a-3p | KLF3 | -0.76 | 0.09613 | 0.42 | 0.13439 | miRNATAP | -0.29 | 0 | NA | |
| 105 | hsa-miR-301a-3p | KLF4 | -0.76 | 0.09613 | 0.88 | 0.08321 | miRNATAP | -0.26 | 0.00133 | NA | |
| 106 | hsa-miR-301a-3p | KLHL20 | -0.76 | 0.09613 | -0.11 | 0.66181 | MirTarget; miRNATAP | -0.14 | 0.00034 | NA | |
| 107 | hsa-miR-301a-3p | LDLR | -0.76 | 0.09613 | 0.91 | 0.1166 | miRNATAP | -0.44 | 0 | NA | |
| 108 | hsa-miR-301a-3p | LONRF3 | -0.76 | 0.09613 | 1.46 | 0.00559 | MirTarget | -0.54 | 0 | NA | |
| 109 | hsa-miR-301a-3p | LPP | -0.76 | 0.09613 | 0.05 | 0.92861 | mirMAP | -0.54 | 0 | NA | |
| 110 | hsa-miR-301a-3p | LRRC58 | -0.76 | 0.09613 | 0.03 | 0.91685 | mirMAP | -0.17 | 0.00022 | NA | |
| 111 | hsa-miR-301a-3p | MACC1 | -0.76 | 0.09613 | 0.54 | 0.4139 | mirMAP | -0.56 | 0 | NA | |
| 112 | hsa-miR-301a-3p | MAP4 | -0.76 | 0.09613 | -0.25 | 0.18178 | MirTarget | -0.16 | 0 | NA | |
| 113 | hsa-miR-301a-3p | MBNL1 | -0.76 | 0.09613 | -0.55 | 0.02896 | MirTarget; miRNATAP | -0.13 | 0.00195 | NA | |
| 114 | hsa-miR-301a-3p | MBNL3 | -0.76 | 0.09613 | -0.26 | 0.67103 | MirTarget; mirMAP; miRNATAP | -0.4 | 8.0E-5 | NA | |
| 115 | hsa-miR-301a-3p | MCFD2 | -0.76 | 0.09613 | 0.22 | 0.37806 | mirMAP | -0.13 | 0.00128 | NA | |
| 116 | hsa-miR-301a-3p | MDFIC | -0.76 | 0.09613 | -0.75 | 0.12581 | MirTarget; miRNATAP | -0.35 | 1.0E-5 | NA | |
| 117 | hsa-miR-301a-3p | MEOX2 | -0.76 | 0.09613 | -0.26 | 0.74062 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.48 | 0.00016 | 22373864 | miR 301a is a candidate oncogene that targets the homeobox gene Gax in human hepatocellular carcinoma; This study aims to characterize the function of upregulated miR-301a in HCC and show how the downstream growth arrest-specific homeobox Gax is negatively regulated by miR-301a; The expression of miR-301a and Gax was detected using real-time PCR on HCC tissues and adjacent non-tumorous tissues; The luciferase reporter assay was used to assess Gax as a target of miR-301a; The nuclear factor κB NF-κB was measured by western blot after inhibiting miR-301a and enhancing Gax; MiR-301a was significantly upregulated and Gax was downregulated in HCC samples compared with in the matching nontumoral tissues; Inhibiting miR-301a expression caused the upregulation of Gax and repressed NF-κB expression; miR-301a is frequently upregulated in HCC and modulates NF-κB expression by negatively regulating Gax |
| 118 | hsa-miR-301a-3p | MIPOL1 | -0.76 | 0.09613 | 0.93 | 0.01754 | mirMAP | -0.27 | 1.0E-5 | NA | |
| 119 | hsa-miR-301a-3p | MTAP | -0.76 | 0.09613 | -0.36 | 0.47652 | mirMAP | -0.26 | 0.00155 | NA | |
| 120 | hsa-miR-301a-3p | MUC15 | -0.76 | 0.09613 | -0.74 | 0.62324 | MirTarget | -0.63 | 0.00936 | NA | |
| 121 | hsa-miR-301a-3p | MUC20 | -0.76 | 0.09613 | 0.85 | 0.28357 | mirMAP | -0.58 | 0 | NA | |
| 122 | hsa-miR-301a-3p | NAA50 | -0.76 | 0.09613 | 0.14 | 0.50217 | MirTarget; miRNATAP | -0.17 | 0 | NA | |
| 123 | hsa-miR-301a-3p | NFIA | -0.76 | 0.09613 | -0.29 | 0.37644 | miRNATAP | -0.27 | 0 | NA | |
| 124 | hsa-miR-301a-3p | NFIB | -0.76 | 0.09613 | -0.28 | 0.48253 | mirMAP; miRNATAP | -0.34 | 0 | NA | |
| 125 | hsa-miR-301a-3p | NLN | -0.76 | 0.09613 | 0.02 | 0.96757 | mirMAP | -0.22 | 0.00032 | NA | |
| 126 | hsa-miR-301a-3p | NOTCH2NL | -0.76 | 0.09613 | -0.52 | 0.32099 | mirMAP | -0.55 | 0 | NA | |
| 127 | hsa-miR-301a-3p | NOX4 | -0.76 | 0.09613 | 0.8 | 0.32066 | MirTarget | -0.61 | 0 | NA | |
| 128 | hsa-miR-301a-3p | NPTX1 | -0.76 | 0.09613 | -0.19 | 0.83181 | miRNATAP | -0.61 | 1.0E-5 | NA | |
| 129 | hsa-miR-301a-3p | NRBF2 | -0.76 | 0.09613 | -0.1 | 0.53207 | MirTarget; miRNATAP | -0.13 | 0 | NA | |
| 130 | hsa-miR-301a-3p | PARD3B | -0.76 | 0.09613 | 0.01 | 0.98353 | mirMAP | -0.48 | 1.0E-5 | NA | |
| 131 | hsa-miR-301a-3p | PAWR | -0.76 | 0.09613 | 0.1 | 0.76694 | mirMAP | -0.23 | 1.0E-5 | NA | |
| 132 | hsa-miR-301a-3p | PDGFRA | -0.76 | 0.09613 | -0.09 | 0.88305 | MirTarget; miRNATAP | -0.58 | 0 | NA | |
| 133 | hsa-miR-301a-3p | PELI1 | -0.76 | 0.09613 | 0.09 | 0.76823 | MirTarget; miRNATAP | -0.16 | 0.00151 | NA | |
| 134 | hsa-miR-301a-3p | PGM2L1 | -0.76 | 0.09613 | 0.65 | 0.12319 | MirTarget; miRNATAP | -0.29 | 2.0E-5 | NA | |
| 135 | hsa-miR-301a-3p | PHACTR2 | -0.76 | 0.09613 | -0.2 | 0.56161 | miRNATAP | -0.17 | 0.00192 | NA | |
| 136 | hsa-miR-301a-3p | PHF14 | -0.76 | 0.09613 | 0.15 | 0.36284 | MirTarget | -0.13 | 0 | NA | |
| 137 | hsa-miR-301a-3p | PIK3CB | -0.76 | 0.09613 | 0.27 | 0.23582 | miRNATAP | -0.13 | 0.00029 | NA | |
| 138 | hsa-miR-301a-3p | PKHD1 | -0.76 | 0.09613 | -0.13 | 0.92023 | MirTarget | -0.82 | 9.0E-5 | NA | |
| 139 | hsa-miR-301a-3p | PLEKHA2 | -0.76 | 0.09613 | -0.7 | 0.02823 | mirMAP | -0.15 | 0.00449 | NA | |
| 140 | hsa-miR-301a-3p | PLEKHA5 | -0.76 | 0.09613 | 1.1 | 0.00209 | mirMAP | -0.18 | 0.00209 | NA | |
| 141 | hsa-miR-301a-3p | PMEPA1 | -0.76 | 0.09613 | 0.39 | 0.51922 | MirTarget; miRNATAP | -0.45 | 0 | NA | |
| 142 | hsa-miR-301a-3p | POLR1A | -0.76 | 0.09613 | 0.01 | 0.97058 | mirMAP | -0.11 | 0.00842 | NA | |
| 143 | hsa-miR-301a-3p | PPARD | -0.76 | 0.09613 | -0.54 | 0.06122 | mirMAP | -0.17 | 0.00036 | NA | |
| 144 | hsa-miR-301a-3p | PPARG | -0.76 | 0.09613 | 1.21 | 0.08927 | MirTarget; miRNATAP | -0.61 | 0 | NA | |
| 145 | hsa-miR-301a-3p | PRKG1 | -0.76 | 0.09613 | 0 | 0.99515 | miRNATAP | -0.52 | 1.0E-5 | NA | |
| 146 | hsa-miR-301a-3p | PRR5L | -0.76 | 0.09613 | -0.34 | 0.49786 | miRNATAP | -0.58 | 0 | NA | |
| 147 | hsa-miR-301a-3p | PTGFRN | -0.76 | 0.09613 | 0.59 | 0.16374 | miRNATAP | -0.51 | 0 | NA | |
| 148 | hsa-miR-301a-3p | PTP4A1 | -0.76 | 0.09613 | 0.16 | 0.63317 | miRNATAP | -0.17 | 0.00226 | NA | |
| 149 | hsa-miR-301a-3p | PTPN14 | -0.76 | 0.09613 | 0.52 | 0.29139 | miRNATAP | -0.51 | 0 | NA | |
| 150 | hsa-miR-301a-3p | PTPRG | -0.76 | 0.09613 | 0.08 | 0.84464 | MirTarget; miRNATAP | -0.23 | 0.00083 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | TISSUE DEVELOPMENT | 48 | 1518 | 2.285e-11 | 1.063e-07 |
| 2 | NEGATIVE REGULATION OF CELL PROLIFERATION | 29 | 643 | 1.34e-10 | 3.117e-07 |
| 3 | CELL DEVELOPMENT | 44 | 1426 | 4.049e-10 | 6.281e-07 |
| 4 | REGULATION OF CELL PROLIFERATION | 45 | 1496 | 5.538e-10 | 6.442e-07 |
| 5 | SKELETAL SYSTEM DEVELOPMENT | 23 | 455 | 1.452e-09 | 1.352e-06 |
| 6 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 33 | 957 | 6.123e-09 | 4.748e-06 |
| 7 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 48 | 1805 | 7.53e-09 | 5.006e-06 |
| 8 | VASCULATURE DEVELOPMENT | 22 | 469 | 1.311e-08 | 7.623e-06 |
| 9 | EMBRYO DEVELOPMENT | 31 | 894 | 1.641e-08 | 8.486e-06 |
| 10 | BLOOD VESSEL MORPHOGENESIS | 19 | 364 | 2.506e-08 | 1.166e-05 |
| 11 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 28 | 788 | 5.232e-08 | 2.029e-05 |
| 12 | CIRCULATORY SYSTEM DEVELOPMENT | 28 | 788 | 5.232e-08 | 2.029e-05 |
| 13 | EPITHELIUM DEVELOPMENT | 31 | 945 | 5.79e-08 | 2.072e-05 |
| 14 | REGULATION OF CELL ADHESION | 24 | 629 | 1.392e-07 | 4.626e-05 |
| 15 | REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 10 | 100 | 1.61e-07 | 4.996e-05 |
| 16 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 36 | 1275 | 1.891e-07 | 5.175e-05 |
| 17 | ANGIOGENESIS | 16 | 293 | 1.796e-07 | 5.175e-05 |
| 18 | TUBE DEVELOPMENT | 22 | 552 | 2.281e-07 | 5.895e-05 |
| 19 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 26 | 771 | 4.455e-07 | 0.0001091 |
| 20 | NEGATIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 10 | 115 | 6.005e-07 | 0.0001397 |
| 21 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 21 | 554 | 9.79e-07 | 0.0002169 |
| 22 | CONNECTIVE TISSUE DEVELOPMENT | 12 | 194 | 1.765e-06 | 0.0003734 |
| 23 | TISSUE MORPHOGENESIS | 20 | 533 | 2.096e-06 | 0.0004063 |
| 24 | LOCOMOTION | 31 | 1114 | 2.024e-06 | 0.0004063 |
| 25 | POSITIVE REGULATION OF GENE EXPRESSION | 41 | 1733 | 2.481e-06 | 0.000444 |
| 26 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 35 | 1360 | 2.433e-06 | 0.000444 |
| 27 | CHONDROCYTE DEVELOPMENT | 5 | 21 | 2.774e-06 | 0.000478 |
| 28 | RESPONSE TO STEROID HORMONE | 19 | 497 | 2.908e-06 | 0.0004833 |
| 29 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 14 | 285 | 3.716e-06 | 0.0005962 |
| 30 | POSITIVE REGULATION OF OSSIFICATION | 8 | 84 | 4.151e-06 | 0.0006438 |
| 31 | POSITIVE REGULATION OF OSTEOBLAST DIFFERENTIATION | 7 | 60 | 4.297e-06 | 0.000645 |
| 32 | RESPONSE TO WOUNDING | 20 | 563 | 4.772e-06 | 0.0006938 |
| 33 | WOUND HEALING | 18 | 470 | 5.223e-06 | 0.0007365 |
| 34 | CARTILAGE DEVELOPMENT | 10 | 147 | 5.627e-06 | 0.0007701 |
| 35 | CELLULAR RESPONSE TO STEROID HORMONE STIMULUS | 12 | 218 | 5.901e-06 | 0.0007845 |
| 36 | ORGAN MORPHOGENESIS | 25 | 841 | 7.062e-06 | 0.0009128 |
| 37 | EMBRYONIC SKELETAL SYSTEM DEVELOPMENT | 9 | 122 | 8.64e-06 | 0.001087 |
| 38 | EMBRYONIC MORPHOGENESIS | 19 | 539 | 9.279e-06 | 0.001136 |
| 39 | MORPHOGENESIS OF AN EPITHELIUM | 16 | 400 | 1.035e-05 | 0.001196 |
| 40 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 26 | 917 | 1.054e-05 | 0.001196 |
| 41 | DEVELOPMENTAL MATURATION | 11 | 193 | 1.046e-05 | 0.001196 |
| 42 | POSITIVE REGULATION OF CELL COMMUNICATION | 36 | 1532 | 1.31e-05 | 0.001452 |
| 43 | TUBE MORPHOGENESIS | 14 | 323 | 1.541e-05 | 0.001668 |
| 44 | REGULATION OF SMOOTH MUSCLE CELL MIGRATION | 6 | 49 | 1.6e-05 | 0.001692 |
| 45 | TAXIS | 17 | 464 | 1.703e-05 | 0.001761 |
| 46 | NEGATIVE REGULATION OF GENE EXPRESSION | 35 | 1493 | 1.85e-05 | 0.001793 |
| 47 | RESPONSE TO LIPID | 25 | 888 | 1.777e-05 | 0.001793 |
| 48 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 27 | 1004 | 1.812e-05 | 0.001793 |
| 49 | POSITIVE REGULATION OF CELL ADHESION | 15 | 376 | 2.028e-05 | 0.001926 |
| 50 | NEGATIVE REGULATION OF CELL COMMUNICATION | 30 | 1192 | 2.125e-05 | 0.001977 |
| 51 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 35 | 1517 | 2.583e-05 | 0.002348 |
| 52 | MULTI MULTICELLULAR ORGANISM PROCESS | 11 | 213 | 2.624e-05 | 0.002348 |
| 53 | POSITIVE REGULATION OF CELL PROLIFERATION | 23 | 814 | 3.713e-05 | 0.003211 |
| 54 | PROTEOGLYCAN METABOLIC PROCESS | 7 | 83 | 3.726e-05 | 0.003211 |
| 55 | EPITHELIAL CELL DIFFERENTIATION | 17 | 495 | 3.854e-05 | 0.00326 |
| 56 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 36 | 1618 | 4.188e-05 | 0.003479 |
| 57 | NEGATIVE REGULATION OF WOUND HEALING | 6 | 58 | 4.263e-05 | 0.00348 |
| 58 | CELLULAR RESPONSE TO LIPID | 16 | 457 | 5.208e-05 | 0.003943 |
| 59 | POSITIVE REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 6 | 60 | 5.177e-05 | 0.003943 |
| 60 | CHONDROCYTE DIFFERENTIATION | 6 | 60 | 5.177e-05 | 0.003943 |
| 61 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 9 | 153 | 5.254e-05 | 0.003943 |
| 62 | TISSUE REMODELING | 7 | 87 | 5.053e-05 | 0.003943 |
| 63 | RESPONSE TO ENDOGENOUS STIMULUS | 33 | 1450 | 5.911e-05 | 0.004366 |
| 64 | NEGATIVE REGULATION OF RESPONSE TO WOUNDING | 9 | 156 | 6.111e-05 | 0.004443 |
| 65 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 16 | 465 | 6.39e-05 | 0.004574 |
| 66 | HEART DEVELOPMENT | 16 | 466 | 6.553e-05 | 0.00462 |
| 67 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 32 | 1395 | 6.68e-05 | 0.004639 |
| 68 | REGULATION OF WOUND HEALING | 8 | 126 | 8.045e-05 | 0.005505 |
| 69 | REGULATION OF DNA BIOSYNTHETIC PROCESS | 7 | 94 | 8.292e-05 | 0.005592 |
| 70 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 20 | 689 | 8.468e-05 | 0.005629 |
| 71 | REGULATION OF CELL CELL ADHESION | 14 | 380 | 9.015e-05 | 0.005908 |
| 72 | REGULATION OF EPITHELIAL CELL MIGRATION | 9 | 166 | 9.862e-05 | 0.006268 |
| 73 | NEURON PROJECTION GUIDANCE | 10 | 205 | 9.834e-05 | 0.006268 |
| 74 | REPRODUCTION | 30 | 1297 | 9.969e-05 | 0.006268 |
| 75 | REGULATION OF CELL DIFFERENTIATION | 33 | 1492 | 0.0001021 | 0.006335 |
| 76 | CELL MATURATION | 8 | 131 | 0.0001057 | 0.006472 |
| 77 | MEMORY | 7 | 98 | 0.000108 | 0.006525 |
| 78 | APPENDAGE DEVELOPMENT | 9 | 169 | 0.0001131 | 0.00666 |
| 79 | LIMB DEVELOPMENT | 9 | 169 | 0.0001131 | 0.00666 |
| 80 | RESPONSE TO MECHANICAL STIMULUS | 10 | 210 | 0.0001199 | 0.006976 |
| 81 | NEURON PROJECTION DEVELOPMENT | 17 | 545 | 0.0001249 | 0.007172 |
| 82 | SPROUTING ANGIOGENESIS | 5 | 45 | 0.0001342 | 0.007524 |
| 83 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 18 | 602 | 0.000134 | 0.007524 |
| 84 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 8 | 138 | 0.0001518 | 0.007798 |
| 85 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 39 | 1929 | 0.0001525 | 0.007798 |
| 86 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 35 | 1656 | 0.0001503 | 0.007798 |
| 87 | CELL MOTILITY | 22 | 835 | 0.0001516 | 0.007798 |
| 88 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 37 | 1791 | 0.0001505 | 0.007798 |
| 89 | RESPONSE TO HORMONE | 23 | 893 | 0.000149 | 0.007798 |
| 90 | LOCALIZATION OF CELL | 22 | 835 | 0.0001516 | 0.007798 |
| 91 | POSITIVE REGULATION OF EPITHELIAL CELL MIGRATION | 7 | 103 | 0.0001476 | 0.007798 |
| 92 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 20 | 724 | 0.0001645 | 0.008092 |
| 93 | NEURON PROJECTION MORPHOGENESIS | 14 | 402 | 0.0001624 | 0.008092 |
| 94 | CELLULAR COMPONENT MORPHOGENESIS | 23 | 900 | 0.000167 | 0.008092 |
| 95 | REGULATION OF HOMOTYPIC CELL CELL ADHESION | 12 | 307 | 0.0001666 | 0.008092 |
| 96 | GLYCOPROTEIN METABOLIC PROCESS | 13 | 353 | 0.0001614 | 0.008092 |
| 97 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 35 | 1672 | 0.0001809 | 0.008678 |
| 98 | REPRODUCTIVE SYSTEM DEVELOPMENT | 14 | 408 | 0.0001892 | 0.008982 |
| 99 | RESPONSE TO KETONE | 9 | 182 | 0.0001978 | 0.009208 |
| 100 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 16 | 513 | 0.0001979 | 0.009208 |
| 101 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 25 | 1036 | 0.0002088 | 0.009525 |
| 102 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 25 | 1036 | 0.0002088 | 0.009525 |
| 103 | REGULATION OF RESPONSE TO WOUNDING | 14 | 413 | 0.0002143 | 0.009683 |
| 104 | RESPONSE TO PROGESTERONE | 5 | 50 | 0.0002226 | 0.009958 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | GOLGI APPARATUS | 37 | 1445 | 1.35e-06 | 0.0007882 |
Over-represented Pathway
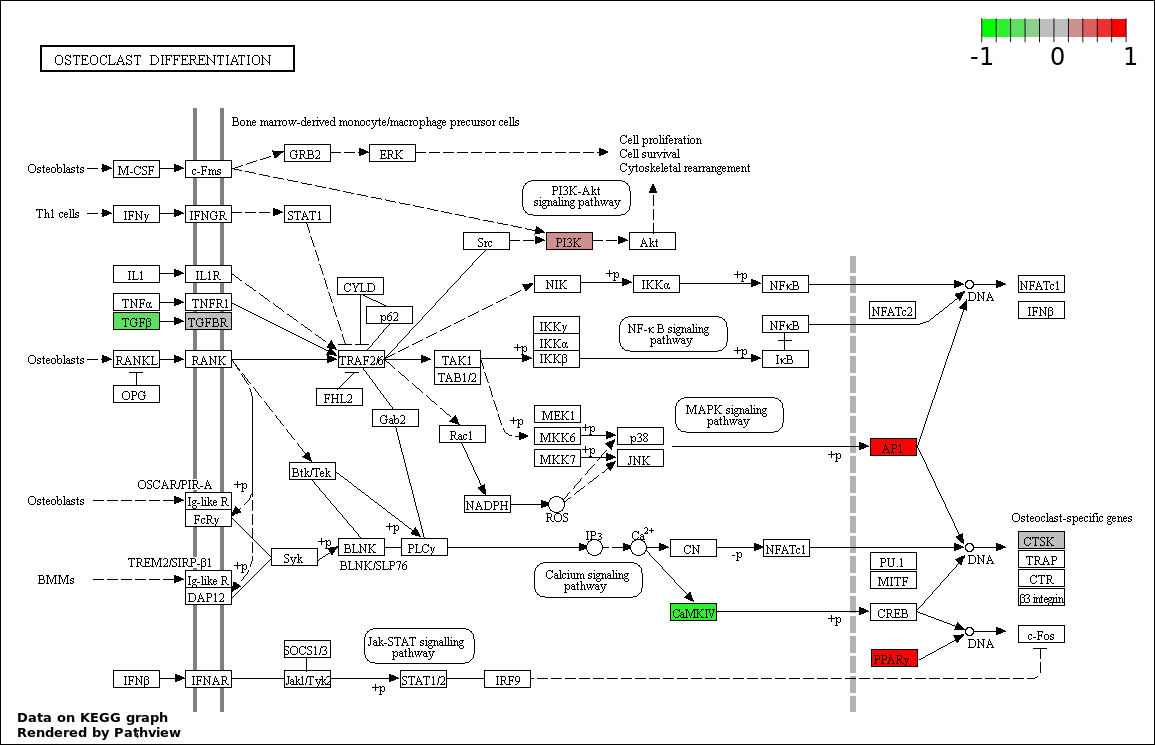
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04380_Osteoclast_differentiation | 7 | 128 | 0.0005583 | 0.1005 | |
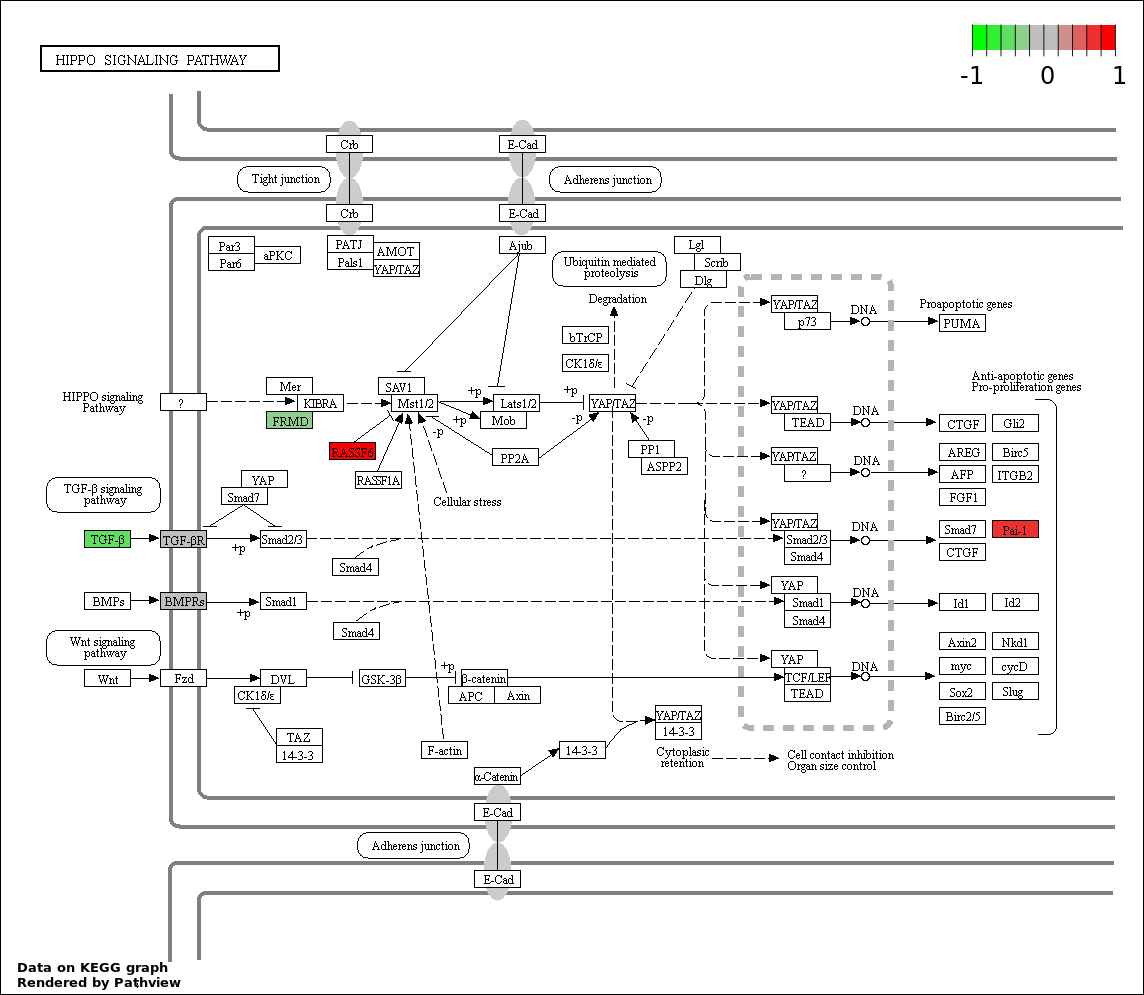
| 2 | hsa04390_Hippo_signaling_pathway | 7 | 154 | 0.001648 | 0.1483 | |
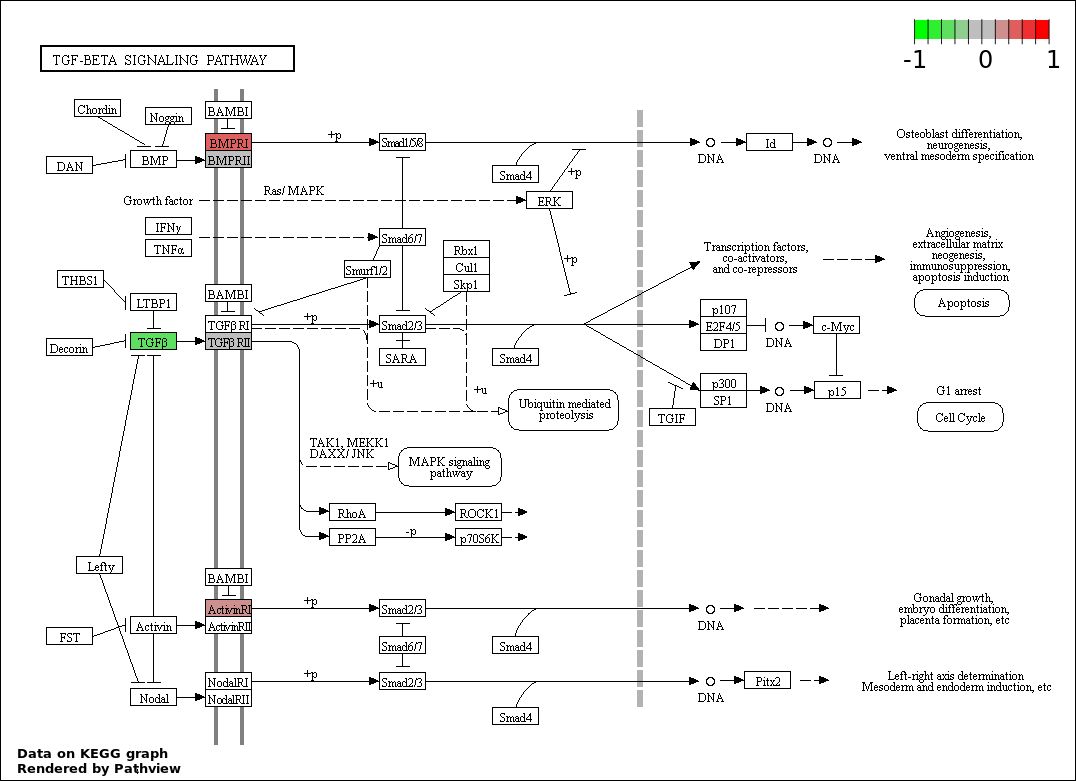
| 3 | hsa04350_TGF.beta_signaling_pathway | 5 | 85 | 0.002522 | 0.1513 | |
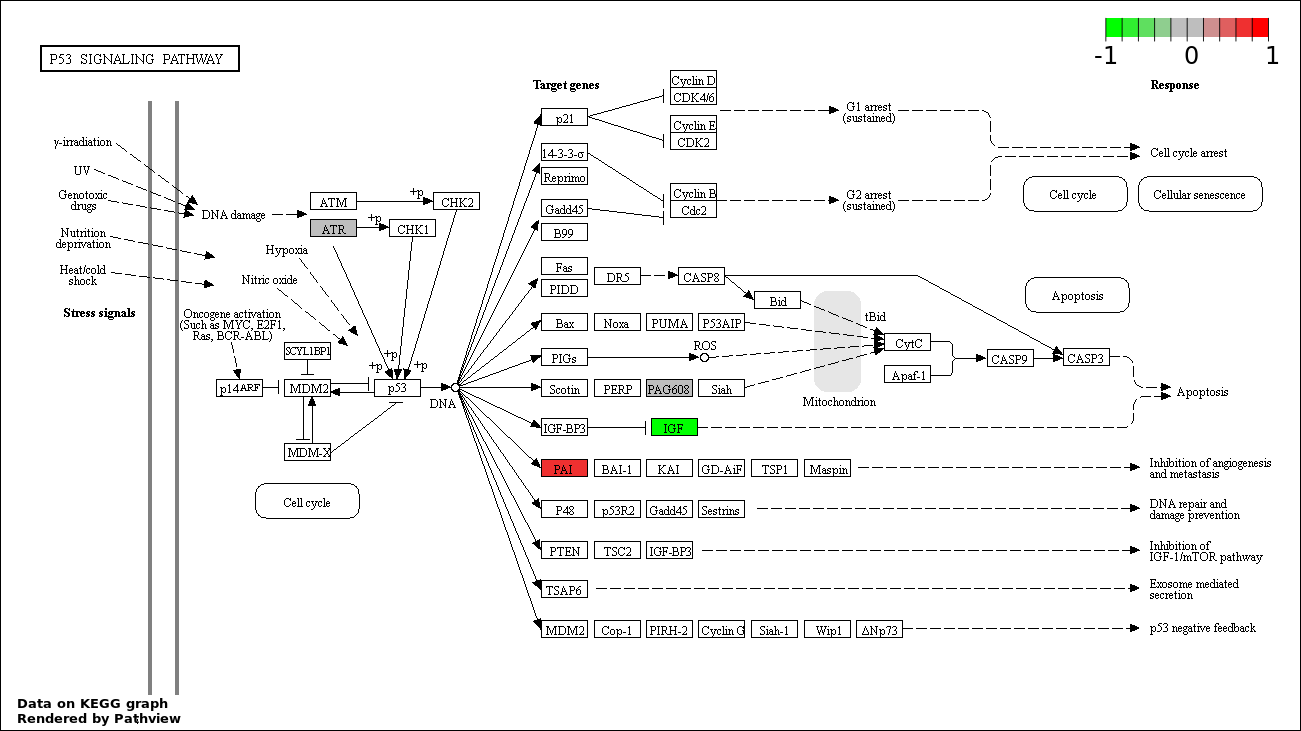
| 4 | hsa04115_p53_signaling_pathway | 4 | 69 | 0.007162 | 0.3223 | |
| 5 | hsa04210_Apoptosis | 4 | 89 | 0.01707 | 0.5314 | |
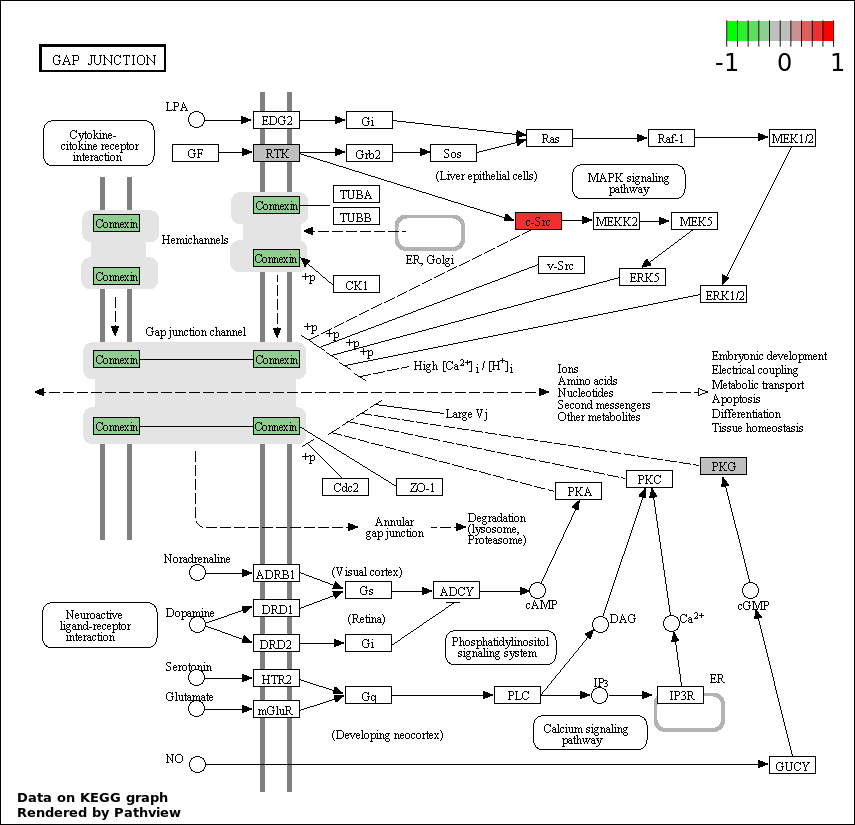
| 6 | hsa04540_Gap_junction | 4 | 90 | 0.01771 | 0.5314 | |
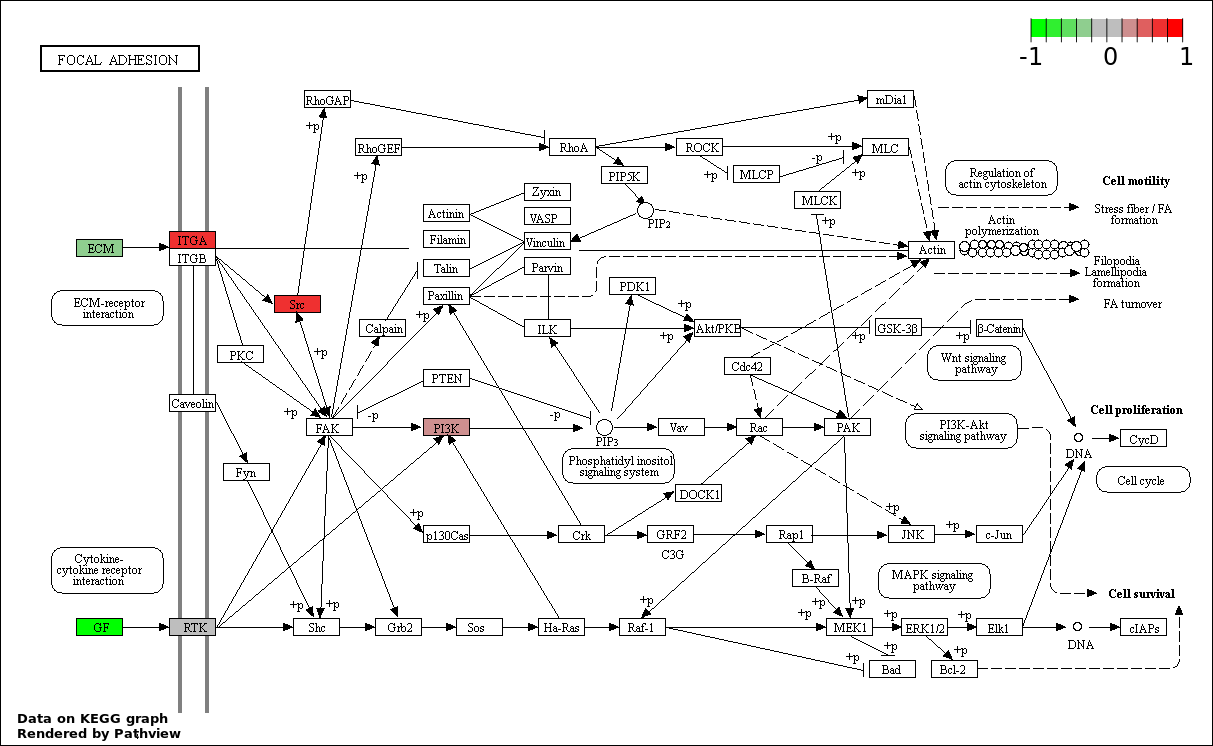
| 7 | hsa04510_Focal_adhesion | 6 | 200 | 0.02398 | 0.5751 | |
| 8 | hsa04144_Endocytosis | 6 | 203 | 0.02556 | 0.5751 | |
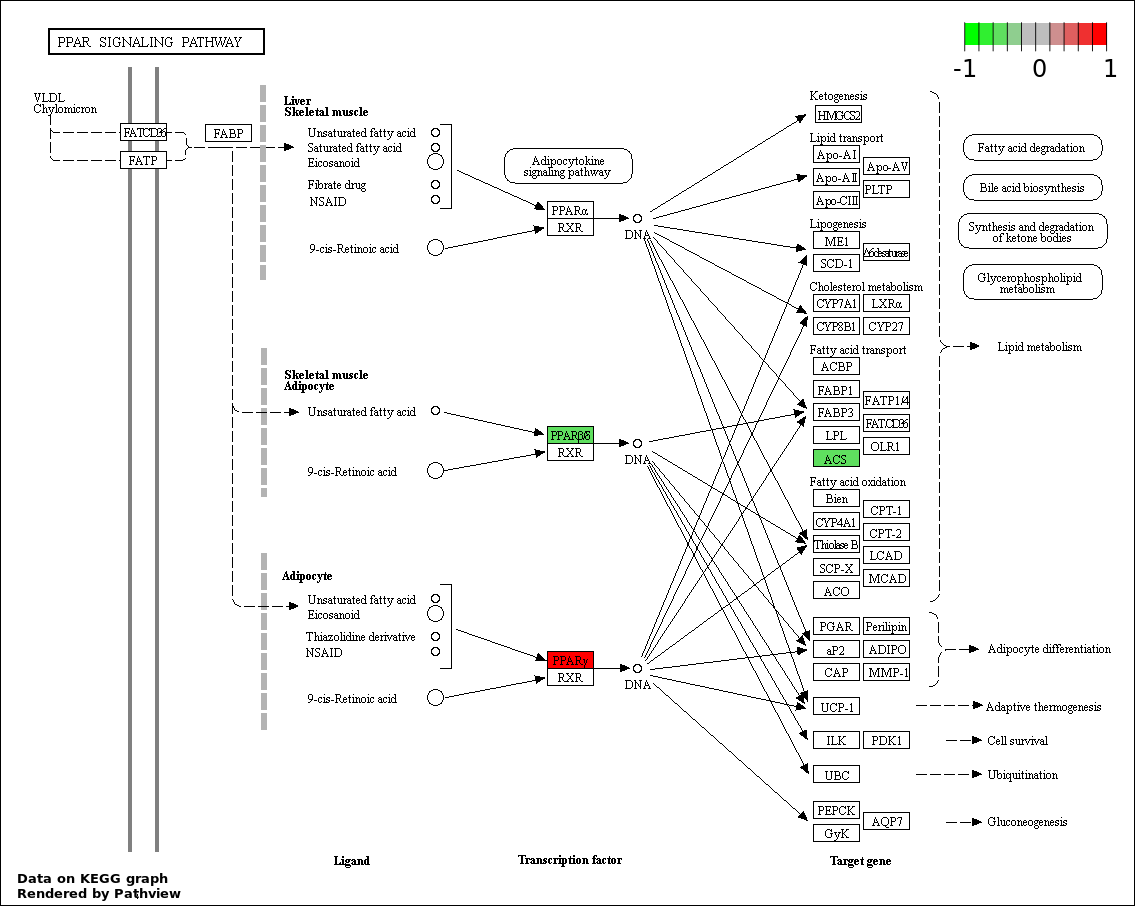
| 9 | hsa03320_PPAR_signaling_pathway | 3 | 70 | 0.04245 | 0.7635 | |
| 10 | hsa00512_Mucin_type_O.Glycan_biosynthesis | 2 | 30 | 0.04318 | 0.7635 | |
| 11 | hsa04014_Ras_signaling_pathway | 6 | 236 | 0.04759 | 0.7635 | |
| 12 | hsa04110_Cell_cycle | 4 | 128 | 0.05377 | 0.7635 | |
| 13 | hsa04360_Axon_guidance | 4 | 130 | 0.05631 | 0.7635 | |
| 14 | hsa04512_ECM.receptor_interaction | 3 | 85 | 0.06806 | 0.7635 | |
| 15 | hsa04012_ErbB_signaling_pathway | 3 | 87 | 0.07191 | 0.7635 | |
| 16 | hsa04914_Progesterone.mediated_oocyte_maturation | 3 | 87 | 0.07191 | 0.7635 | |
| 17 | hsa00600_Sphingolipid_metabolism | 2 | 40 | 0.0721 | 0.7635 | |
| 18 | hsa04960_Aldosterone.regulated_sodium_reabsorption | 2 | 42 | 0.07847 | 0.7847 | |
| 19 | hsa04310_Wnt_signaling_pathway | 4 | 151 | 0.08672 | 0.8037 | |
| 20 | hsa04810_Regulation_of_actin_cytoskeleton | 5 | 214 | 0.0893 | 0.8037 | |
| 21 | hsa04150_mTOR_signaling_pathway | 2 | 52 | 0.1127 | 0.9656 | |
| 22 | hsa04020_Calcium_signaling_pathway | 4 | 177 | 0.1331 | 1 | |
| 23 | hsa04610_Complement_and_coagulation_cascades | 2 | 69 | 0.1771 | 1 | |
| 24 | hsa04730_Long.term_depression | 2 | 70 | 0.1811 | 1 | |
| 25 | hsa04520_Adherens_junction | 2 | 73 | 0.1931 | 1 | |
| 26 | hsa04151_PI3K_AKT_signaling_pathway | 6 | 351 | 0.1937 | 1 | |
| 27 | hsa04370_VEGF_signaling_pathway | 2 | 76 | 0.2051 | 1 | |
| 28 | hsa04974_Protein_digestion_and_absorption | 2 | 81 | 0.2254 | 1 | |
| 29 | hsa04630_Jak.STAT_signaling_pathway | 3 | 155 | 0.2454 | 1 | |
| 30 | hsa00240_Pyrimidine_metabolism | 2 | 99 | 0.299 | 1 | |
| 31 | hsa04620_Toll.like_receptor_signaling_pathway | 2 | 102 | 0.3112 | 1 | |
| 32 | hsa04114_Oocyte_meiosis | 2 | 114 | 0.3596 | 1 | |
| 33 | hsa04722_Neurotrophin_signaling_pathway | 2 | 127 | 0.4105 | 1 | |
| 34 | hsa00230_Purine_metabolism | 2 | 162 | 0.5365 | 1 | |
| 35 | hsa04010_MAPK_signaling_pathway | 3 | 268 | 0.5708 | 1 |