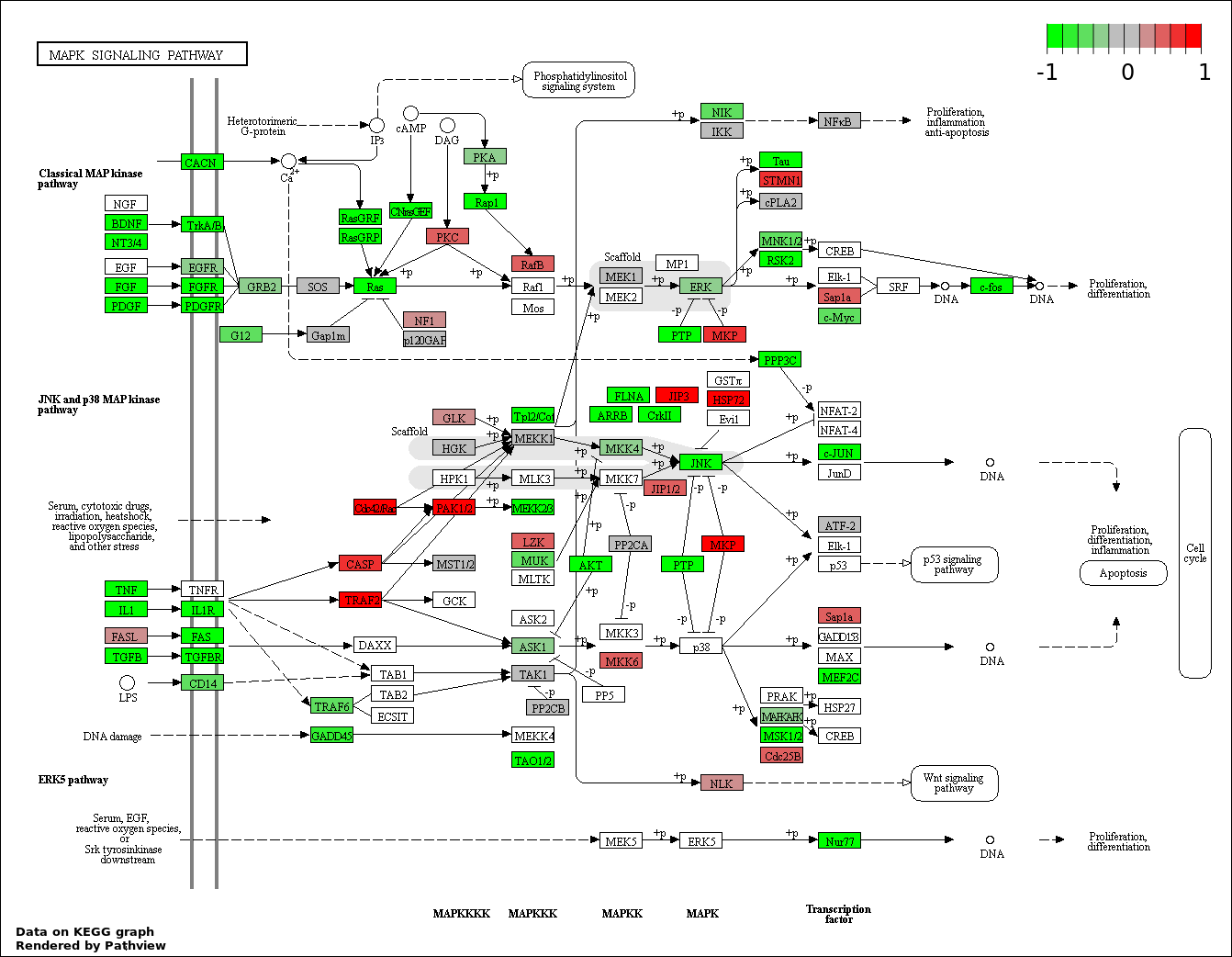
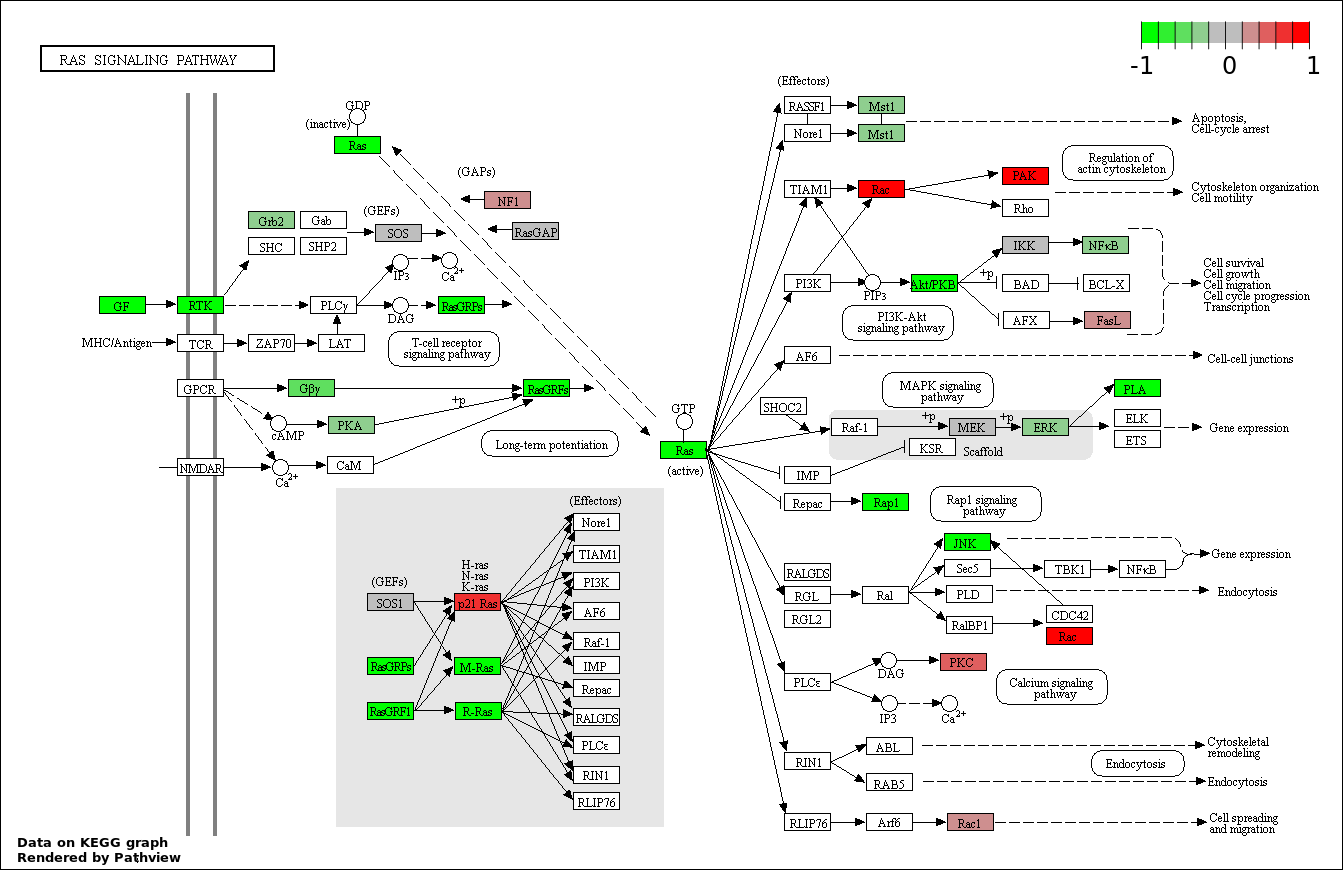
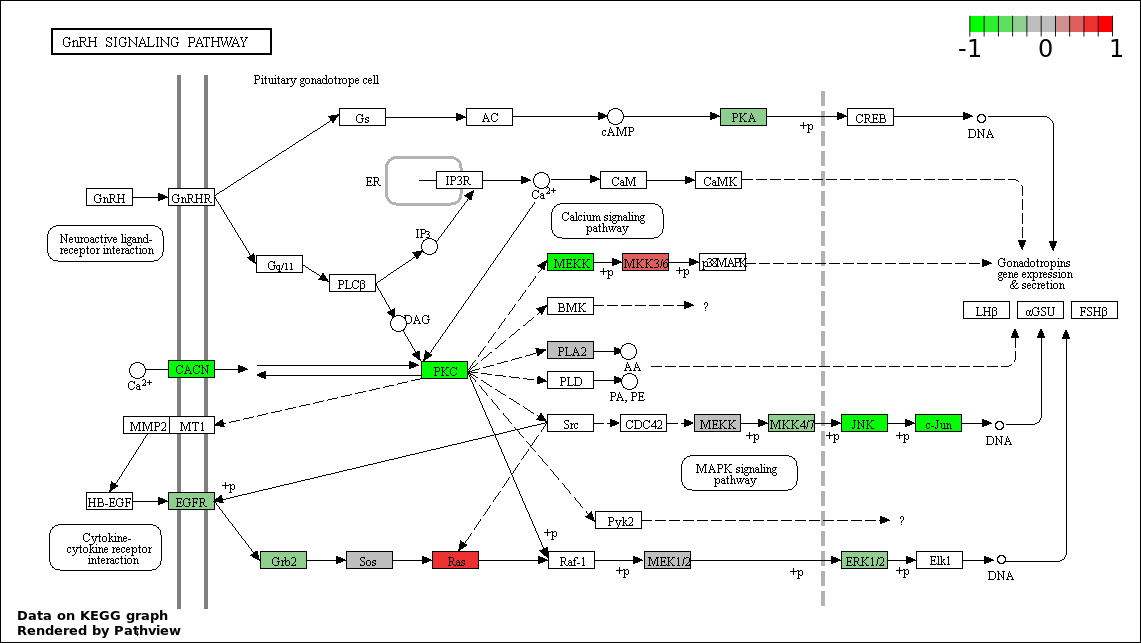
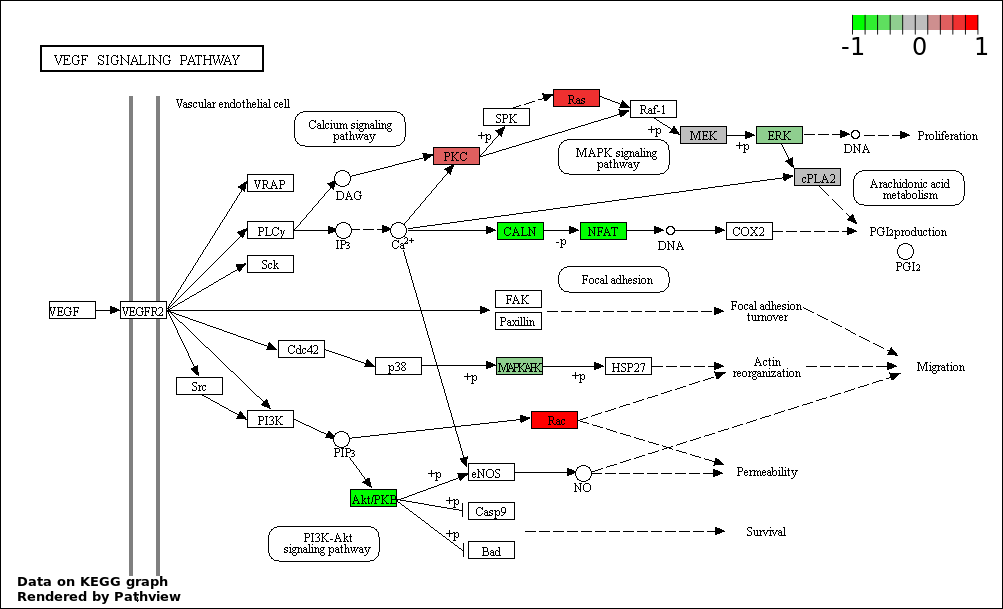
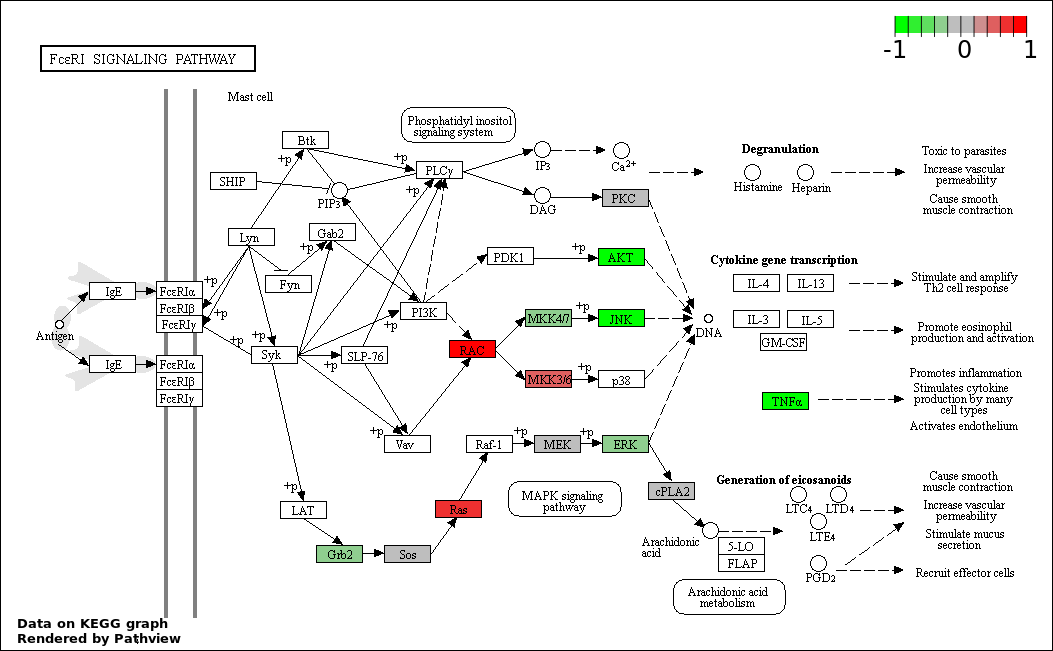
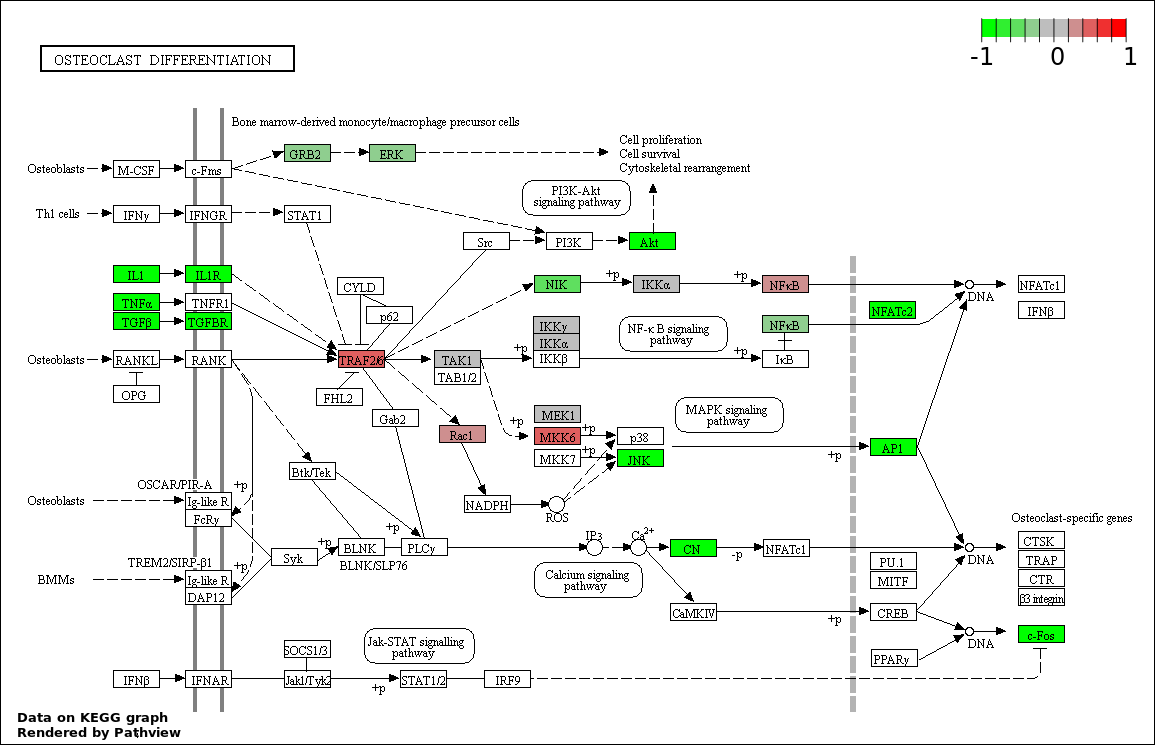
This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-let-7a-5p | AKT2 | -1.37 | 0 | 0.16 | 0.1135 | TargetScan | -0.12 | 0 | NA | |
| 2 | hsa-miR-29a-3p | AKT2 | 0.1 | 0.5732 | 0.16 | 0.1135 | MirTarget | -0.1 | 4.0E-5 | 24076586 | Furthermore a feed-back loop comprising of c-Myc miR-29 family and Akt2 were found in myeloid leukemogenesis |
| 3 | hsa-miR-106b-5p | AKT3 | 1.47 | 0 | -1.44 | 0 | miRNATAP | -0.16 | 0.00426 | NA | |
| 4 | hsa-miR-107 | AKT3 | 0.66 | 0 | -1.44 | 0 | PITA; miRanda | -0.26 | 0.0031 | NA | |
| 5 | hsa-miR-146b-5p | AKT3 | 1.09 | 1.0E-5 | -1.44 | 0 | miRNAWalker2 validate | -0.15 | 0.00189 | NA | |
| 6 | hsa-miR-15a-5p | AKT3 | 1.63 | 0 | -1.44 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.41 | 0 | NA | |
| 7 | hsa-miR-16-5p | AKT3 | 0.75 | 0 | -1.44 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.27 | 0.0001 | NA | |
| 8 | hsa-miR-17-5p | AKT3 | 2.07 | 0 | -1.44 | 0 | TargetScan; miRNATAP | -0.22 | 0 | NA | |
| 9 | hsa-miR-181a-5p | AKT3 | -0.38 | 0.05621 | -1.44 | 0 | miRNATAP | -0.23 | 9.0E-5 | NA | |
| 10 | hsa-miR-181b-5p | AKT3 | 0.67 | 0.00024 | -1.44 | 0 | miRNATAP | -0.37 | 0 | NA | |
| 11 | hsa-miR-20a-5p | AKT3 | 2.65 | 0 | -1.44 | 0 | miRNATAP | -0.24 | 0 | NA | |
| 12 | hsa-miR-22-3p | AKT3 | 1.43 | 0 | -1.44 | 0 | miRNATAP | -0.26 | 0.00109 | NA | |
| 13 | hsa-miR-29b-2-5p | AKT3 | 0.35 | 0.19484 | -1.44 | 0 | mirMAP | -0.18 | 2.0E-5 | NA | |
| 14 | hsa-miR-29b-3p | AKT3 | 3.11 | 0 | -1.44 | 0 | miRNATAP | -0.27 | 0 | 26512921 | MicroRNA 29B mir 29b regulates the Warburg effect in ovarian cancer by targeting AKT2 and AKT3 |
| 15 | hsa-miR-3065-5p | AKT3 | 0.65 | 0.09995 | -1.44 | 0 | mirMAP | -0.18 | 0 | NA | |
| 16 | hsa-miR-362-5p | AKT3 | 0.66 | 0.02433 | -1.44 | 0 | PITA; TargetScan; miRNATAP | -0.23 | 0 | NA | |
| 17 | hsa-miR-542-3p | AKT3 | 1.62 | 0 | -1.44 | 0 | miRanda | -0.19 | 1.0E-5 | NA | |
| 18 | hsa-miR-93-5p | AKT3 | 1.51 | 0 | -1.44 | 0 | miRNATAP | -0.25 | 0 | NA | |
| 19 | hsa-miR-146b-5p | ARRB1 | 1.09 | 1.0E-5 | -1.4 | 0 | miRanda | -0.11 | 0.00436 | NA | |
| 20 | hsa-miR-219a-1-3p | ARRB1 | 0.43 | 0.08924 | -1.4 | 0 | MirTarget | -0.11 | 0.0058 | NA | |
| 21 | hsa-miR-22-3p | ARRB1 | 1.43 | 0 | -1.4 | 0 | MirTarget | -0.28 | 5.0E-5 | NA | |
| 22 | hsa-miR-361-5p | ARRB1 | 0.21 | 0.0801 | -1.4 | 0 | MirTarget; miRanda | -0.22 | 0.00729 | NA | |
| 23 | hsa-miR-455-3p | ARRB1 | 0.59 | 0.04448 | -1.4 | 0 | MirTarget | -0.13 | 0.0001 | NA | |
| 24 | hsa-miR-550a-5p | ARRB1 | 0.6 | 0.03148 | -1.4 | 0 | MirTarget | -0.12 | 0.0006 | NA | |
| 25 | hsa-miR-671-5p | ARRB1 | 2.24 | 0 | -1.4 | 0 | MirTarget | -0.23 | 0 | NA | |
| 26 | hsa-let-7a-3p | ATF2 | 0.17 | 0.43183 | 0.01 | 0.93689 | MirTarget; mirMAP; miRNATAP | -0.13 | 0.0001 | NA | |
| 27 | hsa-miR-140-3p | ATF2 | -1.11 | 0 | 0.01 | 0.93689 | MirTarget; PITA; miRNATAP | -0.16 | 0.00016 | NA | |
| 28 | hsa-miR-15a-5p | ATF2 | 1.63 | 0 | 0.01 | 0.93689 | miRNAWalker2 validate | -0.19 | 0 | NA | |
| 29 | hsa-miR-181a-5p | ATF2 | -0.38 | 0.05621 | 0.01 | 0.93689 | mirMAP; miRNATAP | -0.15 | 0 | NA | |
| 30 | hsa-miR-222-3p | ATF2 | 0.03 | 0.88194 | 0.01 | 0.93689 | MirTarget | -0.13 | 0 | NA | |
| 31 | hsa-miR-26b-5p | ATF2 | 0.72 | 5.0E-5 | 0.01 | 0.93689 | MirTarget; miRNATAP | -0.12 | 0.00049 | 21901137 | Coordinated regulation of ATF2 by miR 26b in γ irradiated lung cancer cells; Concurrent analysis of time-series mRNA and microRNA profiles uncovered that expression of miR-26b was down regulated and its target activating transcription factor 2 ATF2 mRNA was up regulated in γ-irradiated H1299 cells; IR in miR-26b overexpressed H1299 cells could not induce expression of ATF2; From these results we concluded that IR-induced up-regulation of ATF2 was coordinately enhanced by suppression of miR-26b in lung cancer cells which may enhance the effect of IR in the MAPK signaling pathway |
| 32 | hsa-miR-30b-5p | ATF2 | 0.36 | 0.13803 | 0.01 | 0.93689 | mirMAP | -0.18 | 0 | NA | |
| 33 | hsa-miR-30c-5p | ATF2 | -0.33 | 0.1236 | 0.01 | 0.93689 | mirMAP | -0.26 | 0 | NA | |
| 34 | hsa-miR-374a-5p | ATF2 | -0.2 | 0.29808 | 0.01 | 0.93689 | mirMAP | -0.16 | 1.0E-5 | NA | |
| 35 | hsa-miR-10a-5p | BDNF | 0.79 | 0.00059 | -3.58 | 0 | MirTarget; miRNATAP | -0.28 | 0.00181 | NA | |
| 36 | hsa-miR-146b-5p | BDNF | 1.09 | 1.0E-5 | -3.58 | 0 | miRanda | -0.33 | 0.00012 | NA | |
| 37 | hsa-miR-148a-5p | BDNF | 1.46 | 0 | -3.58 | 0 | mirMAP | -0.24 | 0.00212 | NA | |
| 38 | hsa-miR-155-5p | BDNF | 0.81 | 0.00061 | -3.58 | 0 | miRNATAP | -0.23 | 0.00731 | NA | |
| 39 | hsa-miR-15a-5p | BDNF | 1.63 | 0 | -3.58 | 0 | MirTarget; miRNATAP | -0.3 | 0.00719 | 26581909 | MicroRNA 15a 5p suppresses cancer proliferation and division in human hepatocellular carcinoma by targeting BDNF; BDNF was then overexpressed in HepG2 and SNU-182 cells to evaluate its selective effect on miR-15a-5p in HCC modulation; MiR-15a-5p selectively and negatively regulated BDNF at both gene and protein levels in HCC cells; Forced overexpression of BDNF effectively reversed the tumor suppressive functions of miR-15a-5p on HCC proliferation and cell division in vitro; Our study demonstrated that miR-15a-5p is a tumor suppressor in HCC and its regulation is through BDNF in HCC |
| 40 | hsa-miR-182-5p | BDNF | 3.22 | 0 | -3.58 | 0 | MirTarget; miRNATAP | -0.49 | 0 | NA | |
| 41 | hsa-miR-210-3p | BDNF | 4.89 | 0 | -3.58 | 0 | miRNAWalker2 validate; miRTarBase | -0.22 | 0 | NA | |
| 42 | hsa-miR-29b-1-5p | BDNF | 1.71 | 0 | -3.58 | 0 | mirMAP | -0.25 | 0.00028 | NA | |
| 43 | hsa-miR-30e-5p | BDNF | 1.6 | 0 | -3.58 | 0 | miRNATAP | -0.44 | 9.0E-5 | NA | |
| 44 | hsa-miR-7-1-3p | BDNF | 2.61 | 0 | -3.58 | 0 | MirTarget | -0.3 | 0.00109 | NA | |
| 45 | hsa-miR-199a-5p | BRAF | 1.31 | 0 | 0.43 | 0.00363 | miRanda | -0.13 | 1.0E-5 | NA | |
| 46 | hsa-miR-199b-5p | BRAF | 2.14 | 0 | 0.43 | 0.00363 | miRanda | -0.11 | 1.0E-5 | NA | |
| 47 | hsa-miR-29b-2-5p | CACNA1A | 0.35 | 0.19484 | 0.08 | 0.81462 | mirMAP | -0.19 | 0.00112 | NA | |
| 48 | hsa-miR-107 | CACNA1C | 0.66 | 0 | -1.13 | 2.0E-5 | miRNATAP | -0.43 | 0 | NA | |
| 49 | hsa-miR-148b-5p | CACNA1C | 1.39 | 0 | -1.13 | 2.0E-5 | mirMAP | -0.25 | 0 | NA | |
| 50 | hsa-miR-186-5p | CACNA1C | 0.85 | 0 | -1.13 | 2.0E-5 | mirMAP | -0.36 | 4.0E-5 | NA | |
| 51 | hsa-miR-19a-3p | CACNA1C | 2.12 | 0 | -1.13 | 2.0E-5 | MirTarget | -0.17 | 3.0E-5 | NA | |
| 52 | hsa-miR-19b-1-5p | CACNA1C | 1.71 | 0 | -1.13 | 2.0E-5 | MirTarget | -0.32 | 0 | NA | |
| 53 | hsa-miR-19b-3p | CACNA1C | 2.11 | 0 | -1.13 | 2.0E-5 | MirTarget | -0.19 | 0.00017 | NA | |
| 54 | hsa-miR-200c-3p | CACNA1C | 0.38 | 0.08422 | -1.13 | 2.0E-5 | MirTarget | -0.21 | 0.00011 | NA | |
| 55 | hsa-miR-33a-5p | CACNA1C | 2.8 | 0 | -1.13 | 2.0E-5 | MirTarget | -0.18 | 0 | NA | |
| 56 | hsa-miR-421 | CACNA1C | 0.17 | 0.53528 | -1.13 | 2.0E-5 | MirTarget; PITA; miRanda; miRNATAP | -0.32 | 0 | NA | |
| 57 | hsa-miR-450b-5p | CACNA1C | 1.69 | 0 | -1.13 | 2.0E-5 | PITA; miRNATAP | -0.19 | 1.0E-5 | NA | |
| 58 | hsa-miR-542-3p | CACNA1C | 1.62 | 0 | -1.13 | 2.0E-5 | miRanda | -0.2 | 2.0E-5 | NA | |
| 59 | hsa-miR-589-3p | CACNA1C | 1.34 | 2.0E-5 | -1.13 | 2.0E-5 | MirTarget; mirMAP | -0.12 | 0.00331 | NA | |
| 60 | hsa-miR-7-1-3p | CACNA1C | 2.61 | 0 | -1.13 | 2.0E-5 | MirTarget | -0.22 | 6.0E-5 | NA | |
| 61 | hsa-miR-877-5p | CACNA1C | -0.37 | 0.20671 | -1.13 | 2.0E-5 | MirTarget | -0.15 | 0.00044 | NA | |
| 62 | hsa-miR-93-3p | CACNA1C | 1.08 | 2.0E-5 | -1.13 | 2.0E-5 | mirMAP | -0.21 | 1.0E-5 | NA | |
| 63 | hsa-miR-96-5p | CACNA1C | 3.04 | 0 | -1.13 | 2.0E-5 | MirTarget; TargetScan | -0.37 | 0 | NA | |
| 64 | hsa-let-7a-2-3p | CACNA1D | 1.34 | 0.00037 | -1.18 | 0.00432 | MirTarget | -0.35 | 0 | NA | |
| 65 | hsa-let-7i-5p | CACNA1D | 0.61 | 0 | -1.18 | 0.00432 | miRNATAP | -0.48 | 0.00147 | NA | |
| 66 | hsa-miR-134-5p | CACNA1D | 0.17 | 0.62286 | -1.18 | 0.00432 | MirTarget | -0.2 | 0.00026 | NA | |
| 67 | hsa-miR-142-3p | CACNA1D | 3.98 | 0 | -1.18 | 0.00432 | miRanda | -0.25 | 0 | NA | |
| 68 | hsa-miR-542-3p | CACNA1D | 1.62 | 0 | -1.18 | 0.00432 | miRanda | -0.34 | 0 | NA | |
| 69 | hsa-miR-590-3p | CACNA1D | 0.84 | 0.00129 | -1.18 | 0.00432 | miRanda; miRNATAP | -0.37 | 2.0E-5 | NA | |
| 70 | hsa-miR-629-5p | CACNA1D | 1.32 | 0 | -1.18 | 0.00432 | MirTarget | -0.48 | 0 | NA | |
| 71 | hsa-let-7a-5p | CACNA1E | -1.37 | 0 | 1.62 | 0.0002 | TargetScan; miRNATAP | -0.57 | 0 | NA | |
| 72 | hsa-let-7b-5p | CACNA1E | -1.62 | 0 | 1.62 | 0.0002 | miRNATAP | -0.4 | 3.0E-5 | NA | |
| 73 | hsa-let-7e-5p | CACNA1E | -0.75 | 7.0E-5 | 1.62 | 0.0002 | miRNATAP | -0.41 | 0.00013 | NA | |
| 74 | hsa-let-7f-5p | CACNA1E | -0.05 | 0.83408 | 1.62 | 0.0002 | miRNATAP | -0.25 | 0.00167 | NA | |
| 75 | hsa-miR-125b-2-3p | CACNA1E | -1.71 | 0 | 1.62 | 0.0002 | mirMAP | -0.24 | 0.00118 | NA | |
| 76 | hsa-miR-1266-5p | CACNA1E | -0.29 | 0.2815 | 1.62 | 0.0002 | MirTarget | -0.33 | 1.0E-5 | NA | |
| 77 | hsa-miR-143-3p | CACNA1E | -1.21 | 1.0E-5 | 1.62 | 0.0002 | miRNATAP | -0.22 | 0.00282 | NA | |
| 78 | hsa-miR-195-3p | CACNA1E | -1.33 | 0 | 1.62 | 0.0002 | mirMAP | -0.24 | 0.00527 | NA | |
| 79 | hsa-miR-195-5p | CACNA1E | -1.02 | 5.0E-5 | 1.62 | 0.0002 | MirTarget | -0.24 | 0.00385 | NA | |
| 80 | hsa-miR-2110 | CACNA1E | -1.92 | 0 | 1.62 | 0.0002 | mirMAP | -0.23 | 0.00207 | NA | |
| 81 | hsa-miR-3065-3p | CACNA1E | 0.38 | 0.30796 | 1.62 | 0.0002 | MirTarget; miRNATAP | -0.17 | 0.00132 | NA | |
| 82 | hsa-miR-30a-3p | CACNA1E | -2.54 | 0 | 1.62 | 0.0002 | MirTarget; mirMAP; miRNATAP | -0.32 | 0 | NA | |
| 83 | hsa-miR-30b-3p | CACNA1E | -1.21 | 0 | 1.62 | 0.0002 | mirMAP | -0.26 | 0.00078 | NA | |
| 84 | hsa-miR-30d-3p | CACNA1E | 0 | 0.98646 | 1.62 | 0.0002 | MirTarget; mirMAP; miRNATAP | -0.32 | 0.00105 | NA | |
| 85 | hsa-miR-30e-3p | CACNA1E | -0.1 | 0.52624 | 1.62 | 0.0002 | MirTarget; mirMAP; miRNATAP | -0.47 | 0.00014 | NA | |
| 86 | hsa-miR-664a-5p | CACNA1E | -0.09 | 0.66227 | 1.62 | 0.0002 | mirMAP | -0.28 | 0.00463 | NA | |
| 87 | hsa-miR-151a-3p | CACNA1G | 0.67 | 0.00028 | -0.03 | 0.95247 | MirTarget | -0.34 | 0.0014 | NA | |
| 88 | hsa-miR-590-3p | CACNA1G | 0.84 | 0.00129 | -0.03 | 0.95247 | miRanda | -0.44 | 0 | NA | |
| 89 | hsa-miR-590-5p | CACNA1G | 2.07 | 0 | -0.03 | 0.95247 | miRanda | -0.23 | 0.00866 | NA | |
| 90 | hsa-miR-629-3p | CACNA1G | 1.32 | 0.00011 | -0.03 | 0.95247 | MirTarget | -0.22 | 0.00032 | NA | |
| 91 | hsa-let-7a-5p | CACNA1I | -1.37 | 0 | 1.05 | 0.02434 | TargetScan; miRNATAP | -0.52 | 1.0E-5 | NA | |
| 92 | hsa-let-7b-5p | CACNA1I | -1.62 | 0 | 1.05 | 0.02434 | miRNATAP | -0.39 | 0.00013 | NA | |
| 93 | hsa-let-7e-5p | CACNA1I | -0.75 | 7.0E-5 | 1.05 | 0.02434 | miRNATAP | -0.33 | 0.00387 | NA | |
| 94 | hsa-let-7f-5p | CACNA1I | -0.05 | 0.83408 | 1.05 | 0.02434 | miRNATAP | -0.28 | 0.00132 | NA | |
| 95 | hsa-miR-664a-5p | CACNA1I | -0.09 | 0.66227 | 1.05 | 0.02434 | mirMAP | -0.34 | 0.00109 | NA | |
| 96 | hsa-miR-103a-3p | CACNA2D1 | 0.54 | 2.0E-5 | -1.27 | 0.00469 | miRNATAP | -0.61 | 0.00023 | NA | |
| 97 | hsa-miR-107 | CACNA2D1 | 0.66 | 0 | -1.27 | 0.00469 | PITA; miRanda; miRNATAP | -0.55 | 0.00047 | NA | |
| 98 | hsa-miR-141-3p | CACNA2D1 | 3.37 | 0 | -1.27 | 0.00469 | TargetScan | -0.47 | 0 | NA | |
| 99 | hsa-miR-16-2-3p | CACNA2D1 | 0.5 | 0.02636 | -1.27 | 0.00469 | mirMAP | -0.27 | 0.00622 | NA | |
| 100 | hsa-miR-16-5p | CACNA2D1 | 0.75 | 0 | -1.27 | 0.00469 | miRNAWalker2 validate | -0.35 | 0.00593 | NA | |
| 101 | hsa-miR-181a-5p | CACNA2D1 | -0.38 | 0.05621 | -1.27 | 0.00469 | mirMAP | -0.32 | 0.00252 | NA | |
| 102 | hsa-miR-181b-5p | CACNA2D1 | 0.67 | 0.00024 | -1.27 | 0.00469 | mirMAP | -0.51 | 1.0E-5 | NA | |
| 103 | hsa-miR-182-5p | CACNA2D1 | 3.22 | 0 | -1.27 | 0.00469 | miRNATAP | -0.28 | 0.00028 | NA | |
| 104 | hsa-miR-200b-3p | CACNA2D1 | 1.55 | 0 | -1.27 | 0.00469 | TargetScan | -0.42 | 0 | NA | |
| 105 | hsa-miR-26b-5p | CACNA2D1 | 0.72 | 5.0E-5 | -1.27 | 0.00469 | mirMAP | -0.53 | 0 | NA | |
| 106 | hsa-miR-3065-5p | CACNA2D1 | 0.65 | 0.09995 | -1.27 | 0.00469 | mirMAP | -0.28 | 0 | NA | |
| 107 | hsa-miR-30d-3p | CACNA2D1 | 0 | 0.98646 | -1.27 | 0.00469 | mirMAP | -0.26 | 0.0086 | NA | |
| 108 | hsa-miR-338-3p | CACNA2D1 | 0.73 | 0.05063 | -1.27 | 0.00469 | miRanda | -0.19 | 0.00049 | NA | |
| 109 | hsa-miR-362-3p | CACNA2D1 | 0.19 | 0.52808 | -1.27 | 0.00469 | miRanda | -0.33 | 0.00011 | NA | |
| 110 | hsa-miR-450b-5p | CACNA2D1 | 1.69 | 0 | -1.27 | 0.00469 | mirMAP | -0.25 | 0.00033 | NA | |
| 111 | hsa-miR-576-5p | CACNA2D1 | 1.03 | 0 | -1.27 | 0.00469 | mirMAP | -0.27 | 0.00614 | NA | |
| 112 | hsa-miR-590-3p | CACNA2D1 | 0.84 | 0.00129 | -1.27 | 0.00469 | mirMAP | -0.25 | 0.0093 | NA | |
| 113 | hsa-miR-96-5p | CACNA2D1 | 3.04 | 0 | -1.27 | 0.00469 | TargetScan; miRNATAP | -0.43 | 0 | NA | |
| 114 | hsa-miR-10a-5p | CACNA2D2 | 0.79 | 0.00059 | -4.05 | 0 | mirMAP | -0.41 | 0.00076 | NA | |
| 115 | hsa-miR-1271-5p | CACNA2D2 | -0.02 | 0.9511 | -4.05 | 0 | MirTarget | -0.28 | 0.00618 | NA | |
| 116 | hsa-miR-142-3p | CACNA2D2 | 3.98 | 0 | -4.05 | 0 | miRanda | -0.53 | 0 | NA | |
| 117 | hsa-miR-182-5p | CACNA2D2 | 3.22 | 0 | -4.05 | 0 | miRNATAP | -0.34 | 0.00135 | NA | |
| 118 | hsa-miR-193a-3p | CACNA2D2 | 0.55 | 0.0319 | -4.05 | 0 | miRanda | -0.81 | 0 | NA | |
| 119 | hsa-miR-205-5p | CACNA2D2 | 1.45 | 0.03618 | -4.05 | 0 | miRNATAP | -0.12 | 0.00281 | NA | |
| 120 | hsa-miR-22-5p | CACNA2D2 | 1.71 | 0 | -4.05 | 0 | mirMAP | -0.68 | 0 | NA | |
| 121 | hsa-miR-3607-3p | CACNA2D2 | 2.41 | 0 | -4.05 | 0 | miRNATAP | -0.17 | 0.00635 | NA | |
| 122 | hsa-miR-455-3p | CACNA2D2 | 0.59 | 0.04448 | -4.05 | 0 | PITA; miRNATAP | -0.27 | 0.00467 | NA | |
| 123 | hsa-miR-590-3p | CACNA2D2 | 0.84 | 0.00129 | -4.05 | 0 | miRanda | -0.91 | 0 | NA | |
| 124 | hsa-miR-96-5p | CACNA2D2 | 3.04 | 0 | -4.05 | 0 | MirTarget; TargetScan; miRNATAP | -0.46 | 1.0E-5 | NA | |
| 125 | hsa-miR-421 | CACNA2D3 | 0.17 | 0.53528 | -0.94 | 0.01221 | miRanda | -0.21 | 0.00273 | NA | |
| 126 | hsa-miR-590-3p | CACNA2D3 | 0.84 | 0.00129 | -0.94 | 0.01221 | miRanda | -0.38 | 0 | NA | |
| 127 | hsa-miR-335-3p | CACNA2D4 | 1.51 | 0 | -0.25 | 0.17008 | mirMAP | -0.2 | 0 | NA | |
| 128 | hsa-miR-335-5p | CACNA2D4 | -0.47 | 0.0677 | -0.25 | 0.17008 | miRNAWalker2 validate | -0.1 | 0.00569 | NA | |
| 129 | hsa-miR-429 | CACNA2D4 | 2.38 | 0 | -0.25 | 0.17008 | miRanda | -0.12 | 1.0E-5 | NA | |
| 130 | hsa-miR-24-3p | CACNB1 | -0.04 | 0.78587 | 0.36 | 0.20122 | miRNATAP | -0.35 | 0.00051 | NA | |
| 131 | hsa-miR-320b | CACNB1 | 0.23 | 0.37882 | 0.36 | 0.20122 | miRanda | -0.19 | 0.00021 | NA | |
| 132 | hsa-miR-181b-5p | CACNB2 | 0.67 | 0.00024 | -0.6 | 0.03617 | miRNATAP | -0.22 | 0.00304 | NA | |
| 133 | hsa-miR-182-5p | CACNB2 | 3.22 | 0 | -0.6 | 0.03617 | miRNATAP | -0.22 | 1.0E-5 | NA | |
| 134 | hsa-miR-29a-5p | CACNB2 | 1.9 | 0 | -0.6 | 0.03617 | MirTarget | -0.15 | 0.00665 | NA | |
| 135 | hsa-miR-96-5p | CACNB2 | 3.04 | 0 | -0.6 | 0.03617 | miRNATAP | -0.18 | 0.00019 | NA | |
| 136 | hsa-miR-140-3p | CACNB3 | -1.11 | 0 | 0.91 | 1.0E-5 | PITA; miRNATAP | -0.33 | 0 | NA | |
| 137 | hsa-miR-150-5p | CACNB3 | -0.7 | 0.02153 | 0.91 | 1.0E-5 | miRNATAP | -0.16 | 0 | NA | |
| 138 | hsa-miR-342-5p | CACNB3 | -0.38 | 0.0897 | 0.91 | 1.0E-5 | PITA | -0.15 | 0.0004 | NA | |
| 139 | hsa-let-7g-3p | CACNB4 | 2.38 | 0 | -2.29 | 0 | miRNATAP | -0.24 | 1.0E-5 | NA | |
| 140 | hsa-miR-132-3p | CACNB4 | 0.01 | 0.93403 | -2.29 | 0 | miRNATAP | -0.31 | 0.00417 | NA | |
| 141 | hsa-miR-182-5p | CACNB4 | 3.22 | 0 | -2.29 | 0 | miRNATAP | -0.21 | 0.00049 | NA | |
| 142 | hsa-miR-183-5p | CACNB4 | 2.39 | 0 | -2.29 | 0 | miRNATAP | -0.26 | 2.0E-5 | NA | |
| 143 | hsa-miR-196b-5p | CACNB4 | 1.86 | 0.00094 | -2.29 | 0 | miRNATAP | -0.11 | 7.0E-5 | NA | |
| 144 | hsa-miR-20a-3p | CACNB4 | 2.52 | 0 | -2.29 | 0 | miRNATAP | -0.22 | 0.00017 | NA | |
| 145 | hsa-miR-212-3p | CACNB4 | 0.4 | 0.10355 | -2.29 | 0 | miRNATAP | -0.35 | 0 | NA | |
| 146 | hsa-miR-222-3p | CACNB4 | 0.03 | 0.88194 | -2.29 | 0 | miRNATAP | -0.21 | 0.00378 | NA | |
| 147 | hsa-miR-29a-5p | CACNB4 | 1.9 | 0 | -2.29 | 0 | mirMAP | -0.27 | 4.0E-5 | NA | |
| 148 | hsa-miR-29b-1-5p | CACNB4 | 1.71 | 0 | -2.29 | 0 | mirMAP | -0.26 | 0 | NA | |
| 149 | hsa-miR-335-3p | CACNB4 | 1.51 | 0 | -2.29 | 0 | mirMAP | -0.23 | 0.00183 | NA | |
| 150 | hsa-miR-424-5p | CACNB4 | 1.26 | 1.0E-5 | -2.29 | 0 | miRNATAP | -0.26 | 0 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | SIGNAL TRANSDUCTION BY PROTEIN PHOSPHORYLATION | 79 | 404 | 1.158e-87 | 5.389e-84 |
| 2 | INTRACELLULAR SIGNAL TRANSDUCTION | 114 | 1572 | 2.151e-82 | 5.005e-79 |
| 3 | PROTEIN PHOSPHORYLATION | 96 | 944 | 1.768e-80 | 2.742e-77 |
| 4 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 121 | 1977 | 2.629e-80 | 3.058e-77 |
| 5 | REGULATION OF MAPK CASCADE | 83 | 660 | 9.311e-76 | 8.665e-73 |
| 6 | PHOSPHORYLATION | 96 | 1228 | 1.656e-69 | 1.284e-66 |
| 7 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 105 | 1656 | 3.472e-68 | 2.308e-65 |
| 8 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 109 | 1929 | 3.117e-66 | 1.813e-63 |
| 9 | REGULATION OF MAP KINASE ACTIVITY | 61 | 319 | 1.113e-65 | 5.753e-63 |
| 10 | POSITIVE REGULATION OF MAPK CASCADE | 67 | 470 | 1.526e-63 | 7.101e-61 |
| 11 | REGULATION OF PROTEIN MODIFICATION PROCESS | 101 | 1710 | 6.322e-62 | 2.674e-59 |
| 12 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 65 | 470 | 1.3e-60 | 5.041e-58 |
| 13 | REGULATION OF KINASE ACTIVITY | 76 | 776 | 2.809e-60 | 1.005e-57 |
| 14 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 79 | 876 | 3.864e-60 | 1.284e-57 |
| 15 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 101 | 1791 | 5.622e-60 | 1.744e-57 |
| 16 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 97 | 1618 | 1.746e-59 | 5.076e-57 |
| 17 | POSITIVE REGULATION OF KINASE ACTIVITY | 64 | 482 | 1.924e-58 | 5.265e-56 |
| 18 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 82 | 1036 | 3.681e-58 | 9.014e-56 |
| 19 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 82 | 1036 | 3.681e-58 | 9.014e-56 |
| 20 | POSITIVE REGULATION OF CELL COMMUNICATION | 93 | 1532 | 5.65e-57 | 1.314e-54 |
| 21 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 90 | 1518 | 7.051e-54 | 1.562e-51 |
| 22 | REGULATION OF TRANSFERASE ACTIVITY | 76 | 946 | 8.206e-54 | 1.71e-51 |
| 23 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 81 | 1135 | 8.453e-54 | 1.71e-51 |
| 24 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 64 | 616 | 1.599e-51 | 3.1e-49 |
| 25 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 85 | 1492 | 6.25e-49 | 1.163e-46 |
| 26 | ACTIVATION OF PROTEIN KINASE ACTIVITY | 46 | 279 | 8.593e-46 | 1.538e-43 |
| 27 | POSITIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 41 | 289 | 6.209e-38 | 1.07e-35 |
| 28 | POSITIVE REGULATION OF MAP KINASE ACTIVITY | 34 | 207 | 1.141e-33 | 1.896e-31 |
| 29 | REGULATION OF CELL DEATH | 69 | 1472 | 4.021e-33 | 6.451e-31 |
| 30 | RESPONSE TO GROWTH FACTOR | 43 | 475 | 2.062e-31 | 3.198e-29 |
| 31 | REGULATION OF RESPONSE TO STRESS | 66 | 1468 | 1.946e-30 | 2.92e-28 |
| 32 | REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 31 | 197 | 4.06e-30 | 5.904e-28 |
| 33 | STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 25 | 103 | 1.906e-29 | 2.687e-27 |
| 34 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 47 | 689 | 6.963e-29 | 9.529e-27 |
| 35 | REGULATION OF IMMUNE SYSTEM PROCESS | 62 | 1403 | 5.477e-28 | 7.282e-26 |
| 36 | REGULATION OF IMMUNE RESPONSE | 50 | 858 | 1.277e-27 | 1.651e-25 |
| 37 | REGULATION OF JNK CASCADE | 26 | 159 | 8.4e-26 | 1.056e-23 |
| 38 | IMMUNE RESPONSE REGULATING CELL SURFACE RECEPTOR SIGNALING PATHWAY | 33 | 323 | 8.913e-26 | 1.091e-23 |
| 39 | FC EPSILON RECEPTOR SIGNALING PATHWAY | 25 | 142 | 1.114e-25 | 1.329e-23 |
| 40 | CELL DEATH | 51 | 1001 | 1.723e-25 | 2.005e-23 |
| 41 | JNK CASCADE | 21 | 82 | 2.048e-25 | 2.324e-23 |
| 42 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 38 | 498 | 5.247e-25 | 5.813e-23 |
| 43 | ACTIVATION OF MAPKK ACTIVITY | 18 | 52 | 1.22e-24 | 1.32e-22 |
| 44 | REGULATION OF TRANSPORT | 65 | 1804 | 1.832e-24 | 1.938e-22 |
| 45 | FC RECEPTOR SIGNALING PATHWAY | 27 | 206 | 4.239e-24 | 4.383e-22 |
| 46 | POSITIVE REGULATION OF CELL DEATH | 40 | 605 | 5.465e-24 | 5.528e-22 |
| 47 | REGULATION OF ERK1 AND ERK2 CASCADE | 28 | 238 | 1.207e-23 | 1.195e-21 |
| 48 | POSITIVE REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 23 | 135 | 2.4e-23 | 2.327e-21 |
| 49 | POSITIVE REGULATION OF HYDROLASE ACTIVITY | 46 | 905 | 7.471e-23 | 7.095e-21 |
| 50 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 63 | 1848 | 2.4e-22 | 2.233e-20 |
| 51 | RESPONSE TO ENDOGENOUS STIMULUS | 56 | 1450 | 3.281e-22 | 2.993e-20 |
| 52 | RESPONSE TO EXTERNAL STIMULUS | 62 | 1821 | 6.334e-22 | 5.668e-20 |
| 53 | NEGATIVE REGULATION OF CELL DEATH | 44 | 872 | 1.04e-21 | 9.126e-20 |
| 54 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 54 | 1381 | 1.302e-21 | 1.122e-19 |
| 55 | RESPONSE TO ABIOTIC STIMULUS | 47 | 1024 | 1.622e-21 | 1.372e-19 |
| 56 | CELL DEVELOPMENT | 54 | 1426 | 5.707e-21 | 4.742e-19 |
| 57 | REGULATION OF CELLULAR RESPONSE TO STRESS | 39 | 691 | 6.365e-21 | 5.196e-19 |
| 58 | REGULATION OF HYDROLASE ACTIVITY | 52 | 1327 | 8.323e-21 | 6.677e-19 |
| 59 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 48 | 1142 | 2.17e-20 | 1.683e-18 |
| 60 | ACTIVATION OF MAPK ACTIVITY | 21 | 137 | 2.15e-20 | 1.683e-18 |
| 61 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 45 | 1008 | 4.225e-20 | 3.223e-18 |
| 62 | POSITIVE REGULATION OF IMMUNE SYSTEM PROCESS | 42 | 867 | 4.791e-20 | 3.595e-18 |
| 63 | REGULATION OF CELL PROLIFERATION | 54 | 1496 | 5.09e-20 | 3.76e-18 |
| 64 | RAS PROTEIN SIGNAL TRANSDUCTION | 21 | 143 | 5.432e-20 | 3.949e-18 |
| 65 | CELLULAR RESPONSE TO STRESS | 55 | 1565 | 7.031e-20 | 5.033e-18 |
| 66 | NEGATIVE REGULATION OF CELL COMMUNICATION | 48 | 1192 | 1.274e-19 | 8.98e-18 |
| 67 | POSITIVE REGULATION OF ERK1 AND ERK2 CASCADE | 22 | 172 | 1.487e-19 | 1.033e-17 |
| 68 | REGULATION OF NEURON DEATH | 25 | 252 | 2.329e-19 | 1.593e-17 |
| 69 | REGULATION OF CELL DIFFERENTIATION | 53 | 1492 | 2.564e-19 | 1.729e-17 |
| 70 | IMMUNE SYSTEM PROCESS | 61 | 1984 | 2.636e-19 | 1.752e-17 |
| 71 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 50 | 1360 | 8.556e-19 | 5.607e-17 |
| 72 | REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 28 | 365 | 1.383e-18 | 8.936e-17 |
| 73 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 57 | 1805 | 1.814e-18 | 1.156e-16 |
| 74 | CELL CELL SIGNALING | 38 | 767 | 1.849e-18 | 1.163e-16 |
| 75 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 50 | 1395 | 2.479e-18 | 1.538e-16 |
| 76 | POSITIVE REGULATION OF IMMUNE RESPONSE | 33 | 563 | 3.149e-18 | 1.928e-16 |
| 77 | SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 27 | 352 | 6.024e-18 | 3.64e-16 |
| 78 | POSITIVE REGULATION OF GENE EXPRESSION | 55 | 1733 | 7.181e-18 | 4.283e-16 |
| 79 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 54 | 1672 | 7.381e-18 | 4.347e-16 |
| 80 | REGULATION OF GTPASE ACTIVITY | 35 | 673 | 1.128e-17 | 6.562e-16 |
| 81 | VASCULATURE DEVELOPMENT | 30 | 469 | 1.168e-17 | 6.707e-16 |
| 82 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 19 | 138 | 1.216e-17 | 6.901e-16 |
| 83 | POSITIVE REGULATION OF CELL PROLIFERATION | 38 | 814 | 1.351e-17 | 7.572e-16 |
| 84 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 42 | 1021 | 1.882e-17 | 1.042e-15 |
| 85 | PHOSPHATIDYLINOSITOL METABOLIC PROCESS | 21 | 193 | 3.083e-17 | 1.687e-15 |
| 86 | NEGATIVE REGULATION OF MAPK CASCADE | 19 | 145 | 3.155e-17 | 1.707e-15 |
| 87 | PEPTIDYL AMINO ACID MODIFICATION | 38 | 841 | 3.981e-17 | 2.129e-15 |
| 88 | PEPTIDYL SERINE MODIFICATION | 19 | 148 | 4.673e-17 | 2.471e-15 |
| 89 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 36 | 771 | 1.103e-16 | 5.766e-15 |
| 90 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 36 | 788 | 2.186e-16 | 1.118e-14 |
| 91 | CIRCULATORY SYSTEM DEVELOPMENT | 36 | 788 | 2.186e-16 | 1.118e-14 |
| 92 | PROTEIN DEPHOSPHORYLATION | 20 | 190 | 3.502e-16 | 1.771e-14 |
| 93 | LOCOMOTION | 42 | 1114 | 4.165e-16 | 2.084e-14 |
| 94 | REGULATION OF NEURON APOPTOTIC PROCESS | 20 | 192 | 4.301e-16 | 2.107e-14 |
| 95 | PROTEIN AUTOPHOSPHORYLATION | 20 | 192 | 4.301e-16 | 2.107e-14 |
| 96 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 30 | 541 | 5.742e-16 | 2.754e-14 |
| 97 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 30 | 541 | 5.742e-16 | 2.754e-14 |
| 98 | INOSITOL LIPID MEDIATED SIGNALING | 17 | 124 | 7.208e-16 | 3.422e-14 |
| 99 | ACTIVATION OF IMMUNE RESPONSE | 27 | 427 | 7.721e-16 | 3.629e-14 |
| 100 | ORGAN MORPHOGENESIS | 36 | 841 | 1.652e-15 | 7.668e-14 |
| 101 | GLYCEROPHOSPHOLIPID METABOLIC PROCESS | 23 | 297 | 1.664e-15 | 7.668e-14 |
| 102 | POSITIVE REGULATION OF NF KAPPAB TRANSCRIPTION FACTOR ACTIVITY | 17 | 132 | 2.113e-15 | 9.639e-14 |
| 103 | NEGATIVE REGULATION OF MAP KINASE ACTIVITY | 14 | 73 | 2.151e-15 | 9.717e-14 |
| 104 | RESPONSE TO NITROGEN COMPOUND | 36 | 859 | 3.17e-15 | 1.418e-13 |
| 105 | POSITIVE REGULATION OF LOCOMOTION | 26 | 420 | 4.607e-15 | 2.041e-13 |
| 106 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 35 | 823 | 5.291e-15 | 2.323e-13 |
| 107 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 41 | 1152 | 7.166e-15 | 3.116e-13 |
| 108 | LIPID PHOSPHORYLATION | 15 | 99 | 8.364e-15 | 3.603e-13 |
| 109 | TISSUE DEVELOPMENT | 47 | 1518 | 9.262e-15 | 3.954e-13 |
| 110 | REGULATION OF CELLULAR LOCALIZATION | 43 | 1277 | 9.471e-15 | 4.005e-13 |
| 111 | INACTIVATION OF MAPK ACTIVITY | 10 | 26 | 9.555e-15 | 4.005e-13 |
| 112 | NEUROGENESIS | 45 | 1402 | 1.102e-14 | 4.577e-13 |
| 113 | POSITIVE REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 20 | 228 | 1.208e-14 | 4.975e-13 |
| 114 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 30 | 616 | 1.819e-14 | 7.423e-13 |
| 115 | MODULATION OF SYNAPTIC TRANSMISSION | 22 | 301 | 2.355e-14 | 9.53e-13 |
| 116 | NEURON PROJECTION DEVELOPMENT | 28 | 545 | 4.016e-14 | 1.611e-12 |
| 117 | POSITIVE REGULATION OF TRANSPORT | 36 | 936 | 4.306e-14 | 1.713e-12 |
| 118 | NEGATIVE REGULATION OF PHOSPHORYLATION | 25 | 422 | 4.4e-14 | 1.735e-12 |
| 119 | PEPTIDYL TYROSINE MODIFICATION | 18 | 186 | 4.966e-14 | 1.942e-12 |
| 120 | TUBE DEVELOPMENT | 28 | 552 | 5.506e-14 | 2.135e-12 |
| 121 | CELL PROJECTION ORGANIZATION | 35 | 902 | 7.976e-14 | 3.067e-12 |
| 122 | GLYCEROLIPID METABOLIC PROCESS | 23 | 356 | 8.055e-14 | 3.072e-12 |
| 123 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 33 | 799 | 8.132e-14 | 3.076e-12 |
| 124 | RESPONSE TO WOUNDING | 28 | 563 | 8.954e-14 | 3.36e-12 |
| 125 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 25 | 437 | 9.679e-14 | 3.603e-12 |
| 126 | REGULATION OF CELL ACTIVATION | 26 | 484 | 1.281e-13 | 4.713e-12 |
| 127 | PHOSPHOLIPID METABOLIC PROCESS | 23 | 364 | 1.286e-13 | 4.713e-12 |
| 128 | MEMBRANE DEPOLARIZATION | 12 | 61 | 1.653e-13 | 5.973e-12 |
| 129 | NEGATIVE REGULATION OF NEURON DEATH | 17 | 171 | 1.656e-13 | 5.973e-12 |
| 130 | NEURON DIFFERENTIATION | 34 | 874 | 1.785e-13 | 6.389e-12 |
| 131 | PEPTIDYL TYROSINE DEPHOSPHORYLATION | 14 | 100 | 2.096e-13 | 7.446e-12 |
| 132 | REGULATION OF ION TRANSPORT | 28 | 592 | 3.06e-13 | 1.079e-11 |
| 133 | NEURON DEVELOPMENT | 30 | 687 | 3.096e-13 | 1.083e-11 |
| 134 | NEGATIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 15 | 126 | 3.26e-13 | 1.132e-11 |
| 135 | TAXIS | 25 | 464 | 3.702e-13 | 1.271e-11 |
| 136 | REGULATION OF ORGANELLE ORGANIZATION | 39 | 1178 | 3.715e-13 | 1.271e-11 |
| 137 | FIBROBLAST GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 13 | 84 | 4.101e-13 | 1.393e-11 |
| 138 | REGULATION OF TRANSMEMBRANE TRANSPORT | 24 | 426 | 4.386e-13 | 1.479e-11 |
| 139 | REGULATION OF SECRETION | 30 | 699 | 4.825e-13 | 1.615e-11 |
| 140 | WOUND HEALING | 25 | 470 | 4.926e-13 | 1.637e-11 |
| 141 | RESPONSE TO DRUG | 24 | 431 | 5.637e-13 | 1.86e-11 |
| 142 | CALCIUM ION TRANSMEMBRANE TRANSPORT | 16 | 159 | 7.295e-13 | 2.39e-11 |
| 143 | NEGATIVE REGULATION OF KINASE ACTIVITY | 19 | 250 | 7.648e-13 | 2.488e-11 |
| 144 | RESPONSE TO MOLECULE OF BACTERIAL ORIGIN | 21 | 321 | 8.225e-13 | 2.658e-11 |
| 145 | DEPHOSPHORYLATION | 20 | 286 | 8.71e-13 | 2.795e-11 |
| 146 | NEGATIVE REGULATION OF NEURON APOPTOTIC PROCESS | 15 | 135 | 9.104e-13 | 2.901e-11 |
| 147 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 22 | 363 | 1.05e-12 | 3.323e-11 |
| 148 | BLOOD VESSEL MORPHOGENESIS | 22 | 364 | 1.109e-12 | 3.463e-11 |
| 149 | POSITIVE REGULATION OF DEFENSE RESPONSE | 22 | 364 | 1.109e-12 | 3.463e-11 |
| 150 | CALCIUM ION TRANSPORT | 18 | 223 | 1.139e-12 | 3.534e-11 |
| 151 | REGULATION OF CELL ADHESION | 28 | 629 | 1.326e-12 | 4.086e-11 |
| 152 | EPITHELIUM DEVELOPMENT | 34 | 945 | 1.607e-12 | 4.92e-11 |
| 153 | RESPONSE TO FIBROBLAST GROWTH FACTOR | 14 | 116 | 1.702e-12 | 5.176e-11 |
| 154 | RESPONSE TO RADIATION | 23 | 413 | 1.78e-12 | 5.379e-11 |
| 155 | REGULATION OF PROTEIN LOCALIZATION | 34 | 950 | 1.862e-12 | 5.588e-11 |
| 156 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 34 | 957 | 2.283e-12 | 6.808e-11 |
| 157 | ACTIVATION OF INNATE IMMUNE RESPONSE | 17 | 204 | 2.974e-12 | 8.815e-11 |
| 158 | SINGLE ORGANISM BEHAVIOR | 22 | 384 | 3.209e-12 | 9.449e-11 |
| 159 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 36 | 1087 | 3.54e-12 | 1.036e-10 |
| 160 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 33 | 917 | 3.587e-12 | 1.043e-10 |
| 161 | MULTICELLULAR ORGANISMAL SIGNALING | 14 | 123 | 3.848e-12 | 1.112e-10 |
| 162 | REGULATION OF DEFENSE RESPONSE | 30 | 759 | 3.885e-12 | 1.116e-10 |
| 163 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 39 | 1275 | 4.318e-12 | 1.233e-10 |
| 164 | RESPONSE TO INORGANIC SUBSTANCE | 24 | 479 | 5.326e-12 | 1.511e-10 |
| 165 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 15 | 153 | 5.731e-12 | 1.616e-10 |
| 166 | REGULATION OF JUN KINASE ACTIVITY | 12 | 81 | 5.77e-12 | 1.617e-10 |
| 167 | POSITIVE REGULATION OF INNATE IMMUNE RESPONSE | 18 | 246 | 6.001e-12 | 1.672e-10 |
| 168 | REGULATION OF INNATE IMMUNE RESPONSE | 21 | 357 | 6.278e-12 | 1.739e-10 |
| 169 | CARDIAC CONDUCTION | 12 | 82 | 6.708e-12 | 1.847e-10 |
| 170 | RESPONSE TO LIPID | 32 | 888 | 7.743e-12 | 2.119e-10 |
| 171 | REGULATION OF PHOSPHOLIPASE ACTIVITY | 11 | 64 | 8.348e-12 | 2.272e-10 |
| 172 | IMMUNE SYSTEM DEVELOPMENT | 26 | 582 | 8.464e-12 | 2.29e-10 |
| 173 | EMBRYO DEVELOPMENT | 32 | 894 | 9.233e-12 | 2.483e-10 |
| 174 | APOPTOTIC SIGNALING PATHWAY | 19 | 289 | 9.881e-12 | 2.642e-10 |
| 175 | POSITIVE REGULATION OF CELL DIVISION | 14 | 132 | 1.02e-11 | 2.712e-10 |
| 176 | PHOSPHATIDYLINOSITOL 3 PHOSPHATE BIOSYNTHETIC PROCESS | 10 | 49 | 1.243e-11 | 3.287e-10 |
| 177 | PHOSPHATIDYLETHANOLAMINE ACYL CHAIN REMODELING | 8 | 23 | 1.283e-11 | 3.372e-10 |
| 178 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 35 | 1079 | 1.335e-11 | 3.49e-10 |
| 179 | HEART DEVELOPMENT | 23 | 466 | 2.075e-11 | 5.394e-10 |
| 180 | DIVALENT INORGANIC CATION TRANSPORT | 18 | 268 | 2.508e-11 | 6.482e-10 |
| 181 | REGULATION OF HOMOTYPIC CELL CELL ADHESION | 19 | 307 | 2.821e-11 | 7.253e-10 |
| 182 | POSITIVE REGULATION OF PHOSPHOLIPASE ACTIVITY | 10 | 53 | 2.862e-11 | 7.285e-10 |
| 183 | POSITIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 15 | 171 | 2.865e-11 | 7.285e-10 |
| 184 | REGULATION OF PROTEIN SECRETION | 21 | 389 | 3.154e-11 | 7.975e-10 |
| 185 | RESPONSE TO TRANSFORMING GROWTH FACTOR BETA | 14 | 144 | 3.35e-11 | 8.425e-10 |
| 186 | REGULATION OF INTRACELLULAR TRANSPORT | 26 | 621 | 3.556e-11 | 8.895e-10 |
| 187 | PHOSPHATIDYLCHOLINE ACYL CHAIN REMODELING | 8 | 26 | 3.997e-11 | 9.944e-10 |
| 188 | POSITIVE REGULATION OF SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 9 | 39 | 4.135e-11 | 1.023e-09 |
| 189 | RESPONSE TO MECHANICAL STIMULUS | 16 | 210 | 5.172e-11 | 1.273e-09 |
| 190 | MORPHOGENESIS OF AN EPITHELIUM | 21 | 400 | 5.298e-11 | 1.297e-09 |
| 191 | GLYCEROLIPID BIOSYNTHETIC PROCESS | 16 | 211 | 5.554e-11 | 1.353e-09 |
| 192 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 18 | 285 | 6.918e-11 | 1.676e-09 |
| 193 | COGNITION | 17 | 251 | 8.081e-11 | 1.948e-09 |
| 194 | REGULATION OF RAS PROTEIN SIGNAL TRANSDUCTION | 15 | 184 | 8.162e-11 | 1.958e-09 |
| 195 | POSITIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 14 | 154 | 8.306e-11 | 1.982e-09 |
| 196 | CELL MORPHOGENESIS INVOLVED IN NEURON DIFFERENTIATION | 20 | 368 | 8.516e-11 | 2.022e-09 |
| 197 | POSITIVE REGULATION OF LIPID METABOLIC PROCESS | 13 | 128 | 9.948e-11 | 2.35e-09 |
| 198 | ANGIOGENESIS | 18 | 293 | 1.089e-10 | 2.558e-09 |
| 199 | HEAD DEVELOPMENT | 27 | 709 | 1.213e-10 | 2.837e-09 |
| 200 | REGULATION OF CELLULAR COMPONENT BIOGENESIS | 28 | 767 | 1.394e-10 | 3.243e-09 |
| 201 | INNATE IMMUNE RESPONSE ACTIVATING CELL SURFACE RECEPTOR SIGNALING PATHWAY | 12 | 106 | 1.486e-10 | 3.441e-09 |
| 202 | REGULATION OF CELL CELL ADHESION | 20 | 380 | 1.504e-10 | 3.464e-09 |
| 203 | REGULATION OF LIPASE ACTIVITY | 11 | 83 | 1.57e-10 | 3.592e-09 |
| 204 | BEHAVIOR | 23 | 516 | 1.575e-10 | 3.592e-09 |
| 205 | POSITIVE REGULATION OF JUN KINASE ACTIVITY | 10 | 63 | 1.74e-10 | 3.95e-09 |
| 206 | CELL ACTIVATION | 24 | 568 | 1.806e-10 | 4.079e-09 |
| 207 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 32 | 1004 | 1.817e-10 | 4.084e-09 |
| 208 | CELL MOTILITY | 29 | 835 | 2.023e-10 | 4.504e-09 |
| 209 | LOCALIZATION OF CELL | 29 | 835 | 2.023e-10 | 4.504e-09 |
| 210 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 9 | 46 | 2.04e-10 | 4.52e-09 |
| 211 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 24 | 573 | 2.157e-10 | 4.757e-09 |
| 212 | REGULATION OF VASCULATURE DEVELOPMENT | 16 | 233 | 2.433e-10 | 5.34e-09 |
| 213 | RESPONSE TO BACTERIUM | 23 | 528 | 2.472e-10 | 5.401e-09 |
| 214 | PHOSPHOLIPID BIOSYNTHETIC PROCESS | 16 | 235 | 2.76e-10 | 6.002e-09 |
| 215 | NEGATIVE REGULATION OF TRANSFERASE ACTIVITY | 19 | 351 | 2.778e-10 | 6.012e-09 |
| 216 | POSITIVE REGULATION OF LIPASE ACTIVITY | 10 | 66 | 2.808e-10 | 6.05e-09 |
| 217 | REGULATION OF CELL DIVISION | 17 | 272 | 2.824e-10 | 6.056e-09 |
| 218 | GLAND DEVELOPMENT | 20 | 395 | 2.973e-10 | 6.346e-09 |
| 219 | REGULATION OF SYNAPTIC PLASTICITY | 13 | 140 | 3.079e-10 | 6.543e-09 |
| 220 | REGULATION OF SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 17 | 278 | 3.956e-10 | 8.367e-09 |
| 221 | NEURON PROJECTION MORPHOGENESIS | 20 | 402 | 4.044e-10 | 8.513e-09 |
| 222 | RESPONSE TO LIGHT STIMULUS | 17 | 280 | 4.418e-10 | 9.259e-09 |
| 223 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 31 | 983 | 4.739e-10 | 9.887e-09 |
| 224 | REGULATION OF RESPONSE TO EXTERNAL STIMULUS | 30 | 926 | 4.991e-10 | 1.037e-08 |
| 225 | I KAPPAB KINASE NF KAPPAB SIGNALING | 10 | 70 | 5.124e-10 | 1.06e-08 |
| 226 | LIPID MODIFICATION | 15 | 210 | 5.245e-10 | 1.08e-08 |
| 227 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 29 | 872 | 5.54e-10 | 1.136e-08 |
| 228 | POSITIVE REGULATION OF I KAPPAB KINASE NF KAPPAB SIGNALING | 14 | 179 | 6.164e-10 | 1.258e-08 |
| 229 | CELLULAR RESPONSE TO NITROGEN COMPOUND | 22 | 505 | 6.26e-10 | 1.272e-08 |
| 230 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 11 | 95 | 6.938e-10 | 1.404e-08 |
| 231 | REGULATION OF ORGAN GROWTH | 10 | 73 | 7.846e-10 | 1.58e-08 |
| 232 | NEGATIVE REGULATION OF CATALYTIC ACTIVITY | 28 | 829 | 8.093e-10 | 1.623e-08 |
| 233 | REGULATION OF PROTEIN IMPORT | 14 | 183 | 8.245e-10 | 1.647e-08 |
| 234 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 22 | 513 | 8.394e-10 | 1.669e-08 |
| 235 | POSITIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 22 | 514 | 8.704e-10 | 1.723e-08 |
| 236 | REGULATION OF CELL CYCLE | 30 | 949 | 8.909e-10 | 1.757e-08 |
| 237 | REGULATION OF CELL DEVELOPMENT | 28 | 836 | 9.765e-10 | 1.917e-08 |
| 238 | REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 18 | 337 | 1.041e-09 | 2.036e-08 |
| 239 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 11 | 99 | 1.087e-09 | 2.116e-08 |
| 240 | CELLULAR COMPONENT MORPHOGENESIS | 29 | 900 | 1.145e-09 | 2.22e-08 |
| 241 | MEMBRANE DEPOLARIZATION DURING ACTION POTENTIAL | 8 | 39 | 1.429e-09 | 2.758e-08 |
| 242 | REGULATION OF CYTOPLASMIC TRANSPORT | 21 | 481 | 1.522e-09 | 2.927e-08 |
| 243 | ANTIGEN RECEPTOR MEDIATED SIGNALING PATHWAY | 14 | 195 | 1.892e-09 | 3.623e-08 |
| 244 | CELLULAR RESPONSE TO MECHANICAL STIMULUS | 10 | 80 | 1.973e-09 | 3.762e-08 |
| 245 | REGULATION OF SYNAPSE STRUCTURE OR ACTIVITY | 15 | 232 | 2.08e-09 | 3.951e-08 |
| 246 | POSITIVE REGULATION OF VASCULATURE DEVELOPMENT | 12 | 133 | 2.113e-09 | 3.996e-08 |
| 247 | REGULATION OF I KAPPAB KINASE NF KAPPAB SIGNALING | 15 | 233 | 2.207e-09 | 4.157e-08 |
| 248 | ACTIN FILAMENT BASED PROCESS | 20 | 450 | 2.836e-09 | 5.321e-08 |
| 249 | PHOSPHATIDYLINOSITOL ACYL CHAIN REMODELING | 6 | 16 | 3.067e-09 | 5.731e-08 |
| 250 | EMBRYONIC ORGAN DEVELOPMENT | 19 | 406 | 3.126e-09 | 5.819e-08 |
| 251 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 22 | 554 | 3.471e-09 | 6.434e-08 |
| 252 | RESPONSE TO BIOTIC STIMULUS | 28 | 886 | 3.528e-09 | 6.488e-08 |
| 253 | RESPONSE TO CYTOKINE | 25 | 714 | 3.519e-09 | 6.488e-08 |
| 254 | POSITIVE REGULATION OF DNA REPLICATION | 10 | 86 | 4.055e-09 | 7.399e-08 |
| 255 | POSITIVE REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 10 | 86 | 4.055e-09 | 7.399e-08 |
| 256 | RESPONSE TO HORMONE | 28 | 893 | 4.191e-09 | 7.617e-08 |
| 257 | REGULATION OF VESICLE MEDIATED TRANSPORT | 20 | 462 | 4.437e-09 | 8.033e-08 |
| 258 | PHOSPHATIDYLCHOLINE METABOLIC PROCESS | 9 | 64 | 4.456e-09 | 8.037e-08 |
| 259 | REGULATION OF CYTOKINE PRODUCTION | 22 | 563 | 4.658e-09 | 8.335e-08 |
| 260 | REGULATION OF CALCIUM ION TRANSPORT | 14 | 209 | 4.642e-09 | 8.335e-08 |
| 261 | PHOSPHATIDYLSERINE ACYL CHAIN REMODELING | 6 | 17 | 4.706e-09 | 8.357e-08 |
| 262 | PHOSPHATIDYLGLYCEROL ACYL CHAIN REMODELING | 6 | 17 | 4.706e-09 | 8.357e-08 |
| 263 | PEPTIDYL THREONINE MODIFICATION | 8 | 46 | 5.752e-09 | 1.018e-07 |
| 264 | POSITIVE REGULATION OF CELL ADHESION | 18 | 376 | 5.85e-09 | 1.031e-07 |
| 265 | POSITIVE REGULATION OF CELL DEVELOPMENT | 20 | 472 | 6.373e-09 | 1.119e-07 |
| 266 | PHOSPHATIDYLGLYCEROL METABOLIC PROCESS | 7 | 31 | 7.67e-09 | 1.342e-07 |
| 267 | REGULATION OF GROWTH | 23 | 633 | 7.95e-09 | 1.379e-07 |
| 268 | REGULATION OF PEPTIDE TRANSPORT | 15 | 256 | 7.941e-09 | 1.379e-07 |
| 269 | REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 14 | 218 | 7.974e-09 | 1.379e-07 |
| 270 | RHYTHMIC PROCESS | 16 | 298 | 8.582e-09 | 1.479e-07 |
| 271 | TISSUE MORPHOGENESIS | 21 | 533 | 9.3e-09 | 1.595e-07 |
| 272 | POSITIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 11 | 121 | 9.324e-09 | 1.595e-07 |
| 273 | POSITIVE REGULATION OF DNA METABOLIC PROCESS | 13 | 185 | 9.472e-09 | 1.612e-07 |
| 274 | REGULATION OF MEMBRANE POTENTIAL | 17 | 343 | 9.492e-09 | 1.612e-07 |
| 275 | SALIVARY GLAND DEVELOPMENT | 7 | 32 | 9.745e-09 | 1.638e-07 |
| 276 | REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 12 | 152 | 9.703e-09 | 1.638e-07 |
| 277 | ACTION POTENTIAL | 10 | 94 | 9.749e-09 | 1.638e-07 |
| 278 | EMBRYONIC MORPHOGENESIS | 21 | 539 | 1.13e-08 | 1.891e-07 |
| 279 | RESPONSE TO OXIDATIVE STRESS | 17 | 352 | 1.392e-08 | 2.321e-07 |
| 280 | ORGANOPHOSPHATE METABOLIC PROCESS | 27 | 885 | 1.458e-08 | 2.423e-07 |
| 281 | CELLULAR RESPONSE TO CARBOHYDRATE STIMULUS | 9 | 74 | 1.659e-08 | 2.748e-07 |
| 282 | REGULATION OF CELL MORPHOGENESIS | 21 | 552 | 1.706e-08 | 2.816e-07 |
| 283 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 11 | 129 | 1.829e-08 | 3.007e-07 |
| 284 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 12 | 162 | 1.988e-08 | 3.257e-07 |
| 285 | RESPIRATORY SYSTEM DEVELOPMENT | 13 | 197 | 2.012e-08 | 3.284e-07 |
| 286 | REGULATION OF TRANSPORTER ACTIVITY | 13 | 198 | 2.137e-08 | 3.476e-07 |
| 287 | ACTIVATION OF MAPKKK ACTIVITY | 5 | 11 | 2.145e-08 | 3.477e-07 |
| 288 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 40 | 1784 | 2.155e-08 | 3.481e-07 |
| 289 | CELL PROLIFERATION | 23 | 672 | 2.406e-08 | 3.874e-07 |
| 290 | POSITIVE REGULATION OF GROWTH | 14 | 238 | 2.43e-08 | 3.898e-07 |
| 291 | REGULATION OF EPITHELIAL CELL MIGRATION | 12 | 166 | 2.611e-08 | 4.161e-07 |
| 292 | POSITIVE REGULATION OF PROTEIN IMPORT | 10 | 104 | 2.611e-08 | 4.161e-07 |
| 293 | TUBE MORPHOGENESIS | 16 | 323 | 2.663e-08 | 4.229e-07 |
| 294 | MORPHOGENESIS OF A BRANCHING STRUCTURE | 12 | 167 | 2.792e-08 | 4.419e-07 |
| 295 | REGULATION OF LIPID METABOLIC PROCESS | 15 | 282 | 2.895e-08 | 4.566e-07 |
| 296 | ERBB SIGNALING PATHWAY | 9 | 79 | 2.977e-08 | 4.679e-07 |
| 297 | NEUROTROPHIN SIGNALING PATHWAY | 6 | 23 | 3.675e-08 | 5.757e-07 |
| 298 | CELL PART MORPHOGENESIS | 22 | 633 | 3.806e-08 | 5.943e-07 |
| 299 | RESPONSE TO METAL ION | 16 | 333 | 4.068e-08 | 6.31e-07 |
| 300 | REGULATION OF PEPTIDE SECRETION | 13 | 209 | 4.058e-08 | 6.31e-07 |
| 301 | REGULATION OF PHOSPHOLIPASE C ACTIVITY | 7 | 39 | 4.229e-08 | 6.516e-07 |
| 302 | ERBB2 SIGNALING PATHWAY | 7 | 39 | 4.229e-08 | 6.516e-07 |
| 303 | NEGATIVE REGULATION OF CELL PROLIFERATION | 22 | 643 | 5.015e-08 | 7.701e-07 |
| 304 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 13 | 213 | 5.074e-08 | 7.766e-07 |
| 305 | POSITIVE REGULATION OF RESPONSE TO EXTERNAL STIMULUS | 15 | 296 | 5.486e-08 | 8.369e-07 |
| 306 | ETHANOLAMINE CONTAINING COMPOUND METABOLIC PROCESS | 9 | 85 | 5.693e-08 | 8.657e-07 |
| 307 | REGULATION OF VOLTAGE GATED CALCIUM CHANNEL ACTIVITY | 6 | 25 | 6.354e-08 | 9.63e-07 |
| 308 | REGULATION OF CHEMOTAXIS | 12 | 180 | 6.412e-08 | 9.687e-07 |
| 309 | T CELL RECEPTOR SIGNALING PATHWAY | 11 | 146 | 6.618e-08 | 9.965e-07 |
| 310 | SENSORY ORGAN DEVELOPMENT | 19 | 493 | 7.014e-08 | 1.053e-06 |
| 311 | REGULATION OF HEART GROWTH | 7 | 42 | 7.256e-08 | 1.086e-06 |
| 312 | REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 13 | 220 | 7.416e-08 | 1.106e-06 |
| 313 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 15 | 303 | 7.451e-08 | 1.108e-06 |
| 314 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 21 | 602 | 7.481e-08 | 1.109e-06 |
| 315 | RESPONSE TO TEMPERATURE STIMULUS | 11 | 148 | 7.612e-08 | 1.124e-06 |
| 316 | REGULATION OF HEART CONTRACTION | 13 | 221 | 7.82e-08 | 1.152e-06 |
| 317 | CELLULAR RESPONSE TO ABIOTIC STIMULUS | 14 | 263 | 8.455e-08 | 1.241e-06 |
| 318 | RESPONSE TO HEAT | 9 | 89 | 8.53e-08 | 1.248e-06 |
| 319 | REGULATION OF PROTEIN TARGETING | 15 | 307 | 8.841e-08 | 1.29e-06 |
| 320 | MEMBRANE DEPOLARIZATION DURING CARDIAC MUSCLE CELL ACTION POTENTIAL | 5 | 14 | 9.097e-08 | 1.323e-06 |
| 321 | REGULATION OF CYTOSKELETON ORGANIZATION | 19 | 502 | 9.301e-08 | 1.348e-06 |
| 322 | REGULATION OF CELL PROJECTION ORGANIZATION | 20 | 558 | 1.012e-07 | 1.462e-06 |
| 323 | PHOSPHATIDYLINOSITOL BIOSYNTHETIC PROCESS | 10 | 120 | 1.029e-07 | 1.478e-06 |
| 324 | POSITIVE REGULATION OF CHEMOTAXIS | 10 | 120 | 1.029e-07 | 1.478e-06 |
| 325 | IN UTERO EMBRYONIC DEVELOPMENT | 15 | 311 | 1.046e-07 | 1.489e-06 |
| 326 | POSITIVE REGULATION OF CELL ACTIVATION | 15 | 311 | 1.046e-07 | 1.489e-06 |
| 327 | POSITIVE REGULATION OF MESENCHYMAL CELL PROLIFERATION | 6 | 27 | 1.047e-07 | 1.489e-06 |
| 328 | REGULATION OF SYSTEM PROCESS | 19 | 507 | 1.085e-07 | 1.539e-06 |
| 329 | POSITIVE REGULATION OF CELLULAR COMPONENT BIOGENESIS | 17 | 406 | 1.105e-07 | 1.562e-06 |
| 330 | REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 13 | 229 | 1.184e-07 | 1.669e-06 |
| 331 | POSITIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 16 | 360 | 1.187e-07 | 1.669e-06 |
| 332 | EXOCRINE SYSTEM DEVELOPMENT | 7 | 45 | 1.194e-07 | 1.673e-06 |
| 333 | POSITIVE REGULATION OF NEURON DEATH | 8 | 67 | 1.23e-07 | 1.713e-06 |
| 334 | RESPONSE TO REACTIVE OXYGEN SPECIES | 12 | 191 | 1.229e-07 | 1.713e-06 |
| 335 | NECROTIC CELL DEATH | 6 | 28 | 1.323e-07 | 1.831e-06 |
| 336 | PHOSPHATIDYLSERINE METABOLIC PROCESS | 6 | 28 | 1.323e-07 | 1.831e-06 |
| 337 | REGULATION OF TRANSCRIPTION FACTOR IMPORT INTO NUCLEUS | 9 | 95 | 1.508e-07 | 2.082e-06 |
| 338 | REGULATION OF CARDIAC MUSCLE CELL PROLIFERATION | 6 | 29 | 1.655e-07 | 2.279e-06 |
| 339 | POSITIVE REGULATION OF INTRACELLULAR TRANSPORT | 16 | 370 | 1.723e-07 | 2.357e-06 |
| 340 | POSITIVE REGULATION OF CYTOKINE PRODUCTION | 16 | 370 | 1.723e-07 | 2.357e-06 |
| 341 | GLAND MORPHOGENESIS | 9 | 97 | 1.807e-07 | 2.465e-06 |
| 342 | REGULATION OF METAL ION TRANSPORT | 15 | 325 | 1.85e-07 | 2.516e-06 |
| 343 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 9 | 98 | 1.974e-07 | 2.678e-06 |
| 344 | SYNAPTIC SIGNALING | 17 | 424 | 2.046e-07 | 2.759e-06 |
| 345 | REGULATION OF IMMUNE EFFECTOR PROCESS | 17 | 424 | 2.046e-07 | 2.759e-06 |
| 346 | REGULATION OF BINDING | 14 | 283 | 2.08e-07 | 2.797e-06 |
| 347 | POSITIVE REGULATION OF CELL CELL ADHESION | 13 | 243 | 2.351e-07 | 3.152e-06 |
| 348 | NEURON PROJECTION GUIDANCE | 12 | 205 | 2.649e-07 | 3.541e-06 |
| 349 | REGULATION OF DEVELOPMENTAL GROWTH | 14 | 289 | 2.685e-07 | 3.579e-06 |
| 350 | RESPONSE TO CARBOHYDRATE | 11 | 168 | 2.765e-07 | 3.676e-06 |
| 351 | REPRODUCTION | 31 | 1297 | 2.877e-07 | 3.813e-06 |
| 352 | POSITIVE REGULATION OF EPITHELIAL CELL MIGRATION | 9 | 103 | 3.033e-07 | 4.01e-06 |
| 353 | POSITIVE REGULATION OF LIPID KINASE ACTIVITY | 6 | 32 | 3.09e-07 | 4.073e-06 |
| 354 | CELLULAR MACROMOLECULE LOCALIZATION | 30 | 1234 | 3.222e-07 | 4.235e-06 |
| 355 | REGULATION OF BLOOD CIRCULATION | 14 | 295 | 3.443e-07 | 4.513e-06 |
| 356 | INOSITOL PHOSPHATE MEDIATED SIGNALING | 5 | 18 | 3.784e-07 | 4.946e-06 |
| 357 | POSITIVE REGULATION OF FIBROBLAST PROLIFERATION | 7 | 53 | 3.824e-07 | 4.97e-06 |
| 358 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 7 | 53 | 3.824e-07 | 4.97e-06 |
| 359 | REGULATION OF NEURON DIFFERENTIATION | 19 | 554 | 4.225e-07 | 5.476e-06 |
| 360 | REGULATION OF MESENCHYMAL CELL PROLIFERATION | 6 | 34 | 4.52e-07 | 5.842e-06 |
| 361 | POSITIVE REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 13 | 258 | 4.661e-07 | 6.008e-06 |
| 362 | PATTERN RECOGNITION RECEPTOR SIGNALING PATHWAY | 9 | 109 | 4.927e-07 | 6.333e-06 |
| 363 | EPIDERMAL GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 7 | 55 | 4.96e-07 | 6.358e-06 |
| 364 | PLATELET ACTIVATION | 10 | 142 | 4.994e-07 | 6.383e-06 |
| 365 | POSITIVE REGULATION OF CARDIAC MUSCLE CELL PROLIFERATION | 5 | 19 | 5.1e-07 | 6.501e-06 |
| 366 | ALDITOL PHOSPHATE METABOLIC PROCESS | 6 | 35 | 5.416e-07 | 6.885e-06 |
| 367 | REGULATION OF FIBROBLAST PROLIFERATION | 8 | 81 | 5.466e-07 | 6.931e-06 |
| 368 | REGULATION OF HORMONE SECRETION | 13 | 262 | 5.55e-07 | 7.017e-06 |
| 369 | NUCLEAR TRANSPORT | 15 | 355 | 5.7e-07 | 7.188e-06 |
| 370 | SYSTEM PROCESS | 37 | 1785 | 5.864e-07 | 7.375e-06 |
| 371 | CELLULAR RESPONSE TO EXTERNAL STIMULUS | 13 | 264 | 6.049e-07 | 7.586e-06 |
| 372 | REPRODUCTIVE SYSTEM DEVELOPMENT | 16 | 408 | 6.371e-07 | 7.969e-06 |
| 373 | NEUROMUSCULAR JUNCTION DEVELOPMENT | 6 | 36 | 6.452e-07 | 8.049e-06 |
| 374 | EMBRYONIC PLACENTA DEVELOPMENT | 8 | 83 | 6.605e-07 | 8.217e-06 |
| 375 | REGULATION OF ACTIN FILAMENT BASED PROCESS | 14 | 312 | 6.75e-07 | 8.376e-06 |
| 376 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 22 | 750 | 6.998e-07 | 8.66e-06 |
| 377 | CARDIAC MUSCLE CELL ACTION POTENTIAL | 6 | 37 | 7.645e-07 | 9.436e-06 |
| 378 | TOLL LIKE RECEPTOR SIGNALING PATHWAY | 8 | 85 | 7.94e-07 | 9.773e-06 |
| 379 | RESPONSE TO ACID CHEMICAL | 14 | 319 | 8.792e-07 | 1.079e-05 |
| 380 | POSITIVE REGULATION OF ORGAN GROWTH | 6 | 38 | 9.013e-07 | 1.104e-05 |
| 381 | POSITIVE REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 7 | 60 | 9.1e-07 | 1.111e-05 |
| 382 | TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 11 | 190 | 9.434e-07 | 1.149e-05 |
| 383 | POSITIVE REGULATION OF SECRETION | 15 | 370 | 9.584e-07 | 1.164e-05 |
| 384 | REGULATION OF LEUKOCYTE DIFFERENTIATION | 12 | 232 | 9.912e-07 | 1.201e-05 |
| 385 | REGULATION OF PHOSPHOLIPID METABOLIC PROCESS | 7 | 61 | 1.02e-06 | 1.233e-05 |
| 386 | POSITIVE REGULATION OF PEPTIDYL SERINE PHOSPHORYLATION | 8 | 88 | 1.037e-06 | 1.247e-05 |
| 387 | REGULATION OF CATION CHANNEL ACTIVITY | 8 | 88 | 1.037e-06 | 1.247e-05 |
| 388 | POSITIVE REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 7 | 62 | 1.142e-06 | 1.369e-05 |
| 389 | CONNECTIVE TISSUE DEVELOPMENT | 11 | 194 | 1.158e-06 | 1.386e-05 |
| 390 | NEUROLOGICAL SYSTEM PROCESS | 29 | 1242 | 1.186e-06 | 1.412e-05 |
| 391 | POSITIVE REGULATION OF ION TRANSPORT | 12 | 236 | 1.187e-06 | 1.412e-05 |
| 392 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 6 | 40 | 1.235e-06 | 1.462e-05 |
| 393 | CALCIUM MEDIATED SIGNALING | 8 | 90 | 1.232e-06 | 1.462e-05 |
| 394 | POSITIVE REGULATION OF CYTOPLASMIC TRANSPORT | 13 | 282 | 1.269e-06 | 1.498e-05 |
| 395 | REGULATION OF ADAPTIVE IMMUNE RESPONSE | 9 | 123 | 1.37e-06 | 1.613e-05 |
| 396 | REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 15 | 381 | 1.38e-06 | 1.622e-05 |
| 397 | CELLULAR LIPID METABOLIC PROCESS | 24 | 913 | 1.471e-06 | 1.724e-05 |
| 398 | REGULATION OF DNA REPLICATION | 10 | 161 | 1.581e-06 | 1.848e-05 |
| 399 | POSITIVE REGULATION OF PHOSPHOLIPID METABOLIC PROCESS | 6 | 42 | 1.664e-06 | 1.94e-05 |
| 400 | HOMEOSTATIC PROCESS | 30 | 1337 | 1.712e-06 | 1.991e-05 |
| 401 | ACID SECRETION | 7 | 66 | 1.756e-06 | 2.037e-05 |
| 402 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 18 | 552 | 1.777e-06 | 2.057e-05 |
| 403 | POSITIVE REGULATION OF NFAT PROTEIN IMPORT INTO NUCLEUS | 4 | 11 | 1.782e-06 | 2.057e-05 |
| 404 | RESPONSE TO STEROID HORMONE | 17 | 497 | 1.841e-06 | 2.12e-05 |
| 405 | REGULATION OF DNA METABOLIC PROCESS | 14 | 340 | 1.865e-06 | 2.142e-05 |
| 406 | REGULATION OF LIPID BIOSYNTHETIC PROCESS | 9 | 128 | 1.911e-06 | 2.19e-05 |
| 407 | CELLULAR RESPONSE TO DRUG | 7 | 67 | 1.946e-06 | 2.225e-05 |
| 408 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 22 | 801 | 2.069e-06 | 2.359e-05 |
| 409 | POSITIVE REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 7 | 68 | 2.154e-06 | 2.45e-05 |
| 410 | ORGANOPHOSPHATE BIOSYNTHETIC PROCESS | 16 | 450 | 2.287e-06 | 2.595e-05 |
| 411 | REGULATION OF PROTEIN BINDING | 10 | 168 | 2.323e-06 | 2.611e-05 |
| 412 | AMINE METABOLIC PROCESS | 9 | 131 | 2.318e-06 | 2.611e-05 |
| 413 | LEARNING | 9 | 131 | 2.318e-06 | 2.611e-05 |
| 414 | BRANCHING MORPHOGENESIS OF AN EPITHELIAL TUBE | 9 | 131 | 2.318e-06 | 2.611e-05 |
| 415 | REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 8 | 98 | 2.358e-06 | 2.643e-05 |
| 416 | AMMONIUM ION METABOLIC PROCESS | 10 | 169 | 2.451e-06 | 2.741e-05 |
| 417 | LUNG MORPHOGENESIS | 6 | 45 | 2.528e-06 | 2.821e-05 |
| 418 | POSITIVE REGULATION OF PROTEIN SECRETION | 11 | 211 | 2.629e-06 | 2.92e-05 |
| 419 | REGULATION OF CALCIUM ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 7 | 70 | 2.624e-06 | 2.92e-05 |
| 420 | REGULATION OF RAC PROTEIN SIGNAL TRANSDUCTION | 4 | 12 | 2.654e-06 | 2.941e-05 |
| 421 | REGULATION OF P38MAPK CASCADE | 5 | 26 | 2.745e-06 | 3.034e-05 |
| 422 | EMBRYONIC CRANIAL SKELETON MORPHOGENESIS | 6 | 46 | 2.886e-06 | 3.182e-05 |
| 423 | CELL GROWTH | 9 | 135 | 2.974e-06 | 3.272e-05 |
| 424 | ESTABLISHMENT OF LOCALIZATION IN CELL | 34 | 1676 | 2.991e-06 | 3.283e-05 |
| 425 | STRIATED MUSCLE CELL DIFFERENTIATION | 10 | 173 | 3.024e-06 | 3.311e-05 |
| 426 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 15 | 408 | 3.201e-06 | 3.497e-05 |
| 427 | THYMUS DEVELOPMENT | 6 | 47 | 3.285e-06 | 3.568e-05 |
| 428 | FOREBRAIN DEVELOPMENT | 14 | 357 | 3.289e-06 | 3.568e-05 |
| 429 | POSITIVE REGULATION OF NEURON APOPTOTIC PROCESS | 6 | 47 | 3.285e-06 | 3.568e-05 |
| 430 | POSITIVE REGULATION OF HEART GROWTH | 5 | 27 | 3.345e-06 | 3.612e-05 |
| 431 | REGULATION OF FATTY ACID TRANSPORT | 5 | 27 | 3.345e-06 | 3.612e-05 |
| 432 | PLACENTA DEVELOPMENT | 9 | 138 | 3.566e-06 | 3.841e-05 |
| 433 | RESPONSE TO ESTROGEN | 11 | 218 | 3.602e-06 | 3.871e-05 |
| 434 | RESPONSE TO CAMP | 8 | 104 | 3.691e-06 | 3.957e-05 |
| 435 | REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 6 | 48 | 3.727e-06 | 3.959e-05 |
| 436 | METAL ION TRANSPORT | 18 | 582 | 3.727e-06 | 3.959e-05 |
| 437 | REGULATION OF LIPID KINASE ACTIVITY | 6 | 48 | 3.727e-06 | 3.959e-05 |
| 438 | REGULATION OF RESPONSE TO WOUNDING | 15 | 413 | 3.712e-06 | 3.959e-05 |
| 439 | RESPONSE TO ORGANOPHOSPHORUS | 9 | 139 | 3.784e-06 | 4.011e-05 |
| 440 | AGING | 12 | 264 | 3.803e-06 | 4.022e-05 |
| 441 | SINGLE ORGANISM CELLULAR LOCALIZATION | 23 | 898 | 3.832e-06 | 4.043e-05 |
| 442 | RESPONSE TO ALCOHOL | 14 | 362 | 3.862e-06 | 4.066e-05 |
| 443 | NUCLEOTIDE BINDING DOMAIN LEUCINE RICH REPEAT CONTAINING RECEPTOR SIGNALING PATHWAY | 5 | 28 | 4.044e-06 | 4.247e-05 |
| 444 | REGULATION OF HEMOPOIESIS | 13 | 314 | 4.148e-06 | 4.347e-05 |
| 445 | SECRETION | 18 | 588 | 4.295e-06 | 4.491e-05 |
| 446 | FACE DEVELOPMENT | 6 | 50 | 4.756e-06 | 4.951e-05 |
| 447 | REGULATION OF TUMOR NECROSIS FACTOR MEDIATED SIGNALING PATHWAY | 6 | 50 | 4.756e-06 | 4.951e-05 |
| 448 | POSITIVE REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 5 | 29 | 4.852e-06 | 5.039e-05 |
| 449 | POSITIVE REGULATION OF METANEPHROS DEVELOPMENT | 4 | 14 | 5.295e-06 | 5.462e-05 |
| 450 | POSITIVE REGULATION OF P38MAPK CASCADE | 4 | 14 | 5.295e-06 | 5.462e-05 |
| 451 | NEGATIVE REGULATION OF IMMUNE SYSTEM PROCESS | 14 | 372 | 5.28e-06 | 5.462e-05 |
| 452 | POSITIVE REGULATION OF TRANSCRIPTION FACTOR IMPORT INTO NUCLEUS | 6 | 51 | 5.352e-06 | 5.497e-05 |
| 453 | REGULATION OF NEUROTRANSMITTER SECRETION | 6 | 51 | 5.352e-06 | 5.497e-05 |
| 454 | PROTEIN LOCALIZATION | 35 | 1805 | 5.692e-06 | 5.834e-05 |
| 455 | REGULATION OF HEART RATE BY CARDIAC CONDUCTION | 5 | 30 | 5.781e-06 | 5.912e-05 |
| 456 | NEGATIVE REGULATION OF ERK1 AND ERK2 CASCADE | 6 | 52 | 6.006e-06 | 6.115e-05 |
| 457 | REGULATION OF T CELL MEDIATED IMMUNITY | 6 | 52 | 6.006e-06 | 6.115e-05 |
| 458 | EYE DEVELOPMENT | 13 | 326 | 6.227e-06 | 6.326e-05 |
| 459 | MUSCLE STRUCTURE DEVELOPMENT | 15 | 432 | 6.385e-06 | 6.473e-05 |
| 460 | NEGATIVE REGULATION OF CELL CYCLE | 15 | 433 | 6.564e-06 | 6.64e-05 |
| 461 | REGULATION OF MORPHOGENESIS OF A BRANCHING STRUCTURE | 6 | 53 | 6.724e-06 | 6.787e-05 |
| 462 | RESPONSE TO TUMOR NECROSIS FACTOR | 11 | 233 | 6.804e-06 | 6.852e-05 |
| 463 | POSITIVE REGULATION OF INTERLEUKIN 2 PRODUCTION | 5 | 31 | 6.844e-06 | 6.878e-05 |
| 464 | OVULATION CYCLE | 8 | 113 | 6.859e-06 | 6.879e-05 |
| 465 | REGULATION OF LYMPHOCYTE MEDIATED IMMUNITY | 8 | 114 | 7.323e-06 | 7.327e-05 |
| 466 | RESPONSE TO CALCIUM ION | 8 | 115 | 7.812e-06 | 7.8e-05 |
| 467 | MUSCLE CELL DIFFERENTIATION | 11 | 237 | 7.994e-06 | 7.965e-05 |
| 468 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 5 | 32 | 8.054e-06 | 7.991e-05 |
| 469 | NEGATIVE REGULATION OF SIGNAL TRANSDUCTION IN ABSENCE OF LIGAND | 5 | 32 | 8.054e-06 | 7.991e-05 |
| 470 | NIK NF KAPPAB SIGNALING | 7 | 83 | 8.263e-06 | 8.18e-05 |
| 471 | REGULATION OF CALCIUM ION TRANSMEMBRANE TRANSPORT | 8 | 116 | 8.329e-06 | 8.228e-05 |
| 472 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 6 | 55 | 8.369e-06 | 8.233e-05 |
| 473 | CRANIAL SKELETAL SYSTEM DEVELOPMENT | 6 | 55 | 8.369e-06 | 8.233e-05 |
| 474 | CELL CYCLE ARREST | 9 | 154 | 8.737e-06 | 8.558e-05 |
| 475 | POSITIVE REGULATION OF PEPTIDASE ACTIVITY | 9 | 154 | 8.737e-06 | 8.558e-05 |
| 476 | EMBRYONIC DIGESTIVE TRACT DEVELOPMENT | 5 | 33 | 9.426e-06 | 9.137e-05 |
| 477 | SIGNAL TRANSDUCTION IN ABSENCE OF LIGAND | 5 | 33 | 9.426e-06 | 9.137e-05 |
| 478 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 5 | 33 | 9.426e-06 | 9.137e-05 |
| 479 | CYTOPLASMIC PATTERN RECOGNITION RECEPTOR SIGNALING PATHWAY | 5 | 33 | 9.426e-06 | 9.137e-05 |
| 480 | PHOSPHATIDIC ACID METABOLIC PROCESS | 5 | 33 | 9.426e-06 | 9.137e-05 |
| 481 | REGULATION OF PEPTIDYL SERINE PHOSPHORYLATION | 8 | 118 | 9.449e-06 | 9.141e-05 |
| 482 | POSITIVE REGULATION OF VASCULAR ENDOTHELIAL GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 4 | 16 | 9.495e-06 | 9.166e-05 |
| 483 | REGULATION OF HOMEOSTATIC PROCESS | 15 | 447 | 9.591e-06 | 9.239e-05 |
| 484 | POSITIVE REGULATION OF DEVELOPMENTAL GROWTH | 9 | 156 | 9.697e-06 | 9.303e-05 |
| 485 | REGULATION OF LEUKOCYTE MEDIATED IMMUNITY | 9 | 156 | 9.697e-06 | 9.303e-05 |
| 486 | REGULATION OF CYTOKINE PRODUCTION INVOLVED IN IMMUNE RESPONSE | 6 | 57 | 1.033e-05 | 9.886e-05 |
| 487 | RESPONSE TO PURINE CONTAINING COMPOUND | 9 | 158 | 1.075e-05 | 0.0001027 |
| 488 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 10 | 200 | 1.092e-05 | 0.0001042 |
| 489 | FOREBRAIN NEURON DEVELOPMENT | 5 | 34 | 1.097e-05 | 0.0001042 |
| 490 | ORGAN FORMATION | 5 | 34 | 1.097e-05 | 0.0001042 |
| 491 | TISSUE REMODELING | 7 | 87 | 1.129e-05 | 0.000107 |
| 492 | SKELETAL SYSTEM MORPHOGENESIS | 10 | 201 | 1.141e-05 | 0.0001079 |
| 493 | INFLAMMATORY RESPONSE | 15 | 454 | 1.153e-05 | 0.0001088 |
| 494 | SECOND MESSENGER MEDIATED SIGNALING | 9 | 160 | 1.189e-05 | 0.000112 |
| 495 | OVULATION CYCLE PROCESS | 7 | 88 | 1.218e-05 | 0.0001143 |
| 496 | REGULATION OF STEM CELL PROLIFERATION | 7 | 88 | 1.218e-05 | 0.0001143 |
| 497 | REGULATION OF NFAT PROTEIN IMPORT INTO NUCLEUS | 4 | 17 | 1.233e-05 | 0.000115 |
| 498 | REGULATION OF ENDOTHELIAL CELL CHEMOTAXIS | 4 | 17 | 1.233e-05 | 0.000115 |
| 499 | REGULATION OF PRI MIRNA TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 4 | 17 | 1.233e-05 | 0.000115 |
| 500 | NEGATIVE REGULATION OF TRANSPORT | 15 | 458 | 1.278e-05 | 0.0001189 |
| 501 | EPITHELIAL CELL PROLIFERATION | 7 | 89 | 1.312e-05 | 0.0001219 |
| 502 | RESPONSE TO PEPTIDE | 14 | 404 | 1.342e-05 | 0.0001242 |
| 503 | UROGENITAL SYSTEM DEVELOPMENT | 12 | 299 | 1.341e-05 | 0.0001242 |
| 504 | CELLULAR RESPONSE TO BIOTIC STIMULUS | 9 | 163 | 1.38e-05 | 0.0001274 |
| 505 | CHONDROCYTE DIFFERENTIATION | 6 | 60 | 1.394e-05 | 0.0001284 |
| 506 | CYTOSKELETON ORGANIZATION | 21 | 838 | 1.453e-05 | 0.0001336 |
| 507 | CELLULAR RESPONSE TO HEAT | 5 | 36 | 1.466e-05 | 0.0001345 |
| 508 | EPIDERMIS DEVELOPMENT | 11 | 253 | 1.476e-05 | 0.0001352 |
| 509 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 15 | 465 | 1.527e-05 | 0.0001396 |
| 510 | POSITIVE REGULATION OF STEM CELL PROLIFERATION | 6 | 61 | 1.535e-05 | 0.0001399 |
| 511 | REGULATION OF CATION TRANSMEMBRANE TRANSPORT | 10 | 208 | 1.536e-05 | 0.0001399 |
| 512 | GROWTH | 14 | 410 | 1.582e-05 | 0.0001437 |
| 513 | REGULATION OF PROTEOLYSIS | 19 | 711 | 1.584e-05 | 0.0001437 |
| 514 | POSITIVE REGULATION OF BINDING | 8 | 127 | 1.62e-05 | 0.0001466 |
| 515 | ESTABLISHMENT OF PROTEIN LOCALIZATION | 29 | 1423 | 1.661e-05 | 0.00015 |
| 516 | RESPONSE TO NERVE GROWTH FACTOR | 5 | 37 | 1.683e-05 | 0.0001518 |
| 517 | MUSCLE CELL DEVELOPMENT | 8 | 128 | 1.715e-05 | 0.0001543 |
| 518 | LEUKOCYTE MIGRATION | 11 | 259 | 1.836e-05 | 0.0001649 |
| 519 | POSITIVE REGULATION OF LEUKOCYTE DIFFERENTIATION | 8 | 131 | 2.029e-05 | 0.0001816 |
| 520 | REGULATION OF NEUROTRANSMITTER TRANSPORT | 6 | 64 | 2.028e-05 | 0.0001816 |
| 521 | WNT SIGNALING PATHWAY CALCIUM MODULATING PATHWAY | 5 | 39 | 2.192e-05 | 0.0001954 |
| 522 | LIPID BIOSYNTHETIC PROCESS | 16 | 539 | 2.189e-05 | 0.0001954 |
| 523 | POSITIVE REGULATION OF LIPID BIOSYNTHETIC PROCESS | 6 | 66 | 2.423e-05 | 0.0002155 |
| 524 | GLIOGENESIS | 9 | 175 | 2.43e-05 | 0.0002158 |
| 525 | REGULATION OF ICOSANOID SECRETION | 4 | 20 | 2.46e-05 | 0.0002176 |
| 526 | MICROVILLUS ORGANIZATION | 4 | 20 | 2.46e-05 | 0.0002176 |
| 527 | GLIAL CELL DIFFERENTIATION | 8 | 136 | 2.661e-05 | 0.0002349 |
| 528 | NEGATIVE REGULATION OF STRESS ACTIVATED MAPK CASCADE | 5 | 41 | 2.812e-05 | 0.0002469 |
| 529 | REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 7 | 100 | 2.812e-05 | 0.0002469 |
| 530 | NEGATIVE REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 5 | 41 | 2.812e-05 | 0.0002469 |
| 531 | CELLULAR RESPONSE TO HORMONE STIMULUS | 16 | 552 | 2.921e-05 | 0.0002559 |
| 532 | NECROPTOTIC PROCESS | 4 | 21 | 3.018e-05 | 0.0002639 |
| 533 | POSITIVE REGULATION OF LYMPHOCYTE MEDIATED IMMUNITY | 6 | 69 | 3.128e-05 | 0.0002731 |
| 534 | NEGATIVE REGULATION OF IMMUNE EFFECTOR PROCESS | 7 | 102 | 3.197e-05 | 0.0002786 |
| 535 | CENTRAL NERVOUS SYSTEM NEURON DEVELOPMENT | 6 | 70 | 3.396e-05 | 0.0002954 |
| 536 | REGULATION OF MYELOID CELL DIFFERENTIATION | 9 | 183 | 3.455e-05 | 3e-04 |
| 537 | CELL CELL SIGNALING INVOLVED IN CARDIAC CONDUCTION | 4 | 22 | 3.663e-05 | 0.0003168 |
| 538 | SOMATIC STEM CELL DIVISION | 4 | 22 | 3.663e-05 | 0.0003168 |
| 539 | SKIN EPIDERMIS DEVELOPMENT | 6 | 71 | 3.683e-05 | 0.000318 |
| 540 | POSITIVE REGULATION OF CELL CYCLE | 12 | 332 | 3.756e-05 | 0.0003236 |
| 541 | CEREBRAL CORTEX DEVELOPMENT | 7 | 105 | 3.855e-05 | 0.0003316 |
| 542 | POSITIVE REGULATION OF NEURON PROJECTION DEVELOPMENT | 10 | 232 | 3.915e-05 | 0.0003361 |
| 543 | POSITIVE REGULATION OF CALCIUM ION TRANSPORT | 7 | 106 | 4.098e-05 | 0.0003511 |
| 544 | SYNAPSE ORGANIZATION | 8 | 145 | 4.212e-05 | 0.0003602 |
| 545 | CELLULAR CHEMICAL HOMEOSTASIS | 16 | 570 | 4.288e-05 | 0.0003661 |
| 546 | POSITIVE REGULATION OF PROTEIN BINDING | 6 | 73 | 4.315e-05 | 0.0003677 |
| 547 | REGULATION OF PEPTIDASE ACTIVITY | 13 | 392 | 4.331e-05 | 0.0003684 |
| 548 | REGULATION OF METANEPHROS DEVELOPMENT | 4 | 23 | 4.404e-05 | 0.0003739 |
| 549 | REGULATION OF MYELOID LEUKOCYTE DIFFERENTIATION | 7 | 108 | 4.62e-05 | 0.0003916 |
| 550 | CARTILAGE DEVELOPMENT | 8 | 147 | 4.643e-05 | 0.0003928 |
| 551 | DIGESTIVE SYSTEM DEVELOPMENT | 8 | 148 | 4.872e-05 | 0.0004115 |
| 552 | RESPONSE TO HYDROGEN PEROXIDE | 7 | 109 | 4.901e-05 | 0.0004131 |
| 553 | INTRASPECIES INTERACTION BETWEEN ORGANISMS | 5 | 46 | 4.966e-05 | 0.0004171 |
| 554 | SOCIAL BEHAVIOR | 5 | 46 | 4.966e-05 | 0.0004171 |
| 555 | RESPONSE TO TOXIC SUBSTANCE | 10 | 241 | 5.395e-05 | 0.0004523 |
| 556 | REGULATION OF ORGAN MORPHOGENESIS | 10 | 242 | 5.585e-05 | 0.0004674 |
| 557 | INORGANIC ION TRANSMEMBRANE TRANSPORT | 16 | 583 | 5.602e-05 | 0.000468 |
| 558 | INTRACELLULAR PROTEIN TRANSPORT | 19 | 781 | 5.678e-05 | 0.0004735 |
| 559 | POSITIVE REGULATION OF REACTIVE OXYGEN SPECIES BIOSYNTHETIC PROCESS | 5 | 48 | 6.116e-05 | 0.0005082 |
| 560 | REGULATION OF INTERLEUKIN 2 PRODUCTION | 5 | 48 | 6.116e-05 | 0.0005082 |
| 561 | PALLIUM DEVELOPMENT | 8 | 153 | 6.165e-05 | 0.0005113 |
| 562 | EPITHELIAL CELL APOPTOTIC PROCESS | 4 | 25 | 6.206e-05 | 0.0005129 |
| 563 | HISTONE PHOSPHORYLATION | 4 | 25 | 6.206e-05 | 0.0005129 |
| 564 | LYMPHOCYTE COSTIMULATION | 6 | 78 | 6.28e-05 | 0.0005181 |
| 565 | POSITIVE REGULATION OF ENDOCYTOSIS | 7 | 114 | 6.525e-05 | 0.0005364 |
| 566 | REGULATION OF ENDOTHELIAL CELL MIGRATION | 7 | 114 | 6.525e-05 | 0.0005364 |
| 567 | REGULATION OF ENDOCYTOSIS | 9 | 199 | 6.629e-05 | 0.000544 |
| 568 | BONE MORPHOGENESIS | 6 | 79 | 6.748e-05 | 0.0005528 |
| 569 | CELLULAR RESPONSE TO CALCIUM ION | 5 | 49 | 6.763e-05 | 0.0005531 |
| 570 | RESPONSE TO INTERLEUKIN 1 | 7 | 115 | 6.897e-05 | 0.0005621 |
| 571 | NEPHRON DEVELOPMENT | 7 | 115 | 6.897e-05 | 0.0005621 |
| 572 | CELLULAR RESPONSE TO INORGANIC SUBSTANCE | 8 | 156 | 7.069e-05 | 0.000575 |
| 573 | DEVELOPMENTAL PROGRAMMED CELL DEATH | 4 | 26 | 7.284e-05 | 0.0005905 |
| 574 | REGULATION OF CELL FATE COMMITMENT | 4 | 26 | 7.284e-05 | 0.0005905 |
| 575 | REGULATION OF ORGANIC ACID TRANSPORT | 5 | 50 | 7.462e-05 | 0.0006001 |
| 576 | REGULATION OF SYNAPTIC TRANSMISSION GLUTAMATERGIC | 5 | 50 | 7.462e-05 | 0.0006001 |
| 577 | LYMPHOCYTE HOMEOSTASIS | 5 | 50 | 7.462e-05 | 0.0006001 |
| 578 | LIPID METABOLIC PROCESS | 24 | 1158 | 7.468e-05 | 0.0006001 |
| 579 | POSITIVE REGULATION OF MYELOID LEUKOCYTE DIFFERENTIATION | 5 | 50 | 7.462e-05 | 0.0006001 |
| 580 | OSSIFICATION | 10 | 251 | 7.574e-05 | 0.0006077 |
| 581 | NEGATIVE REGULATION OF CELL ACTIVATION | 8 | 158 | 7.73e-05 | 0.0006191 |
| 582 | CELL CYCLE | 26 | 1316 | 8.028e-05 | 0.0006418 |
| 583 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 11 | 306 | 8.347e-05 | 0.0006662 |
| 584 | REGULATION OF VASCULAR ENDOTHELIAL GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 4 | 27 | 8.493e-05 | 0.0006767 |
| 585 | HAIR CYCLE | 6 | 83 | 8.902e-05 | 0.0007069 |
| 586 | MOLTING CYCLE | 6 | 83 | 8.902e-05 | 0.0007069 |
| 587 | NEGATIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 18 | 740 | 9.004e-05 | 0.0007137 |
| 588 | CELL CHEMOTAXIS | 8 | 162 | 9.208e-05 | 0.0007287 |
| 589 | REGULATION OF PROTEIN KINASE B SIGNALING | 7 | 121 | 9.513e-05 | 0.0007515 |
| 590 | POSITIVE REGULATION OF HEMOPOIESIS | 8 | 163 | 9.612e-05 | 0.0007575 |
| 591 | RESPONSE TO OXYGEN LEVELS | 11 | 311 | 9.637e-05 | 0.0007575 |
| 592 | HEMOSTASIS | 11 | 311 | 9.637e-05 | 0.0007575 |
| 593 | POSITIVE REGULATION OF LEUKOCYTE APOPTOTIC PROCESS | 4 | 28 | 9.841e-05 | 0.0007696 |
| 594 | REGULATION OF FIBROBLAST MIGRATION | 4 | 28 | 9.841e-05 | 0.0007696 |
| 595 | POSITIVE REGULATION OF SUBSTRATE ADHESION DEPENDENT CELL SPREADING | 4 | 28 | 9.841e-05 | 0.0007696 |
| 596 | REGULATION OF NITRIC OXIDE BIOSYNTHETIC PROCESS | 5 | 53 | 9.895e-05 | 0.0007725 |
| 597 | SECRETION BY CELL | 14 | 486 | 9.978e-05 | 0.0007777 |
| 598 | EMBRYONIC SKELETAL SYSTEM DEVELOPMENT | 7 | 122 | 0.0001002 | 0.0007796 |
| 599 | EPHRIN RECEPTOR SIGNALING PATHWAY | 6 | 85 | 0.0001017 | 0.0007872 |
| 600 | HEART PROCESS | 6 | 85 | 0.0001017 | 0.0007872 |
| 601 | POSITIVE REGULATION OF LEUKOCYTE MEDIATED IMMUNITY | 6 | 85 | 0.0001017 | 0.0007872 |
| 602 | POSITIVE REGULATION OF ENDOTHELIAL CELL CHEMOTAXIS | 3 | 11 | 0.0001032 | 0.0007936 |
| 603 | NEGATIVE REGULATION OF NEUROTRANSMITTER SECRETION | 3 | 11 | 0.0001032 | 0.0007936 |
| 604 | POSITIVE REGULATION OF PRI MIRNA TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 3 | 11 | 0.0001032 | 0.0007936 |
| 605 | REGULATION OF FEVER GENERATION | 3 | 11 | 0.0001032 | 0.0007936 |
| 606 | SKIN DEVELOPMENT | 9 | 211 | 0.0001038 | 0.0007969 |
| 607 | POSITIVE REGULATION OF MITOTIC CELL CYCLE | 7 | 123 | 0.0001055 | 0.0008084 |
| 608 | STEM CELL DIVISION | 4 | 29 | 0.0001134 | 0.0008649 |
| 609 | POSITIVE REGULATION OF NIK NF KAPPAB SIGNALING | 4 | 29 | 0.0001134 | 0.0008649 |
| 610 | CARDIAC MUSCLE CELL CONTRACTION | 4 | 29 | 0.0001134 | 0.0008649 |
| 611 | CELL PROJECTION ASSEMBLY | 10 | 264 | 0.0001149 | 0.0008751 |
| 612 | REGULATION OF KIDNEY DEVELOPMENT | 5 | 55 | 0.0001183 | 0.0008992 |
| 613 | EPITHELIAL CELL DIFFERENTIATION | 14 | 495 | 0.0001209 | 0.000918 |
| 614 | REGULATION OF WOUND HEALING | 7 | 126 | 0.0001226 | 0.0009279 |
| 615 | RESPONSE TO UV | 7 | 126 | 0.0001226 | 0.0009279 |
| 616 | PROTEIN LOCALIZATION TO MEMBRANE | 12 | 376 | 0.0001229 | 0.0009285 |
| 617 | CELLULAR RESPONSE TO INTERLEUKIN 1 | 6 | 88 | 0.0001233 | 0.0009298 |
| 618 | POSITIVE REGULATION OF HOMEOSTATIC PROCESS | 9 | 216 | 0.000124 | 0.0009333 |
| 619 | NEURON NEURON SYNAPTIC TRANSMISSION | 5 | 56 | 0.000129 | 0.0009678 |
| 620 | POSITIVE REGULATION OF MUSCLE TISSUE DEVELOPMENT | 5 | 56 | 0.000129 | 0.0009678 |
| 621 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 13 | 437 | 0.0001293 | 0.0009689 |
| 622 | MYD88 INDEPENDENT TOLL LIKE RECEPTOR SIGNALING PATHWAY | 4 | 30 | 0.0001299 | 0.0009689 |
| 623 | REGULATION OF NEUROTRANSMITTER RECEPTOR ACTIVITY | 4 | 30 | 0.0001299 | 0.0009689 |
| 624 | POSITIVE REGULATION OF ORGANIC ACID TRANSPORT | 4 | 30 | 0.0001299 | 0.0009689 |
| 625 | REGULATION OF VITAMIN METABOLIC PROCESS | 3 | 12 | 0.0001367 | 0.001016 |
| 626 | TRACHEA MORPHOGENESIS | 3 | 12 | 0.0001367 | 0.001016 |
| 627 | NUCLEAR IMPORT | 7 | 129 | 0.000142 | 0.001054 |
| 628 | MULTICELLULAR ORGANISM REPRODUCTION | 18 | 768 | 0.0001432 | 0.001061 |
| 629 | MULTICELLULAR ORGANISMAL HOMEOSTASIS | 10 | 272 | 0.0001466 | 0.001085 |
| 630 | REGULATION OF PROTEIN STABILITY | 9 | 221 | 0.0001473 | 0.001088 |
| 631 | LIPOPOLYSACCHARIDE MEDIATED SIGNALING PATHWAY | 4 | 31 | 0.0001482 | 0.001091 |
| 632 | REGULATION OF PLATELET ACTIVATION | 4 | 31 | 0.0001482 | 0.001091 |
| 633 | CELLULAR RESPONSE TO PEPTIDE | 10 | 274 | 0.0001556 | 0.001144 |
| 634 | HOMEOSTASIS OF NUMBER OF CELLS | 8 | 175 | 0.0001572 | 0.001154 |
| 635 | REGULATED EXOCYTOSIS | 9 | 224 | 0.000163 | 0.001195 |
| 636 | RESPONSE TO CORTICOSTEROID | 8 | 176 | 0.0001635 | 0.001196 |
| 637 | EMBRYONIC SKELETAL SYSTEM MORPHOGENESIS | 6 | 93 | 0.0001673 | 0.001222 |
| 638 | MYD88 DEPENDENT TOLL LIKE RECEPTOR SIGNALING PATHWAY | 4 | 32 | 0.0001682 | 0.001225 |
| 639 | POSITIVE REGULATION OF T CELL MEDIATED IMMUNITY | 4 | 32 | 0.0001682 | 0.001225 |
| 640 | NEURONAL STEM CELL DIVISION | 3 | 13 | 0.0001766 | 0.001264 |
| 641 | REGULATION OF PROSTAGLANDIN SECRETION | 3 | 13 | 0.0001766 | 0.001264 |
| 642 | PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 3 | 13 | 0.0001766 | 0.001264 |
| 643 | POSITIVE REGULATION OF STEROID BIOSYNTHETIC PROCESS | 3 | 13 | 0.0001766 | 0.001264 |
| 644 | REGULATION OF CELL PROLIFERATION INVOLVED IN KIDNEY DEVELOPMENT | 3 | 13 | 0.0001766 | 0.001264 |
| 645 | EYELID DEVELOPMENT IN CAMERA TYPE EYE | 3 | 13 | 0.0001766 | 0.001264 |
| 646 | HEPATOCYTE APOPTOTIC PROCESS | 3 | 13 | 0.0001766 | 0.001264 |
| 647 | INTERLEUKIN 1 MEDIATED SIGNALING PATHWAY | 3 | 13 | 0.0001766 | 0.001264 |
| 648 | NEUROBLAST DIVISION | 3 | 13 | 0.0001766 | 0.001264 |
| 649 | REGULATION OF CELL FATE SPECIFICATION | 3 | 13 | 0.0001766 | 0.001264 |
| 650 | ACTIVATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 13 | 0.0001766 | 0.001264 |
| 651 | REGULATION OF DNA BIOSYNTHETIC PROCESS | 6 | 94 | 0.0001774 | 0.001268 |
| 652 | REGULATION OF MONOOXYGENASE ACTIVITY | 5 | 60 | 0.0001793 | 0.001277 |
| 653 | LEUKOCYTE HOMEOSTASIS | 5 | 60 | 0.0001793 | 0.001277 |
| 654 | EMBRYONIC ORGAN MORPHOGENESIS | 10 | 279 | 0.0001802 | 0.001282 |
| 655 | TELENCEPHALON DEVELOPMENT | 9 | 228 | 0.0001861 | 0.001322 |
| 656 | FC GAMMA RECEPTOR SIGNALING PATHWAY | 6 | 95 | 0.000188 | 0.00133 |
| 657 | ACTIVATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 6 | 95 | 0.000188 | 0.00133 |
| 658 | REGULATION OF LIPID TRANSPORT | 6 | 95 | 0.000188 | 0.00133 |
| 659 | NEGATIVE REGULATION OF JNK CASCADE | 4 | 33 | 0.0001901 | 0.001342 |
| 660 | SKELETAL SYSTEM DEVELOPMENT | 13 | 455 | 0.0001922 | 0.001355 |
| 661 | REGULATION OF INTERLEUKIN 8 PRODUCTION | 5 | 61 | 0.0001939 | 0.001365 |
| 662 | MUSCLE SYSTEM PROCESS | 10 | 282 | 0.0001964 | 0.001381 |
| 663 | FEMALE GAMETE GENERATION | 6 | 96 | 0.0001991 | 0.001397 |
| 664 | PLATELET DERIVED GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 4 | 34 | 0.000214 | 0.001499 |
| 665 | REGULATION OF RESPIRATORY BURST | 3 | 14 | 0.0002233 | 0.001546 |
| 666 | MEMORY | 6 | 98 | 0.0002229 | 0.001546 |
| 667 | CATION TRANSPORT | 18 | 796 | 0.0002223 | 0.001546 |
| 668 | ARACHIDONIC ACID SECRETION | 3 | 14 | 0.0002233 | 0.001546 |
| 669 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 3 | 14 | 0.0002233 | 0.001546 |
| 670 | POSITIVE REGULATION OF T CELL CYTOKINE PRODUCTION | 3 | 14 | 0.0002233 | 0.001546 |
| 671 | ARACHIDONATE TRANSPORT | 3 | 14 | 0.0002233 | 0.001546 |
| 672 | CELLULAR RESPONSE TO OXIDATIVE STRESS | 8 | 184 | 0.0002215 | 0.001546 |
| 673 | STRIATED MUSCLE CONTRACTION | 6 | 99 | 0.0002355 | 0.001628 |
| 674 | BONE REMODELING | 4 | 35 | 0.0002399 | 0.001656 |
| 675 | CHEMICAL HOMEOSTASIS | 19 | 874 | 0.0002438 | 0.00168 |
| 676 | NEGATIVE REGULATION OF GENE EXPRESSION | 27 | 1493 | 0.0002478 | 0.001705 |
| 677 | REGULATION OF MITOTIC CELL CYCLE | 13 | 468 | 0.0002525 | 0.001735 |
| 678 | LEUKOCYTE DIFFERENTIATION | 10 | 292 | 0.0002597 | 0.001783 |
| 679 | CALCIUM ION IMPORT | 5 | 65 | 0.0002617 | 0.001788 |
| 680 | REGULATION OF REACTIVE OXYGEN SPECIES BIOSYNTHETIC PROCESS | 5 | 65 | 0.0002617 | 0.001788 |
| 681 | POSITIVE REGULATION OF INTERFERON GAMMA PRODUCTION | 5 | 65 | 0.0002617 | 0.001788 |
| 682 | REGULATION OF PRODUCTION OF MOLECULAR MEDIATOR OF IMMUNE RESPONSE | 6 | 101 | 0.0002626 | 0.001792 |
| 683 | MYELOID CELL DIFFERENTIATION | 8 | 189 | 0.0002656 | 0.00181 |
| 684 | NEGATIVE REGULATION OF INTRACELLULAR TRANSPORT | 7 | 143 | 0.0002682 | 0.001819 |
| 685 | POSITIVE CHEMOTAXIS | 4 | 36 | 0.0002681 | 0.001819 |
| 686 | HEAD MORPHOGENESIS | 4 | 36 | 0.0002681 | 0.001819 |
| 687 | MESENCHYME DEVELOPMENT | 8 | 190 | 0.0002753 | 0.001864 |
| 688 | REGULATION OF HEAT GENERATION | 3 | 15 | 0.0002773 | 0.001867 |
| 689 | NEGATIVE REGULATION OF DENDRITE MORPHOGENESIS | 3 | 15 | 0.0002773 | 0.001867 |
| 690 | REGULATION OF N METHYL D ASPARTATE SELECTIVE GLUTAMATE RECEPTOR ACTIVITY | 3 | 15 | 0.0002773 | 0.001867 |
| 691 | NEUROTROPHIN TRK RECEPTOR SIGNALING PATHWAY | 3 | 15 | 0.0002773 | 0.001867 |
| 692 | FOREBRAIN GENERATION OF NEURONS | 5 | 66 | 0.0002812 | 0.001891 |
| 693 | REGULATION OF MUSCLE TISSUE DEVELOPMENT | 6 | 103 | 0.000292 | 0.001958 |
| 694 | REGULATION OF MUSCLE ORGAN DEVELOPMENT | 6 | 103 | 0.000292 | 0.001958 |
| 695 | CELLULAR RESPONSE TO CYTOKINE STIMULUS | 15 | 606 | 0.0002945 | 0.001972 |
| 696 | LEUKOCYTE ACTIVATION | 12 | 414 | 0.0002981 | 0.001993 |
| 697 | CELL COMMUNICATION INVOLVED IN CARDIAC CONDUCTION | 4 | 37 | 0.0002986 | 0.001993 |
| 698 | POSITIVE REGULATION OF ENDOTHELIAL CELL MIGRATION | 5 | 67 | 0.0003017 | 0.002011 |
| 699 | RESPONSE TO ESTRADIOL | 7 | 146 | 0.0003044 | 0.002024 |
| 700 | CELLULAR HOMEOSTASIS | 16 | 676 | 0.0003044 | 0.002024 |
| 701 | DEVELOPMENTAL GROWTH INVOLVED IN MORPHOGENESIS | 6 | 104 | 0.0003077 | 0.002042 |
| 702 | REGULATION OF HORMONE LEVELS | 13 | 478 | 0.0003093 | 0.00205 |
| 703 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 15 | 609 | 0.0003103 | 0.002054 |
| 704 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 27 | 1517 | 0.0003192 | 0.00211 |
| 705 | REGULATION OF CELL JUNCTION ASSEMBLY | 5 | 68 | 0.0003234 | 0.002128 |
| 706 | MULTICELLULAR ORGANISMAL RESPONSE TO STRESS | 5 | 68 | 0.0003234 | 0.002128 |
| 707 | ORGAN GROWTH | 5 | 68 | 0.0003234 | 0.002128 |
| 708 | EAR DEVELOPMENT | 8 | 195 | 0.0003279 | 0.002155 |
| 709 | POSITIVE REGULATION OF CELL GROWTH | 7 | 148 | 0.0003306 | 0.00217 |
| 710 | NEGATIVE REGULATION OF NEUROTRANSMITTER TRANSPORT | 3 | 16 | 0.0003391 | 0.002216 |
| 711 | CELLULAR RESPONSE TO ANTIBIOTIC | 3 | 16 | 0.0003391 | 0.002216 |
| 712 | BRANCHING INVOLVED IN SALIVARY GLAND MORPHOGENESIS | 3 | 16 | 0.0003391 | 0.002216 |
| 713 | MEMBRANE ORGANIZATION | 19 | 899 | 0.0003466 | 0.002259 |
| 714 | POSITIVE REGULATION OF AXONOGENESIS | 5 | 69 | 0.0003462 | 0.002259 |
| 715 | ASTROCYTE DIFFERENTIATION | 4 | 39 | 0.0003668 | 0.002384 |
| 716 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 4 | 39 | 0.0003668 | 0.002384 |
| 717 | REGULATION OF MEMBRANE PERMEABILITY | 5 | 70 | 0.0003702 | 0.002403 |
| 718 | PROTEIN LOCALIZATION TO CELL PERIPHERY | 7 | 151 | 0.0003733 | 0.002419 |
| 719 | NEGATIVE REGULATION OF SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 4 | 40 | 0.0004048 | 0.002619 |
| 720 | NEGATIVE REGULATION OF PLATELET ACTIVATION | 3 | 17 | 0.0004091 | 0.002626 |
| 721 | ACTIVATION OF NF KAPPAB INDUCING KINASE ACTIVITY | 3 | 17 | 0.0004091 | 0.002626 |
| 722 | REGULATION OF IMMUNOGLOBULIN SECRETION | 3 | 17 | 0.0004091 | 0.002626 |
| 723 | POSITIVE REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 17 | 0.0004091 | 0.002626 |
| 724 | NEGATIVE REGULATION OF GLUCOSE TRANSPORT | 3 | 17 | 0.0004091 | 0.002626 |
| 725 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 3 | 17 | 0.0004091 | 0.002626 |
| 726 | NEURON MIGRATION | 6 | 110 | 0.000416 | 0.002666 |
| 727 | EXOCYTOSIS | 10 | 310 | 0.0004168 | 0.002668 |
| 728 | REGULATION OF CYTOSOLIC CALCIUM ION CONCENTRATION | 8 | 203 | 0.0004289 | 0.002741 |
| 729 | PROTEIN IMPORT | 7 | 155 | 0.000437 | 0.002789 |
| 730 | POSITIVE REGULATION OF KIDNEY DEVELOPMENT | 4 | 41 | 0.0004455 | 0.002836 |
| 731 | LUNG ALVEOLUS DEVELOPMENT | 4 | 41 | 0.0004455 | 0.002836 |
| 732 | POSITIVE REGULATION OF ADAPTIVE IMMUNE RESPONSE | 5 | 73 | 0.0004499 | 0.00286 |
| 733 | POSITIVE REGULATION OF IMMUNE EFFECTOR PROCESS | 7 | 156 | 0.0004542 | 0.002879 |
| 734 | PROTEIN LOCALIZATION TO NUCLEUS | 7 | 156 | 0.0004542 | 0.002879 |
| 735 | RESPONSE TO AMINO ACID | 6 | 112 | 0.000458 | 0.002896 |
| 736 | ZYMOGEN ACTIVATION | 6 | 112 | 0.000458 | 0.002896 |
| 737 | REGULATION OF LEUKOCYTE PROLIFERATION | 8 | 206 | 0.0004727 | 0.002985 |
| 738 | REGULATION OF PROTEIN COMPLEX ASSEMBLY | 11 | 375 | 0.0004817 | 0.003037 |
| 739 | OVULATION | 3 | 18 | 0.0004878 | 0.003067 |
| 740 | RESPONSE TO WATER | 3 | 18 | 0.0004878 | 0.003067 |
| 741 | REGULATION OF NIK NF KAPPAB SIGNALING | 4 | 42 | 0.0004891 | 0.003071 |
| 742 | REGULATION OF EMBRYONIC DEVELOPMENT | 6 | 114 | 0.0005033 | 0.003156 |
| 743 | CELLULAR GLUCOSE HOMEOSTASIS | 5 | 75 | 0.0005097 | 0.003188 |
| 744 | MULTI ORGANISM BEHAVIOR | 5 | 75 | 0.0005097 | 0.003188 |
| 745 | VESICLE MEDIATED TRANSPORT | 23 | 1239 | 0.000519 | 0.003242 |
| 746 | NEGATIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 6 | 115 | 0.0005272 | 0.003289 |
| 747 | REGULATION OF BODY FLUID LEVELS | 13 | 506 | 0.0005298 | 0.0033 |
| 748 | REGULATION OF SUBSTRATE ADHESION DEPENDENT CELL SPREADING | 4 | 43 | 0.0005356 | 0.003323 |
| 749 | REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 4 | 43 | 0.0005356 | 0.003323 |
| 750 | CEREBRAL CORTEX CELL MIGRATION | 4 | 43 | 0.0005356 | 0.003323 |
| 751 | NEGATIVE REGULATION OF CYTOKINE PRODUCTION | 8 | 211 | 0.0005538 | 0.003431 |
| 752 | RESPONSE TO MUSCLE STRETCH | 3 | 19 | 0.0005756 | 0.003542 |
| 753 | REGULATION OF ALPHA AMINO 3 HYDROXY 5 METHYL 4 ISOXAZOLE PROPIONATE SELECTIVE GLUTAMATE RECEPTOR ACTIVITY | 3 | 19 | 0.0005756 | 0.003542 |
| 754 | KIDNEY VASCULATURE DEVELOPMENT | 3 | 19 | 0.0005756 | 0.003542 |
| 755 | NEGATIVE REGULATION OF MEIOTIC CELL CYCLE | 3 | 19 | 0.0005756 | 0.003542 |
| 756 | RENAL SYSTEM VASCULATURE DEVELOPMENT | 3 | 19 | 0.0005756 | 0.003542 |
| 757 | LABYRINTHINE LAYER DEVELOPMENT | 4 | 44 | 0.0005851 | 0.003592 |
| 758 | BODY MORPHOGENESIS | 4 | 44 | 0.0005851 | 0.003592 |
| 759 | MULTI MULTICELLULAR ORGANISM PROCESS | 8 | 213 | 0.0005893 | 0.003613 |
| 760 | CELLULAR RESPONSE TO DNA DAMAGE STIMULUS | 16 | 720 | 0.000605 | 0.003704 |
| 761 | POSITIVE REGULATION OF INTERLEUKIN 8 PRODUCTION | 4 | 45 | 0.0006379 | 0.00389 |
| 762 | ENDOCHONDRAL BONE MORPHOGENESIS | 4 | 45 | 0.0006379 | 0.00389 |
| 763 | REGULATION OF ALCOHOL BIOSYNTHETIC PROCESS | 4 | 45 | 0.0006379 | 0.00389 |
| 764 | IMMUNE RESPONSE | 21 | 1100 | 0.0006439 | 0.003921 |
| 765 | DEVELOPMENT OF PRIMARY SEXUAL CHARACTERISTICS | 8 | 216 | 0.0006459 | 0.003928 |
| 766 | DENDRITE DEVELOPMENT | 5 | 79 | 0.0006472 | 0.003931 |
| 767 | CENTRAL NERVOUS SYSTEM NEURON DIFFERENTIATION | 7 | 166 | 0.0006579 | 0.003991 |
| 768 | ION TRANSPORT | 23 | 1262 | 0.0006682 | 0.004048 |
| 769 | EMBRYONIC HEMOPOIESIS | 3 | 20 | 0.0006728 | 0.00405 |
| 770 | ICOSANOID TRANSPORT | 3 | 20 | 0.0006728 | 0.00405 |
| 771 | FATTY ACID DERIVATIVE TRANSPORT | 3 | 20 | 0.0006728 | 0.00405 |
| 772 | TRACHEA DEVELOPMENT | 3 | 20 | 0.0006728 | 0.00405 |
| 773 | PROTEIN REFOLDING | 3 | 20 | 0.0006728 | 0.00405 |
| 774 | NEGATIVE REGULATION OF LIPID METABOLIC PROCESS | 5 | 80 | 0.0006855 | 0.004121 |
| 775 | NEGATIVE REGULATION OF IMMUNE RESPONSE | 6 | 121 | 0.0006898 | 0.004142 |
| 776 | REGULATION OF AXONOGENESIS | 7 | 168 | 0.0007062 | 0.004234 |
| 777 | CELLULAR RESPONSE TO LIPID | 12 | 457 | 0.000717 | 0.004294 |
| 778 | POSITIVE REGULATION OF MYELOID CELL DIFFERENTIATION | 5 | 81 | 0.0007255 | 0.004328 |
| 779 | POSITIVE REGULATION OF PROTEIN KINASE B SIGNALING | 5 | 81 | 0.0007255 | 0.004328 |
| 780 | DEVELOPMENTAL GROWTH | 10 | 333 | 0.0007256 | 0.004328 |
| 781 | MUSCLE FIBER DEVELOPMENT | 4 | 47 | 0.0007533 | 0.004476 |
| 782 | ENDOCRINE SYSTEM DEVELOPMENT | 6 | 123 | 0.0007519 | 0.004476 |
| 783 | RESPONSE TO ANTIBIOTIC | 4 | 47 | 0.0007533 | 0.004476 |
| 784 | KIDNEY MORPHOGENESIS | 5 | 82 | 0.0007672 | 0.004553 |
| 785 | B CELL HOMEOSTASIS | 3 | 21 | 0.0007799 | 0.004611 |
| 786 | NEGATIVE REGULATION OF ORGAN GROWTH | 3 | 21 | 0.0007799 | 0.004611 |
| 787 | BONE RESORPTION | 3 | 21 | 0.0007799 | 0.004611 |
| 788 | TISSUE HOMEOSTASIS | 7 | 171 | 0.0007838 | 0.004626 |
| 789 | MONOCARBOXYLIC ACID TRANSPORT | 6 | 124 | 0.0007844 | 0.004626 |
| 790 | REGULATION OF NF KAPPAB IMPORT INTO NUCLEUS | 4 | 48 | 0.0008162 | 0.004807 |
| 791 | KIDNEY EPITHELIUM DEVELOPMENT | 6 | 125 | 0.0008181 | 0.004813 |
| 792 | REGULATION OF CELL SUBSTRATE ADHESION | 7 | 173 | 0.0008392 | 0.00493 |
| 793 | NEGATIVE REGULATION OF DEVELOPMENTAL GROWTH | 5 | 84 | 0.000856 | 0.005023 |
| 794 | REGULATION OF NEURONAL SYNAPTIC PLASTICITY | 4 | 49 | 0.0008828 | 0.00516 |
| 795 | GLOMERULUS DEVELOPMENT | 4 | 49 | 0.0008828 | 0.00516 |
| 796 | REGULATION OF STEROID BIOSYNTHETIC PROCESS | 4 | 49 | 0.0008828 | 0.00516 |
| 797 | NEGATIVE REGULATION OF CYTOKINE PRODUCTION INVOLVED IN IMMUNE RESPONSE | 3 | 22 | 0.0008973 | 0.005215 |
| 798 | POSITIVE REGULATION OF CYTOSKELETON ORGANIZATION | 7 | 175 | 0.0008977 | 0.005215 |
| 799 | REGULATION OF T CELL CYTOKINE PRODUCTION | 3 | 22 | 0.0008973 | 0.005215 |
| 800 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 3 | 22 | 0.0008973 | 0.005215 |
| 801 | SYNAPTIC TRANSMISSION GLUTAMATERGIC | 3 | 22 | 0.0008973 | 0.005215 |
| 802 | PALATE DEVELOPMENT | 5 | 85 | 0.0009031 | 0.00524 |
| 803 | DIVALENT INORGANIC CATION HOMEOSTASIS | 10 | 343 | 0.000909 | 0.005267 |
| 804 | REGULATION OF HEART RATE | 5 | 86 | 0.0009522 | 0.005504 |
| 805 | REGULATION OF CALCIUM ION DEPENDENT EXOCYTOSIS | 5 | 86 | 0.0009522 | 0.005504 |
| 806 | REGULATION OF REPRODUCTIVE PROCESS | 6 | 129 | 0.0009641 | 0.005566 |
| 807 | REGULATION OF OSSIFICATION | 7 | 178 | 0.0009914 | 0.005716 |
| 808 | REGULATION OF REGULATED SECRETORY PATHWAY | 6 | 130 | 0.001004 | 0.005779 |
| 809 | ALCOHOL METABOLIC PROCESS | 10 | 348 | 0.001014 | 0.005833 |
| 810 | POSITIVE REGULATION OF MITOTIC NUCLEAR DIVISION | 4 | 51 | 0.001027 | 0.005857 |
| 811 | POSITIVE REGULATION OF ALCOHOL BIOSYNTHETIC PROCESS | 3 | 23 | 0.001025 | 0.005857 |
| 812 | POSITIVE REGULATION OF LIPID TRANSPORT | 4 | 51 | 0.001027 | 0.005857 |
| 813 | REGULATION OF BLOOD VESSEL ENDOTHELIAL CELL MIGRATION | 4 | 51 | 0.001027 | 0.005857 |
| 814 | RESPONSE TO NICOTINE | 4 | 51 | 0.001027 | 0.005857 |
| 815 | MECHANORECEPTOR DIFFERENTIATION | 4 | 51 | 0.001027 | 0.005857 |
| 816 | POSITIVE REGULATION OF STEROID METABOLIC PROCESS | 3 | 23 | 0.001025 | 0.005857 |
| 817 | PROTEIN STABILIZATION | 6 | 131 | 0.001044 | 0.005947 |
| 818 | NEGATIVE REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 5 | 88 | 0.001056 | 0.006008 |
| 819 | DEFENSE RESPONSE | 22 | 1231 | 0.001134 | 0.006445 |
| 820 | REGULATION OF OXIDOREDUCTASE ACTIVITY | 5 | 90 | 0.001168 | 0.00659 |
| 821 | POSITIVE REGULATION OF OSTEOCLAST DIFFERENTIATION | 3 | 24 | 0.001164 | 0.00659 |
| 822 | ENDOTHELIUM DEVELOPMENT | 5 | 90 | 0.001168 | 0.00659 |
| 823 | POSITIVE REGULATION OF RECEPTOR INTERNALIZATION | 3 | 24 | 0.001164 | 0.00659 |
| 824 | REGULATION OF LONG TERM NEURONAL SYNAPTIC PLASTICITY | 3 | 24 | 0.001164 | 0.00659 |
| 825 | MESONEPHROS DEVELOPMENT | 5 | 90 | 0.001168 | 0.00659 |
| 826 | MESENCHYMAL CELL DIFFERENTIATION | 6 | 134 | 0.001174 | 0.006612 |
| 827 | CYTOSOLIC CALCIUM ION TRANSPORT | 4 | 53 | 0.001187 | 0.006672 |
| 828 | CELLULAR RESPONSE TO AMINO ACID STIMULUS | 4 | 53 | 0.001187 | 0.006672 |
| 829 | CELL JUNCTION ORGANIZATION | 7 | 185 | 0.00124 | 0.006958 |
| 830 | SENSORY ORGAN MORPHOGENESIS | 8 | 239 | 0.001241 | 0.006958 |
| 831 | TRANSMISSION OF NERVE IMPULSE | 4 | 54 | 0.001274 | 0.007115 |
| 832 | B CELL RECEPTOR SIGNALING PATHWAY | 4 | 54 | 0.001274 | 0.007115 |
| 833 | CELL CYCLE PROCESS | 20 | 1081 | 0.001273 | 0.007115 |
| 834 | EPITHELIAL CELL DEVELOPMENT | 7 | 186 | 0.001279 | 0.007128 |
| 835 | REGULATION OF EXOCYTOSIS | 7 | 186 | 0.001279 | 0.007128 |
| 836 | MITOCHONDRIAL MEMBRANE ORGANIZATION | 5 | 92 | 0.001289 | 0.007175 |
| 837 | RESPONSE TO ALKALOID | 6 | 137 | 0.001315 | 0.007287 |
| 838 | CELLULAR RESPONSE TO TOXIC SUBSTANCE | 3 | 25 | 0.001314 | 0.007287 |
| 839 | POSITIVE REGULATION OF BLOOD VESSEL ENDOTHELIAL CELL MIGRATION | 3 | 25 | 0.001314 | 0.007287 |
| 840 | THYROID GLAND DEVELOPMENT | 3 | 25 | 0.001314 | 0.007287 |
| 841 | ESTABLISHMENT OF PROTEIN LOCALIZATION TO ORGANELLE | 10 | 361 | 0.001335 | 0.007385 |
| 842 | REGULATION OF POSTSYNAPTIC MEMBRANE POTENTIAL | 4 | 55 | 0.001364 | 0.007537 |
| 843 | REGULATION OF ANION TRANSPORT | 6 | 138 | 0.001365 | 0.007537 |
| 844 | POSITIVE REGULATION OF PROTEOLYSIS | 10 | 363 | 0.001391 | 0.007667 |
| 845 | REGULATION OF CYTOKINE BIOSYNTHETIC PROCESS | 5 | 94 | 0.001419 | 0.007814 |
| 846 | REGULATION OF NEUROTRANSMITTER LEVELS | 7 | 190 | 0.001446 | 0.007941 |
| 847 | PHAGOCYTOSIS | 7 | 190 | 0.001446 | 0.007941 |
| 848 | SMAD PROTEIN SIGNAL TRANSDUCTION | 4 | 56 | 0.001459 | 0.007998 |
| 849 | OUTFLOW TRACT MORPHOGENESIS | 4 | 56 | 0.001459 | 0.007998 |
| 850 | NON CANONICAL WNT SIGNALING PATHWAY | 6 | 140 | 0.00147 | 0.008044 |
| 851 | REPLACEMENT OSSIFICATION | 3 | 26 | 0.001476 | 0.008044 |
| 852 | REGULATION OF T CELL DIFFERENTIATION IN THYMUS | 3 | 26 | 0.001476 | 0.008044 |
| 853 | REGULATION OF THYMOCYTE AGGREGATION | 3 | 26 | 0.001476 | 0.008044 |
| 854 | ENDOCHONDRAL OSSIFICATION | 3 | 26 | 0.001476 | 0.008044 |
| 855 | LIPID CATABOLIC PROCESS | 8 | 247 | 0.001529 | 0.008321 |
| 856 | MYELOID LEUKOCYTE DIFFERENTIATION | 5 | 96 | 0.001558 | 0.008444 |
| 857 | ENDOTHELIAL CELL MIGRATION | 4 | 57 | 0.001559 | 0.008444 |
| 858 | POSITIVE REGULATION OF CYTOKINE SECRETION | 5 | 96 | 0.001558 | 0.008444 |
| 859 | CARDIOCYTE DIFFERENTIATION | 5 | 96 | 0.001558 | 0.008444 |
| 860 | REGULATION OF INTERFERON GAMMA PRODUCTION | 5 | 97 | 0.001631 | 0.008826 |
| 861 | HIPPO SIGNALING | 3 | 27 | 0.00165 | 0.008868 |
| 862 | EXCITATORY POSTSYNAPTIC POTENTIAL | 3 | 27 | 0.00165 | 0.008868 |
| 863 | POSITIVE REGULATION OF NEUTROPHIL MIGRATION | 3 | 27 | 0.00165 | 0.008868 |
| 864 | INFLAMMATORY RESPONSE TO ANTIGENIC STIMULUS | 3 | 27 | 0.00165 | 0.008868 |
| 865 | RESPONSE TO LITHIUM ION | 3 | 27 | 0.00165 | 0.008868 |
| 866 | REGULATION OF NEUTROPHIL CHEMOTAXIS | 3 | 27 | 0.00165 | 0.008868 |
| 867 | NEGATIVE REGULATION OF WOUND HEALING | 4 | 58 | 0.001663 | 0.008905 |
| 868 | POSITIVE REGULATION OF CYTOKINE BIOSYNTHETIC PROCESS | 4 | 58 | 0.001663 | 0.008905 |
| 869 | POSITIVE REGULATION OF ANION TRANSPORT | 4 | 58 | 0.001663 | 0.008905 |
| 870 | REGULATION OF RESPONSE TO CYTOKINE STIMULUS | 6 | 144 | 0.001697 | 0.009073 |
| 871 | MYELOID LEUKOCYTE ACTIVATION | 5 | 98 | 0.001707 | 0.009119 |
| 872 | CELLULAR PROCESS INVOLVED IN REPRODUCTION IN MULTICELLULAR ORGANISM | 8 | 252 | 0.001734 | 0.009254 |
| 873 | POSITIVE REGULATION OF DNA BIOSYNTHETIC PROCESS | 4 | 59 | 0.001772 | 0.009423 |
| 874 | VASCULOGENESIS | 4 | 59 | 0.001772 | 0.009423 |
| 875 | REGULATION OF GLIAL CELL DIFFERENTIATION | 4 | 59 | 0.001772 | 0.009423 |
| 876 | POSITIVE REGULATION OF PROTEIN COMPLEX ASSEMBLY | 7 | 197 | 0.001777 | 0.009439 |
| 877 | MYELOID LEUKOCYTE MIGRATION | 5 | 99 | 0.001785 | 0.009472 |
| 878 | POSITIVE REGULATION OF ACUTE INFLAMMATORY RESPONSE | 3 | 28 | 0.001837 | 0.009722 |
| 879 | NEGATIVE REGULATION OF DENDRITE DEVELOPMENT | 3 | 28 | 0.001837 | 0.009722 |
| 880 | REGULATION OF GLUCOSE TRANSPORT | 5 | 100 | 0.001866 | 0.009857 |
| 881 | LIMBIC SYSTEM DEVELOPMENT | 5 | 100 | 0.001866 | 0.009857 |
| 882 | REGULATION OF T CELL PROLIFERATION | 6 | 147 | 0.001883 | 0.009936 |
| 883 | STEM CELL PROLIFERATION | 4 | 60 | 0.001886 | 0.009937 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | PROTEIN KINASE ACTIVITY | 59 | 640 | 3.575e-44 | 3.321e-41 |
| 2 | RECEPTOR SIGNALING PROTEIN SERINE THREONINE KINASE ACTIVITY | 31 | 92 | 1.875e-41 | 8.711e-39 |
| 3 | KINASE ACTIVITY | 62 | 842 | 1.047e-40 | 3.243e-38 |
| 4 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 62 | 992 | 1.618e-36 | 3.758e-34 |
| 5 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 46 | 445 | 2.863e-36 | 5.319e-34 |
| 6 | RECEPTOR SIGNALING PROTEIN ACTIVITY | 33 | 172 | 4.854e-35 | 7.515e-33 |
| 7 | RIBONUCLEOTIDE BINDING | 68 | 1860 | 4.15e-26 | 5.508e-24 |
| 8 | KINASE BINDING | 42 | 606 | 5.227e-26 | 6.07e-24 |
| 9 | ENZYME BINDING | 61 | 1737 | 3.179e-22 | 3.281e-20 |
| 10 | ADENYL NUCLEOTIDE BINDING | 56 | 1514 | 2.578e-21 | 2.395e-19 |
| 11 | VOLTAGE GATED CALCIUM CHANNEL ACTIVITY | 14 | 42 | 3.886e-19 | 3.282e-17 |
| 12 | MOLECULAR FUNCTION REGULATOR | 49 | 1353 | 3.906e-18 | 3.024e-16 |
| 13 | RAS GUANYL NUCLEOTIDE EXCHANGE FACTOR ACTIVITY | 23 | 228 | 4.815e-18 | 3.441e-16 |
| 14 | PHOSPHOPROTEIN PHOSPHATASE ACTIVITY | 21 | 178 | 5.706e-18 | 3.787e-16 |
| 15 | SIGNAL TRANSDUCER ACTIVITY | 55 | 1731 | 6.819e-18 | 4.223e-16 |
| 16 | MAP KINASE KINASE KINASE ACTIVITY | 11 | 22 | 1.087e-17 | 6.313e-16 |
| 17 | PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 15 | 70 | 3.487e-17 | 1.906e-15 |
| 18 | GUANYL NUCLEOTIDE EXCHANGE FACTOR ACTIVITY | 24 | 303 | 2.318e-16 | 1.196e-14 |
| 19 | MAP KINASE PHOSPHATASE ACTIVITY | 9 | 14 | 4.712e-16 | 2.304e-14 |
| 20 | HIGH VOLTAGE GATED CALCIUM CHANNEL ACTIVITY | 8 | 11 | 4.72e-15 | 2.192e-13 |
| 21 | PROTEIN TYROSINE PHOSPHATASE ACTIVITY | 15 | 103 | 1.539e-14 | 6.497e-13 |
| 22 | RECEPTOR BINDING | 46 | 1476 | 1.522e-14 | 6.497e-13 |
| 23 | GROWTH FACTOR RECEPTOR BINDING | 16 | 129 | 2.667e-14 | 1.077e-12 |
| 24 | PHOSPHATASE ACTIVITY | 21 | 275 | 4.013e-14 | 1.553e-12 |
| 25 | PROTEIN TYROSINE KINASE ACTIVITY | 17 | 176 | 2.667e-13 | 9.909e-12 |
| 26 | CALCIUM ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 15 | 128 | 4.124e-13 | 1.474e-11 |
| 27 | GROWTH FACTOR ACTIVITY | 16 | 160 | 8.045e-13 | 2.768e-11 |
| 28 | X1 PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 10 | 43 | 3.034e-12 | 1.007e-10 |
| 29 | GROWTH FACTOR BINDING | 14 | 123 | 3.848e-12 | 1.233e-10 |
| 30 | PROTEIN TYROSINE SERINE THREONINE PHOSPHATASE ACTIVITY | 10 | 45 | 4.974e-12 | 1.54e-10 |
| 31 | HYDROLASE ACTIVITY ACTING ON ESTER BONDS | 29 | 739 | 1.107e-11 | 3.234e-10 |
| 32 | PHOSPHORIC ESTER HYDROLASE ACTIVITY | 21 | 368 | 1.114e-11 | 3.234e-10 |
| 33 | VOLTAGE GATED CATION CHANNEL ACTIVITY | 14 | 134 | 1.254e-11 | 3.53e-10 |
| 34 | PHOSPHATIDYLINOSITOL KINASE ACTIVITY | 10 | 51 | 1.904e-11 | 5.203e-10 |
| 35 | DIVALENT INORGANIC CATION TRANSMEMBRANE TRANSPORTER ACTIVITY | 15 | 167 | 2.038e-11 | 5.411e-10 |
| 36 | FIBROBLAST GROWTH FACTOR RECEPTOR BINDING | 8 | 28 | 7.834e-11 | 2.022e-09 |
| 37 | MITOGEN ACTIVATED PROTEIN KINASE KINASE KINASE BINDING | 7 | 18 | 1.022e-10 | 2.565e-09 |
| 38 | VOLTAGE GATED ION CHANNEL ACTIVITY | 15 | 190 | 1.286e-10 | 3.145e-09 |
| 39 | PHOSPHOLIPASE A2 ACTIVITY | 8 | 31 | 1.944e-10 | 4.632e-09 |
| 40 | TRANSMEMBRANE RECEPTOR PROTEIN KINASE ACTIVITY | 10 | 81 | 2.234e-09 | 5.187e-08 |
| 41 | MAGNESIUM ION BINDING | 14 | 199 | 2.463e-09 | 5.582e-08 |
| 42 | CYTOKINE RECEPTOR BINDING | 15 | 271 | 1.705e-08 | 3.772e-07 |
| 43 | DIACYLGLYCEROL BINDING | 5 | 11 | 2.145e-08 | 4.633e-07 |
| 44 | CALCIUM DEPENDENT PROTEIN KINASE ACTIVITY | 5 | 12 | 3.65e-08 | 7.707e-07 |
| 45 | PROTEIN SERINE THREONINE TYROSINE KINASE ACTIVITY | 7 | 39 | 4.229e-08 | 8.731e-07 |
| 46 | CATION CHANNEL ACTIVITY | 15 | 298 | 5.993e-08 | 1.21e-06 |
| 47 | PROTEIN DIMERIZATION ACTIVITY | 30 | 1149 | 6.905e-08 | 1.365e-06 |
| 48 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE ACTIVITY | 8 | 64 | 8.535e-08 | 1.652e-06 |
| 49 | MAP KINASE ACTIVITY | 5 | 14 | 9.097e-08 | 1.725e-06 |
| 50 | PLATELET DERIVED GROWTH FACTOR RECEPTOR BINDING | 5 | 15 | 1.355e-07 | 2.517e-06 |
| 51 | GATED CHANNEL ACTIVITY | 15 | 325 | 1.85e-07 | 3.369e-06 |
| 52 | MITOGEN ACTIVATED PROTEIN KINASE KINASE BINDING | 5 | 16 | 1.957e-07 | 3.496e-06 |
| 53 | MACROMOLECULAR COMPLEX BINDING | 32 | 1399 | 4.764e-07 | 8.35e-06 |
| 54 | IDENTICAL PROTEIN BINDING | 29 | 1209 | 6.873e-07 | 1.182e-05 |
| 55 | ENZYME ACTIVATOR ACTIVITY | 17 | 471 | 8.85e-07 | 1.495e-05 |
| 56 | PHOSPHOLIPASE ACTIVITY | 8 | 94 | 1.718e-06 | 2.85e-05 |
| 57 | PLATELET DERIVED GROWTH FACTOR BINDING | 4 | 11 | 1.782e-06 | 2.904e-05 |
| 58 | MAP KINASE KINASE ACTIVITY | 4 | 12 | 2.654e-06 | 4.251e-05 |
| 59 | ENZYME REGULATOR ACTIVITY | 24 | 959 | 3.439e-06 | 5.414e-05 |
| 60 | PROTEIN HETERODIMERIZATION ACTIVITY | 16 | 468 | 3.778e-06 | 5.849e-05 |
| 61 | METAL ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 15 | 417 | 4.172e-06 | 6.354e-05 |
| 62 | NEUROTROPHIN RECEPTOR BINDING | 4 | 14 | 5.295e-06 | 7.933e-05 |
| 63 | KINASE REGULATOR ACTIVITY | 10 | 186 | 5.77e-06 | 8.508e-05 |
| 64 | PROTEIN COMPLEX BINDING | 23 | 935 | 7.426e-06 | 0.0001078 |
| 65 | LIPASE ACTIVITY | 8 | 117 | 8.874e-06 | 0.0001268 |
| 66 | PROTEIN DOMAIN SPECIFIC BINDING | 18 | 624 | 9.681e-06 | 0.0001363 |
| 67 | HEPARIN BINDING | 9 | 157 | 1.021e-05 | 0.0001416 |
| 68 | GTPASE ACTIVITY | 11 | 246 | 1.136e-05 | 0.0001551 |
| 69 | GLYCOSAMINOGLYCAN BINDING | 10 | 205 | 1.354e-05 | 0.0001824 |
| 70 | PHOSPHOPROTEIN BINDING | 6 | 60 | 1.394e-05 | 0.000185 |
| 71 | PASSIVE TRANSMEMBRANE TRANSPORTER ACTIVITY | 15 | 464 | 1.489e-05 | 0.0001949 |
| 72 | KINASE ACTIVATOR ACTIVITY | 6 | 62 | 1.687e-05 | 0.0002177 |
| 73 | CHANNEL REGULATOR ACTIVITY | 8 | 131 | 2.029e-05 | 0.0002583 |
| 74 | CARBOXYLIC ESTER HYDROLASE ACTIVITY | 8 | 135 | 2.523e-05 | 0.0003167 |
| 75 | CALMODULIN BINDING | 9 | 179 | 2.905e-05 | 0.0003598 |
| 76 | CYSTEINE TYPE ENDOPEPTIDASE REGULATOR ACTIVITY INVOLVED IN APOPTOTIC PROCESS | 5 | 42 | 3.17e-05 | 0.0003875 |
| 77 | SMAD BINDING | 6 | 72 | 3.989e-05 | 0.0004813 |
| 78 | GUANYL NUCLEOTIDE BINDING | 13 | 390 | 4.109e-05 | 0.0004895 |
| 79 | SIGNALING ADAPTOR ACTIVITY | 6 | 74 | 4.662e-05 | 0.0005483 |
| 80 | PROTEIN PHOSPHORYLATED AMINO ACID BINDING | 4 | 24 | 5.249e-05 | 0.0006095 |
| 81 | TUMOR NECROSIS FACTOR RECEPTOR SUPERFAMILY BINDING | 5 | 47 | 5.518e-05 | 0.0006329 |
| 82 | INORGANIC CATION TRANSMEMBRANE TRANSPORTER ACTIVITY | 15 | 527 | 6.416e-05 | 0.0007269 |
| 83 | PROTEIN HOMODIMERIZATION ACTIVITY | 18 | 722 | 6.589e-05 | 0.0007375 |
| 84 | CHEMOATTRACTANT ACTIVITY | 4 | 27 | 8.493e-05 | 0.0009393 |
| 85 | UBIQUITIN LIKE PROTEIN LIGASE BINDING | 10 | 264 | 0.0001149 | 0.001256 |
| 86 | TUMOR NECROSIS FACTOR RECEPTOR BINDING | 4 | 30 | 0.0001299 | 0.001404 |
| 87 | TRANSCRIPTION FACTOR BINDING | 14 | 524 | 0.0002178 | 0.002326 |
| 88 | SULFUR COMPOUND BINDING | 9 | 234 | 0.0002258 | 0.002357 |
| 89 | THIOESTERASE BINDING | 3 | 14 | 0.0002233 | 0.002357 |
| 90 | PROTEIN SERINE THREONINE PHOSPHATASE ACTIVITY | 5 | 64 | 0.0002433 | 0.002511 |
| 91 | HISTONE DEACETYLASE BINDING | 6 | 105 | 0.000324 | 0.003308 |
| 92 | CALCIUM CHANNEL REGULATOR ACTIVITY | 4 | 38 | 0.0003314 | 0.003347 |
| 93 | TRANSFORMING GROWTH FACTOR BETA BINDING | 3 | 16 | 0.0003391 | 0.003351 |
| 94 | PROTEIN KINASE C ACTIVITY | 3 | 16 | 0.0003391 | 0.003351 |
| 95 | CATION TRANSMEMBRANE TRANSPORTER ACTIVITY | 15 | 622 | 0.0003875 | 0.003789 |
| 96 | GTP DEPENDENT PROTEIN BINDING | 3 | 17 | 0.0004091 | 0.003959 |
| 97 | PROTEIN SERINE THREONINE KINASE ACTIVATOR ACTIVITY | 3 | 19 | 0.0005756 | 0.005401 |
| 98 | PHOSPHATASE BINDING | 7 | 162 | 0.0005692 | 0.005401 |
| 99 | HISTONE KINASE ACTIVITY | 3 | 19 | 0.0005756 | 0.005401 |
| 100 | PROTEIN PHOSPHATASE BINDING | 6 | 120 | 0.0006603 | 0.006135 |
| 101 | BINDING BRIDGING | 7 | 173 | 0.0008392 | 0.007719 |
| 102 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 8 | 226 | 0.000867 | 0.007896 |
| 103 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR BINDING | 4 | 50 | 0.000953 | 0.008596 |
| 104 | R SMAD BINDING | 3 | 23 | 0.001025 | 0.008985 |
| 105 | FIBROBLAST GROWTH FACTOR BINDING | 3 | 23 | 0.001025 | 0.008985 |
| 106 | CYSTEINE TYPE ENDOPEPTIDASE INHIBITOR ACTIVITY INVOLVED IN APOPTOTIC PROCESS | 3 | 23 | 0.001025 | 0.008985 |
| 107 | HEAT SHOCK PROTEIN BINDING | 5 | 89 | 0.001111 | 0.009648 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | VOLTAGE GATED CALCIUM CHANNEL COMPLEX | 15 | 40 | 2.438e-21 | 1.424e-18 |
| 2 | CALCIUM CHANNEL COMPLEX | 15 | 62 | 4.778e-18 | 1.395e-15 |
| 3 | PLASMA MEMBRANE PROTEIN COMPLEX | 27 | 510 | 5.862e-14 | 1.141e-11 |
| 4 | CATION CHANNEL COMPLEX | 15 | 167 | 2.038e-11 | 2.976e-09 |
| 5 | SARCOLEMMA | 12 | 125 | 1.031e-09 | 1.204e-07 |
| 6 | NEURON PROJECTION | 28 | 942 | 1.332e-08 | 1.297e-06 |
| 7 | MEMBRANE PROTEIN COMPLEX | 29 | 1020 | 1.886e-08 | 1.573e-06 |
| 8 | CYTOPLASMIC SIDE OF MEMBRANE | 12 | 170 | 3.405e-08 | 2.486e-06 |
| 9 | PERINUCLEAR REGION OF CYTOPLASM | 22 | 642 | 4.88e-08 | 2.597e-06 |
| 10 | NEURON PART | 32 | 1265 | 4.892e-08 | 2.597e-06 |
| 11 | SIDE OF MEMBRANE | 18 | 428 | 4.282e-08 | 2.597e-06 |
| 12 | SOMATODENDRITIC COMPARTMENT | 22 | 650 | 6.062e-08 | 2.95e-06 |
| 13 | CELL SUBSTRATE JUNCTION | 17 | 398 | 8.309e-08 | 3.733e-06 |
| 14 | T TUBULE | 7 | 45 | 1.194e-07 | 4.98e-06 |
| 15 | TRANSPORTER COMPLEX | 15 | 321 | 1.577e-07 | 6.138e-06 |
| 16 | PROTEIN KINASE COMPLEX | 8 | 90 | 1.232e-06 | 4.498e-05 |
| 17 | ANCHORING JUNCTION | 17 | 489 | 1.477e-06 | 5.075e-05 |
| 18 | CELL PROJECTION | 36 | 1786 | 1.664e-06 | 5.4e-05 |
| 19 | IKAPPAB KINASE COMPLEX | 4 | 11 | 1.782e-06 | 5.476e-05 |
| 20 | DENDRITE | 16 | 451 | 2.353e-06 | 6.872e-05 |
| 21 | CELL BODY | 16 | 494 | 7.472e-06 | 0.0002078 |
| 22 | CELL JUNCTION | 26 | 1151 | 8.213e-06 | 0.000218 |
| 23 | MEMBRANE MICRODOMAIN | 12 | 288 | 9.21e-06 | 0.0002339 |
| 24 | MEMBRANE REGION | 25 | 1134 | 1.885e-05 | 0.0004587 |
| 25 | RECEPTOR COMPLEX | 12 | 327 | 3.24e-05 | 0.000757 |
| 26 | ACTIN FILAMENT | 6 | 70 | 3.396e-05 | 0.0007629 |
| 27 | VACUOLE | 25 | 1180 | 3.642e-05 | 0.0007878 |
| 28 | INTRACELLULAR VESICLE | 26 | 1259 | 3.854e-05 | 0.0008038 |
| 29 | EXCITATORY SYNAPSE | 9 | 197 | 6.132e-05 | 0.001235 |
| 30 | AXON | 13 | 418 | 8.306e-05 | 0.001617 |
| 31 | CD40 RECEPTOR COMPLEX | 3 | 11 | 0.0001032 | 0.001944 |
| 32 | SYNAPSE | 18 | 754 | 0.0001139 | 0.002079 |
| 33 | EXTRACELLULAR SPACE | 26 | 1376 | 0.0001646 | 0.002913 |
| 34 | ENDOSOME | 18 | 793 | 0.0002123 | 0.003646 |
| 35 | CELL PROJECTION PART | 20 | 946 | 0.0002396 | 0.003997 |
| 36 | INTRINSIC COMPONENT OF THE CYTOPLASMIC SIDE OF THE PLASMA MEMBRANE | 3 | 15 | 0.0002773 | 0.004499 |
| 37 | SITE OF POLARIZED GROWTH | 7 | 149 | 0.0003444 | 0.005292 |
| 38 | VESICLE LUMEN | 6 | 106 | 0.000341 | 0.005292 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04010_MAPK_signaling_pathway | 175 | 268 | 0 | 0 | |
| 2 | hsa04014_Ras_signaling_pathway | 72 | 236 | 7.357e-95 | 6.622e-93 | |
| 3 | hsa04912_GnRH_signaling_pathway | 32 | 101 | 9.006e-42 | 5.403e-40 | |
| 4 | hsa04722_Neurotrophin_signaling_pathway | 34 | 127 | 1.647e-41 | 7.412e-40 | |
| 5 | hsa04370_VEGF_signaling_pathway | 29 | 76 | 1.074e-40 | 3.868e-39 | |
| 6 | hsa04664_Fc_epsilon_RI_signaling_pathway | 29 | 79 | 4.243e-40 | 1.273e-38 | |
| 7 | hsa04810_Regulation_of_actin_cytoskeleton | 38 | 214 | 5.385e-39 | 1.385e-37 | |
| 8 | hsa04380_Osteoclast_differentiation | 31 | 128 | 2.382e-36 | 5.359e-35 | |
| 9 | hsa04151_PI3K_AKT_signaling_pathway | 41 | 351 | 2.054e-34 | 4.109e-33 | |
| 10 | hsa04660_T_cell_receptor_signaling_pathway | 28 | 108 | 8.472e-34 | 1.525e-32 | |
| 11 | hsa04662_B_cell_receptor_signaling_pathway | 25 | 75 | 2.179e-33 | 3.565e-32 | |
| 12 | hsa04510_Focal_adhesion | 32 | 200 | 2.552e-31 | 3.828e-30 | |
| 13 | hsa04012_ErbB_signaling_pathway | 24 | 87 | 8.526e-30 | 1.18e-28 | |
| 14 | hsa04720_Long.term_potentiation | 22 | 70 | 8.151e-29 | 1.048e-27 | |
| 15 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 25 | 136 | 3.558e-26 | 4.27e-25 | |
| 16 | hsa04730_Long.term_depression | 20 | 70 | 2.525e-25 | 2.841e-24 | |
| 17 | hsa04210_Apoptosis | 21 | 89 | 1.382e-24 | 1.463e-23 | |
| 18 | hsa04062_Chemokine_signaling_pathway | 26 | 189 | 8.67e-24 | 8.67e-23 | |
| 19 | hsa04620_Toll.like_receptor_signaling_pathway | 21 | 102 | 3.127e-23 | 2.963e-22 | |
| 20 | hsa04270_Vascular_smooth_muscle_contraction | 20 | 116 | 1.558e-20 | 1.403e-19 | |
| 21 | hsa04540_Gap_junction | 18 | 90 | 8.151e-20 | 6.986e-19 | |
| 22 | hsa04310_Wnt_signaling_pathway | 21 | 151 | 1.75e-19 | 1.432e-18 | |
| 23 | hsa04914_Progesterone.mediated_oocyte_maturation | 16 | 87 | 4.068e-17 | 3.183e-16 | |
| 24 | hsa04910_Insulin_signaling_pathway | 18 | 138 | 2.425e-16 | 1.819e-15 | |
| 25 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 15 | 95 | 4.418e-15 | 3.181e-14 | |
| 26 | hsa04972_Pancreatic_secretion | 15 | 101 | 1.138e-14 | 7.88e-14 | |
| 27 | hsa04020_Calcium_signaling_pathway | 18 | 177 | 2.081e-14 | 1.387e-13 | |
| 28 | hsa04360_Axon_guidance | 16 | 130 | 3.019e-14 | 1.94e-13 | |
| 29 | hsa00592_alpha.Linolenic_acid_metabolism | 9 | 20 | 3.779e-14 | 2.346e-13 | |
| 30 | hsa00591_Linoleic_acid_metabolism | 9 | 30 | 2.987e-12 | 1.792e-11 | |
| 31 | hsa04621_NOD.like_receptor_signaling_pathway | 11 | 59 | 3.263e-12 | 1.895e-11 | |
| 32 | hsa00565_Ether_lipid_metabolism | 9 | 36 | 1.879e-11 | 1.057e-10 | |
| 33 | hsa04622_RIG.I.like_receptor_signaling_pathway | 11 | 71 | 2.727e-11 | 1.487e-10 | |
| 34 | hsa04260_Cardiac_muscle_contraction | 11 | 77 | 6.8e-11 | 3.6e-10 | |
| 35 | hsa04975_Fat_digestion_and_absorption | 9 | 46 | 2.04e-10 | 1.049e-09 | |
| 36 | hsa04144_Endocytosis | 15 | 203 | 3.266e-10 | 1.633e-09 | |
| 37 | hsa04114_Oocyte_meiosis | 12 | 114 | 3.512e-10 | 1.708e-09 | |
| 38 | hsa04920_Adipocytokine_signaling_pathway | 10 | 68 | 3.812e-10 | 1.806e-09 | |
| 39 | hsa04520_Adherens_junction | 10 | 73 | 7.846e-10 | 3.621e-09 | |
| 40 | hsa00590_Arachidonic_acid_metabolism | 9 | 59 | 2.111e-09 | 9.499e-09 | |
| 41 | hsa04150_mTOR_signaling_pathway | 8 | 52 | 1.586e-08 | 6.965e-08 | |
| 42 | hsa04916_Melanogenesis | 10 | 101 | 1.966e-08 | 8.424e-08 | |
| 43 | hsa04530_Tight_junction | 11 | 133 | 2.517e-08 | 1.054e-07 | |
| 44 | hsa00564_Glycerophospholipid_metabolism | 9 | 80 | 3.329e-08 | 1.362e-07 | |
| 45 | hsa04320_Dorso.ventral_axis_formation | 6 | 25 | 6.354e-08 | 2.541e-07 | |
| 46 | hsa04350_TGF.beta_signaling_pathway | 7 | 85 | 9.68e-06 | 3.788e-05 | |
| 47 | hsa04960_Aldosterone.regulated_sodium_reabsorption | 5 | 42 | 3.17e-05 | 0.0001214 | |
| 48 | hsa04670_Leukocyte_transendothelial_migration | 7 | 117 | 7.694e-05 | 0.0002885 | |
| 49 | hsa04141_Protein_processing_in_endoplasmic_reticulum | 8 | 168 | 0.0001186 | 0.0004357 | |
| 50 | hsa04110_Cell_cycle | 7 | 128 | 0.0001353 | 0.0004872 | |
| 51 | hsa04115_p53_signaling_pathway | 5 | 69 | 0.0003462 | 0.001222 | |
| 52 | hsa04390_Hippo_signaling_pathway | 7 | 154 | 0.0004203 | 0.001455 | |
| 53 | hsa04971_Gastric_acid_secretion | 5 | 74 | 0.0004791 | 0.001627 | |
| 54 | hsa04973_Carbohydrate_digestion_and_absorption | 4 | 44 | 0.0005851 | 0.00195 | |
| 55 | hsa04640_Hematopoietic_cell_lineage | 5 | 88 | 0.001056 | 0.003457 | |
| 56 | hsa04970_Salivary_secretion | 5 | 89 | 0.001111 | 0.003572 | |
| 57 | hsa04623_Cytosolic_DNA.sensing_pathway | 4 | 56 | 0.001459 | 0.004608 | |
| 58 | hsa04612_Antigen_processing_and_presentation | 4 | 78 | 0.004885 | 0.01516 | |
| 59 | hsa04742_Taste_transduction | 3 | 52 | 0.01063 | 0.03244 | |
| 60 | hsa04630_Jak.STAT_signaling_pathway | 5 | 155 | 0.01181 | 0.03544 | |
| 61 | hsa04070_Phosphatidylinositol_signaling_system | 3 | 78 | 0.03106 | 0.09167 | |
| 62 | hsa04962_Vasopressin.regulated_water_reabsorption | 2 | 44 | 0.05668 | 0.1646 | |
| 63 | hsa04340_Hedgehog_signaling_pathway | 2 | 56 | 0.08629 | 0.2465 | |
| 64 | hsa03040_Spliceosome | 3 | 128 | 0.1023 | 0.2876 | |
| 65 | hsa04976_Bile_secretion | 2 | 71 | 0.1282 | 0.3549 | |
| 66 | hsa04120_Ubiquitin_mediated_proteolysis | 2 | 139 | 0.3438 | 0.9238 | |
| 67 | hsa04145_Phagosome | 2 | 156 | 0.3971 | 1 | |
| 68 | hsa04740_Olfactory_transduction | 2 | 388 | 0.8564 | 1 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | RP11-1024P17.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-9-5p | 20 | TGFBR2 | Sponge network | -2.062 | 0 | -1.915 | 0 | 0.762 |
| 2 | TBX5-AS1 |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-3p | 13 | CACNA1C | Sponge network | -2.108 | 0 | -1.133 | 2.0E-5 | 0.692 |
| 3 | TBX5-AS1 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 21 | PDGFRA | Sponge network | -2.108 | 0 | -0.92 | 0.00039 | 0.659 |
| 4 | LINC00702 |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-589-3p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-3p;hsa-miR-96-5p | 15 | CACNA1C | Sponge network | -2.856 | 0 | -1.133 | 2.0E-5 | 0.633 |
| 5 | DNM3OS |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 11 | PDGFRA | Sponge network | 0.053 | 0.85755 | -0.92 | 0.00039 | 0.612 |
| 6 | LINC00968 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-93-5p | 20 | TGFBR2 | Sponge network | -4.19 | 0 | -1.915 | 0 | 0.609 |
| 7 | LINC00702 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-339-5p;hsa-miR-33a-5p;hsa-miR-34a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 29 | PDGFRA | Sponge network | -2.856 | 0 | -0.92 | 0.00039 | 0.589 |
| 8 | TBX5-AS1 |
hsa-miR-107;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-501-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 10 | FGF7 | Sponge network | -2.108 | 0 | -1.351 | 0 | 0.587 |
| 9 | RP11-389C8.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-93-5p | 18 | TGFBR2 | Sponge network | -2.039 | 0 | -1.915 | 0 | 0.579 |
| 10 | TBX5-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-92a-3p | 19 | TGFBR2 | Sponge network | -2.108 | 0 | -1.915 | 0 | 0.563 |
| 11 | AC109642.1 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-96-5p | 17 | PDGFRA | Sponge network | -2.791 | 0 | -0.92 | 0.00039 | 0.557 |
| 12 | GAS6-AS2 |
hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-5p | 12 | FGF2 | Sponge network | -1.761 | 0 | -2.545 | 0 | 0.556 |
| 13 | RP11-354E11.2 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-96-5p | 15 | MEF2C | Sponge network | -2.138 | 0 | -0.862 | 5.0E-5 | 0.554 |
| 14 | AC109642.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 19 | TGFBR2 | Sponge network | -2.791 | 0 | -1.915 | 0 | 0.551 |
| 15 | RP11-399O19.9 |
hsa-miR-107;hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-429;hsa-miR-7-1-3p | 12 | MEF2C | Sponge network | -0.873 | 0.00072 | -0.862 | 5.0E-5 | 0.549 |
| 16 | RP11-284N8.3 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-331-5p;hsa-miR-425-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-96-5p | 17 | MEF2C | Sponge network | -0.761 | 0.05061 | -0.862 | 5.0E-5 | 0.542 |
| 17 | RP11-389C8.2 |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-421;hsa-miR-542-3p;hsa-miR-589-3p;hsa-miR-7-1-3p | 11 | CACNA1C | Sponge network | -2.039 | 0 | -1.133 | 2.0E-5 | 0.54 |
| 18 | AC011899.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181d-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-708-5p | 12 | MAP3K3 | Sponge network | -2.611 | 0 | -1.148 | 0 | 0.537 |
| 19 | RP11-389C8.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 11 | MAP3K3 | Sponge network | -2.039 | 0 | -1.148 | 0 | 0.535 |
| 20 | MAGI2-AS3 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-429;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 24 | PDGFRA | Sponge network | -1.892 | 0 | -0.92 | 0.00039 | 0.534 |
| 21 | LINC00702 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 22 | TGFBR2 | Sponge network | -2.856 | 0 | -1.915 | 0 | 0.534 |
| 22 | MAGI2-AS3 |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-3p | 12 | CACNA1C | Sponge network | -1.892 | 0 | -1.133 | 2.0E-5 | 0.528 |
| 23 | LINC00702 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-93-3p | 18 | FGF2 | Sponge network | -2.856 | 0 | -2.545 | 0 | 0.527 |
| 24 | MAGI2-AS3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 21 | TGFBR2 | Sponge network | -1.892 | 0 | -1.915 | 0 | 0.522 |
| 25 | SH3RF3-AS1 |
hsa-miR-130b-3p;hsa-miR-16-1-3p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-26b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 10 | PDGFRA | Sponge network | -1.583 | 0 | -0.92 | 0.00039 | 0.516 |
| 26 | MAGI2-AS3 |
hsa-miR-107;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-16-2-3p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-429;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 10 | FGF7 | Sponge network | -1.892 | 0 | -1.351 | 0 | 0.514 |
| 27 | RP11-720L2.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -2.305 | 0 | -1.915 | 0 | 0.509 |
| 28 | FENDRR |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-93-5p | 15 | TGFBR2 | Sponge network | -4.222 | 0 | -1.915 | 0 | 0.506 |
| 29 | LINC00702 |
hsa-miR-107;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 11 | FGF7 | Sponge network | -2.856 | 0 | -1.351 | 0 | 0.505 |
| 30 | TBX5-AS1 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-590-5p;hsa-miR-93-3p | 15 | FGF2 | Sponge network | -2.108 | 0 | -2.545 | 0 | 0.504 |
| 31 | LINC00968 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-590-5p;hsa-miR-93-3p | 12 | FGF2 | Sponge network | -4.19 | 0 | -2.545 | 0 | 0.504 |
| 32 | LINC00702 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 12 | MAP3K3 | Sponge network | -2.856 | 0 | -1.148 | 0 | 0.5 |
| 33 | MAGI2-AS3 |
hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-503-5p;hsa-miR-93-3p | 15 | FGF2 | Sponge network | -1.892 | 0 | -2.545 | 0 | 0.499 |
| 34 | LINC00968 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-708-5p;hsa-miR-93-5p;hsa-miR-96-5p | 14 | MAP3K3 | Sponge network | -4.19 | 0 | -1.148 | 0 | 0.499 |
| 35 | LINC00961 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-9-5p | 12 | TGFBR2 | Sponge network | -2.724 | 0 | -1.915 | 0 | 0.497 |
| 36 | AC109642.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-93-5p;hsa-miR-96-5p | 13 | MAP3K3 | Sponge network | -2.791 | 0 | -1.148 | 0 | 0.497 |
| 37 | MIR497HG |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-589-3p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-3p;hsa-miR-96-5p | 12 | CACNA1C | Sponge network | -2.142 | 0 | -1.133 | 2.0E-5 | 0.494 |
| 38 | FENDRR |
hsa-miR-148b-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-542-3p;hsa-miR-7-1-3p;hsa-miR-93-3p;hsa-miR-96-5p | 11 | CACNA1C | Sponge network | -4.222 | 0 | -1.133 | 2.0E-5 | 0.494 |
| 39 | LINC00968 |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-93-3p;hsa-miR-96-5p | 11 | CACNA1C | Sponge network | -4.19 | 0 | -1.133 | 2.0E-5 | 0.492 |
| 40 | FENDRR |
hsa-miR-142-3p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-93-3p | 10 | FGF2 | Sponge network | -4.222 | 0 | -2.545 | 0 | 0.489 |
| 41 | MAGI2-AS3 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-9-5p;hsa-miR-92a-3p | 19 | MEF2C | Sponge network | -1.892 | 0 | -0.862 | 5.0E-5 | 0.487 |
| 42 | LINC00968 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-96-5p | 18 | MEF2C | Sponge network | -4.19 | 0 | -0.862 | 5.0E-5 | 0.48 |
| 43 | LINC00472 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -2.952 | 0 | -1.915 | 0 | 0.479 |
| 44 | TBX5-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-7-1-3p | 12 | MAP3K3 | Sponge network | -2.108 | 0 | -1.148 | 0 | 0.476 |
| 45 | TBX5-AS1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-324-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-550a-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-744-3p;hsa-miR-9-5p;hsa-miR-92a-3p | 21 | MEF2C | Sponge network | -2.108 | 0 | -0.862 | 5.0E-5 | 0.473 |
| 46 | AC109642.1 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-32-5p;hsa-miR-503-5p;hsa-miR-93-3p | 12 | FGF2 | Sponge network | -2.791 | 0 | -2.545 | 0 | 0.467 |
| 47 | RP11-1024P17.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p | 10 | MAP3K3 | Sponge network | -2.062 | 0 | -1.148 | 0 | 0.461 |
| 48 | RP11-166D19.1 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-429;hsa-miR-454-3p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-96-5p | 21 | PDGFRA | Sponge network | -0.582 | 0.05253 | -0.92 | 0.00039 | 0.459 |
| 49 | RP11-389C8.2 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-2355-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-429;hsa-miR-550a-3p;hsa-miR-590-3p;hsa-miR-7-1-3p | 17 | MEF2C | Sponge network | -2.039 | 0 | -0.862 | 5.0E-5 | 0.454 |
| 50 | LINC00472 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -2.952 | 0 | -1.148 | 0 | 0.453 |
| 51 | BZRAP1-AS1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-2355-5p;hsa-miR-335-3p;hsa-miR-425-5p | 11 | MEF2C | Sponge network | -0.785 | 0.00723 | -0.862 | 5.0E-5 | 0.452 |
| 52 | GAS6-AS2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-429;hsa-miR-590-5p;hsa-miR-7-1-3p | 20 | PDGFRA | Sponge network | -1.761 | 0 | -0.92 | 0.00039 | 0.451 |
| 53 | GAS6-AS2 |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-7-1-3p | 10 | CACNA1C | Sponge network | -1.761 | 0 | -1.133 | 2.0E-5 | 0.448 |
| 54 | AC011899.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 14 | TGFBR2 | Sponge network | -2.611 | 0 | -1.915 | 0 | 0.448 |
| 55 | AC109642.1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-590-3p;hsa-miR-744-3p;hsa-miR-9-5p;hsa-miR-92a-3p;hsa-miR-96-5p | 17 | MEF2C | Sponge network | -2.791 | 0 | -0.862 | 5.0E-5 | 0.447 |
| 56 | RP11-1008C21.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-92a-3p;hsa-miR-93-5p | 16 | TGFBR2 | Sponge network | -1.249 | 0 | -1.915 | 0 | 0.446 |
| 57 | RP11-378A13.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-93-5p | 17 | TGFBR2 | Sponge network | -1.713 | 0 | -1.915 | 0 | 0.446 |
| 58 | LINC00702 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-2355-5p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-550a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-744-3p;hsa-miR-92a-3p;hsa-miR-96-5p | 24 | MEF2C | Sponge network | -2.856 | 0 | -0.862 | 5.0E-5 | 0.445 |
| 59 | RP11-399O19.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-93-5p | 15 | TGFBR2 | Sponge network | -0.873 | 0.00072 | -1.915 | 0 | 0.445 |
| 60 | MIR497HG |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-2355-5p;hsa-miR-324-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-9-5p;hsa-miR-96-5p | 17 | MEF2C | Sponge network | -2.142 | 0 | -0.862 | 5.0E-5 | 0.444 |
| 61 | PCED1B-AS1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-2355-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-744-3p;hsa-miR-92a-3p;hsa-miR-96-5p | 14 | MEF2C | Sponge network | -0.672 | 0.02084 | -0.862 | 5.0E-5 | 0.444 |
| 62 | LINC00968 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-339-5p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-96-5p | 22 | PDGFRA | Sponge network | -4.19 | 0 | -0.92 | 0.00039 | 0.443 |
| 63 | CTD-2013N24.2 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-429 | 12 | FGF2 | Sponge network | -1.745 | 0 | -2.545 | 0 | 0.441 |
| 64 | RP5-1042I8.7 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-320b;hsa-miR-331-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-744-3p;hsa-miR-92a-3p;hsa-miR-96-5p | 14 | MEF2C | Sponge network | -0.733 | 0.00018 | -0.862 | 5.0E-5 | 0.44 |
| 65 | AC079630.4 |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-320b;hsa-miR-331-5p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-9-5p | 14 | MEF2C | Sponge network | -3.758 | 0 | -0.862 | 5.0E-5 | 0.437 |
| 66 | AF131215.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-9-5p | 12 | TGFBR2 | Sponge network | -2.09 | 0 | -1.915 | 0 | 0.435 |
| 67 | AC079630.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-93-5p | 16 | TGFBR2 | Sponge network | -3.758 | 0 | -1.915 | 0 | 0.433 |
| 68 | RP11-1024P17.1 |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-33a-5p;hsa-miR-589-3p;hsa-miR-93-3p | 10 | CACNA1C | Sponge network | -2.062 | 0 | -1.133 | 2.0E-5 | 0.431 |
| 69 | RP11-389C8.2 |
hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-589-3p | 11 | FGF2 | Sponge network | -2.039 | 0 | -2.545 | 0 | 0.428 |
| 70 | LINC00472 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p | 10 | NTRK2 | Sponge network | -2.952 | 0 | -2.564 | 0 | 0.427 |
| 71 | LINC00961 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-17-5p;hsa-miR-181d-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-22-3p;hsa-miR-28-5p;hsa-miR-708-5p;hsa-miR-96-5p | 12 | MAP3K3 | Sponge network | -2.724 | 0 | -1.148 | 0 | 0.427 |
| 72 | MIR497HG |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 18 | PDGFRA | Sponge network | -2.142 | 0 | -0.92 | 0.00039 | 0.426 |
| 73 | FENDRR |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-708-5p;hsa-miR-93-5p;hsa-miR-96-5p | 12 | MAP3K3 | Sponge network | -4.222 | 0 | -1.148 | 0 | 0.425 |
| 74 | CTD-2013N24.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-93-5p | 19 | TGFBR2 | Sponge network | -1.745 | 0 | -1.915 | 0 | 0.42 |
| 75 | RP11-354E11.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-96-5p | 10 | MAP3K3 | Sponge network | -2.138 | 0 | -1.148 | 0 | 0.418 |
| 76 | RP11-354E11.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-9-5p | 17 | TGFBR2 | Sponge network | -2.138 | 0 | -1.915 | 0 | 0.418 |
| 77 | SH3RF3-AS1 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-503-5p;hsa-miR-590-5p | 13 | FGF2 | Sponge network | -1.583 | 0 | -2.545 | 0 | 0.417 |
| 78 | RP11-1008C21.1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-744-3p | 12 | MEF2C | Sponge network | -1.826 | 3.0E-5 | -0.862 | 5.0E-5 | 0.415 |
| 79 | MAGI2-AS3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -1.892 | 0 | -1.148 | 0 | 0.414 |
| 80 | LINC00702 |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-21-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p | 16 | TGFB2 | Sponge network | -2.856 | 0 | -1.547 | 1.0E-5 | 0.413 |
| 81 | RP11-401P9.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -3.04 | 0 | -1.915 | 0 | 0.413 |
| 82 | WDFY3-AS2 |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-320b;hsa-miR-331-5p;hsa-miR-335-3p;hsa-miR-425-5p;hsa-miR-550a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-744-3p;hsa-miR-96-5p | 21 | MEF2C | Sponge network | -1.297 | 0 | -0.862 | 5.0E-5 | 0.413 |
| 83 | AF131215.2 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-2355-5p;hsa-miR-324-3p;hsa-miR-9-5p | 10 | MEF2C | Sponge network | -2.09 | 0 | -0.862 | 5.0E-5 | 0.411 |
| 84 | CTD-2013N24.2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 22 | PDGFRA | Sponge network | -1.745 | 0 | -0.92 | 0.00039 | 0.41 |
| 85 | RP11-476D10.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-590-3p | 12 | TGFBR2 | Sponge network | -4.519 | 0 | -1.915 | 0 | 0.41 |
| 86 | RP11-456K23.1 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 22 | PDGFRA | Sponge network | -1.488 | 0 | -0.92 | 0.00039 | 0.409 |
| 87 | AC144831.1 |
hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -2.063 | 0 | -1.148 | 0 | 0.408 |
| 88 | WDFY3-AS2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-590-5p | 16 | TGFBR2 | Sponge network | -1.297 | 0 | -1.915 | 0 | 0.407 |
| 89 | RP11-536K7.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-93-5p | 10 | PDGFRA | Sponge network | -1.239 | 5.0E-5 | -0.92 | 0.00039 | 0.407 |
| 90 | RP11-401P9.4 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-196b-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-29a-5p;hsa-miR-29b-1-5p;hsa-miR-542-3p;hsa-miR-708-3p | 10 | CACNB4 | Sponge network | -3.04 | 0 | -2.288 | 0 | 0.407 |
| 91 | RP11-399O19.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -0.873 | 0.00072 | -1.148 | 0 | 0.404 |
| 92 | SFTA1P | hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-744-3p;hsa-miR-9-5p | 10 | MEF2C | Sponge network | -3.139 | 0 | -0.862 | 5.0E-5 | 0.403 |
| 93 | RP11-476D10.1 |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-21-5p;hsa-miR-425-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-96-5p | 12 | MEF2C | Sponge network | -4.519 | 0 | -0.862 | 5.0E-5 | 0.403 |
| 94 | RP11-401P9.4 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-590-5p;hsa-miR-93-3p | 10 | FGF2 | Sponge network | -3.04 | 0 | -2.545 | 0 | 0.402 |
| 95 | LINC00968 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-421;hsa-miR-590-3p;hsa-miR-590-5p | 14 | NTRK2 | Sponge network | -4.19 | 0 | -2.564 | 0 | 0.4 |
| 96 | MIR497HG |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 14 | MAP3K3 | Sponge network | -2.142 | 0 | -1.148 | 0 | 0.393 |
| 97 | RP11-456K23.1 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-589-3p;hsa-miR-590-5p | 14 | FGF2 | Sponge network | -1.488 | 0 | -2.545 | 0 | 0.393 |
| 98 | LINC00472 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-29a-5p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p | 10 | CACNB4 | Sponge network | -2.952 | 0 | -2.288 | 0 | 0.392 |
| 99 | RP11-532F6.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-9-5p | 12 | TGFBR2 | Sponge network | -2.028 | 0 | -1.915 | 0 | 0.391 |
| 100 | AC079630.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -3.758 | 0 | -1.148 | 0 | 0.39 |
| 101 | RP11-352D13.6 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-93-5p | 10 | TGFBR2 | Sponge network | -4.634 | 0 | -1.915 | 0 | 0.39 |
| 102 | MIR22HG |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -1.704 | 0 | -1.915 | 0 | 0.39 |
| 103 | RP11-456K23.1 |
hsa-miR-106b-5p;hsa-miR-146b-5p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-29b-3p;hsa-miR-362-5p;hsa-miR-542-3p;hsa-miR-93-5p | 10 | AKT3 | Sponge network | -1.488 | 0 | -1.44 | 0 | 0.389 |
| 104 | MIAT | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-miR-125b-2-3p;hsa-miR-1266-5p;hsa-miR-143-3p;hsa-miR-195-5p;hsa-miR-2110;hsa-miR-3065-3p;hsa-miR-30a-3p;hsa-miR-30b-3p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-664a-5p | 15 | CACNA1E | Sponge network | 1.49 | 0.00047 | 1.62 | 0.0002 | 0.388 |
| 105 | RP11-720L2.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-93-5p | 10 | PDGFRA | Sponge network | -2.305 | 0 | -0.92 | 0.00039 | 0.388 |
| 106 | SNHG18 |
hsa-miR-107;hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-19a-3p;hsa-miR-200c-5p;hsa-miR-2355-5p;hsa-miR-590-3p | 10 | MEF2C | Sponge network | -1.073 | 0.00533 | -0.862 | 5.0E-5 | 0.388 |
| 107 | FENDRR |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-3p | 14 | NTRK2 | Sponge network | -4.222 | 0 | -2.564 | 0 | 0.387 |
| 108 | MIR497HG |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-93-5p | 18 | TGFBR2 | Sponge network | -2.142 | 0 | -1.915 | 0 | 0.387 |
| 109 | LL22NC03-86G7.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | MAP3K3 | Sponge network | -1.177 | 3.0E-5 | -1.148 | 0 | 0.387 |
| 110 | HHIP-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-22-3p;hsa-miR-28-5p;hsa-miR-7-1-3p | 10 | MAP3K3 | Sponge network | -2.807 | 0 | -1.148 | 0 | 0.386 |
| 111 | RP11-293M10.6 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-590-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -1.199 | 0.00063 | -1.915 | 0 | 0.386 |
| 112 | RP5-839B4.8 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -5.037 | 0 | -1.915 | 0 | 0.384 |
| 113 | RP11-1008C21.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-590-3p | 14 | TGFBR2 | Sponge network | -1.826 | 3.0E-5 | -1.915 | 0 | 0.382 |
| 114 | BDNF-AS |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-2355-5p;hsa-miR-331-5p;hsa-miR-335-3p;hsa-miR-425-5p;hsa-miR-590-3p | 10 | MEF2C | Sponge network | -0.568 | 0.02011 | -0.862 | 5.0E-5 | 0.382 |
| 115 | RP11-401P9.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-708-5p;hsa-miR-93-5p | 12 | MAP3K3 | Sponge network | -3.04 | 0 | -1.148 | 0 | 0.381 |
| 116 | GAS6-AS2 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-2355-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 17 | MEF2C | Sponge network | -1.761 | 0 | -0.862 | 5.0E-5 | 0.381 |
| 117 | AC011899.9 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-7-1-3p | 14 | MEF2C | Sponge network | -2.611 | 0 | -0.862 | 5.0E-5 | 0.38 |
| 118 | CTA-373H7.7 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-miR-1266-5p;hsa-miR-143-3p;hsa-miR-195-5p;hsa-miR-2110;hsa-miR-3065-3p;hsa-miR-30a-3p;hsa-miR-30b-3p;hsa-miR-664a-5p | 12 | CACNA1E | Sponge network | 1.49 | 0.00107 | 1.62 | 0.0002 | 0.38 |
| 119 | RP4-639F20.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-22-3p;hsa-miR-28-5p;hsa-miR-96-5p | 10 | MAP3K3 | Sponge network | -1.312 | 0 | -1.148 | 0 | 0.378 |
| 120 | RP11-389C8.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-3p | 13 | NTRK2 | Sponge network | -2.039 | 0 | -2.564 | 0 | 0.377 |
| 121 | AC003090.1 |
hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-22-3p;hsa-miR-7-1-3p;hsa-miR-708-5p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | MAP3K3 | Sponge network | -3.16 | 2.0E-5 | -1.148 | 0 | 0.376 |
| 122 | AC093627.10 |
hsa-miR-130b-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-19a-3p;hsa-miR-200c-5p;hsa-miR-324-3p;hsa-miR-421;hsa-miR-590-5p;hsa-miR-744-3p;hsa-miR-92a-3p | 10 | MEF2C | Sponge network | -0.174 | 0.61136 | -0.862 | 5.0E-5 | 0.375 |
| 123 | RP11-166D19.1 |
hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-93-3p | 11 | FGF2 | Sponge network | -0.582 | 0.05253 | -2.545 | 0 | 0.375 |
| 124 | AC007743.1 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-320b;hsa-miR-429;hsa-miR-590-5p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -2.595 | 0 | -1.915 | 0 | 0.375 |
| 125 | HHIP-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-590-3p | 14 | TGFBR2 | Sponge network | -2.807 | 0 | -1.915 | 0 | 0.374 |
| 126 | RP11-1024P17.1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-9-5p | 15 | MEF2C | Sponge network | -2.062 | 0 | -0.862 | 5.0E-5 | 0.373 |
| 127 | RP11-166D19.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-1266-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-30d-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-5p | 10 | RASGRF2 | Sponge network | -0.582 | 0.05253 | -0.287 | 0.25734 | 0.373 |
| 128 | AC007743.1 |
hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-212-3p;hsa-miR-22-3p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-708-5p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -2.595 | 0 | -1.148 | 0 | 0.373 |
| 129 | RP11-456K23.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-93-5p | 19 | TGFBR2 | Sponge network | -1.488 | 0 | -1.915 | 0 | 0.372 |
| 130 | GAS6-AS2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-5p;hsa-miR-92a-3p | 18 | TGFBR2 | Sponge network | -1.761 | 0 | -1.915 | 0 | 0.371 |
| 131 | CTD-2013N24.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -1.745 | 0 | -1.148 | 0 | 0.37 |
| 132 | LINC00702 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p | 17 | NTRK2 | Sponge network | -2.856 | 0 | -2.564 | 0 | 0.369 |
| 133 | CYP1B1-AS1 |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-2355-5p;hsa-miR-320b;hsa-miR-331-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-744-3p | 15 | MEF2C | Sponge network | -1.073 | 0.00045 | -0.862 | 5.0E-5 | 0.369 |
| 134 | RP11-532F6.3 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-550a-3p;hsa-miR-744-3p;hsa-miR-9-5p | 14 | MEF2C | Sponge network | -2.028 | 0 | -0.862 | 5.0E-5 | 0.367 |
| 135 | LINC00261 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p | 15 | TGFBR2 | Sponge network | -2.566 | 0.00025 | -1.915 | 0 | 0.367 |
| 136 | C1orf132 |
hsa-miR-132-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-196b-5p;hsa-miR-212-3p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-542-3p;hsa-miR-590-3p;hsa-miR-708-3p | 11 | CACNB4 | Sponge network | -0.86 | 0.02429 | -2.288 | 0 | 0.364 |
| 137 | RP11-532F6.3 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-576-5p | 11 | PDGFRA | Sponge network | -2.028 | 0 | -0.92 | 0.00039 | 0.363 |
| 138 | RP11-378A13.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-708-5p;hsa-miR-93-5p | 11 | MAP3K3 | Sponge network | -1.713 | 0 | -1.148 | 0 | 0.362 |
| 139 | FENDRR |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-196b-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-542-3p;hsa-miR-590-3p;hsa-miR-96-5p | 13 | CACNB4 | Sponge network | -4.222 | 0 | -2.288 | 0 | 0.361 |
| 140 | RP11-10C24.3 | hsa-miR-106a-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-5p | 11 | TGFBR2 | Sponge network | -1.013 | 0 | -1.915 | 0 | 0.361 |
| 141 | RP11-284N8.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-22-3p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 12 | MAP3K3 | Sponge network | -0.761 | 0.05061 | -1.148 | 0 | 0.36 |
| 142 | AC004947.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p | 13 | NTRK2 | Sponge network | -3.94 | 0 | -2.564 | 0 | 0.358 |
| 143 | AF131215.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-454-3p | 11 | NTRK2 | Sponge network | -2.09 | 0 | -2.564 | 0 | 0.358 |
| 144 | RP1-78O14.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p | 12 | TGFBR2 | Sponge network | -4.409 | 0 | -1.915 | 0 | 0.358 |
| 145 | CTD-2269F5.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | PDGFRA | Sponge network | -1.576 | 0.00334 | -0.92 | 0.00039 | 0.358 |
| 146 | RP11-354E11.2 |
hsa-miR-107;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-590-3p | 10 | TGFB2 | Sponge network | -2.138 | 0 | -1.547 | 1.0E-5 | 0.356 |
| 147 | RP5-839B4.8 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 11 | MAP3K3 | Sponge network | -5.037 | 0 | -1.148 | 0 | 0.355 |
| 148 | AC109642.1 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-196b-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-29a-5p;hsa-miR-29b-1-5p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-96-5p | 11 | CACNB4 | Sponge network | -2.791 | 0 | -2.288 | 0 | 0.355 |
| 149 | RP11-354E11.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-576-5p;hsa-miR-590-3p | 12 | NTRK2 | Sponge network | -2.138 | 0 | -2.564 | 0 | 0.355 |
| 150 | RP11-1024P17.1 |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-335-3p;hsa-miR-589-3p;hsa-miR-590-3p | 13 | TGFB2 | Sponge network | -2.062 | 0 | -1.547 | 1.0E-5 | 0.354 |
| 151 | NR2F1-AS1 |
hsa-miR-107;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 14 | MEF2C | Sponge network | -0.427 | 0.1559 | -0.862 | 5.0E-5 | 0.354 |
| 152 | LINC00968 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-2355-3p;hsa-miR-29a-5p;hsa-miR-590-3p;hsa-miR-9-5p | 10 | NFATC2 | Sponge network | -4.19 | 0 | -1.179 | 0.0004 | 0.353 |
| 153 | LINC00961 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-22-5p;hsa-miR-9-5p;hsa-miR-96-5p | 11 | MEF2C | Sponge network | -2.724 | 0 | -0.862 | 5.0E-5 | 0.352 |
| 154 | RP11-238K6.1 | hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-331-5p;hsa-miR-425-5p;hsa-miR-590-3p;hsa-miR-744-3p | 12 | MEF2C | Sponge network | -5.195 | 0 | -0.862 | 5.0E-5 | 0.351 |
| 155 | LINC00702 |
hsa-miR-103a-3p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-182-5p;hsa-miR-200b-3p;hsa-miR-450b-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-96-5p | 11 | CACNA2D1 | Sponge network | -2.856 | 0 | -1.269 | 0.00469 | 0.35 |
| 156 | RP11-77A13.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -6.738 | 0 | -1.915 | 0 | 0.35 |
| 157 | RP11-365O16.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -2.765 | 0.00017 | -1.915 | 0 | 0.349 |
| 158 | AC007743.1 |
hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-320b;hsa-miR-421;hsa-miR-425-5p;hsa-miR-429;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-744-3p | 14 | MEF2C | Sponge network | -2.595 | 0 | -0.862 | 5.0E-5 | 0.348 |
| 159 | PCED1B-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-93-5p;hsa-miR-96-5p | 13 | PDGFRA | Sponge network | -0.672 | 0.02084 | -0.92 | 0.00039 | 0.348 |
| 160 | LINC00920 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p | 10 | TGFBR2 | Sponge network | -0.998 | 0.0011 | -1.915 | 0 | 0.347 |
| 161 | RP11-389C8.2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-429;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 19 | PDGFRA | Sponge network | -2.039 | 0 | -0.92 | 0.00039 | 0.347 |
| 162 | AC003090.1 |
hsa-miR-130b-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-425-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-744-3p;hsa-miR-96-5p | 11 | MEF2C | Sponge network | -3.16 | 2.0E-5 | -0.862 | 5.0E-5 | 0.347 |
| 163 | RP4-639F20.1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-200c-5p;hsa-miR-22-5p;hsa-miR-320b;hsa-miR-96-5p | 10 | MEF2C | Sponge network | -1.312 | 0 | -0.862 | 5.0E-5 | 0.347 |
| 164 | RP11-88I21.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -8.789 | 0 | -1.915 | 0 | 0.346 |
| 165 | RP5-1042I8.7 |
hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-93-5p | 10 | TGFBR2 | Sponge network | -0.733 | 0.00018 | -1.915 | 0 | 0.345 |
| 166 | AC109642.1 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-590-3p | 11 | NTRK2 | Sponge network | -2.791 | 0 | -2.564 | 0 | 0.344 |
| 167 | RP11-456K23.1 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-335-3p;hsa-miR-550a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-744-3p;hsa-miR-9-5p | 14 | MEF2C | Sponge network | -1.488 | 0 | -0.862 | 5.0E-5 | 0.344 |
| 168 | RP11-517P14.2 | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.795 | 6.0E-5 | -1.915 | 0 | 0.344 |
| 169 | LIPE-AS1 |
hsa-miR-106a-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-5p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.734 | 0.00039 | -1.915 | 0 | 0.343 |
| 170 | CTD-2377D24.6 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-miR-125b-2-3p;hsa-miR-143-3p;hsa-miR-195-3p;hsa-miR-195-5p;hsa-miR-2110;hsa-miR-3065-3p;hsa-miR-30a-3p;hsa-miR-30b-3p;hsa-miR-30d-3p;hsa-miR-30e-3p | 14 | CACNA1E | Sponge network | 3.878 | 1.0E-5 | 1.62 | 0.0002 | 0.343 |
| 171 | AC109642.1 |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-590-3p | 11 | TGFB2 | Sponge network | -2.791 | 0 | -1.547 | 1.0E-5 | 0.343 |
| 172 | MAGI2-AS3 |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-193b-3p;hsa-miR-200a-3p;hsa-miR-21-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p | 14 | TGFB2 | Sponge network | -1.892 | 0 | -1.547 | 1.0E-5 | 0.342 |
| 173 | RP11-680F8.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -1.628 | 0 | -1.915 | 0 | 0.34 |
| 174 | CYP1B1-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20b-5p;hsa-miR-28-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -1.073 | 0.00045 | -1.915 | 0 | 0.34 |
| 175 | AC109642.1 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-2355-3p;hsa-miR-29a-5p;hsa-miR-590-3p;hsa-miR-9-5p | 10 | NFATC2 | Sponge network | -2.791 | 0 | -1.179 | 0.0004 | 0.339 |
| 176 | RP11-166D19.1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-429;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-96-5p | 20 | MEF2C | Sponge network | -0.582 | 0.05253 | -0.862 | 5.0E-5 | 0.339 |
| 177 | CTC-366B18.4 | hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-200c-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-425-5p;hsa-miR-590-3p | 10 | MEF2C | Sponge network | -0.652 | 0.01265 | -0.862 | 5.0E-5 | 0.338 |
| 178 | AC079630.4 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-182-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-3p | 11 | NTRK2 | Sponge network | -3.758 | 0 | -2.564 | 0 | 0.338 |
| 179 | RP11-1024P17.1 |
hsa-miR-142-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-589-3p;hsa-miR-93-3p | 13 | FGF2 | Sponge network | -2.062 | 0 | -2.545 | 0 | 0.338 |
| 180 | RP11-378A13.1 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-744-3p | 13 | MEF2C | Sponge network | -1.713 | 0 | -0.862 | 5.0E-5 | 0.337 |
| 181 | LINC00472 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-425-5p;hsa-miR-7-1-3p;hsa-miR-744-3p | 12 | MEF2C | Sponge network | -2.952 | 0 | -0.862 | 5.0E-5 | 0.335 |
| 182 | LINC01024 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-200c-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-421;hsa-miR-550a-3p;hsa-miR-590-3p;hsa-miR-7-1-3p | 11 | MEF2C | Sponge network | -0.659 | 0.02711 | -0.862 | 5.0E-5 | 0.334 |
| 183 | CTD-2013N24.2 |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-21-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p | 13 | TGFB2 | Sponge network | -1.745 | 0 | -1.547 | 1.0E-5 | 0.334 |
| 184 | MAGI2-AS3 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p | 16 | NTRK2 | Sponge network | -1.892 | 0 | -2.564 | 0 | 0.333 |
| 185 | RP11-1008C21.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-22-3p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -1.249 | 0 | -1.148 | 0 | 0.329 |
| 186 | RP11-389C8.2 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-196b-5p;hsa-miR-20a-3p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-542-3p;hsa-miR-589-3p;hsa-miR-590-3p | 10 | CACNB4 | Sponge network | -2.039 | 0 | -2.288 | 0 | 0.328 |
| 187 | LINC00968 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-196b-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-29a-5p;hsa-miR-29b-1-5p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-590-3p;hsa-miR-96-5p | 12 | CACNB4 | Sponge network | -4.19 | 0 | -2.288 | 0 | 0.327 |
| 188 | CASC2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p | 12 | NTRK2 | Sponge network | -1.086 | 0 | -2.564 | 0 | 0.326 |
| 189 | RP11-456K23.1 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-196b-5p;hsa-miR-20a-3p;hsa-miR-222-3p;hsa-miR-29a-5p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-542-3p;hsa-miR-589-3p;hsa-miR-590-3p | 13 | CACNB4 | Sponge network | -1.488 | 0 | -2.288 | 0 | 0.325 |
| 190 | MIR497HG |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-5p | 10 | NTRK2 | Sponge network | -2.142 | 0 | -2.564 | 0 | 0.325 |
| 191 | LINC00607 | hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-320b;hsa-miR-331-5p;hsa-miR-429;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-744-3p | 13 | MEF2C | Sponge network | -2.277 | 0 | -0.862 | 5.0E-5 | 0.324 |
| 192 | RP11-354E11.2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-96-5p | 16 | PDGFRA | Sponge network | -2.138 | 0 | -0.92 | 0.00039 | 0.323 |
| 193 | RP11-1008C21.2 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-320b;hsa-miR-92a-3p | 12 | MEF2C | Sponge network | -1.249 | 0 | -0.862 | 5.0E-5 | 0.322 |
| 194 | FGF14-AS2 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-331-5p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-96-5p | 12 | MEF2C | Sponge network | -2.159 | 0 | -0.862 | 5.0E-5 | 0.322 |
| 195 | RP11-88I21.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | MAP3K3 | Sponge network | -8.789 | 0 | -1.148 | 0 | 0.321 |
| 196 | RP11-401P9.4 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-93-5p | 16 | PDGFRA | Sponge network | -3.04 | 0 | -0.92 | 0.00039 | 0.32 |
| 197 | LL22NC03-86G7.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -1.177 | 3.0E-5 | -1.915 | 0 | 0.32 |
| 198 | TBX5-AS1 |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-335-3p;hsa-miR-590-5p | 12 | TGFB2 | Sponge network | -2.108 | 0 | -1.547 | 1.0E-5 | 0.32 |
| 199 | AC004947.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-7-1-3p;hsa-miR-708-5p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -3.94 | 0 | -1.148 | 0 | 0.318 |
| 200 | DIO3OS | hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-2355-5p;hsa-miR-324-3p;hsa-miR-331-5p;hsa-miR-550a-3p | 11 | MEF2C | Sponge network | -1.936 | 0.00085 | -0.862 | 5.0E-5 | 0.316 |
| 201 | RP11-77A13.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | MAP3K3 | Sponge network | -6.738 | 0 | -1.148 | 0 | 0.315 |
| 202 | AC011899.9 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-26b-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 15 | PDGFRA | Sponge network | -2.611 | 0 | -0.92 | 0.00039 | 0.315 |
| 203 | PSMG3-AS1 |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-200c-5p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-590-3p | 11 | MEF2C | Sponge network | -0.522 | 0.09584 | -0.862 | 5.0E-5 | 0.313 |
| 204 | LINC00443 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p | 11 | TGFBR2 | Sponge network | -3.704 | 0.0003 | -1.915 | 0 | 0.311 |
| 205 | RP11-378A13.1 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-29a-5p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-708-3p | 11 | CACNB4 | Sponge network | -1.713 | 0 | -2.288 | 0 | 0.311 |
| 206 | CTD-2013N24.2 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-7-1-3p | 15 | MEF2C | Sponge network | -1.745 | 0 | -0.862 | 5.0E-5 | 0.311 |
| 207 | RP11-736K20.5 | hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 11 | PDGFRA | Sponge network | -1.84 | 0 | -0.92 | 0.00039 | 0.311 |
| 208 | RP11-399O19.9 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-576-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 14 | PDGFRA | Sponge network | -0.873 | 0.00072 | -0.92 | 0.00039 | 0.31 |
| 209 | WDFY3-AS2 |
hsa-let-7g-3p;hsa-miR-132-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-96-5p | 11 | CACNB4 | Sponge network | -1.297 | 0 | -2.288 | 0 | 0.31 |
| 210 | AC011899.9 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 11 | NTRK2 | Sponge network | -2.611 | 0 | -2.564 | 0 | 0.31 |
| 211 | RP11-352D13.6 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p | 10 | NTRK2 | Sponge network | -4.634 | 0 | -2.564 | 0 | 0.31 |
| 212 | RP11-476D10.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181d-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-7-1-3p;hsa-miR-96-5p | 10 | MAP3K3 | Sponge network | -4.519 | 0 | -1.148 | 0 | 0.309 |
| 213 | BZRAP1-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-335-3p | 10 | TGFBR2 | Sponge network | -0.785 | 0.00723 | -1.915 | 0 | 0.309 |
| 214 | SH3RF3-AS1 |
hsa-miR-107;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-26b-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 12 | MEF2C | Sponge network | -1.583 | 0 | -0.862 | 5.0E-5 | 0.309 |
| 215 | LINC00968 |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-193b-3p;hsa-miR-200a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-590-5p | 12 | TGFB2 | Sponge network | -4.19 | 0 | -1.547 | 1.0E-5 | 0.308 |
| 216 | LINC00092 | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-590-3p | 10 | TGFBR2 | Sponge network | -2.383 | 0 | -1.915 | 0 | 0.308 |
| 217 | TBX5-AS1 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-196b-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-29a-5p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-450b-5p | 10 | CACNB4 | Sponge network | -2.108 | 0 | -2.288 | 0 | 0.308 |
| 218 | RP5-839B4.8 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-21-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-425-5p;hsa-miR-7-1-3p | 13 | MEF2C | Sponge network | -5.037 | 0 | -0.862 | 5.0E-5 | 0.308 |
| 219 | AF131215.9 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 11 | NTRK2 | Sponge network | -1.808 | 0 | -2.564 | 0 | 0.307 |
| 220 | HLA-F-AS1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-96-5p | 17 | MEF2C | Sponge network | -0.495 | 0.12126 | -0.862 | 5.0E-5 | 0.306 |
| 221 | AC004947.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -3.94 | 0 | -1.915 | 0 | 0.306 |
| 222 | RP11-1223D19.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -0.862 | 0.05389 | -1.915 | 0 | 0.303 |
| 223 | RBPMS-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130a-3p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-93-5p | 15 | TGFBR2 | Sponge network | -1.548 | 1.0E-5 | -1.915 | 0 | 0.301 |
| 224 | RP11-452C13.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p | 10 | TGFBR2 | Sponge network | -2.725 | 1.0E-5 | -1.915 | 0 | 0.3 |
| 225 | CASC2 |
hsa-miR-106a-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 15 | TGFBR2 | Sponge network | -1.086 | 0 | -1.915 | 0 | 0.299 |
| 226 | TBX5-AS1 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-5p | 14 | NTRK2 | Sponge network | -2.108 | 0 | -2.564 | 0 | 0.299 |
| 227 | RP11-378A13.1 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-3p | 14 | NTRK2 | Sponge network | -1.713 | 0 | -2.564 | 0 | 0.298 |
| 228 | RP11-401P9.4 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-5p | 13 | NTRK2 | Sponge network | -3.04 | 0 | -2.564 | 0 | 0.298 |
| 229 | LINC00922 |
hsa-miR-107;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-324-3p;hsa-miR-421;hsa-miR-7-1-3p;hsa-miR-744-3p;hsa-miR-9-5p;hsa-miR-92a-3p | 12 | MEF2C | Sponge network | -0.842 | 0.11239 | -0.862 | 5.0E-5 | 0.298 |
| 230 | RP11-400K9.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-93-5p | 11 | PDGFRA | Sponge network | -1.193 | 0.00359 | -0.92 | 0.00039 | 0.297 |
| 231 | RP11-166D19.1 |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-589-3p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-3p;hsa-miR-96-5p | 14 | CACNA1C | Sponge network | -0.582 | 0.05253 | -1.133 | 2.0E-5 | 0.295 |
| 232 | PCED1B-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | MAP3K3 | Sponge network | -0.672 | 0.02084 | -1.148 | 0 | 0.295 |
| 233 | WDFY3-AS2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-22-3p;hsa-miR-7-1-3p;hsa-miR-708-5p;hsa-miR-96-5p | 12 | MAP3K3 | Sponge network | -1.297 | 0 | -1.148 | 0 | 0.295 |
| 234 | CTC-297N7.9 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-2355-5p;hsa-miR-590-5p;hsa-miR-744-3p;hsa-miR-9-5p | 10 | MEF2C | Sponge network | -1.562 | 6.0E-5 | -0.862 | 5.0E-5 | 0.295 |
| 235 | PSMG3-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20b-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-590-3p | 10 | TGFBR2 | Sponge network | -0.522 | 0.09584 | -1.915 | 0 | 0.294 |
| 236 | RP11-166D19.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-92a-3p | 18 | TGFBR2 | Sponge network | -0.582 | 0.05253 | -1.915 | 0 | 0.294 |
| 237 | RP11-456K23.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-155-5p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20a-5p;hsa-miR-542-3p;hsa-miR-590-3p;hsa-miR-93-5p | 11 | RAPGEF2 | Sponge network | -1.488 | 0 | -0.845 | 0 | 0.293 |
| 238 | LINC00968 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20a-5p;hsa-miR-339-5p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-93-5p | 11 | RAPGEF2 | Sponge network | -4.19 | 0 | -0.845 | 0 | 0.292 |
| 239 | AF131215.9 |
hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 10 | TGFBR2 | Sponge network | -1.808 | 0 | -1.915 | 0 | 0.292 |
| 240 | LINC00702 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-339-5p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-93-5p | 10 | RAPGEF2 | Sponge network | -2.856 | 0 | -0.845 | 0 | 0.291 |
| 241 | RP11-462G12.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -1.071 | 0.01175 | -1.915 | 0 | 0.291 |
| 242 | RP11-400K9.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -1.193 | 0.00359 | -1.915 | 0 | 0.291 |
| 243 | FENDRR |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-331-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-550a-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-744-3p;hsa-miR-96-5p | 19 | MEF2C | Sponge network | -4.222 | 0 | -0.862 | 5.0E-5 | 0.29 |
| 244 | C1orf132 |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-2355-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-550a-3p;hsa-miR-590-3p;hsa-miR-9-5p | 13 | MEF2C | Sponge network | -0.86 | 0.02429 | -0.862 | 5.0E-5 | 0.289 |
| 245 | GAS6-AS2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-5p | 13 | NTRK2 | Sponge network | -1.761 | 0 | -2.564 | 0 | 0.287 |
| 246 | RP11-716O23.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-92a-3p | 11 | TGFBR2 | Sponge network | -5.465 | 0 | -1.915 | 0 | 0.286 |
| 247 | LINC00619 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-576-5p;hsa-miR-7-1-3p | 13 | PDGFRA | Sponge network | -2.307 | 0.02217 | -0.92 | 0.00039 | 0.285 |
| 248 | AF131215.2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-589-3p | 11 | PDGFRA | Sponge network | -2.09 | 0 | -0.92 | 0.00039 | 0.285 |
| 249 | LINC00922 |
hsa-miR-107;hsa-miR-148b-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-421;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-3p | 10 | CACNA1C | Sponge network | -0.842 | 0.11239 | -1.133 | 2.0E-5 | 0.284 |
| 250 | HLA-F-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -0.495 | 0.12126 | -1.915 | 0 | 0.283 |
| 251 | TPRG1-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-9-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.947 | 0.04384 | -1.915 | 0 | 0.282 |
| 252 | RP11-532F6.3 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-576-5p | 11 | NTRK2 | Sponge network | -2.028 | 0 | -2.564 | 0 | 0.281 |
| 253 | AF131215.9 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-196b-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-590-3p | 10 | CACNB4 | Sponge network | -1.808 | 0 | -2.288 | 0 | 0.281 |
| 254 | CTC-297N7.9 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-590-5p | 10 | NTRK2 | Sponge network | -1.562 | 6.0E-5 | -2.564 | 0 | 0.28 |
| 255 | RP11-284N8.3 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 14 | PDGFRA | Sponge network | -0.761 | 0.05061 | -0.92 | 0.00039 | 0.278 |
| 256 | CTD-2135D7.5 | hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-421;hsa-miR-590-5p | 12 | MEF2C | Sponge network | -3.162 | 0 | -0.862 | 5.0E-5 | 0.277 |
| 257 | AF131215.9 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-200c-5p;hsa-miR-21-5p;hsa-miR-2355-5p;hsa-miR-335-3p;hsa-miR-590-3p | 10 | MEF2C | Sponge network | -1.808 | 0 | -0.862 | 5.0E-5 | 0.276 |
| 258 | RP11-246K15.1 | hsa-miR-107;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-5p;hsa-miR-22-5p;hsa-miR-320b;hsa-miR-9-5p | 10 | MEF2C | Sponge network | -5.663 | 0 | -0.862 | 5.0E-5 | 0.273 |
| 259 | RP11-77A13.1 |
hsa-miR-107;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-96-5p | 10 | MEF2C | Sponge network | -6.738 | 0 | -0.862 | 5.0E-5 | 0.273 |
| 260 | BAIAP2-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.182 | 0.51705 | -1.915 | 0 | 0.272 |
| 261 | RP11-389C8.2 |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-589-3p;hsa-miR-590-3p | 10 | TGFB2 | Sponge network | -2.039 | 0 | -1.547 | 1.0E-5 | 0.272 |
| 262 | MIR497HG |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-503-5p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-93-3p | 13 | FGF2 | Sponge network | -2.142 | 0 | -2.545 | 0 | 0.271 |
| 263 | CTD-2008P7.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130a-3p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-9-5p | 15 | TGFBR2 | Sponge network | -1.912 | 1.0E-5 | -1.915 | 0 | 0.27 |
| 264 | NR2F1-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 12 | PDGFRA | Sponge network | -0.427 | 0.1559 | -0.92 | 0.00039 | 0.27 |
| 265 | RP11-159H10.3 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-miR-125b-2-3p;hsa-miR-1266-5p;hsa-miR-143-3p;hsa-miR-195-5p;hsa-miR-2110;hsa-miR-3065-3p;hsa-miR-30a-3p;hsa-miR-30e-3p | 12 | CACNA1E | Sponge network | 3.889 | 0 | 1.62 | 0.0002 | 0.268 |
| 266 | LINC01010 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.54 | 0.27666 | -1.915 | 0 | 0.267 |
| 267 | RP11-352D13.6 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-96-5p | 10 | MEF2C | Sponge network | -4.634 | 0 | -0.862 | 5.0E-5 | 0.266 |
| 268 | CTD-2003C8.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-454-3p | 11 | TGFBR2 | Sponge network | -3.403 | 0 | -1.915 | 0 | 0.266 |
| 269 | NR2F1-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | MAP3K3 | Sponge network | -0.427 | 0.1559 | -1.148 | 0 | 0.265 |
| 270 | BDNF-AS |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.568 | 0.02011 | -1.915 | 0 | 0.263 |
| 271 | RP11-401P9.4 |
hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-21-5p;hsa-miR-2355-5p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-590-5p | 10 | MEF2C | Sponge network | -3.04 | 0 | -0.862 | 5.0E-5 | 0.263 |
| 272 | SAP30L-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-590-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 10 | TGFBR2 | Sponge network | 0.631 | 0.02291 | -1.915 | 0 | 0.263 |
| 273 | RP4-639F20.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-320b | 10 | TGFBR2 | Sponge network | -1.312 | 0 | -1.915 | 0 | 0.263 |
| 274 | CTC-523E23.4 | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-9-5p | 10 | TGFBR2 | Sponge network | -1.636 | 0.00051 | -1.915 | 0 | 0.261 |
| 275 | HLA-F-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181d-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 11 | MAP3K3 | Sponge network | -0.495 | 0.12126 | -1.148 | 0 | 0.26 |
| 276 | LINC00922 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-9-5p;hsa-miR-92a-3p | 11 | TGFBR2 | Sponge network | -0.842 | 0.11239 | -1.915 | 0 | 0.26 |
| 277 | LINC00261 |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-550a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 15 | MEF2C | Sponge network | -2.566 | 0.00025 | -0.862 | 5.0E-5 | 0.26 |
| 278 | RP11-1024P17.1 |
hsa-let-7g-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-196b-5p;hsa-miR-20a-3p;hsa-miR-212-3p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-589-3p;hsa-miR-590-3p | 11 | CACNB4 | Sponge network | -2.062 | 0 | -2.288 | 0 | 0.26 |
| 279 | LL22NC03-86G7.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-96-5p | 11 | PDGFRA | Sponge network | -1.177 | 3.0E-5 | -0.92 | 0.00039 | 0.257 |
| 280 | AC003991.3 |
hsa-miR-10a-3p;hsa-miR-130b-5p;hsa-miR-18a-5p;hsa-miR-194-5p;hsa-miR-200c-5p;hsa-miR-320b;hsa-miR-331-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-425-5p;hsa-miR-7-1-3p;hsa-miR-744-3p;hsa-miR-9-5p;hsa-miR-92a-3p | 14 | MEF2C | Sponge network | -0.787 | 0.08132 | -0.862 | 5.0E-5 | 0.256 |
| 281 | AC011526.1 | hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-93-5p;hsa-miR-96-5p | 12 | PDGFRA | Sponge network | -2.783 | 0 | -0.92 | 0.00039 | 0.254 |
| 282 | LINC00619 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p | 10 | TGFBR2 | Sponge network | -2.307 | 0.02217 | -1.915 | 0 | 0.253 |
| 283 | CASC2 |
hsa-miR-130b-5p;hsa-miR-183-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-2355-5p;hsa-miR-335-3p;hsa-miR-425-5p;hsa-miR-550a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-744-3p | 15 | MEF2C | Sponge network | -1.086 | 0 | -0.862 | 5.0E-5 | 0.252 |
| 284 | CTD-2013N24.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p | 14 | NTRK2 | Sponge network | -1.745 | 0 | -2.564 | 0 | 0.252 |
| 285 | PCED1B-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-92a-3p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -0.672 | 0.02084 | -1.915 | 0 | 0.252 |
| 286 | LUCAT1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-125b-2-3p;hsa-miR-143-3p;hsa-miR-195-3p;hsa-miR-195-5p;hsa-miR-2110;hsa-miR-3065-3p;hsa-miR-30a-3p;hsa-miR-30b-3p | 11 | CACNA1E | Sponge network | 3.001 | 0 | 1.62 | 0.0002 | 0.251 |
| 287 | AC093495.4 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -1.583 | 0.00599 | -1.915 | 0 | 0.25 |