This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-148a-5p | CD44 | 1.16 | 0.00015 | -0.8 | 0.04276 | mirMAP | -0.18 | 0.00281 | 23861222 | In MHCC97H and MHCC97L cells over-expression of miR-148a blocked the EMT process attenuated the expression of CD90 and CD44 biomarkers for liver cancer stem cells and inhibited their migratory capacity |
| 2 | hsa-miR-15b-5p | CD44 | 1.57 | 0 | -0.8 | 0.04276 | miRNAWalker2 validate | -0.45 | 0 | NA | |
| 3 | hsa-miR-16-5p | CD44 | 1.76 | 0 | -0.8 | 0.04276 | miRNAWalker2 validate | -0.23 | 0.00779 | NA | |
| 4 | hsa-miR-181a-2-3p | CD44 | 2.19 | 0 | -0.8 | 0.04276 | MirTarget | -0.23 | 0.00019 | NA | |
| 5 | hsa-miR-185-5p | CD44 | 2.34 | 0 | -0.8 | 0.04276 | MirTarget | -0.37 | 1.0E-5 | NA | |
| 6 | hsa-miR-28-5p | CD44 | 0.23 | 0.07429 | -0.8 | 0.04276 | miRanda | -0.92 | 0 | NA | |
| 7 | hsa-miR-30e-3p | CD44 | -0.22 | 0.23538 | -0.8 | 0.04276 | mirMAP | -0.79 | 0 | NA | |
| 8 | hsa-miR-320a | CD44 | 0.44 | 0.03902 | -0.8 | 0.04276 | miRNAWalker2 validate | -0.27 | 0.00272 | NA | |
| 9 | hsa-miR-326 | CD44 | 0.72 | 0.06103 | -0.8 | 0.04276 | miRanda | -0.35 | 0 | NA | |
| 10 | hsa-miR-328-3p | CD44 | 0.6 | 0.01386 | -0.8 | 0.04276 | miRNAWalker2 validate; miRTarBase | -0.39 | 0 | NA | |
| 11 | hsa-miR-330-3p | CD44 | 0.88 | 0.00074 | -0.8 | 0.04276 | miRNAWalker2 validate; miRTarBase | -0.4 | 0 | NA | |
| 12 | hsa-miR-330-5p | CD44 | 1.15 | 0 | -0.8 | 0.04276 | miRanda | -0.23 | 0.00452 | NA | |
| 13 | hsa-miR-33a-3p | CD44 | 1.73 | 0 | -0.8 | 0.04276 | mirMAP | -0.39 | 0 | NA | |
| 14 | hsa-miR-34a-5p | CD44 | 1.9 | 0 | -0.8 | 0.04276 | miRNAWalker2 validate; miRTarBase | -0.28 | 0.00023 | 25572695; 24423412; 25551284; 25044638; 21240262; 26231042; 27497057; 23314380 | The c-Myc and CD44 were confirmed as direct targets of miR-34a in EJ cell apoptosis induced by PRE;Furthermore we identified CD44 as being targeted by miR-34a in MIBC cells following cisplatin treatment and increased CD44 expression could efficiently reverse the effect of miR-34a on MIBC cell proliferation colongenic potential and chemosensitivity; Cisplatin-based chemotherapy induced demethylation of miR-34a promoter and increased miR-34a expression which in turn sensitized MIBC cells to cisplatin and decreased the tumorigenicity and proliferation of cancer cells that by reducing the production of CD44;MicroRNA 34a functions as an anti metastatic microRNA and suppresses angiogenesis in bladder cancer by directly targeting CD44; In this study we focus on it that microRNA-34a functions as an anti-metastatic microRNA and suppress angiogenesis in bladder cancer by directly targeting CD44; Our study defines a major metastasis and angiogenesis suppressive role for mir-34a a microRNA functions as a tumor suppressor in bladder cancer by directly targeting CD44 which would be helpful as a therapeutic approach to block bladder cancer metastasis;Nanocomplex-assisted delivery of miR-34a induces cell apoptosis and suppresses migration proliferation and tumor growth of breast cancer cells via targeting CD44 and a Notch-1-signaling pathway;The microRNA miR 34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44; We identified and validated CD44 as a direct and functional target of miR-34a and found that CD44 knockdown phenocopied miR-34a overexpression in inhibiting prostate cancer regeneration and metastasis;Registered report: the microRNA miR 34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44; Tumors with exogenous miR-34a showed reduced levels of CD44 expression Figure 4A and mutation of two putative miR-34a binding sites in the CD33 3' UTR partially abrogated signal repression in a luciferase assay Figure 4D;Nanovesicle mediated systemic delivery of microRNA 34a for CD44 overexpressing gastric cancer stem cell therapy; MicroRNA-34a miR-34a is a promising candidate for CD44 repression-based cancer therapy as it has been reported to inhibit proliferation metastasis and survival of CD44-positive CSCs; Here we used nanovesicles containing PLI/miR complexes NVs/miR to systemically deliver miR-34a and induce miR-34a-triggered CD44 suppression in orthotopically and subcutaneously implanted tumors in nude mice;miR 34a inhibits the metastasis of osteosarcoma cells by repressing the expression of CD44; The ectopic overexpression of miR-34a significantly inhibited the migration and invasive ability of osteosarcoma cells by repressing the expression of CD44; These data suggest that miR-34a plays a tumor suppressor role in the metastasis of osteosarcoma cells by repressing the expression of CD44; Therefore it can be concluded that through the inhibition of CD44 expression levels miR-34a plays a significant role in the migration and invasion of osteosarcoma cells |
| 15 | hsa-miR-421 | CD44 | 1.18 | 1.0E-5 | -0.8 | 0.04276 | miRanda | -0.37 | 0 | NA | |
| 16 | hsa-miR-532-3p | CD44 | 1.97 | 0 | -0.8 | 0.04276 | miRNATAP | -0.44 | 0 | NA | |
| 17 | hsa-miR-590-3p | CD44 | 2.59 | 0 | -0.8 | 0.04276 | MirTarget; miRanda; miRNATAP | -0.17 | 0.01369 | NA | |
| 18 | hsa-miR-744-5p | CD44 | 1.48 | 0 | -0.8 | 0.04276 | miRNAWalker2 validate | -0.46 | 0 | NA | |
| 19 | hsa-miR-942-5p | CD44 | 2.35 | 0 | -0.8 | 0.04276 | MirTarget | -0.13 | 0.03557 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | REGULATION OF CELL DEATH | 36 | 1472 | 2.999e-17 | 1.395e-13 |
| 2 | REGULATION OF CELL PROLIFERATION | 35 | 1496 | 3.827e-16 | 8.904e-13 |
| 3 | POSITIVE REGULATION OF GENE EXPRESSION | 35 | 1733 | 3.292e-14 | 5.106e-11 |
| 4 | RESPONSE TO ENDOGENOUS STIMULUS | 32 | 1450 | 5.472e-14 | 6.366e-11 |
| 5 | NEGATIVE REGULATION OF CELL DEATH | 24 | 872 | 1.679e-12 | 1.562e-09 |
| 6 | NEGATIVE REGULATION OF CELL PROLIFERATION | 21 | 643 | 2.135e-12 | 1.656e-09 |
| 7 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 32 | 1784 | 1.474e-11 | 9.798e-09 |
| 8 | RESPONSE TO HORMONE | 23 | 893 | 2.044e-11 | 1.189e-08 |
| 9 | NEUROGENESIS | 28 | 1402 | 3.48e-11 | 1.799e-08 |
| 10 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 29 | 1517 | 4.01e-11 | 1.866e-08 |
| 11 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 31 | 1805 | 1.036e-10 | 4.383e-08 |
| 12 | NEGATIVE REGULATION OF GENE EXPRESSION | 28 | 1493 | 1.503e-10 | 5.379e-08 |
| 13 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 28 | 1492 | 1.48e-10 | 5.379e-08 |
| 14 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 31 | 1848 | 1.87e-10 | 6.215e-08 |
| 15 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 22 | 917 | 2.35e-10 | 6.431e-08 |
| 16 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 23 | 1008 | 2.266e-10 | 6.431e-08 |
| 17 | NEGATIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 20 | 740 | 2.255e-10 | 6.431e-08 |
| 18 | GLAND DEVELOPMENT | 15 | 395 | 5.465e-10 | 1.413e-07 |
| 19 | REGULATION OF CELL DIFFERENTIATION | 27 | 1492 | 7.758e-10 | 1.805e-07 |
| 20 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 26 | 1381 | 7.541e-10 | 1.805e-07 |
| 21 | RESPONSE TO ABIOTIC STIMULUS | 22 | 1024 | 1.853e-09 | 4.107e-07 |
| 22 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 23 | 1142 | 2.538e-09 | 5.368e-07 |
| 23 | POSITIVE REGULATION OF CELL DEATH | 17 | 605 | 3.411e-09 | 6.9e-07 |
| 24 | REPRODUCTIVE SYSTEM DEVELOPMENT | 14 | 408 | 7.891e-09 | 1.53e-06 |
| 25 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 19 | 823 | 9.036e-09 | 1.682e-06 |
| 26 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 24 | 1395 | 2.295e-08 | 4.108e-06 |
| 27 | RESPONSE TO NUTRIENT | 10 | 191 | 2.493e-08 | 4.296e-06 |
| 28 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 21 | 1087 | 2.954e-08 | 4.909e-06 |
| 29 | RESPONSE TO VITAMIN A | 5 | 20 | 3.186e-08 | 5.111e-06 |
| 30 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 14 | 465 | 4.069e-08 | 6.107e-06 |
| 31 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 26 | 1672 | 4.026e-08 | 6.107e-06 |
| 32 | REGULATION OF GROWTH | 16 | 633 | 4.553e-08 | 6.62e-06 |
| 33 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 21 | 1135 | 6.18e-08 | 8.662e-06 |
| 34 | REGULATION OF PROTEIN MODIFICATION PROCESS | 26 | 1710 | 6.33e-08 | 8.662e-06 |
| 35 | RESPONSE TO ESTROGEN | 10 | 218 | 8.68e-08 | 1.154e-05 |
| 36 | RESPONSE TO STEROID HORMONE | 14 | 497 | 9.254e-08 | 1.196e-05 |
| 37 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 17 | 771 | 1.18e-07 | 1.484e-05 |
| 38 | NEURON DIFFERENTIATION | 18 | 874 | 1.319e-07 | 1.615e-05 |
| 39 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 18 | 876 | 1.365e-07 | 1.628e-05 |
| 40 | CELL DEVELOPMENT | 23 | 1426 | 1.558e-07 | 1.813e-05 |
| 41 | RESPONSE TO EXTRACELLULAR STIMULUS | 13 | 441 | 1.652e-07 | 1.851e-05 |
| 42 | RESPONSE TO LIPID | 18 | 888 | 1.67e-07 | 1.851e-05 |
| 43 | EMBRYO DEVELOPMENT | 18 | 894 | 1.846e-07 | 1.997e-05 |
| 44 | CELLULAR RESPONSE TO HORMONE STIMULUS | 14 | 552 | 3.317e-07 | 3.508e-05 |
| 45 | CELL MOTILITY | 17 | 835 | 3.64e-07 | 3.682e-05 |
| 46 | LOCALIZATION OF CELL | 17 | 835 | 3.64e-07 | 3.682e-05 |
| 47 | CELLULAR RESPONSE TO OXYGEN LEVELS | 8 | 143 | 4.037e-07 | 3.997e-05 |
| 48 | RESPONSE TO ESTRADIOL | 8 | 146 | 4.732e-07 | 4.587e-05 |
| 49 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 25 | 1791 | 6.171e-07 | 5.86e-05 |
| 50 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 13 | 498 | 6.57e-07 | 6.072e-05 |
| 51 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 17 | 872 | 6.656e-07 | 6.072e-05 |
| 52 | CELL DEATH | 18 | 1001 | 9.6e-07 | 8.281e-05 |
| 53 | CELLULAR RESPONSE TO STEROID HORMONE STIMULUS | 9 | 218 | 9.61e-07 | 8.281e-05 |
| 54 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 14 | 602 | 9.341e-07 | 8.281e-05 |
| 55 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 18 | 1004 | 1.002e-06 | 8.478e-05 |
| 56 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 16 | 799 | 1.045e-06 | 8.53e-05 |
| 57 | FOREBRAIN DEVELOPMENT | 11 | 357 | 1.029e-06 | 8.53e-05 |
| 58 | RESPONSE TO ALCOHOL | 11 | 362 | 1.179e-06 | 9.458e-05 |
| 59 | INTRACELLULAR RECEPTOR SIGNALING PATHWAY | 8 | 168 | 1.371e-06 | 0.0001081 |
| 60 | DIENCEPHALON DEVELOPMENT | 6 | 77 | 1.769e-06 | 0.0001372 |
| 61 | REGULATION OF CELL DEVELOPMENT | 16 | 836 | 1.884e-06 | 0.0001437 |
| 62 | ERBB SIGNALING PATHWAY | 6 | 79 | 2.058e-06 | 0.0001545 |
| 63 | POSITIVE REGULATION OF CELL COMMUNICATION | 22 | 1532 | 2.174e-06 | 0.0001602 |
| 64 | STEROID HORMONE MEDIATED SIGNALING PATHWAY | 7 | 125 | 2.204e-06 | 0.0001602 |
| 65 | RESPONSE TO OXYGEN LEVELS | 10 | 311 | 2.25e-06 | 0.0001611 |
| 66 | RESPONSE TO NITROGEN COMPOUND | 16 | 859 | 2.674e-06 | 0.0001885 |
| 67 | RESPONSE TO EXTERNAL STIMULUS | 24 | 1821 | 3.04e-06 | 0.0002111 |
| 68 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 17 | 983 | 3.404e-06 | 0.0002329 |
| 69 | RESPONSE TO INCREASED OXYGEN LEVELS | 4 | 23 | 3.947e-06 | 0.0002624 |
| 70 | RESPONSE TO HYPEROXIA | 4 | 23 | 3.947e-06 | 0.0002624 |
| 71 | PLACENTA DEVELOPMENT | 7 | 138 | 4.259e-06 | 0.0002791 |
| 72 | LOCOMOTION | 18 | 1114 | 4.355e-06 | 0.0002815 |
| 73 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 14 | 689 | 4.507e-06 | 0.0002873 |
| 74 | REGULATION OF BODY FLUID LEVELS | 12 | 506 | 4.911e-06 | 0.0003088 |
| 75 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 15 | 801 | 5.346e-06 | 0.0003317 |
| 76 | EPITHELIAL CELL APOPTOTIC PROCESS | 4 | 25 | 5.598e-06 | 0.0003427 |
| 77 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 13 | 609 | 6.023e-06 | 0.000364 |
| 78 | HEAD DEVELOPMENT | 14 | 709 | 6.251e-06 | 0.0003729 |
| 79 | MYELOID LEUKOCYTE DIFFERENTIATION | 6 | 96 | 6.434e-06 | 0.000379 |
| 80 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 18 | 1152 | 6.926e-06 | 0.0003996 |
| 81 | REGULATION OF CHROMOSOME ORGANIZATION | 9 | 278 | 7.042e-06 | 0.0003996 |
| 82 | TISSUE DEVELOPMENT | 21 | 1518 | 6.97e-06 | 0.0003996 |
| 83 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 19 | 1275 | 7.284e-06 | 0.0004035 |
| 84 | RESPONSE TO VITAMIN | 6 | 98 | 7.25e-06 | 0.0004035 |
| 85 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 22 | 1656 | 7.674e-06 | 0.0004201 |
| 86 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 14 | 724 | 7.93e-06 | 0.0004291 |
| 87 | REGULATION OF CHROMATIN ORGANIZATION | 7 | 152 | 8.054e-06 | 0.0004298 |
| 88 | REGULATION OF BINDING | 9 | 283 | 8.128e-06 | 0.0004298 |
| 89 | REGULATION OF CATABOLIC PROCESS | 14 | 731 | 8.842e-06 | 0.0004623 |
| 90 | CHROMATIN MODIFICATION | 12 | 539 | 9.298e-06 | 0.0004754 |
| 91 | REPRODUCTION | 19 | 1297 | 9.296e-06 | 0.0004754 |
| 92 | HORMONE MEDIATED SIGNALING PATHWAY | 7 | 158 | 1.038e-05 | 0.0005247 |
| 93 | REGULATION OF FAT CELL DIFFERENTIATION | 6 | 106 | 1.14e-05 | 0.0005702 |
| 94 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 14 | 750 | 1.181e-05 | 0.0005844 |
| 95 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 12 | 554 | 1.224e-05 | 0.0005993 |
| 96 | UROGENITAL SYSTEM DEVELOPMENT | 9 | 299 | 1.262e-05 | 0.0006116 |
| 97 | POSITIVE REGULATION OF MAPK CASCADE | 11 | 470 | 1.419e-05 | 0.000667 |
| 98 | WOUND HEALING | 11 | 470 | 1.419e-05 | 0.000667 |
| 99 | NEURON MIGRATION | 6 | 110 | 1.409e-05 | 0.000667 |
| 100 | RESPONSE TO WOUNDING | 12 | 563 | 1.436e-05 | 0.0006683 |
| 101 | CHROMATIN ORGANIZATION | 13 | 663 | 1.49e-05 | 0.0006865 |
| 102 | RESPONSE TO INORGANIC SUBSTANCE | 11 | 479 | 1.692e-05 | 0.0007644 |
| 103 | REGULATION OF CELL GROWTH | 10 | 391 | 1.677e-05 | 0.0007644 |
| 104 | IN UTERO EMBRYONIC DEVELOPMENT | 9 | 311 | 1.724e-05 | 0.0007715 |
| 105 | REGULATION OF ERK1 AND ERK2 CASCADE | 8 | 238 | 1.783e-05 | 0.0007903 |
| 106 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 19 | 1360 | 1.813e-05 | 0.000796 |
| 107 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 21 | 1618 | 1.843e-05 | 0.0008014 |
| 108 | NEURON DEVELOPMENT | 13 | 687 | 2.165e-05 | 0.0009327 |
| 109 | INTRACELLULAR STEROID HORMONE RECEPTOR SIGNALING PATHWAY | 5 | 71 | 2.21e-05 | 0.0009366 |
| 110 | FOREBRAIN NEURON FATE COMMITMENT | 3 | 12 | 2.214e-05 | 0.0009366 |
| 111 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 16 | 1021 | 2.331e-05 | 0.0009771 |
| 112 | ACUTE INFLAMMATORY RESPONSE | 5 | 73 | 2.53e-05 | 0.001051 |
| 113 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 23 | 1929 | 2.642e-05 | 0.001088 |
| 114 | CELLULAR RESPONSE TO NITROGEN COMPOUND | 11 | 505 | 2.752e-05 | 0.001123 |
| 115 | HEPATOCYTE APOPTOTIC PROCESS | 3 | 13 | 2.869e-05 | 0.001157 |
| 116 | LIPID METABOLIC PROCESS | 17 | 1158 | 2.885e-05 | 0.001157 |
| 117 | REGULATION OF PROTEOLYSIS | 13 | 711 | 3.094e-05 | 0.001231 |
| 118 | POSITIVE REGULATION OF BINDING | 6 | 127 | 3.188e-05 | 0.001257 |
| 119 | MYELOID CELL DIFFERENTIATION | 7 | 189 | 3.3e-05 | 0.001291 |
| 120 | STEM CELL DIFFERENTIATION | 7 | 190 | 3.414e-05 | 0.001324 |
| 121 | ERBB2 SIGNALING PATHWAY | 4 | 39 | 3.459e-05 | 0.00133 |
| 122 | REGULATION OF NEURON APOPTOTIC PROCESS | 7 | 192 | 3.649e-05 | 0.001392 |
| 123 | EPITHELIUM DEVELOPMENT | 15 | 945 | 3.739e-05 | 0.001411 |
| 124 | REGULATION OF TRANSFERASE ACTIVITY | 15 | 946 | 3.785e-05 | 0.001411 |
| 125 | CELLULAR RESPONSE TO STRESS | 20 | 1565 | 3.79e-05 | 0.001411 |
| 126 | NEGATIVE REGULATION OF CELL COMMUNICATION | 17 | 1192 | 4.156e-05 | 0.001535 |
| 127 | GRANULOCYTE DIFFERENTIATION | 3 | 15 | 4.532e-05 | 0.001661 |
| 128 | RESPONSE TO ETHANOL | 6 | 136 | 4.684e-05 | 0.001686 |
| 129 | PITUITARY GLAND DEVELOPMENT | 4 | 42 | 4.656e-05 | 0.001686 |
| 130 | EMBRYONIC PLACENTA DEVELOPMENT | 5 | 83 | 4.712e-05 | 0.001686 |
| 131 | NEGATIVE REGULATION OF HISTONE METHYLATION | 3 | 16 | 5.559e-05 | 0.00196 |
| 132 | REGULATION OF PROTEIN HOMOOLIGOMERIZATION | 3 | 16 | 5.559e-05 | 0.00196 |
| 133 | TUBE DEVELOPMENT | 11 | 552 | 6.159e-05 | 0.002155 |
| 134 | CELLULAR RESPONSE TO LIPID | 10 | 457 | 6.277e-05 | 0.00218 |
| 135 | REGULATION OF NEURON DIFFERENTIATION | 11 | 554 | 6.362e-05 | 0.002193 |
| 136 | NUCLEOSOME DISASSEMBLY | 3 | 17 | 6.727e-05 | 0.002229 |
| 137 | REGULATION OF MAPK CASCADE | 12 | 660 | 6.755e-05 | 0.002229 |
| 138 | RESPONSE TO COCAINE | 4 | 46 | 6.69e-05 | 0.002229 |
| 139 | PROTEIN DNA COMPLEX DISASSEMBLY | 3 | 17 | 6.727e-05 | 0.002229 |
| 140 | CHROMATIN DISASSEMBLY | 3 | 17 | 6.727e-05 | 0.002229 |
| 141 | REGULATION OF HISTONE H3 K9 METHYLATION | 3 | 17 | 6.727e-05 | 0.002229 |
| 142 | CELLULAR RESPONSE TO INSULIN STIMULUS | 6 | 146 | 6.956e-05 | 0.002279 |
| 143 | HEART DEVELOPMENT | 10 | 466 | 7.381e-05 | 0.002402 |
| 144 | REGULATION OF KINASE ACTIVITY | 13 | 776 | 7.568e-05 | 0.002445 |
| 145 | NEGATIVE REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY BY P53 CLASS MEDIATOR | 3 | 18 | 8.045e-05 | 0.002564 |
| 146 | CELL PROLIFERATION | 12 | 672 | 8.018e-05 | 0.002564 |
| 147 | RESPONSE TO GROWTH FACTOR | 10 | 475 | 8.646e-05 | 0.002737 |
| 148 | NEGATIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 12 | 684 | 9.48e-05 | 0.002981 |
| 149 | ORGANIC HYDROXY COMPOUND METABOLIC PROCESS | 10 | 482 | 9.752e-05 | 0.003046 |
| 150 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 15 | 1036 | 0.0001055 | 0.00323 |
| 151 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 15 | 1036 | 0.0001055 | 0.00323 |
| 152 | CELL FATE COMMITMENT | 7 | 227 | 0.0001048 | 0.00323 |
| 153 | REGULATION OF TRANSCRIPTION INVOLVED IN CELL FATE COMMITMENT | 3 | 20 | 0.0001116 | 0.003372 |
| 154 | INTRACELLULAR ESTROGEN RECEPTOR SIGNALING PATHWAY | 3 | 20 | 0.0001116 | 0.003372 |
| 155 | POSITIVE REGULATION OF CELL PROLIFERATION | 13 | 814 | 0.0001221 | 0.003667 |
| 156 | REGULATION OF ORGANELLE ORGANIZATION | 16 | 1178 | 0.0001275 | 0.003804 |
| 157 | EMBRYONIC ORGAN DEVELOPMENT | 9 | 406 | 0.000135 | 0.004001 |
| 158 | RESPONSE TO ACID CHEMICAL | 8 | 319 | 0.00014 | 0.004102 |
| 159 | CENTRAL NERVOUS SYSTEM NEURON DIFFERENTIATION | 6 | 166 | 0.0001411 | 0.004102 |
| 160 | POSITIVE REGULATION OF GROWTH | 7 | 238 | 0.0001405 | 0.004102 |
| 161 | REGULATION OF HISTONE METHYLATION | 4 | 56 | 0.0001452 | 0.004197 |
| 162 | REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY BY P53 CLASS MEDIATOR | 3 | 22 | 0.0001497 | 0.004301 |
| 163 | RESPONSE TO TOXIC SUBSTANCE | 7 | 241 | 0.0001518 | 0.004334 |
| 164 | PEPTIDYL AMINO ACID MODIFICATION | 13 | 841 | 0.0001686 | 0.004784 |
| 165 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE III PROMOTER | 3 | 23 | 0.0001716 | 0.00481 |
| 166 | POSITIVE REGULATION OF ERK1 AND ERK2 CASCADE | 6 | 172 | 0.0001712 | 0.00481 |
| 167 | REGULATION OF CELL ADHESION | 11 | 629 | 0.0001944 | 0.005416 |
| 168 | REGULATION OF NEURON DEATH | 7 | 252 | 0.0001997 | 0.005531 |
| 169 | REGULATION OF PHOSPHOLIPID METABOLIC PROCESS | 4 | 61 | 0.0002026 | 0.005579 |
| 170 | DEFENSE RESPONSE | 16 | 1231 | 0.0002111 | 0.005743 |
| 171 | RESPONSE TO DRUG | 9 | 431 | 0.0002109 | 0.005743 |
| 172 | POSITIVE REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 4 | 62 | 0.0002158 | 0.005838 |
| 173 | NEGATIVE REGULATION OF CELL CYCLE | 9 | 433 | 0.0002183 | 0.005871 |
| 174 | CELLULAR RESPONSE TO ALCOHOL | 5 | 115 | 0.000221 | 0.005885 |
| 175 | PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 3 | 25 | 0.0002213 | 0.005885 |
| 176 | MAMMARY GLAND DEVELOPMENT | 5 | 117 | 0.0002395 | 0.006331 |
| 177 | PEPTIDYL TYROSINE MODIFICATION | 6 | 186 | 0.0002613 | 0.00687 |
| 178 | FOREBRAIN GENERATION OF NEURONS | 4 | 66 | 0.0002748 | 0.007135 |
| 179 | REGULATION OF HOMEOSTATIC PROCESS | 9 | 447 | 0.000276 | 0.007135 |
| 180 | RESPONSE TO OXIDATIVE STRESS | 8 | 352 | 0.0002732 | 0.007135 |
| 181 | REGULATION OF HISTONE H3 K4 METHYLATION | 3 | 27 | 0.0002796 | 0.007147 |
| 182 | NEGATIVE REGULATION OF SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 3 | 27 | 0.0002796 | 0.007147 |
| 183 | REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 5 | 122 | 0.0002908 | 0.007394 |
| 184 | INTERSPECIES INTERACTION BETWEEN ORGANISMS | 11 | 662 | 0.0003014 | 0.007555 |
| 185 | ENDOCRINE SYSTEM DEVELOPMENT | 5 | 123 | 0.000302 | 0.007555 |
| 186 | SYMBIOSIS ENCOMPASSING MUTUALISM THROUGH PARASITISM | 11 | 662 | 0.0003014 | 0.007555 |
| 187 | INFLAMMATORY RESPONSE | 9 | 454 | 0.0003093 | 0.007669 |
| 188 | PROTEIN AUTOPHOSPHORYLATION | 6 | 192 | 0.0003098 | 0.007669 |
| 189 | REGULATION OF MEGAKARYOCYTE DIFFERENTIATION | 3 | 28 | 0.000312 | 0.007682 |
| 190 | REGULATION OF CELLULAR LOCALIZATION | 16 | 1277 | 0.0003189 | 0.007768 |
| 191 | DEVELOPMENTAL MATURATION | 6 | 193 | 0.0003186 | 0.007768 |
| 192 | CELLULAR COMPONENT MORPHOGENESIS | 13 | 900 | 0.0003256 | 0.007892 |
| 193 | CELLULAR RESPONSE TO PEPTIDE | 7 | 274 | 0.000332 | 0.008005 |
| 194 | EAR DEVELOPMENT | 6 | 195 | 0.0003366 | 0.008032 |
| 195 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 8 | 363 | 0.0003357 | 0.008032 |
| 196 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 12 | 788 | 0.0003502 | 0.008272 |
| 197 | CIRCULATORY SYSTEM DEVELOPMENT | 12 | 788 | 0.0003502 | 0.008272 |
| 198 | CIRCULATORY SYSTEM PROCESS | 8 | 366 | 0.0003547 | 0.008335 |
| 199 | BODY FLUID SECRETION | 4 | 71 | 0.0003638 | 0.008507 |
| 200 | CELLULAR LIPID METABOLIC PROCESS | 13 | 913 | 0.0003735 | 0.008689 |
| 201 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 6 | 200 | 0.0003852 | 0.008918 |
| 202 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 10 | 573 | 0.0003918 | 0.009026 |
| 203 | CELL MATURATION | 5 | 131 | 0.0004038 | 0.009229 |
| 204 | REGULATION OF NEURAL PRECURSOR CELL PROLIFERATION | 4 | 73 | 0.0004046 | 0.009229 |
| 205 | POSITIVE REGULATION OF CELL DEVELOPMENT | 9 | 472 | 0.0004106 | 0.00932 |
| 206 | RNA STABILIZATION | 3 | 31 | 0.0004238 | 0.009525 |
| 207 | BILE ACID AND BILE SALT TRANSPORT | 3 | 31 | 0.0004238 | 0.009525 |
| 208 | RESPONSE TO INSULIN | 6 | 205 | 0.0004392 | 0.009825 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | MACROMOLECULAR COMPLEX BINDING | 28 | 1399 | 3.31e-11 | 3.075e-08 |
| 2 | ENZYME BINDING | 30 | 1737 | 2.016e-10 | 5.075e-08 |
| 3 | CHROMATIN BINDING | 16 | 435 | 2.185e-10 | 5.075e-08 |
| 4 | TRANSCRIPTION COACTIVATOR ACTIVITY | 14 | 296 | 1.243e-10 | 5.075e-08 |
| 5 | RECEPTOR BINDING | 27 | 1476 | 6.117e-10 | 1.137e-07 |
| 6 | REGULATORY REGION NUCLEIC ACID BINDING | 20 | 818 | 1.287e-09 | 1.992e-07 |
| 7 | TRANSCRIPTION FACTOR ACTIVITY PROTEIN BINDING | 17 | 588 | 2.225e-09 | 2.717e-07 |
| 8 | SEQUENCE SPECIFIC DNA BINDING | 22 | 1037 | 2.34e-09 | 2.717e-07 |
| 9 | DOUBLE STRANDED DNA BINDING | 17 | 764 | 1.036e-07 | 1.069e-05 |
| 10 | RNA POLYMERASE II TRANSCRIPTION FACTOR ACTIVITY SEQUENCE SPECIFIC DNA BINDING | 15 | 629 | 2.656e-07 | 2.468e-05 |
| 11 | PROTEIN COMPLEX BINDING | 18 | 935 | 3.571e-07 | 3.015e-05 |
| 12 | NUCLEIC ACID BINDING TRANSCRIPTION FACTOR ACTIVITY | 19 | 1199 | 2.995e-06 | 0.0002319 |
| 13 | IDENTICAL PROTEIN BINDING | 19 | 1209 | 3.381e-06 | 0.0002416 |
| 14 | ENHANCER BINDING | 6 | 93 | 5.351e-06 | 0.0003314 |
| 15 | LIGAND DEPENDENT NUCLEAR RECEPTOR TRANSCRIPTION COACTIVATOR ACTIVITY | 5 | 53 | 5.21e-06 | 0.0003314 |
| 16 | KINASE BINDING | 13 | 606 | 5.711e-06 | 0.0003316 |
| 17 | NUCLEOSOMAL DNA BINDING | 4 | 30 | 1.191e-05 | 0.0006507 |
| 18 | RNA POLYMERASE II DISTAL ENHANCER SEQUENCE SPECIFIC DNA BINDING | 5 | 65 | 1.434e-05 | 0.00074 |
| 19 | TRANSCRIPTION FACTOR BINDING | 11 | 524 | 3.853e-05 | 0.00183 |
| 20 | PROTEIN DOMAIN SPECIFIC BINDING | 12 | 624 | 3.941e-05 | 0.00183 |
| 21 | STEROID HORMONE RECEPTOR BINDING | 5 | 81 | 4.189e-05 | 0.001853 |
| 22 | NUCLEOSOME BINDING | 4 | 45 | 6.131e-05 | 0.002589 |
| 23 | CORE PROMOTER PROXIMAL REGION DNA BINDING | 9 | 371 | 6.812e-05 | 0.002752 |
| 24 | TRANSCRIPTION FACTOR ACTIVITY DIRECT LIGAND REGULATED SEQUENCE SPECIFIC DNA BINDING | 4 | 48 | 7.92e-05 | 0.003066 |
| 25 | CORE PROMOTER BINDING | 6 | 152 | 8.694e-05 | 0.003231 |
| 26 | PROTEIN KINASE C BINDING | 4 | 50 | 9.306e-05 | 0.003325 |
| 27 | CORE PROMOTER SEQUENCE SPECIFIC DNA BINDING | 5 | 101 | 0.0001202 | 0.004135 |
| 28 | LIGAND DEPENDENT NUCLEAR RECEPTOR BINDING | 3 | 23 | 0.0001716 | 0.005498 |
| 29 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 8 | 328 | 0.0001694 | 0.005498 |
| 30 | STEROID HORMONE RECEPTOR ACTIVITY | 4 | 59 | 0.000178 | 0.005512 |
| 31 | PROTEIN TYROSINE KINASE ACTIVITY | 6 | 176 | 0.0001939 | 0.005811 |
| 32 | CELL ADHESION MOLECULE BINDING | 6 | 186 | 0.0002613 | 0.007586 |
| 33 | PROTEIN DIMERIZATION ACTIVITY | 15 | 1149 | 0.000325 | 0.009151 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CHROMATIN | 14 | 441 | 2.101e-08 | 6.136e-06 |
| 2 | NUCLEAR CHROMATIN | 12 | 291 | 1.339e-08 | 6.136e-06 |
| 3 | NUCLEAR PERIPHERY | 7 | 121 | 1.772e-06 | 0.000345 |
| 4 | NUCLEAR CHROMOSOME | 12 | 523 | 6.865e-06 | 0.0007057 |
| 5 | NUCLEAR MATRIX | 6 | 98 | 7.25e-06 | 0.0007057 |
| 6 | PLASMA MEMBRANE REGION | 16 | 929 | 7.239e-06 | 0.0007057 |
| 7 | NPBAF COMPLEX | 3 | 12 | 2.214e-05 | 0.001847 |
| 8 | NBAF COMPLEX | 3 | 14 | 3.638e-05 | 0.002656 |
| 9 | SWI SNF COMPLEX | 3 | 15 | 4.532e-05 | 0.002941 |
| 10 | PERINUCLEAR REGION OF CYTOPLASM | 12 | 642 | 5.184e-05 | 0.003028 |
| 11 | CHROMOSOME | 14 | 880 | 6.814e-05 | 0.003618 |
| 12 | NEURON PART | 17 | 1265 | 8.681e-05 | 0.0039 |
| 13 | MEMBRANE REGION | 16 | 1134 | 8.186e-05 | 0.0039 |
| 14 | BAF TYPE COMPLEX | 3 | 23 | 0.0001716 | 0.007159 |
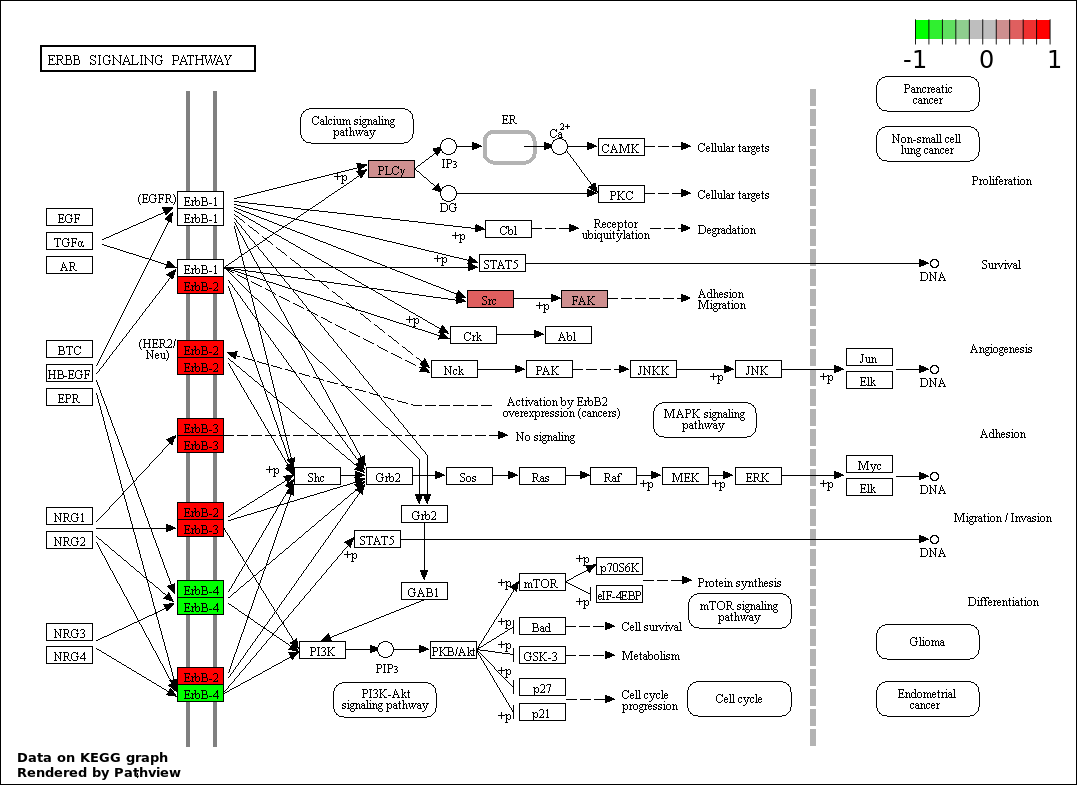
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04012_ErbB_signaling_pathway | 6 | 87 | 3.627e-06 | 0.0006529 | |
| 2 | hsa04670_Leukocyte_transendothelial_migration | 5 | 117 | 0.0002395 | 0.02155 | |
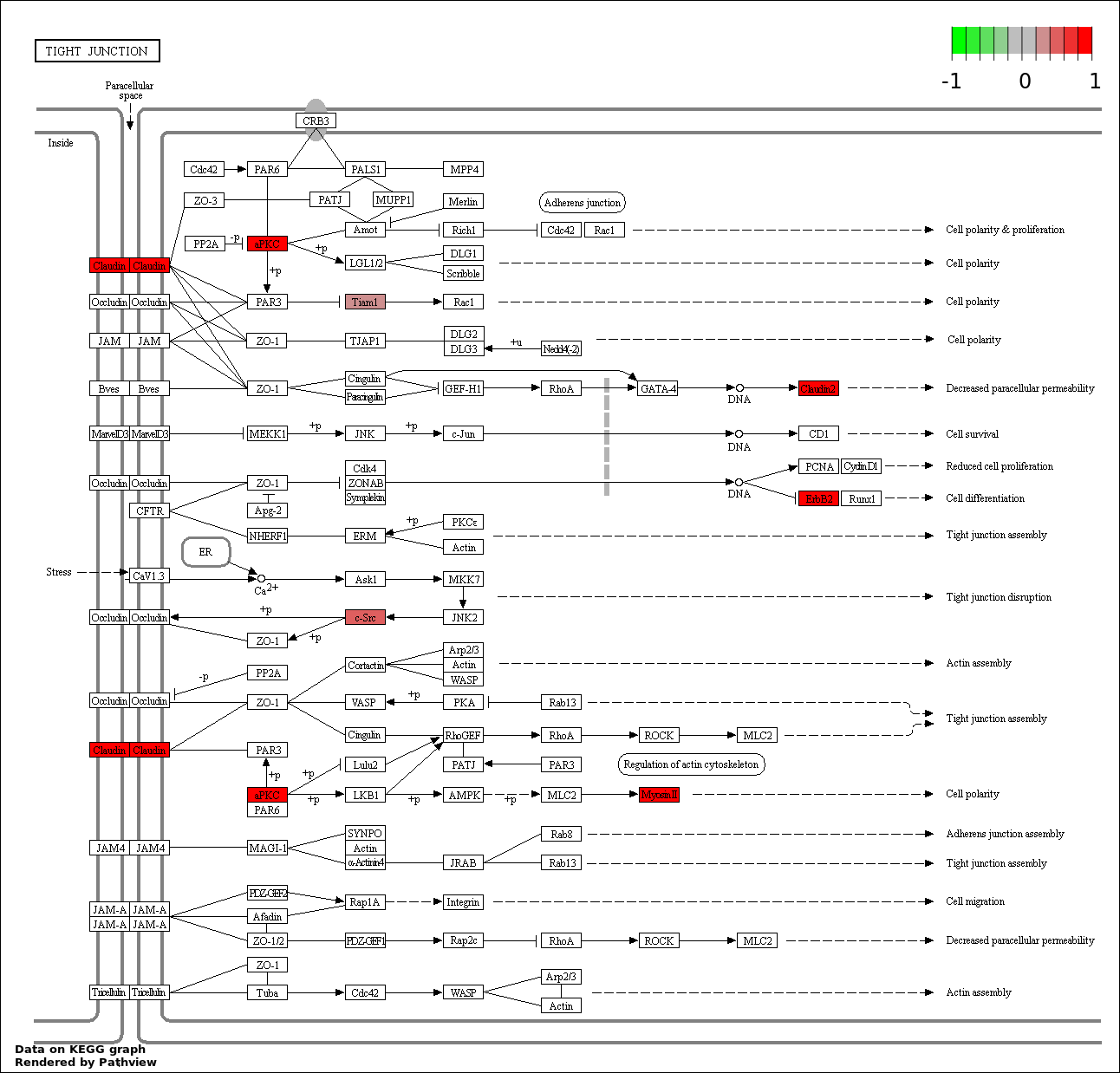
| 3 | hsa04530_Tight_junction | 5 | 133 | 0.0004328 | 0.02597 | |
| 4 | hsa04144_Endocytosis | 5 | 203 | 0.002839 | 0.1263 | |
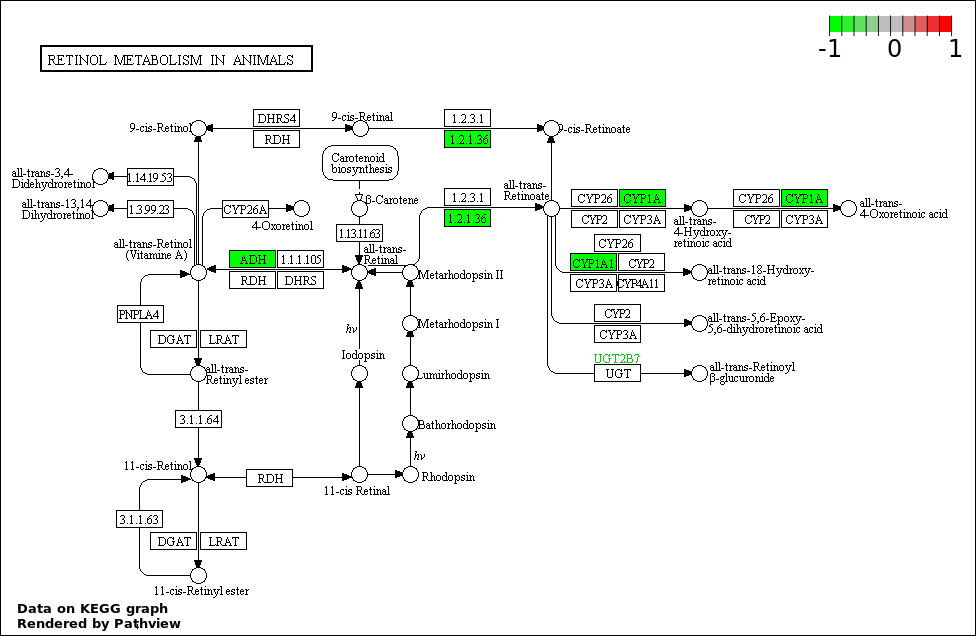
| 5 | hsa00830_Retinol_metabolism | 3 | 64 | 0.003508 | 0.1263 | |
| 6 | hsa04115_p53_signaling_pathway | 3 | 69 | 0.004337 | 0.1301 | |
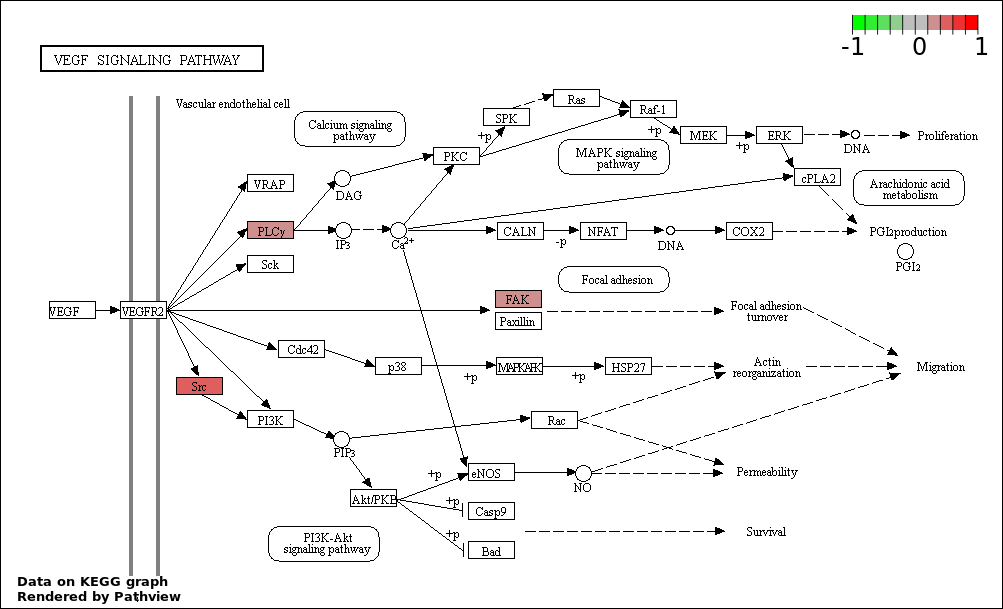
| 7 | hsa04370_VEGF_signaling_pathway | 3 | 76 | 0.005681 | 0.1461 | |
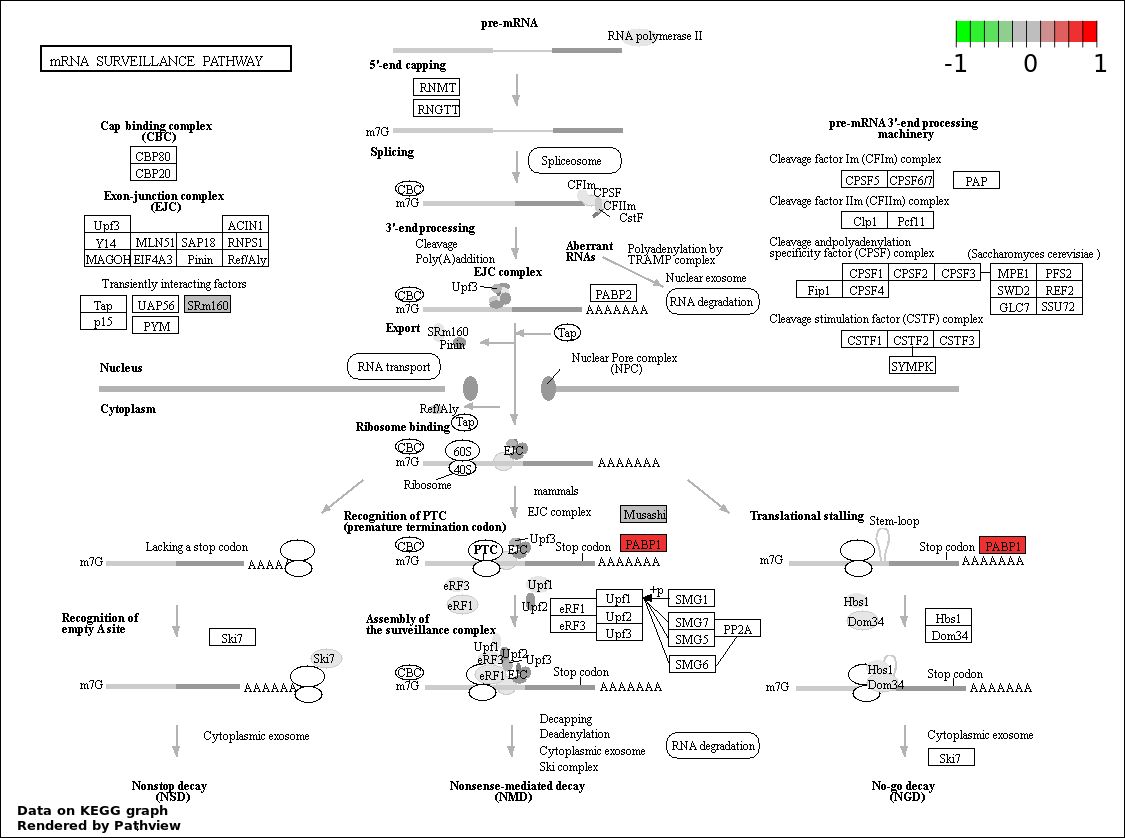
| 8 | hsa03015_mRNA_surveillance_pathway | 3 | 83 | 0.007251 | 0.1631 | |
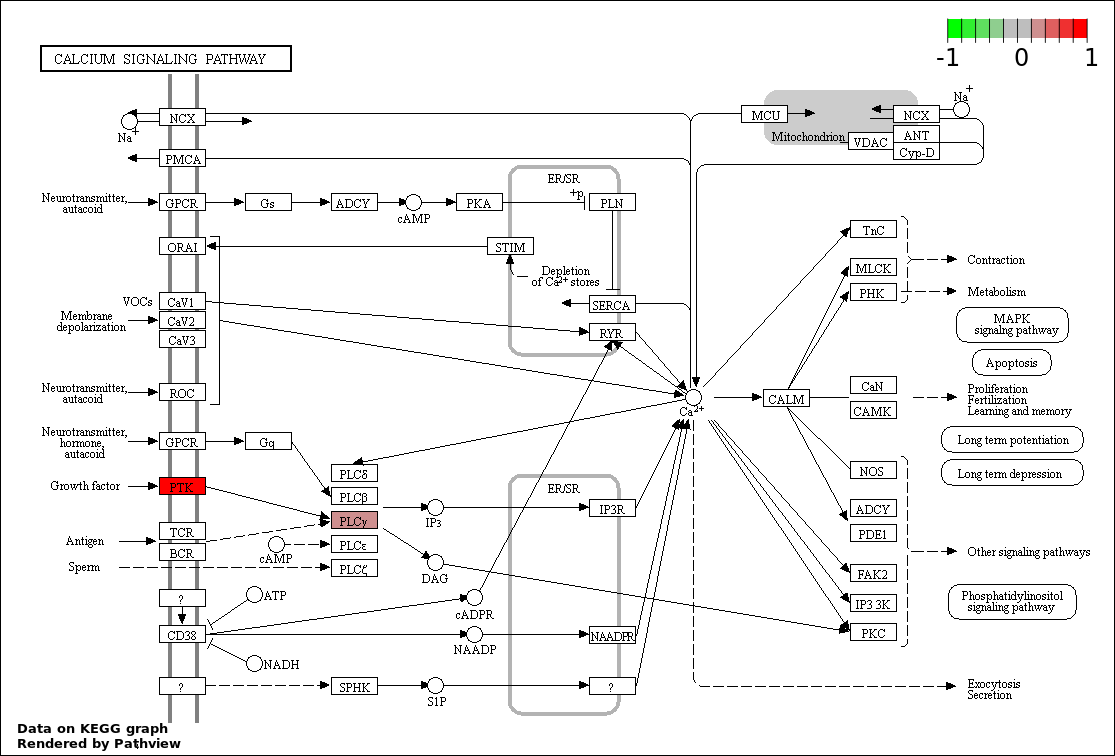
| 9 | hsa04020_Calcium_signaling_pathway | 4 | 177 | 0.01011 | 0.2022 | |
| 10 | hsa00270_Cysteine_and_methionine_metabolism | 2 | 36 | 0.01266 | 0.228 | |
| 11 | hsa04810_Regulation_of_actin_cytoskeleton | 4 | 214 | 0.01905 | 0.3117 | |
| 12 | hsa00980_Metabolism_of_xenobiotics_by_cytochrome_P450 | 2 | 71 | 0.0449 | 0.6068 | |
| 13 | hsa03018_RNA_degradation | 2 | 71 | 0.0449 | 0.6068 | |
| 14 | hsa04520_Adherens_junction | 2 | 73 | 0.04719 | 0.6068 | |
| 15 | hsa04062_Chemokine_signaling_pathway | 3 | 189 | 0.06119 | 0.7343 | |
| 16 | hsa04510_Focal_adhesion | 3 | 200 | 0.06998 | 0.7872 | |
| 17 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 2 | 95 | 0.07504 | 0.7946 | |
| 18 | hsa04620_Toll.like_receptor_signaling_pathway | 2 | 102 | 0.08477 | 0.8477 | |
| 19 | hsa04014_Ras_signaling_pathway | 3 | 236 | 0.1023 | 0.9211 | |
| 20 | hsa04514_Cell_adhesion_molecules_.CAMs. | 2 | 136 | 0.1366 | 1 | |
| 21 | hsa04120_Ubiquitin_mediated_proteolysis | 2 | 139 | 0.1414 | 1 | |
| 22 | hsa03013_RNA_transport | 2 | 152 | 0.1629 | 1 | |
| 23 | hsa04390_Hippo_signaling_pathway | 2 | 154 | 0.1662 | 1 | |
| 24 | hsa04151_PI3K_AKT_signaling_pathway | 3 | 351 | 0.2332 | 1 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | FLG-AS1 | hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-28-5p;hsa-miR-30e-3p;hsa-miR-320a;hsa-miR-326;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-532-3p;hsa-miR-590-3p;hsa-miR-942-5p | 15 | CD44 | Sponge network | -1.812 | 0.00363 | -0.797 | 0.04276 | 0.407 |
| 2 | RP11-532F6.3 | hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-30e-3p;hsa-miR-326;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-590-3p;hsa-miR-744-5p | 10 | CD44 | Sponge network | -1.772 | 1.0E-5 | -0.797 | 0.04276 | 0.356 |
| 3 | LINC00163 | hsa-miR-148a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-28-5p;hsa-miR-326;hsa-miR-328-3p;hsa-miR-330-5p;hsa-miR-33a-3p | 10 | CD44 | Sponge network | -4.312 | 0.00014 | -0.797 | 0.04276 | 0.346 |
| 4 | RP11-356J5.12 | hsa-miR-148a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-30e-3p;hsa-miR-320a;hsa-miR-326;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-532-3p;hsa-miR-590-3p;hsa-miR-942-5p | 13 | CD44 | Sponge network | -2.015 | 0 | -0.797 | 0.04276 | 0.317 |
| 5 | APCDD1L-AS1 | hsa-miR-148a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-28-5p;hsa-miR-320a;hsa-miR-328-3p;hsa-miR-330-3p;hsa-miR-532-3p | 10 | CD44 | Sponge network | -2.022 | 0.00702 | -0.797 | 0.04276 | 0.316 |
| 6 | LINC00702 | hsa-miR-148a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-30e-3p;hsa-miR-320a;hsa-miR-326;hsa-miR-328-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-34a-5p;hsa-miR-421;hsa-miR-532-3p;hsa-miR-590-3p;hsa-miR-942-5p | 16 | CD44 | Sponge network | -2.704 | 0 | -0.797 | 0.04276 | 0.309 |
| 7 | CTD-2013N24.2 | hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-30e-3p;hsa-miR-326;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-532-3p;hsa-miR-590-3p;hsa-miR-942-5p | 11 | CD44 | Sponge network | -1.002 | 1.0E-5 | -0.797 | 0.04276 | 0.287 |
| 8 | LINC00565 | hsa-miR-148a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-34a-5p;hsa-miR-744-5p;hsa-miR-942-5p | 10 | CD44 | Sponge network | -1.493 | 0.05998 | -0.797 | 0.04276 | 0.272 |
| 9 | LINC00242 | hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-185-5p;hsa-miR-28-5p;hsa-miR-326;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-34a-5p;hsa-miR-421;hsa-miR-744-5p | 10 | CD44 | Sponge network | -0.58 | 0.1728 | -0.797 | 0.04276 | 0.27 |
| 10 | RP11-166D19.1 | hsa-miR-148a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-30e-3p;hsa-miR-320a;hsa-miR-328-3p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-532-3p;hsa-miR-590-3p;hsa-miR-744-5p;hsa-miR-942-5p | 16 | CD44 | Sponge network | -3.855 | 0 | -0.797 | 0.04276 | 0.267 |
| 11 | AC093627.10 | hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-2-3p;hsa-miR-185-5p;hsa-miR-30e-3p;hsa-miR-326;hsa-miR-328-3p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-421;hsa-miR-532-3p;hsa-miR-590-3p;hsa-miR-744-5p;hsa-miR-942-5p | 14 | CD44 | Sponge network | -2.338 | 8.0E-5 | -0.797 | 0.04276 | 0.255 |