This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-141-3p | ABL1 | 1.13 | 0.09667 | -0.28 | 0.25357 | MirTarget | -0.12 | 0 | NA | |
| 2 | hsa-miR-141-3p | ABL2 | 1.13 | 0.09667 | -0.13 | 0.63539 | MirTarget; TargetScan; miRNATAP | -0.16 | 0 | NA | |
| 3 | hsa-miR-141-3p | ADAM22 | 1.13 | 0.09667 | -0.88 | 0.25527 | MirTarget | -0.33 | 7.0E-5 | NA | |
| 4 | hsa-miR-141-3p | ADARB1 | 1.13 | 0.09667 | -0.6 | 0.08985 | MirTarget | -0.29 | 0 | NA | |
| 5 | hsa-miR-141-3p | AKAP2 | 1.13 | 0.09667 | -0.87 | 0.26357 | TargetScan | -0.48 | 0 | NA | |
| 6 | hsa-miR-141-3p | AMOTL1 | 1.13 | 0.09667 | -0.52 | 0.23224 | TargetScan | -0.26 | 0 | NA | |
| 7 | hsa-miR-141-3p | AMZ1 | 1.13 | 0.09667 | -0.84 | 0.19343 | mirMAP | -0.21 | 0.00226 | NA | |
| 8 | hsa-miR-141-3p | ANGPTL1 | 1.13 | 0.09667 | -0.94 | 0.28852 | MirTarget | -0.52 | 0 | NA | |
| 9 | hsa-miR-141-3p | ANK2 | 1.13 | 0.09667 | -0.87 | 0.25994 | TargetScan; miRNATAP | -0.48 | 0 | NA | |
| 10 | hsa-miR-141-3p | ANKRD13A | 1.13 | 0.09667 | -0.83 | 0.00018 | MirTarget | -0.11 | 0 | NA | |
| 11 | hsa-miR-141-3p | ANKRD44 | 1.13 | 0.09667 | -1.3 | 0.04083 | TargetScan | -0.4 | 0 | NA | |
| 12 | hsa-miR-141-3p | ANO6 | 1.13 | 0.09667 | 0.05 | 0.90255 | MirTarget; TargetScan | -0.13 | 0.00132 | NA | |
| 13 | hsa-miR-141-3p | APBB2 | 1.13 | 0.09667 | -0.11 | 0.68147 | MirTarget; TargetScan | -0.14 | 0 | NA | |
| 14 | hsa-miR-141-3p | APOLD1 | 1.13 | 0.09667 | 0.05 | 0.92161 | TargetScan | -0.29 | 0 | NA | |
| 15 | hsa-miR-141-3p | AR | 1.13 | 0.09667 | -0.52 | 0.55352 | mirMAP | -0.46 | 0 | 26062412; 22314666 | miR 141 3p regulates the expression of androgen receptor by targeting its 3'UTR in prostate cancer LNCaP cells; After prostate cancer cell line LNCaP was transfected with miR-141-3p mimics expression levels of AR mRNA and protein in the LNCaP cells were detected by reverse transcription PCR and Western blotting respectively; The 3'untranslated regions 3'UTR of AR mRNA containing the binding site of miR-141-3p was amplified by PCR and inserted into pmiR-report vector a 3'downstream luciferase reporter gene; Transfection of miR-141-3p mimics decreased both mRNA and protein expression levels of AR in LNCaP cells; AR is a direct target gene of miR-141-3p;miR 141 modulates androgen receptor transcriptional activity in human prostate cancer cells through targeting the small heterodimer partner protein; Here we investigated the correlation of Shp expression with the cellular level of miR-141 and its effects on AR transcriptional activity in non-malignant and malignant human prostate epithelial cell lines; Phenethyl isothiocyanate a natural constituent of many edible cruciferous vegetables increased Shp expression downregulated miR-141 and inhibited AR transcriptional activity in LNCaP cells; Shp is a target for miR-141 and it is downregulated in cultured human PCa cells with the involvement of upregulation of miR-141 which promotes AR transcriptional activity |
| 16 | hsa-miR-141-3p | ARHGAP23 | 1.13 | 0.09667 | 0.01 | 0.98674 | TargetScan | -0.19 | 0.00104 | NA | |
| 17 | hsa-miR-141-3p | ARHGAP24 | 1.13 | 0.09667 | -0.91 | 0.02885 | TargetScan; miRNATAP | -0.16 | 0.00023 | NA | |
| 18 | hsa-miR-141-3p | ARID5B | 1.13 | 0.09667 | -0.68 | 0.07996 | TargetScan; miRNATAP | -0.31 | 0 | NA | |
| 19 | hsa-miR-141-3p | ARL10 | 1.13 | 0.09667 | -0.61 | 0.19487 | mirMAP | -0.23 | 0 | NA | |
| 20 | hsa-miR-141-3p | ASTN1 | 1.13 | 0.09667 | -1.5 | 0.17933 | MirTarget; miRNATAP | -0.25 | 0.04128 | NA | |
| 21 | hsa-miR-141-3p | ATP6V1B2 | 1.13 | 0.09667 | -0.68 | 0.00779 | MirTarget; TargetScan; miRNATAP | -0.11 | 0.00011 | NA | |
| 22 | hsa-miR-141-3p | ATP8A2 | 1.13 | 0.09667 | -0.04 | 0.95305 | TargetScan | -0.18 | 0.01465 | NA | |
| 23 | hsa-miR-141-3p | ATXN1 | 1.13 | 0.09667 | -0.51 | 0.09797 | MirTarget; TargetScan; miRNATAP | -0.18 | 0 | NA | |
| 24 | hsa-miR-141-3p | BACH2 | 1.13 | 0.09667 | -1.75 | 0.00364 | TargetScan | -0.43 | 0 | NA | |
| 25 | hsa-miR-141-3p | BEND4 | 1.13 | 0.09667 | -1.9 | 0.08641 | TargetScan; miRNATAP | -0.51 | 2.0E-5 | NA | |
| 26 | hsa-miR-141-3p | BICD2 | 1.13 | 0.09667 | -0.53 | 0.06121 | MirTarget; miRNATAP | -0.18 | 0 | NA | |
| 27 | hsa-miR-15a-5p | BMI1 | -0.74 | 0.01074 | -0.18 | 0.45133 | miRNAWalker2 validate; miRTarBase | -0.14 | 0.0172 | 27596816 | Our results suggest that overexpression of miR-15a and miR-16 mediates down-regulation of BMI1 and leads to mitochondrial mediated apoptosis |
| 28 | hsa-miR-362-3p | BMI1 | -1.21 | 0.01257 | -0.18 | 0.45133 | miRanda | -0.11 | 0.00173 | NA | |
| 29 | hsa-miR-455-5p | BMI1 | -0.23 | 0.54044 | -0.18 | 0.45133 | miRanda | -0.11 | 0.01889 | NA | |
| 30 | hsa-miR-590-3p | BMI1 | -0.92 | 0.03456 | -0.18 | 0.45133 | miRanda; mirMAP | -0.12 | 0.00201 | NA | |
| 31 | hsa-miR-141-3p | BNC2 | 1.13 | 0.09667 | -0.15 | 0.8368 | TargetScan | -0.57 | 0 | NA | |
| 32 | hsa-miR-141-3p | BTN2A2 | 1.13 | 0.09667 | -1.12 | 0.00207 | mirMAP | -0.22 | 0 | NA | |
| 33 | hsa-miR-141-3p | BVES | 1.13 | 0.09667 | -0.04 | 0.94915 | mirMAP | -0.28 | 1.0E-5 | NA | |
| 34 | hsa-miR-141-3p | C11orf21 | 1.13 | 0.09667 | -1.67 | 0.02483 | mirMAP | -0.39 | 0 | NA | |
| 35 | hsa-miR-141-3p | C20orf194 | 1.13 | 0.09667 | -0.45 | 0.19186 | TargetScan; mirMAP | -0.11 | 0.00365 | NA | |
| 36 | hsa-miR-141-3p | C21orf91 | 1.13 | 0.09667 | -0.85 | 0.01519 | TargetScan | -0.23 | 0 | NA | |
| 37 | hsa-miR-141-3p | C6orf120 | 1.13 | 0.09667 | -0.4 | 0.05976 | TargetScan | -0.11 | 0 | NA | |
| 38 | hsa-miR-141-3p | CACNA1E | 1.13 | 0.09667 | -0.51 | 0.44626 | MirTarget; TargetScan; miRNATAP | -0.45 | 0 | NA | |
| 39 | hsa-miR-141-3p | CACNA1I | 1.13 | 0.09667 | -0.24 | 0.78112 | TargetScan | -0.31 | 0.00117 | NA | |
| 40 | hsa-miR-141-3p | CACNA2D1 | 1.13 | 0.09667 | -0.16 | 0.86886 | TargetScan | -0.49 | 0 | NA | |
| 41 | hsa-miR-141-3p | CADM1 | 1.13 | 0.09667 | -0.53 | 0.36556 | MirTarget; TargetScan | -0.15 | 0.01458 | NA | |
| 42 | hsa-miR-141-3p | CALU | 1.13 | 0.09667 | 0.41 | 0.28312 | TargetScan | -0.18 | 0 | NA | |
| 43 | hsa-miR-141-3p | CAMK4 | 1.13 | 0.09667 | -0.61 | 0.35184 | mirMAP | -0.28 | 5.0E-5 | NA | |
| 44 | hsa-miR-141-3p | CBL | 1.13 | 0.09667 | -0.48 | 0.10129 | MirTarget; TargetScan; mirMAP | -0.17 | 0 | NA | |
| 45 | hsa-miR-141-3p | CCDC80 | 1.13 | 0.09667 | 0.02 | 0.97481 | MirTarget; TargetScan | -0.57 | 0 | NA | |
| 46 | hsa-miR-141-3p | CD274 | 1.13 | 0.09667 | -0.72 | 0.22253 | MirTarget | -0.23 | 0.0003 | NA | |
| 47 | hsa-miR-141-3p | CDC14A | 1.13 | 0.09667 | -0.8 | 0.05646 | TargetScan; miRNATAP | -0.23 | 0 | NA | |
| 48 | hsa-miR-141-3p | CDC42EP3 | 1.13 | 0.09667 | 0.05 | 0.88181 | MirTarget; TargetScan | -0.12 | 0.00065 | NA | |
| 49 | hsa-miR-141-3p | CDON | 1.13 | 0.09667 | -0.13 | 0.81496 | TargetScan; miRNATAP | -0.18 | 0.00203 | NA | |
| 50 | hsa-miR-141-3p | CDV3 | 1.13 | 0.09667 | -0.37 | 0.06194 | TargetScan; miRNATAP | -0.1 | 0 | NA | |
| 51 | hsa-miR-141-3p | CELF2 | 1.13 | 0.09667 | -1.75 | 0.00444 | mirMAP; miRNATAP | -0.53 | 0 | NA | |
| 52 | hsa-miR-141-3p | CEP170 | 1.13 | 0.09667 | -0.54 | 0.15049 | TargetScan | -0.2 | 0 | NA | |
| 53 | hsa-miR-141-3p | CFL2 | 1.13 | 0.09667 | -0.4 | 0.27702 | TargetScan | -0.21 | 0 | NA | |
| 54 | hsa-miR-141-3p | CHRDL1 | 1.13 | 0.09667 | -0.73 | 0.53577 | MirTarget; TargetScan | -0.9 | 0 | NA | |
| 55 | hsa-miR-141-3p | CLIC4 | 1.13 | 0.09667 | -0.37 | 0.38824 | MirTarget | -0.3 | 0 | NA | |
| 56 | hsa-miR-141-3p | CLSPN | 1.13 | 0.09667 | 0.41 | 0.57124 | MirTarget | -0.25 | 0.00127 | NA | |
| 57 | hsa-miR-141-3p | CNTN1 | 1.13 | 0.09667 | -0.06 | 0.94437 | MirTarget; TargetScan; miRNATAP | -0.29 | 0.00358 | NA | |
| 58 | hsa-miR-141-3p | COL11A1 | 1.13 | 0.09667 | 1.03 | 0.52179 | miRNATAP | -0.61 | 0.00037 | NA | |
| 59 | hsa-miR-141-3p | COL15A1 | 1.13 | 0.09667 | 0.39 | 0.50342 | MirTarget | -0.32 | 0 | NA | |
| 60 | hsa-miR-141-3p | COL8A1 | 1.13 | 0.09667 | 0.61 | 0.5252 | mirMAP | -0.5 | 0 | NA | |
| 61 | hsa-miR-141-3p | CORO1C | 1.13 | 0.09667 | -0.17 | 0.56501 | TargetScan | -0.17 | 0 | NA | |
| 62 | hsa-miR-141-3p | CREB3L2 | 1.13 | 0.09667 | 0.1 | 0.72448 | MirTarget | -0.13 | 6.0E-5 | NA | |
| 63 | hsa-miR-141-3p | CREB5 | 1.13 | 0.09667 | -1.59 | 0.00384 | mirMAP | -0.22 | 0.0002 | NA | |
| 64 | hsa-miR-141-3p | CRLF3 | 1.13 | 0.09667 | -0.46 | 0.08918 | TargetScan | -0.15 | 0 | NA | |
| 65 | hsa-miR-141-3p | CRYBG3 | 1.13 | 0.09667 | -0.21 | 0.58758 | MirTarget | -0.16 | 8.0E-5 | NA | |
| 66 | hsa-miR-141-3p | CSF3 | 1.13 | 0.09667 | -0.85 | 0.51687 | TargetScan; miRNATAP | -0.48 | 0.00061 | NA | |
| 67 | hsa-miR-141-3p | CSTA | 1.13 | 0.09667 | -0.68 | 0.38523 | MirTarget | -0.21 | 0.01287 | NA | |
| 68 | hsa-miR-141-3p | CTSB | 1.13 | 0.09667 | -0.34 | 0.3287 | mirMAP | -0.17 | 1.0E-5 | NA | |
| 69 | hsa-miR-141-3p | CXCL12 | 1.13 | 0.09667 | -1.19 | 0.1171 | miRNATAP | -0.59 | 0 | NA | |
| 70 | hsa-miR-141-3p | CYLD | 1.13 | 0.09667 | -0.58 | 0.0188 | mirMAP | -0.12 | 0 | NA | |
| 71 | hsa-miR-141-3p | CYP26B1 | 1.13 | 0.09667 | -0.14 | 0.81629 | MirTarget; TargetScan; miRNATAP | -0.33 | 0 | NA | |
| 72 | hsa-miR-141-3p | DCP2 | 1.13 | 0.09667 | -0.45 | 0.02171 | MirTarget; TargetScan | -0.12 | 0 | NA | |
| 73 | hsa-miR-141-3p | DCUN1D3 | 1.13 | 0.09667 | -0.02 | 0.94021 | MirTarget; TargetScan | -0.13 | 0 | NA | |
| 74 | hsa-miR-141-3p | DDI2 | 1.13 | 0.09667 | -0.23 | 0.72424 | mirMAP | -0.17 | 0.01803 | NA | |
| 75 | hsa-miR-141-3p | DIO2 | 1.13 | 0.09667 | 1.45 | 0.03366 | miRNATAP | -0.31 | 3.0E-5 | NA | |
| 76 | hsa-miR-141-3p | DLC1 | 1.13 | 0.09667 | -0.71 | 0.10958 | TargetScan; miRNATAP | -0.32 | 0 | 26278151 | MicroRNA 141 regulates the tumour suppressor DLC1 in colorectal cancer; Luciferase reporter assays and Western blots showed that DLC1 was a direct target of miR-141 in CRC; The expression levels of miR-141 were obviously up-regulated in CRC tissues compared to non-cancerous tissues while DLC1 expression levels were down-regulated in a high proportion of clinical samples 14/18; In addition correlation analyses revealed negative correlation between miR-141 levels and DLC1 expression levels in CRC tissues; In conclusion we demonstrated that miR-141 is up-regulated in CRC and acts as a functional oncogene by targeting DLC1 |
| 77 | hsa-miR-141-3p | DLX5 | 1.13 | 0.09667 | -0.22 | 0.77457 | miRNAWalker2 validate; miRTarBase | -0.2 | 0.0163 | NA | |
| 78 | hsa-miR-141-3p | DOCK4 | 1.13 | 0.09667 | -0.58 | 0.16909 | MirTarget; TargetScan | -0.18 | 5.0E-5 | NA | |
| 79 | hsa-miR-141-3p | DPYSL3 | 1.13 | 0.09667 | -0.46 | 0.37566 | TargetScan | -0.31 | 0 | NA | |
| 80 | hsa-miR-141-3p | DR1 | 1.13 | 0.09667 | 0.07 | 0.74852 | MirTarget; TargetScan | -0.1 | 1.0E-5 | NA | |
| 81 | hsa-miR-141-3p | DSEL | 1.13 | 0.09667 | -0.27 | 0.62922 | TargetScan | -0.23 | 0.00015 | NA | |
| 82 | hsa-miR-141-3p | DUSP1 | 1.13 | 0.09667 | -0.09 | 0.86695 | TargetScan | -0.33 | 0 | NA | |
| 83 | hsa-miR-141-3p | DYNC1I1 | 1.13 | 0.09667 | 0.74 | 0.31036 | MirTarget | -0.18 | 0.02299 | NA | |
| 84 | hsa-miR-141-3p | DZIP1 | 1.13 | 0.09667 | -0.6 | 0.27307 | TargetScan; miRNATAP | -0.21 | 0.00025 | NA | |
| 85 | hsa-miR-141-3p | E2F3 | 1.13 | 0.09667 | -0.2 | 0.50008 | miRNAWalker2 validate; TargetScan; miRNATAP | -0.12 | 0.00019 | 26233544 | MiR 141 Inhibits Gastric Cancer Proliferation by Interacting with Long Noncoding RNA MEG3 and Down Regulating E2F3 Expression; QRT-PCR was used to detect miR-141 MEG3 and E2F3 in gastric tissues and cells; Western blot and luciferase activity were used to identify E2F3 as one of the direct targets of miR-141; E2F3 was identified as a target of miR-141 and its inhibition significantly reduced MEG3 expression; E2F3 expression was also found to be negatively associated with both MEG3 and miR-141; E2F3 over-expression partly reversed the changes caused by transfection of miR-141 mimic and inhibition of miR-141 or MEG3 overrides MEG3- or miR-141-induced modulation of cell growth in GC; These findings together suggested that miR-141 could be interacting with MEG3 and targeting E2F3 and these factors may play important anti-tumor effects in GC pathogenesis and provide therapeutic targets in the clinics |
| 86 | hsa-miR-141-3p | EGR2 | 1.13 | 0.09667 | -0.6 | 0.36333 | TargetScan; miRNATAP | -0.39 | 0 | NA | |
| 87 | hsa-miR-141-3p | EHD3 | 1.13 | 0.09667 | -0.58 | 0.13605 | mirMAP | -0.27 | 0 | NA | |
| 88 | hsa-miR-141-3p | ELK3 | 1.13 | 0.09667 | -0.28 | 0.5575 | TargetScan | -0.26 | 0 | NA | |
| 89 | hsa-miR-141-3p | EPB41L3 | 1.13 | 0.09667 | -0.67 | 0.22434 | miRNATAP | -0.29 | 0 | NA | |
| 90 | hsa-miR-141-3p | ERG | 1.13 | 0.09667 | -1 | 0.04239 | MirTarget; TargetScan | -0.37 | 0 | NA | |
| 91 | hsa-miR-141-3p | ETS1 | 1.13 | 0.09667 | -0.99 | 0.01795 | mirMAP | -0.31 | 0 | NA | |
| 92 | hsa-miR-141-3p | FAM133A | 1.13 | 0.09667 | 0.37 | 0.63896 | mirMAP | -0.25 | 0.00298 | NA | |
| 93 | hsa-miR-141-3p | FAM168A | 1.13 | 0.09667 | -0.35 | 0.46871 | TargetScan | -0.22 | 2.0E-5 | NA | |
| 94 | hsa-miR-141-3p | FAM20C | 1.13 | 0.09667 | 0.18 | 0.60037 | MirTarget | -0.13 | 0.00033 | NA | |
| 95 | hsa-miR-141-3p | FAM26E | 1.13 | 0.09667 | -0.42 | 0.45631 | mirMAP | -0.43 | 0 | NA | |
| 96 | hsa-miR-141-3p | FAT3 | 1.13 | 0.09667 | -1.22 | 0.10046 | MirTarget; miRNATAP | -0.45 | 0 | NA | |
| 97 | hsa-miR-141-3p | FER | 1.13 | 0.09667 | -0.09 | 0.82691 | mirMAP | -0.13 | 0.0023 | NA | |
| 98 | hsa-miR-141-3p | FGF13 | 1.13 | 0.09667 | -0.98 | 0.1122 | TargetScan | -0.15 | 0.02579 | NA | |
| 99 | hsa-miR-141-3p | FKBP5 | 1.13 | 0.09667 | -0.64 | 0.31693 | TargetScan | -0.32 | 0 | NA | |
| 100 | hsa-miR-141-3p | FLI1 | 1.13 | 0.09667 | -1.49 | 0.00419 | TargetScan | -0.45 | 0 | NA | |
| 101 | hsa-miR-141-3p | FOXN3 | 1.13 | 0.09667 | -0.48 | 0.10355 | MirTarget; TargetScan; miRNATAP | -0.2 | 0 | NA | |
| 102 | hsa-miR-141-3p | FOXO1 | 1.13 | 0.09667 | -0.73 | 0.05833 | mirMAP | -0.26 | 0 | NA | |
| 103 | hsa-miR-141-3p | FOXP2 | 1.13 | 0.09667 | 0.26 | 0.75107 | TargetScan | -0.34 | 0.0001 | NA | |
| 104 | hsa-miR-141-3p | FRMD4A | 1.13 | 0.09667 | -1.3 | 0.00053 | MirTarget; TargetScan; miRNATAP | -0.24 | 0 | NA | |
| 105 | hsa-miR-141-3p | FRMD6 | 1.13 | 0.09667 | -0.32 | 0.5949 | MirTarget; TargetScan; miRNATAP | -0.33 | 0 | NA | |
| 106 | hsa-miR-141-3p | FZD4 | 1.13 | 0.09667 | -0.61 | 0.11853 | TargetScan | -0.28 | 0 | NA | |
| 107 | hsa-miR-141-3p | GAB2 | 1.13 | 0.09667 | -0.37 | 0.19352 | mirMAP | -0.12 | 3.0E-5 | NA | |
| 108 | hsa-miR-141-3p | GATA3 | 1.13 | 0.09667 | 0.31 | 0.69983 | TargetScan | -0.25 | 0.00281 | NA | |
| 109 | hsa-miR-141-3p | GDF6 | 1.13 | 0.09667 | 0.41 | 0.59382 | TargetScan | -0.46 | 0 | NA | |
| 110 | hsa-miR-141-3p | GEM | 1.13 | 0.09667 | 0.03 | 0.9651 | MirTarget | -0.4 | 0 | NA | |
| 111 | hsa-miR-141-3p | GFRA1 | 1.13 | 0.09667 | -0.09 | 0.91461 | mirMAP | -0.36 | 8.0E-5 | NA | |
| 112 | hsa-miR-141-3p | GFRA2 | 1.13 | 0.09667 | -1.27 | 0.07593 | mirMAP | -0.45 | 0 | NA | |
| 113 | hsa-miR-141-3p | GIT2 | 1.13 | 0.09667 | -0.68 | 0.00253 | MirTarget | -0.14 | 0 | NA | |
| 114 | hsa-miR-141-3p | GJC1 | 1.13 | 0.09667 | 0.22 | 0.60528 | MirTarget; TargetScan; miRNATAP | -0.27 | 0 | NA | |
| 115 | hsa-miR-141-3p | GLCCI1 | 1.13 | 0.09667 | -0.47 | 0.23746 | MirTarget; TargetScan; miRNATAP | -0.15 | 0.00028 | NA | |
| 116 | hsa-miR-141-3p | GLDN | 1.13 | 0.09667 | -1.2 | 0.08392 | MirTarget | -0.19 | 0.01159 | NA | |
| 117 | hsa-miR-141-3p | GLI2 | 1.13 | 0.09667 | 0.04 | 0.95178 | MirTarget | -0.36 | 0 | NA | |
| 118 | hsa-miR-141-3p | GLRX | 1.13 | 0.09667 | -0.37 | 0.29901 | MirTarget; TargetScan; miRNATAP | -0.13 | 0.00035 | NA | |
| 119 | hsa-miR-141-3p | GNA12 | 1.13 | 0.09667 | -0.27 | 0.27167 | mirMAP | -0.13 | 0 | NA | |
| 120 | hsa-miR-141-3p | GNA13 | 1.13 | 0.09667 | -0.51 | 0.05072 | MirTarget; TargetScan | -0.14 | 0 | NA | |
| 121 | hsa-miR-141-3p | GNG7 | 1.13 | 0.09667 | -1.53 | 0.0195 | MirTarget; TargetScan; miRNATAP | -0.15 | 0.03175 | NA | |
| 122 | hsa-miR-141-3p | GPAM | 1.13 | 0.09667 | -0.86 | 0.0486 | MirTarget | -0.19 | 7.0E-5 | NA | |
| 123 | hsa-miR-141-3p | GPC6 | 1.13 | 0.09667 | 0.23 | 0.70012 | TargetScan | -0.3 | 0 | NA | |
| 124 | hsa-miR-141-3p | GPM6B | 1.13 | 0.09667 | -0.6 | 0.33946 | TargetScan; miRNATAP | -0.38 | 0 | NA | |
| 125 | hsa-miR-141-3p | GPR137B | 1.13 | 0.09667 | -0.13 | 0.6871 | TargetScan | -0.17 | 0 | NA | |
| 126 | hsa-miR-141-3p | GPR137C | 1.13 | 0.09667 | -0.74 | 0.0544 | TargetScan; miRNATAP | -0.14 | 0.00063 | NA | |
| 127 | hsa-miR-141-3p | GPR174 | 1.13 | 0.09667 | -1.4 | 0.13498 | TargetScan | -0.66 | 0 | NA | |
| 128 | hsa-miR-141-3p | GRAP2 | 1.13 | 0.09667 | -1.06 | 0.10477 | mirMAP | -0.38 | 0 | NA | |
| 129 | hsa-miR-141-3p | GRIK3 | 1.13 | 0.09667 | -0.66 | 0.41704 | mirMAP | -0.26 | 0.00292 | NA | |
| 130 | hsa-miR-141-3p | GRIN3A | 1.13 | 0.09667 | -0.82 | 0.03097 | TargetScan; miRNATAP | -0.15 | 0.00025 | NA | |
| 131 | hsa-miR-141-3p | HAS2 | 1.13 | 0.09667 | 0.52 | 0.52522 | MirTarget | -0.41 | 0 | NA | |
| 132 | hsa-miR-141-3p | HECW1 | 1.13 | 0.09667 | 0.52 | 0.58204 | TargetScan | -0.4 | 7.0E-5 | NA | |
| 133 | hsa-miR-141-3p | HEYL | 1.13 | 0.09667 | 0.19 | 0.73062 | mirMAP | -0.22 | 0.00021 | NA | |
| 134 | hsa-miR-141-3p | HGF | 1.13 | 0.09667 | -0.89 | 0.25864 | MirTarget; TargetScan | -0.57 | 0 | NA | |
| 135 | hsa-miR-141-3p | HIPK3 | 1.13 | 0.09667 | -0.25 | 0.7029 | MirTarget; TargetScan | -0.19 | 0.00673 | NA | |
| 136 | hsa-miR-141-3p | HLF | 1.13 | 0.09667 | -0.66 | 0.37924 | TargetScan | -0.17 | 0.03646 | NA | |
| 137 | hsa-miR-141-3p | HS3ST3B1 | 1.13 | 0.09667 | -0.6 | 0.33145 | mirMAP | -0.17 | 0.00957 | NA | |
| 138 | hsa-miR-141-3p | HSPA13 | 1.13 | 0.09667 | 0.02 | 0.95259 | TargetScan | -0.1 | 0.00769 | NA | |
| 139 | hsa-miR-141-3p | HTR2A | 1.13 | 0.09667 | 1.09 | 0.2119 | TargetScan | -0.56 | 0 | NA | |
| 140 | hsa-miR-141-3p | ICOSLG | 1.13 | 0.09667 | -0.7 | 0.05851 | mirMAP | -0.18 | 0 | NA | |
| 141 | hsa-miR-141-3p | IGDCC4 | 1.13 | 0.09667 | -0.95 | 0.14689 | TargetScan; miRNATAP | -0.54 | 0 | NA | |
| 142 | hsa-miR-141-3p | IGF2 | 1.13 | 0.09667 | 0.45 | 0.58211 | MirTarget; TargetScan | -0.38 | 1.0E-5 | NA | |
| 143 | hsa-miR-141-3p | IGSF6 | 1.13 | 0.09667 | -1.79 | 0.00917 | TargetScan | -0.46 | 0 | NA | |
| 144 | hsa-miR-141-3p | IL18R1 | 1.13 | 0.09667 | -0.25 | 0.60736 | MirTarget | -0.25 | 0 | NA | |
| 145 | hsa-miR-141-3p | IL6R | 1.13 | 0.09667 | -0.25 | 0.69611 | MirTarget | -0.17 | 0.01362 | NA | |
| 146 | hsa-miR-141-3p | IRAK3 | 1.13 | 0.09667 | -0.55 | 0.29063 | mirMAP | -0.21 | 0.00013 | NA | |
| 147 | hsa-miR-141-3p | ITGAV | 1.13 | 0.09667 | -0.33 | 0.42648 | MirTarget | -0.13 | 0.00399 | NA | |
| 148 | hsa-miR-141-3p | ITSN1 | 1.13 | 0.09667 | -0.16 | 0.58414 | TargetScan; mirMAP | -0.18 | 0 | NA | |
| 149 | hsa-miR-141-3p | JAZF1 | 1.13 | 0.09667 | -0.56 | 0.09201 | MirTarget; miRNATAP | -0.14 | 0.00013 | NA | |
| 150 | hsa-miR-141-3p | KATNAL1 | 1.13 | 0.09667 | -0.34 | 0.35357 | TargetScan | -0.14 | 0.00058 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 72 | 1395 | 1.616e-15 | 7.519e-12 |
| 2 | REGULATION OF CELL DIFFERENTIATION | 71 | 1492 | 1.52e-13 | 3.536e-10 |
| 3 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 60 | 1142 | 2.565e-13 | 3.978e-10 |
| 4 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 49 | 823 | 5.863e-13 | 6.82e-10 |
| 5 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 53 | 957 | 1.024e-12 | 9.499e-10 |
| 6 | CELLULAR COMPONENT MORPHOGENESIS | 51 | 900 | 1.225e-12 | 9.499e-10 |
| 7 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 47 | 788 | 1.718e-12 | 9.994e-10 |
| 8 | CIRCULATORY SYSTEM DEVELOPMENT | 47 | 788 | 1.718e-12 | 9.994e-10 |
| 9 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 37 | 513 | 2.02e-12 | 1.044e-09 |
| 10 | CELL DEVELOPMENT | 66 | 1426 | 4.504e-12 | 2.096e-09 |
| 11 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 60 | 1275 | 2.568e-11 | 1.086e-08 |
| 12 | TISSUE DEVELOPMENT | 66 | 1518 | 6.875e-11 | 2.666e-08 |
| 13 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 70 | 1672 | 8.759e-11 | 3.135e-08 |
| 14 | NEUROGENESIS | 62 | 1402 | 1.463e-10 | 4.862e-08 |
| 15 | VASCULATURE DEVELOPMENT | 32 | 469 | 2.789e-10 | 8.651e-08 |
| 16 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 25 | 306 | 6.911e-10 | 1.913e-07 |
| 17 | REGULATION OF CELL ADHESION | 37 | 629 | 6.991e-10 | 1.913e-07 |
| 18 | REGULATION OF NEURON DIFFERENTIATION | 34 | 554 | 1.213e-09 | 2.687e-07 |
| 19 | BLOOD VESSEL MORPHOGENESIS | 27 | 364 | 1.202e-09 | 2.687e-07 |
| 20 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 49 | 1021 | 1.136e-09 | 2.687e-07 |
| 21 | REGULATION OF CELL MORPHOGENESIS | 34 | 552 | 1.105e-09 | 2.687e-07 |
| 22 | LOCOMOTION | 51 | 1114 | 2.494e-09 | 5.276e-07 |
| 23 | REGULATION OF CELL PROJECTION ORGANIZATION | 33 | 558 | 5.389e-09 | 1.09e-06 |
| 24 | NEURON PROJECTION MORPHOGENESIS | 27 | 402 | 1.015e-08 | 1.968e-06 |
| 25 | CELL PROJECTION ORGANIZATION | 43 | 902 | 1.592e-08 | 2.893e-06 |
| 26 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 39 | 771 | 1.616e-08 | 2.893e-06 |
| 27 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 30 | 498 | 1.822e-08 | 3.139e-06 |
| 28 | POSITIVE REGULATION OF CELL DEVELOPMENT | 29 | 472 | 2.065e-08 | 3.432e-06 |
| 29 | POSITIVE REGULATION OF CELL PROLIFERATION | 40 | 814 | 2.315e-08 | 3.714e-06 |
| 30 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 38 | 750 | 2.396e-08 | 3.716e-06 |
| 31 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 36 | 689 | 2.647e-08 | 3.973e-06 |
| 32 | CELL MORPHOGENESIS INVOLVED IN NEURON DIFFERENTIATION | 25 | 368 | 2.899e-08 | 4.215e-06 |
| 33 | EMBRYONIC MORPHOGENESIS | 31 | 539 | 3.032e-08 | 4.276e-06 |
| 34 | CELL PART MORPHOGENESIS | 34 | 633 | 3.393e-08 | 4.643e-06 |
| 35 | NEURON DIFFERENTIATION | 41 | 874 | 5.497e-08 | 7.307e-06 |
| 36 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 27 | 437 | 5.752e-08 | 7.435e-06 |
| 37 | POSITIVE REGULATION OF GENE EXPRESSION | 64 | 1733 | 8.275e-08 | 1.041e-05 |
| 38 | SKELETAL SYSTEM DEVELOPMENT | 27 | 455 | 1.304e-07 | 1.588e-05 |
| 39 | NEURON PROJECTION DEVELOPMENT | 30 | 545 | 1.331e-07 | 1.588e-05 |
| 40 | ANGIOGENESIS | 21 | 293 | 1.556e-07 | 1.81e-05 |
| 41 | REGULATION OF CELL CELL ADHESION | 24 | 380 | 2.139e-07 | 2.369e-05 |
| 42 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 25 | 408 | 2.111e-07 | 2.369e-05 |
| 43 | GROWTH | 25 | 410 | 2.315e-07 | 2.505e-05 |
| 44 | NEURON DEVELOPMENT | 34 | 687 | 2.369e-07 | 2.505e-05 |
| 45 | REGULATION OF EXTENT OF CELL GROWTH | 12 | 101 | 3.871e-07 | 3.915e-05 |
| 46 | REGULATION OF CELL DEVELOPMENT | 38 | 836 | 3.844e-07 | 3.915e-05 |
| 47 | REGULATION OF CELL ACTIVATION | 27 | 484 | 4.438e-07 | 4.302e-05 |
| 48 | POSITIVE REGULATION OF OSSIFICATION | 11 | 84 | 4.387e-07 | 4.302e-05 |
| 49 | REGULATION OF AXONOGENESIS | 15 | 168 | 6.087e-07 | 5.78e-05 |
| 50 | HEART DEVELOPMENT | 26 | 466 | 7.267e-07 | 6.763e-05 |
| 51 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 63 | 1805 | 7.715e-07 | 7.039e-05 |
| 52 | REGULATION OF CELL SIZE | 15 | 172 | 8.227e-07 | 7.361e-05 |
| 53 | CELL MOTILITY | 37 | 835 | 1.016e-06 | 8.751e-05 |
| 54 | LOCALIZATION OF CELL | 37 | 835 | 1.016e-06 | 8.751e-05 |
| 55 | MUSCLE TISSUE DEVELOPMENT | 19 | 275 | 1.062e-06 | 8.983e-05 |
| 56 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 20 | 303 | 1.129e-06 | 9.379e-05 |
| 57 | ORGAN MORPHOGENESIS | 37 | 841 | 1.203e-06 | 9.821e-05 |
| 58 | REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 21 | 337 | 1.538e-06 | 0.0001234 |
| 59 | EMBRYO DEVELOPMENT | 38 | 894 | 1.954e-06 | 0.0001541 |
| 60 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 41 | 1004 | 2.055e-06 | 0.0001593 |
| 61 | MUSCLE STRUCTURE DEVELOPMENT | 24 | 432 | 2.108e-06 | 0.0001608 |
| 62 | REGULATION OF DEVELOPMENTAL GROWTH | 19 | 289 | 2.222e-06 | 0.0001667 |
| 63 | POSITIVE REGULATION OF IMMUNE SYSTEM PROCESS | 37 | 867 | 2.451e-06 | 0.000181 |
| 64 | MESENCHYME DEVELOPMENT | 15 | 190 | 2.876e-06 | 0.0002091 |
| 65 | BIOLOGICAL ADHESION | 41 | 1032 | 4.047e-06 | 0.0002897 |
| 66 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 60 | 1784 | 4.829e-06 | 0.0003404 |
| 67 | NEGATIVE REGULATION OF CELL COMMUNICATION | 45 | 1192 | 5.037e-06 | 0.0003498 |
| 68 | REGULATION OF HOMOTYPIC CELL CELL ADHESION | 19 | 307 | 5.358e-06 | 0.0003666 |
| 69 | REGULATION OF CELLULAR COMPONENT SIZE | 20 | 337 | 5.693e-06 | 0.0003839 |
| 70 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 32 | 724 | 5.961e-06 | 0.0003963 |
| 71 | TAXIS | 24 | 464 | 7.13e-06 | 0.0004673 |
| 72 | NEURON PROJECTION GUIDANCE | 15 | 205 | 7.294e-06 | 0.0004714 |
| 73 | POSITIVE REGULATION OF NEURON PROJECTION DEVELOPMENT | 16 | 232 | 7.72e-06 | 0.000492 |
| 74 | MESENCHYMAL CELL DIFFERENTIATION | 12 | 134 | 7.899e-06 | 0.0004967 |
| 75 | INTRACELLULAR SIGNAL TRANSDUCTION | 54 | 1572 | 8.509e-06 | 0.000521 |
| 76 | POSITIVE REGULATION OF CELL ADHESION | 21 | 376 | 8.48e-06 | 0.000521 |
| 77 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 56 | 1656 | 8.972e-06 | 0.0005422 |
| 78 | TUBE MORPHOGENESIS | 19 | 323 | 1.105e-05 | 0.0006591 |
| 79 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 28 | 609 | 1.133e-05 | 0.0006673 |
| 80 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 62 | 1929 | 1.363e-05 | 0.0007735 |
| 81 | POSITIVE REGULATION OF CELL CELL ADHESION | 16 | 243 | 1.38e-05 | 0.0007735 |
| 82 | UROGENITAL SYSTEM DEVELOPMENT | 18 | 299 | 1.378e-05 | 0.0007735 |
| 83 | STEM CELL DIFFERENTIATION | 14 | 190 | 1.357e-05 | 0.0007735 |
| 84 | POSITIVE REGULATION OF LOCOMOTION | 22 | 420 | 1.416e-05 | 0.000778 |
| 85 | REGULATION OF EPITHELIAL CELL MIGRATION | 13 | 166 | 1.438e-05 | 0.000778 |
| 86 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 48 | 1360 | 1.428e-05 | 0.000778 |
| 87 | DEVELOPMENTAL GROWTH | 19 | 333 | 1.694e-05 | 0.0009058 |
| 88 | SENSORY ORGAN DEVELOPMENT | 24 | 493 | 1.938e-05 | 0.001013 |
| 89 | REGULATION OF CELL PROLIFERATION | 51 | 1496 | 1.917e-05 | 0.001013 |
| 90 | OSSIFICATION | 16 | 251 | 2.059e-05 | 0.001065 |
| 91 | RESPONSE TO WOUNDING | 26 | 563 | 2.171e-05 | 0.00111 |
| 92 | POSITIVE REGULATION OF EPITHELIAL CELL MIGRATION | 10 | 103 | 2.232e-05 | 0.001129 |
| 93 | DEVELOPMENTAL GROWTH INVOLVED IN MORPHOGENESIS | 10 | 104 | 2.429e-05 | 0.001215 |
| 94 | DIGESTIVE TRACT MORPHOGENESIS | 7 | 48 | 2.78e-05 | 0.001376 |
| 95 | REGULATION OF FAT CELL DIFFERENTIATION | 10 | 106 | 2.869e-05 | 0.001376 |
| 96 | CARDIAC SEPTUM DEVELOPMENT | 9 | 85 | 2.868e-05 | 0.001376 |
| 97 | REGULATION OF ANATOMICAL STRUCTURE SIZE | 23 | 472 | 2.846e-05 | 0.001376 |
| 98 | REGULATION OF OSSIFICATION | 13 | 178 | 3.016e-05 | 0.001432 |
| 99 | POSITIVE REGULATION OF ENDOTHELIAL CELL MIGRATION | 8 | 67 | 3.328e-05 | 0.001564 |
| 100 | PLATELET DERIVED GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 6 | 34 | 3.443e-05 | 0.001589 |
| 101 | SYMPATHETIC NERVOUS SYSTEM DEVELOPMENT | 5 | 21 | 3.45e-05 | 0.001589 |
| 102 | ESTABLISHMENT OF CELL POLARITY | 9 | 88 | 3.79e-05 | 0.001726 |
| 103 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 53 | 1618 | 3.822e-05 | 0.001726 |
| 104 | NEURON MIGRATION | 10 | 110 | 3.955e-05 | 0.00177 |
| 105 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 34 | 872 | 4.102e-05 | 0.001818 |
| 106 | EYE DEVELOPMENT | 18 | 326 | 4.33e-05 | 0.001901 |
| 107 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 32 | 801 | 4.391e-05 | 0.001902 |
| 108 | REGULATION OF PROTEIN MODIFICATION PROCESS | 55 | 1710 | 4.416e-05 | 0.001902 |
| 109 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 58 | 1848 | 5.303e-05 | 0.002243 |
| 110 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 12 | 162 | 5.292e-05 | 0.002243 |
| 111 | INNER EAR MORPHOGENESIS | 9 | 92 | 5.399e-05 | 0.002263 |
| 112 | ESTABLISHMENT OR MAINTENANCE OF CELL POLARITY | 11 | 141 | 6.764e-05 | 0.00281 |
| 113 | EMBRYONIC ORGAN MORPHOGENESIS | 16 | 279 | 7.347e-05 | 0.003009 |
| 114 | SEMAPHORIN PLEXIN SIGNALING PATHWAY INVOLVED IN NEURON PROJECTION GUIDANCE | 4 | 13 | 7.373e-05 | 0.003009 |
| 115 | REGULATION OF WNT SIGNALING PATHWAY | 17 | 310 | 7.651e-05 | 0.003096 |
| 116 | MEMBRANE DEPOLARIZATION DURING ACTION POTENTIAL | 6 | 39 | 7.723e-05 | 0.003098 |
| 117 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 56 | 1791 | 8.038e-05 | 0.003197 |
| 118 | MEMBRANE ASSEMBLY | 5 | 25 | 8.473e-05 | 0.003341 |
| 119 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 36 | 983 | 8.75e-05 | 0.003421 |
| 120 | REGULATION OF ACTIVATED T CELL PROLIFERATION | 6 | 40 | 8.945e-05 | 0.003469 |
| 121 | VASCULAR ENDOTHELIAL GROWTH FACTOR SIGNALING PATHWAY | 4 | 14 | 0.0001017 | 0.003911 |
| 122 | NEGATIVE REGULATION OF FAT CELL DIFFERENTIATION | 6 | 41 | 0.0001032 | 0.003934 |
| 123 | VASCULOGENESIS | 7 | 59 | 0.0001081 | 0.004089 |
| 124 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 40 | 1152 | 0.0001109 | 0.004162 |
| 125 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 37 | 1036 | 0.0001159 | 0.004247 |
| 126 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 37 | 1036 | 0.0001159 | 0.004247 |
| 127 | TUBE DEVELOPMENT | 24 | 552 | 0.0001144 | 0.004247 |
| 128 | AUTONOMIC NERVOUS SYSTEM DEVELOPMENT | 6 | 42 | 0.0001185 | 0.004307 |
| 129 | REGULATION OF LEUKOCYTE DIFFERENTIATION | 14 | 232 | 0.0001206 | 0.004351 |
| 130 | MEMBRANE DEPOLARIZATION | 7 | 61 | 0.0001339 | 0.004793 |
| 131 | REGULATION OF KINASE ACTIVITY | 30 | 776 | 0.0001363 | 0.004843 |
| 132 | POSITIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 11 | 154 | 0.0001491 | 0.005256 |
| 133 | SINGLE ORGANISM CELL ADHESION | 21 | 459 | 0.0001527 | 0.005341 |
| 134 | HEAD DEVELOPMENT | 28 | 709 | 0.0001607 | 0.005579 |
| 135 | REGULATION OF CARTILAGE DEVELOPMENT | 7 | 63 | 0.0001645 | 0.005635 |
| 136 | SENSORY ORGAN MORPHOGENESIS | 14 | 239 | 0.0001647 | 0.005635 |
| 137 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 39 | 1135 | 0.0001681 | 0.005711 |
| 138 | REGULATION OF T CELL DIFFERENTIATION | 9 | 107 | 0.0001746 | 0.005887 |
| 139 | STEM CELL DIVISION | 5 | 29 | 0.0001782 | 0.005903 |
| 140 | HEART MORPHOGENESIS | 13 | 212 | 0.0001789 | 0.005903 |
| 141 | RETINA VASCULATURE DEVELOPMENT IN CAMERA TYPE EYE | 4 | 16 | 0.0001795 | 0.005903 |
| 142 | REGULATION OF TRANSPORT | 55 | 1804 | 0.0001801 | 0.005903 |
| 143 | REGULATION OF ORGAN MORPHOGENESIS | 14 | 242 | 0.0001875 | 0.0061 |
| 144 | IMMUNE SYSTEM PROCESS | 59 | 1984 | 0.0001986 | 0.006378 |
| 145 | REGULATION OF CHONDROCYTE DIFFERENTIATION | 6 | 46 | 0.0001987 | 0.006378 |
| 146 | MEMBRANE BIOGENESIS | 5 | 30 | 0.0002106 | 0.006711 |
| 147 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 20 | 437 | 0.0002174 | 0.00688 |
| 148 | REGULATION OF BODY FLUID LEVELS | 22 | 506 | 0.0002195 | 0.006886 |
| 149 | CELL GROWTH | 10 | 135 | 0.0002205 | 0.006886 |
| 150 | MUSCLE ORGAN DEVELOPMENT | 15 | 277 | 0.0002294 | 0.007117 |
| 151 | EMBRYONIC ORGAN DEVELOPMENT | 19 | 406 | 0.0002352 | 0.007248 |
| 152 | HINDBRAIN DEVELOPMENT | 10 | 137 | 0.0002485 | 0.00746 |
| 153 | SKELETAL MUSCLE ORGAN DEVELOPMENT | 10 | 137 | 0.0002485 | 0.00746 |
| 154 | REGULATION OF IMMUNE SYSTEM PROCESS | 45 | 1403 | 0.0002481 | 0.00746 |
| 155 | EAR MORPHOGENESIS | 9 | 112 | 0.0002467 | 0.00746 |
| 156 | IMMUNE SYSTEM DEVELOPMENT | 24 | 582 | 0.0002519 | 0.00748 |
| 157 | REGULATION OF INTERLEUKIN 2 PRODUCTION | 6 | 48 | 0.0002524 | 0.00748 |
| 158 | POSITIVE REGULATION OF CELL ACTIVATION | 16 | 311 | 0.0002557 | 0.007529 |
| 159 | REGULATION OF ACTIN FILAMENT BASED PROCESS | 16 | 312 | 0.000265 | 0.007707 |
| 160 | PROTEIN AUTOPHOSPHORYLATION | 12 | 192 | 0.0002644 | 0.007707 |
| 161 | REGULATION OF CELL SHAPE | 10 | 139 | 0.0002794 | 0.008075 |
| 162 | REGULATION OF HEMOPOIESIS | 16 | 314 | 0.0002846 | 0.008169 |
| 163 | CONNECTIVE TISSUE DEVELOPMENT | 12 | 194 | 0.0002908 | 0.008169 |
| 164 | REGULATION OF ACTIN CYTOSKELETON REORGANIZATION | 5 | 32 | 0.0002886 | 0.008169 |
| 165 | NEGATIVE REGULATION OF CELL ADHESION | 13 | 223 | 0.0002932 | 0.008169 |
| 166 | BEHAVIOR | 22 | 516 | 0.0002878 | 0.008169 |
| 167 | POSITIVE REGULATION OF AXONOGENESIS | 7 | 69 | 0.0002916 | 0.008169 |
| 168 | CARDIAC MUSCLE TISSUE DEVELOPMENT | 10 | 140 | 0.000296 | 0.008198 |
| 169 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 15 | 285 | 0.0003111 | 0.008565 |
| 170 | CENTRAL NERVOUS SYSTEM NEURON DEVELOPMENT | 7 | 70 | 0.0003189 | 0.008729 |
| 171 | REGULATION OF ION TRANSPORT | 24 | 592 | 0.0003228 | 0.008783 |
| 172 | REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 12 | 197 | 0.0003345 | 0.00901 |
| 173 | EMBRYONIC DIGESTIVE TRACT DEVELOPMENT | 5 | 33 | 0.000335 | 0.00901 |
| 174 | ACTIN FILAMENT BASED MOVEMENT | 8 | 93 | 0.0003419 | 0.009144 |
| 175 | REGULATION OF GROWTH | 25 | 633 | 0.0003591 | 0.009548 |
| 176 | TELENCEPHALON DEVELOPMENT | 13 | 228 | 0.0003629 | 0.009594 |
| 177 | REGULATION OF NON CANONICAL WNT SIGNALING PATHWAY | 4 | 19 | 0.0003659 | 0.009618 |
| 178 | CARDIAC CHAMBER DEVELOPMENT | 10 | 144 | 0.000371 | 0.009697 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | RNA POLYMERASE II TRANSCRIPTION FACTOR ACTIVITY SEQUENCE SPECIFIC DNA BINDING | 36 | 629 | 2.5e-09 | 1.822e-06 |
| 2 | NUCLEIC ACID BINDING TRANSCRIPTION FACTOR ACTIVITY | 53 | 1199 | 3.922e-09 | 1.822e-06 |
| 3 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION REGULATORY REGION SEQUENCE SPECIFIC BINDING | 22 | 315 | 1.235e-07 | 3.824e-05 |
| 4 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 22 | 328 | 2.487e-07 | 5.775e-05 |
| 5 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 17 | 226 | 1.232e-06 | 0.0002289 |
| 6 | REGULATORY REGION NUCLEIC ACID BINDING | 36 | 818 | 1.678e-06 | 0.0002598 |
| 7 | SEQUENCE SPECIFIC DNA BINDING | 40 | 1037 | 1.071e-05 | 0.001422 |
| 8 | SEMAPHORIN RECEPTOR ACTIVITY | 4 | 11 | 3.505e-05 | 0.00407 |
| 9 | DOUBLE STRANDED DNA BINDING | 31 | 764 | 4.341e-05 | 0.004481 |
| 10 | TRANSCRIPTIONAL REPRESSOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION REGULATORY REGION SEQUENCE SPECIFIC BINDING | 12 | 168 | 7.527e-05 | 0.006992 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CELL JUNCTION | 48 | 1151 | 1.362e-07 | 7.952e-05 |
| 2 | CYTOPLASMIC SIDE OF MEMBRANE | 13 | 170 | 1.855e-05 | 0.005416 |
| 3 | SEMAPHORIN RECEPTOR COMPLEX | 4 | 11 | 3.505e-05 | 0.00571 |
| 4 | CELL PROJECTION | 57 | 1786 | 3.911e-05 | 0.00571 |
| 5 | SITE OF POLARIZED GROWTH | 11 | 149 | 0.0001112 | 0.009278 |
| 6 | EXTRINSIC COMPONENT OF CYTOPLASMIC SIDE OF PLASMA MEMBRANE | 9 | 98 | 8.87e-05 | 0.009278 |
| 7 | CELL LEADING EDGE | 18 | 350 | 0.0001074 | 0.009278 |
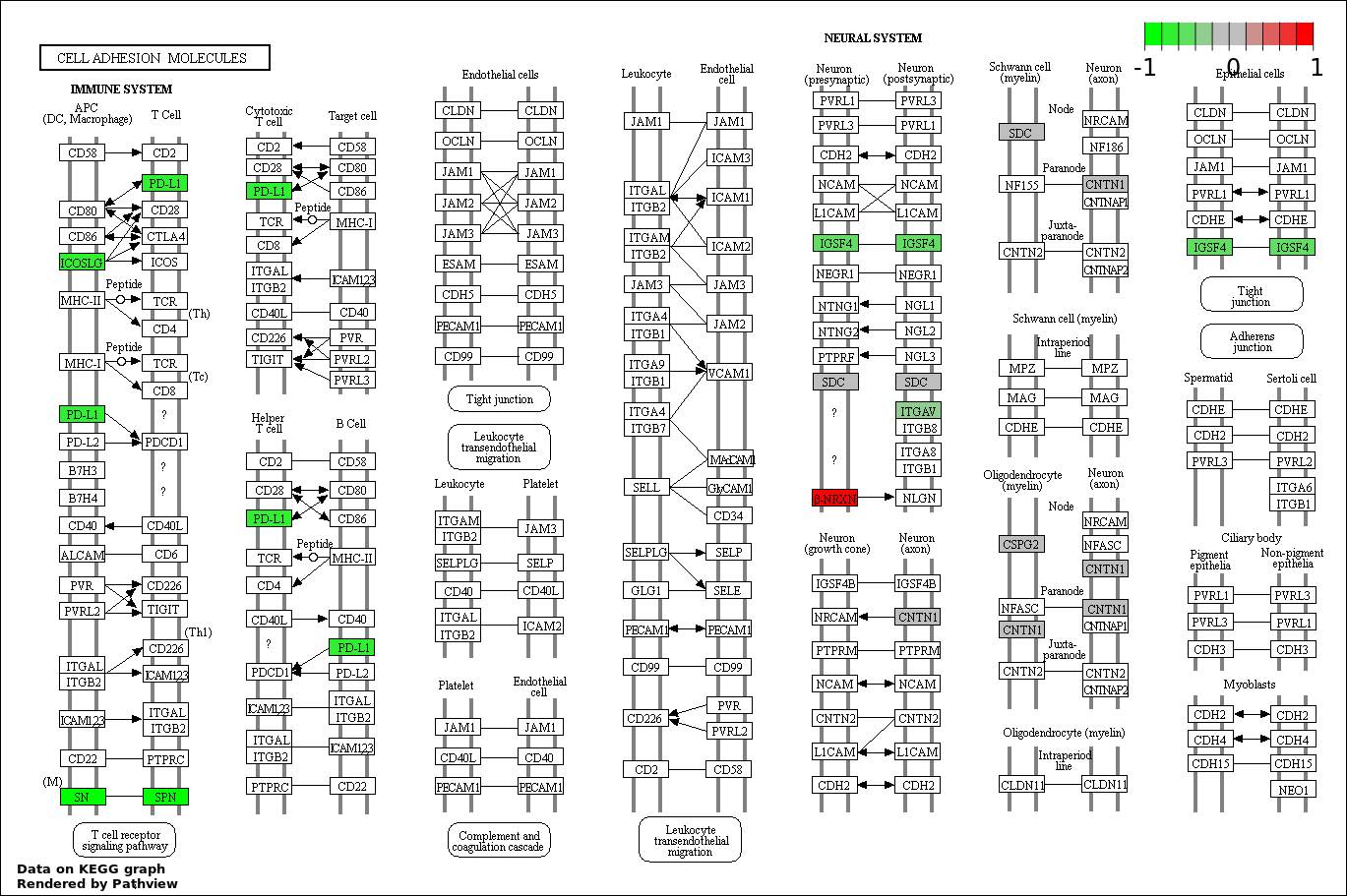
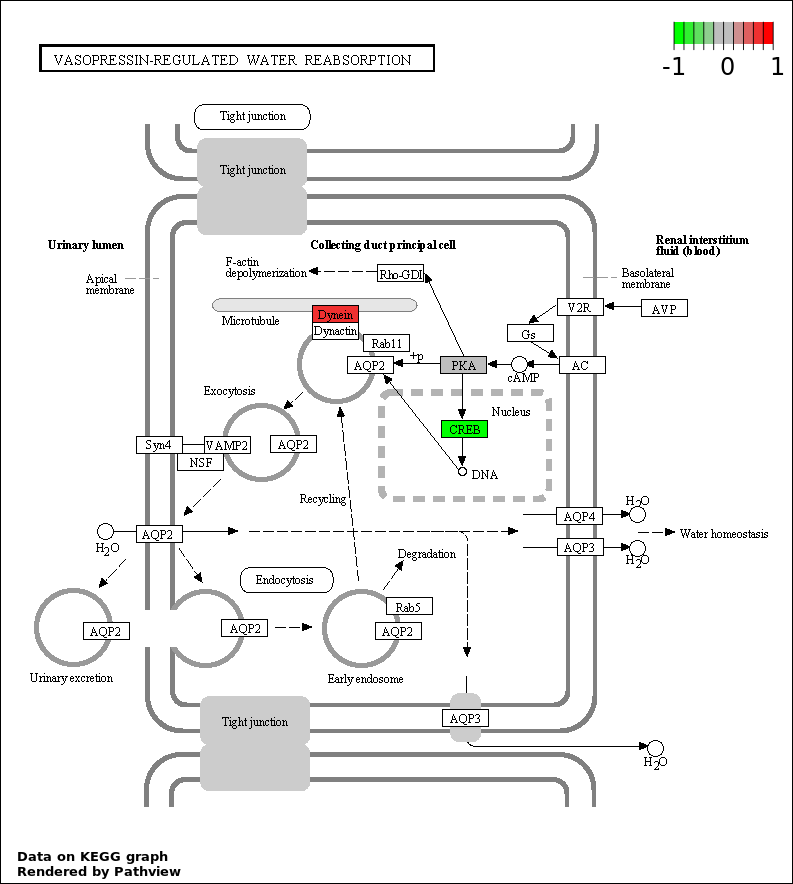
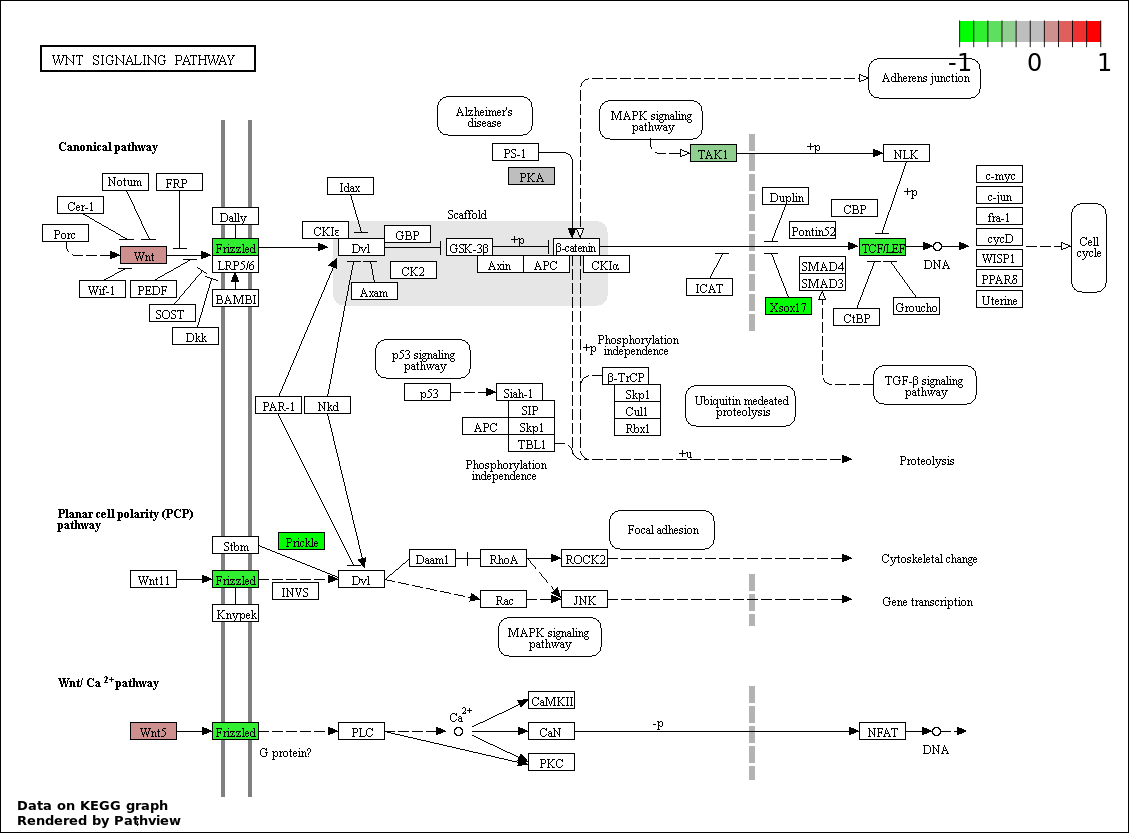
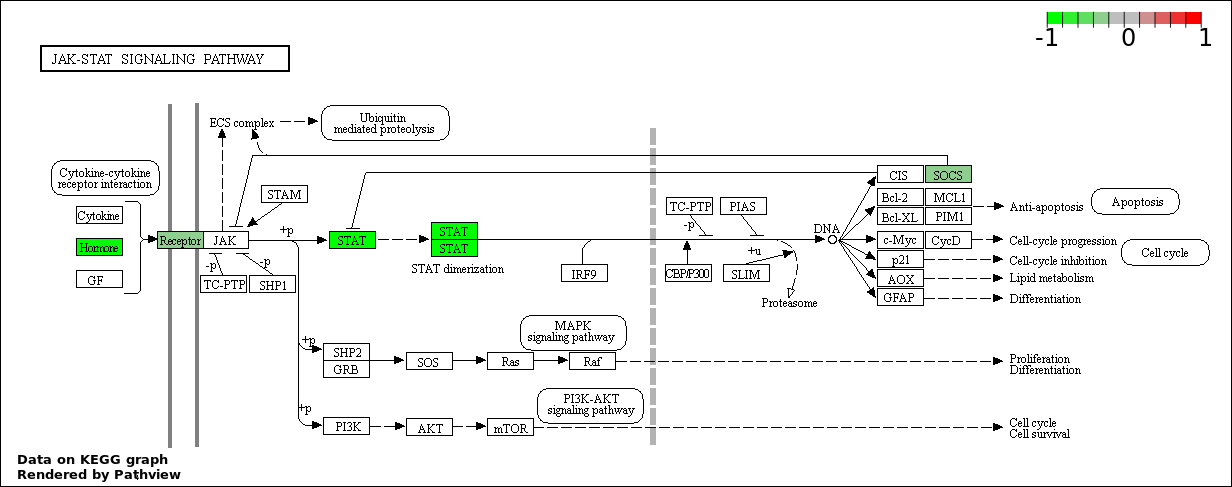
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04514_Cell_adhesion_molecules_.CAMs. | 10 | 136 | 0.0002342 | 0.04215 | |
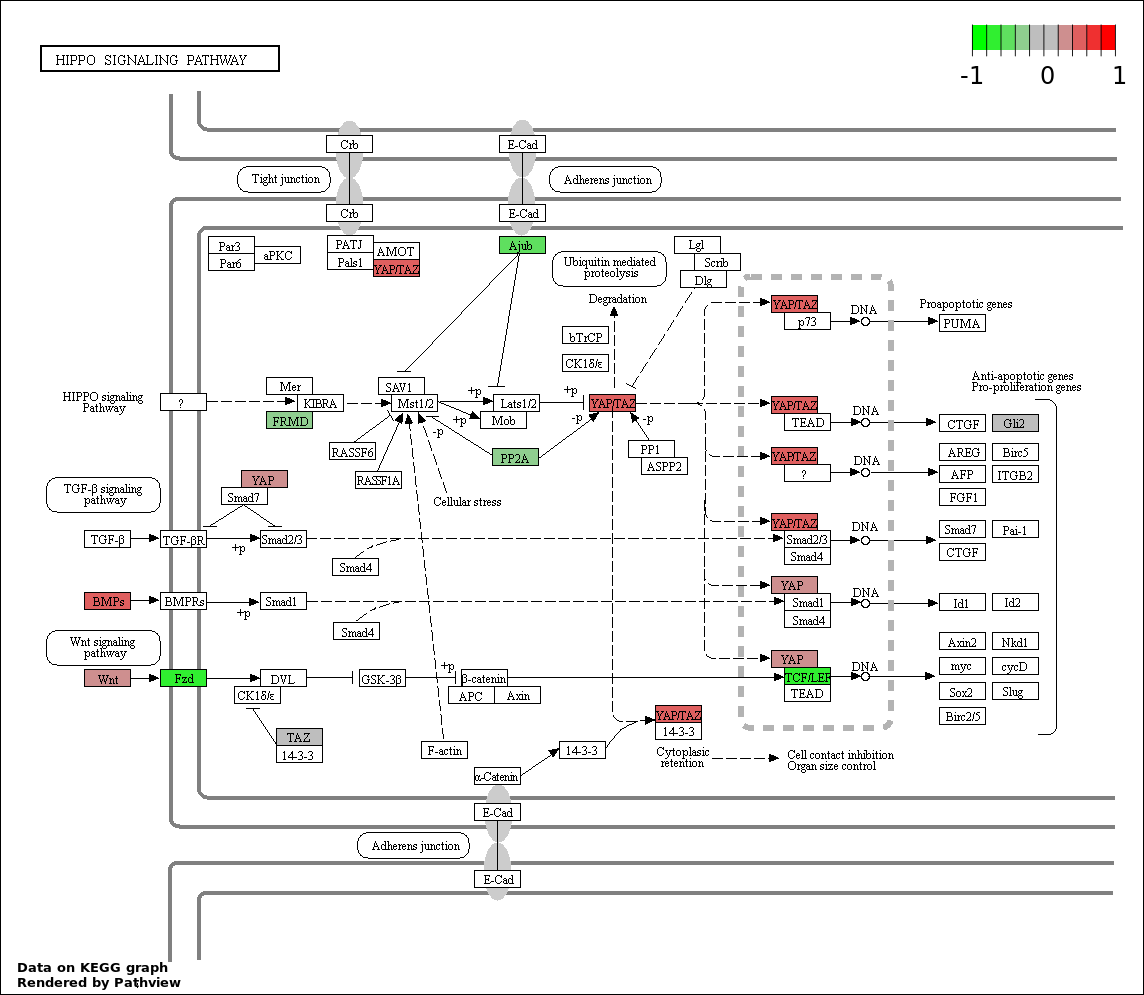
| 2 | hsa04390_Hippo_signaling_pathway | 10 | 154 | 0.0006299 | 0.05669 | |
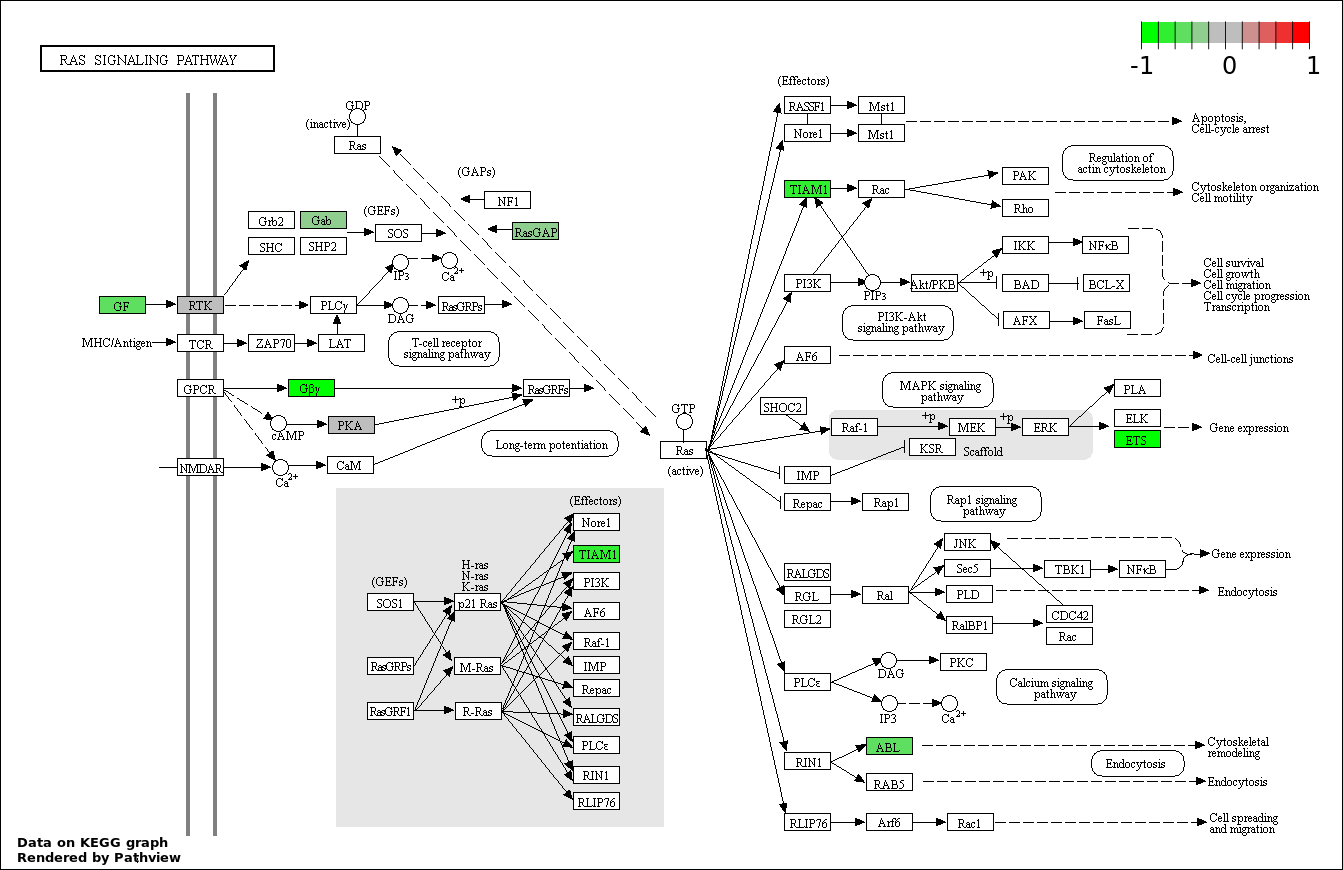
| 3 | hsa04014_Ras_signaling_pathway | 12 | 236 | 0.001628 | 0.09765 | |
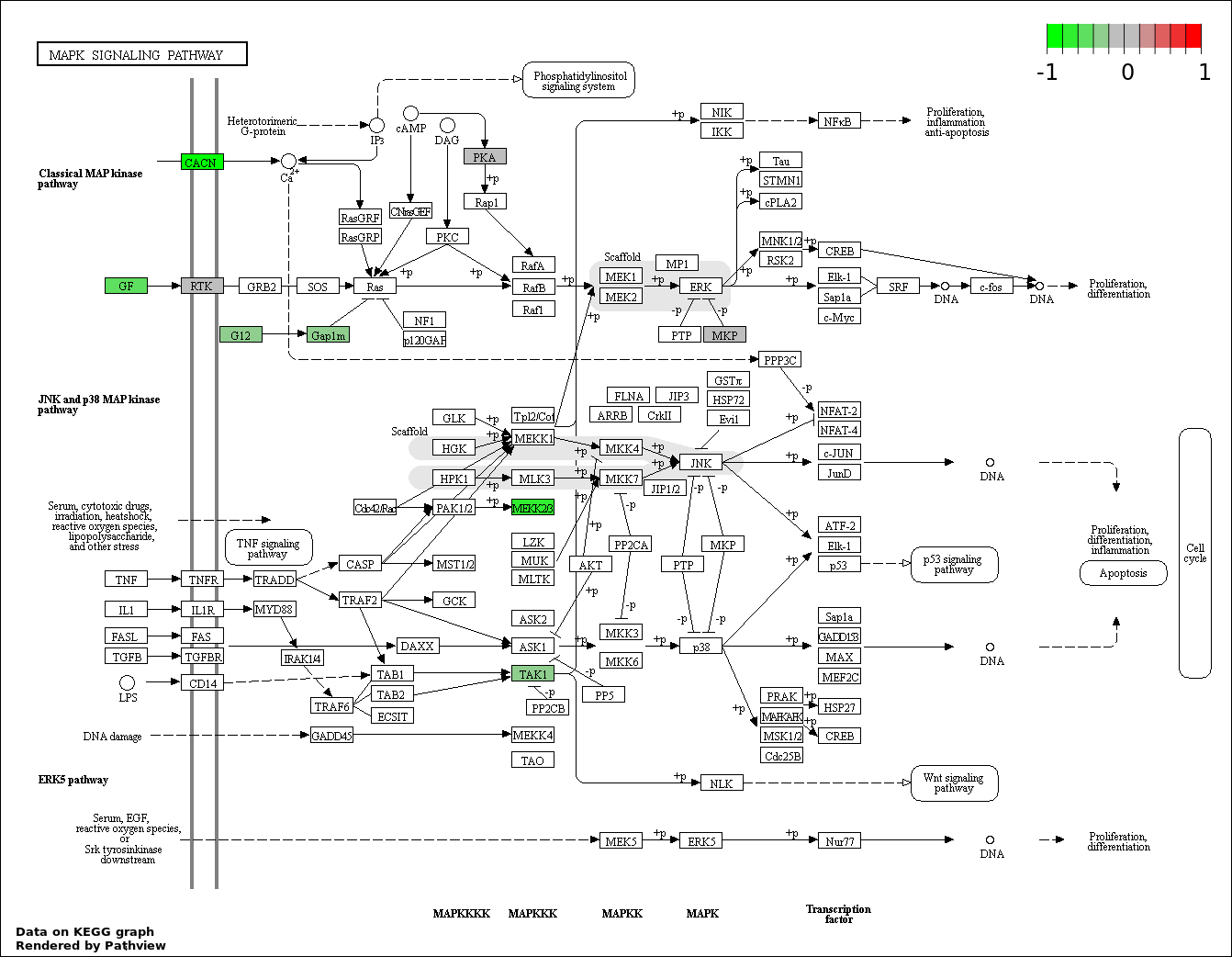
| 4 | hsa04010_MAPK_signaling_pathway | 12 | 268 | 0.004583 | 0.2062 | |
| 5 | hsa04962_Vasopressin.regulated_water_reabsorption | 4 | 44 | 0.008898 | 0.3203 | |
| 6 | hsa04310_Wnt_signaling_pathway | 7 | 151 | 0.02306 | 0.4816 | |
| 7 | hsa04012_ErbB_signaling_pathway | 5 | 87 | 0.02316 | 0.4816 | |
| 8 | hsa04630_Jak.STAT_signaling_pathway | 7 | 155 | 0.02613 | 0.4816 | |
| 9 | hsa04130_SNARE_interactions_in_vesicular_transport | 3 | 36 | 0.02898 | 0.4816 | |
| 10 | hsa00604_Glycosphingolipid_biosynthesis_._ganglio_series | 2 | 15 | 0.03088 | 0.4816 | |
| 11 | hsa04722_Neurotrophin_signaling_pathway | 6 | 127 | 0.03155 | 0.4816 | |
| 12 | hsa04151_PI3K_AKT_signaling_pathway | 12 | 351 | 0.03214 | 0.4816 | |
| 13 | hsa04360_Axon_guidance | 6 | 130 | 0.03478 | 0.4816 | |
| 14 | hsa04916_Melanogenesis | 5 | 101 | 0.04037 | 0.519 | |
| 15 | hsa04810_Regulation_of_actin_cytoskeleton | 8 | 214 | 0.04738 | 0.54 | |
| 16 | hsa04020_Calcium_signaling_pathway | 7 | 177 | 0.048 | 0.54 | |
| 17 | hsa04340_Hedgehog_signaling_pathway | 3 | 56 | 0.08611 | 0.8615 | |
| 18 | hsa04540_Gap_junction | 4 | 90 | 0.08706 | 0.8615 | |
| 19 | hsa04380_Osteoclast_differentiation | 5 | 128 | 0.09094 | 0.8615 | |
| 20 | hsa04530_Tight_junction | 5 | 133 | 0.1028 | 0.925 | |
| 21 | hsa04062_Chemokine_signaling_pathway | 6 | 189 | 0.1416 | 1 | |
| 22 | hsa04660_T_cell_receptor_signaling_pathway | 4 | 108 | 0.1423 | 1 | |
| 23 | hsa04520_Adherens_junction | 3 | 73 | 0.1547 | 1 | |
| 24 | hsa04145_Phagosome | 5 | 156 | 0.1661 | 1 | |
| 25 | hsa04512_ECM.receptor_interaction | 3 | 85 | 0.2105 | 1 | |
| 26 | hsa04110_Cell_cycle | 4 | 128 | 0.2156 | 1 | |
| 27 | hsa04672_Intestinal_immune_network_for_IgA_production | 2 | 49 | 0.2313 | 1 | |
| 28 | hsa04912_GnRH_signaling_pathway | 3 | 101 | 0.2901 | 1 | |
| 29 | hsa04621_NOD.like_receptor_signaling_pathway | 2 | 59 | 0.3004 | 1 | |
| 30 | hsa04510_Focal_adhesion | 5 | 200 | 0.3158 | 1 | |
| 31 | hsa04144_Endocytosis | 5 | 203 | 0.3268 | 1 | |
| 32 | hsa04270_Vascular_smooth_muscle_contraction | 3 | 116 | 0.3664 | 1 | |
| 33 | hsa04720_Long.term_potentiation | 2 | 70 | 0.3751 | 1 | |
| 34 | hsa04730_Long.term_depression | 2 | 70 | 0.3751 | 1 | |
| 35 | hsa04622_RIG.I.like_receptor_signaling_pathway | 2 | 71 | 0.3817 | 1 | |
| 36 | hsa04612_Antigen_processing_and_presentation | 2 | 78 | 0.4273 | 1 | |
| 37 | hsa04664_Fc_epsilon_RI_signaling_pathway | 2 | 79 | 0.4337 | 1 | |
| 38 | hsa04974_Protein_digestion_and_absorption | 2 | 81 | 0.4463 | 1 | |
| 39 | hsa04350_TGF.beta_signaling_pathway | 2 | 85 | 0.4711 | 1 | |
| 40 | hsa04910_Insulin_signaling_pathway | 3 | 138 | 0.475 | 1 | |
| 41 | hsa04640_Hematopoietic_cell_lineage | 2 | 88 | 0.4892 | 1 | |
| 42 | hsa04210_Apoptosis | 2 | 89 | 0.4952 | 1 | |
| 43 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 2 | 95 | 0.53 | 1 | |
| 44 | hsa04114_Oocyte_meiosis | 2 | 114 | 0.6291 | 1 | |
| 45 | hsa04120_Ubiquitin_mediated_proteolysis | 2 | 139 | 0.7337 | 1 | |
| 46 | hsa04740_Olfactory_transduction | 2 | 388 | 0.9946 | 1 |