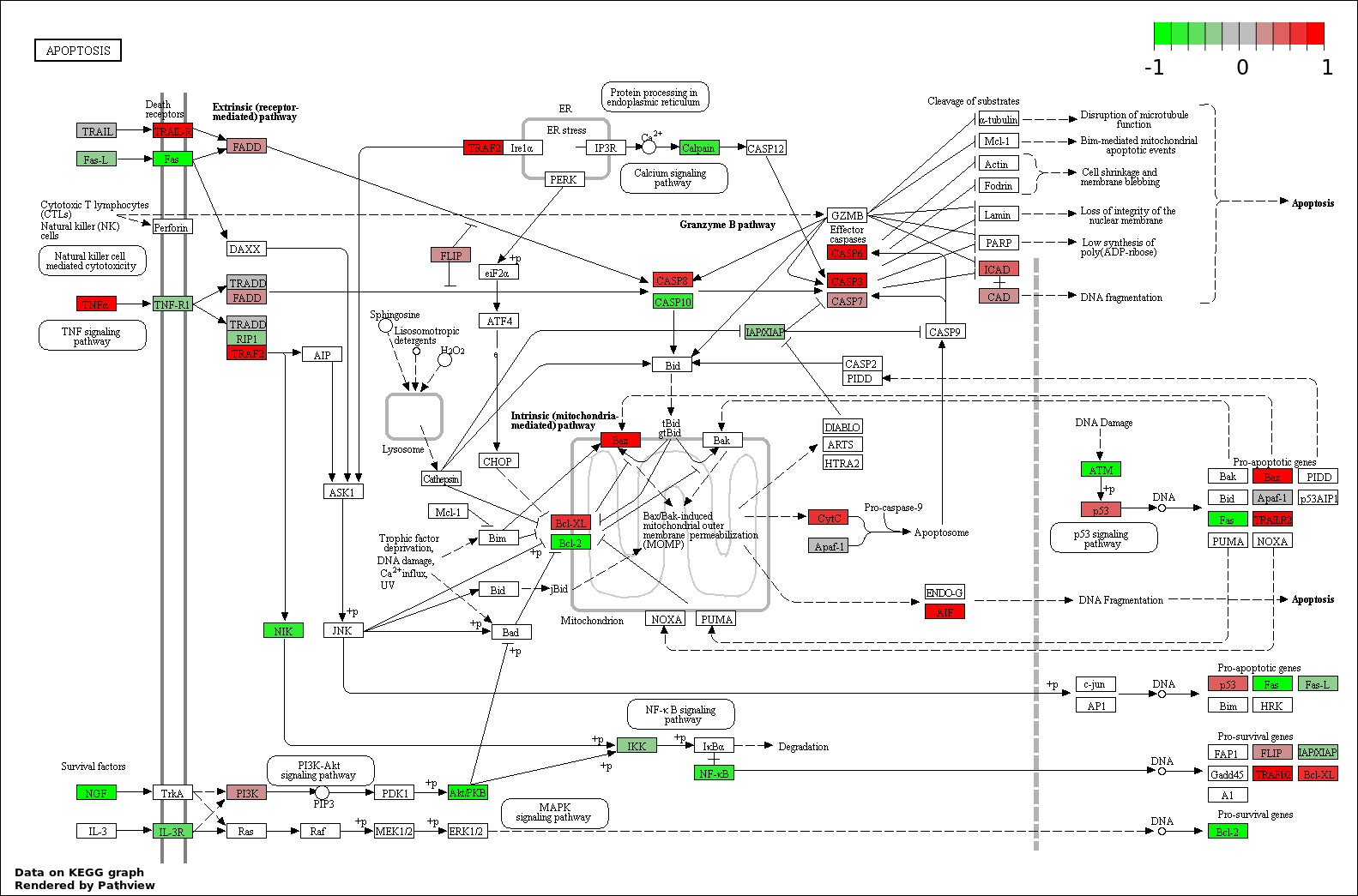
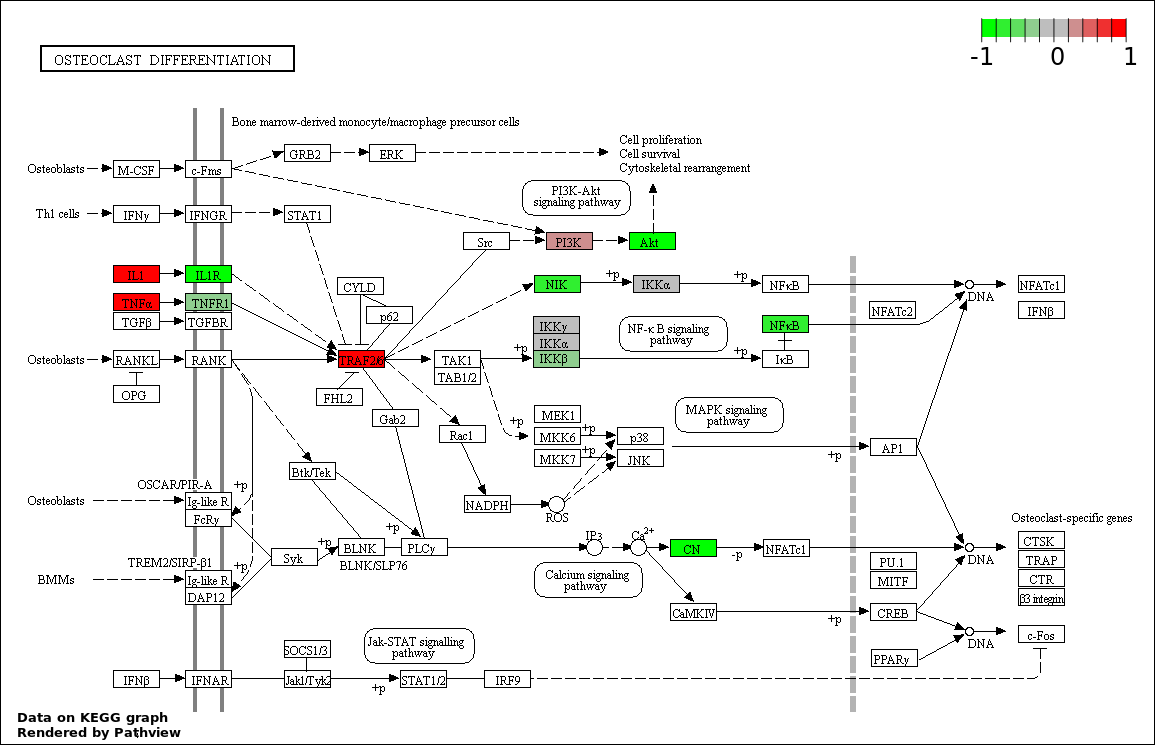
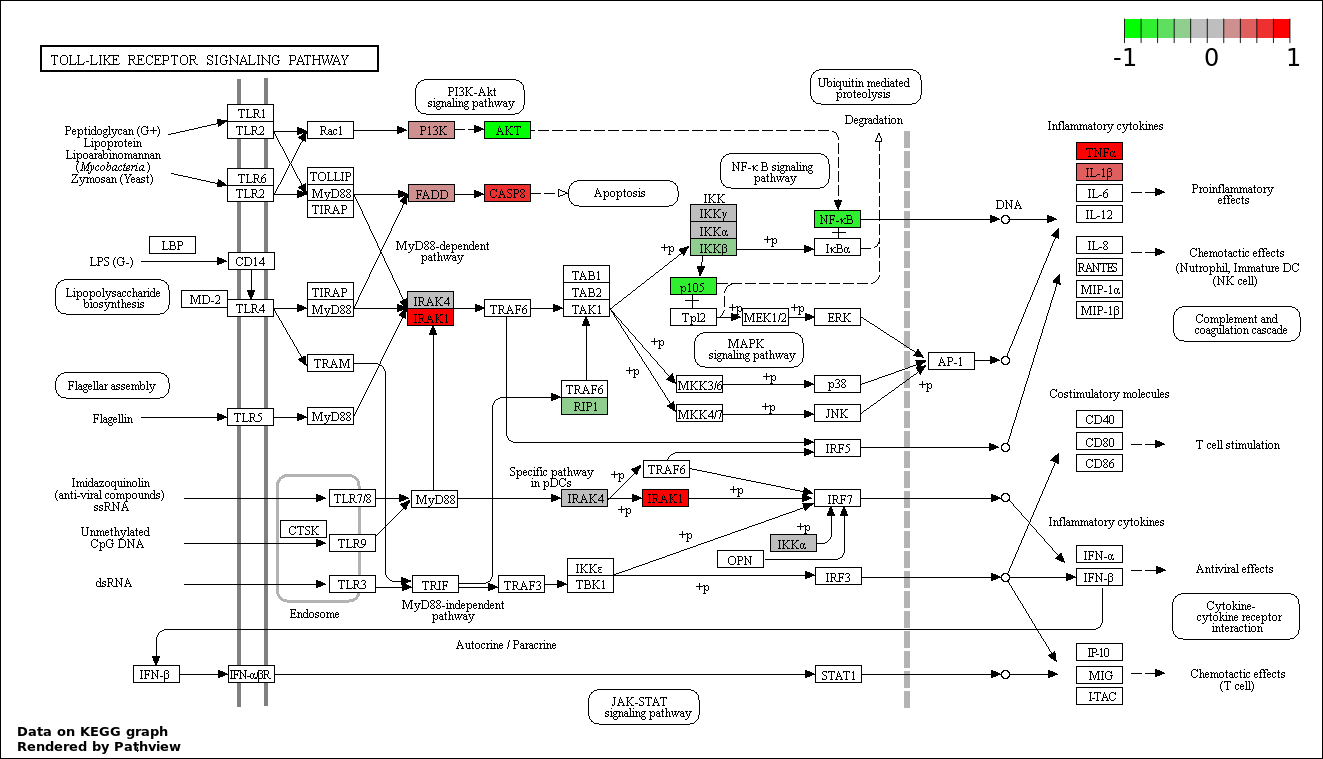
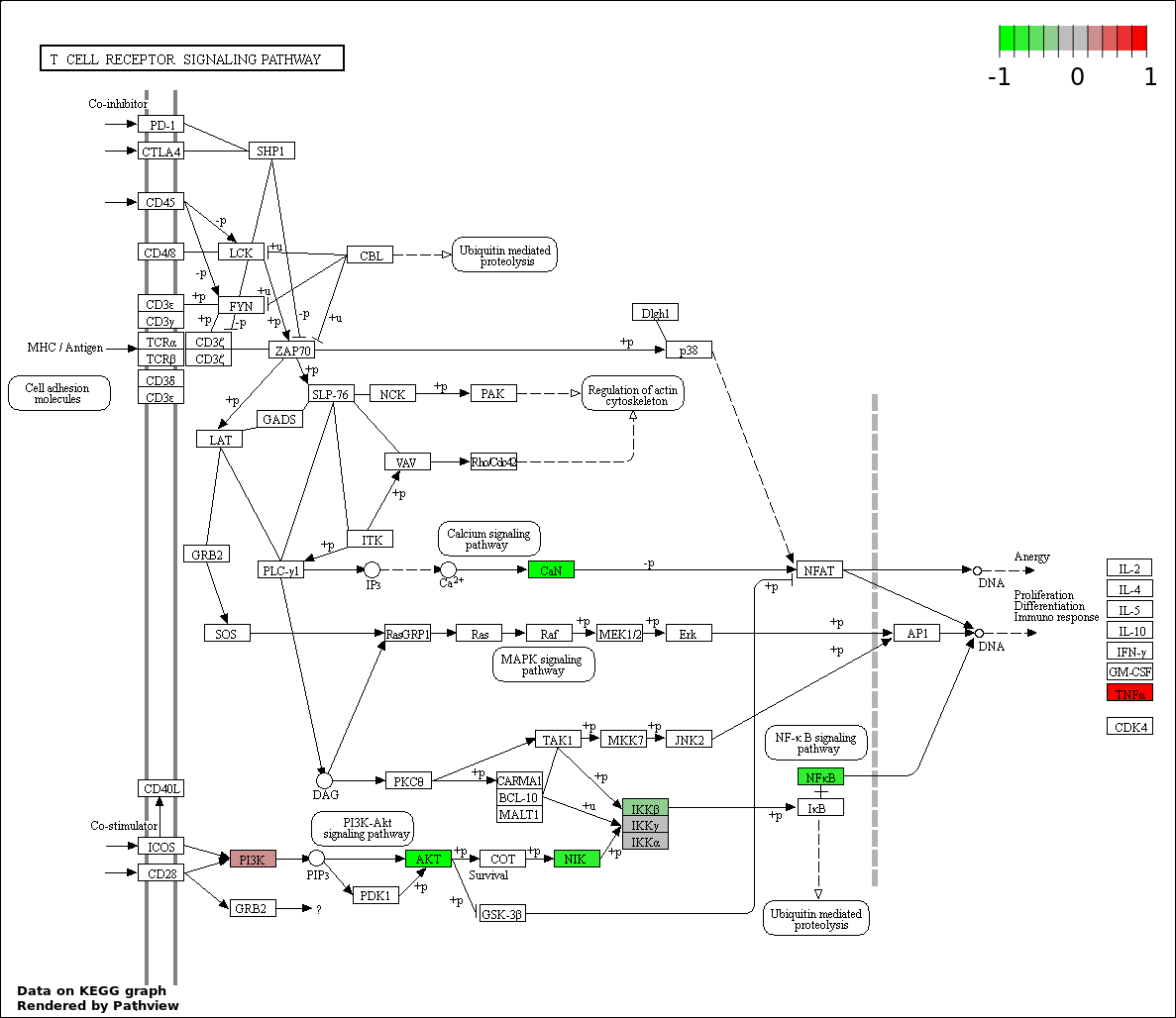
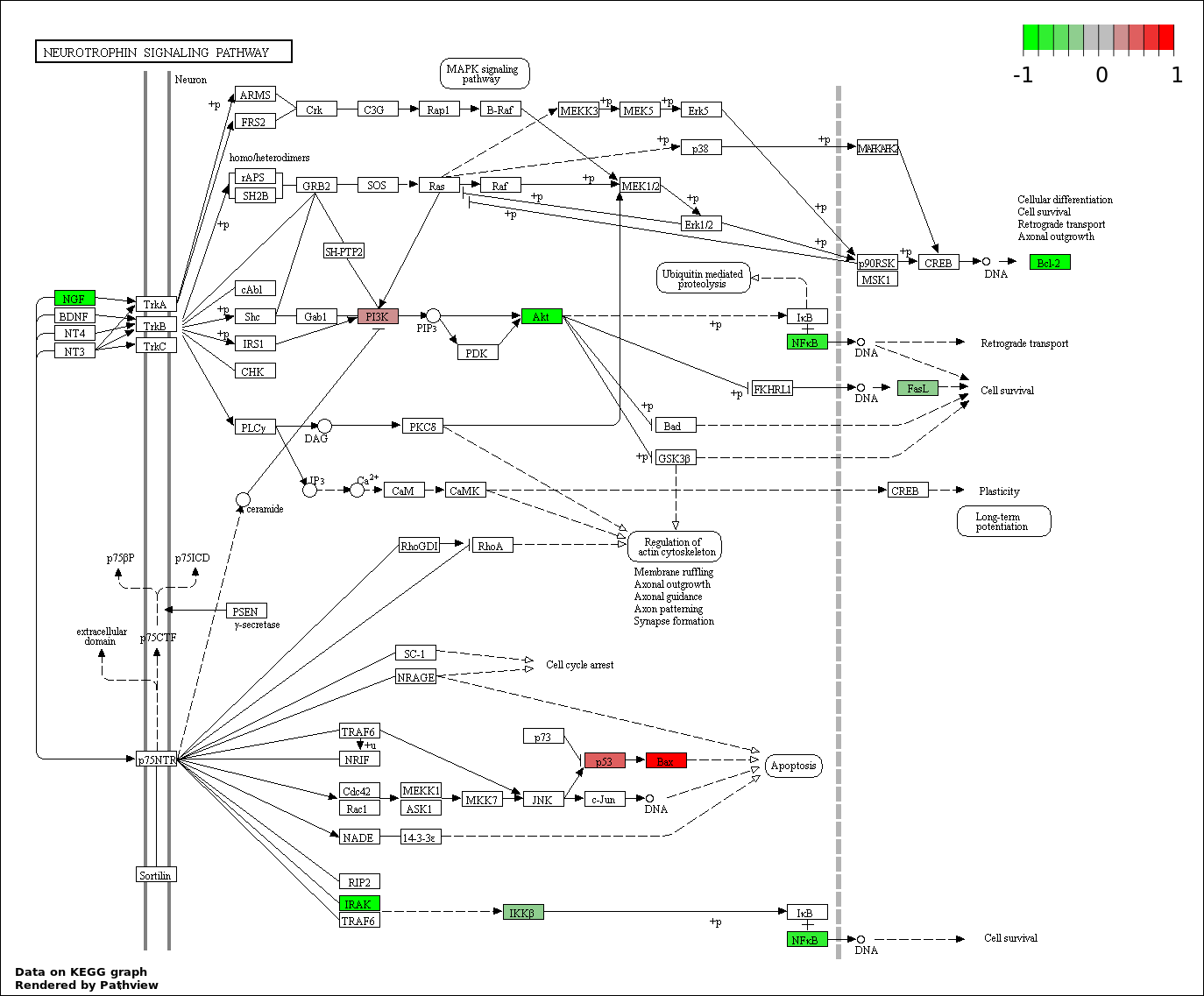
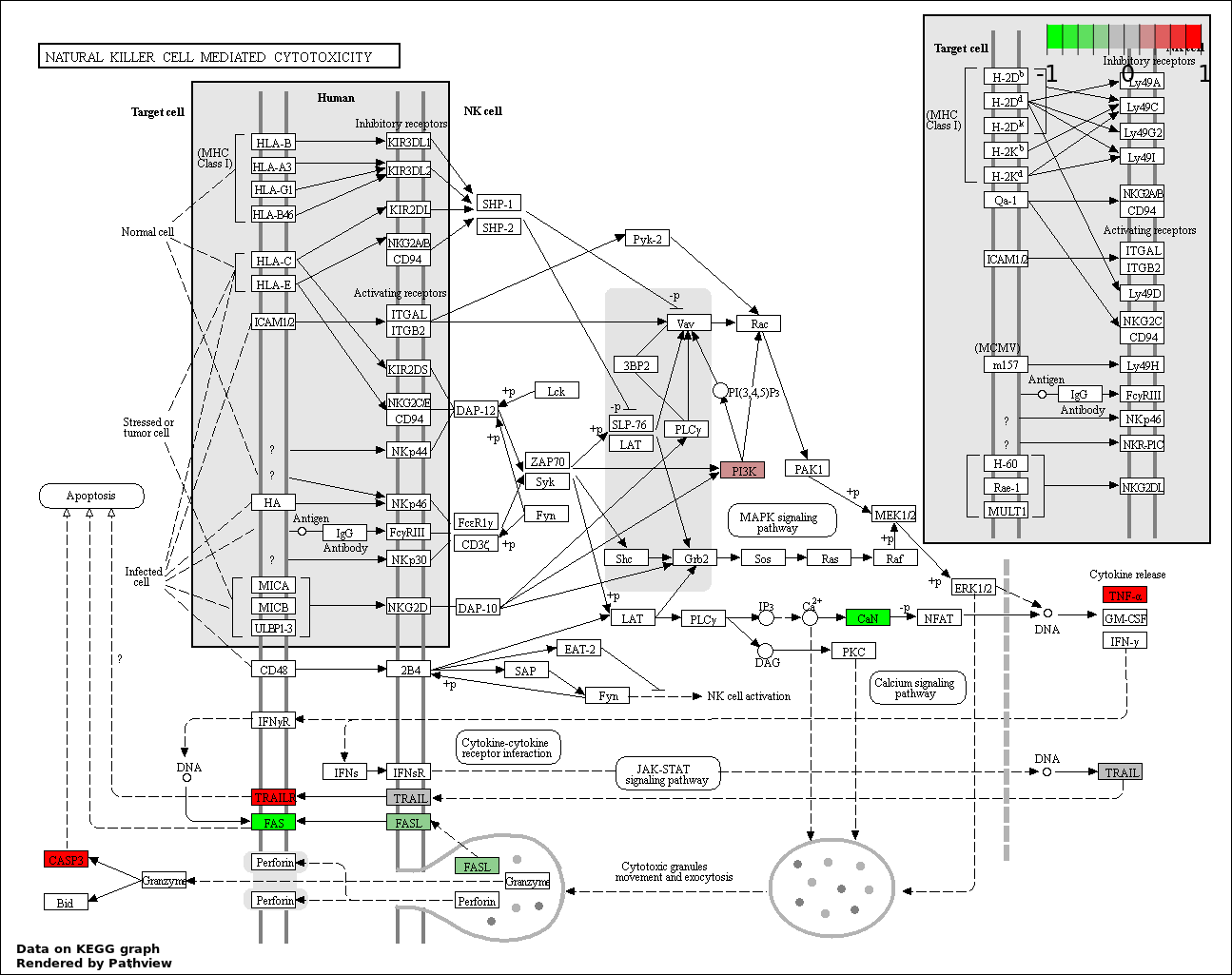
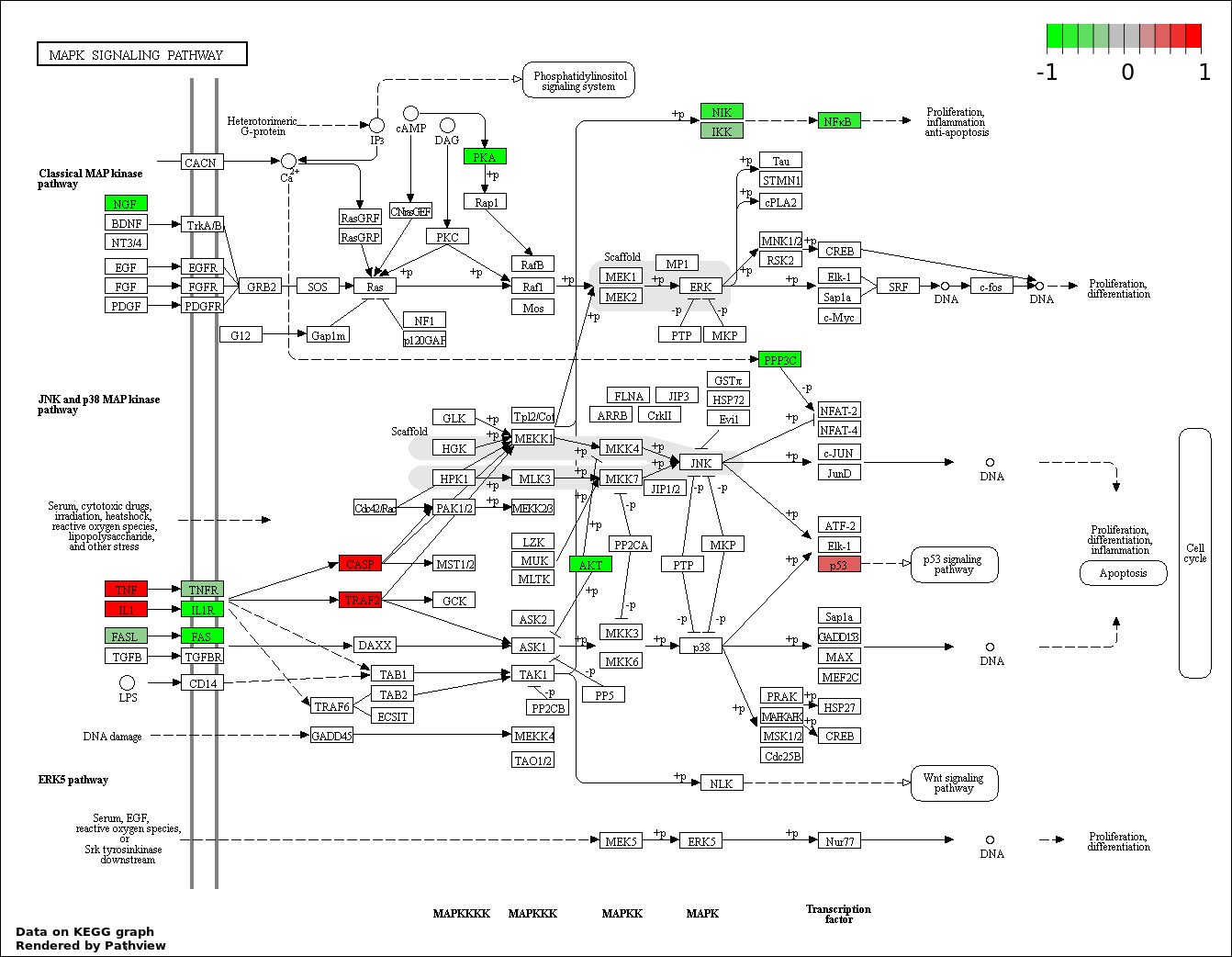
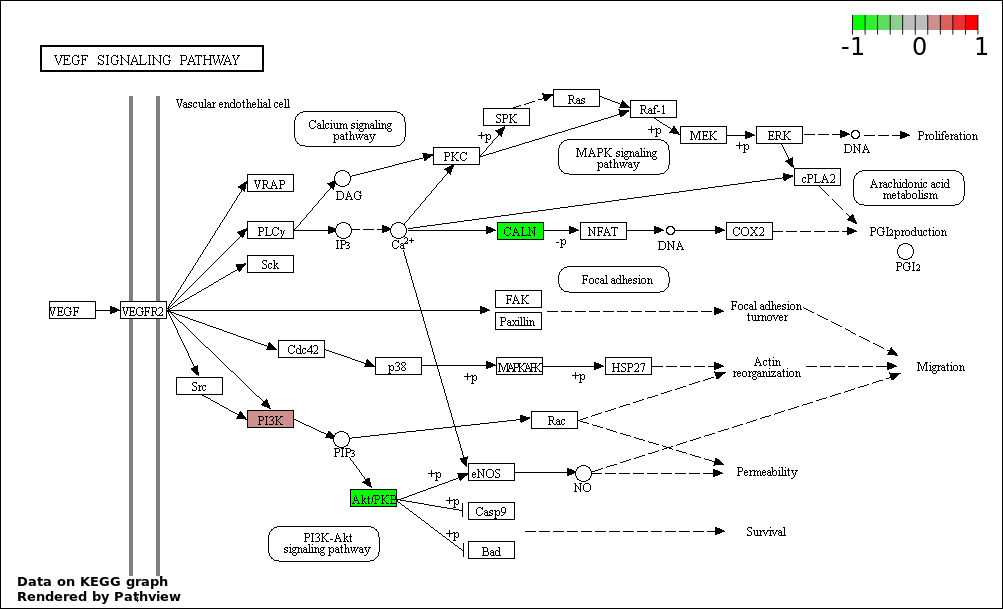
This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-125b-5p | AIFM1 | -1.42 | 0 | 1.21 | 0 | miRNATAP | -0.12 | 1.0E-5 | NA | |
| 2 | hsa-miR-143-5p | AIFM1 | -1.5 | 0 | 1.21 | 0 | miRNATAP | -0.13 | 1.0E-5 | NA | |
| 3 | hsa-miR-145-5p | AIFM1 | -2.62 | 0 | 1.21 | 0 | miRNATAP | -0.17 | 0 | 20332243 | Artificial overexpression of miR145 by using adenoviral vectors in prostate cancer PC-3 and DU145 cells significantly downregulated BNIP3 together with the upregulation of AIF reduced cell growth and increased cell death |
| 4 | hsa-miR-199a-5p | AIFM1 | -0.9 | 0.00031 | 1.21 | 0 | miRanda | -0.11 | 0.0001 | NA | |
| 5 | hsa-miR-143-3p | AKT1 | -2.59 | 0 | 0.52 | 0 | miRNAWalker2 validate | -0.05 | 0.00136 | NA | |
| 6 | hsa-let-7a-5p | AKT2 | 0.1 | 0.43289 | 0.09 | 0.4188 | TargetScan | -0.2 | 0 | NA | |
| 7 | hsa-let-7b-3p | AKT2 | 0.35 | 0.07056 | 0.09 | 0.4188 | MirTarget | -0.08 | 0.00243 | NA | |
| 8 | hsa-miR-22-3p | AKT2 | 0.74 | 0 | 0.09 | 0.4188 | mirMAP | -0.08 | 0.04433 | NA | |
| 9 | hsa-miR-106a-5p | AKT3 | 2.94 | 0 | -3.43 | 0 | miRNATAP | -0.32 | 0 | NA | |
| 10 | hsa-miR-106b-5p | AKT3 | 2.18 | 0 | -3.43 | 0 | miRNATAP | -0.77 | 0 | NA | |
| 11 | hsa-miR-107 | AKT3 | 1.81 | 0 | -3.43 | 0 | PITA; miRanda | -0.75 | 0 | NA | |
| 12 | hsa-miR-135a-5p | AKT3 | 0.31 | 0.47441 | -3.43 | 0 | miRNATAP | -0.09 | 0.00382 | NA | |
| 13 | hsa-miR-142-3p | AKT3 | 2.07 | 0 | -3.43 | 0 | miRanda | -0.18 | 0.00013 | NA | |
| 14 | hsa-miR-146b-5p | AKT3 | 1.31 | 0 | -3.43 | 0 | miRNAWalker2 validate | -0.29 | 0 | NA | |
| 15 | hsa-miR-15a-5p | AKT3 | 1.78 | 0 | -3.43 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.75 | 0 | NA | |
| 16 | hsa-miR-15b-5p | AKT3 | 2.5 | 0 | -3.43 | 0 | miRNATAP | -0.81 | 0 | NA | |
| 17 | hsa-miR-16-5p | AKT3 | 1.88 | 0 | -3.43 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.74 | 0 | NA | |
| 18 | hsa-miR-17-3p | AKT3 | 1.09 | 0 | -3.43 | 0 | miRNATAP | -0.46 | 0 | NA | |
| 19 | hsa-miR-17-5p | AKT3 | 2.27 | 0 | -3.43 | 0 | TargetScan; miRNATAP | -0.64 | 0 | NA | |
| 20 | hsa-miR-181a-5p | AKT3 | 0.97 | 0 | -3.43 | 0 | miRNATAP | -0.24 | 0.00024 | NA | |
| 21 | hsa-miR-181b-5p | AKT3 | 1.3 | 0 | -3.43 | 0 | miRNATAP | -0.37 | 0 | NA | |
| 22 | hsa-miR-181c-5p | AKT3 | 0.6 | 0.01077 | -3.43 | 0 | miRNATAP | -0.17 | 0.00212 | NA | |
| 23 | hsa-miR-20a-5p | AKT3 | 1.51 | 0 | -3.43 | 0 | miRNATAP | -0.48 | 0 | NA | |
| 24 | hsa-miR-20b-5p | AKT3 | 3.36 | 0 | -3.43 | 0 | miRNATAP | -0.17 | 0 | NA | |
| 25 | hsa-miR-29b-2-5p | AKT3 | 0.3 | 0.11355 | -3.43 | 0 | mirMAP | -0.14 | 0.03379 | NA | |
| 26 | hsa-miR-3065-5p | AKT3 | 2.75 | 0 | -3.43 | 0 | mirMAP | -0.26 | 0 | NA | |
| 27 | hsa-miR-32-3p | AKT3 | 2.4 | 0 | -3.43 | 0 | mirMAP | -0.54 | 0 | NA | |
| 28 | hsa-miR-320a | AKT3 | 0.5 | 0.00226 | -3.43 | 0 | PITA; miRanda; miRNATAP | -0.37 | 0 | NA | |
| 29 | hsa-miR-320b | AKT3 | 1.1 | 0 | -3.43 | 0 | PITA; miRanda; miRNATAP | -0.38 | 0 | NA | |
| 30 | hsa-miR-320c | AKT3 | 0.73 | 0.00058 | -3.43 | 0 | PITA; miRanda; miRNATAP | -0.29 | 0 | NA | |
| 31 | hsa-miR-335-3p | AKT3 | 1.32 | 0 | -3.43 | 0 | mirMAP | -0.44 | 0 | NA | |
| 32 | hsa-miR-33a-3p | AKT3 | 1.39 | 0 | -3.43 | 0 | mirMAP | -0.46 | 0 | NA | |
| 33 | hsa-miR-340-5p | AKT3 | 1.35 | 0 | -3.43 | 0 | mirMAP | -0.56 | 0 | NA | |
| 34 | hsa-miR-34c-3p | AKT3 | 0.24 | 0.65945 | -3.43 | 0 | PITA | -0.06 | 0.01923 | NA | |
| 35 | hsa-miR-362-3p | AKT3 | 0.54 | 0.00242 | -3.43 | 0 | miRanda | -0.21 | 0.00314 | NA | |
| 36 | hsa-miR-362-5p | AKT3 | 0.45 | 0.02925 | -3.43 | 0 | PITA; TargetScan; miRNATAP | -0.2 | 0.0011 | NA | |
| 37 | hsa-miR-421 | AKT3 | 2.1 | 0 | -3.43 | 0 | miRanda; mirMAP | -0.44 | 0 | NA | |
| 38 | hsa-miR-501-3p | AKT3 | 2.19 | 0 | -3.43 | 0 | miRNATAP | -0.52 | 0 | NA | |
| 39 | hsa-miR-502-3p | AKT3 | 0.43 | 0.01008 | -3.43 | 0 | miRNATAP | -0.2 | 0.00914 | NA | |
| 40 | hsa-miR-505-3p | AKT3 | 1.38 | 0 | -3.43 | 0 | mirMAP | -0.49 | 0 | 22051041 | We also find that Akt3 correlate inversely with miR-505 modulates drug sensitivity in MCF7-ADR |
| 41 | hsa-miR-548v | AKT3 | 1.7 | 0 | -3.43 | 0 | miRNATAP | -0.21 | 0 | NA | |
| 42 | hsa-miR-577 | AKT3 | 3.23 | 0 | -3.43 | 0 | mirMAP | -0.18 | 0 | NA | |
| 43 | hsa-miR-616-5p | AKT3 | 1.97 | 0 | -3.43 | 0 | mirMAP | -0.34 | 0 | NA | |
| 44 | hsa-miR-769-5p | AKT3 | 1.41 | 0 | -3.43 | 0 | PITA; miRNATAP | -0.67 | 0 | NA | |
| 45 | hsa-miR-93-5p | AKT3 | 2.58 | 0 | -3.43 | 0 | miRNATAP | -0.67 | 0 | NA | |
| 46 | hsa-miR-335-3p | APAF1 | 1.32 | 0 | -0 | 0.97011 | mirMAP | -0.07 | 0.00422 | NA | |
| 47 | hsa-miR-335-5p | APAF1 | 0.35 | 0.09622 | -0 | 0.97011 | miRNAWalker2 validate | -0.13 | 0 | NA | |
| 48 | hsa-miR-125a-3p | ATM | 0.77 | 6.0E-5 | -0.95 | 0 | miRanda | -0.11 | 0.00094 | NA | |
| 49 | hsa-miR-146b-5p | ATM | 1.31 | 0 | -0.95 | 0 | miRanda | -0.09 | 0.00381 | 27602131 | The role of microRNA 146b miR-146b in ATC remains to be elucidated; In order to characterize the role of miR-146b in ATC overexpression or interference of miR-146b was induced in ATC cell lines and cell proliferation and migration were evaluated; The potential targets of miR-146b were searched in the Gene Expression Omnibus database for ATC and matched non-tumor control samples; The expression level of potential targets was detected following overexpression or interference of miR-146b in ATC cell lines; In addition cell migration of ATC was also affected by miR-146b; During the search for potential targets of miR-146b in ATC p21 also known as p21Waf1/Cip1 or CDKN1A was noted for its role in cell cycle progression and tumor pathogenesis; In conclusion p21 may participate in the regulation of ATC cell proliferation by miR-146b |
| 50 | hsa-miR-186-5p | ATM | 1.01 | 0 | -0.95 | 0 | mirMAP | -0.13 | 0.00632 | NA | |
| 51 | hsa-miR-18a-5p | ATM | 3.22 | 0 | -0.95 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.12 | 0 | 23437304; 25963391; 23857602; 23229340 | MicroRNA 18a attenuates DNA damage repair through suppressing the expression of ataxia telangiectasia mutated in colorectal cancer; Through in silico search the 3'UTR of Ataxia telangiectasia mutated ATM contains a conserved miR-18a binding site; Expression of ATM was down-regulated in CRC tumors p<0.0001 and inversely correlated with miR-18a expression r = -0.4562 p<0.01; This was further confirmed by the down-regulation of ATM protein by miR-18a; As ATM is a key enzyme in DNA damage repair we evaluated the effect of miR-18a on DNA double-strand breaks; miR-18a attenuates cellular repair of DNA double-strand breaks by directly suppressing ATM a key enzyme in DNA damage repair;However the upregulation of miR-18a suppressed the level of ataxia-telangiectasia mutated and attenuated DNA double-strand break repair after irradiation which re-sensitized the cervical cancer cells to radiotherapy by promoting apoptosis;Furthermore we used antisense oligonucleotides against micro RNAs miRNA or miRNA overexpression plasmids to study the role of miR-18a and -106a on ATM expression; Furthermore we identified that ERα activates miR-18a and -106a to downregulate ATM expression; We reveal a novel mechanism involving ERα and miR-18a and -106a regulation of ATM in breast cancer;MicroRNA 18a upregulates autophagy and ataxia telangiectasia mutated gene expression in HCT116 colon cancer cells; Previous studies showed that certain microRNAs including miR-18a potentially regulate ATM in cancer cells; However the mechanisms behind the modulation of ATM by miR-18a remain to be elucidated in colon cancer cells; In the present study we explored the impact of miR-18a on the autophagy process and ATM expression in HCT116 colon cancer cells; Western blotting and luciferase assays were implemented to explore the impact of miR-18a on ATM gene expression in HCT116 cells; Moreover miR-18a overexpression led to the upregulation of ATM expression and suppression of mTORC1 activity; Results of the present study pertaining to the role of miR-18a in regulating autophagy and ATM gene expression in colon cancer cells revealed a novel function for miR-18a in a critical cellular event and on a crucial gene with significant impacts in cancer development progression treatment and in other diseases |
| 52 | hsa-miR-203a-3p | ATM | 3.14 | 0 | -0.95 | 0 | MirTarget | -0.09 | 0 | 24145123; 27542403 | miR 203 induces oxaliplatin resistance in colorectal cancer cells by negatively regulating ATM kinase; In silico analysis identified ataxia telangiectasia mutated ATM a primary mediator of the DNA damage response as a potential target of miR-203; Using TCGA database we identified a significant reverse correlation of miR-203 and ATM expression in CRC tissues; We validated ATM as a bona fide target of miR-203 in CRC cells; Mutation of the putative miR-203 binding site in the 3' untranslated region 3'UTR of the ATM mRNA abolished the inhibitory effect of miR-203 on ATM;MiR 203 inhibits tumor invasion and metastasis in gastric cancer by ATM; Our results showed that miR-203 was significantly downregulated in gastric cancer tissues and cells while ataxia telangiectasia mutated kinase ATM was upregulated in gastric cancer tissues and cells and was directly regulated by miR-203; ATM knockdown phenocopied the effect of miR-203 overexpression |
| 53 | hsa-miR-21-5p | ATM | 1.51 | 0 | -0.95 | 0 | mirMAP | -0.1 | 0.00811 | 26289851 | MiR-21 is an oncomiR that is overexpressed in nearly all cancers including ATC; Hence suppression of miR-21 could pave the way for ATC therapy |
| 54 | hsa-miR-324-5p | ATM | 2.15 | 0 | -0.95 | 0 | miRanda | -0.16 | 0 | NA | |
| 55 | hsa-miR-335-3p | ATM | 1.32 | 0 | -0.95 | 0 | mirMAP | -0.09 | 0.00248 | NA | |
| 56 | hsa-miR-339-5p | ATM | 1.79 | 0 | -0.95 | 0 | miRanda | -0.16 | 0 | NA | |
| 57 | hsa-miR-421 | ATM | 2.1 | 0 | -0.95 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget; miRanda | -0.12 | 1.0E-5 | NA | |
| 58 | hsa-miR-590-3p | ATM | 1.73 | 0 | -0.95 | 0 | miRanda; mirMAP | -0.12 | 0.00045 | NA | |
| 59 | hsa-miR-590-5p | ATM | 1.46 | 0 | -0.95 | 0 | mirMAP | -0.13 | 0.00013 | NA | |
| 60 | hsa-miR-766-3p | ATM | 2.7 | 0 | -0.95 | 0 | MirTarget | -0.15 | 0 | NA | |
| 61 | hsa-miR-92a-3p | ATM | 1.69 | 0 | -0.95 | 0 | miRNAWalker2 validate | -0.12 | 0.00206 | NA | |
| 62 | hsa-miR-939-5p | ATM | 2.15 | 0 | -0.95 | 0 | MirTarget | -0.12 | 1.0E-5 | NA | |
| 63 | hsa-miR-30a-5p | BAX | 0.2 | 0.42032 | 0.87 | 0 | miRNAWalker2 validate | -0.07 | 0.00399 | NA | |
| 64 | hsa-miR-103a-3p | BCL2 | 1.51 | 0 | -2.27 | 0 | miRNAWalker2 validate | -0.32 | 3.0E-5 | NA | |
| 65 | hsa-miR-130b-5p | BCL2 | 3.62 | 0 | -2.27 | 0 | mirMAP | -0.25 | 0 | 27364335 | The level of microRNA-130b in relationship with the expression of PPARγ VEGF-A BCL-2 and apoptosis were analyzed in 91 lung cancer patient samples using immunohistochemistry and terminal deoxynucleotidyl transferase dUTP nick end labeling TUNEL assay on tissue microarrays; In vitro and in vivo miR-130b enrichment associated with down-regulation of PPARγ up-regulation of VEGF-A and BCL-2 and decreased apoptosis |
| 66 | hsa-miR-142-3p | BCL2 | 2.07 | 0 | -2.27 | 0 | miRanda | -0.25 | 0 | NA | |
| 67 | hsa-miR-15a-5p | BCL2 | 1.78 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase | -0.47 | 0 | 25594541; 26915294; 18931683; 25623762; 22335947 | MicroRNAs miRNAs encoded by the miR-15 cluster are known to induce G1 arrest and apoptosis by targeting G1 checkpoints and the anti-apoptotic B cell lymphoma 2 BCL-2 gene;As a result transcript levels of the tumor-suppressive miR-15 and let-7 families increased which targeted and decreased the expression of the crucial prosurvival genes BCL-2 and BCL-XL respectively;MicroRNAs miRNAs are noncoding small RNAs that repress protein translation by targeting specific messenger RNAs miR-15a and miR-16-1 act as putative tumor suppressors by targeting the oncogene BCL2;miR 15a and miR 16 modulate drug resistance by targeting bcl 2 in human colon cancer cells; To investigate the reversal effect of targeted modulation of bcl-2 expression by miR-15a and miR-16 on drug resistance of human colon cancer cells;The expression of MiR-15a was significantly inhibited by Bcl-2 P < 0.05 |
| 68 | hsa-miR-15b-3p | BCL2 | 3.25 | 0 | -2.27 | 0 | mirMAP | -0.41 | 0 | 25594541; 26915294; 26884837; 18449891 | MicroRNAs miRNAs encoded by the miR-15 cluster are known to induce G1 arrest and apoptosis by targeting G1 checkpoints and the anti-apoptotic B cell lymphoma 2 BCL-2 gene;As a result transcript levels of the tumor-suppressive miR-15 and let-7 families increased which targeted and decreased the expression of the crucial prosurvival genes BCL-2 and BCL-XL respectively;MiR 15b mediates liver cancer cells proliferation through targeting BCL 2; MiR-15b overexpression downregulated BCL2 mRNA and protein expression obviously P < 0.05; On the contrary miR-15b inhibitor transfection markedly reduced miR-15b expression in liver cancer cells P < 0.05 promoted tumor cell proliferation and increased BCL2 mRNA and protein expression; MiR-15b can inhibit HepG2 cell proliferation and down-regulate BCL2 mRNA and protein expression;miR 15b and miR 16 modulate multidrug resistance by targeting BCL2 in human gastric cancer cells; The downregulation of miR-15b and miR-16 in SGC7901/VCR cells was concurrent with the upregulation of Bcl-2 protein; Taken together our findings suggest that miR-15b and miR-16 could play a role in the development of MDR in gastric cancer cells at least in part by modulation of apoptosis via targeting BCL2 |
| 69 | hsa-miR-15b-5p | BCL2 | 2.5 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase | -0.53 | 0 | 25594541; 26915294; 26884837; 18449891 | MicroRNAs miRNAs encoded by the miR-15 cluster are known to induce G1 arrest and apoptosis by targeting G1 checkpoints and the anti-apoptotic B cell lymphoma 2 BCL-2 gene;As a result transcript levels of the tumor-suppressive miR-15 and let-7 families increased which targeted and decreased the expression of the crucial prosurvival genes BCL-2 and BCL-XL respectively;MiR 15b mediates liver cancer cells proliferation through targeting BCL 2; MiR-15b overexpression downregulated BCL2 mRNA and protein expression obviously P < 0.05; On the contrary miR-15b inhibitor transfection markedly reduced miR-15b expression in liver cancer cells P < 0.05 promoted tumor cell proliferation and increased BCL2 mRNA and protein expression; MiR-15b can inhibit HepG2 cell proliferation and down-regulate BCL2 mRNA and protein expression;miR 15b and miR 16 modulate multidrug resistance by targeting BCL2 in human gastric cancer cells; The downregulation of miR-15b and miR-16 in SGC7901/VCR cells was concurrent with the upregulation of Bcl-2 protein; Taken together our findings suggest that miR-15b and miR-16 could play a role in the development of MDR in gastric cancer cells at least in part by modulation of apoptosis via targeting BCL2 |
| 70 | hsa-miR-16-1-3p | BCL2 | 2.09 | 0 | -2.27 | 0 | mirMAP | -0.31 | 0 | 18931683 | MicroRNAs miRNAs are noncoding small RNAs that repress protein translation by targeting specific messenger RNAs miR-15a and miR-16-1 act as putative tumor suppressors by targeting the oncogene BCL2 |
| 71 | hsa-miR-16-2-3p | BCL2 | 2.34 | 0 | -2.27 | 0 | mirMAP | -0.43 | 0 | NA | |
| 72 | hsa-miR-16-5p | BCL2 | 1.88 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase | -0.57 | 0 | 21336967; 24447552; 18449891; 25435430; 24598659; 18931683; 22966344; 25623762 | P glycoprotein enhances radiation induced apoptotic cell death through the regulation of miR 16 and Bcl 2 expressions in hepatocellular carcinoma cells; RHepG2 cells the multidrug resistant subline of human hepatocellular carcinoma HepG2 cells expressed higher levels of Pgp as well as miR-16 and lower level of Bcl-2 than the parental cells; On the other hand ectopic mdr1 expression enhanced radiation-induced apoptosis in HepG2 cells SK-HEP-1 cells MiHa cells and furthermore induced miR-16 and suppressed its target gene Bcl-2 in HepG2 cells; Moreover the enhancement effects of Pgp and miR-16 on radiation-induced apoptosis were counteracted by overexpression of Bcl-2;To study the expression of miR-16 and bcl-2 in T lymphoblastic lymphoma/leukemia T-LBL/ALL and its relationship to prognosis; The relationship of miR-16 and bcl-2 was significantP = 0.042χ2 = 4.147; The relationship of miR-16 and bcl-2 might suggested that gene regulation may be influenced by them;miR 15b and miR 16 modulate multidrug resistance by targeting BCL2 in human gastric cancer cells; The downregulation of miR-15b and miR-16 in SGC7901/VCR cells was concurrent with the upregulation of Bcl-2 protein; Taken together our findings suggest that miR-15b and miR-16 could play a role in the development of MDR in gastric cancer cells at least in part by modulation of apoptosis via targeting BCL2;We demonstrated that anti-apoptotic protein Bcl-2 was directly targeted miR-16 in paclitaxel resistant lung cancer cells; Combined overexpression of miR-16 and miR-17 greatly reduced Beclin-1 and Bcl-2 expressions respectively; miR-17 overexpression reduced cytoprotective autophagy by targeting Beclin-1 whereas overexpression of miR-16 potentiated paclitaxel induced apoptotic cell death by inhibiting anti-apoptotic protein Bcl-2;The miR-16 expression correlated with BCL-2 protein r = 0.51 P < 0.05;MicroRNAs miRNAs are noncoding small RNAs that repress protein translation by targeting specific messenger RNAs miR-15a and miR-16-1 act as putative tumor suppressors by targeting the oncogene BCL2;The overall objective of our investigation was to assess whether miRNA-16 miR-16 is involved in the regulation of critical genes such as BCL2 that control the sensitivity of pancreatic cancer cells to apoptosis; This study showed that the ectopic overexpression of miR-16 may be therapeutically beneficial as is evidenced by impaired cell survival with concomitant attenuation of anti-apoptotic protein Bcl-2; Moreover the luciferase reporter assay suggested that miR-16 post-transcriptionally regulates Bcl-2 expression in pancreatic cancer cells through the target sites of the 3' untranslated region of this gene;miR 15a and miR 16 modulate drug resistance by targeting bcl 2 in human colon cancer cells; To investigate the reversal effect of targeted modulation of bcl-2 expression by miR-15a and miR-16 on drug resistance of human colon cancer cells |
| 73 | hsa-miR-17-5p | BCL2 | 2.27 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase | -0.32 | 0 | 25435430 | Combined overexpression of miR-16 and miR-17 greatly reduced Beclin-1 and Bcl-2 expressions respectively; miR-17 overexpression reduced cytoprotective autophagy by targeting Beclin-1 whereas overexpression of miR-16 potentiated paclitaxel induced apoptotic cell death by inhibiting anti-apoptotic protein Bcl-2 |
| 74 | hsa-miR-182-5p | BCL2 | 5.18 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase; mirMAP | -0.27 | 0 | 23936432; 26870290 | Inhibition of proliferation and induction of autophagy by atorvastatin in PC3 prostate cancer cells correlate with downregulation of Bcl2 and upregulation of miR 182 and p21; Bcl2 and p21 were identified to be potential target genes of miR-182 in PC3 cells;The expression levels of B-cell lymphoma-2 Bcl-2 and microRNA-182 miR-182 were detected using western blot analysis and quantitative reverse transcription-polymerase chain reaction respectively; Mangiferin treatment was also able to significantly reduce Bcl-2 expression levels and enhance miR-182 expression in PC3 cells; Finally it was observed that mangiferin inhibited proliferation and induced apoptosis in PC3 human prostate cancer cells and this effect was correlated with downregulation of Bcl-2 and upregulation of miR-182 |
| 75 | hsa-miR-186-5p | BCL2 | 1.01 | 0 | -2.27 | 0 | mirMAP | -0.47 | 0 | NA | |
| 76 | hsa-miR-192-5p | BCL2 | 1.98 | 0 | -2.27 | 0 | miRNAWalker2 validate | -0.31 | 0 | 26550150 | MicroRNA 192 regulates chemo resistance of lung adenocarcinoma for gemcitabine and cisplatin combined therapy by targeting Bcl 2; In this paper we try to test whether miR-192 regulates chemo-resistance in human carcinoma A549 mice model by targeting Bcl-2; MTT assay real-time RT-PCR western blotting assay were used to investigate miR-192 expression levels cell viability ratio and Bcl-2 protein expression levels; Bcl-2 mRNA and protein expression levels up-regulated in miR-192 inhibitor treated tumor; Bcl-2 is a key regulator for miR-192 related chemotherapy resistance; In this study we demonstrate that miR-192 regulates chemoresistance for gemcitabine and cisplatin combined chemotherapy in human adenocarcinoma lung cancer A549 cells and Bcl-2 is the target of miR-192 |
| 77 | hsa-miR-200a-5p | BCL2 | 4.89 | 0 | -2.27 | 0 | mirMAP | -0.26 | 0 | NA | |
| 78 | hsa-miR-200b-3p | BCL2 | 4.57 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase; TargetScan; mirMAP | -0.23 | 0 | NA | |
| 79 | hsa-miR-200b-5p | BCL2 | 4.84 | 0 | -2.27 | 0 | mirMAP | -0.28 | 0 | NA | |
| 80 | hsa-miR-200c-3p | BCL2 | 4.2 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase; mirMAP | -0.23 | 0 | NA | |
| 81 | hsa-miR-20a-3p | BCL2 | 1.64 | 0 | -2.27 | 0 | mirMAP | -0.11 | 0.04522 | NA | |
| 82 | hsa-miR-20a-5p | BCL2 | 1.51 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase | -0.17 | 0.00288 | NA | |
| 83 | hsa-miR-21-5p | BCL2 | 1.51 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase | -0.32 | 1.0E-5 | 21468550; 25994220; 25381586; 26555418; 23359184; 22964582; 21376256 | BCL-2 up-regulation could be achieved by miR-21 overexpression which prevented T24 cells from apoptosis induced by doxorubicin; Furthermore the miR-21 induced BCL-2 up-regulation could be cancelled by the PI3K inhibitor LY294002;Meanwhile miR-21 loss reduced STAT3 and Bcl-2 activation causing an increase in the apoptosis of tumour cells in CAC mice;Changes in the sensitivity of osteosarcoma cells to CDDP were examined after transfection with miR-21 mimics or anti-miR-21 or bcl-2 siRNA in combination with CDDP;The expression of Bax Bcl-2 and miR-21 in parental and paclitaxel-resistant cells was detected by RT-PCR and Western blotting;Resveratrol induces apoptosis of pancreatic cancers cells by inhibiting miR 21 regulation of BCL 2 expression; We also used Western blot to measure BCL-2 protein levels after down-regulation of miR-21 expression; Besides down-regulation of miR-21 expression can inhibit BCL-2 expression in PANC-1 CFPAC-1 and MIA Paca-2 cells; Over-expression of miR-21 expression can reverse down-regulation of BCL-2 expression and apoptosis induced by resveratrol; In this study we demonstrated that the effect of resveratrol on apoptosis is due to inhibiting miR-21 regulation of BCL-2 expression;Tumors harvested from these lungs have elevated levels of oncogenic miRNAs miR-21 and miR-155; are deficient for p53-regulated miRNAs; and have heightened expression of miR-34 target genes such as Met and Bcl-2;Bcl 2 upregulation induced by miR 21 via a direct interaction is associated with apoptosis and chemoresistance in MIA PaCa 2 pancreatic cancer cells; However the roles and mechanisms of miRNA miR-21 in regulation of Bcl-2 in pancreatic cancer remain to be elucidated; Then luciferase activity was observed after miR-21 mimics and pRL-TK plasmids containing wild-type and mutant 3'UTRs of Bcl-2 mRNA were co-transfected; Cells transfected with miR-21 inhibitor revealed an opposite trend. There was a significant increase in luciferase activity in the cells transfected with the wild-type pRL-TK plasmid in contrast to those transfected with the mutant one indicating that miR-21 promotes Bcl-2 expression by binding directly to the 3'UTR of Bcl-2 mRNA; Upregulation of Bcl-2 directly induced by miR-21 is associated with apoptosis chemoresistance and proliferation of MIA PaCa-2 pancreatic cancer cells |
| 84 | hsa-miR-215-5p | BCL2 | 2.38 | 0 | -2.27 | 0 | miRNAWalker2 validate | -0.33 | 0 | NA | |
| 85 | hsa-miR-224-5p | BCL2 | 1.55 | 1.0E-5 | -2.27 | 0 | mirMAP | -0.1 | 0.00523 | 24796455 | In addition the expressions of Bcl2 mRNA and protein were 1.05 ± 0.04 and 0.21 ± 0.03 in the miR-224 ASO group significantly lower than that in the control group 4.87 ± 0.96 and 0.88 ± 0.09 P < 0.01 |
| 86 | hsa-miR-24-2-5p | BCL2 | 1.52 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase | -0.41 | 0 | NA | |
| 87 | hsa-miR-3065-5p | BCL2 | 2.75 | 0 | -2.27 | 0 | mirMAP | -0.27 | 0 | NA | |
| 88 | hsa-miR-32-3p | BCL2 | 2.4 | 0 | -2.27 | 0 | mirMAP | -0.32 | 0 | NA | |
| 89 | hsa-miR-335-3p | BCL2 | 1.32 | 0 | -2.27 | 0 | mirMAP | -0.17 | 0.00274 | NA | |
| 90 | hsa-miR-338-5p | BCL2 | 0.59 | 0.04471 | -2.27 | 0 | PITA | -0.17 | 6.0E-5 | NA | |
| 91 | hsa-miR-33a-5p | BCL2 | 1.05 | 4.0E-5 | -2.27 | 0 | mirMAP | -0.14 | 0.00486 | NA | |
| 92 | hsa-miR-33b-5p | BCL2 | 2.26 | 0 | -2.27 | 0 | miRTarBase; mirMAP | -0.23 | 0 | NA | |
| 93 | hsa-miR-342-3p | BCL2 | 0.91 | 0 | -2.27 | 0 | miRanda | -0.19 | 0.01633 | NA | |
| 94 | hsa-miR-365a-3p | BCL2 | 0.82 | 1.0E-5 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase | -0.33 | 0 | NA | |
| 95 | hsa-miR-375 | BCL2 | 2.57 | 0 | -2.27 | 0 | miRNAWalker2 validate | -0.06 | 0.00889 | 26381132; 26697569; 25613180 | The levels of miR-375 Bax and Bcl-2 protein expression in treated cells were determined by Western blot and RT-PCR; Moreover compared to control group the expression of Bcl-2 and miR-375 decreases with formononetin in the U2OS cells while Bax increases;Exosome Carried microRNA 375 Inhibits Cell Progression and Dissemination via Bcl 2 Blocking in Colon Cancer; RT-PCR for Bcl-2 expression showed that Bcl-2 is down-regulated for miR-375 inhibitor and up-regulated for the miR-375 mimic a result confirmed by Western blotting; The present study brings to the forefront new data that suggest miR-375 as a new player in controlling the pathways responsible for inhibiting the natural history of CRC tumor cells via the Bcl-2 pathway;mRNA levels of ERα Bcl-2 and miR-375 were quantified using real-time polymerase chain reaction; After treatment with biochanin A ERα miR-375 and Bcl-2 expression was significantly upregulated |
| 96 | hsa-miR-429 | BCL2 | 5.04 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase; PITA; mirMAP | -0.23 | 0 | 23999873; 26513239; 26511969 | MiR 429 up regulation induces apoptosis and suppresses invasion by targeting Bcl 2 and SP 1 in esophageal carcinoma; Subsequent Western blotting and luciferase reporter assays showed that miR-429 can bind to putative binding sites within the Bcl-2 and SP1 mRNA 3' untranslated regions UTRs to reduce their expression; Up-regulation of miR-429 inhibits invasion and promotes apoptosis in EC cells by targeting Bcl-2 and SP1; Our findings suggest that Bcl-2 and SP1 may serve as major targets of miR-429;MiR 429 Induces Gastric Carcinoma Cell Apoptosis Through Bcl 2; Here we studied the levels of miR-429 and anti-apoptotic protein Bcl-2 in GC specimens; We performed bioinformatics analyses and used luciferase-reporter assay to analyze the relationship between miR-429 and Bcl-2 in GC cells; MiR-429 levels were significantly decreased and Bcl-2 levels were significantly increased in GC specimens compared to the paired adjacent non-tumor gastric tissue; Moreover the levels of miR-429 and Bcl-2 inversely correlated in GC specimens; Bioinformatics analyses showed that miR-429 targeted the 3'-UTR of Bcl-2 mRNA to inhibit its translation which was confirmed by luciferase-reporter assay;MiR 429 induces apoptosis of glioblastoma cell through Bcl 2; Here we analyzed the levels of miR-429 and anti-apoptotic protein Bcl-2 in GBM specimens; We combined bioinformatics analyses and luciferase reporter assay to determine the relationship between miR-429 and Bcl-2 in GBM cells; We found that miR-429 levels were significantly decreased and Bcl-2 levels were significantly increased in GBM specimens compared to the paired adjacent non-tumor brain tissue; Moreover the levels of miR-429 and Bcl-2 inversely correlated; MiR-429 targeted the 3'-UTR of Bcl-2 mRNA to inhibit its translation |
| 97 | hsa-miR-577 | BCL2 | 3.23 | 0 | -2.27 | 0 | PITA | -0.12 | 2.0E-5 | NA | |
| 98 | hsa-miR-582-5p | BCL2 | -0.13 | 0.57451 | -2.27 | 0 | PITA | -0.14 | 0.00591 | NA | |
| 99 | hsa-miR-590-3p | BCL2 | 1.73 | 0 | -2.27 | 0 | miRanda; mirMAP | -0.44 | 0 | NA | |
| 100 | hsa-miR-590-5p | BCL2 | 1.46 | 0 | -2.27 | 0 | miRanda | -0.46 | 0 | NA | |
| 101 | hsa-miR-616-5p | BCL2 | 1.97 | 0 | -2.27 | 0 | mirMAP | -0.31 | 0 | NA | |
| 102 | hsa-miR-629-5p | BCL2 | 1.41 | 0 | -2.27 | 0 | mirMAP | -0.22 | 4.0E-5 | NA | |
| 103 | hsa-miR-7-1-3p | BCL2 | 1.84 | 0 | -2.27 | 0 | mirMAP | -0.21 | 0.00021 | NA | |
| 104 | hsa-miR-7-5p | BCL2 | 3.57 | 0 | -2.27 | 0 | miRNAWalker2 validate; miRTarBase; mirMAP | -0.24 | 0 | 26464649; 25862909; 21750649 | Western blotting was used to evaluate the effect of miR-7 on Bcl2 in A549 and H460 cells; Moreover subsequent experiments showed that BCL-2 was downregulated by miR-7 at both transcriptional and translational levels; This study further extends the biological role of miR-7 in NSCLC A549 and H460 cells and identifies BCL-2 as a novel target possibly involved in miR-7-mediated growth suppression and apoptosis induction of NSCLC cells;miR-7 overexpression correlated with diminished BCL2 expression but there was no relationship between miR-7 and EGFR expression neither in tumour samples nor in the cell lines; Of the two postulated miR-7 target genes we examined BCL2 but not EGFR seems to be a possible miR-7 target in OC;Bioinformatics predictions revealed a potential binding site of miR-7 on 3'UTR of BCL-2 and it was further confirmed by luciferase assay; Moreover subsequent experiments showed that BCL-2 was downregulated by miR-7 at both transcriptional and translational levels; These results suggest that miR-7 regulates the expression of BCL-2 through direct 3'UTR interactions |
| 105 | hsa-miR-96-5p | BCL2 | 5.73 | 0 | -2.27 | 0 | miRNAWalker2 validate; TargetScan | -0.26 | 0 | NA | |
| 106 | hsa-let-7a-5p | BCL2L1 | 0.1 | 0.43289 | 0.65 | 0 | TargetScan; miRNATAP | -0.2 | 1.0E-5 | 26915294; 20347499 | As a result transcript levels of the tumor-suppressive miR-15 and let-7 families increased which targeted and decreased the expression of the crucial prosurvival genes BCL-2 and BCL-XL respectively;The let 7 family of microRNAs inhibits Bcl xL expression and potentiates sorafenib induced apoptosis in human hepatocellular carcinoma; The effect of let-7 on Bcl-xL expression was examined by Western blot and a reporter assay; Microarray analysis followed by in silico target prediction identified let-7 microRNAs as being downregulated in Huh7 hepatoma cells in comparison with primary human hepatocytes as well as possessing a putative target site in the bcl-xl mRNA |
| 107 | hsa-let-7b-5p | BCL2L1 | -0.07 | 0.65185 | 0.65 | 0 | miRNATAP | -0.12 | 0.00057 | 26915294; 20347499 | As a result transcript levels of the tumor-suppressive miR-15 and let-7 families increased which targeted and decreased the expression of the crucial prosurvival genes BCL-2 and BCL-XL respectively;The let 7 family of microRNAs inhibits Bcl xL expression and potentiates sorafenib induced apoptosis in human hepatocellular carcinoma; The effect of let-7 on Bcl-xL expression was examined by Western blot and a reporter assay; Microarray analysis followed by in silico target prediction identified let-7 microRNAs as being downregulated in Huh7 hepatoma cells in comparison with primary human hepatocytes as well as possessing a putative target site in the bcl-xl mRNA |
| 108 | hsa-let-7g-5p | BCL2L1 | 1.16 | 0 | 0.65 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.07 | 0.03609 | 20347499 | Over-expression of let-7c or let-7g led to a clear decrease of Bcl-xL expression in Huh7 and HepG2 cell lines; Reporter assays revealed direct post-transcriptional regulation involving let-7c or let-7g and the 3'-untranslated region of bcl-xl mRNA |
| 109 | hsa-miR-140-5p | BCL2L1 | -0.89 | 0 | 0.65 | 0 | PITA; miRanda; miRNATAP | -0.1 | 0.00286 | NA | |
| 110 | hsa-miR-664a-5p | BCL2L1 | 0.23 | 0.25436 | 0.65 | 0 | mirMAP | -0.08 | 0.0042 | NA | |
| 111 | hsa-miR-16-2-3p | BIRC3 | 2.34 | 0 | -0.22 | 0.47948 | mirMAP | -0.13 | 0.04984 | NA | |
| 112 | hsa-miR-374b-5p | BIRC3 | -0.16 | 0.22606 | -0.22 | 0.47948 | mirMAP | -0.22 | 0.04644 | NA | |
| 113 | hsa-miR-651-5p | BIRC3 | 1.64 | 0 | -0.22 | 0.47948 | MirTarget | -0.17 | 0.00447 | NA | |
| 114 | hsa-miR-98-5p | BIRC3 | 1.06 | 0 | -0.22 | 0.47948 | miRNAWalker2 validate | -0.28 | 0.00772 | NA | |
| 115 | hsa-miR-107 | CAPN2 | 1.81 | 0 | -0.79 | 0 | miRanda | -0.19 | 0 | NA | |
| 116 | hsa-miR-16-2-3p | CAPN2 | 2.34 | 0 | -0.79 | 0 | mirMAP | -0.18 | 0 | NA | |
| 117 | hsa-miR-20a-3p | CAPN2 | 1.64 | 0 | -0.79 | 0 | MirTarget | -0.15 | 0 | NA | |
| 118 | hsa-miR-30a-5p | CAPN2 | 0.2 | 0.42032 | -0.79 | 0 | miRNAWalker2 validate | -0.08 | 0.00167 | NA | |
| 119 | hsa-miR-320a | CAPN2 | 0.5 | 0.00226 | -0.79 | 0 | miRanda | -0.11 | 0.0029 | NA | |
| 120 | hsa-miR-320b | CAPN2 | 1.1 | 0 | -0.79 | 0 | miRanda | -0.12 | 2.0E-5 | NA | |
| 121 | hsa-miR-320c | CAPN2 | 0.73 | 0.00058 | -0.79 | 0 | miRanda | -0.12 | 3.0E-5 | NA | |
| 122 | hsa-miR-421 | CAPN2 | 2.1 | 0 | -0.79 | 0 | miRanda | -0.17 | 0 | NA | |
| 123 | hsa-miR-590-3p | CAPN2 | 1.73 | 0 | -0.79 | 0 | miRanda | -0.11 | 0.00028 | NA | |
| 124 | hsa-miR-590-5p | CAPN2 | 1.46 | 0 | -0.79 | 0 | miRanda | -0.1 | 0.00306 | NA | |
| 125 | hsa-miR-7-5p | CAPN2 | 3.57 | 0 | -0.79 | 0 | miRNAWalker2 validate | -0.11 | 0 | NA | |
| 126 | hsa-miR-16-2-3p | CASP10 | 2.34 | 0 | -0.72 | 0.00202 | mirMAP | -0.14 | 0.00675 | NA | |
| 127 | hsa-miR-181c-5p | CASP10 | 0.6 | 0.01077 | -0.72 | 0.00202 | mirMAP | -0.11 | 0.02227 | NA | |
| 128 | hsa-miR-186-5p | CASP10 | 1.01 | 0 | -0.72 | 0.00202 | MirTarget; mirMAP | -0.23 | 0.00505 | NA | |
| 129 | hsa-miR-195-3p | CASP10 | -1.78 | 0 | -0.72 | 0.00202 | mirMAP | -0.12 | 0.0036 | NA | |
| 130 | hsa-miR-19a-3p | CASP10 | 1.85 | 0 | -0.72 | 0.00202 | MirTarget; mirMAP | -0.14 | 0.0111 | NA | |
| 131 | hsa-miR-19b-3p | CASP10 | 1.34 | 0 | -0.72 | 0.00202 | MirTarget; mirMAP | -0.22 | 0.00084 | NA | |
| 132 | hsa-miR-200b-3p | CASP10 | 4.57 | 0 | -0.72 | 0.00202 | mirMAP | -0.12 | 0.00031 | NA | |
| 133 | hsa-miR-200c-3p | CASP10 | 4.2 | 0 | -0.72 | 0.00202 | mirMAP | -0.09 | 0.00801 | NA | |
| 134 | hsa-miR-30b-3p | CASP10 | 0.97 | 0 | -0.72 | 0.00202 | MirTarget | -0.2 | 0.00064 | NA | |
| 135 | hsa-miR-320a | CASP10 | 0.5 | 0.00226 | -0.72 | 0.00202 | miRanda | -0.21 | 0.0015 | NA | |
| 136 | hsa-miR-320b | CASP10 | 1.1 | 0 | -0.72 | 0.00202 | miRanda | -0.18 | 0.00058 | NA | |
| 137 | hsa-miR-320c | CASP10 | 0.73 | 0.00058 | -0.72 | 0.00202 | miRanda | -0.15 | 0.00479 | NA | |
| 138 | hsa-miR-33a-3p | CASP10 | 1.39 | 0 | -0.72 | 0.00202 | mirMAP | -0.27 | 0 | NA | |
| 139 | hsa-miR-3614-5p | CASP10 | 3.22 | 0 | -0.72 | 0.00202 | mirMAP | -0.06 | 0.04578 | NA | |
| 140 | hsa-miR-429 | CASP10 | 5.04 | 0 | -0.72 | 0.00202 | mirMAP; miRNATAP | -0.09 | 0.0024 | NA | |
| 141 | hsa-miR-532-5p | CASP10 | 0.31 | 0.06801 | -0.72 | 0.00202 | mirMAP | -0.22 | 0.00094 | NA | |
| 142 | hsa-miR-589-3p | CASP10 | 1.7 | 0 | -0.72 | 0.00202 | mirMAP | -0.2 | 1.0E-5 | NA | |
| 143 | hsa-miR-616-5p | CASP10 | 1.97 | 0 | -0.72 | 0.00202 | mirMAP | -0.11 | 0.01291 | NA | |
| 144 | hsa-miR-744-3p | CASP10 | 2.12 | 0 | -0.72 | 0.00202 | mirMAP | -0.29 | 0 | NA | |
| 145 | hsa-miR-766-3p | CASP10 | 2.7 | 0 | -0.72 | 0.00202 | mirMAP | -0.1 | 0.04345 | NA | |
| 146 | hsa-let-7c-5p | CASP3 | -1.72 | 0 | 0.88 | 0 | MirTarget | -0.11 | 0 | NA | |
| 147 | hsa-miR-101-3p | CASP3 | -1.48 | 0 | 0.88 | 0 | MirTarget | -0.2 | 0 | NA | |
| 148 | hsa-miR-139-5p | CASP3 | -2.41 | 0 | 0.88 | 0 | miRanda | -0.13 | 0 | NA | |
| 149 | hsa-miR-140-5p | CASP3 | -0.89 | 0 | 0.88 | 0 | miRanda | -0.13 | 2.0E-5 | NA | |
| 150 | hsa-miR-195-3p | CASP3 | -1.78 | 0 | 0.88 | 0 | MirTarget | -0.11 | 0 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 50 | 1929 | 6.578e-35 | 3.061e-31 |
| 2 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 22 | 99 | 1.609e-34 | 3.743e-31 |
| 3 | CELL DEATH | 40 | 1001 | 1.274e-33 | 1.976e-30 |
| 4 | APOPTOTIC SIGNALING PATHWAY | 28 | 289 | 2.112e-33 | 2.456e-30 |
| 5 | INTRACELLULAR SIGNAL TRANSDUCTION | 45 | 1572 | 2.552e-32 | 2.375e-29 |
| 6 | REGULATION OF CELL DEATH | 44 | 1472 | 3.153e-32 | 2.445e-29 |
| 7 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 46 | 1848 | 1.555e-30 | 1.034e-27 |
| 8 | POSITIVE REGULATION OF CELL COMMUNICATION | 43 | 1532 | 3.363e-30 | 1.889e-27 |
| 9 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 16 | 39 | 3.653e-30 | 1.889e-27 |
| 10 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 42 | 1492 | 2.133e-29 | 9.927e-27 |
| 11 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 44 | 1791 | 1.256e-28 | 5.312e-26 |
| 12 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 34 | 876 | 1.545e-27 | 5.992e-25 |
| 13 | ACTIVATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 18 | 95 | 7e-27 | 2.505e-24 |
| 14 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 22 | 213 | 1.043e-26 | 3.372e-24 |
| 15 | POSITIVE REGULATION OF I KAPPAB KINASE NF KAPPAB SIGNALING | 21 | 179 | 1.087e-26 | 3.372e-24 |
| 16 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 41 | 1656 | 2.238e-26 | 6.507e-24 |
| 17 | NEGATIVE REGULATION OF CELL DEATH | 33 | 872 | 2.713e-26 | 7.424e-24 |
| 18 | ACTIVATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 11 | 13 | 3.882e-26 | 1.003e-23 |
| 19 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 38 | 1381 | 9.575e-26 | 2.345e-23 |
| 20 | ZYMOGEN ACTIVATION | 18 | 112 | 1.711e-25 | 3.982e-23 |
| 21 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 39 | 1518 | 1.948e-25 | 4.316e-23 |
| 22 | RESPONSE TO CYTOKINE | 30 | 714 | 5.04e-25 | 1.066e-22 |
| 23 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 17 | 98 | 1.052e-24 | 2.127e-22 |
| 24 | IMMUNE SYSTEM PROCESS | 42 | 1984 | 1.821e-24 | 3.53e-22 |
| 25 | POSITIVE REGULATION OF CELL DEATH | 28 | 605 | 2.132e-24 | 3.969e-22 |
| 26 | REGULATION OF I KAPPAB KINASE NF KAPPAB SIGNALING | 21 | 233 | 3.243e-24 | 5.804e-22 |
| 27 | POSITIVE REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 11 | 17 | 6.092e-24 | 1.05e-21 |
| 28 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 18 | 153 | 6.336e-23 | 1.053e-20 |
| 29 | POSITIVE REGULATION OF PEPTIDASE ACTIVITY | 18 | 154 | 7.157e-23 | 1.148e-20 |
| 30 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 11 | 22 | 3.425e-22 | 5.312e-20 |
| 31 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 22 | 363 | 1.454e-21 | 2.183e-19 |
| 32 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 32 | 1135 | 1.636e-21 | 2.378e-19 |
| 33 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 36 | 1618 | 4.52e-21 | 6.373e-19 |
| 34 | REGULATION OF PEPTIDASE ACTIVITY | 22 | 392 | 7.731e-21 | 1.058e-18 |
| 35 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 18 | 200 | 8.99e-21 | 1.195e-18 |
| 36 | CELLULAR RESPONSE TO CYTOKINE STIMULUS | 25 | 606 | 1.804e-20 | 2.332e-18 |
| 37 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 30 | 1036 | 2.223e-20 | 2.653e-18 |
| 38 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 30 | 1036 | 2.223e-20 | 2.653e-18 |
| 39 | POSITIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 17 | 171 | 2.154e-20 | 2.653e-18 |
| 40 | REGULATION OF TRANSFERASE ACTIVITY | 29 | 946 | 2.506e-20 | 2.915e-18 |
| 41 | REGULATION OF KINASE ACTIVITY | 27 | 776 | 2.963e-20 | 3.362e-18 |
| 42 | PROTEIN MATURATION | 19 | 265 | 5.14e-20 | 5.694e-18 |
| 43 | REGULATION OF IMMUNE SYSTEM PROCESS | 33 | 1403 | 7.964e-20 | 8.618e-18 |
| 44 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 11 | 34 | 1.344e-19 | 1.421e-17 |
| 45 | CYTOKINE MEDIATED SIGNALING PATHWAY | 22 | 452 | 1.667e-19 | 1.723e-17 |
| 46 | REGULATION OF PROTEIN MODIFICATION PROCESS | 35 | 1710 | 3.125e-19 | 3.161e-17 |
| 47 | REGULATION OF IMMUNE RESPONSE | 27 | 858 | 3.892e-19 | 3.853e-17 |
| 48 | RESPONSE TO NITROGEN COMPOUND | 27 | 859 | 4.01e-19 | 3.887e-17 |
| 49 | POSITIVE REGULATION OF KINASE ACTIVITY | 22 | 482 | 6.59e-19 | 6.258e-17 |
| 50 | REGULATION OF PROTEOLYSIS | 25 | 711 | 8.314e-19 | 7.737e-17 |
| 51 | POSITIVE REGULATION OF PROTEOLYSIS | 20 | 363 | 8.667e-19 | 7.907e-17 |
| 52 | POSITIVE REGULATION OF IMMUNE RESPONSE | 23 | 563 | 1.051e-18 | 9.409e-17 |
| 53 | PHOSPHORYLATION | 30 | 1228 | 2.609e-18 | 2.29e-16 |
| 54 | ACTIVATION OF PROTEIN KINASE ACTIVITY | 18 | 279 | 3.699e-18 | 3.187e-16 |
| 55 | RESPONSE TO TUMOR NECROSIS FACTOR | 17 | 233 | 4.419e-18 | 3.739e-16 |
| 56 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 21 | 470 | 7.369e-18 | 6.123e-16 |
| 57 | I KAPPAB KINASE NF KAPPAB SIGNALING | 12 | 70 | 1.364e-17 | 1.113e-15 |
| 58 | ACTIVATION OF IMMUNE RESPONSE | 20 | 427 | 2.064e-17 | 1.656e-15 |
| 59 | RESPONSE TO MOLECULE OF BACTERIAL ORIGIN | 18 | 321 | 4.47e-17 | 3.525e-15 |
| 60 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 11 | 55 | 5.304e-17 | 4.113e-15 |
| 61 | PROTEIN PHOSPHORYLATION | 26 | 944 | 5.503e-17 | 4.198e-15 |
| 62 | POSITIVE REGULATION OF IMMUNE SYSTEM PROCESS | 25 | 867 | 8.948e-17 | 6.611e-15 |
| 63 | REGULATION OF RESPONSE TO CYTOKINE STIMULUS | 14 | 144 | 8.951e-17 | 6.611e-15 |
| 64 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 22 | 616 | 1.173e-16 | 8.528e-15 |
| 65 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 24 | 799 | 1.741e-16 | 1.246e-14 |
| 66 | TUMOR NECROSIS FACTOR MEDIATED SIGNALING PATHWAY | 13 | 118 | 2.322e-16 | 1.637e-14 |
| 67 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 34 | 1977 | 2.778e-16 | 1.929e-14 |
| 68 | CELLULAR RESPONSE TO NITROGEN COMPOUND | 20 | 505 | 5.233e-16 | 3.581e-14 |
| 69 | POSITIVE REGULATION OF NF KAPPAB TRANSCRIPTION FACTOR ACTIVITY | 13 | 132 | 1.035e-15 | 6.982e-14 |
| 70 | REGULATION OF TUMOR NECROSIS FACTOR MEDIATED SIGNALING PATHWAY | 10 | 50 | 1.507e-15 | 1.002e-13 |
| 71 | RESPONSE TO BIOTIC STIMULUS | 24 | 886 | 1.755e-15 | 1.15e-13 |
| 72 | POSITIVE REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 15 | 228 | 2.336e-15 | 1.51e-13 |
| 73 | NEURON APOPTOTIC PROCESS | 9 | 35 | 3.472e-15 | 2.213e-13 |
| 74 | RESPONSE TO ABIOTIC STIMULUS | 25 | 1024 | 4.213e-15 | 2.649e-13 |
| 75 | INTRINSIC APOPTOTIC SIGNALING PATHWAY | 13 | 152 | 6.668e-15 | 4.137e-13 |
| 76 | POSITIVE REGULATION OF DEFENSE RESPONSE | 17 | 364 | 7.708e-15 | 4.719e-13 |
| 77 | TOLL LIKE RECEPTOR SIGNALING PATHWAY | 11 | 85 | 8.706e-15 | 5.261e-13 |
| 78 | INFLAMMATORY RESPONSE | 18 | 454 | 1.885e-14 | 1.124e-12 |
| 79 | CELLULAR RESPONSE TO ABIOTIC STIMULUS | 15 | 263 | 1.933e-14 | 1.138e-12 |
| 80 | REGULATION OF HYDROLASE ACTIVITY | 27 | 1327 | 2.053e-14 | 1.194e-12 |
| 81 | RESPONSE TO ENDOGENOUS STIMULUS | 28 | 1450 | 2.086e-14 | 1.199e-12 |
| 82 | REGULATION OF RESPONSE TO STRESS | 28 | 1468 | 2.841e-14 | 1.612e-12 |
| 83 | POSITIVE REGULATION OF HYDROLASE ACTIVITY | 23 | 905 | 3.055e-14 | 1.713e-12 |
| 84 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 9 | 46 | 5.253e-14 | 2.91e-12 |
| 85 | NEURON DEATH | 9 | 47 | 6.479e-14 | 3.547e-12 |
| 86 | RESPONSE TO EXTERNAL STIMULUS | 30 | 1821 | 1.123e-13 | 6.077e-12 |
| 87 | PATTERN RECOGNITION RECEPTOR SIGNALING PATHWAY | 11 | 109 | 1.465e-13 | 7.837e-12 |
| 88 | IMMUNE RESPONSE | 24 | 1100 | 2.026e-13 | 1.071e-11 |
| 89 | SIGNAL TRANSDUCTION IN ABSENCE OF LIGAND | 8 | 33 | 2.172e-13 | 1.123e-11 |
| 90 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 8 | 33 | 2.172e-13 | 1.123e-11 |
| 91 | CELLULAR RESPONSE TO MECHANICAL STIMULUS | 10 | 80 | 2.223e-13 | 1.137e-11 |
| 92 | REGULATION OF NEURON DEATH | 14 | 252 | 2.251e-13 | 1.138e-11 |
| 93 | RESPONSE TO BACTERIUM | 18 | 528 | 2.479e-13 | 1.24e-11 |
| 94 | ACTIVATION OF INNATE IMMUNE RESPONSE | 13 | 204 | 3.045e-13 | 1.507e-11 |
| 95 | IMMUNE RESPONSE REGULATING CELL SURFACE RECEPTOR SIGNALING PATHWAY | 15 | 323 | 3.881e-13 | 1.901e-11 |
| 96 | CELLULAR RESPONSE TO STRESS | 27 | 1565 | 1.07e-12 | 5.188e-11 |
| 97 | PROTEOLYSIS | 24 | 1208 | 1.517e-12 | 7.279e-11 |
| 98 | HOMEOSTATIC PROCESS | 25 | 1337 | 1.71e-12 | 8.036e-11 |
| 99 | POSITIVE REGULATION OF GENE EXPRESSION | 28 | 1733 | 1.702e-12 | 8.036e-11 |
| 100 | RESPONSE TO LIPID | 21 | 888 | 2.074e-12 | 9.65e-11 |
| 101 | REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 15 | 365 | 2.257e-12 | 1.04e-10 |
| 102 | FC EPSILON RECEPTOR SIGNALING PATHWAY | 11 | 142 | 2.779e-12 | 1.268e-10 |
| 103 | REGULATION OF NECROTIC CELL DEATH | 7 | 26 | 3.269e-12 | 1.477e-10 |
| 104 | POSITIVE REGULATION OF INNATE IMMUNE RESPONSE | 13 | 246 | 3.315e-12 | 1.483e-10 |
| 105 | NECROTIC CELL DEATH | 7 | 28 | 5.852e-12 | 2.593e-10 |
| 106 | PHOSPHATIDYLINOSITOL 3 PHOSPHATE BIOSYNTHETIC PROCESS | 8 | 49 | 6.745e-12 | 2.961e-10 |
| 107 | T CELL APOPTOTIC PROCESS | 6 | 15 | 7.87e-12 | 3.422e-10 |
| 108 | CELLULAR RESPONSE TO EXTERNAL STIMULUS | 13 | 264 | 8.084e-12 | 3.483e-10 |
| 109 | RESPONSE TO WOUNDING | 17 | 563 | 8.728e-12 | 3.726e-10 |
| 110 | MYD88 INDEPENDENT TOLL LIKE RECEPTOR SIGNALING PATHWAY | 7 | 30 | 1e-11 | 4.232e-10 |
| 111 | REGULATION OF DEFENSE RESPONSE | 19 | 759 | 1.097e-11 | 4.598e-10 |
| 112 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 7 | 32 | 1.645e-11 | 6.773e-10 |
| 113 | NEGATIVE REGULATION OF SIGNAL TRANSDUCTION IN ABSENCE OF LIGAND | 7 | 32 | 1.645e-11 | 6.773e-10 |
| 114 | EXECUTION PHASE OF APOPTOSIS | 8 | 55 | 1.791e-11 | 7.309e-10 |
| 115 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 24 | 1360 | 1.861e-11 | 7.528e-10 |
| 116 | REGULATION OF CELL PROLIFERATION | 25 | 1496 | 1.997e-11 | 8.009e-10 |
| 117 | RESPONSE TO HORMONE | 20 | 893 | 2.091e-11 | 8.315e-10 |
| 118 | REGULATION OF INNATE IMMUNE RESPONSE | 14 | 357 | 2.483e-11 | 9.789e-10 |
| 119 | POSITIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 13 | 289 | 2.514e-11 | 9.83e-10 |
| 120 | T CELL HOMEOSTASIS | 7 | 34 | 2.614e-11 | 1.014e-09 |
| 121 | POSITIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 14 | 360 | 2.775e-11 | 1.067e-09 |
| 122 | LYMPHOCYTE APOPTOTIC PROCESS | 6 | 18 | 2.895e-11 | 1.104e-09 |
| 123 | POSITIVE REGULATION OF INTRACELLULAR TRANSPORT | 14 | 370 | 3.991e-11 | 1.51e-09 |
| 124 | NEGATIVE REGULATION OF CELL COMMUNICATION | 22 | 1192 | 7.015e-11 | 2.632e-09 |
| 125 | WOUND HEALING | 15 | 470 | 8.081e-11 | 2.984e-09 |
| 126 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 21 | 1079 | 8.039e-11 | 2.984e-09 |
| 127 | REGULATION OF MAP KINASE ACTIVITY | 13 | 319 | 8.587e-11 | 3.146e-09 |
| 128 | ANTIGEN RECEPTOR MEDIATED SIGNALING PATHWAY | 11 | 195 | 8.772e-11 | 3.189e-09 |
| 129 | STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 9 | 103 | 1.02e-10 | 3.68e-09 |
| 130 | CHEMICAL HOMEOSTASIS | 19 | 874 | 1.229e-10 | 4.399e-09 |
| 131 | RESPONSE TO PEPTIDE | 14 | 404 | 1.273e-10 | 4.523e-09 |
| 132 | DEFENSE RESPONSE | 22 | 1231 | 1.301e-10 | 4.587e-09 |
| 133 | INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO DNA DAMAGE | 8 | 71 | 1.496e-10 | 5.234e-09 |
| 134 | LEUKOCYTE APOPTOTIC PROCESS | 6 | 23 | 1.552e-10 | 5.39e-09 |
| 135 | FC RECEPTOR SIGNALING PATHWAY | 11 | 206 | 1.581e-10 | 5.448e-09 |
| 136 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 20 | 1008 | 1.806e-10 | 6.18e-09 |
| 137 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 26 | 1805 | 1.852e-10 | 6.291e-09 |
| 138 | REGULATION OF EXECUTION PHASE OF APOPTOSIS | 6 | 24 | 2.064e-10 | 6.959e-09 |
| 139 | CELLULAR RESPONSE TO PEPTIDE | 12 | 274 | 2.146e-10 | 7.183e-09 |
| 140 | PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 6 | 25 | 2.708e-10 | 9.001e-09 |
| 141 | POSITIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 15 | 514 | 2.804e-10 | 9.253e-09 |
| 142 | PHOSPHATIDYLINOSITOL BIOSYNTHETIC PROCESS | 9 | 120 | 4.053e-10 | 1.328e-08 |
| 143 | POSITIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 9 | 121 | 4.367e-10 | 1.421e-08 |
| 144 | LYMPHOCYTE HOMEOSTASIS | 7 | 50 | 4.641e-10 | 1.5e-08 |
| 145 | POSITIVE REGULATION OF TRANSCRIPTION FACTOR IMPORT INTO NUCLEUS | 7 | 51 | 5.364e-10 | 1.71e-08 |
| 146 | NIK NF KAPPAB SIGNALING | 8 | 83 | 5.363e-10 | 1.71e-08 |
| 147 | INOSITOL LIPID MEDIATED SIGNALING | 9 | 124 | 5.439e-10 | 1.722e-08 |
| 148 | NUCLEOTIDE BINDING DOMAIN LEUCINE RICH REPEAT CONTAINING RECEPTOR SIGNALING PATHWAY | 6 | 28 | 5.713e-10 | 1.796e-08 |
| 149 | INTERLEUKIN 1 MEDIATED SIGNALING PATHWAY | 5 | 13 | 6.15e-10 | 1.92e-08 |
| 150 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 15 | 552 | 7.489e-10 | 2.323e-08 |
| 151 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 9 | 129 | 7.746e-10 | 2.387e-08 |
| 152 | POSITIVE REGULATION OF MAPK CASCADE | 14 | 470 | 9.152e-10 | 2.802e-08 |
| 153 | REGULATION OF MAPK CASCADE | 16 | 660 | 9.779e-10 | 2.974e-08 |
| 154 | CELL ACTIVATION | 15 | 568 | 1.107e-09 | 3.346e-08 |
| 155 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 19 | 1004 | 1.258e-09 | 3.778e-08 |
| 156 | INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO ENDOPLASMIC RETICULUM STRESS | 6 | 32 | 1.359e-09 | 4.053e-08 |
| 157 | REGULATION OF NEURON APOPTOTIC PROCESS | 10 | 192 | 1.461e-09 | 4.329e-08 |
| 158 | LEUKOCYTE CELL CELL ADHESION | 11 | 255 | 1.521e-09 | 4.48e-08 |
| 159 | CYTOPLASMIC PATTERN RECOGNITION RECEPTOR SIGNALING PATHWAY | 6 | 33 | 1.656e-09 | 4.847e-08 |
| 160 | LEUKOCYTE HOMEOSTASIS | 7 | 60 | 1.745e-09 | 5.074e-08 |
| 161 | T CELL RECEPTOR SIGNALING PATHWAY | 9 | 146 | 2.328e-09 | 6.728e-08 |
| 162 | POSITIVE REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 6 | 36 | 2.889e-09 | 8.296e-08 |
| 163 | ACTIVATION OF PROTEIN KINASE A ACTIVITY | 5 | 17 | 2.925e-09 | 8.348e-08 |
| 164 | POSITIVE REGULATION OF MAP KINASE ACTIVITY | 10 | 207 | 3.025e-09 | 8.582e-08 |
| 165 | POSITIVE REGULATION OF PROTEIN IMPORT | 8 | 104 | 3.301e-09 | 9.31e-08 |
| 166 | NEGATIVE REGULATION OF CATALYTIC ACTIVITY | 17 | 829 | 3.379e-09 | 9.472e-08 |
| 167 | RESPONSE TO MECHANICAL STIMULUS | 10 | 210 | 3.475e-09 | 9.682e-08 |
| 168 | RESPONSE TO OXIDATIVE STRESS | 12 | 352 | 3.695e-09 | 1.023e-07 |
| 169 | POSITIVE REGULATION OF NEURON DEATH | 7 | 67 | 3.853e-09 | 1.061e-07 |
| 170 | REGULATION OF CATABOLIC PROCESS | 16 | 731 | 4.227e-09 | 1.157e-07 |
| 171 | REGULATION OF LIPID METABOLIC PROCESS | 11 | 282 | 4.362e-09 | 1.187e-07 |
| 172 | REGULATION OF MEMBRANE PERMEABILITY | 7 | 70 | 5.266e-09 | 1.425e-07 |
| 173 | LEUKOCYTE DIFFERENTIATION | 11 | 292 | 6.268e-09 | 1.686e-07 |
| 174 | CELLULAR RESPONSE TO HORMONE STIMULUS | 14 | 552 | 7.169e-09 | 1.917e-07 |
| 175 | CELLULAR GLUCOSE HOMEOSTASIS | 7 | 75 | 8.598e-09 | 2.286e-07 |
| 176 | REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 6 | 43 | 8.866e-09 | 2.331e-07 |
| 177 | CELLULAR COMPONENT DISASSEMBLY INVOLVED IN EXECUTION PHASE OF APOPTOSIS | 6 | 43 | 8.866e-09 | 2.331e-07 |
| 178 | NECROPTOTIC PROCESS | 5 | 21 | 9.512e-09 | 2.486e-07 |
| 179 | RESPONSE TO INORGANIC SUBSTANCE | 13 | 479 | 1.19e-08 | 3.092e-07 |
| 180 | RESPONSE TO OXYGEN LEVELS | 11 | 311 | 1.204e-08 | 3.094e-07 |
| 181 | HEMOSTASIS | 11 | 311 | 1.204e-08 | 3.094e-07 |
| 182 | RESPONSE TO TOXIC SUBSTANCE | 10 | 241 | 1.299e-08 | 3.321e-07 |
| 183 | POSITIVE REGULATION OF NEURON APOPTOTIC PROCESS | 6 | 47 | 1.544e-08 | 3.926e-07 |
| 184 | SIGNAL TRANSDUCTION BY PROTEIN PHOSPHORYLATION | 12 | 404 | 1.715e-08 | 4.313e-07 |
| 185 | REGULATION OF ORGANELLE ORGANIZATION | 19 | 1178 | 1.715e-08 | 4.313e-07 |
| 186 | POSITIVE REGULATION OF TRANSPORT | 17 | 936 | 2.044e-08 | 5.114e-07 |
| 187 | RESPONSE TO GAMMA RADIATION | 6 | 50 | 2.266e-08 | 5.639e-07 |
| 188 | POSITIVE REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 10 | 258 | 2.481e-08 | 6.14e-07 |
| 189 | REGULATION OF PROTEIN LOCALIZATION | 17 | 950 | 2.541e-08 | 6.255e-07 |
| 190 | CELLULAR RESPONSE TO INTERLEUKIN 1 | 7 | 88 | 2.651e-08 | 6.493e-07 |
| 191 | PHOSPHATIDYLINOSITOL METABOLIC PROCESS | 9 | 193 | 2.677e-08 | 6.521e-07 |
| 192 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 18 | 1087 | 2.981e-08 | 7.224e-07 |
| 193 | REGULATION OF INTRACELLULAR TRANSPORT | 14 | 621 | 3.15e-08 | 7.593e-07 |
| 194 | LYMPHOCYTE ACTIVATION | 11 | 342 | 3.193e-08 | 7.657e-07 |
| 195 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 6 | 53 | 3.247e-08 | 7.748e-07 |
| 196 | MITOCHONDRIAL MEMBRANE ORGANIZATION | 7 | 92 | 3.618e-08 | 8.589e-07 |
| 197 | REGULATION OF CELL ADHESION | 14 | 629 | 3.694e-08 | 8.724e-07 |
| 198 | MULTICELLULAR ORGANISMAL HOMEOSTASIS | 10 | 272 | 4.084e-08 | 9.597e-07 |
| 199 | NEGATIVE REGULATION OF CATABOLIC PROCESS | 9 | 203 | 4.14e-08 | 9.68e-07 |
| 200 | REGULATION OF TRANSCRIPTION FACTOR IMPORT INTO NUCLEUS | 7 | 95 | 4.525e-08 | 1.053e-06 |
| 201 | REGULATION OF NECROPTOTIC PROCESS | 4 | 11 | 4.722e-08 | 1.088e-06 |
| 202 | POSITIVE REGULATION OF NFAT PROTEIN IMPORT INTO NUCLEUS | 4 | 11 | 4.722e-08 | 1.088e-06 |
| 203 | PEPTIDYL SERINE MODIFICATION | 8 | 148 | 5.335e-08 | 1.223e-06 |
| 204 | REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 5 | 29 | 5.43e-08 | 1.227e-06 |
| 205 | POSITIVE REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 5 | 29 | 5.43e-08 | 1.227e-06 |
| 206 | POSITIVE REGULATION OF NIK NF KAPPAB SIGNALING | 5 | 29 | 5.43e-08 | 1.227e-06 |
| 207 | POSITIVE REGULATION OF CYTOPLASMIC TRANSPORT | 10 | 282 | 5.733e-08 | 1.289e-06 |
| 208 | GLYCEROLIPID BIOSYNTHETIC PROCESS | 9 | 211 | 5.772e-08 | 1.291e-06 |
| 209 | LIPID PHOSPHORYLATION | 7 | 99 | 6.029e-08 | 1.342e-06 |
| 210 | REGULATION OF GLUCOSE TRANSPORT | 7 | 100 | 6.464e-08 | 1.432e-06 |
| 211 | REGULATION OF GLUCOSE IMPORT | 6 | 60 | 6.944e-08 | 1.526e-06 |
| 212 | SINGLE ORGANISM CELL ADHESION | 12 | 459 | 6.951e-08 | 1.526e-06 |
| 213 | REGULATION OF ENDOTHELIAL CELL DEVELOPMENT | 4 | 12 | 7.063e-08 | 1.529e-06 |
| 214 | REGULATION OF ESTABLISHMENT OF ENDOTHELIAL BARRIER | 4 | 12 | 7.063e-08 | 1.529e-06 |
| 215 | POSITIVE REGULATION OF GLUCOSE IMPORT IN RESPONSE TO INSULIN STIMULUS | 4 | 12 | 7.063e-08 | 1.529e-06 |
| 216 | RENAL SYSTEM PROCESS | 7 | 102 | 7.416e-08 | 1.598e-06 |
| 217 | REGULATION OF MITOCHONDRION ORGANIZATION | 9 | 218 | 7.635e-08 | 1.63e-06 |
| 218 | REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 9 | 218 | 7.635e-08 | 1.63e-06 |
| 219 | LIPOPOLYSACCHARIDE MEDIATED SIGNALING PATHWAY | 5 | 31 | 7.727e-08 | 1.642e-06 |
| 220 | REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 9 | 220 | 8.256e-08 | 1.746e-06 |
| 221 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 16 | 917 | 9.999e-08 | 2.105e-06 |
| 222 | CELLULAR RESPONSE TO BIOTIC STIMULUS | 8 | 163 | 1.127e-07 | 2.363e-06 |
| 223 | IMMUNE SYSTEM DEVELOPMENT | 13 | 582 | 1.162e-07 | 2.424e-06 |
| 224 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 22 | 1784 | 1.21e-07 | 2.514e-06 |
| 225 | REGULATION OF CELL ACTIVATION | 12 | 484 | 1.235e-07 | 2.554e-06 |
| 226 | PROTEIN KINASE B SIGNALING | 5 | 34 | 1.255e-07 | 2.572e-06 |
| 227 | RENAL WATER HOMEOSTASIS | 5 | 34 | 1.255e-07 | 2.572e-06 |
| 228 | RESPONSE TO ENDOPLASMIC RETICULUM STRESS | 9 | 233 | 1.347e-07 | 2.749e-06 |
| 229 | RESPONSE TO AMINO ACID | 7 | 112 | 1.415e-07 | 2.874e-06 |
| 230 | REGULATION OF CAMP DEPENDENT PROTEIN KINASE ACTIVITY | 4 | 14 | 1.421e-07 | 2.874e-06 |
| 231 | PHOSPHOLIPID BIOSYNTHETIC PROCESS | 9 | 235 | 1.449e-07 | 2.918e-06 |
| 232 | REGULATION OF TRANSPORT | 22 | 1804 | 1.472e-07 | 2.952e-06 |
| 233 | GLUCOSE HOMEOSTASIS | 8 | 170 | 1.559e-07 | 3.099e-06 |
| 234 | CARBOHYDRATE HOMEOSTASIS | 8 | 170 | 1.559e-07 | 3.099e-06 |
| 235 | NEGATIVE REGULATION OF NEURON DEATH | 8 | 171 | 1.631e-07 | 3.229e-06 |
| 236 | RESPONSE TO INTERLEUKIN 1 | 7 | 115 | 1.696e-07 | 3.345e-06 |
| 237 | RESPONSE TO ACID CHEMICAL | 10 | 319 | 1.807e-07 | 3.549e-06 |
| 238 | LEUKOCYTE ACTIVATION | 11 | 414 | 2.199e-07 | 4.3e-06 |
| 239 | CELLULAR RESPONSE TO GLUCAGON STIMULUS | 5 | 38 | 2.239e-07 | 4.359e-06 |
| 240 | CELLULAR RESPONSE TO CARBOHYDRATE STIMULUS | 6 | 74 | 2.471e-07 | 4.791e-06 |
| 241 | REGULATION OF PROTEIN IMPORT | 8 | 183 | 2.743e-07 | 5.295e-06 |
| 242 | NEGATIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 7 | 126 | 3.17e-07 | 6.095e-06 |
| 243 | RESPONSE TO DRUG | 11 | 431 | 3.283e-07 | 6.287e-06 |
| 244 | LEUKOCYTE MIGRATION | 9 | 259 | 3.297e-07 | 6.287e-06 |
| 245 | REGULATION OF NFAT PROTEIN IMPORT INTO NUCLEUS | 4 | 17 | 3.351e-07 | 6.29e-06 |
| 246 | ACTIVATION OF NF KAPPAB INDUCING KINASE ACTIVITY | 4 | 17 | 3.351e-07 | 6.29e-06 |
| 247 | REGULATION OF CELLULAR LOCALIZATION | 18 | 1277 | 3.352e-07 | 6.29e-06 |
| 248 | REGULATION OF GLUCOSE IMPORT IN RESPONSE TO INSULIN STIMULUS | 4 | 17 | 3.351e-07 | 6.29e-06 |
| 249 | PROTEIN OLIGOMERIZATION | 11 | 434 | 3.517e-07 | 6.572e-06 |
| 250 | REGULATION OF NIK NF KAPPAB SIGNALING | 5 | 42 | 3.753e-07 | 6.957e-06 |
| 251 | POSITIVE REGULATION OF GLUCOSE TRANSPORT | 5 | 42 | 3.753e-07 | 6.957e-06 |
| 252 | RESPONSE TO REACTIVE OXYGEN SPECIES | 8 | 191 | 3.801e-07 | 7.019e-06 |
| 253 | AGING | 9 | 264 | 3.872e-07 | 7.121e-06 |
| 254 | INSULIN RECEPTOR SIGNALING PATHWAY | 6 | 80 | 3.942e-07 | 7.221e-06 |
| 255 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 17 | 1152 | 4.037e-07 | 7.366e-06 |
| 256 | JNK CASCADE | 6 | 82 | 4.568e-07 | 8.302e-06 |
| 257 | REGULATION OF RELEASE OF CYTOCHROME C FROM MITOCHONDRIA | 5 | 44 | 4.765e-07 | 8.627e-06 |
| 258 | RESPONSE TO ALKALOID | 7 | 137 | 5.599e-07 | 1.006e-05 |
| 259 | ACTIVATION OF MAPK ACTIVITY | 7 | 137 | 5.599e-07 | 1.006e-05 |
| 260 | POSITIVE REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 6 | 86 | 6.065e-07 | 1.085e-05 |
| 261 | CELLULAR HOMEOSTASIS | 13 | 676 | 6.401e-07 | 1.141e-05 |
| 262 | NEGATIVE REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 6 | 88 | 6.952e-07 | 1.235e-05 |
| 263 | CELLULAR CHEMICAL HOMEOSTASIS | 12 | 570 | 7.074e-07 | 1.252e-05 |
| 264 | RESPONSE TO GLUCAGON | 5 | 48 | 7.431e-07 | 1.31e-05 |
| 265 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 12 | 573 | 7.477e-07 | 1.313e-05 |
| 266 | LYMPHOCYTE DIFFERENTIATION | 8 | 209 | 7.525e-07 | 1.316e-05 |
| 267 | RESPONSE TO IONIZING RADIATION | 7 | 145 | 8.217e-07 | 1.432e-05 |
| 268 | NEGATIVE REGULATION OF LIPID CATABOLIC PROCESS | 4 | 21 | 8.336e-07 | 1.447e-05 |
| 269 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 20 | 1672 | 8.773e-07 | 1.517e-05 |
| 270 | REGULATION OF CELL CELL ADHESION | 10 | 380 | 8.972e-07 | 1.546e-05 |
| 271 | REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 10 | 381 | 9.188e-07 | 1.578e-05 |
| 272 | REGULATION OF CYTOPLASMIC TRANSPORT | 11 | 481 | 9.682e-07 | 1.656e-05 |
| 273 | GLYCEROPHOSPHOLIPID METABOLIC PROCESS | 9 | 297 | 1.035e-06 | 1.763e-05 |
| 274 | MITOCHONDRION ORGANIZATION | 12 | 594 | 1.091e-06 | 1.853e-05 |
| 275 | REGULATION OF LIPID CATABOLIC PROCESS | 5 | 52 | 1.116e-06 | 1.887e-05 |
| 276 | REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 7 | 152 | 1.128e-06 | 1.902e-05 |
| 277 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 18 | 1395 | 1.216e-06 | 2.042e-05 |
| 278 | POSITIVE REGULATION OF CELLULAR RESPONSE TO INSULIN STIMULUS | 4 | 23 | 1.227e-06 | 2.053e-05 |
| 279 | REGULATION OF HOMOTYPIC CELL CELL ADHESION | 9 | 307 | 1.361e-06 | 2.261e-05 |
| 280 | REGULATION OF PROTEIN TARGETING | 9 | 307 | 1.361e-06 | 2.261e-05 |
| 281 | CELL CELL ADHESION | 12 | 608 | 1.391e-06 | 2.304e-05 |
| 282 | REGULATION OF BODY FLUID LEVELS | 11 | 506 | 1.586e-06 | 2.617e-05 |
| 283 | PROTEIN COMPLEX BIOGENESIS | 16 | 1132 | 1.672e-06 | 2.739e-05 |
| 284 | PROTEIN COMPLEX ASSEMBLY | 16 | 1132 | 1.672e-06 | 2.739e-05 |
| 285 | RESPONSE TO RADIATION | 10 | 413 | 1.899e-06 | 3.101e-05 |
| 286 | MULTICELLULAR ORGANISMAL WATER HOMEOSTASIS | 5 | 58 | 1.935e-06 | 3.147e-05 |
| 287 | DEVELOPMENTAL PROGRAMMED CELL DEATH | 4 | 26 | 2.055e-06 | 3.331e-05 |
| 288 | REGULATION OF CELLULAR RESPONSE TO INSULIN STIMULUS | 5 | 59 | 2.108e-06 | 3.406e-05 |
| 289 | POSITIVE REGULATION OF MITOCHONDRION ORGANIZATION | 7 | 167 | 2.118e-06 | 3.41e-05 |
| 290 | RESPONSE TO CARBOHYDRATE | 7 | 168 | 2.204e-06 | 3.536e-05 |
| 291 | POSITIVE REGULATION OF CELL CELL ADHESION | 8 | 243 | 2.328e-06 | 3.723e-05 |
| 292 | REGULATION OF ENDOTHELIAL CELL DIFFERENTIATION | 4 | 27 | 2.406e-06 | 3.82e-05 |
| 293 | DNA CATABOLIC PROCESS | 4 | 27 | 2.406e-06 | 3.82e-05 |
| 294 | RESPONSE TO VIRUS | 8 | 247 | 2.628e-06 | 4.16e-05 |
| 295 | RESPONSE TO METAL ION | 9 | 333 | 2.654e-06 | 4.186e-05 |
| 296 | NEGATIVE REGULATION OF KINASE ACTIVITY | 8 | 250 | 2.874e-06 | 4.519e-05 |
| 297 | HOMEOSTASIS OF NUMBER OF CELLS | 7 | 175 | 2.89e-06 | 4.528e-05 |
| 298 | RESPONSE TO CORTICOSTEROID | 7 | 176 | 3.002e-06 | 4.673e-05 |
| 299 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 11 | 541 | 3.023e-06 | 4.673e-05 |
| 300 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 11 | 541 | 3.023e-06 | 4.673e-05 |
| 301 | PROTEIN HETEROOLIGOMERIZATION | 6 | 113 | 3.023e-06 | 4.673e-05 |
| 302 | MITOCHONDRIAL TRANSPORT | 7 | 177 | 3.116e-06 | 4.801e-05 |
| 303 | REGULATION OF PEPTIDYL SERINE PHOSPHORYLATION | 6 | 118 | 3.888e-06 | 5.971e-05 |
| 304 | CELL PROLIFERATION | 12 | 672 | 3.912e-06 | 5.988e-05 |
| 305 | NEGATIVE REGULATION OF TRANSFERASE ACTIVITY | 9 | 351 | 4.075e-06 | 6.217e-05 |
| 306 | REGULATION OF CYTOKINE PRODUCTION | 11 | 563 | 4.425e-06 | 6.729e-05 |
| 307 | GLYCEROLIPID METABOLIC PROCESS | 9 | 356 | 4.57e-06 | 6.927e-05 |
| 308 | NEGATIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 12 | 684 | 4.687e-06 | 7.081e-05 |
| 309 | REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 6 | 122 | 4.717e-06 | 7.103e-05 |
| 310 | MYD88 DEPENDENT TOLL LIKE RECEPTOR SIGNALING PATHWAY | 4 | 32 | 4.863e-06 | 7.3e-05 |
| 311 | WATER HOMEOSTASIS | 5 | 70 | 4.944e-06 | 7.397e-05 |
| 312 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 10 | 465 | 5.442e-06 | 8.116e-05 |
| 313 | PHOSPHOLIPID METABOLIC PROCESS | 9 | 364 | 5.47e-06 | 8.132e-05 |
| 314 | RESPONSE TO UV | 6 | 126 | 5.683e-06 | 8.422e-05 |
| 315 | POSITIVE REGULATION OF CYTOKINE PRODUCTION | 9 | 370 | 6.242e-06 | 9.22e-05 |
| 316 | NEGATIVE REGULATION OF IMMUNE SYSTEM PROCESS | 9 | 372 | 6.519e-06 | 9.599e-05 |
| 317 | B CELL ACTIVATION | 6 | 132 | 7.43e-06 | 0.000109 |
| 318 | CELL DEVELOPMENT | 17 | 1426 | 7.448e-06 | 0.000109 |
| 319 | CELLULAR RESPONSE TO DNA DAMAGE STIMULUS | 12 | 720 | 7.887e-06 | 0.000115 |
| 320 | IMMUNE EFFECTOR PROCESS | 10 | 486 | 8.014e-06 | 0.0001165 |
| 321 | REGULATION OF VITAMIN METABOLIC PROCESS | 3 | 12 | 9.218e-06 | 0.0001336 |
| 322 | REGULATION OF INFLAMMATORY RESPONSE | 8 | 294 | 9.452e-06 | 0.0001366 |
| 323 | LIPID MODIFICATION | 7 | 210 | 9.583e-06 | 0.0001373 |
| 324 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 6 | 138 | 9.589e-06 | 0.0001373 |
| 325 | NEGATIVE REGULATION OF LIPID METABOLIC PROCESS | 5 | 80 | 9.554e-06 | 0.0001373 |
| 326 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 11 | 616 | 1.035e-05 | 0.0001478 |
| 327 | NEGATIVE REGULATION OF HYDROLASE ACTIVITY | 9 | 397 | 1.098e-05 | 0.0001562 |
| 328 | PLATELET ACTIVATION | 6 | 142 | 1.129e-05 | 0.0001602 |
| 329 | CELLULAR RESPONSE TO OXYGEN LEVELS | 6 | 143 | 1.175e-05 | 0.0001662 |
| 330 | HEPATOCYTE APOPTOTIC PROCESS | 3 | 13 | 1.195e-05 | 0.0001675 |
| 331 | RESPONSE TO COBALT ION | 3 | 13 | 1.195e-05 | 0.0001675 |
| 332 | POSITIVE REGULATION OF MACROPHAGE DIFFERENTIATION | 3 | 13 | 1.195e-05 | 0.0001675 |
| 333 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 4 | 40 | 1.21e-05 | 0.000169 |
| 334 | CELLULAR RESPONSE TO INSULIN STIMULUS | 6 | 146 | 1.323e-05 | 0.0001843 |
| 335 | REGULATION OF CELL DIFFERENTIATION | 17 | 1492 | 1.351e-05 | 0.0001876 |
| 336 | POSITIVE REGULATION OF CELL ACTIVATION | 8 | 311 | 1.42e-05 | 0.0001967 |
| 337 | RESPONSE TO TEMPERATURE STIMULUS | 6 | 148 | 1.43e-05 | 0.0001974 |
| 338 | TOLL LIKE RECEPTOR 9 SIGNALING PATHWAY | 3 | 14 | 1.517e-05 | 0.0002089 |
| 339 | POSITIVE REGULATION OF PEPTIDYL SERINE PHOSPHORYLATION | 5 | 88 | 1.523e-05 | 0.0002091 |
| 340 | RESPONSE TO HEAT | 5 | 89 | 1.609e-05 | 0.0002202 |
| 341 | CELLULAR LIPID METABOLIC PROCESS | 13 | 913 | 1.705e-05 | 0.0002327 |
| 342 | NEGATIVE REGULATION OF PHOSPHORYLATION | 9 | 422 | 1.783e-05 | 0.0002426 |
| 343 | REGULATION OF LEUKOCYTE DIFFERENTIATION | 7 | 232 | 1.827e-05 | 0.0002471 |
| 344 | PROTEIN COMPLEX SUBUNIT ORGANIZATION | 17 | 1527 | 1.826e-05 | 0.0002471 |
| 345 | APOPTOTIC DNA FRAGMENTATION | 3 | 15 | 1.892e-05 | 0.0002552 |
| 346 | NEGATIVE REGULATION OF CELLULAR CATABOLIC PROCESS | 6 | 156 | 1.928e-05 | 0.0002593 |
| 347 | POSITIVE REGULATION OF INTERLEUKIN 8 PRODUCTION | 4 | 45 | 1.946e-05 | 0.000261 |
| 348 | REGULATION OF JNK CASCADE | 6 | 159 | 2.147e-05 | 0.0002871 |
| 349 | RESPONSE TO ANTIBIOTIC | 4 | 47 | 2.317e-05 | 0.000309 |
| 350 | MACROMOLECULAR COMPLEX ASSEMBLY | 16 | 1398 | 2.426e-05 | 0.0003225 |
| 351 | POSITIVE REGULATION OF REACTIVE OXYGEN SPECIES BIOSYNTHETIC PROCESS | 4 | 48 | 2.521e-05 | 0.0003333 |
| 352 | REGULATION OF LIPID KINASE ACTIVITY | 4 | 48 | 2.521e-05 | 0.0003333 |
| 353 | REGULATION OF DNA METABOLIC PROCESS | 8 | 340 | 2.692e-05 | 0.0003549 |
| 354 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 13 | 957 | 2.794e-05 | 0.0003673 |
| 355 | NEGATIVE REGULATION OF ANOIKIS | 3 | 17 | 2.813e-05 | 0.0003687 |
| 356 | REGULATION OF AUTOPHAGY | 7 | 249 | 2.876e-05 | 0.0003759 |
| 357 | INTRACELLULAR RECEPTOR SIGNALING PATHWAY | 6 | 168 | 2.929e-05 | 0.0003818 |
| 358 | ORGANOPHOSPHATE BIOSYNTHETIC PROCESS | 9 | 450 | 2.954e-05 | 0.0003839 |
| 359 | POSITIVE REGULATION OF MYELOID LEUKOCYTE DIFFERENTIATION | 4 | 50 | 2.968e-05 | 0.0003847 |
| 360 | REGULATION OF CELLULAR RESPONSE TO STRESS | 11 | 691 | 3e-05 | 0.0003878 |
| 361 | CELLULAR PROCESS INVOLVED IN REPRODUCTION IN MULTICELLULAR ORGANISM | 7 | 252 | 3.105e-05 | 0.0004002 |
| 362 | RESPONSE TO NICOTINE | 4 | 51 | 3.212e-05 | 0.0004129 |
| 363 | NEGATIVE REGULATION OF ORGANIC ACID TRANSPORT | 3 | 18 | 3.367e-05 | 0.0004304 |
| 364 | INOSITOL PHOSPHATE MEDIATED SIGNALING | 3 | 18 | 3.367e-05 | 0.0004304 |
| 365 | STRIATED MUSCLE CELL DIFFERENTIATION | 6 | 173 | 3.454e-05 | 0.00044 |
| 366 | REGULATION OF T CELL MEDIATED IMMUNITY | 4 | 52 | 3.471e-05 | 0.00044 |
| 367 | POSITIVE REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY | 4 | 52 | 3.471e-05 | 0.00044 |
| 368 | CELLULAR RESPONSE TO ACID CHEMICAL | 6 | 175 | 3.683e-05 | 0.0004657 |
| 369 | REGULATION OF NITRIC OXIDE BIOSYNTHETIC PROCESS | 4 | 53 | 3.744e-05 | 0.0004703 |
| 370 | INNATE IMMUNE RESPONSE ACTIVATING CELL SURFACE RECEPTOR SIGNALING PATHWAY | 5 | 106 | 3.75e-05 | 0.0004703 |
| 371 | CELLULAR RESPONSE TO AMINO ACID STIMULUS | 4 | 53 | 3.744e-05 | 0.0004703 |
| 372 | DNA CATABOLIC PROCESS ENDONUCLEOLYTIC | 3 | 19 | 3.988e-05 | 0.0004988 |
| 373 | REGULATION OF LYMPHOCYTE APOPTOTIC PROCESS | 4 | 54 | 4.032e-05 | 0.000503 |
| 374 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 14 | 1142 | 4.084e-05 | 0.000508 |
| 375 | REGULATION OF MYELOID LEUKOCYTE DIFFERENTIATION | 5 | 108 | 4.102e-05 | 0.000509 |
| 376 | RESPONSE TO GROWTH FACTOR | 9 | 475 | 4.498e-05 | 0.0005566 |
| 377 | RESPONSE TO KETONE | 6 | 182 | 4.586e-05 | 0.000566 |
| 378 | POSITIVE REGULATION OF LYMPHOCYTE APOPTOTIC PROCESS | 3 | 20 | 4.68e-05 | 0.0005746 |
| 379 | REGULATION OF MACROPHAGE DIFFERENTIATION | 3 | 20 | 4.68e-05 | 0.0005746 |
| 380 | REGULATION OF MYELOID CELL DIFFERENTIATION | 6 | 183 | 4.728e-05 | 0.000579 |
| 381 | APOPTOTIC MITOCHONDRIAL CHANGES | 4 | 57 | 4.997e-05 | 0.0006102 |
| 382 | REGULATION OF LYMPHOCYTE MEDIATED IMMUNITY | 5 | 114 | 5.315e-05 | 0.0006473 |
| 383 | B CELL HOMEOSTASIS | 3 | 21 | 5.446e-05 | 0.0006599 |
| 384 | RESPONSE TO NITRIC OXIDE | 3 | 21 | 5.446e-05 | 0.0006599 |
| 385 | POSITIVE REGULATION OF CELL ADHESION | 8 | 376 | 5.487e-05 | 0.0006631 |
| 386 | MYELOID CELL DIFFERENTIATION | 6 | 189 | 5.658e-05 | 0.0006821 |
| 387 | PROTEIN DEPHOSPHORYLATION | 6 | 190 | 5.827e-05 | 0.0007005 |
| 388 | BIOLOGICAL ADHESION | 13 | 1032 | 6.086e-05 | 0.0007298 |
| 389 | REGULATION OF MONOOXYGENASE ACTIVITY | 4 | 60 | 6.119e-05 | 0.0007319 |
| 390 | PROTEIN AUTOPHOSPHORYLATION | 6 | 192 | 6.175e-05 | 0.0007367 |
| 391 | REGULATION OF PROTEIN HOMODIMERIZATION ACTIVITY | 3 | 22 | 6.29e-05 | 0.0007447 |
| 392 | POSITIVE REGULATION OF PROTEIN OLIGOMERIZATION | 3 | 22 | 6.29e-05 | 0.0007447 |
| 393 | RELEASE OF CYTOCHROME C FROM MITOCHONDRIA | 3 | 22 | 6.29e-05 | 0.0007447 |
| 394 | INNATE IMMUNE RESPONSE | 10 | 619 | 6.324e-05 | 0.0007468 |
| 395 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 9 | 498 | 6.476e-05 | 0.0007628 |
| 396 | REGULATION OF PHOSPHOLIPID METABOLIC PROCESS | 4 | 61 | 6.531e-05 | 0.0007654 |
| 397 | REGULATION OF INTERLEUKIN 8 PRODUCTION | 4 | 61 | 6.531e-05 | 0.0007654 |
| 398 | EMBRYO DEVELOPMENT | 12 | 894 | 6.639e-05 | 0.0007762 |
| 399 | MEMBRANE ORGANIZATION | 12 | 899 | 7.003e-05 | 0.0008166 |
| 400 | REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 6 | 197 | 7.12e-05 | 0.0008282 |
| 401 | POSITIVE REGULATION OF JUN KINASE ACTIVITY | 4 | 63 | 7.415e-05 | 0.0008604 |
| 402 | REGULATION OF ADAPTIVE IMMUNE RESPONSE | 5 | 123 | 7.632e-05 | 0.0008811 |
| 403 | T CELL DIFFERENTIATION | 5 | 123 | 7.632e-05 | 0.0008811 |
| 404 | REGULATION OF ANOIKIS | 3 | 24 | 8.225e-05 | 0.0009473 |
| 405 | REGULATION OF REACTIVE OXYGEN SPECIES BIOSYNTHETIC PROCESS | 4 | 65 | 8.383e-05 | 0.0009631 |
| 406 | RESPONSE TO INSULIN | 6 | 205 | 8.869e-05 | 0.001016 |
| 407 | POSITIVE REGULATION OF LIPID BIOSYNTHETIC PROCESS | 4 | 66 | 8.9e-05 | 0.001017 |
| 408 | REGULATION OF LEUKOCYTE PROLIFERATION | 6 | 206 | 9.109e-05 | 0.001039 |
| 409 | REGULATION OF LIPID BIOSYNTHETIC PROCESS | 5 | 128 | 9.217e-05 | 0.001043 |
| 410 | REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION TO MITOCHONDRION | 5 | 128 | 9.217e-05 | 0.001043 |
| 411 | POSITIVE REGULATION OF LIPID METABOLIC PROCESS | 5 | 128 | 9.217e-05 | 0.001043 |
| 412 | CELLULAR EXTRAVASATION | 3 | 25 | 9.323e-05 | 0.001048 |
| 413 | APOPTOTIC NUCLEAR CHANGES | 3 | 25 | 9.323e-05 | 0.001048 |
| 414 | EPITHELIAL CELL APOPTOTIC PROCESS | 3 | 25 | 9.323e-05 | 0.001048 |
| 415 | CELLULAR RESPONSE TO DRUG | 4 | 67 | 9.44e-05 | 0.001058 |
| 416 | POSITIVE REGULATION OF LEUKOCYTE DIFFERENTIATION | 5 | 131 | 0.0001028 | 0.00115 |
| 417 | NEGATIVE REGULATION OF LIPID TRANSPORT | 3 | 26 | 0.0001051 | 0.001173 |
| 418 | REGULATION OF RESPONSE TO WOUNDING | 8 | 413 | 0.0001055 | 0.001174 |
| 419 | POSITIVE REGULATION OF LYMPHOCYTE MEDIATED IMMUNITY | 4 | 69 | 0.0001059 | 0.001176 |
| 420 | REGULATION OF CELL CYCLE | 12 | 949 | 0.000117 | 0.001297 |
| 421 | NEGATIVE REGULATION OF NEURON APOPTOTIC PROCESS | 5 | 135 | 0.0001185 | 0.001297 |
| 422 | POSITIVE REGULATION OF LOCOMOTION | 8 | 420 | 0.0001184 | 0.001297 |
| 423 | POSITIVE REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 5 | 135 | 0.0001185 | 0.001297 |
| 424 | REGULATION OF FATTY ACID TRANSPORT | 3 | 27 | 0.000118 | 0.001297 |
| 425 | LIPID BIOSYNTHETIC PROCESS | 9 | 539 | 0.0001183 | 0.001297 |
| 426 | REGULATION OF HEMOPOIESIS | 7 | 314 | 0.0001237 | 0.001352 |
| 427 | CELLULAR RESPONSE TO RADIATION | 5 | 137 | 0.000127 | 0.001384 |
| 428 | POSITIVE REGULATION OF ADAPTIVE IMMUNE RESPONSE | 4 | 73 | 0.0001319 | 0.001418 |
| 429 | POSITIVE REGULATION OF LEUKOCYTE APOPTOTIC PROCESS | 3 | 28 | 0.0001318 | 0.001418 |
| 430 | RESPONSE TO COPPER ION | 3 | 28 | 0.0001318 | 0.001418 |
| 431 | POSITIVE REGULATION OF ACUTE INFLAMMATORY RESPONSE | 3 | 28 | 0.0001318 | 0.001418 |
| 432 | POSITIVE REGULATION OF RELEASE OF CYTOCHROME C FROM MITOCHONDRIA | 3 | 28 | 0.0001318 | 0.001418 |
| 433 | POSITIVE REGULATION OF CELL PROLIFERATION | 11 | 814 | 0.0001304 | 0.001418 |
| 434 | VASCULAR ENDOTHELIAL GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 4 | 74 | 0.0001391 | 0.001491 |
| 435 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 11 | 823 | 0.0001436 | 0.001536 |
| 436 | MUSCLE ADAPTATION | 3 | 29 | 0.0001466 | 0.001554 |
| 437 | GRANULOCYTE MIGRATION | 4 | 75 | 0.0001465 | 0.001554 |
| 438 | POSITIVE REGULATION OF MONOOXYGENASE ACTIVITY | 3 | 29 | 0.0001466 | 0.001554 |
| 439 | NEGATIVE REGULATION OF CELL CYCLE | 8 | 433 | 0.000146 | 0.001554 |
| 440 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 10 | 689 | 0.0001528 | 0.001616 |
| 441 | REGULATION OF CELL CYCLE PROCESS | 9 | 558 | 0.0001535 | 0.001619 |
| 442 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 8 | 437 | 0.0001555 | 0.001637 |
| 443 | NEGATIVE REGULATION OF B CELL ACTIVATION | 3 | 30 | 0.0001625 | 0.0017 |
| 444 | CELL MOTILITY | 11 | 835 | 0.000163 | 0.0017 |
| 445 | LOCALIZATION OF CELL | 11 | 835 | 0.000163 | 0.0017 |
| 446 | REGULATION OF ANION TRANSMEMBRANE TRANSPORT | 3 | 30 | 0.0001625 | 0.0017 |
| 447 | NEGATIVE REGULATION OF PROTEOLYSIS | 7 | 329 | 0.000165 | 0.001718 |
| 448 | REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY | 5 | 145 | 0.0001657 | 0.001721 |
| 449 | LYMPHOCYTE COSTIMULATION | 4 | 78 | 0.0001706 | 0.001768 |
| 450 | REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 5 | 147 | 0.0001766 | 0.001826 |
| 451 | REGULATION OF LEUKOCYTE APOPTOTIC PROCESS | 4 | 79 | 0.0001792 | 0.001847 |
| 452 | POSITIVE REGULATION OF INTERLEUKIN 2 PRODUCTION | 3 | 31 | 0.0001794 | 0.001847 |
| 453 | LIPID METABOLIC PROCESS | 13 | 1158 | 0.0001931 | 0.001984 |
| 454 | MUSCLE CELL DIFFERENTIATION | 6 | 237 | 0.0001957 | 0.002006 |
| 455 | POSITIVE REGULATION OF MYELOID CELL DIFFERENTIATION | 4 | 81 | 0.0001974 | 0.002011 |
| 456 | POSITIVE REGULATION OF T CELL MEDIATED IMMUNITY | 3 | 32 | 0.0001975 | 0.002011 |
| 457 | REGULATION OF JUN KINASE ACTIVITY | 4 | 81 | 0.0001974 | 0.002011 |
| 458 | ESTABLISHMENT OF LOCALIZATION IN CELL | 16 | 1676 | 0.0002092 | 0.002125 |
| 459 | CELLULAR RESPONSE TO LIPID | 8 | 457 | 0.0002109 | 0.002138 |
| 460 | NEGATIVE REGULATION OF ANION TRANSPORT | 3 | 33 | 0.0002167 | 0.002192 |
| 461 | REGULATION OF LEUKOCYTE MEDIATED IMMUNITY | 5 | 156 | 0.0002329 | 0.00235 |
| 462 | NEGATIVE REGULATION OF PEPTIDASE ACTIVITY | 6 | 245 | 0.0002341 | 0.002357 |
| 463 | CELLULAR RESPONSE TO ALKALOID | 3 | 34 | 0.0002371 | 0.002382 |
| 464 | POSITIVE REGULATION OF LEUKOCYTE MEDIATED IMMUNITY | 4 | 85 | 0.0002377 | 0.002383 |
| 465 | GAMETE GENERATION | 9 | 595 | 0.0002475 | 0.002473 |
| 466 | REGULATION OF MITOTIC CELL CYCLE | 8 | 468 | 0.0002477 | 0.002473 |
| 467 | REGULATION OF PROTEIN OLIGOMERIZATION | 3 | 35 | 0.0002586 | 0.002577 |
| 468 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 9 | 602 | 0.0002698 | 0.002682 |
| 469 | LEUKOCYTE PROLIFERATION | 4 | 88 | 0.0002715 | 0.002694 |
| 470 | T CELL PROLIFERATION | 3 | 36 | 0.0002814 | 0.002786 |
| 471 | B CELL DIFFERENTIATION | 4 | 89 | 0.0002835 | 0.002795 |
| 472 | EPITHELIAL CELL PROLIFERATION | 4 | 89 | 0.0002835 | 0.002795 |
| 473 | POSITIVE REGULATION OF HEMOPOIESIS | 5 | 163 | 0.0002853 | 0.002806 |
| 474 | REGULATION OF OXIDOREDUCTASE ACTIVITY | 4 | 90 | 0.0002959 | 0.002899 |
| 475 | RESPONSE TO ALCOHOL | 7 | 362 | 0.0002955 | 0.002899 |
| 476 | SINGLE ORGANISM CELLULAR LOCALIZATION | 11 | 898 | 0.0003053 | 0.002985 |
| 477 | REGULATION OF CYTOKINE BIOSYNTHETIC PROCESS | 4 | 94 | 0.0003494 | 0.003409 |
| 478 | WNT SIGNALING PATHWAY CALCIUM MODULATING PATHWAY | 3 | 39 | 0.0003575 | 0.003465 |
| 479 | ERBB2 SIGNALING PATHWAY | 3 | 39 | 0.0003575 | 0.003465 |
| 480 | SPLEEN DEVELOPMENT | 3 | 39 | 0.0003575 | 0.003465 |
| 481 | FC GAMMA RECEPTOR SIGNALING PATHWAY | 4 | 95 | 0.0003638 | 0.003512 |
| 482 | REGULATION OF LIPID TRANSPORT | 4 | 95 | 0.0003638 | 0.003512 |
| 483 | MULTICELLULAR ORGANISM REPRODUCTION | 10 | 768 | 0.0003647 | 0.003513 |
| 484 | RESPONSE TO STEROID HORMONE | 8 | 497 | 0.0003706 | 0.003563 |
| 485 | MYELOID LEUKOCYTE DIFFERENTIATION | 4 | 96 | 0.0003786 | 0.003625 |
| 486 | FEMALE GAMETE GENERATION | 4 | 96 | 0.0003786 | 0.003625 |
| 487 | REGULATION OF ACTIVATED T CELL PROLIFERATION | 3 | 40 | 0.0003855 | 0.003683 |
| 488 | MACROMOLECULE CATABOLIC PROCESS | 11 | 926 | 0.0003964 | 0.003779 |
| 489 | NEGATIVE REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 4 | 98 | 0.0004095 | 0.003897 |
| 490 | MYELOID LEUKOCYTE MIGRATION | 4 | 99 | 0.0004257 | 0.004041 |
| 491 | SYSTEM PROCESS | 16 | 1785 | 0.0004264 | 0.004041 |
| 492 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 10 | 788 | 0.0004464 | 0.004213 |
| 493 | CIRCULATORY SYSTEM DEVELOPMENT | 10 | 788 | 0.0004464 | 0.004213 |
| 494 | REGULATION OF TUMOR NECROSIS FACTOR SUPERFAMILY CYTOKINE PRODUCTION | 4 | 101 | 0.0004592 | 0.004325 |
| 495 | CELLULAR COMPONENT DISASSEMBLY | 8 | 515 | 0.0004693 | 0.004411 |
| 496 | MYELOID LEUKOCYTE MEDIATED IMMUNITY | 3 | 43 | 0.0004778 | 0.004465 |
| 497 | REGULATION OF POLYSACCHARIDE METABOLIC PROCESS | 3 | 43 | 0.0004778 | 0.004465 |
| 498 | RESPONSE TO LIGHT STIMULUS | 6 | 280 | 0.0004769 | 0.004465 |
| 499 | MUSCLE SYSTEM PROCESS | 6 | 282 | 0.0004952 | 0.004617 |
| 500 | CELLULAR RESPONSE TO OXIDATIVE STRESS | 5 | 184 | 0.0004973 | 0.004628 |
| 501 | GLAND DEVELOPMENT | 7 | 395 | 0.0004986 | 0.004631 |
| 502 | LOCOMOTION | 12 | 1114 | 0.0005099 | 0.004727 |
| 503 | DEPHOSPHORYLATION | 6 | 286 | 0.0005333 | 0.004933 |
| 504 | INTERSPECIES INTERACTION BETWEEN ORGANISMS | 9 | 662 | 0.0005387 | 0.004963 |
| 505 | SYMBIOSIS ENCOMPASSING MUTUALISM THROUGH PARASITISM | 9 | 662 | 0.0005387 | 0.004963 |
| 506 | THYMOCYTE AGGREGATION | 3 | 45 | 0.0005467 | 0.005007 |
| 507 | T CELL DIFFERENTIATION IN THYMUS | 3 | 45 | 0.0005467 | 0.005007 |
| 508 | REGULATION OF ALCOHOL BIOSYNTHETIC PROCESS | 3 | 45 | 0.0005467 | 0.005007 |
| 509 | REGULATION OF CELL CYCLE ARREST | 4 | 108 | 0.0005916 | 0.005408 |
| 510 | RESPONSE TO HYDROGEN PEROXIDE | 4 | 109 | 0.0006125 | 0.005588 |
| 511 | POSITIVE REGULATION OF OXIDOREDUCTASE ACTIVITY | 3 | 47 | 0.0006215 | 0.005659 |
| 512 | REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION TO PLASMA MEMBRANE | 3 | 48 | 0.0006612 | 0.005998 |
| 513 | REGULATION OF INTERLEUKIN 2 PRODUCTION | 3 | 48 | 0.0006612 | 0.005998 |
| 514 | REGULATION OF CERAMIDE BIOSYNTHETIC PROCESS | 2 | 11 | 0.0006694 | 0.006025 |
| 515 | POSITIVE REGULATION OF HAIR CYCLE | 2 | 11 | 0.0006694 | 0.006025 |
| 516 | REGULATION OF FEVER GENERATION | 2 | 11 | 0.0006694 | 0.006025 |
| 517 | NEGATIVE REGULATION OF NECROTIC CELL DEATH | 2 | 11 | 0.0006694 | 0.006025 |
| 518 | OVULATION CYCLE | 4 | 113 | 0.0007014 | 0.006287 |
| 519 | REGULATION OF STEROID BIOSYNTHETIC PROCESS | 3 | 49 | 0.0007026 | 0.006287 |
| 520 | POSITIVE REGULATION OF INFLAMMATORY RESPONSE | 4 | 113 | 0.0007014 | 0.006287 |
| 521 | NEGATIVE REGULATION OF MITOTIC CELL CYCLE | 5 | 199 | 0.0007092 | 0.006334 |
| 522 | PEPTIDYL AMINO ACID MODIFICATION | 10 | 841 | 0.0007399 | 0.006595 |
| 523 | REGULATION OF ORGANIC ACID TRANSPORT | 3 | 50 | 0.0007455 | 0.006633 |
| 524 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 8 | 554 | 0.000757 | 0.006715 |
| 525 | REGULATION OF IMMUNE EFFECTOR PROCESS | 7 | 424 | 0.0007577 | 0.006715 |
| 526 | FEMALE SEX DIFFERENTIATION | 4 | 116 | 0.0007738 | 0.006845 |
| 527 | PLASMA MEMBRANE ORGANIZATION | 5 | 203 | 0.0007757 | 0.006849 |
| 528 | TISSUE DEVELOPMENT | 14 | 1518 | 0.0007878 | 0.006942 |
| 529 | REGULATION OF CHEMOKINE BIOSYNTHETIC PROCESS | 2 | 12 | 0.0008014 | 0.006983 |
| 530 | I KAPPAB PHOSPHORYLATION | 2 | 12 | 0.0008014 | 0.006983 |
| 531 | REPLICATIVE SENESCENCE | 2 | 12 | 0.0008014 | 0.006983 |
| 532 | MYELIN MAINTENANCE | 2 | 12 | 0.0008014 | 0.006983 |
| 533 | POSITIVE REGULATION OF INTERLEUKIN 2 BIOSYNTHETIC PROCESS | 2 | 12 | 0.0008014 | 0.006983 |
| 534 | POSITIVE REGULATION OF EXECUTION PHASE OF APOPTOSIS | 2 | 12 | 0.0008014 | 0.006983 |
| 535 | CELLULAR RESPONSE TO IONIZING RADIATION | 3 | 52 | 0.0008364 | 0.007274 |
| 536 | GLUCOSE METABOLIC PROCESS | 4 | 119 | 0.0008513 | 0.00739 |
| 537 | CYTOKINE PRODUCTION | 4 | 120 | 0.0008783 | 0.007611 |
| 538 | NEGATIVE REGULATION OF AUTOPHAGY | 3 | 53 | 0.0008843 | 0.007634 |
| 539 | GERM CELL DEVELOPMENT | 5 | 209 | 0.000884 | 0.007634 |
| 540 | REGULATION OF B CELL ACTIVATION | 4 | 121 | 0.000906 | 0.007806 |
| 541 | REGULATION OF MITOCHONDRIAL MEMBRANE POTENTIAL | 3 | 54 | 0.000934 | 0.008018 |
| 542 | B CELL RECEPTOR SIGNALING PATHWAY | 3 | 54 | 0.000934 | 0.008018 |
| 543 | REGULATION OF SPHINGOLIPID BIOSYNTHETIC PROCESS | 2 | 13 | 0.000945 | 0.008024 |
| 544 | REGULATION OF MEMBRANE LIPID METABOLIC PROCESS | 2 | 13 | 0.000945 | 0.008024 |
| 545 | POSITIVE REGULATION OF STEROID BIOSYNTHETIC PROCESS | 2 | 13 | 0.000945 | 0.008024 |
| 546 | POSITIVE REGULATION OF ENDOPLASMIC RETICULUM UNFOLDED PROTEIN RESPONSE | 2 | 13 | 0.000945 | 0.008024 |
| 547 | REGULATION OF HISTONE PHOSPHORYLATION | 2 | 13 | 0.000945 | 0.008024 |
| 548 | REGULATION OF BICELLULAR TIGHT JUNCTION ASSEMBLY | 2 | 13 | 0.000945 | 0.008024 |
| 549 | RESPONSE TO EXTRACELLULAR STIMULUS | 7 | 441 | 0.0009532 | 0.008079 |
| 550 | NEGATIVE REGULATION OF CELL CYCLE PROCESS | 5 | 214 | 0.0009825 | 0.008312 |
| 551 | REGULATION OF B CELL PROLIFERATION | 3 | 55 | 0.0009853 | 0.008321 |
| 552 | POSITIVE REGULATION OF HOMEOSTATIC PROCESS | 5 | 216 | 0.001024 | 0.008633 |
| 553 | REGULATION OF HOMEOSTATIC PROCESS | 7 | 447 | 0.001031 | 0.008675 |
| 554 | RESPONSE TO ESTROGEN | 5 | 218 | 0.001067 | 0.008962 |
| 555 | SEXUAL REPRODUCTION | 9 | 730 | 0.001078 | 0.009036 |
| 556 | ORGANOPHOSPHATE METABOLIC PROCESS | 10 | 885 | 0.001091 | 0.009075 |
| 557 | POSITIVE REGULATION OF TUMOR NECROSIS FACTOR SUPERFAMILY CYTOKINE PRODUCTION | 3 | 57 | 0.001093 | 0.009075 |
| 558 | REGULATION OF CYTOKINE PRODUCTION INVOLVED IN IMMUNE RESPONSE | 3 | 57 | 0.001093 | 0.009075 |
| 559 | REGULATION OF FIBROBLAST APOPTOTIC PROCESS | 2 | 14 | 0.0011 | 0.009075 |
| 560 | INSULIN LIKE GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 2 | 14 | 0.0011 | 0.009075 |
| 561 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 2 | 14 | 0.0011 | 0.009075 |
| 562 | T CELL MIGRATION | 2 | 14 | 0.0011 | 0.009075 |
| 563 | DETERMINATION OF ADULT LIFESPAN | 2 | 14 | 0.0011 | 0.009075 |
| 564 | NEGATIVE REGULATION OF ION TRANSPORT | 4 | 127 | 0.001085 | 0.009075 |
| 565 | NUCLEAR IMPORT | 4 | 129 | 0.001149 | 0.009455 |
| 566 | POSITIVE REGULATION OF CYTOKINE BIOSYNTHETIC PROCESS | 3 | 58 | 0.00115 | 0.009455 |
| 567 | POSITIVE REGULATION OF CELL CYCLE | 6 | 332 | 0.001157 | 0.009496 |
| 568 | NEGATIVE REGULATION OF TRANSPORT | 7 | 458 | 0.001187 | 0.009721 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | TUMOR NECROSIS FACTOR RECEPTOR SUPERFAMILY BINDING | 11 | 47 | 7.893e-18 | 7.332e-15 |
| 2 | CYTOKINE RECEPTOR BINDING | 16 | 271 | 1.362e-15 | 6.328e-13 |
| 3 | KINASE ACTIVITY | 23 | 842 | 6.616e-15 | 2.049e-12 |
| 4 | DEATH RECEPTOR BINDING | 7 | 18 | 1.618e-13 | 3.757e-11 |
| 5 | ENZYME BINDING | 29 | 1737 | 2.481e-13 | 3.841e-11 |
| 6 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 23 | 992 | 2.099e-13 | 3.841e-11 |
| 7 | X1 PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 8 | 43 | 2.206e-12 | 2.927e-10 |
| 8 | PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 9 | 70 | 2.9e-12 | 2.993e-10 |
| 9 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 16 | 445 | 2.881e-12 | 2.993e-10 |
| 10 | CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC PROCESS | 6 | 15 | 7.87e-12 | 7.311e-10 |
| 11 | PHOSPHATIDYLINOSITOL KINASE ACTIVITY | 8 | 51 | 9.471e-12 | 7.745e-10 |
| 12 | TUMOR NECROSIS FACTOR RECEPTOR BINDING | 7 | 30 | 1e-11 | 7.745e-10 |
| 13 | PROTEIN KINASE ACTIVITY | 17 | 640 | 6.498e-11 | 4.643e-09 |
| 14 | PROTEIN HETERODIMERIZATION ACTIVITY | 15 | 468 | 7.613e-11 | 5.052e-09 |
| 15 | PROTEASE BINDING | 9 | 104 | 1.114e-10 | 6.898e-09 |
| 16 | ADENYL NUCLEOTIDE BINDING | 24 | 1514 | 1.72e-10 | 9.988e-09 |
| 17 | DEATH RECEPTOR ACTIVITY | 6 | 24 | 2.064e-10 | 1.128e-08 |
| 18 | KINASE REGULATOR ACTIVITY | 10 | 186 | 1.073e-09 | 5.537e-08 |
| 19 | UBIQUITIN LIKE PROTEIN LIGASE BINDING | 11 | 264 | 2.19e-09 | 1.071e-07 |
| 20 | IDENTICAL PROTEIN BINDING | 20 | 1209 | 4.24e-09 | 1.97e-07 |
| 21 | RIBONUCLEOTIDE BINDING | 24 | 1860 | 1.07e-08 | 4.734e-07 |
| 22 | KINASE BINDING | 14 | 606 | 2.321e-08 | 8.985e-07 |
| 23 | CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 7 | 86 | 2.257e-08 | 8.985e-07 |
| 24 | RECEPTOR BINDING | 21 | 1476 | 2.24e-08 | 8.985e-07 |
| 25 | PROTEIN SERINE THREONINE KINASE INHIBITOR ACTIVITY | 5 | 30 | 6.498e-08 | 2.415e-06 |
| 26 | PROTEIN DIMERIZATION ACTIVITY | 18 | 1149 | 6.926e-08 | 2.475e-06 |
| 27 | ENZYME REGULATOR ACTIVITY | 16 | 959 | 1.84e-07 | 6.329e-06 |
| 28 | PROTEIN KINASE A CATALYTIC SUBUNIT BINDING | 4 | 15 | 1.932e-07 | 6.411e-06 |
| 29 | PROTEIN PHOSPHATASE BINDING | 7 | 120 | 2.271e-07 | 7.276e-06 |
| 30 | INTERLEUKIN 1 RECEPTOR BINDING | 4 | 16 | 2.569e-07 | 7.7e-06 |
| 31 | PHOSPHATIDYLINOSITOL PHOSPHATE KINASE ACTIVITY | 4 | 16 | 2.569e-07 | 7.7e-06 |
| 32 | MOLECULAR FUNCTION REGULATOR | 18 | 1353 | 7.812e-07 | 2.268e-05 |
| 33 | CAMP BINDING | 4 | 23 | 1.227e-06 | 3.454e-05 |
| 34 | PHOSPHATASE BINDING | 7 | 162 | 1.729e-06 | 4.724e-05 |
| 35 | PROTEIN DOMAIN SPECIFIC BINDING | 12 | 624 | 1.823e-06 | 4.839e-05 |
| 36 | CYSTEINE TYPE PEPTIDASE ACTIVITY | 7 | 184 | 4.028e-06 | 0.0001039 |
| 37 | SIGNAL TRANSDUCER ACTIVITY | 19 | 1731 | 6.308e-06 | 0.0001584 |
| 38 | INSULIN RECEPTOR SUBSTRATE BINDING | 3 | 11 | 6.931e-06 | 0.0001694 |
| 39 | CYCLIC NUCLEOTIDE BINDING | 4 | 36 | 7.882e-06 | 0.0001877 |
| 40 | PROTEIN KINASE A BINDING | 4 | 42 | 1.474e-05 | 0.0003339 |
| 41 | CYSTEINE TYPE ENDOPEPTIDASE REGULATOR ACTIVITY INVOLVED IN APOPTOTIC PROCESS | 4 | 42 | 1.474e-05 | 0.0003339 |
| 42 | KINASE INHIBITOR ACTIVITY | 5 | 89 | 1.609e-05 | 0.000356 |
| 43 | ENZYME INHIBITOR ACTIVITY | 8 | 378 | 5.695e-05 | 0.00123 |
| 44 | PROTEIN SERINE THREONINE PHOSPHATASE ACTIVITY | 4 | 64 | 7.888e-05 | 0.001665 |
| 45 | GROWTH FACTOR RECEPTOR BINDING | 5 | 129 | 9.563e-05 | 0.001974 |
| 46 | PROTEIN COMPLEX BINDING | 12 | 935 | 0.0001017 | 0.002054 |
| 47 | PROTEIN PHOSPHATASE 2A BINDING | 3 | 28 | 0.0001318 | 0.002605 |
| 48 | PROTEIN HOMODIMERIZATION ACTIVITY | 10 | 722 | 0.000223 | 0.004315 |
| 49 | HEAT SHOCK PROTEIN BINDING | 4 | 89 | 0.0002835 | 0.005376 |
| 50 | MACROMOLECULAR COMPLEX BINDING | 14 | 1399 | 0.0003483 | 0.006471 |
| 51 | PHOSPHOPROTEIN PHOSPHATASE ACTIVITY | 5 | 178 | 0.0004275 | 0.007787 |
| 52 | SCAFFOLD PROTEIN BINDING | 3 | 45 | 0.0005467 | 0.009766 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | TRANSFERASE COMPLEX TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 17 | 237 | 5.902e-18 | 3.447e-15 |
| 2 | CATALYTIC COMPLEX | 27 | 1038 | 4.801e-17 | 1.402e-14 |
| 3 | MEMBRANE MICRODOMAIN | 17 | 288 | 1.581e-16 | 3.078e-14 |
| 4 | PHOSPHATIDYLINOSITOL 3 KINASE COMPLEX | 8 | 20 | 2.044e-15 | 2.984e-13 |
| 5 | MEMBRANE PROTEIN COMPLEX | 23 | 1020 | 3.748e-13 | 4.378e-11 |
| 6 | TRANSFERASE COMPLEX | 18 | 703 | 2.939e-11 | 2.86e-09 |
| 7 | PROTEIN KINASE COMPLEX | 8 | 90 | 1.033e-09 | 8.621e-08 |
| 8 | CILIARY BASE | 5 | 23 | 1.564e-08 | 1.142e-06 |
| 9 | PLASMA MEMBRANE PROTEIN COMPLEX | 13 | 510 | 2.494e-08 | 1.618e-06 |
| 10 | MEMBRANE REGION | 18 | 1134 | 5.677e-08 | 3.316e-06 |
| 11 | CYTOSOLIC PART | 9 | 223 | 9.269e-08 | 4.921e-06 |
| 12 | EXTRINSIC COMPONENT OF MEMBRANE | 8 | 252 | 3.049e-06 | 0.0001484 |
| 13 | CD40 RECEPTOR COMPLEX | 3 | 11 | 6.931e-06 | 0.0002891 |
| 14 | IKAPPAB KINASE COMPLEX | 3 | 11 | 6.931e-06 | 0.0002891 |
| 15 | RECEPTOR COMPLEX | 8 | 327 | 2.038e-05 | 0.0007933 |
| 16 | PHOSPHATASE COMPLEX | 4 | 48 | 2.521e-05 | 0.0009202 |
| 17 | MITOCHONDRION | 16 | 1633 | 0.0001551 | 0.005327 |
| 18 | INTRINSIC COMPONENT OF PLASMA MEMBRANE | 16 | 1649 | 0.0001736 | 0.005631 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04210_Apoptosis | 71 | 89 | 1.237e-185 | 2.189e-183 | |
| 2 | hsa04380_Osteoclast_differentiation | 25 | 128 | 9.337e-38 | 8.264e-36 | |
| 3 | hsa04620_Toll.like_receptor_signaling_pathway | 22 | 102 | 3.339e-34 | 1.97e-32 | |
| 4 | hsa04660_T_cell_receptor_signaling_pathway | 22 | 108 | 1.34e-33 | 5.929e-32 | |
| 5 | hsa04662_B_cell_receptor_signaling_pathway | 20 | 75 | 3.708e-33 | 1.313e-31 | |
| 6 | hsa04722_Neurotrophin_signaling_pathway | 22 | 127 | 6.528e-32 | 1.926e-30 | |
| 7 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 22 | 136 | 3.3e-31 | 8.344e-30 | |
| 8 | hsa04010_MAPK_signaling_pathway | 26 | 268 | 6.085e-31 | 1.346e-29 | |
| 9 | hsa04370_VEGF_signaling_pathway | 16 | 76 | 9.53e-25 | 1.874e-23 | |
| 10 | hsa04910_Insulin_signaling_pathway | 18 | 138 | 9.127e-24 | 1.615e-22 | |
| 11 | hsa04062_Chemokine_signaling_pathway | 17 | 189 | 1.222e-19 | 1.966e-18 | |
| 12 | hsa04973_Carbohydrate_digestion_and_absorption | 11 | 44 | 3.504e-18 | 5.058e-17 | |
| 13 | hsa04914_Progesterone.mediated_oocyte_maturation | 13 | 87 | 3.715e-18 | 5.058e-17 | |
| 14 | hsa04150_mTOR_signaling_pathway | 11 | 52 | 2.7e-17 | 3.413e-16 | |
| 15 | hsa04664_Fc_epsilon_RI_signaling_pathway | 12 | 79 | 6.406e-17 | 7.559e-16 | |
| 16 | hsa04510_Focal_adhesion | 15 | 200 | 3.295e-16 | 3.645e-15 | |
| 17 | hsa04920_Adipocytokine_signaling_pathway | 11 | 68 | 6.556e-16 | 6.826e-15 | |
| 18 | hsa04622_RIG.I.like_receptor_signaling_pathway | 11 | 71 | 1.086e-15 | 1.068e-14 | |
| 19 | hsa04630_Jak.STAT_signaling_pathway | 13 | 155 | 8.617e-15 | 8.028e-14 | |
| 20 | hsa04012_ErbB_signaling_pathway | 11 | 87 | 1.136e-14 | 1.006e-13 | |
| 21 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 11 | 95 | 3.099e-14 | 2.612e-13 | |
| 22 | hsa04621_NOD.like_receptor_signaling_pathway | 9 | 59 | 5.778e-13 | 4.648e-12 | |
| 23 | hsa04960_Aldosterone.regulated_sodium_reabsorption | 8 | 42 | 1.8e-12 | 1.386e-11 | |
| 24 | hsa04115_p53_signaling_pathway | 9 | 69 | 2.534e-12 | 1.869e-11 | |
| 25 | hsa04070_Phosphatidylinositol_signaling_system | 8 | 78 | 3.233e-10 | 2.289e-09 | |
| 26 | hsa04720_Long.term_potentiation | 7 | 70 | 5.266e-09 | 3.585e-08 | |
| 27 | hsa04670_Leukocyte_transendothelial_migration | 8 | 117 | 8.427e-09 | 5.524e-08 | |
| 28 | hsa04623_Cytosolic_DNA.sensing_pathway | 6 | 56 | 4.554e-08 | 2.879e-07 | |
| 29 | hsa04310_Wnt_signaling_pathway | 8 | 151 | 6.236e-08 | 3.806e-07 | |
| 30 | hsa04114_Oocyte_meiosis | 7 | 114 | 1.598e-07 | 9.426e-07 | |
| 31 | hsa04810_Regulation_of_actin_cytoskeleton | 8 | 214 | 8.993e-07 | 5.135e-06 | |
| 32 | hsa04020_Calcium_signaling_pathway | 7 | 177 | 3.116e-06 | 1.724e-05 | |
| 33 | hsa00562_Inositol_phosphate_metabolism | 4 | 57 | 4.997e-05 | 0.000268 | |
| 34 | hsa04360_Axon_guidance | 5 | 130 | 9.918e-05 | 0.0005163 | |
| 35 | hsa04640_Hematopoietic_cell_lineage | 4 | 88 | 0.0002715 | 0.001373 | |
| 36 | hsa04141_Protein_processing_in_endoplasmic_reticulum | 4 | 168 | 0.003014 | 0.01482 | |
| 37 | hsa04962_Vasopressin.regulated_water_reabsorption | 2 | 44 | 0.01068 | 0.05107 | |
| 38 | hsa04530_Tight_junction | 3 | 133 | 0.01183 | 0.05508 | |
| 39 | hsa04742_Taste_transduction | 2 | 52 | 0.01469 | 0.06669 | |
| 40 | hsa04340_Hedgehog_signaling_pathway | 2 | 56 | 0.01691 | 0.07484 | |
| 41 | hsa04976_Bile_secretion | 2 | 71 | 0.02638 | 0.1139 | |
| 42 | hsa04971_Gastric_acid_secretion | 2 | 74 | 0.02848 | 0.12 | |
| 43 | hsa04970_Salivary_secretion | 2 | 89 | 0.03991 | 0.1638 | |
| 44 | hsa04540_Gap_junction | 2 | 90 | 0.04073 | 0.1638 | |
| 45 | hsa04912_GnRH_signaling_pathway | 2 | 101 | 0.0501 | 0.1928 | |
| 46 | hsa04916_Melanogenesis | 2 | 101 | 0.0501 | 0.1928 | |
| 47 | hsa04270_Vascular_smooth_muscle_contraction | 2 | 116 | 0.06399 | 0.241 | |
| 48 | hsa04110_Cell_cycle | 2 | 128 | 0.07591 | 0.2799 | |
| 49 | hsa04120_Ubiquitin_mediated_proteolysis | 2 | 139 | 0.08741 | 0.3158 | |
| 50 | hsa04740_Olfactory_transduction | 2 | 388 | 0.4018 | 1 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | MAGI2-AS3 |
hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-320c;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-505-3p;hsa-miR-548v;hsa-miR-616-5p;hsa-miR-769-5p;hsa-miR-93-5p | 17 | AKT3 | Sponge network | -3.29 | 0.00119 | -3.432 | 0 | 0.642 |
| 2 | RP11-696N14.1 |
hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-29a-3p;hsa-miR-338-5p;hsa-miR-584-5p | 10 | PIK3R1 | Sponge network | -0.342 | 0.704 | -0.896 | 0.00072 | 0.469 |
| 3 | HAND2-AS1 |
hsa-miR-106b-5p;hsa-miR-107;hsa-miR-146b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-17-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-320b;hsa-miR-421;hsa-miR-616-5p;hsa-miR-93-5p | 13 | AKT3 | Sponge network | -3.916 | 0.01209 | -3.432 | 0 | 0.459 |
| 4 | LINC00641 |
hsa-let-7f-1-3p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-let-7i-5p;hsa-miR-106b-5p;hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-186-5p;hsa-miR-590-3p;hsa-miR-628-3p | 10 | PPP3CA | Sponge network | -1.526 | 0.02432 | -0.565 | 4.0E-5 | 0.446 |
| 5 | MAGI2-AS3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-107;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-32-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-92a-3p | 11 | PRKAR2B | Sponge network | -3.29 | 0.00119 | -2.601 | 0 | 0.439 |
| 6 | MAGI2-AS3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7d-3p;hsa-let-7f-1-3p;hsa-miR-141-3p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-194-3p;hsa-miR-200a-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-32-3p;hsa-miR-339-5p;hsa-miR-429;hsa-miR-548v;hsa-miR-590-3p;hsa-miR-616-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 23 | PRKACB | Sponge network | -3.29 | 0.00119 | -1.289 | 0 | 0.42 |
| 7 | HAND2-AS1 |
hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-144-3p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-324-3p;hsa-miR-330-3p;hsa-miR-338-5p;hsa-miR-486-5p;hsa-miR-584-5p;hsa-miR-629-3p;hsa-miR-93-5p | 19 | PIK3R1 | Sponge network | -3.916 | 0.01209 | -0.896 | 0.00072 | 0.374 |
| 8 | EMX2OS |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-186-5p;hsa-miR-215-5p;hsa-miR-24-2-5p;hsa-miR-32-3p;hsa-miR-33b-5p;hsa-miR-342-3p;hsa-miR-590-3p;hsa-miR-629-5p | 12 | BCL2 | Sponge network | -2.112 | 0.14094 | -2.274 | 0 | 0.368 |
| 9 | PLAC4 | hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-338-5p;hsa-miR-409-3p;hsa-miR-93-5p | 11 | PIK3R1 | Sponge network | 1.014 | 0.01175 | -0.896 | 0.00072 | 0.347 |
| 10 | RP1-60O19.1 |
hsa-miR-16-1-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-205-5p;hsa-miR-20a-3p;hsa-miR-25-3p;hsa-miR-3200-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-320c;hsa-miR-33a-5p;hsa-miR-361-5p;hsa-miR-590-3p;hsa-miR-9-3p;hsa-miR-92a-3p | 15 | IRAK3 | Sponge network | -3.138 | 0.16483 | -1.695 | 0 | 0.328 |
| 11 | RP11-166D19.1 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-15b-3p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-186-5p;hsa-miR-192-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-215-5p;hsa-miR-3065-5p;hsa-miR-32-3p;hsa-miR-33a-5p;hsa-miR-33b-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-616-5p;hsa-miR-629-5p | 20 | BCL2 | Sponge network | -3.146 | 0.02178 | -2.274 | 0 | 0.324 |
| 12 | LINC00667 |
hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-141-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-200a-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-32-3p;hsa-miR-429;hsa-miR-548v;hsa-miR-96-5p | 12 | PRKACB | Sponge network | -2.558 | 0.01903 | -1.289 | 0 | 0.31 |
| 13 | USP3-AS1 |
hsa-miR-103a-3p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-144-3p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-335-3p;hsa-miR-93-5p | 14 | PIK3R1 | Sponge network | -1.35 | 0.3108 | -0.896 | 0.00072 | 0.303 |
| 14 | RP11-166D19.1 |
hsa-let-7d-3p;hsa-let-7f-1-3p;hsa-miR-16-2-3p;hsa-miR-194-3p;hsa-miR-200a-3p;hsa-miR-3065-5p;hsa-miR-32-3p;hsa-miR-330-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-616-5p | 12 | PRKACB | Sponge network | -3.146 | 0.02178 | -1.289 | 0 | 0.295 |
| 15 | RP11-696N14.1 |
hsa-miR-142-3p;hsa-miR-15b-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-224-5p;hsa-miR-24-2-5p;hsa-miR-338-5p;hsa-miR-342-3p;hsa-miR-582-5p | 11 | BCL2 | Sponge network | -0.342 | 0.704 | -2.274 | 0 | 0.291 |
| 16 | RP11-161M6.2 |
hsa-miR-107;hsa-miR-15b-5p;hsa-miR-17-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-769-5p;hsa-miR-93-5p | 10 | AKT3 | Sponge network | -0.737 | 0.61297 | -3.432 | 0 | 0.278 |
| 17 | CASC2 |
hsa-miR-1301-3p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-324-3p;hsa-miR-330-3p;hsa-miR-335-3p;hsa-miR-486-5p;hsa-miR-629-3p | 13 | PIK3R1 | Sponge network | -2.463 | 0.00078 | -0.896 | 0.00072 | 0.274 |
| 18 | ZNF883 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7d-3p;hsa-let-7f-1-3p;hsa-miR-16-2-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-194-3p;hsa-miR-2110;hsa-miR-590-5p;hsa-miR-616-5p | 11 | PRKACB | Sponge network | -1.835 | 2.0E-5 | -1.289 | 0 | 0.272 |
| 19 | MAGI2-AS3 |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-192-5p;hsa-miR-200b-3p;hsa-miR-25-3p;hsa-miR-30e-5p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-320c;hsa-miR-429;hsa-miR-590-3p;hsa-miR-616-5p | 17 | PRKAR1A | Sponge network | -3.29 | 0.00119 | -0.941 | 0 | 0.267 |
| 20 | LINC00473 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-20a-5p;hsa-miR-320a;hsa-miR-320c;hsa-miR-335-3p;hsa-miR-616-5p;hsa-miR-93-5p | 14 | AKT3 | Sponge network | -6.267 | 0.02511 | -3.432 | 0 | 0.265 |
| 21 | MAGI2-AS3 |
hsa-miR-130b-5p;hsa-miR-15a-5p;hsa-miR-15b-3p;hsa-miR-15b-5p;hsa-miR-16-1-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-192-5p;hsa-miR-200a-5p;hsa-miR-200b-3p;hsa-miR-200b-5p;hsa-miR-200c-3p;hsa-miR-32-3p;hsa-miR-33a-5p;hsa-miR-33b-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-616-5p;hsa-miR-629-5p;hsa-miR-7-1-3p;hsa-miR-7-5p | 24 | BCL2 | Sponge network | -3.29 | 0.00119 | -2.274 | 0 | 0.262 |
| 22 | MAGI2-AS3 |
hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-3065-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-33a-3p;hsa-miR-590-3p;hsa-miR-616-5p;hsa-miR-628-3p;hsa-miR-7-1-3p;hsa-miR-93-3p;hsa-miR-940 | 16 | PPP3CA | Sponge network | -3.29 | 0.00119 | -0.565 | 4.0E-5 | 0.26 |
| 23 | LINC00663 | hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-33b-5p;hsa-miR-590-3p;hsa-miR-629-5p | 10 | BCL2 | Sponge network | -2.496 | 4.0E-5 | -2.274 | 0 | 0.258 |