This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-181b-5p | ABCA5 | 0.49 | 0.00105 | -0.14 | 0.30791 | mirMAP | -0.26 | 0 | NA | |
| 2 | hsa-miR-181b-5p | ABHD13 | 0.49 | 0.00105 | -0.68 | 0 | MirTarget | -0.11 | 0.00024 | NA | |
| 3 | hsa-miR-181b-5p | ABHD15 | 0.49 | 0.00105 | -1.3 | 0 | mirMAP | -0.44 | 0 | NA | |
| 4 | hsa-miR-181b-5p | ABTB2 | 0.49 | 0.00105 | -0.43 | 0.0031 | MirTarget; miRNATAP | -0.34 | 0 | NA | |
| 5 | hsa-miR-181b-5p | ACAD8 | 0.49 | 0.00105 | -0.52 | 0 | MirTarget | -0.15 | 0 | NA | |
| 6 | hsa-miR-181b-5p | ACSL1 | 0.49 | 0.00105 | -2.33 | 0 | MirTarget | -0.69 | 0 | NA | |
| 7 | hsa-miR-181b-5p | ACSL6 | 0.49 | 0.00105 | 0.39 | 0.26458 | miRNATAP | -0.75 | 0 | NA | |
| 8 | hsa-miR-181b-5p | ACSS3 | 0.49 | 0.00105 | -1.23 | 0.00019 | MirTarget | -1 | 0 | NA | |
| 9 | hsa-miR-181b-5p | ACVR1C | 0.49 | 0.00105 | -2.2 | 0 | miRNATAP | -0.67 | 0 | NA | |
| 10 | hsa-miR-181b-5p | ADCY1 | 0.49 | 0.00105 | -2.7 | 0 | miRNATAP | -0.49 | 9.0E-5 | NA | |
| 11 | hsa-miR-181b-5p | ADCY9 | 0.49 | 0.00105 | -0.11 | 0.30518 | miRNATAP | -0.14 | 7.0E-5 | 24269684 | miR 181b promotes cell proliferation and reduces apoptosis by repressing the expression of adenylyl cyclase 9 AC9 in cervical cancer cells; Phenotypic experiments indicated that miR-181b and AC9 exerted opposite effects on cell proliferation and apoptosis |
| 12 | hsa-miR-181b-5p | ADM | 0.49 | 0.00105 | -1.26 | 0 | MirTarget; miRNATAP | -0.18 | 0.00297 | NA | |
| 13 | hsa-miR-181b-5p | AFF4 | 0.49 | 0.00105 | -0.64 | 0 | mirMAP; miRNATAP | -0.14 | 1.0E-5 | NA | |
| 14 | hsa-miR-181b-5p | AFG3L2 | 0.49 | 0.00105 | -0.29 | 0.0003 | MirTarget; miRNATAP | -0.14 | 0 | NA | |
| 15 | hsa-miR-181b-5p | AGFG2 | 0.49 | 0.00105 | -0.02 | 0.87061 | mirMAP | -0.19 | 0 | NA | |
| 16 | hsa-miR-181b-5p | AGGF1 | 0.49 | 0.00105 | 0.14 | 0.03026 | mirMAP | -0.12 | 0 | NA | |
| 17 | hsa-miR-181b-5p | AGPAT3 | 0.49 | 0.00105 | -0.18 | 0.05507 | MirTarget | -0.11 | 0.00016 | NA | |
| 18 | hsa-miR-181b-5p | AKR1C1 | 0.49 | 0.00105 | -0.12 | 0.65413 | mirMAP | -0.52 | 0 | NA | |
| 19 | hsa-miR-21-5p | AKT2 | 1.51 | 0 | -0.34 | 1.0E-5 | miRNAWalker2 validate | -0.25 | 0 | NA | |
| 20 | hsa-miR-330-5p | AKT2 | 0.44 | 0.00533 | -0.34 | 1.0E-5 | miRanda | -0.1 | 1.0E-5 | NA | |
| 21 | hsa-miR-342-3p | AKT2 | -0.32 | 0.04498 | -0.34 | 1.0E-5 | mirMAP | -0.12 | 0 | NA | |
| 22 | hsa-miR-106b-5p | AKT3 | 0.65 | 0 | -0.66 | 0.00047 | miRNATAP | -0.26 | 0.00148 | NA | |
| 23 | hsa-miR-107 | AKT3 | 0.24 | 0.01708 | -0.66 | 0.00047 | PITA; miRanda | -0.6 | 0 | NA | |
| 24 | hsa-miR-122-5p | AKT3 | -1.24 | 0 | -0.66 | 0.00047 | miRNAWalker2 validate; miRTarBase | -0.28 | 0 | 24244539 | miR 122 regulates tumorigenesis in hepatocellular carcinoma by targeting AKT3; Here we identify AKT3 as a novel and direct target of miR-122; Restoration of miR-122 expression in HCC cell lines decreases AKT3 levels inhibits cell migration and proliferation and induces apoptosis; These anti-tumor phenotypes can be rescued by reconstitution of AKT3 expression indicating the essential role of AKT3 in miR-122 mediated HCC transformation; Our data strongly suggest that miR-122 is a tumor suppressor that targets AKT3 to regulate tumorigenesis in HCCs and a potential therapeutic candidate for liver cancer |
| 25 | hsa-miR-146b-5p | AKT3 | 0.42 | 0.04574 | -0.66 | 0.00047 | miRNAWalker2 validate | -0.16 | 0.00026 | NA | |
| 26 | hsa-miR-15a-5p | AKT3 | 0.35 | 0.00077 | -0.66 | 0.00047 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.27 | 0.00291 | NA | |
| 27 | hsa-miR-17-3p | AKT3 | 0.41 | 0.00422 | -0.66 | 0.00047 | miRNATAP | -0.35 | 0 | NA | |
| 28 | hsa-miR-17-5p | AKT3 | 0.7 | 2.0E-5 | -0.66 | 0.00047 | TargetScan; miRNATAP | -0.29 | 0 | NA | |
| 29 | hsa-miR-20a-5p | AKT3 | 0.85 | 0 | -0.66 | 0.00047 | miRNATAP | -0.29 | 0 | NA | |
| 30 | hsa-miR-32-3p | AKT3 | 0.22 | 0.20722 | -0.66 | 0.00047 | mirMAP | -0.23 | 4.0E-5 | NA | |
| 31 | hsa-miR-33a-3p | AKT3 | -0.68 | 1.0E-5 | -0.66 | 0.00047 | mirMAP | -0.23 | 0.0001 | NA | |
| 32 | hsa-miR-362-3p | AKT3 | 0.81 | 0 | -0.66 | 0.00047 | miRanda | -0.22 | 0.00083 | NA | |
| 33 | hsa-miR-362-5p | AKT3 | 0.72 | 2.0E-5 | -0.66 | 0.00047 | PITA; TargetScan; miRNATAP | -0.15 | 0.00734 | NA | |
| 34 | hsa-miR-374a-5p | AKT3 | 0.02 | 0.86978 | -0.66 | 0.00047 | mirMAP | -0.28 | 0.00294 | NA | |
| 35 | hsa-miR-421 | AKT3 | 0.94 | 0 | -0.66 | 0.00047 | miRanda; mirMAP | -0.13 | 0.01091 | NA | |
| 36 | hsa-miR-501-3p | AKT3 | 1 | 0 | -0.66 | 0.00047 | miRNATAP | -0.12 | 0.04823 | NA | |
| 37 | hsa-miR-502-3p | AKT3 | 0.66 | 0 | -0.66 | 0.00047 | miRNATAP | -0.26 | 0.0008 | NA | |
| 38 | hsa-miR-616-5p | AKT3 | 0.15 | 0.40284 | -0.66 | 0.00047 | mirMAP | -0.2 | 0.0001 | NA | |
| 39 | hsa-miR-93-5p | AKT3 | 1.4 | 0 | -0.66 | 0.00047 | miRNATAP | -0.28 | 1.0E-5 | NA | |
| 40 | hsa-miR-181b-5p | ANKRD50 | 0.49 | 0.00105 | -0.25 | 0.04433 | MirTarget; miRNATAP | -0.11 | 0.00648 | NA | |
| 41 | hsa-miR-181b-5p | AP1G1 | 0.49 | 0.00105 | -0.48 | 0 | miRNATAP | -0.14 | 0 | NA | |
| 42 | hsa-miR-181b-5p | API5 | 0.49 | 0.00105 | -0.32 | 0 | MirTarget | -0.16 | 0 | NA | |
| 43 | hsa-miR-181b-5p | AR | 0.49 | 0.00105 | -2.66 | 0 | miRNATAP | -1.3 | 0 | NA | |
| 44 | hsa-miR-181b-5p | ARFGEF2 | 0.49 | 0.00105 | 0.04 | 0.70711 | MirTarget; miRNATAP | -0.11 | 0.00402 | NA | |
| 45 | hsa-miR-181b-5p | ARHGAP32 | 0.49 | 0.00105 | -0.2 | 0.09921 | mirMAP | -0.15 | 0.00021 | NA | |
| 46 | hsa-miR-181b-5p | ARHGAP5 | 0.49 | 0.00105 | -0.39 | 8.0E-5 | mirMAP | -0.16 | 0 | NA | |
| 47 | hsa-miR-181b-5p | ARIH1 | 0.49 | 0.00105 | -0.13 | 0.06993 | MirTarget; mirMAP | -0.13 | 0 | NA | |
| 48 | hsa-miR-181b-5p | ARRDC3 | 0.49 | 0.00105 | -0.7 | 0 | MirTarget; mirMAP; miRNATAP | -0.2 | 7.0E-5 | NA | |
| 49 | hsa-miR-181b-5p | ASB13 | 0.49 | 0.00105 | -0.76 | 0 | miRNAWalker2 validate | -0.38 | 0 | NA | |
| 50 | hsa-miR-181b-5p | ASPA | 0.49 | 0.00105 | -2.88 | 0 | mirMAP | -0.46 | 0 | NA | |
| 51 | hsa-miR-181b-5p | ATE1 | 0.49 | 0.00105 | -0.6 | 0.0009 | mirMAP | -0.15 | 0.01316 | NA | |
| 52 | hsa-miR-181b-5p | ATP11C | 0.49 | 0.00105 | -1.6 | 0 | MirTarget; miRNATAP | -0.56 | 0 | NA | |
| 53 | hsa-miR-181b-5p | ATXN1 | 0.49 | 0.00105 | -0.15 | 0.16412 | miRNATAP | -0.11 | 0.00167 | NA | |
| 54 | hsa-miR-181b-5p | ATXN7 | 0.49 | 0.00105 | -0.47 | 0 | MirTarget | -0.11 | 0.00015 | NA | |
| 55 | hsa-miR-181b-5p | BCL6 | 0.49 | 0.00105 | -0.38 | 0.00305 | miRNATAP | -0.19 | 0 | NA | |
| 56 | hsa-miR-181b-5p | BHLHE40 | 0.49 | 0.00105 | -1.43 | 0 | MirTarget; miRNATAP | -0.13 | 0.01684 | NA | |
| 57 | hsa-miR-181b-5p | BRWD1 | 0.49 | 0.00105 | -0.47 | 0 | MirTarget | -0.15 | 0 | NA | |
| 58 | hsa-miR-181b-5p | C16orf87 | 0.49 | 0.00105 | -0.85 | 0 | miRNATAP | -0.23 | 0 | NA | |
| 59 | hsa-miR-181b-5p | C17orf51 | 0.49 | 0.00105 | -0.67 | 0.00164 | mirMAP | -0.15 | 0.03042 | NA | |
| 60 | hsa-miR-181b-5p | C2orf69 | 0.49 | 0.00105 | -0.19 | 0.04079 | miRNATAP | -0.25 | 0 | NA | |
| 61 | hsa-miR-181b-5p | C5orf51 | 0.49 | 0.00105 | 0.25 | 0.00285 | mirMAP | -0.11 | 2.0E-5 | NA | |
| 62 | hsa-miR-181b-5p | CA13 | 0.49 | 0.00105 | -1.26 | 0 | mirMAP | -0.29 | 2.0E-5 | NA | |
| 63 | hsa-miR-15a-5p | CAB39 | 0.35 | 0.00077 | -0.15 | 0.00963 | miRNATAP | -0.11 | 8.0E-5 | NA | |
| 64 | hsa-miR-16-1-3p | CAB39 | 0.39 | 0.00112 | -0.15 | 0.00963 | MirTarget | -0.12 | 0 | NA | |
| 65 | hsa-miR-193a-3p | CAB39 | -0.12 | 0.30939 | -0.15 | 0.00963 | MirTarget; miRanda | -0.12 | 0 | NA | |
| 66 | hsa-miR-192-3p | CAB39L | -0.64 | 0.00027 | -0.5 | 1.0E-5 | mirMAP | -0.12 | 0.00012 | NA | |
| 67 | hsa-miR-192-5p | CAB39L | -0.5 | 0.00345 | -0.5 | 1.0E-5 | miRNAWalker2 validate | -0.17 | 0 | NA | |
| 68 | hsa-miR-361-5p | CAB39L | 0.23 | 0.00962 | -0.5 | 1.0E-5 | MirTarget; miRanda | -0.2 | 0.00208 | NA | |
| 69 | hsa-miR-362-3p | CAB39L | 0.81 | 0 | -0.5 | 1.0E-5 | miRanda | -0.15 | 0.00018 | NA | |
| 70 | hsa-miR-374a-3p | CAB39L | -0.21 | 0.06235 | -0.5 | 1.0E-5 | MirTarget | -0.12 | 0.02355 | NA | |
| 71 | hsa-miR-590-3p | CAB39L | -0.47 | 2.0E-5 | -0.5 | 1.0E-5 | miRanda | -0.1 | 0.04487 | NA | |
| 72 | hsa-miR-590-5p | CAB39L | -0.1 | 0.31003 | -0.5 | 1.0E-5 | miRanda | -0.14 | 0.01477 | NA | |
| 73 | hsa-miR-618 | CAB39L | 0.14 | 0.51715 | -0.5 | 1.0E-5 | mirMAP | -0.1 | 0.0003 | NA | |
| 74 | hsa-miR-181b-5p | CABLES1 | 0.49 | 0.00105 | -0.24 | 0.1576 | mirMAP | -0.13 | 0.01656 | NA | |
| 75 | hsa-miR-181b-5p | CASK | 0.49 | 0.00105 | 0.43 | 0.00011 | mirMAP | -0.11 | 0.0027 | NA | |
| 76 | hsa-miR-181b-5p | CAT | 0.49 | 0.00105 | -1.65 | 0 | miRNAWalker2 validate | -0.5 | 0 | NA | |
| 77 | hsa-miR-181b-5p | CBX7 | 0.49 | 0.00105 | -0.08 | 0.52937 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.3 | 0 | NA | |
| 78 | hsa-miR-181b-5p | CDC42BPA | 0.49 | 0.00105 | 0.05 | 0.70443 | MirTarget; miRNATAP | -0.22 | 0 | NA | |
| 79 | hsa-miR-181b-5p | CDC5L | 0.49 | 0.00105 | 0.18 | 0.02901 | mirMAP | -0.11 | 4.0E-5 | NA | |
| 80 | hsa-miR-181b-5p | CEP120 | 0.49 | 0.00105 | -0.39 | 2.0E-5 | MirTarget | -0.11 | 0.00024 | NA | |
| 81 | hsa-miR-181b-5p | CHD9 | 0.49 | 0.00105 | -0.51 | 3.0E-5 | miRNATAP | -0.15 | 0.00011 | NA | |
| 82 | hsa-miR-181b-5p | CLCN5 | 0.49 | 0.00105 | -0.78 | 0 | mirMAP | -0.15 | 0.00129 | NA | |
| 83 | hsa-miR-181b-5p | CLIP1 | 0.49 | 0.00105 | -0.28 | 0.00918 | MirTarget; miRNATAP | -0.15 | 2.0E-5 | NA | |
| 84 | hsa-miR-181b-5p | CLOCK | 0.49 | 0.00105 | -0.53 | 0.00745 | mirMAP | -0.13 | 0.0421 | NA | |
| 85 | hsa-miR-181b-5p | CNKSR2 | 0.49 | 0.00105 | -0.55 | 0.15771 | MirTarget; miRNATAP | -0.69 | 0 | NA | |
| 86 | hsa-miR-181b-5p | CNNM2 | 0.49 | 0.00105 | 0.05 | 0.64877 | miRNATAP | -0.18 | 0 | NA | |
| 87 | hsa-miR-181b-5p | CNST | 0.49 | 0.00105 | -0.2 | 0.03829 | mirMAP | -0.18 | 0 | NA | |
| 88 | hsa-miR-181b-5p | CNTNAP3 | 0.49 | 0.00105 | 0.11 | 0.74285 | MirTarget | -0.34 | 0.0013 | NA | |
| 89 | hsa-miR-181b-5p | COBL | 0.49 | 0.00105 | -0.63 | 0.00023 | mirMAP | -0.17 | 0.00249 | NA | |
| 90 | hsa-miR-181b-5p | CPEB4 | 0.49 | 0.00105 | -1.06 | 0 | miRNATAP | -0.29 | 0 | NA | |
| 91 | hsa-miR-181b-5p | CPOX | 0.49 | 0.00105 | -0.65 | 0 | MirTarget | -0.17 | 0 | NA | |
| 92 | hsa-miR-181b-5p | CREBL2 | 0.49 | 0.00105 | -0.86 | 0 | MirTarget | -0.28 | 0 | NA | |
| 93 | hsa-miR-181b-5p | CSNK1G3 | 0.49 | 0.00105 | -0.37 | 0 | mirMAP; miRNATAP | -0.12 | 0 | NA | |
| 94 | hsa-miR-181b-5p | CXADR | 0.49 | 0.00105 | -0.23 | 0.19668 | miRNATAP | -0.17 | 0.00344 | NA | |
| 95 | hsa-miR-181b-5p | CYP4F3 | 0.49 | 0.00105 | -1.55 | 0 | mirMAP | -0.81 | 0 | NA | |
| 96 | hsa-miR-181b-5p | DAAM1 | 0.49 | 0.00105 | -0.82 | 3.0E-5 | mirMAP | -0.24 | 0.00015 | NA | |
| 97 | hsa-miR-30d-5p | DDIT4 | 0.72 | 0 | 0.23 | 0.23684 | MirTarget; miRNATAP | -0.21 | 0.00439 | NA | |
| 98 | hsa-miR-32-5p | DDIT4 | 0.08 | 0.54898 | 0.23 | 0.23684 | miRNATAP | -0.2 | 0.00446 | NA | |
| 99 | hsa-miR-618 | DDIT4 | 0.14 | 0.51715 | 0.23 | 0.23684 | mirMAP | -0.1 | 0.02699 | NA | |
| 100 | hsa-miR-92a-3p | DDIT4 | 0.21 | 0.13429 | 0.23 | 0.23684 | miRNATAP | -0.18 | 0.00604 | NA | |
| 101 | hsa-miR-181b-5p | DDX60L | 0.49 | 0.00105 | -0.81 | 0 | MirTarget | -0.17 | 0.00123 | NA | |
| 102 | hsa-miR-181b-5p | DENND4C | 0.49 | 0.00105 | -0.45 | 0 | MirTarget | -0.12 | 0.00024 | NA | |
| 103 | hsa-miR-181b-5p | DENND5B | 0.49 | 0.00105 | -0.62 | 0 | MirTarget; mirMAP | -0.25 | 0 | NA | |
| 104 | hsa-miR-181b-5p | DERL1 | 0.49 | 0.00105 | 0.12 | 0.20253 | MirTarget; miRNATAP | -0.14 | 0 | NA | |
| 105 | hsa-miR-181b-5p | DIAPH1 | 0.49 | 0.00105 | -0.4 | 2.0E-5 | mirMAP | -0.28 | 0 | NA | |
| 106 | hsa-miR-181b-5p | DIP2C | 0.49 | 0.00105 | -0.78 | 5.0E-5 | MirTarget; miRNATAP | -0.28 | 1.0E-5 | NA | |
| 107 | hsa-miR-181b-5p | DIXDC1 | 0.49 | 0.00105 | -1.03 | 0 | mirMAP | -0.18 | 0.00085 | NA | |
| 108 | hsa-miR-181b-5p | DLC1 | 0.49 | 0.00105 | -1.42 | 0 | mirMAP | -0.13 | 0.00829 | NA | |
| 109 | hsa-miR-181b-5p | EAF1 | 0.49 | 0.00105 | -0.37 | 2.0E-5 | mirMAP | -0.11 | 0.00016 | NA | |
| 110 | hsa-miR-330-5p | EIF4B | 0.44 | 0.00533 | -0.21 | 0.00928 | miRNATAP | -0.12 | 0 | NA | |
| 111 | hsa-miR-339-5p | EIF4B | 0.28 | 0.03557 | -0.21 | 0.00928 | miRanda | -0.13 | 1.0E-5 | NA | |
| 112 | hsa-miR-342-3p | EIF4B | -0.32 | 0.04498 | -0.21 | 0.00928 | miRanda | -0.13 | 0 | NA | |
| 113 | hsa-miR-362-3p | EIF4B | 0.81 | 0 | -0.21 | 0.00928 | miRanda | -0.12 | 3.0E-5 | NA | |
| 114 | hsa-miR-484 | EIF4B | 0.09 | 0.45398 | -0.21 | 0.00928 | miRNAWalker2 validate | -0.16 | 1.0E-5 | NA | |
| 115 | hsa-miR-497-5p | EIF4B | -1.41 | 0 | -0.21 | 0.00928 | miRNATAP | -0.1 | 0 | NA | |
| 116 | hsa-miR-107 | EIF4E | 0.24 | 0.01708 | -0.42 | 0 | miRanda | -0.15 | 5.0E-5 | NA | |
| 117 | hsa-miR-25-3p | EIF4E | 0.63 | 0 | -0.42 | 0 | mirMAP | -0.12 | 0.00065 | NA | |
| 118 | hsa-miR-500a-3p | EIF4E | 1.19 | 0 | -0.42 | 0 | MirTarget | -0.1 | 3.0E-5 | NA | |
| 119 | hsa-miR-660-5p | EIF4E | 0.99 | 0 | -0.42 | 0 | mirMAP | -0.16 | 0 | NA | |
| 120 | hsa-miR-140-5p | EIF4E2 | -0.22 | 0.01407 | -0.03 | 0.66056 | miRanda | -0.14 | 0.00026 | NA | |
| 121 | hsa-miR-29a-3p | EIF4E2 | -0.86 | 0 | -0.03 | 0.66056 | MirTarget | -0.11 | 2.0E-5 | NA | |
| 122 | hsa-miR-125b-5p | EIF4EBP1 | -1.36 | 0 | -0.03 | 0.83808 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.12 | 0.00778 | 17891175; 26646586 | Using a bioinformatics approach we have identified additional potential mRNA targets of one of the miRNAs miR-125b that are upregulated in prostate cancer and confirmed increased expression of one of these targets EIF4EBP1 in prostate cancer tissues;Increased expression of miR-125a and miR-125b inhibited invasion and migration of SKOV3 and OVCAR-429 ovarian cancer cells and was associated with a decrease in EIF4EBP1 expression |
| 123 | hsa-miR-181b-5p | ELOVL6 | 0.49 | 0.00105 | -1.01 | 0 | mirMAP | -0.41 | 0 | NA | |
| 124 | hsa-miR-181b-5p | EMP2 | 0.49 | 0.00105 | -0.3 | 0.02441 | mirMAP | -0.3 | 0 | NA | |
| 125 | hsa-miR-181b-5p | EPB41L4B | 0.49 | 0.00105 | -1.97 | 0 | miRNATAP | -0.36 | 0 | NA | |
| 126 | hsa-miR-181b-5p | EPS15 | 0.49 | 0.00105 | -0.3 | 1.0E-5 | miRNATAP | -0.13 | 0 | NA | |
| 127 | hsa-miR-181b-5p | ERLIN2 | 0.49 | 0.00105 | -0.77 | 0 | MirTarget | -0.18 | 1.0E-5 | NA | |
| 128 | hsa-miR-181b-5p | ESR1 | 0.49 | 0.00105 | -4.12 | 0 | miRNATAP | -0.93 | 0 | NA | |
| 129 | hsa-miR-181b-5p | EVI5 | 0.49 | 0.00105 | -0.66 | 0 | mirMAP | -0.11 | 0.00086 | NA | |
| 130 | hsa-miR-181b-5p | FAM169A | 0.49 | 0.00105 | 0.37 | 0.13291 | mirMAP | -0.51 | 0 | NA | |
| 131 | hsa-miR-181b-5p | FAM46A | 0.49 | 0.00105 | -1.45 | 0 | mirMAP; miRNATAP | -0.24 | 1.0E-5 | NA | |
| 132 | hsa-miR-181b-5p | FAM47E | 0.49 | 0.00105 | 0.2 | 0.23251 | mirMAP | -0.22 | 3.0E-5 | NA | |
| 133 | hsa-miR-181b-5p | FGD4 | 0.49 | 0.00105 | -1.16 | 0 | mirMAP | -0.21 | 0.00013 | NA | |
| 134 | hsa-miR-181b-5p | FGF2 | 0.49 | 0.00105 | -1.09 | 0.00032 | mirMAP | -0.52 | 0 | NA | |
| 135 | hsa-miR-181b-5p | FNDC3A | 0.49 | 0.00105 | -1 | 0 | MirTarget; mirMAP; miRNATAP | -0.17 | 0.00013 | NA | |
| 136 | hsa-miR-181b-5p | GAN | 0.49 | 0.00105 | -0.68 | 0 | mirMAP | -0.16 | 5.0E-5 | NA | |
| 137 | hsa-miR-181b-5p | GANC | 0.49 | 0.00105 | -0.01 | 0.89064 | MirTarget | -0.15 | 0 | NA | |
| 138 | hsa-miR-181b-5p | GATM | 0.49 | 0.00105 | -1.68 | 0 | MirTarget | -0.45 | 0 | NA | |
| 139 | hsa-miR-181b-5p | GDA | 0.49 | 0.00105 | -1.85 | 0 | miRNATAP | -0.39 | 0.0029 | NA | |
| 140 | hsa-miR-181b-5p | GFOD1 | 0.49 | 0.00105 | -0.74 | 0 | mirMAP | -0.39 | 0 | NA | |
| 141 | hsa-miR-181b-5p | GFRA1 | 0.49 | 0.00105 | -2.55 | 0 | mirMAP | -0.89 | 0 | NA | |
| 142 | hsa-miR-181b-5p | GHITM | 0.49 | 0.00105 | -0.68 | 0 | MirTarget; miRNATAP | -0.24 | 0 | NA | |
| 143 | hsa-miR-181b-5p | GOLGA1 | 0.49 | 0.00105 | -0.06 | 0.44214 | MirTarget; miRNATAP | -0.14 | 0 | NA | |
| 144 | hsa-miR-181b-5p | GPAM | 0.49 | 0.00105 | -0.47 | 0.0638 | MirTarget; mirMAP | -0.46 | 0 | NA | |
| 145 | hsa-miR-181b-5p | GTPBP10 | 0.49 | 0.00105 | -0.21 | 0.01214 | mirMAP | -0.18 | 0 | NA | |
| 146 | hsa-miR-181b-5p | HEMK1 | 0.49 | 0.00105 | 0.06 | 0.44892 | mirMAP | -0.16 | 0 | NA | |
| 147 | hsa-miR-181b-5p | HEY2 | 0.49 | 0.00105 | -1.35 | 0 | miRNATAP | -0.33 | 0 | NA | |
| 148 | hsa-miR-107 | HIF1A | 0.24 | 0.01708 | -0.37 | 0.0111 | miRNAWalker2 validate; miRTarBase; miRanda | -0.44 | 0 | NA | |
| 149 | hsa-miR-17-5p | HIF1A | 0.7 | 2.0E-5 | -0.37 | 0.0111 | miRTarBase; MirTarget; TargetScan | -0.21 | 0 | NA | |
| 150 | hsa-miR-20a-5p | HIF1A | 0.85 | 0 | -0.37 | 0.0111 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.25 | 0 | 22901144 | Correlation analysis showed that the key miRNAs miR-20a and miR-20b negatively correlated with the target proteins VEGF-A and HIF-1alpha |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | RESPONSE TO INSULIN | 23 | 205 | 6.183e-12 | 2.877e-08 |
| 2 | PROTEIN PHOSPHORYLATION | 50 | 944 | 2.679e-11 | 6.232e-08 |
| 3 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 72 | 1791 | 3.124e-10 | 2.423e-07 |
| 4 | CELLULAR RESPONSE TO HORMONE STIMULUS | 35 | 552 | 3.077e-10 | 2.423e-07 |
| 5 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 77 | 1977 | 2.859e-10 | 2.423e-07 |
| 6 | PHOSPHORYLATION | 57 | 1228 | 1.643e-10 | 2.423e-07 |
| 7 | RESPONSE TO PEPTIDE | 29 | 404 | 6.552e-10 | 4.121e-07 |
| 8 | REGULATION OF LIPID METABOLIC PROCESS | 24 | 282 | 7.086e-10 | 4.121e-07 |
| 9 | REGULATION OF GLUCOSE IMPORT | 12 | 60 | 9.287e-10 | 4.321e-07 |
| 10 | INTRACELLULAR SIGNAL TRANSDUCTION | 65 | 1572 | 8.819e-10 | 4.321e-07 |
| 11 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 66 | 1618 | 1.112e-09 | 4.703e-07 |
| 12 | RESPONSE TO HORMONE | 45 | 893 | 1.411e-09 | 5.472e-07 |
| 13 | NEGATIVE REGULATION OF CELL DEATH | 43 | 872 | 6.433e-09 | 2.302e-06 |
| 14 | REGULATION OF GENERATION OF PRECURSOR METABOLITES AND ENERGY | 13 | 89 | 1.049e-08 | 3.488e-06 |
| 15 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 64 | 1656 | 1.69e-08 | 5.241e-06 |
| 16 | REGULATION OF CELL DEATH | 59 | 1472 | 1.872e-08 | 5.443e-06 |
| 17 | REGULATION OF COENZYME METABOLIC PROCESS | 10 | 50 | 2.388e-08 | 6.158e-06 |
| 18 | REGULATION OF CARBOHYDRATE METABOLIC PROCESS | 17 | 172 | 2.515e-08 | 6.158e-06 |
| 19 | REGULATION OF COFACTOR METABOLIC PROCESS | 10 | 50 | 2.388e-08 | 6.158e-06 |
| 20 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 59 | 1518 | 5.555e-08 | 1.175e-05 |
| 21 | NEGATIVE REGULATION OF TOR SIGNALING | 8 | 30 | 5.553e-08 | 1.175e-05 |
| 22 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 45 | 1008 | 5.44e-08 | 1.175e-05 |
| 23 | REGULATION OF CELLULAR CARBOHYDRATE CATABOLIC PROCESS | 9 | 42 | 6.42e-08 | 1.245e-05 |
| 24 | REGULATION OF CARBOHYDRATE CATABOLIC PROCESS | 9 | 42 | 6.42e-08 | 1.245e-05 |
| 25 | INOSITOL LIPID MEDIATED SIGNALING | 14 | 124 | 8.216e-08 | 1.529e-05 |
| 26 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 35 | 689 | 8.89e-08 | 1.591e-05 |
| 27 | CELLULAR RESPONSE TO INSULIN STIMULUS | 15 | 146 | 1.005e-07 | 1.72e-05 |
| 28 | RESPONSE TO OXYGEN LEVELS | 22 | 311 | 1.035e-07 | 1.72e-05 |
| 29 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 45 | 1036 | 1.198e-07 | 1.808e-05 |
| 30 | PEPTIDYL SERINE MODIFICATION | 15 | 148 | 1.204e-07 | 1.808e-05 |
| 31 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 45 | 1036 | 1.198e-07 | 1.808e-05 |
| 32 | REGULATION OF CELLULAR RESPONSE TO INSULIN STIMULUS | 10 | 59 | 1.258e-07 | 1.829e-05 |
| 33 | RESPONSE TO ENDOGENOUS STIMULUS | 56 | 1450 | 1.575e-07 | 2.22e-05 |
| 34 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 47 | 1135 | 2.48e-07 | 3.205e-05 |
| 35 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 28 | 498 | 2.432e-07 | 3.205e-05 |
| 36 | CELLULAR RESPONSE TO PEPTIDE | 20 | 274 | 2.4e-07 | 3.205e-05 |
| 37 | REGULATION OF NUCLEOTIDE CATABOLIC PROCESS | 8 | 36 | 2.604e-07 | 3.274e-05 |
| 38 | RESPONSE TO EXTRACELLULAR STIMULUS | 26 | 441 | 2.676e-07 | 3.276e-05 |
| 39 | ACTIVATION OF PROTEIN KINASE ACTIVITY | 20 | 279 | 3.207e-07 | 3.826e-05 |
| 40 | REGULATION OF GLUCOSE TRANSPORT | 12 | 100 | 3.569e-07 | 4.151e-05 |
| 41 | NEGATIVE REGULATION OF CELL COMMUNICATION | 48 | 1192 | 4.122e-07 | 4.678e-05 |
| 42 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 20 | 285 | 4.501e-07 | 4.986e-05 |
| 43 | REGULATION OF TOR SIGNALING | 10 | 68 | 5.016e-07 | 5.304e-05 |
| 44 | POSITIVE REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 10 | 68 | 5.016e-07 | 5.304e-05 |
| 45 | REGULATION OF FATTY ACID METABOLIC PROCESS | 11 | 87 | 6.462e-07 | 6.398e-05 |
| 46 | REGULATION OF FATTY ACID OXIDATION | 7 | 28 | 6.267e-07 | 6.398e-05 |
| 47 | REGULATION OF PROTEIN MODIFICATION PROCESS | 61 | 1710 | 6.336e-07 | 6.398e-05 |
| 48 | REGULATION OF GLUCOSE METABOLIC PROCESS | 12 | 106 | 6.764e-07 | 6.556e-05 |
| 49 | RESPONSE TO NITROGEN COMPOUND | 38 | 859 | 7.999e-07 | 7.596e-05 |
| 50 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 63 | 1805 | 8.483e-07 | 7.894e-05 |
| 51 | POSITIVE REGULATION OF GLUCOSE TRANSPORT | 8 | 42 | 9.211e-07 | 8.242e-05 |
| 52 | REGULATION OF CELL MATRIX ADHESION | 11 | 90 | 9.133e-07 | 8.242e-05 |
| 53 | POSITIVE REGULATION OF CARBOHYDRATE METABOLIC PROCESS | 10 | 75 | 1.274e-06 | 0.0001118 |
| 54 | REGULATION OF CELL CYCLE | 40 | 949 | 1.359e-06 | 0.0001171 |
| 55 | REGULATION OF KINASE ACTIVITY | 35 | 776 | 1.444e-06 | 0.0001222 |
| 56 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 54 | 1492 | 2.002e-06 | 0.0001663 |
| 57 | POSITIVE REGULATION OF GENE EXPRESSION | 60 | 1733 | 2.105e-06 | 0.0001719 |
| 58 | PROTEIN KINASE B SIGNALING | 7 | 34 | 2.586e-06 | 0.0002074 |
| 59 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 35 | 799 | 2.779e-06 | 0.0002192 |
| 60 | CELLULAR RESPONSE TO OXYGEN LEVELS | 13 | 143 | 2.887e-06 | 0.0002239 |
| 61 | CHEMICAL HOMEOSTASIS | 37 | 874 | 3.142e-06 | 0.0002284 |
| 62 | PHOSPHATIDYLINOSITOL 3 PHOSPHATE BIOSYNTHETIC PROCESS | 8 | 49 | 3.141e-06 | 0.0002284 |
| 63 | REGULATION OF NUCLEOSIDE METABOLIC PROCESS | 8 | 49 | 3.141e-06 | 0.0002284 |
| 64 | REGULATION OF ATP METABOLIC PROCESS | 8 | 49 | 3.141e-06 | 0.0002284 |
| 65 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 50 | 1360 | 3.252e-06 | 0.0002328 |
| 66 | REGULATION OF CIRCADIAN RHYTHM | 11 | 103 | 3.528e-06 | 0.0002487 |
| 67 | POSITIVE REGULATION OF GLUCOSE METABOLIC PROCESS | 7 | 36 | 3.885e-06 | 0.0002698 |
| 68 | REGULATION OF CELLULAR KETONE METABOLIC PROCESS | 14 | 173 | 4.763e-06 | 0.0003259 |
| 69 | PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 6 | 25 | 5.303e-06 | 0.0003525 |
| 70 | RESPONSE TO ACTIVITY | 9 | 69 | 5.238e-06 | 0.0003525 |
| 71 | REGULATION OF NEURON DEATH | 17 | 252 | 5.597e-06 | 0.0003668 |
| 72 | CELL CYCLE ARREST | 13 | 154 | 6.559e-06 | 0.000418 |
| 73 | POSITIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 13 | 154 | 6.559e-06 | 0.000418 |
| 74 | TOR SIGNALING | 5 | 16 | 8.101e-06 | 0.0005026 |
| 75 | REGULATION OF FATTY ACID BETA OXIDATION | 5 | 16 | 8.101e-06 | 0.0005026 |
| 76 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 12 | 138 | 1.097e-05 | 0.0006573 |
| 77 | POSITIVE REGULATION OF GLYCOGEN METABOLIC PROCESS | 5 | 17 | 1.13e-05 | 0.0006573 |
| 78 | REGULATION OF GLUCOSE IMPORT IN RESPONSE TO INSULIN STIMULUS | 5 | 17 | 1.13e-05 | 0.0006573 |
| 79 | POSITIVE REGULATION OF NUCLEOTIDE CATABOLIC PROCESS | 5 | 17 | 1.13e-05 | 0.0006573 |
| 80 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 49 | 1381 | 1.076e-05 | 0.0006573 |
| 81 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 39 | 1004 | 1.262e-05 | 0.0007248 |
| 82 | PROTEIN CATABOLIC PROCESS | 27 | 579 | 1.33e-05 | 0.0007547 |
| 83 | RHYTHMIC PROCESS | 18 | 298 | 1.365e-05 | 0.0007653 |
| 84 | POSITIVE REGULATION OF KINASE ACTIVITY | 24 | 482 | 1.401e-05 | 0.0007763 |
| 85 | REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 10 | 98 | 1.472e-05 | 0.0008059 |
| 86 | PHOSPHATIDYLINOSITOL BIOSYNTHETIC PROCESS | 11 | 120 | 1.548e-05 | 0.0008281 |
| 87 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 60 | 1848 | 1.545e-05 | 0.0008281 |
| 88 | INSULIN RECEPTOR SIGNALING PATHWAY | 9 | 80 | 1.79e-05 | 0.0009465 |
| 89 | REGULATION OF TRANSFERASE ACTIVITY | 37 | 946 | 1.839e-05 | 0.0009613 |
| 90 | GLUCOSE HOMEOSTASIS | 13 | 170 | 1.908e-05 | 0.0009754 |
| 91 | CARBOHYDRATE HOMEOSTASIS | 13 | 170 | 1.908e-05 | 0.0009754 |
| 92 | RESPONSE TO ABIOTIC STIMULUS | 39 | 1024 | 1.968e-05 | 0.0009955 |
| 93 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 58 | 1784 | 2.108e-05 | 0.001055 |
| 94 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 32 | 771 | 2.215e-05 | 0.001096 |
| 95 | NEGATIVE REGULATION OF CELL CYCLE | 22 | 433 | 2.356e-05 | 0.001154 |
| 96 | ESTABLISHMENT OF LOCALIZATION IN CELL | 55 | 1676 | 2.747e-05 | 0.001304 |
| 97 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 22 | 437 | 2.709e-05 | 0.001304 |
| 98 | MACROMOLECULE CATABOLIC PROCESS | 36 | 926 | 2.722e-05 | 0.001304 |
| 99 | REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 15 | 229 | 2.801e-05 | 0.001316 |
| 100 | REGULATION OF LIPID BIOSYNTHETIC PROCESS | 11 | 128 | 2.843e-05 | 0.001323 |
| 101 | CELLULAR RESPONSE TO NITROGEN COMPOUND | 24 | 505 | 2.982e-05 | 0.001374 |
| 102 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 26 | 573 | 3.063e-05 | 0.001397 |
| 103 | REGULATION OF DEVELOPMENTAL GROWTH | 17 | 289 | 3.307e-05 | 0.001494 |
| 104 | PROTEIN STABILIZATION | 11 | 131 | 3.527e-05 | 0.001578 |
| 105 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 27 | 616 | 3.921e-05 | 0.001737 |
| 106 | REGULATION OF CELLULAR AMIDE METABOLIC PROCESS | 19 | 354 | 4.065e-05 | 0.001784 |
| 107 | REGULATION OF GLYCOGEN METABOLIC PROCESS | 6 | 35 | 4.153e-05 | 0.001792 |
| 108 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 38 | 1021 | 4.159e-05 | 0.001792 |
| 109 | ACTIN FILAMENT BASED PROCESS | 22 | 450 | 4.206e-05 | 0.001796 |
| 110 | GLYCEROLIPID BIOSYNTHETIC PROCESS | 14 | 211 | 4.484e-05 | 0.001897 |
| 111 | POSITIVE REGULATION OF LOCOMOTION | 21 | 420 | 4.547e-05 | 0.001906 |
| 112 | REGULATION OF PROTEIN LOCALIZATION | 36 | 950 | 4.629e-05 | 0.001923 |
| 113 | REGULATION OF LIPID CATABOLIC PROCESS | 7 | 52 | 4.82e-05 | 0.001985 |
| 114 | FATTY ACID HOMEOSTASIS | 4 | 12 | 5.235e-05 | 0.002118 |
| 115 | POSITIVE REGULATION OF GLUCOSE IMPORT IN RESPONSE TO INSULIN STIMULUS | 4 | 12 | 5.235e-05 | 0.002118 |
| 116 | CIRCADIAN RHYTHM | 11 | 137 | 5.329e-05 | 0.002138 |
| 117 | POSITIVE REGULATION OF CELLULAR RESPONSE TO INSULIN STIMULUS | 5 | 23 | 5.604e-05 | 0.00221 |
| 118 | REGULATION OF TRANSPORT | 57 | 1804 | 5.573e-05 | 0.00221 |
| 119 | NEUROGENESIS | 47 | 1402 | 6.758e-05 | 0.002642 |
| 120 | REGULATION OF POSITIVE CHEMOTAXIS | 5 | 24 | 6.971e-05 | 0.002659 |
| 121 | POSITIVE REGULATION OF NUCLEOSIDE METABOLIC PROCESS | 5 | 24 | 6.971e-05 | 0.002659 |
| 122 | POSITIVE REGULATION OF ATP METABOLIC PROCESS | 5 | 24 | 6.971e-05 | 0.002659 |
| 123 | REGULATION OF SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 16 | 278 | 7.268e-05 | 0.00275 |
| 124 | RESPONSE TO CARBOHYDRATE | 12 | 168 | 7.721e-05 | 0.002874 |
| 125 | CELLULAR GLUCOSE HOMEOSTASIS | 8 | 75 | 7.71e-05 | 0.002874 |
| 126 | REGULATION OF CELL PROLIFERATION | 49 | 1496 | 8.293e-05 | 0.003062 |
| 127 | CELLULAR CARBOHYDRATE METABOLIC PROCESS | 11 | 144 | 8.382e-05 | 0.003071 |
| 128 | CIRCADIAN REGULATION OF GENE EXPRESSION | 7 | 57 | 8.794e-05 | 0.003197 |
| 129 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 53 | 1672 | 9.675e-05 | 0.00349 |
| 130 | LIPID PHOSPHORYLATION | 9 | 99 | 9.797e-05 | 0.003507 |
| 131 | NEGATIVE REGULATION OF RHO PROTEIN SIGNAL TRANSDUCTION | 4 | 14 | 0.0001028 | 0.003623 |
| 132 | REGULATION OF CELL SUBSTRATE ADHESION | 12 | 173 | 0.0001022 | 0.003623 |
| 133 | CYTOSKELETON ORGANIZATION | 32 | 838 | 0.0001075 | 0.003762 |
| 134 | POSTTRANSCRIPTIONAL REGULATION OF GENE EXPRESSION | 21 | 448 | 0.0001133 | 0.003936 |
| 135 | WNT SIGNALING PATHWAY | 18 | 351 | 0.0001151 | 0.003937 |
| 136 | POSITIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 16 | 289 | 0.0001143 | 0.003937 |
| 137 | REGULATION OF POLYSACCHARIDE METABOLIC PROCESS | 6 | 43 | 0.0001376 | 0.004672 |
| 138 | PHOSPHOLIPID BIOSYNTHETIC PROCESS | 14 | 235 | 0.000142 | 0.004787 |
| 139 | POSITIVE REGULATION OF CELL PROLIFERATION | 31 | 814 | 0.0001447 | 0.004844 |
| 140 | NEGATIVE REGULATION OF GENE EXPRESSION | 48 | 1493 | 0.0001533 | 0.005096 |
| 141 | TUBE FORMATION | 10 | 129 | 0.0001552 | 0.005123 |
| 142 | POSITIVE REGULATION OF GROWTH | 14 | 238 | 0.0001621 | 0.005311 |
| 143 | RESPONSE TO OSMOTIC STRESS | 7 | 63 | 0.0001672 | 0.005441 |
| 144 | REGULATION OF NUCLEOTIDE METABOLIC PROCESS | 13 | 211 | 0.0001753 | 0.005664 |
| 145 | HOMEOSTATIC PROCESS | 44 | 1337 | 0.0001771 | 0.005684 |
| 146 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 58 | 1929 | 0.0001826 | 0.005819 |
| 147 | POSITIVE REGULATION OF VASCULATURE DEVELOPMENT | 10 | 133 | 0.0001995 | 0.006314 |
| 148 | SMALL MOLECULE METABOLIC PROCESS | 54 | 1767 | 0.0002096 | 0.006589 |
| 149 | NEGATIVE REGULATION OF CELL MATRIX ADHESION | 5 | 30 | 0.0002132 | 0.006658 |
| 150 | LIPID HOMEOSTASIS | 9 | 110 | 0.0002197 | 0.006735 |
| 151 | REGULATION OF CARBOHYDRATE BIOSYNTHETIC PROCESS | 8 | 87 | 0.00022 | 0.006735 |
| 152 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 21 | 470 | 0.0002186 | 0.006735 |
| 153 | CELLULAR LIPID METABOLIC PROCESS | 33 | 913 | 0.0002316 | 0.007042 |
| 154 | REGULATION OF PRI MIRNA TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 4 | 17 | 0.0002337 | 0.007062 |
| 155 | NEGATIVE REGULATION OF CELLULAR RESPONSE TO INSULIN STIMULUS | 5 | 31 | 0.0002503 | 0.007515 |
| 156 | REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION TO PLASMA MEMBRANE | 6 | 48 | 0.000256 | 0.007637 |
| 157 | CELLULAR CATABOLIC PROCESS | 43 | 1322 | 0.0002706 | 0.00802 |
| 158 | REGULATION OF CATABOLIC PROCESS | 28 | 731 | 0.0002765 | 0.008092 |
| 159 | REGULATION OF PROTEIN STABILITY | 13 | 221 | 0.0002757 | 0.008092 |
| 160 | PHOSPHATIDYLINOSITOL METABOLIC PROCESS | 12 | 193 | 0.0002842 | 0.008213 |
| 161 | POSITIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 22 | 514 | 0.0002831 | 0.008213 |
| 162 | CARDIAC CELL DEVELOPMENT | 6 | 49 | 0.0002872 | 0.00825 |
| 163 | REGULATION OF HYDROLASE ACTIVITY | 43 | 1327 | 0.0002932 | 0.008368 |
| 164 | QUATERNARY AMMONIUM GROUP TRANSPORT | 4 | 18 | 0.0002961 | 0.008402 |
| 165 | CARDIAC MUSCLE TISSUE DEVELOPMENT | 10 | 140 | 0.0003024 | 0.008527 |
| 166 | ORGANOPHOSPHATE BIOSYNTHETIC PROCESS | 20 | 450 | 0.0003291 | 0.009224 |
| 167 | ESTABLISHMENT OF PROTEIN LOCALIZATION | 45 | 1423 | 0.0003583 | 0.009982 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | KINASE ACTIVITY | 50 | 842 | 4.055e-13 | 3.767e-10 |
| 2 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 33 | 445 | 1.728e-11 | 8.028e-09 |
| 3 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 50 | 992 | 1.537e-10 | 4.76e-08 |
| 4 | PROTEIN KINASE ACTIVITY | 38 | 640 | 3.357e-10 | 7.798e-08 |
| 5 | ENZYME BINDING | 70 | 1737 | 5.251e-10 | 9.757e-08 |
| 6 | ADENYL NUCLEOTIDE BINDING | 62 | 1514 | 3.369e-09 | 5.217e-07 |
| 7 | RIBONUCLEOTIDE BINDING | 66 | 1860 | 2.428e-07 | 3.223e-05 |
| 8 | PHOSPHATIDYLINOSITOL KINASE ACTIVITY | 9 | 51 | 3.778e-07 | 4.387e-05 |
| 9 | KINASE BINDING | 31 | 606 | 4.318e-07 | 4.457e-05 |
| 10 | X1 PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 8 | 43 | 1.113e-06 | 0.0001034 |
| 11 | PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 9 | 70 | 5.913e-06 | 0.0004994 |
| 12 | STEROID HORMONE RECEPTOR ACTIVITY | 8 | 59 | 1.313e-05 | 0.001016 |
| 13 | PROTEIN DOMAIN SPECIFIC BINDING | 28 | 624 | 1.852e-05 | 0.001323 |
| 14 | INSULIN RECEPTOR SUBSTRATE BINDING | 4 | 11 | 3.542e-05 | 0.00235 |
| 15 | PHOSPHATASE BINDING | 12 | 162 | 5.43e-05 | 0.003363 |
| 16 | MOLECULAR FUNCTION REGULATOR | 46 | 1353 | 5.825e-05 | 0.003382 |
| 17 | ENZYME ACTIVATOR ACTIVITY | 22 | 471 | 8.228e-05 | 0.004496 |
| 18 | ENZYME REGULATOR ACTIVITY | 35 | 959 | 0.000124 | 0.006398 |
| 19 | KINASE REGULATOR ACTIVITY | 12 | 186 | 0.0002021 | 0.009883 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | PHOSPHATIDYLINOSITOL 3 KINASE COMPLEX | 8 | 20 | 1.407e-09 | 8.217e-07 |
| 2 | TRANSFERASE COMPLEX TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 19 | 237 | 1.123e-07 | 3.28e-05 |
| 3 | ENDOSOME | 37 | 793 | 3.13e-07 | 6.093e-05 |
| 4 | TRANSFERASE COMPLEX | 34 | 703 | 4.29e-07 | 6.264e-05 |
| 5 | VACUOLE | 47 | 1180 | 7.505e-07 | 8.766e-05 |
| 6 | VACUOLAR MEMBRANE | 29 | 587 | 2.064e-06 | 0.0002009 |
| 7 | VACUOLAR PART | 30 | 694 | 1.888e-05 | 0.001575 |
| 8 | CATALYTIC COMPLEX | 39 | 1038 | 2.662e-05 | 0.001943 |
| 9 | CELL LEADING EDGE | 19 | 350 | 3.485e-05 | 0.002261 |
| 10 | PROTEIN KINASE COMPLEX | 9 | 90 | 4.63e-05 | 0.002704 |
| 11 | ENDOSOMAL PART | 21 | 430 | 6.368e-05 | 0.003381 |
| 12 | GOLGI APPARATUS | 47 | 1445 | 0.0001381 | 0.006719 |
| 13 | NEURON PART | 42 | 1265 | 0.0002079 | 0.009342 |
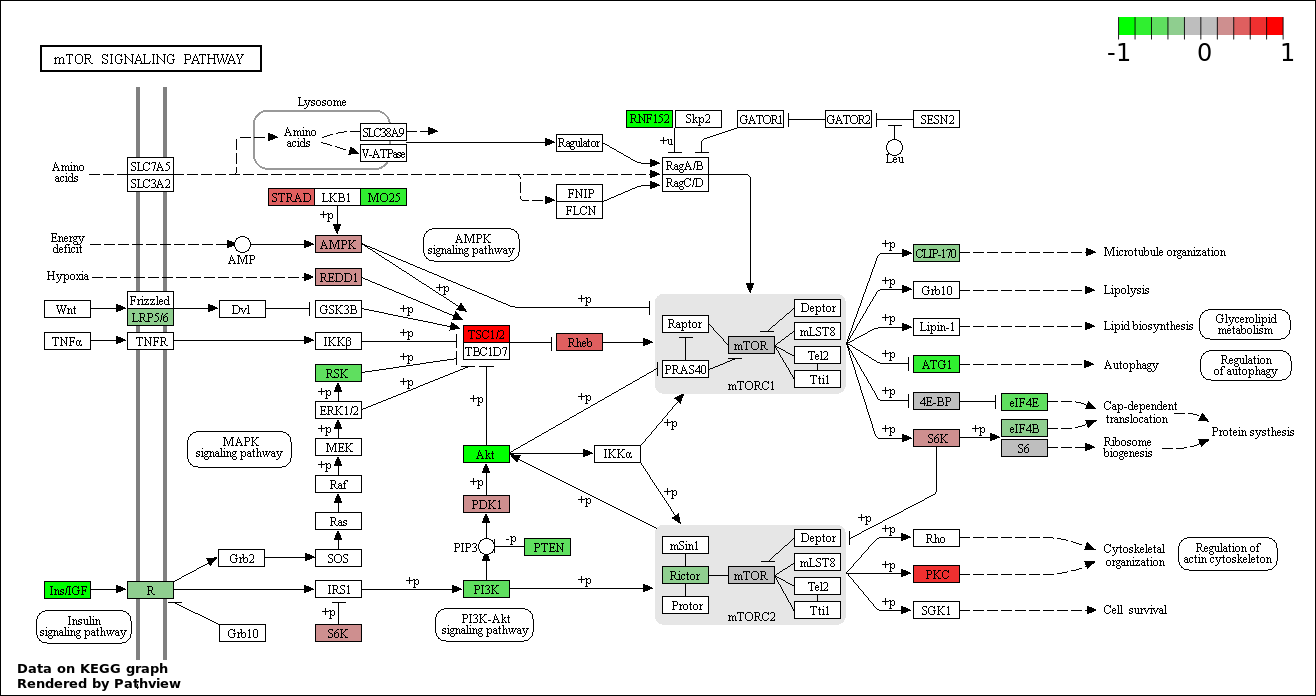
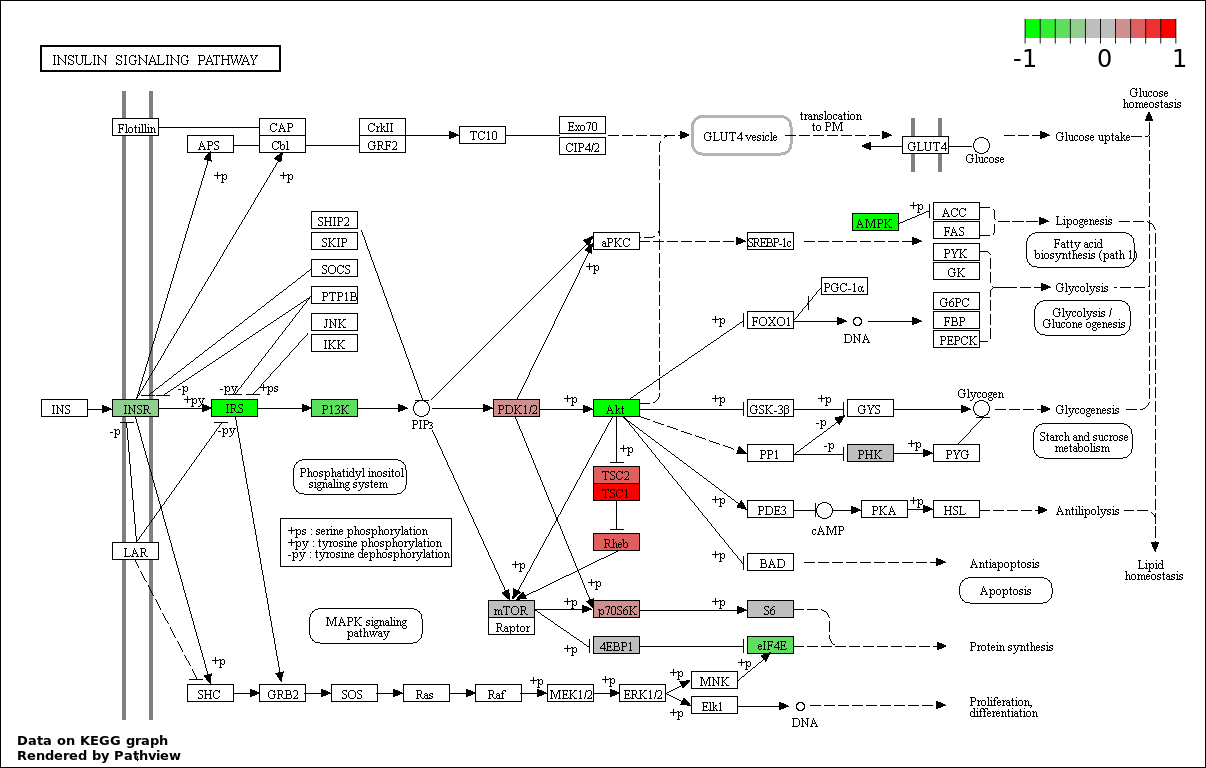
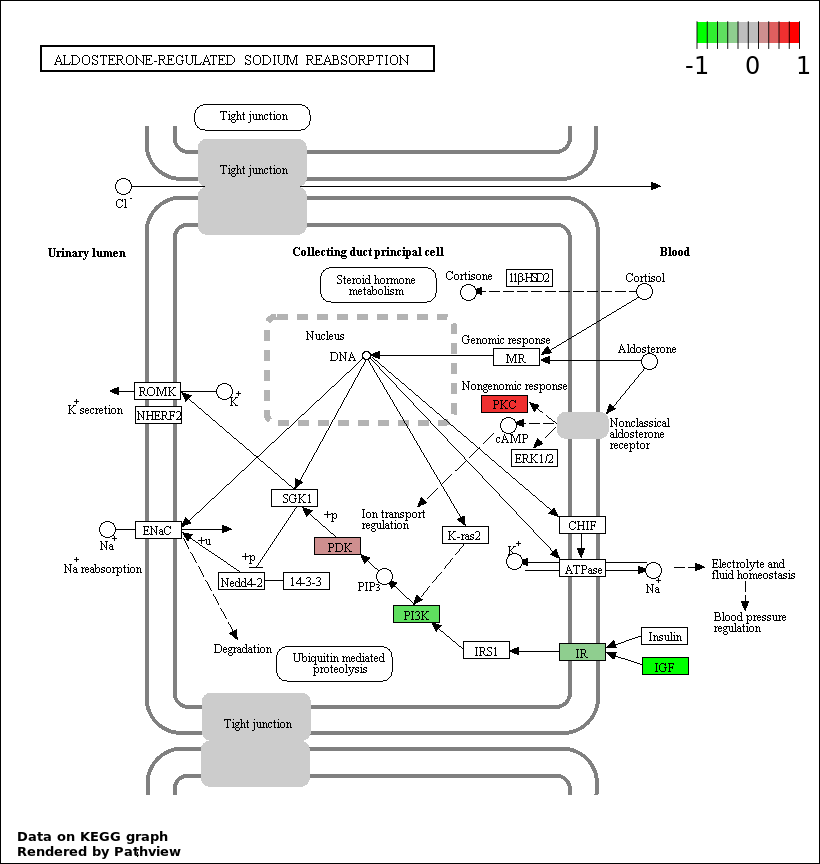
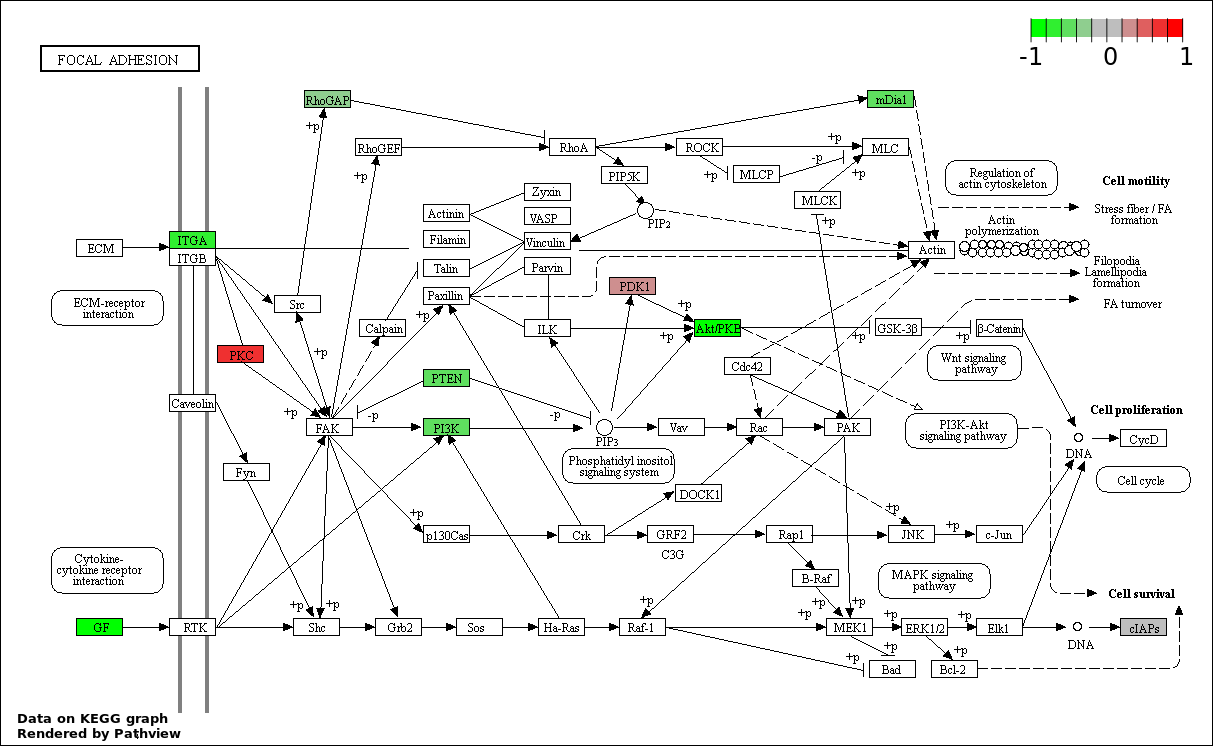
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04150_mTOR_signaling_pathway | 36 | 52 | 7.953e-51 | 1.432e-48 | |
| 2 | hsa04910_Insulin_signaling_pathway | 25 | 138 | 7.709e-18 | 6.938e-16 | |
| 3 | hsa04151_PI3K_AKT_signaling_pathway | 33 | 351 | 2.393e-14 | 1.436e-12 | |
| 4 | hsa04960_Aldosterone.regulated_sodium_reabsorption | 12 | 42 | 9.926e-12 | 4.466e-10 | |
| 5 | hsa04914_Progesterone.mediated_oocyte_maturation | 15 | 87 | 6.757e-11 | 2.433e-09 | |
| 6 | hsa04510_Focal_adhesion | 20 | 200 | 1.184e-09 | 3.551e-08 | |
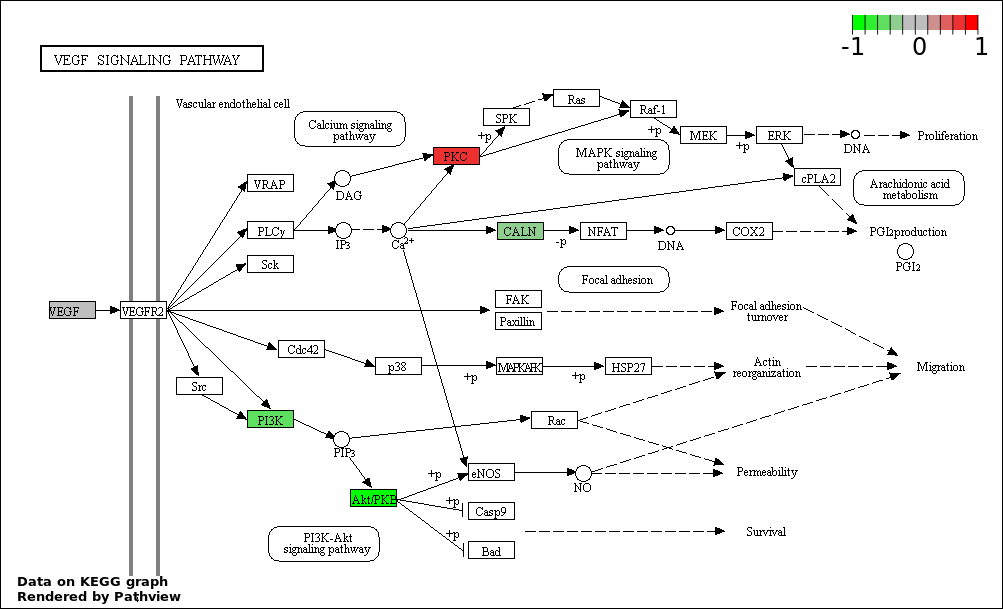
| 7 | hsa04370_VEGF_signaling_pathway | 13 | 76 | 1.418e-09 | 3.647e-08 | |
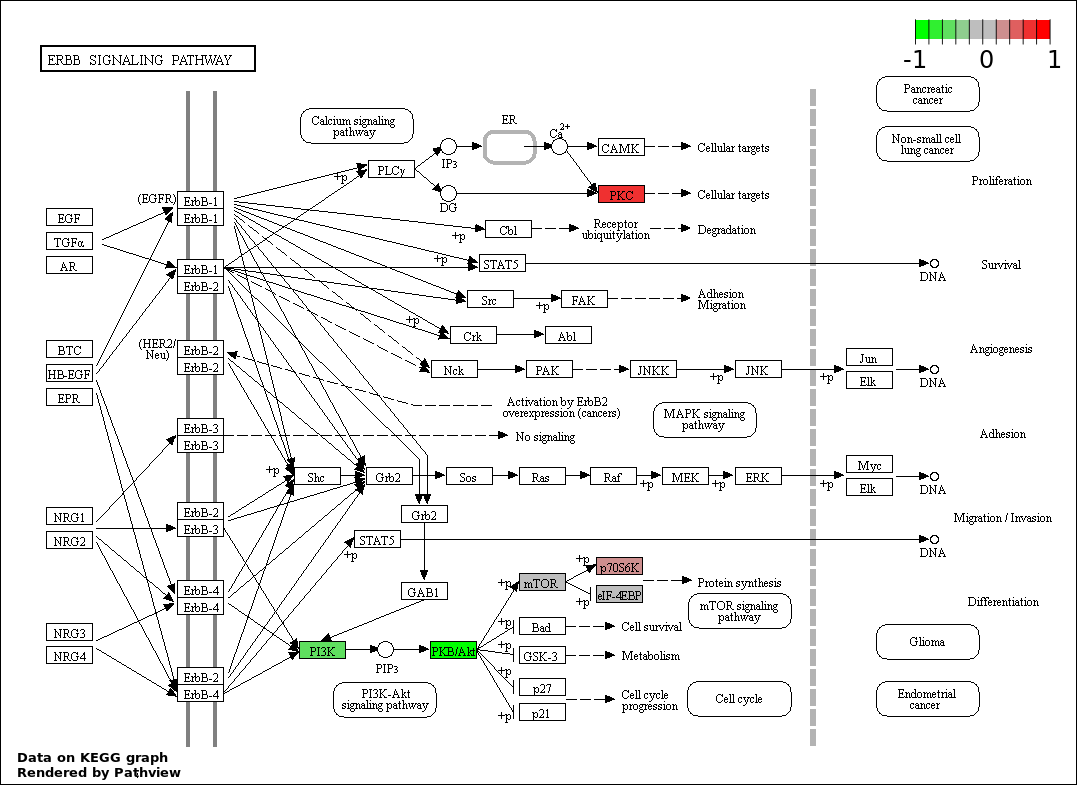
| 8 | hsa04012_ErbB_signaling_pathway | 13 | 87 | 7.898e-09 | 1.777e-07 | |
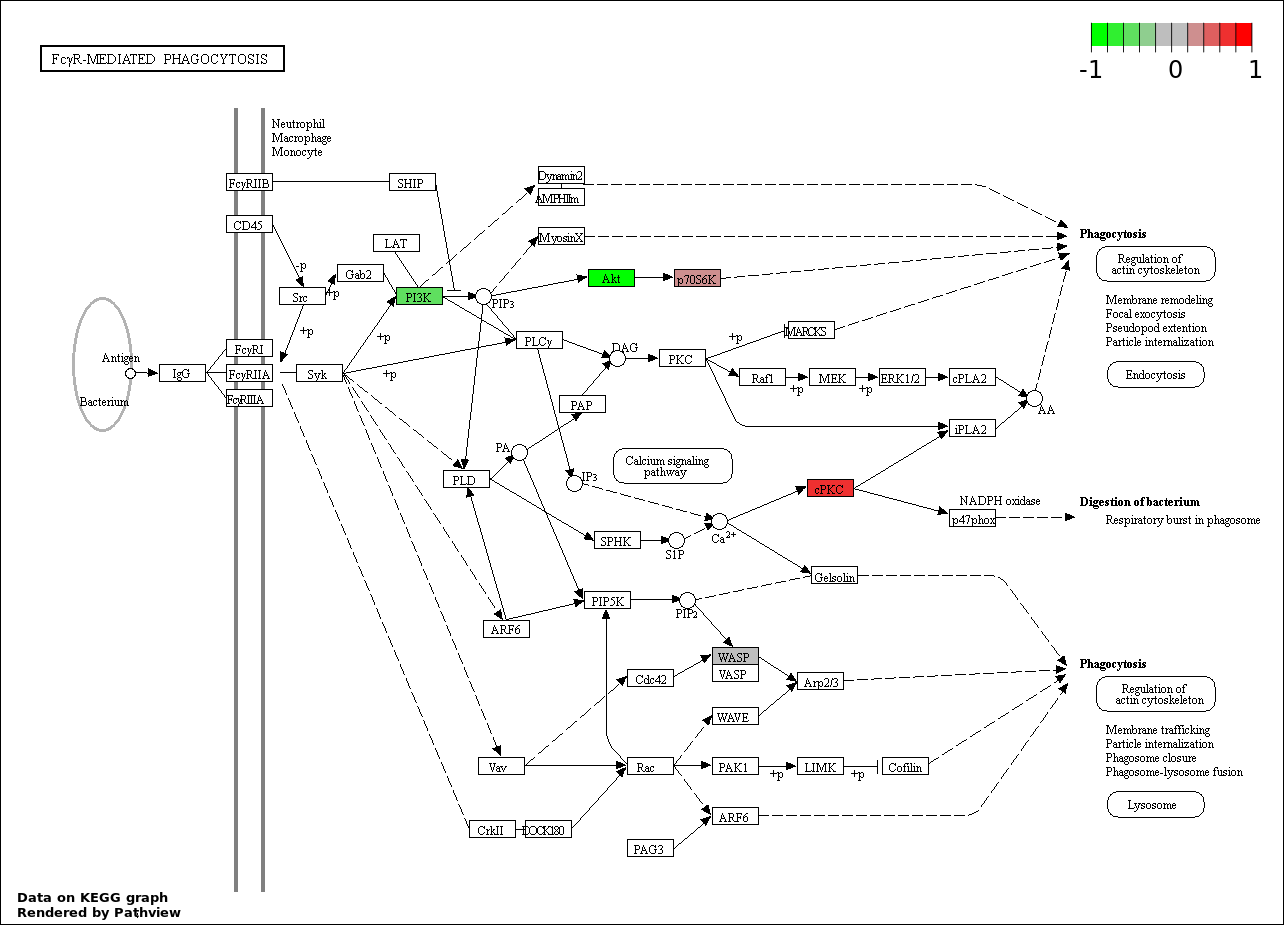
| 9 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 13 | 95 | 2.355e-08 | 4.709e-07 | |
| 10 | hsa04210_Apoptosis | 12 | 89 | 9.693e-08 | 1.616e-06 | |
| 11 | hsa04973_Carbohydrate_digestion_and_absorption | 9 | 44 | 9.878e-08 | 1.616e-06 | |
| 12 | hsa04664_Fc_epsilon_RI_signaling_pathway | 11 | 79 | 2.382e-07 | 3.573e-06 | |
| 13 | hsa04722_Neurotrophin_signaling_pathway | 13 | 127 | 7.501e-07 | 1.039e-05 | |
| 14 | hsa04662_B_cell_receptor_signaling_pathway | 10 | 75 | 1.274e-06 | 1.638e-05 | |
| 15 | hsa04070_Phosphatidylinositol_signaling_system | 10 | 78 | 1.84e-06 | 2.208e-05 | |
| 16 | hsa04920_Adipocytokine_signaling_pathway | 9 | 68 | 4.63e-06 | 5.209e-05 | |
| 17 | hsa04014_Ras_signaling_pathway | 16 | 236 | 9.901e-06 | 0.0001048 | |
| 18 | hsa04380_Osteoclast_differentiation | 11 | 128 | 2.843e-05 | 0.0002843 | |
| 19 | hsa04660_T_cell_receptor_signaling_pathway | 10 | 108 | 3.452e-05 | 0.0003271 | |
| 20 | hsa04710_Circadian_rhythm_._mammal | 5 | 23 | 5.604e-05 | 0.0005044 | |
| 21 | hsa04620_Toll.like_receptor_signaling_pathway | 9 | 102 | 0.0001234 | 0.001058 | |
| 22 | hsa04630_Jak.STAT_signaling_pathway | 11 | 155 | 0.0001616 | 0.001322 | |
| 23 | hsa04062_Chemokine_signaling_pathway | 12 | 189 | 0.0002344 | 0.001794 | |
| 24 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 10 | 136 | 0.0002392 | 0.001794 | |
| 25 | hsa04670_Leukocyte_transendothelial_migration | 9 | 117 | 0.0003487 | 0.002511 | |
| 26 | hsa04810_Regulation_of_actin_cytoskeleton | 12 | 214 | 0.0007179 | 0.00497 | |
| 27 | hsa04114_Oocyte_meiosis | 8 | 114 | 0.001343 | 0.00895 | |
| 28 | hsa04010_MAPK_signaling_pathway | 13 | 268 | 0.001652 | 0.01062 | |
| 29 | hsa04720_Long.term_potentiation | 6 | 70 | 0.001941 | 0.01205 | |
| 30 | hsa00562_Inositol_phosphate_metabolism | 5 | 57 | 0.004156 | 0.02493 | |
| 31 | hsa04972_Pancreatic_secretion | 6 | 101 | 0.01163 | 0.06752 | |
| 32 | hsa04530_Tight_junction | 7 | 133 | 0.01246 | 0.07011 | |
| 33 | hsa04141_Protein_processing_in_endoplasmic_reticulum | 8 | 168 | 0.01373 | 0.07488 | |
| 34 | hsa04140_Regulation_of_autophagy | 3 | 34 | 0.02512 | 0.133 | |
| 35 | hsa04115_p53_signaling_pathway | 4 | 69 | 0.03993 | 0.2031 | |
| 36 | hsa04912_GnRH_signaling_pathway | 5 | 101 | 0.04076 | 0.2031 | |
| 37 | hsa03320_PPAR_signaling_pathway | 4 | 70 | 0.04175 | 0.2031 | |
| 38 | hsa04146_Peroxisome | 4 | 79 | 0.06021 | 0.2852 | |
| 39 | hsa04270_Vascular_smooth_muscle_contraction | 5 | 116 | 0.06622 | 0.3056 | |
| 40 | hsa04977_Vitamin_digestion_and_absorption | 2 | 24 | 0.07314 | 0.3291 | |
| 41 | hsa04970_Salivary_secretion | 4 | 89 | 0.085 | 0.3732 | |
| 42 | hsa04540_Gap_junction | 4 | 90 | 0.08772 | 0.3759 | |
| 43 | hsa00340_Histidine_metabolism | 2 | 29 | 0.1013 | 0.4146 | |
| 44 | hsa00020_Citrate_cycle_.TCA_cycle. | 2 | 30 | 0.1073 | 0.4292 | |
| 45 | hsa00640_Propanoate_metabolism | 2 | 32 | 0.1195 | 0.4675 | |
| 46 | hsa04730_Long.term_depression | 3 | 70 | 0.1423 | 0.5451 | |
| 47 | hsa04976_Bile_secretion | 3 | 71 | 0.1467 | 0.5501 | |
| 48 | hsa04310_Wnt_signaling_pathway | 5 | 151 | 0.1524 | 0.5597 | |
| 49 | hsa04520_Adherens_junction | 3 | 73 | 0.1555 | 0.5599 | |
| 50 | hsa04971_Gastric_acid_secretion | 3 | 74 | 0.16 | 0.5648 | |
| 51 | hsa00380_Tryptophan_metabolism | 2 | 42 | 0.1844 | 0.6255 | |
| 52 | hsa00564_Glycerophospholipid_metabolism | 3 | 80 | 0.1877 | 0.6255 | |
| 53 | hsa00071_Fatty_acid_metabolism | 2 | 43 | 0.1911 | 0.6255 | |
| 54 | hsa00860_Porphyrin_and_chlorophyll_metabolism | 2 | 43 | 0.1911 | 0.6255 | |
| 55 | hsa04350_TGF.beta_signaling_pathway | 3 | 85 | 0.2116 | 0.6801 | |
| 56 | hsa04020_Calcium_signaling_pathway | 5 | 177 | 0.2358 | 0.7296 | |
| 57 | hsa00561_Glycerolipid_metabolism | 2 | 50 | 0.2391 | 0.7296 | |
| 58 | hsa04120_Ubiquitin_mediated_proteolysis | 4 | 139 | 0.2612 | 0.7706 | |
| 59 | hsa04916_Melanogenesis | 3 | 101 | 0.2915 | 0.8198 | |
| 60 | hsa00590_Arachidonic_acid_metabolism | 2 | 59 | 0.3015 | 0.8349 | |
| 61 | hsa03013_RNA_transport | 4 | 152 | 0.3153 | 0.8471 | |
| 62 | hsa04144_Endocytosis | 5 | 203 | 0.3288 | 0.8577 | |
| 63 | hsa04514_Cell_adhesion_molecules_.CAMs. | 3 | 136 | 0.4672 | 1 | |
| 64 | hsa04512_ECM.receptor_interaction | 2 | 85 | 0.4725 | 1 | |
| 65 | hsa04640_Hematopoietic_cell_lineage | 2 | 88 | 0.4906 | 1 | |
| 66 | hsa00230_Purine_metabolism | 3 | 162 | 0.5846 | 1 | |
| 67 | hsa04142_Lysosome | 2 | 121 | 0.6627 | 1 | |
| 68 | hsa04360_Axon_guidance | 2 | 130 | 0.7007 | 1 | |
| 69 | hsa04390_Hippo_signaling_pathway | 2 | 154 | 0.7849 | 1 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HCG11 |
hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15a-5p;hsa-miR-17-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-362-3p;hsa-miR-421;hsa-miR-501-3p;hsa-miR-502-3p;hsa-miR-93-5p | 12 | AKT3 | Sponge network | -0.781 | 0 | -0.659 | 0.00047 | 0.574 |
| 2 | DHRS4-AS1 | hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-582-5p;hsa-miR-589-3p | 10 | PIK3R1 | Sponge network | -0.646 | 0.01829 | -0.892 | 0 | 0.557 |
| 3 | DNM3OS | hsa-miR-106b-5p;hsa-miR-107;hsa-miR-122-5p;hsa-miR-15a-5p;hsa-miR-17-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-32-3p;hsa-miR-502-3p;hsa-miR-616-5p;hsa-miR-93-5p | 11 | AKT3 | Sponge network | -2.094 | 1.0E-5 | -0.659 | 0.00047 | 0.541 |
| 4 | RP11-434D9.1 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-409-3p;hsa-miR-629-3p | 13 | PIK3R1 | Sponge network | -2.913 | 0 | -0.892 | 0 | 0.525 |
| 5 | RP11-290F5.1 |
hsa-miR-103a-2-5p;hsa-miR-15b-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-301a-3p;hsa-miR-3913-5p;hsa-miR-421;hsa-miR-485-3p;hsa-miR-940 | 10 | IGF1 | Sponge network | -1.679 | 5.0E-5 | -3.091 | 0 | 0.523 |
| 6 | LINC00261 | hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-15b-5p;hsa-miR-188-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-324-3p;hsa-miR-330-3p;hsa-miR-589-3p;hsa-miR-629-3p | 13 | PIK3R1 | Sponge network | -1.194 | 0 | -0.892 | 0 | 0.513 |
| 7 | RP11-513G11.3 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-93-5p | 10 | PIK3R1 | Sponge network | -2.342 | 5.0E-5 | -0.892 | 0 | 0.492 |
| 8 | RP11-12A2.3 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-188-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-93-5p | 14 | PIK3R1 | Sponge network | -4.779 | 0 | -0.892 | 0 | 0.489 |
| 9 | LINC01018 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-188-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-324-3p;hsa-miR-338-5p;hsa-miR-409-3p;hsa-miR-589-3p;hsa-miR-629-3p;hsa-miR-93-5p | 18 | PIK3R1 | Sponge network | -3.231 | 0 | -0.892 | 0 | 0.472 |
| 10 | LDLRAD4-AS1 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-188-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-589-3p;hsa-miR-93-5p | 13 | PIK3R1 | Sponge network | -3.366 | 0 | -0.892 | 0 | 0.45 |
| 11 | RP11-119D9.1 | hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-188-5p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-629-3p;hsa-miR-93-5p | 11 | PIK3R1 | Sponge network | -2.765 | 0 | -0.892 | 0 | 0.418 |
| 12 | LINC01057 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-26a-5p | 10 | PRKAA2 | Sponge network | 1.427 | 3.0E-5 | 0.91 | 0.01333 | 0.409 |
| 13 | RP11-290F5.1 |
hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-188-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-324-3p;hsa-miR-493-5p | 13 | PIK3R1 | Sponge network | -1.679 | 5.0E-5 | -0.892 | 0 | 0.402 |
| 14 | RP11-115J16.1 | hsa-miR-103a-3p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-185-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-324-3p;hsa-miR-338-5p;hsa-miR-589-3p;hsa-miR-93-5p | 12 | PIK3R1 | Sponge network | -2.038 | 7.0E-5 | -0.892 | 0 | 0.382 |
| 15 | LINC00238 | hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-188-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-629-3p | 12 | PIK3R1 | Sponge network | -4.997 | 0 | -0.892 | 0 | 0.369 |
| 16 | MAGI2-AS3 |
hsa-miR-17-5p;hsa-miR-192-3p;hsa-miR-193a-3p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-30d-5p;hsa-miR-3127-5p;hsa-miR-532-3p;hsa-miR-589-5p;hsa-miR-93-3p | 10 | RPS6KA2 | Sponge network | -1.801 | 0 | -0.43 | 0.0094 | 0.363 |
| 17 | AC004862.6 | hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-188-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-589-3p;hsa-miR-629-3p;hsa-miR-93-5p | 13 | PIK3R1 | Sponge network | -2.202 | 0.00081 | -0.892 | 0 | 0.357 |
| 18 | PART1 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-26a-5p;hsa-miR-30e-3p | 10 | PRKAA2 | Sponge network | 3.525 | 1.0E-5 | 0.91 | 0.01333 | 0.351 |
| 19 | RP11-250B2.6 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-188-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-93-5p | 12 | PIK3R1 | Sponge network | -0.98 | 2.0E-5 | -0.892 | 0 | 0.342 |
| 20 | AP000473.5 | hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-338-5p | 11 | PIK3R1 | Sponge network | -1.157 | 0.00884 | -0.892 | 0 | 0.341 |
| 21 | RP11-963H4.3 | hsa-miR-106b-5p;hsa-miR-15b-5p;hsa-miR-17-5p;hsa-miR-21-5p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-324-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p | 10 | PIK3R1 | Sponge network | -1.857 | 0.00116 | -0.892 | 0 | 0.328 |
| 22 | RP11-440G9.1 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-26a-5p | 10 | PRKAA2 | Sponge network | 4.78 | 0 | 0.91 | 0.01333 | 0.324 |
| 23 | RP11-407B7.1 | hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-188-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-324-3p;hsa-miR-338-5p;hsa-miR-582-5p;hsa-miR-629-3p;hsa-miR-93-5p | 13 | PIK3R1 | Sponge network | -0.818 | 0.00584 | -0.892 | 0 | 0.324 |
| 24 | RP1-140K8.5 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-26a-5p | 10 | PRKAA2 | Sponge network | 1.954 | 0 | 0.91 | 0.01333 | 0.323 |
| 25 | LINC00864 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-15b-5p;hsa-miR-185-5p;hsa-miR-21-5p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-629-3p | 10 | PIK3R1 | Sponge network | -3.954 | 0 | -0.892 | 0 | 0.322 |
| 26 | MAGI2-AS3 |
hsa-miR-106b-5p;hsa-miR-107;hsa-miR-17-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-362-3p;hsa-miR-362-5p;hsa-miR-421;hsa-miR-616-5p;hsa-miR-93-5p | 10 | AKT3 | Sponge network | -1.801 | 0 | -0.659 | 0.00047 | 0.322 |
| 27 | RP11-166D19.1 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-15b-5p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-188-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-222-3p;hsa-miR-324-3p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-629-3p | 14 | PIK3R1 | Sponge network | -0.244 | 0.28835 | -0.892 | 0 | 0.32 |
| 28 | LINC00511 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-26a-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-628-3p | 12 | PRKAA2 | Sponge network | 2.468 | 0 | 0.91 | 0.01333 | 0.309 |
| 29 | DLGAP1-AS2 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-26a-5p;hsa-miR-30e-3p | 11 | PRKAA2 | Sponge network | 1.357 | 0 | 0.91 | 0.01333 | 0.306 |
| 30 | LINC00885 | hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-188-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-324-3p | 10 | PIK3R1 | Sponge network | -4.686 | 0 | -0.892 | 0 | 0.3 |
| 31 | MAGI2-AS3 |
hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-15b-5p;hsa-miR-17-5p;hsa-miR-188-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-589-3p;hsa-miR-629-3p;hsa-miR-93-5p | 13 | PIK3R1 | Sponge network | -1.801 | 0 | -0.892 | 0 | 0.294 |
| 32 | GAS5 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p;hsa-miR-576-5p | 13 | PRKAA2 | Sponge network | 1.966 | 0 | 0.91 | 0.01333 | 0.294 |
| 33 | DLGAP1-AS1 | hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-5p | 10 | PRKAA2 | Sponge network | 0.91 | 0 | 0.91 | 0.01333 | 0.293 |
| 34 | AC005562.1 | hsa-let-7b-3p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-628-3p | 13 | PRKAA2 | Sponge network | 1.127 | 0 | 0.91 | 0.01333 | 0.29 |
| 35 | AC016747.3 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-5p;hsa-miR-576-5p | 13 | PRKAA2 | Sponge network | 1.235 | 0 | 0.91 | 0.01333 | 0.29 |
| 36 | RP4-717I23.3 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p;hsa-miR-628-3p | 13 | PRKAA2 | Sponge network | 1.867 | 0 | 0.91 | 0.01333 | 0.285 |
| 37 | RP1-228H13.5 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-576-5p | 14 | PRKAA2 | Sponge network | 1.554 | 0 | 0.91 | 0.01333 | 0.285 |
| 38 | AC005550.3 |
hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-188-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-629-3p | 10 | PIK3R1 | Sponge network | -2.571 | 0.00132 | -0.892 | 0 | 0.281 |
| 39 | AP001469.9 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-576-5p | 13 | PRKAA2 | Sponge network | 2.428 | 0 | 0.91 | 0.01333 | 0.279 |
| 40 | AC074117.10 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p | 12 | PRKAA2 | Sponge network | 1.254 | 0 | 0.91 | 0.01333 | 0.275 |
| 41 | RP11-121C2.2 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p;hsa-miR-30e-5p | 12 | PRKAA2 | Sponge network | 1.286 | 0 | 0.91 | 0.01333 | 0.273 |
| 42 | RP11-7F17.3 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-188-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-324-3p | 11 | PIK3R1 | Sponge network | -0.873 | 0.00204 | -0.892 | 0 | 0.271 |
| 43 | SMIM2-AS1 | hsa-miR-103a-3p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-589-3p;hsa-miR-629-3p | 13 | PIK3R1 | Sponge network | -0.66 | 0.00587 | -0.892 | 0 | 0.27 |
| 44 | RP11-133K1.6 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-576-5p | 10 | PRKAA2 | Sponge network | 1.056 | 0 | 0.91 | 0.01333 | 0.269 |
| 45 | TMCC1-AS1 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p | 11 | PRKAA2 | Sponge network | 2.298 | 0 | 0.91 | 0.01333 | 0.258 |
| 46 | NUTM2A-AS1 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-139-5p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p;hsa-miR-576-5p | 10 | PRKAA2 | Sponge network | 0.788 | 0 | 0.91 | 0.01333 | 0.256 |
| 47 | WAC-AS1 | hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p | 10 | PRKAA2 | Sponge network | 0.869 | 0 | 0.91 | 0.01333 | 0.255 |
| 48 | HCG20 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-5p | 10 | PRKAA2 | Sponge network | 3.632 | 0 | 0.91 | 0.01333 | 0.253 |
| 49 | KB-1460A1.5 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p | 11 | PRKAA2 | Sponge network | 1.949 | 0 | 0.91 | 0.01333 | 0.253 |
| 50 | LINC00680 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p | 11 | PRKAA2 | Sponge network | 1.404 | 0 | 0.91 | 0.01333 | 0.252 |
| 51 | AC073283.4 | hsa-let-7b-3p;hsa-let-7b-5p;hsa-let-7c-5p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-195-3p;hsa-miR-22-5p;hsa-miR-26a-5p;hsa-miR-30e-3p;hsa-miR-30e-5p | 13 | PRKAA2 | Sponge network | 1.514 | 0 | 0.91 | 0.01333 | 0.251 |