This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-192-5p | ABCA8 | 2.1 | 0.00233 | -0.99 | 0.28561 | miRNAWalker2 validate | -0.45 | 0 | NA | |
| 2 | hsa-miR-103a-2-5p | ABCD2 | -2.06 | 0.00116 | -1.98 | 0.01409 | MirTarget | -0.26 | 0.00558 | NA | |
| 3 | hsa-miR-142-3p | ABHD11 | -2.4 | 4.0E-5 | 0.87 | 0.03845 | miRanda | -0.18 | 0.00067 | NA | |
| 4 | hsa-miR-142-3p | ABHD12 | -2.4 | 4.0E-5 | 0.22 | 0.40184 | miRanda | -0.15 | 0 | NA | |
| 5 | hsa-miR-103a-2-5p | ABL2 | -2.06 | 0.00116 | -0.13 | 0.63539 | MirTarget | -0.13 | 6.0E-5 | NA | |
| 6 | hsa-miR-141-5p | ABL2 | 2.11 | 0.00092 | -0.13 | 0.63539 | MirTarget; miRNATAP | -0.14 | 1.0E-5 | NA | |
| 7 | hsa-miR-200a-5p | ABL2 | 2.65 | 1.0E-5 | -0.13 | 0.63539 | mirMAP | -0.16 | 0 | NA | |
| 8 | hsa-miR-194-3p | ACAD11 | 2.07 | 0.00253 | -0.57 | 0.03547 | MirTarget | -0.11 | 0.00017 | NA | |
| 9 | hsa-miR-192-5p | ACSL1 | 2.1 | 0.00233 | -0.25 | 0.53807 | miRNAWalker2 validate | -0.15 | 0.00051 | NA | |
| 10 | hsa-miR-142-3p | ACTR3B | -2.4 | 4.0E-5 | 0.29 | 0.32393 | miRanda | -0.1 | 0.00359 | NA | |
| 11 | hsa-miR-141-5p | ACVRL1 | 2.11 | 0.00092 | -0.37 | 0.23741 | MirTarget | -0.23 | 0 | NA | |
| 12 | hsa-miR-194-3p | ADAM11 | 2.07 | 0.00253 | -0.08 | 0.89144 | mirMAP | -0.22 | 0.00035 | NA | |
| 13 | hsa-miR-103a-2-5p | ADAM22 | -2.06 | 0.00116 | -0.88 | 0.25527 | mirMAP | -0.31 | 0.00069 | NA | |
| 14 | hsa-miR-194-3p | ADAMTS15 | 2.07 | 0.00253 | 0.16 | 0.85349 | MirTarget | -0.38 | 2.0E-5 | NA | |
| 15 | hsa-miR-194-5p | ADAMTS5 | 2.55 | 0.00024 | -0.73 | 0.15695 | mirMAP | -0.22 | 3.0E-5 | NA | |
| 16 | hsa-miR-192-5p | ADARB1 | 2.1 | 0.00233 | -0.6 | 0.08985 | mirMAP | -0.26 | 0 | NA | |
| 17 | hsa-miR-141-5p | ADCY1 | 2.11 | 0.00092 | -0.03 | 0.96543 | mirMAP | -0.24 | 0.0093 | NA | |
| 18 | hsa-miR-200a-5p | ADD2 | 2.65 | 1.0E-5 | -1.1 | 0.14442 | mirMAP | -0.53 | 0 | NA | |
| 19 | hsa-miR-192-5p | AFAP1 | 2.1 | 0.00233 | 0.11 | 0.75207 | mirMAP | -0.1 | 0.0041 | NA | |
| 20 | hsa-miR-192-5p | AFAP1L1 | 2.1 | 0.00233 | -0.13 | 0.74099 | miRNAWalker2 validate | -0.27 | 0 | NA | |
| 21 | hsa-miR-141-5p | AFF2 | 2.11 | 0.00092 | -0.79 | 0.24367 | mirMAP | -0.34 | 1.0E-5 | NA | |
| 22 | hsa-miR-192-5p | AFF2 | 2.1 | 0.00233 | -0.79 | 0.24367 | mirMAP | -0.23 | 0.00104 | NA | |
| 23 | hsa-miR-194-3p | AFF2 | 2.07 | 0.00253 | -0.79 | 0.24367 | MirTarget | -0.17 | 0.01833 | NA | |
| 24 | hsa-miR-194-5p | AFF2 | 2.55 | 0.00024 | -0.79 | 0.24367 | mirMAP | -0.19 | 0.00694 | NA | |
| 25 | hsa-miR-141-5p | AFF3 | 2.11 | 0.00092 | -0.98 | 0.22174 | mirMAP | -0.43 | 0 | NA | |
| 26 | hsa-miR-200a-5p | AFF3 | 2.65 | 1.0E-5 | -0.98 | 0.22174 | MirTarget | -0.44 | 0 | NA | |
| 27 | hsa-miR-142-3p | AGXT | -2.4 | 4.0E-5 | 1.09 | 0.31324 | miRanda | -0.31 | 0.02223 | NA | |
| 28 | hsa-miR-141-5p | AIF1L | 2.11 | 0.00092 | -1.32 | 0.02648 | MirTarget | -0.31 | 0 | NA | |
| 29 | hsa-miR-142-3p | AK5 | -2.4 | 4.0E-5 | 0.41 | 0.61895 | miRanda | -0.33 | 0.00111 | NA | |
| 30 | hsa-miR-200a-5p | AKAP2 | 2.65 | 1.0E-5 | -0.87 | 0.26357 | mirMAP | -0.22 | 0.0164 | NA | |
| 31 | hsa-miR-103a-2-5p | AKR1C1 | -2.06 | 0.00116 | -0.93 | 0.17886 | mirMAP | -0.27 | 0.00074 | NA | |
| 32 | hsa-miR-142-3p | AKT1S1 | -2.4 | 4.0E-5 | 0.36 | 0.14789 | MirTarget; miRanda; miRNATAP | -0.15 | 0 | NA | |
| 33 | hsa-miR-192-5p | ALCAM | 2.1 | 0.00233 | 0.09 | 0.85523 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.12 | 0.02294 | 21119604 | MicroRNA 192 and 215 are upregulated in human gastric cancer in vivo and suppress ALCAM expression in vitro; Both transfection of mimics of microRNA-192 or -215 and ALCAM knockdown by an ALCAM-specific siRNA significantly increased the migration of HFE145 cells |
| 34 | hsa-miR-103a-2-5p | ALDH1L2 | -2.06 | 0.00116 | 0.27 | 0.69125 | mirMAP | -0.34 | 1.0E-5 | NA | |
| 35 | hsa-miR-194-3p | ALDH1L2 | 2.07 | 0.00253 | 0.27 | 0.69125 | MirTarget | -0.28 | 8.0E-5 | NA | |
| 36 | hsa-miR-194-5p | ALDH1L2 | 2.55 | 0.00024 | 0.27 | 0.69125 | mirMAP | -0.31 | 1.0E-5 | NA | |
| 37 | hsa-miR-142-3p | AMN1 | -2.4 | 4.0E-5 | 0.09 | 0.76433 | miRanda | -0.15 | 3.0E-5 | NA | |
| 38 | hsa-miR-192-5p | AMOTL1 | 2.1 | 0.00233 | -0.52 | 0.23224 | mirMAP | -0.29 | 0 | NA | |
| 39 | hsa-miR-141-5p | AMZ1 | 2.11 | 0.00092 | -0.84 | 0.19343 | mirMAP | -0.28 | 0.00012 | NA | |
| 40 | hsa-miR-194-3p | AMZ1 | 2.07 | 0.00253 | -0.84 | 0.19343 | mirMAP | -0.28 | 3.0E-5 | NA | |
| 41 | hsa-miR-144-5p | ANG | -2.21 | 0.00517 | 1.15 | 0.04302 | MirTarget | -0.14 | 0.00945 | NA | |
| 42 | hsa-miR-194-5p | ANGPTL1 | 2.55 | 0.00024 | -0.94 | 0.28852 | MirTarget | -0.43 | 0 | NA | |
| 43 | hsa-miR-194-3p | ANGPTL6 | 2.07 | 0.00253 | -1.69 | 0.01351 | MirTarget | -0.29 | 5.0E-5 | NA | |
| 44 | hsa-miR-192-5p | ANKRD6 | 2.1 | 0.00233 | -0.42 | 0.28815 | miRNAWalker2 validate | -0.16 | 0.00014 | NA | |
| 45 | hsa-miR-194-3p | ANTXR1 | 2.07 | 0.00253 | -0.08 | 0.90778 | MirTarget | -0.23 | 0.00113 | NA | |
| 46 | hsa-miR-103a-2-5p | ANTXR2 | -2.06 | 0.00116 | 0.45 | 0.35344 | MirTarget | -0.18 | 0.00119 | NA | |
| 47 | hsa-miR-141-5p | AP1S2 | 2.11 | 0.00092 | -0.98 | 0.02184 | MirTarget | -0.36 | 0 | NA | |
| 48 | hsa-miR-192-5p | AP1S2 | 2.1 | 0.00233 | -0.98 | 0.02184 | miRNAWalker2 validate | -0.27 | 0 | NA | |
| 49 | hsa-miR-215-5p | AP1S2 | 3.07 | 0.00158 | -0.98 | 0.02184 | miRNAWalker2 validate | -0.13 | 8.0E-5 | NA | |
| 50 | hsa-miR-192-5p | APOLD1 | 2.1 | 0.00233 | 0.05 | 0.92161 | miRNAWalker2 validate | -0.29 | 0 | NA | |
| 51 | hsa-miR-194-3p | APOLD1 | 2.07 | 0.00253 | 0.05 | 0.92161 | MirTarget | -0.25 | 1.0E-5 | NA | |
| 52 | hsa-miR-215-5p | APOLD1 | 3.07 | 0.00158 | 0.05 | 0.92161 | miRNAWalker2 validate | -0.18 | 0 | NA | |
| 53 | hsa-miR-200a-5p | AR | 2.65 | 1.0E-5 | -0.52 | 0.55352 | miRNATAP | -0.22 | 0.03873 | 24391862 | We identified miR-200 b as a downstream target of androgen receptor and linked its expression to decreased tumorigenicity and metastatic capacity of the prostate cancer cells |
| 54 | hsa-miR-142-3p | ARF5 | -2.4 | 4.0E-5 | 0.16 | 0.49632 | miRanda | -0.13 | 0 | NA | |
| 55 | hsa-miR-141-5p | ARHGAP19 | 2.11 | 0.00092 | -0.39 | 0.08267 | mirMAP | -0.12 | 0 | NA | |
| 56 | hsa-miR-192-5p | ARHGAP19 | 2.1 | 0.00233 | -0.39 | 0.08267 | miRNAWalker2 validate | -0.11 | 1.0E-5 | NA | |
| 57 | hsa-miR-192-5p | ARHGAP29 | 2.1 | 0.00233 | -0.8 | 0.07773 | miRNAWalker2 validate | -0.21 | 1.0E-5 | NA | |
| 58 | hsa-miR-215-5p | ARHGAP29 | 3.07 | 0.00158 | -0.8 | 0.07773 | miRNAWalker2 validate | -0.11 | 0.0008 | NA | |
| 59 | hsa-miR-192-5p | ARID5B | 2.1 | 0.00233 | -0.68 | 0.07996 | mirMAP | -0.23 | 0 | NA | |
| 60 | hsa-miR-142-3p | ARL1 | -2.4 | 4.0E-5 | 0.29 | 0.19981 | miRanda; miRNATAP | -0.14 | 0 | NA | |
| 61 | hsa-miR-192-5p | ARL4C | 2.1 | 0.00233 | -0.63 | 0.25561 | miRNAWalker2 validate | -0.29 | 0 | NA | |
| 62 | hsa-miR-215-5p | ARL4C | 3.07 | 0.00158 | -0.63 | 0.25561 | miRNAWalker2 validate | -0.11 | 0.00667 | NA | |
| 63 | hsa-miR-142-3p | ARPC1A | -2.4 | 4.0E-5 | 0.57 | 0.03866 | miRanda | -0.16 | 0 | NA | |
| 64 | hsa-miR-194-5p | ASAP1 | 2.55 | 0.00024 | -0.35 | 0.27587 | MirTarget; miRNATAP | -0.23 | 0 | NA | |
| 65 | hsa-miR-142-3p | ATG4D | -2.4 | 4.0E-5 | 0.16 | 0.62544 | miRanda | -0.16 | 3.0E-5 | NA | |
| 66 | hsa-miR-103a-2-5p | ATP10B | -2.06 | 0.00116 | 2.38 | 0.05795 | mirMAP | -0.44 | 0.00317 | NA | |
| 67 | hsa-miR-192-5p | ATP10D | 2.1 | 0.00233 | -0.5 | 0.15756 | miRNAWalker2 validate | -0.14 | 0.00017 | NA | |
| 68 | hsa-miR-142-3p | ATP1B1 | -2.4 | 4.0E-5 | 1.43 | 0.00225 | PITA; miRanda; miRNATAP | -0.18 | 0.00175 | NA | |
| 69 | hsa-miR-192-5p | ATP8B4 | 2.1 | 0.00233 | -1.37 | 0.01009 | MirTarget | -0.29 | 0 | NA | |
| 70 | hsa-miR-194-5p | ATP8B4 | 2.55 | 0.00024 | -1.37 | 0.01009 | MirTarget | -0.28 | 0 | NA | |
| 71 | hsa-miR-200a-5p | ATP8B4 | 2.65 | 1.0E-5 | -1.37 | 0.01009 | MirTarget | -0.34 | 0 | NA | |
| 72 | hsa-miR-215-5p | ATP8B4 | 3.07 | 0.00158 | -1.37 | 0.01009 | MirTarget | -0.11 | 0.00536 | NA | |
| 73 | hsa-miR-142-3p | ATRN | -2.4 | 4.0E-5 | -0.04 | 0.87648 | miRanda | -0.1 | 0.00261 | NA | |
| 74 | hsa-miR-142-3p | ATXN2 | -2.4 | 4.0E-5 | 0.19 | 0.36948 | miRanda | -0.12 | 1.0E-5 | NA | |
| 75 | hsa-miR-192-5p | B3GALNT1 | 2.1 | 0.00233 | 0.12 | 0.70806 | miRNAWalker2 validate | -0.12 | 0.00054 | NA | |
| 76 | hsa-miR-141-5p | BACH2 | 2.11 | 0.00092 | -1.75 | 0.00364 | miRNATAP | -0.42 | 0 | NA | |
| 77 | hsa-miR-103a-2-5p | BAG4 | -2.06 | 0.00116 | -0.02 | 0.95003 | MirTarget | -0.14 | 0.00125 | NA | |
| 78 | hsa-miR-103a-2-5p | BARD1 | -2.06 | 0.00116 | -0.02 | 0.97017 | MirTarget | -0.17 | 0.02781 | NA | |
| 79 | hsa-miR-103a-2-5p | BCAS1 | -2.06 | 0.00116 | 3.1 | 0.00836 | mirMAP | -0.32 | 0.02129 | NA | |
| 80 | hsa-miR-192-5p | BCL2 | 2.1 | 0.00233 | -1.63 | 0.00261 | miRNAWalker2 validate | -0.31 | 0 | 26550150 | MicroRNA 192 regulates chemo resistance of lung adenocarcinoma for gemcitabine and cisplatin combined therapy by targeting Bcl 2; In this paper we try to test whether miR-192 regulates chemo-resistance in human carcinoma A549 mice model by targeting Bcl-2; MTT assay real-time RT-PCR western blotting assay were used to investigate miR-192 expression levels cell viability ratio and Bcl-2 protein expression levels; Bcl-2 mRNA and protein expression levels up-regulated in miR-192 inhibitor treated tumor; Bcl-2 is a key regulator for miR-192 related chemotherapy resistance; In this study we demonstrate that miR-192 regulates chemoresistance for gemcitabine and cisplatin combined chemotherapy in human adenocarcinoma lung cancer A549 cells and Bcl-2 is the target of miR-192 |
| 81 | hsa-miR-200a-5p | BCL2 | 2.65 | 1.0E-5 | -1.63 | 0.00261 | mirMAP | -0.37 | 0 | NA | |
| 82 | hsa-miR-215-5p | BCL2 | 3.07 | 0.00158 | -1.63 | 0.00261 | miRNAWalker2 validate | -0.11 | 0.00607 | NA | |
| 83 | hsa-miR-192-5p | BEND4 | 2.1 | 0.00233 | -1.9 | 0.08641 | mirMAP | -0.36 | 0.00188 | NA | |
| 84 | hsa-miR-200a-5p | BEND4 | 2.65 | 1.0E-5 | -1.9 | 0.08641 | mirMAP | -0.49 | 0.00018 | NA | |
| 85 | hsa-miR-142-3p | BHLHB9 | -2.4 | 4.0E-5 | 0.16 | 0.58081 | miRanda | -0.12 | 0.0007 | NA | |
| 86 | hsa-miR-192-5p | BHLHE22 | 2.1 | 0.00233 | -0.66 | 0.3246 | MirTarget; miRNATAP | -0.43 | 0 | NA | |
| 87 | hsa-miR-215-5p | BHLHE22 | 3.07 | 0.00158 | -0.66 | 0.3246 | MirTarget | -0.2 | 5.0E-5 | NA | |
| 88 | hsa-miR-194-5p | BICD2 | 2.55 | 0.00024 | -0.53 | 0.06121 | miRNATAP | -0.24 | 0 | NA | |
| 89 | hsa-miR-194-3p | BMF | 2.07 | 0.00253 | -1.61 | 2.0E-5 | mirMAP | -0.19 | 0 | NA | |
| 90 | hsa-miR-192-5p | BMP2K | 2.1 | 0.00233 | -1.07 | 0.00527 | miRNAWalker2 validate | -0.11 | 0.00682 | NA | |
| 91 | hsa-miR-194-3p | BMP8A | 2.07 | 0.00253 | -0.62 | 0.21754 | MirTarget | -0.24 | 0 | NA | |
| 92 | hsa-miR-103a-2-5p | BMPR2 | -2.06 | 0.00116 | -0.03 | 0.9248 | MirTarget | -0.12 | 0.00039 | NA | |
| 93 | hsa-miR-194-5p | BNIP2 | 2.55 | 0.00024 | -0.4 | 0.1487 | MirTarget | -0.11 | 0.00011 | NA | |
| 94 | hsa-miR-142-3p | BNIP3 | -2.4 | 4.0E-5 | -0.11 | 0.85548 | miRanda | -0.15 | 0.03481 | NA | |
| 95 | hsa-miR-142-3p | BOD1 | -2.4 | 4.0E-5 | 0.19 | 0.28558 | miRNAWalker2 validate; MirTarget; miRanda; miRNATAP | -0.12 | 0 | NA | |
| 96 | hsa-miR-142-3p | BOLA1 | -2.4 | 4.0E-5 | 0.2 | 0.53027 | miRanda | -0.12 | 0.00135 | NA | |
| 97 | hsa-miR-192-5p | BTF3L4 | 2.1 | 0.00233 | -0.07 | 0.72387 | miRNAWalker2 validate | -0.1 | 0 | NA | |
| 98 | hsa-miR-192-5p | BVES | 2.1 | 0.00233 | -0.04 | 0.94915 | mirMAP | -0.21 | 0.001 | NA | |
| 99 | hsa-miR-194-3p | C10orf105 | 2.07 | 0.00253 | -1.54 | 0.0032 | mirMAP | -0.14 | 0.01047 | NA | |
| 100 | hsa-miR-192-5p | C15orf52 | 2.1 | 0.00233 | 0.11 | 0.85147 | mirMAP | -0.16 | 0.0093 | NA | |
| 101 | hsa-miR-194-3p | C1QTNF1 | 2.07 | 0.00253 | -0.43 | 0.26096 | mirMAP | -0.13 | 0.00076 | NA | |
| 102 | hsa-miR-192-5p | C1QTNF3 | 2.1 | 0.00233 | -0.15 | 0.8394 | miRNAWalker2 validate | -0.27 | 0.0004 | NA | |
| 103 | hsa-miR-194-3p | C20orf194 | 2.07 | 0.00253 | -0.45 | 0.19186 | miRNATAP | -0.11 | 0.00251 | NA | |
| 104 | hsa-miR-192-5p | CAB39L | 2.1 | 0.00233 | 0.39 | 0.24273 | miRNAWalker2 validate | -0.12 | 0.00077 | NA | |
| 105 | hsa-miR-194-3p | CABP7 | 2.07 | 0.00253 | 0.79 | 0.4844 | mirMAP; miRNATAP | -0.27 | 0.02249 | NA | |
| 106 | hsa-miR-194-3p | CACHD1 | 2.07 | 0.00253 | -0.83 | 0.06065 | MirTarget; miRNATAP | -0.18 | 0.00016 | NA | |
| 107 | hsa-miR-142-3p | CACNA1D | -2.4 | 4.0E-5 | 0.6 | 0.3219 | miRanda | -0.21 | 0.00465 | NA | |
| 108 | hsa-miR-141-5p | CACNA1E | 2.11 | 0.00092 | -0.51 | 0.44626 | mirMAP | -0.43 | 0 | NA | |
| 109 | hsa-miR-200a-5p | CACNA1E | 2.65 | 1.0E-5 | -0.51 | 0.44626 | mirMAP | -0.28 | 0.00026 | NA | |
| 110 | hsa-miR-142-3p | CACNA2D2 | -2.4 | 4.0E-5 | 0.74 | 0.45001 | miRanda | -0.32 | 0.00842 | NA | |
| 111 | hsa-miR-192-5p | CADM1 | 2.1 | 0.00233 | -0.53 | 0.36556 | miRNAWalker2 validate | -0.22 | 0.00028 | NA | |
| 112 | hsa-miR-194-5p | CADM1 | 2.55 | 0.00024 | -0.53 | 0.36556 | miRNATAP | -0.22 | 0.00023 | NA | |
| 113 | hsa-miR-103a-2-5p | CADM2 | -2.06 | 0.00116 | -0.65 | 0.48039 | mirMAP | -0.3 | 0.0055 | NA | |
| 114 | hsa-miR-142-3p | CALM1 | -2.4 | 4.0E-5 | 0.06 | 0.79466 | miRanda; miRNATAP | -0.12 | 4.0E-5 | NA | |
| 115 | hsa-miR-141-5p | CAMK1 | 2.11 | 0.00092 | -0.66 | 0.03634 | MirTarget | -0.24 | 0 | NA | |
| 116 | hsa-miR-141-5p | CAMK1D | 2.11 | 0.00092 | -1.41 | 0.01231 | mirMAP | -0.2 | 0.0022 | NA | |
| 117 | hsa-miR-141-5p | CARD8 | 2.11 | 0.00092 | -0.89 | 0.00259 | MirTarget | -0.23 | 0 | NA | |
| 118 | hsa-miR-194-5p | CARD8 | 2.55 | 0.00024 | -0.89 | 0.00259 | mirMAP | -0.13 | 2.0E-5 | NA | |
| 119 | hsa-miR-194-3p | CAV1 | 2.07 | 0.00253 | -0.35 | 0.54059 | MirTarget | -0.46 | 0 | NA | |
| 120 | hsa-miR-141-5p | CBL | 2.11 | 0.00092 | -0.48 | 0.10129 | mirMAP | -0.12 | 0.00022 | NA | |
| 121 | hsa-miR-192-5p | CBL | 2.1 | 0.00233 | -0.48 | 0.10129 | mirMAP | -0.13 | 2.0E-5 | NA | |
| 122 | hsa-miR-194-3p | CBX6 | 2.07 | 0.00253 | -0.4 | 0.42264 | mirMAP | -0.19 | 0.00022 | NA | |
| 123 | hsa-miR-142-3p | CBY1 | -2.4 | 4.0E-5 | 0.18 | 0.35088 | miRanda | -0.13 | 0 | NA | |
| 124 | hsa-miR-142-3p | CCDC148 | -2.4 | 4.0E-5 | 0.71 | 0.15403 | miRanda | -0.25 | 3.0E-5 | NA | |
| 125 | hsa-miR-192-5p | CCDC18 | 2.1 | 0.00233 | -0.54 | 0.19501 | miRNAWalker2 validate | -0.15 | 0.00058 | NA | |
| 126 | hsa-miR-192-5p | CCDC69 | 2.1 | 0.00233 | -1.15 | 0.05012 | mirMAP | -0.29 | 0 | NA | |
| 127 | hsa-miR-200a-5p | CCDC88A | 2.65 | 1.0E-5 | -0.54 | 0.20423 | miRNATAP | -0.17 | 0.00097 | NA | |
| 128 | hsa-miR-142-3p | CCND1 | -2.4 | 4.0E-5 | 1.04 | 0.01709 | miRanda | -0.11 | 0.04224 | 23619912 | Transfection of miR-142-3p mimics in colon cancer cells downregulated cyclin D1 expression induced G1 phase cell cycle arrest and elevated the sensitivity of the cells to 5-fluorouracil |
| 129 | hsa-miR-141-5p | CD226 | 2.11 | 0.00092 | -1.6 | 0.03423 | MirTarget | -0.44 | 0 | NA | |
| 130 | hsa-miR-192-5p | CD83 | 2.1 | 0.00233 | -1.46 | 0.00797 | miRNAWalker2 validate | -0.28 | 0 | NA | |
| 131 | hsa-miR-215-5p | CD83 | 3.07 | 0.00158 | -1.46 | 0.00797 | miRNAWalker2 validate | -0.12 | 0.00447 | NA | |
| 132 | hsa-miR-192-5p | CD93 | 2.1 | 0.00233 | -0.36 | 0.42563 | MirTarget | -0.27 | 0 | NA | |
| 133 | hsa-miR-215-5p | CD93 | 3.07 | 0.00158 | -0.36 | 0.42563 | MirTarget | -0.11 | 0.00149 | NA | |
| 134 | hsa-miR-192-5p | CDC14A | 2.1 | 0.00233 | -0.8 | 0.05646 | miRNAWalker2 validate | -0.15 | 0.00099 | NA | |
| 135 | hsa-miR-194-5p | CDC14A | 2.55 | 0.00024 | -0.8 | 0.05646 | miRNATAP | -0.14 | 0.00173 | NA | |
| 136 | hsa-miR-144-5p | CDC42BPA | -2.21 | 0.00517 | 1.03 | 0.00092 | MirTarget | -0.11 | 0.00024 | NA | |
| 137 | hsa-miR-192-5p | CDCA4 | 2.1 | 0.00233 | 0.25 | 0.42479 | miRNAWalker2 validate | -0.11 | 0.0009 | NA | |
| 138 | hsa-miR-150-3p | CDCP1 | -2.15 | 0.0055 | 1.81 | 0.00037 | MirTarget | -0.22 | 1.0E-5 | NA | |
| 139 | hsa-miR-194-5p | CDH2 | 2.55 | 0.00024 | -0.86 | 0.27079 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.35 | 1.0E-5 | 21845495; 20979124 | The results of real-time PCR and western blot highlighted that miR-194 interacted with N-cadherin and negatively regulated its expression at the translational level;The overexpression of miR-194 in liver mesenchymal-like cancer cells reduced the expression of the mesenchymal cell marker N-cadherin and suppressed invasion and migration of the mesenchymal-like cancer cells both in vitro and in vivo |
| 140 | hsa-miR-194-3p | CDH23 | 2.07 | 0.00253 | -1.68 | 0.00275 | mirMAP | -0.27 | 1.0E-5 | NA | |
| 141 | hsa-miR-192-5p | CDK14 | 2.1 | 0.00233 | -0.45 | 0.37362 | miRNAWalker2 validate | -0.35 | 0 | NA | |
| 142 | hsa-miR-194-5p | CDK14 | 2.55 | 0.00024 | -0.45 | 0.37362 | MirTarget | -0.33 | 0 | NA | |
| 143 | hsa-miR-215-5p | CDK14 | 3.07 | 0.00158 | -0.45 | 0.37362 | miRNAWalker2 validate | -0.18 | 0 | NA | |
| 144 | hsa-miR-142-3p | CDKL2 | -2.4 | 4.0E-5 | -0.3 | 0.66224 | MirTarget; miRanda | -0.27 | 0.00122 | NA | |
| 145 | hsa-miR-141-5p | CDO1 | 2.11 | 0.00092 | -0.75 | 0.28667 | MirTarget | -0.32 | 6.0E-5 | NA | |
| 146 | hsa-miR-192-5p | CDON | 2.1 | 0.00233 | -0.13 | 0.81496 | miRNAWalker2 validate | -0.36 | 0 | NA | |
| 147 | hsa-miR-194-3p | CDON | 2.07 | 0.00253 | -0.13 | 0.81496 | MirTarget | -0.3 | 0 | NA | |
| 148 | hsa-miR-141-5p | CDR1 | 2.11 | 0.00092 | -0.32 | 0.68902 | MirTarget | -0.27 | 0.00269 | NA | |
| 149 | hsa-miR-142-3p | CDS1 | -2.4 | 4.0E-5 | 1.72 | 8.0E-5 | miRanda | -0.24 | 1.0E-5 | NA | |
| 150 | hsa-miR-194-3p | CENPP | 2.07 | 0.00253 | 0.24 | 0.49727 | mirMAP | -0.15 | 5.0E-5 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 82 | 788 | 3.834e-14 | 8.92e-11 |
| 2 | CIRCULATORY SYSTEM DEVELOPMENT | 82 | 788 | 3.834e-14 | 8.92e-11 |
| 3 | REGULATION OF CELL DIFFERENTIATION | 121 | 1492 | 1.994e-12 | 3.092e-09 |
| 4 | NEUROGENESIS | 115 | 1402 | 3.706e-12 | 4.311e-09 |
| 5 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 130 | 1672 | 5.187e-12 | 4.827e-09 |
| 6 | CELL DEVELOPMENT | 112 | 1426 | 1.068e-10 | 7.102e-08 |
| 7 | BIOLOGICAL ADHESION | 89 | 1032 | 1.055e-10 | 7.102e-08 |
| 8 | VASCULATURE DEVELOPMENT | 52 | 469 | 2.296e-10 | 1.336e-07 |
| 9 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 109 | 1395 | 2.664e-10 | 1.377e-07 |
| 10 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 77 | 872 | 7.277e-10 | 3.386e-07 |
| 11 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 82 | 957 | 8.289e-10 | 3.506e-07 |
| 12 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 69 | 750 | 1.07e-09 | 3.83e-07 |
| 13 | TISSUE DEVELOPMENT | 114 | 1518 | 1.034e-09 | 3.83e-07 |
| 14 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 100 | 1275 | 1.311e-09 | 4.357e-07 |
| 15 | REGULATION OF CELL PROJECTION ORGANIZATION | 56 | 558 | 1.962e-09 | 5.705e-07 |
| 16 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 53 | 513 | 1.955e-09 | 5.705e-07 |
| 17 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 38 | 306 | 2.847e-09 | 7.791e-07 |
| 18 | BLOOD VESSEL MORPHOGENESIS | 42 | 364 | 3.913e-09 | 9.769e-07 |
| 19 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 72 | 823 | 3.989e-09 | 9.769e-07 |
| 20 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 90 | 1142 | 7.596e-09 | 1.767e-06 |
| 21 | REGULATION OF NEURON DIFFERENTIATION | 54 | 554 | 1.06e-08 | 2.349e-06 |
| 22 | CELLULAR COMPONENT MORPHOGENESIS | 75 | 900 | 1.505e-08 | 3.183e-06 |
| 23 | LOCOMOTION | 87 | 1114 | 2.101e-08 | 4.25e-06 |
| 24 | HEAD DEVELOPMENT | 63 | 709 | 2.195e-08 | 4.256e-06 |
| 25 | CELL MOTILITY | 70 | 835 | 3.695e-08 | 6.275e-06 |
| 26 | LOCALIZATION OF CELL | 70 | 835 | 3.695e-08 | 6.275e-06 |
| 27 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 43 | 408 | 3.776e-08 | 6.275e-06 |
| 28 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 81 | 1021 | 3.411e-08 | 6.275e-06 |
| 29 | POSITIVE REGULATION OF NEURON PROJECTION DEVELOPMENT | 30 | 232 | 5.312e-08 | 8.524e-06 |
| 30 | REGULATION OF CELL DEVELOPMENT | 69 | 836 | 8.479e-08 | 1.315e-05 |
| 31 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 44 | 437 | 9.913e-08 | 1.488e-05 |
| 32 | CELL MORPHOGENESIS INVOLVED IN NEURON DIFFERENTIATION | 39 | 368 | 1.405e-07 | 2.043e-05 |
| 33 | ACTIN FILAMENT BASED PROCESS | 44 | 450 | 2.278e-07 | 3.211e-05 |
| 34 | ANGIOGENESIS | 33 | 293 | 3.203e-07 | 4.384e-05 |
| 35 | NEURON DIFFERENTIATION | 69 | 874 | 4.482e-07 | 5.795e-05 |
| 36 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 63 | 771 | 4.484e-07 | 5.795e-05 |
| 37 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 32 | 285 | 5.168e-07 | 6.499e-05 |
| 38 | REGULATION OF DENDRITE DEVELOPMENT | 19 | 120 | 6.736e-07 | 8.244e-05 |
| 39 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 33 | 303 | 6.91e-07 | 8.244e-05 |
| 40 | TUBE DEVELOPMENT | 49 | 552 | 8.605e-07 | 0.0001001 |
| 41 | NEURON DEVELOPMENT | 57 | 687 | 9.923e-07 | 0.0001126 |
| 42 | PROTEIN LOCALIZATION | 118 | 1805 | 1.07e-06 | 0.0001185 |
| 43 | NEURAL PRECURSOR CELL PROLIFERATION | 14 | 70 | 1.122e-06 | 0.0001215 |
| 44 | MUSCLE STRUCTURE DEVELOPMENT | 41 | 432 | 1.271e-06 | 0.0001344 |
| 45 | NEURON PROJECTION MORPHOGENESIS | 39 | 402 | 1.342e-06 | 0.0001388 |
| 46 | NEURON PROJECTION DEVELOPMENT | 48 | 545 | 1.391e-06 | 0.0001407 |
| 47 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 94 | 1360 | 1.532e-06 | 0.0001517 |
| 48 | TISSUE MORPHOGENESIS | 47 | 533 | 1.732e-06 | 0.0001679 |
| 49 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 76 | 1036 | 1.955e-06 | 0.0001779 |
| 50 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 76 | 1036 | 1.955e-06 | 0.0001779 |
| 51 | ORGAN MORPHOGENESIS | 65 | 841 | 2.017e-06 | 0.0001779 |
| 52 | HETEROPHILIC CELL CELL ADHESION VIA PLASMA MEMBRANE CELL ADHESION MOLECULES | 11 | 45 | 1.972e-06 | 0.0001779 |
| 53 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 73 | 983 | 2.027e-06 | 0.0001779 |
| 54 | POSITIVE REGULATION OF CELL DEVELOPMENT | 43 | 472 | 2.067e-06 | 0.0001781 |
| 55 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 109 | 1656 | 2.13e-06 | 0.0001802 |
| 56 | POSITIVE REGULATION OF DENDRITE DEVELOPMENT | 13 | 64 | 2.25e-06 | 0.0001869 |
| 57 | REGULATION OF CELLULAR COMPONENT SIZE | 34 | 337 | 2.685e-06 | 0.0002191 |
| 58 | TUBE MORPHOGENESIS | 33 | 323 | 2.858e-06 | 0.0002293 |
| 59 | MORPHOGENESIS OF AN EPITHELIUM | 38 | 400 | 3.025e-06 | 0.0002386 |
| 60 | REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 16 | 98 | 3.352e-06 | 0.00026 |
| 61 | EPITHELIUM DEVELOPMENT | 70 | 945 | 3.622e-06 | 0.0002675 |
| 62 | CYTOSKELETON ORGANIZATION | 64 | 838 | 3.588e-06 | 0.0002675 |
| 63 | HEART DEVELOPMENT | 42 | 466 | 3.56e-06 | 0.0002675 |
| 64 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 106 | 1618 | 3.695e-06 | 0.0002687 |
| 65 | POSITIVE REGULATION OF ENDOTHELIAL CELL MIGRATION | 13 | 67 | 3.864e-06 | 0.0002766 |
| 66 | CELL PART MORPHOGENESIS | 52 | 633 | 3.968e-06 | 0.0002797 |
| 67 | RESPONSE TO ENDOGENOUS STIMULUS | 97 | 1450 | 4.198e-06 | 0.0002915 |
| 68 | TELENCEPHALON DEVELOPMENT | 26 | 228 | 4.467e-06 | 0.0003057 |
| 69 | POSITIVE REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 13 | 68 | 4.595e-06 | 0.0003098 |
| 70 | REGULATION OF TRANSPORT | 115 | 1804 | 4.96e-06 | 0.0003297 |
| 71 | CELL PROJECTION ORGANIZATION | 67 | 902 | 5.379e-06 | 0.0003525 |
| 72 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 61 | 801 | 6.666e-06 | 0.0004308 |
| 73 | REGULATION OF CELL ADHESION | 51 | 629 | 7.079e-06 | 0.0004512 |
| 74 | NEGATIVE REGULATION OF LOCOMOTION | 28 | 263 | 7.346e-06 | 0.0004619 |
| 75 | RESPONSE TO WOUNDING | 47 | 563 | 7.583e-06 | 0.0004705 |
| 76 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 72 | 1008 | 9.265e-06 | 0.0005672 |
| 77 | FOREBRAIN DEVELOPMENT | 34 | 357 | 9.413e-06 | 0.0005688 |
| 78 | CELL PROLIFERATION | 53 | 672 | 1.028e-05 | 0.000613 |
| 79 | REGULATION OF ANATOMICAL STRUCTURE SIZE | 41 | 472 | 1.133e-05 | 0.0006674 |
| 80 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 60 | 799 | 1.215e-05 | 0.0007065 |
| 81 | RESPONSE TO GROWTH FACTOR | 41 | 475 | 1.317e-05 | 0.0007474 |
| 82 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 49 | 609 | 1.312e-05 | 0.0007474 |
| 83 | NEGATIVE REGULATION OF CELL COMMUNICATION | 81 | 1192 | 1.583e-05 | 0.0008872 |
| 84 | MULTICELLULAR ORGANISMAL SIGNALING | 17 | 123 | 1.676e-05 | 0.0009286 |
| 85 | CELLULAR MACROMOLECULE LOCALIZATION | 83 | 1234 | 1.794e-05 | 0.0009823 |
| 86 | NEUROBLAST PROLIFERATION | 8 | 29 | 1.927e-05 | 0.001043 |
| 87 | CELL SUBSTRATE ADHESION | 20 | 164 | 2.083e-05 | 0.001114 |
| 88 | SECRETION | 47 | 588 | 2.322e-05 | 0.001228 |
| 89 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 114 | 1848 | 2.386e-05 | 0.001247 |
| 90 | COGNITION | 26 | 251 | 2.491e-05 | 0.00126 |
| 91 | REGULATION OF EPITHELIAL CELL MIGRATION | 20 | 166 | 2.49e-05 | 0.00126 |
| 92 | SKELETAL SYSTEM DEVELOPMENT | 39 | 455 | 2.492e-05 | 0.00126 |
| 93 | REGULATION OF ACTIN FILAMENT BASED PROCESS | 30 | 312 | 2.593e-05 | 0.001267 |
| 94 | CELL CELL ADHESION | 48 | 608 | 2.614e-05 | 0.001267 |
| 95 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 111 | 1791 | 2.539e-05 | 0.001267 |
| 96 | CONNECTIVE TISSUE DEVELOPMENT | 22 | 194 | 2.596e-05 | 0.001267 |
| 97 | POSITIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 19 | 154 | 2.825e-05 | 0.001355 |
| 98 | REGULATION OF TRANSMEMBRANE TRANSPORT | 37 | 426 | 3.012e-05 | 0.00143 |
| 99 | RESPONSE TO TRANSFORMING GROWTH FACTOR BETA | 18 | 144 | 3.819e-05 | 0.001795 |
| 100 | REGULATION OF PROTEIN MODIFICATION PROCESS | 106 | 1710 | 3.912e-05 | 0.00182 |
| 101 | ACTION POTENTIAL | 14 | 94 | 3.953e-05 | 0.001821 |
| 102 | SYNAPSE ORGANIZATION | 18 | 145 | 4.192e-05 | 0.001912 |
| 103 | AMINOGLYCAN BIOSYNTHETIC PROCESS | 15 | 107 | 4.384e-05 | 0.001981 |
| 104 | REGULATION OF CELL MORPHOGENESIS | 44 | 552 | 4.494e-05 | 0.00201 |
| 105 | RESPONSE TO HORMONE | 63 | 893 | 4.897e-05 | 0.00217 |
| 106 | MEMBRANE ASSEMBLY | 7 | 25 | 5.82e-05 | 0.002555 |
| 107 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 51 | 689 | 7.879e-05 | 0.003426 |
| 108 | TAXIS | 38 | 464 | 8.355e-05 | 0.0036 |
| 109 | REGULATION OF ENDOTHELIAL CELL MIGRATION | 15 | 114 | 9.212e-05 | 0.003932 |
| 110 | SPROUTING ANGIOGENESIS | 9 | 45 | 9.34e-05 | 0.003951 |
| 111 | SINGLE ORGANISM BEHAVIOR | 33 | 384 | 9.776e-05 | 0.004098 |
| 112 | POSITIVE REGULATION OF CELL COMMUNICATION | 95 | 1532 | 0.0001006 | 0.004179 |
| 113 | RESPONSE TO CALCIUM ION | 15 | 115 | 0.0001019 | 0.004195 |
| 114 | REGULATION OF DENDRITIC SPINE DEVELOPMENT | 10 | 56 | 0.0001057 | 0.004228 |
| 115 | POSITIVE REGULATION OF AXON EXTENSION | 8 | 36 | 0.0001045 | 0.004228 |
| 116 | POSITIVE REGULATION OF MUSCLE TISSUE DEVELOPMENT | 10 | 56 | 0.0001057 | 0.004228 |
| 117 | GLYCOPROTEIN METABOLIC PROCESS | 31 | 353 | 0.0001063 | 0.004228 |
| 118 | EMBRYONIC MORPHOGENESIS | 42 | 539 | 0.0001102 | 0.004347 |
| 119 | MUSCLE TISSUE DEVELOPMENT | 26 | 275 | 0.0001157 | 0.004523 |
| 120 | NEGATIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 25 | 262 | 0.0001344 | 0.005212 |
| 121 | SYNAPSE ASSEMBLY | 11 | 69 | 0.0001405 | 0.005402 |
| 122 | SINGLE ORGANISM CELL ADHESION | 37 | 459 | 0.0001421 | 0.005419 |
| 123 | PROTEOGLYCAN BIOSYNTHETIC PROCESS | 10 | 58 | 0.0001435 | 0.005428 |
| 124 | REGULATION OF MEMBRANE POTENTIAL | 30 | 343 | 0.0001466 | 0.005503 |
| 125 | RESPONSE TO OXYGEN LEVELS | 28 | 311 | 0.0001487 | 0.005535 |
| 126 | POSITIVE REGULATION OF VASCULATURE DEVELOPMENT | 16 | 133 | 0.0001598 | 0.005853 |
| 127 | CELL CELL ADHESION VIA PLASMA MEMBRANE ADHESION MOLECULES | 21 | 204 | 0.0001591 | 0.005853 |
| 128 | POSITIVE REGULATION OF CARTILAGE DEVELOPMENT | 7 | 29 | 0.000163 | 0.005925 |
| 129 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 60 | 876 | 0.0001659 | 0.005982 |
| 130 | NEURON PROJECTION GUIDANCE | 21 | 205 | 0.0001704 | 0.006098 |
| 131 | REGULATION OF CELLULAR COMPONENT BIOGENESIS | 54 | 767 | 0.0001776 | 0.006217 |
| 132 | BEHAVIOR | 40 | 516 | 0.0001777 | 0.006217 |
| 133 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 18 | 162 | 0.0001777 | 0.006217 |
| 134 | PROTEOGLYCAN METABOLIC PROCESS | 12 | 83 | 0.0001854 | 0.006344 |
| 135 | CELLULAR RESPONSE TO HORMONE STIMULUS | 42 | 552 | 0.0001848 | 0.006344 |
| 136 | UROGENITAL SYSTEM DEVELOPMENT | 27 | 299 | 0.0001846 | 0.006344 |
| 137 | STEM CELL PROLIFERATION | 10 | 60 | 0.0001921 | 0.006524 |
| 138 | MEMBRANE BIOGENESIS | 7 | 30 | 0.0002049 | 0.006909 |
| 139 | REGULATION OF POTASSIUM ION TRANSMEMBRANE TRANSPORT | 10 | 61 | 0.0002212 | 0.007403 |
| 140 | HINDBRAIN DEVELOPMENT | 16 | 137 | 0.0002258 | 0.007405 |
| 141 | NEGATIVE REGULATION OF CELL DEVELOPMENT | 27 | 303 | 0.000229 | 0.007405 |
| 142 | POSITIVE REGULATION OF HYDROLASE ACTIVITY | 61 | 905 | 0.0002254 | 0.007405 |
| 143 | SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 30 | 352 | 0.0002302 | 0.007405 |
| 144 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 73 | 1135 | 0.0002308 | 0.007405 |
| 145 | REGULATION OF FILOPODIUM ASSEMBLY | 8 | 40 | 0.0002286 | 0.007405 |
| 146 | POSITIVE REGULATION OF LOCOMOTION | 34 | 420 | 0.0002411 | 0.007638 |
| 147 | AMINOGLYCAN METABOLIC PROCESS | 18 | 166 | 0.0002413 | 0.007638 |
| 148 | REGULATION OF SYSTEM PROCESS | 39 | 507 | 0.0002492 | 0.007833 |
| 149 | REGULATION OF DEVELOPMENTAL GROWTH | 26 | 289 | 0.0002557 | 0.007984 |
| 150 | PALLIUM DEVELOPMENT | 17 | 153 | 0.0002661 | 0.008255 |
| 151 | REGULATION OF CELL PROLIFERATION | 91 | 1496 | 0.0002735 | 0.008321 |
| 152 | LUNG ALVEOLUS DEVELOPMENT | 8 | 41 | 0.0002736 | 0.008321 |
| 153 | LACTATION | 8 | 41 | 0.0002736 | 0.008321 |
| 154 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 92 | 1518 | 0.0002861 | 0.008645 |
| 155 | REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 22 | 229 | 0.0003017 | 0.009057 |
| 156 | POSITIVE REGULATION OF DENDRITE MORPHOGENESIS | 7 | 32 | 0.0003147 | 0.009387 |
| 157 | GROWTH | 33 | 410 | 0.000328 | 0.009722 |
| 158 | REGULATION OF VESICLE MEDIATED TRANSPORT | 36 | 462 | 0.0003337 | 0.009752 |
| 159 | REGULATION OF ACTIN FILAMENT LENGTH | 17 | 156 | 0.0003352 | 0.009752 |
| 160 | REGULATION OF SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 25 | 278 | 0.0003354 | 0.009752 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CYTOSKELETAL PROTEIN BINDING | 77 | 819 | 3.86e-11 | 3.586e-08 |
| 2 | ACTIN BINDING | 44 | 393 | 4.232e-09 | 1.966e-06 |
| 3 | CALCIUM ION BINDING | 62 | 697 | 2.742e-08 | 8.49e-06 |
| 4 | CELL ADHESION MOLECULE BINDING | 26 | 186 | 8.52e-08 | 1.979e-05 |
| 5 | PROTEIN DOMAIN SPECIFIC BINDING | 55 | 624 | 2.298e-07 | 4.269e-05 |
| 6 | MACROMOLECULAR COMPLEX BINDING | 96 | 1399 | 1.605e-06 | 0.0002486 |
| 7 | PROTEIN COMPLEX BINDING | 70 | 935 | 2.502e-06 | 0.000332 |
| 8 | GROWTH FACTOR BINDING | 18 | 123 | 4.211e-06 | 0.000489 |
| 9 | RNA POLYMERASE II TRANSCRIPTION FACTOR ACTIVITY SEQUENCE SPECIFIC DNA BINDING | 50 | 629 | 1.483e-05 | 0.001517 |
| 10 | ALPHA ACTININ BINDING | 7 | 21 | 1.633e-05 | 0.001517 |
| 11 | INTEGRIN BINDING | 15 | 105 | 3.5e-05 | 0.002956 |
| 12 | GTPASE BINDING | 28 | 295 | 5.994e-05 | 0.00464 |
| 13 | CALMODULIN BINDING | 20 | 179 | 7.347e-05 | 0.004875 |
| 14 | CATION CHANNEL ACTIVITY | 28 | 298 | 7.154e-05 | 0.004875 |
| 15 | PDZ DOMAIN BINDING | 13 | 90 | 0.0001022 | 0.006328 |
| 16 | ENZYME BINDING | 105 | 1737 | 0.0001146 | 0.006653 |
| 17 | VOLTAGE GATED ION CHANNEL ACTIVITY | 20 | 190 | 0.0001673 | 0.007773 |
| 18 | STEROID HORMONE RECEPTOR ACTIVITY | 10 | 59 | 0.0001663 | 0.007773 |
| 19 | ACTININ BINDING | 7 | 29 | 0.000163 | 0.007773 |
| 20 | RAB GTPASE BINDING | 15 | 120 | 0.0001653 | 0.007773 |
| 21 | METAL ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 34 | 417 | 0.000211 | 0.009332 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CELL PROJECTION | 136 | 1786 | 6.823e-12 | 3.984e-09 |
| 2 | CELL JUNCTION | 97 | 1151 | 5.073e-11 | 1.481e-08 |
| 3 | NEURON PART | 103 | 1265 | 8.803e-11 | 1.714e-08 |
| 4 | NEURON PROJECTION | 83 | 942 | 1.67e-10 | 2.439e-08 |
| 5 | CELL PROJECTION PART | 78 | 946 | 1.208e-08 | 1.411e-06 |
| 6 | SYNAPSE | 66 | 754 | 1.794e-08 | 1.746e-06 |
| 7 | CELL LEADING EDGE | 39 | 350 | 3.675e-08 | 3.066e-06 |
| 8 | SOMATODENDRITIC COMPARTMENT | 57 | 650 | 1.636e-07 | 1.194e-05 |
| 9 | CELL SURFACE | 63 | 757 | 2.366e-07 | 1.535e-05 |
| 10 | LAMELLIPODIUM | 24 | 172 | 2.75e-07 | 1.606e-05 |
| 11 | ANCHORING JUNCTION | 45 | 489 | 9.106e-07 | 4.834e-05 |
| 12 | ACTIN CYTOSKELETON | 41 | 444 | 2.543e-06 | 0.0001238 |
| 13 | DENDRITE | 41 | 451 | 3.754e-06 | 0.0001687 |
| 14 | MEMBRANE REGION | 79 | 1134 | 8.37e-06 | 0.0003492 |
| 15 | FILOPODIUM | 15 | 94 | 9.003e-06 | 0.0003505 |
| 16 | SYNAPSE PART | 49 | 610 | 1.37e-05 | 0.0005002 |
| 17 | CATION CHANNEL COMPLEX | 20 | 167 | 2.718e-05 | 0.0009338 |
| 18 | GOLGI APPARATUS PART | 63 | 893 | 4.897e-05 | 0.001505 |
| 19 | PLASMA MEMBRANE REGION | 65 | 929 | 4.761e-05 | 0.001505 |
| 20 | SITE OF POLARIZED GROWTH | 18 | 149 | 6.024e-05 | 0.001759 |
| 21 | CELL SUBSTRATE JUNCTION | 34 | 398 | 8.661e-05 | 0.002409 |
| 22 | INTRINSIC COMPONENT OF PLASMA MEMBRANE | 101 | 1649 | 9.573e-05 | 0.002541 |
| 23 | CYTOSKELETON | 116 | 1967 | 0.0001347 | 0.003421 |
| 24 | CELL BODY | 39 | 494 | 0.0001466 | 0.003567 |
| 25 | INTRACELLULAR VESICLE | 80 | 1259 | 0.0001702 | 0.003976 |
| 26 | NEURON SPINE | 15 | 121 | 0.0001815 | 0.004077 |
| 27 | CELL CELL JUNCTION | 32 | 383 | 0.0002072 | 0.00442 |
| 28 | GOLGI APPARATUS | 89 | 1445 | 0.0002119 | 0.00442 |
| 29 | ACTIN BASED CELL PROJECTION | 19 | 181 | 0.00025 | 0.005034 |
| 30 | EXTRACELLULAR MATRIX COMPONENT | 15 | 125 | 0.0002606 | 0.005073 |
| 31 | POSTSYNAPSE | 31 | 378 | 0.0003571 | 0.006726 |
| 32 | CELL CELL ADHERENS JUNCTION | 9 | 54 | 0.0003989 | 0.00728 |
| 33 | SECRETORY GRANULE MEMBRANE | 11 | 78 | 0.0004243 | 0.007509 |
| 34 | AXON | 33 | 418 | 0.0004622 | 0.00794 |
| 35 | BASEMENT MEMBRANE | 12 | 93 | 0.0005425 | 0.008932 |
| 36 | PLASMA MEMBRANE PROTEIN COMPLEX | 38 | 510 | 0.0005506 | 0.008932 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
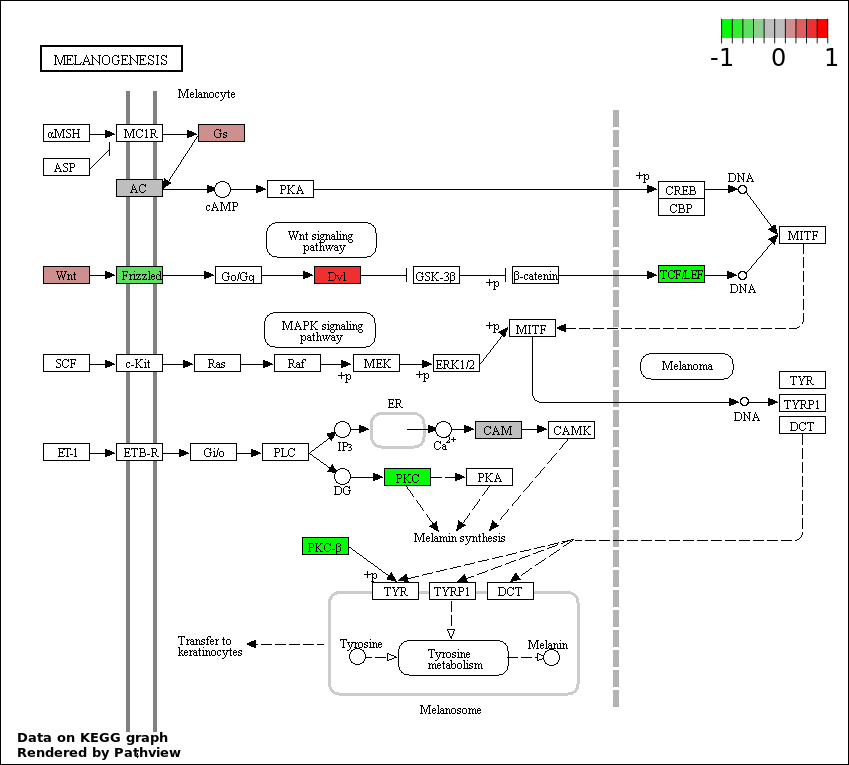
| 1 | hsa04916_Melanogenesis | 14 | 101 | 8.853e-05 | 0.00686 | |
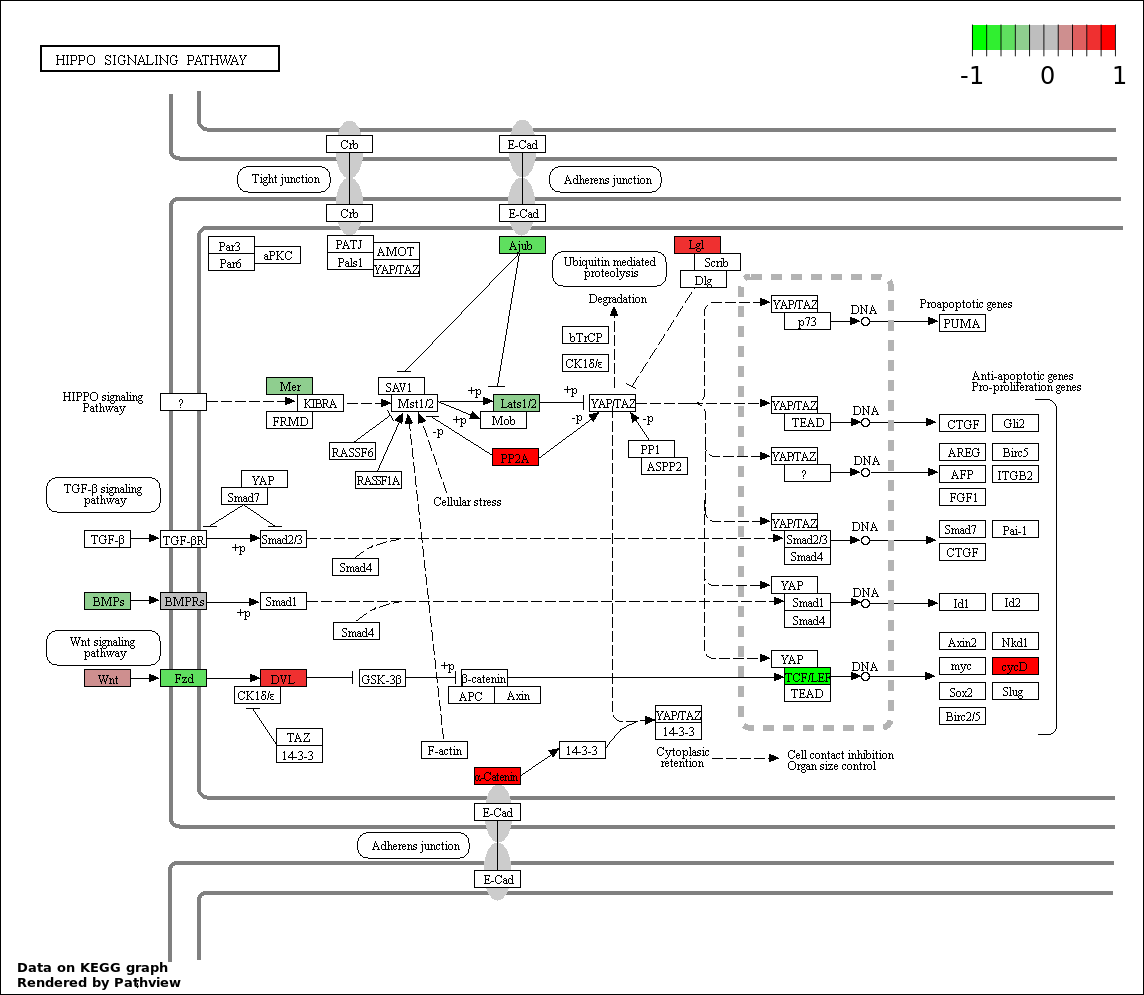
| 2 | hsa04390_Hippo_signaling_pathway | 18 | 154 | 9.286e-05 | 0.00686 | |
| 3 | hsa04810_Regulation_of_actin_cytoskeleton | 22 | 214 | 0.0001143 | 0.00686 | |
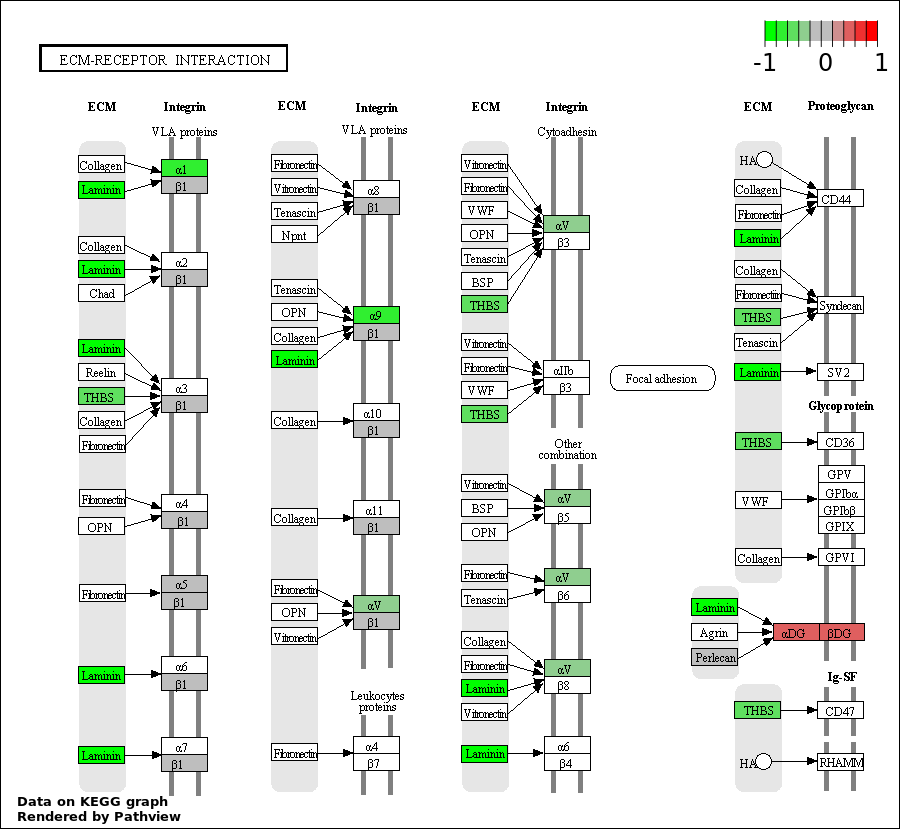
| 4 | hsa04512_ECM.receptor_interaction | 12 | 85 | 0.0002332 | 0.008441 | |
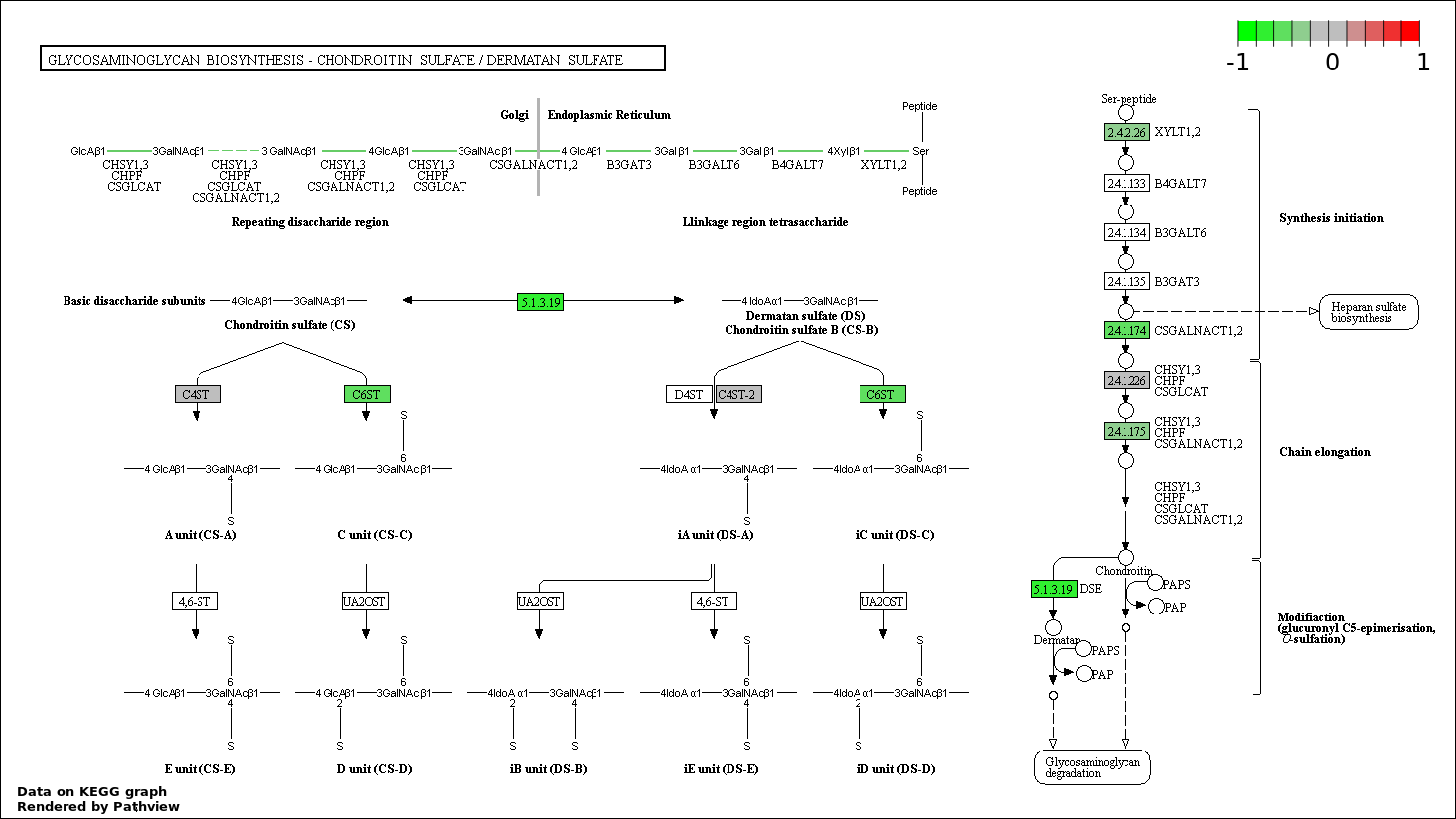
| 5 | hsa00532_Glycosaminoglycan_biosynthesis_._chondroitin_sulfate | 6 | 22 | 0.0002345 | 0.008441 | |
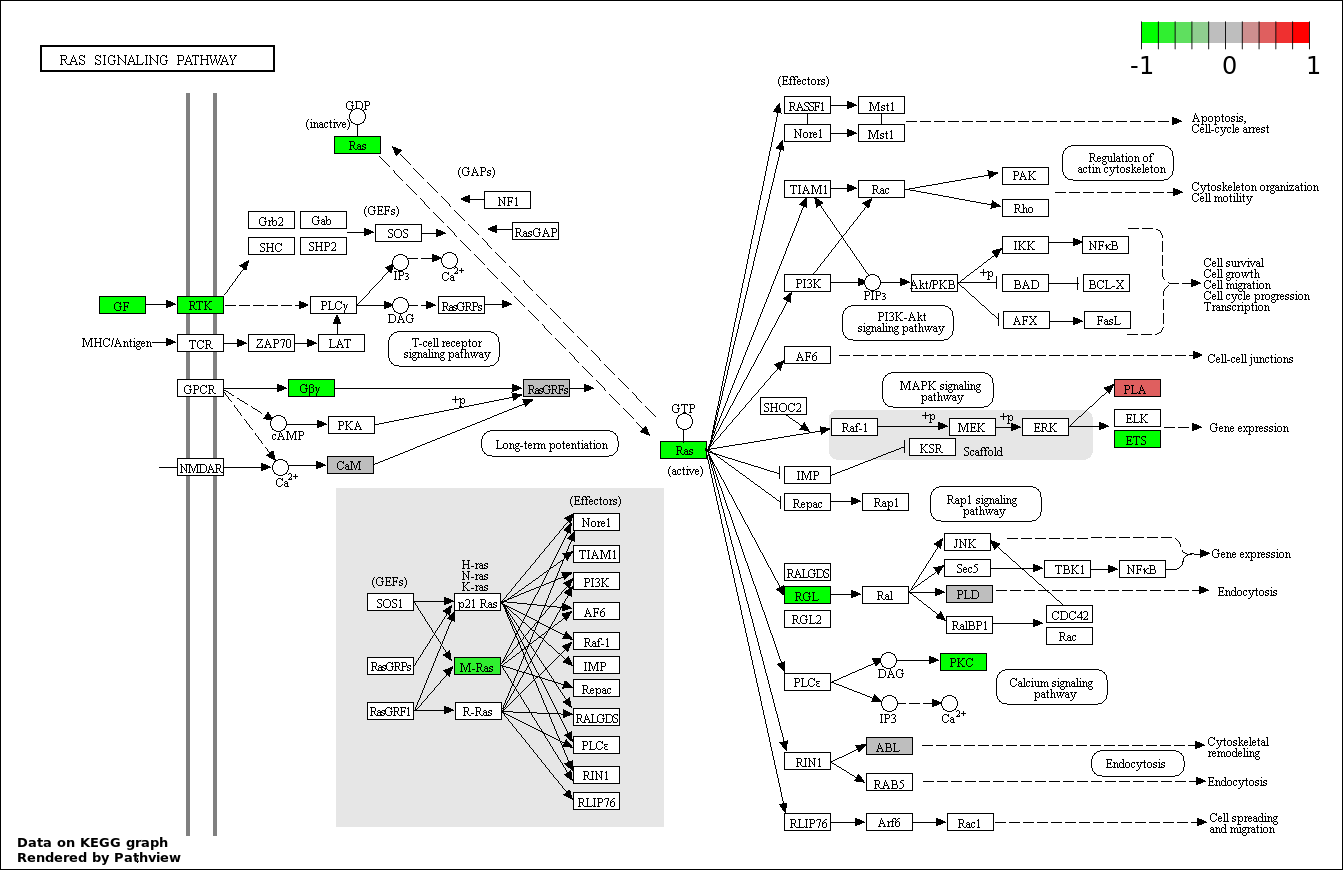
| 6 | hsa04014_Ras_signaling_pathway | 22 | 236 | 0.0004577 | 0.01373 | |
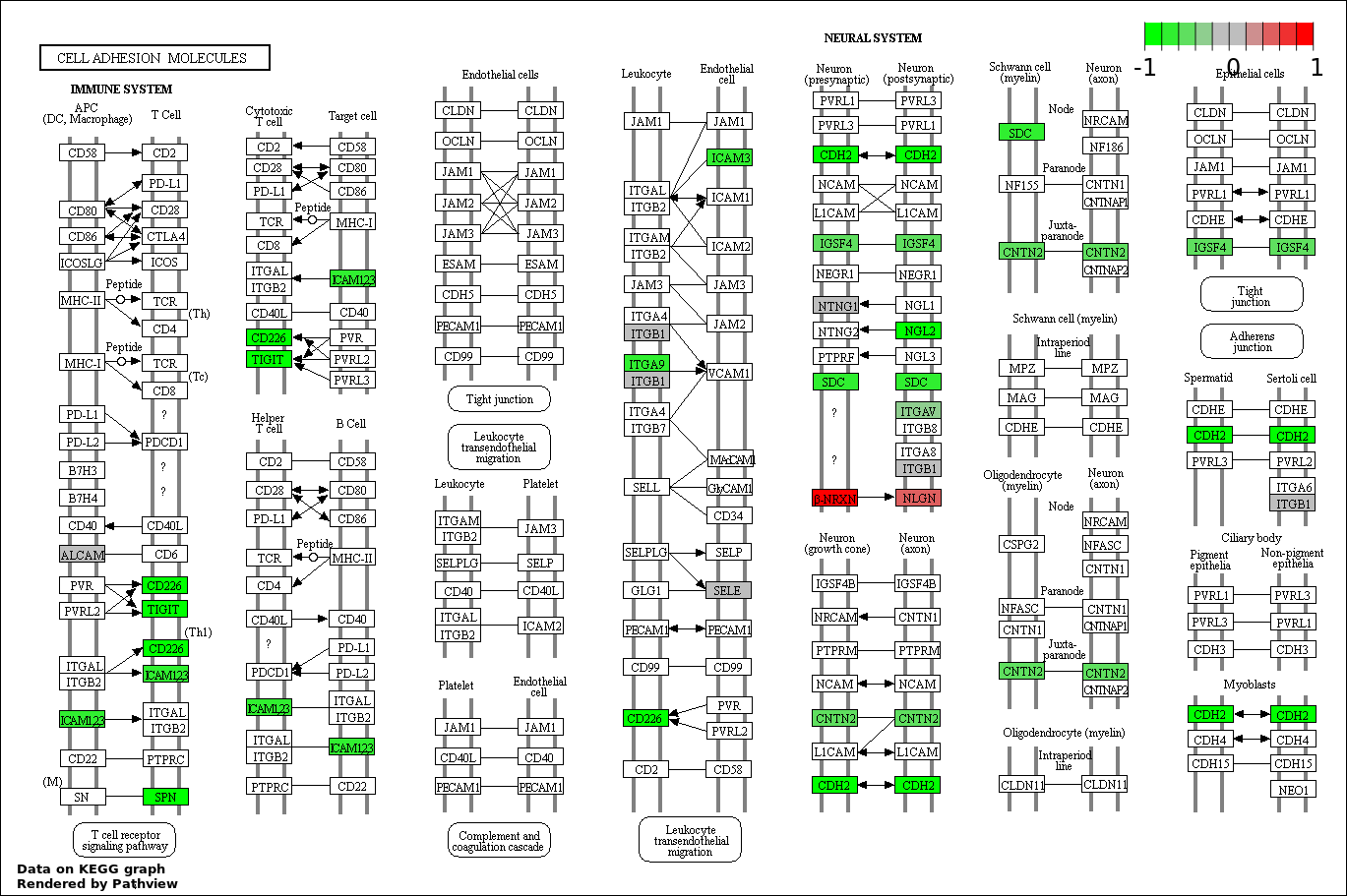
| 7 | hsa04514_Cell_adhesion_molecules_.CAMs. | 15 | 136 | 0.0006471 | 0.01521 | |
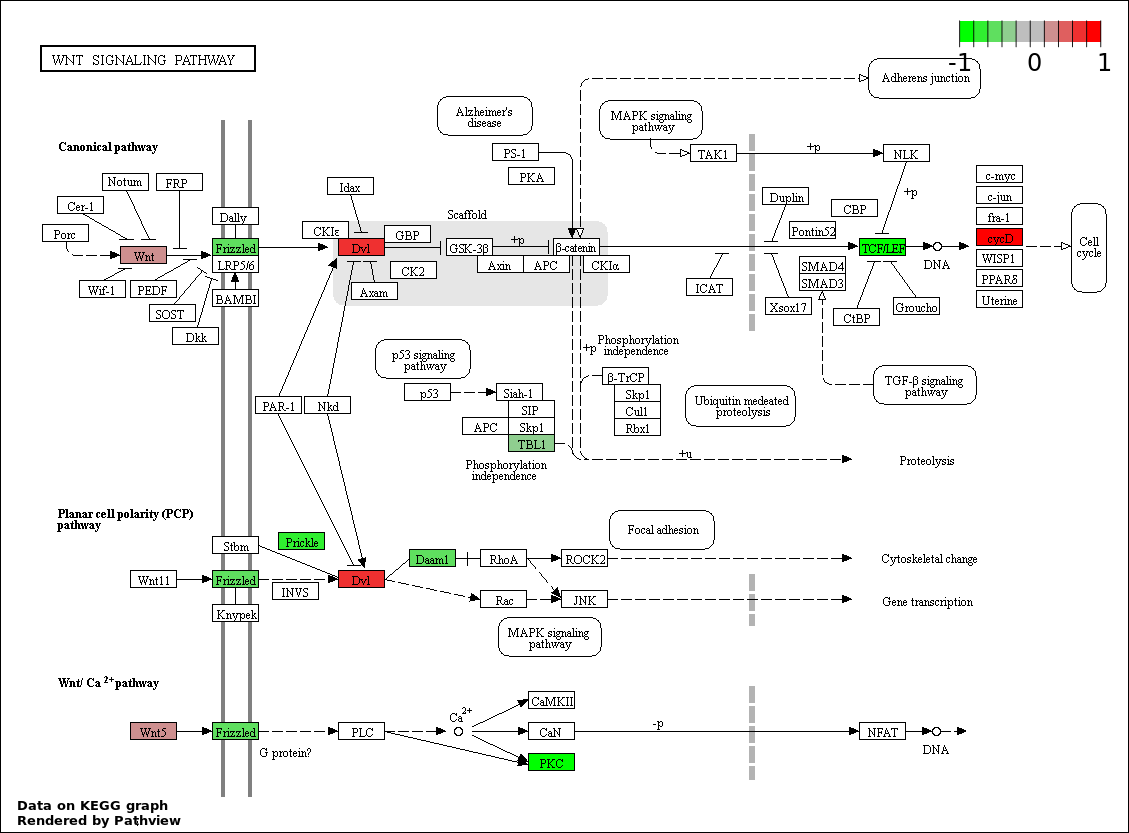
| 8 | hsa04310_Wnt_signaling_pathway | 16 | 151 | 0.0006758 | 0.01521 | |
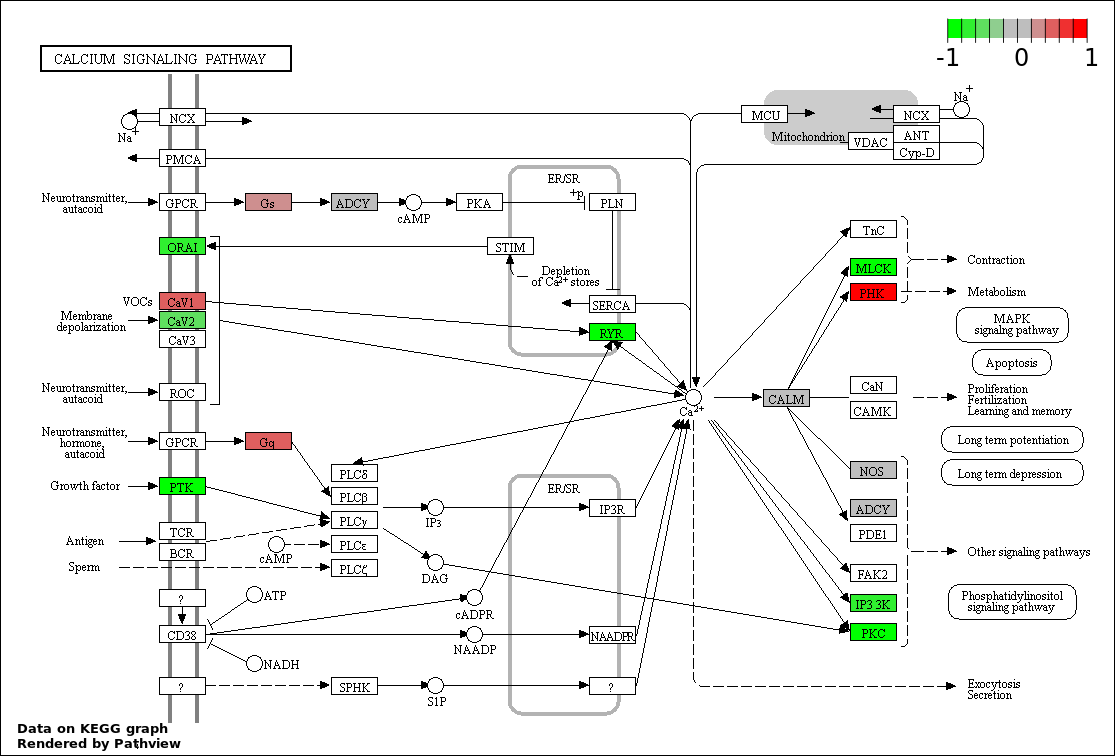
| 9 | hsa04020_Calcium_signaling_pathway | 17 | 177 | 0.001408 | 0.02815 | |
| 10 | hsa04151_PI3K_AKT_signaling_pathway | 27 | 351 | 0.00212 | 0.03515 | |
| 11 | hsa04510_Focal_adhesion | 18 | 200 | 0.002148 | 0.03515 | |
| 12 | hsa04912_GnRH_signaling_pathway | 11 | 101 | 0.003609 | 0.05137 | |
| 13 | hsa04270_Vascular_smooth_muscle_contraction | 12 | 116 | 0.00371 | 0.05137 | |
| 14 | hsa04730_Long.term_depression | 8 | 70 | 0.009238 | 0.1188 | |
| 15 | hsa04971_Gastric_acid_secretion | 8 | 74 | 0.01274 | 0.1451 | |
| 16 | hsa04970_Salivary_secretion | 9 | 89 | 0.0129 | 0.1451 | |
| 17 | hsa04010_MAPK_signaling_pathway | 19 | 268 | 0.01988 | 0.2105 | |
| 18 | hsa00534_Glycosaminoglycan_biosynthesis_._heparan_sulfate | 4 | 26 | 0.02264 | 0.2264 | |
| 19 | hsa04972_Pancreatic_secretion | 9 | 101 | 0.02724 | 0.2581 | |
| 20 | hsa04520_Adherens_junction | 7 | 73 | 0.03444 | 0.31 | |
| 21 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 8 | 95 | 0.04796 | 0.4031 | |
| 22 | hsa04360_Axon_guidance | 10 | 130 | 0.04926 | 0.4031 | |
| 23 | hsa00510_N.Glycan_biosynthesis | 5 | 49 | 0.05519 | 0.4192 | |
| 24 | hsa04530_Tight_junction | 10 | 133 | 0.05589 | 0.4192 | |
| 25 | hsa00565_Ether_lipid_metabolism | 4 | 36 | 0.06424 | 0.4625 | |
| 26 | hsa04350_TGF.beta_signaling_pathway | 7 | 85 | 0.0682 | 0.4722 | |
| 27 | hsa04710_Circadian_rhythm_._mammal | 3 | 23 | 0.07124 | 0.4749 | |
| 28 | hsa04976_Bile_secretion | 6 | 71 | 0.07951 | 0.5112 | |
| 29 | hsa04960_Aldosterone.regulated_sodium_reabsorption | 4 | 42 | 0.1005 | 0.624 | |
| 30 | hsa04114_Oocyte_meiosis | 8 | 114 | 0.1103 | 0.6514 | |
| 31 | hsa04070_Phosphatidylinositol_signaling_system | 6 | 78 | 0.1122 | 0.6514 | |
| 32 | hsa00603_Glycosphingolipid_biosynthesis_._globo_series | 2 | 14 | 0.1163 | 0.6541 | |
| 33 | hsa00604_Glycosphingolipid_biosynthesis_._ganglio_series | 2 | 15 | 0.1306 | 0.7122 | |
| 34 | hsa04330_Notch_signaling_pathway | 4 | 47 | 0.1365 | 0.7228 | |
| 35 | hsa04012_ErbB_signaling_pathway | 6 | 87 | 0.1626 | 0.836 | |
| 36 | hsa04720_Long.term_potentiation | 5 | 70 | 0.1741 | 0.8362 | |
| 37 | hsa00670_One_carbon_pool_by_folate | 2 | 18 | 0.1754 | 0.8362 | |
| 38 | hsa04062_Chemokine_signaling_pathway | 11 | 189 | 0.1783 | 0.8362 | |
| 39 | hsa04540_Gap_junction | 6 | 90 | 0.1812 | 0.8362 | |
| 40 | hsa04130_SNARE_interactions_in_vesicular_transport | 3 | 36 | 0.1941 | 0.8734 | |
| 41 | hsa04340_Hedgehog_signaling_pathway | 4 | 56 | 0.2117 | 0.9294 | |
| 42 | hsa01040_Biosynthesis_of_unsaturated_fatty_acids | 2 | 21 | 0.2223 | 0.9525 | |
| 43 | hsa04910_Insulin_signaling_pathway | 8 | 138 | 0.23 | 0.9627 | |
| 44 | hsa04146_Peroxisome | 5 | 79 | 0.2421 | 0.9904 | |
| 45 | hsa00564_Glycerophospholipid_metabolism | 5 | 80 | 0.2501 | 1 | |
| 46 | hsa04141_Protein_processing_in_endoplasmic_reticulum | 9 | 168 | 0.2817 | 1 | |
| 47 | hsa04962_Vasopressin.regulated_water_reabsorption | 3 | 44 | 0.2849 | 1 | |
| 48 | hsa04973_Carbohydrate_digestion_and_absorption | 3 | 44 | 0.2849 | 1 | |
| 49 | hsa04320_Dorso.ventral_axis_formation | 2 | 25 | 0.2858 | 1 | |
| 50 | hsa04914_Progesterone.mediated_oocyte_maturation | 5 | 87 | 0.3074 | 1 | |
| 51 | hsa00030_Pentose_phosphate_pathway | 2 | 27 | 0.3175 | 1 | |
| 52 | hsa04115_p53_signaling_pathway | 4 | 69 | 0.3341 | 1 | |
| 53 | hsa00512_Mucin_type_O.Glycan_biosynthesis | 2 | 30 | 0.3643 | 1 | |
| 54 | hsa04670_Leukocyte_transendothelial_migration | 6 | 117 | 0.3748 | 1 | |
| 55 | hsa04150_mTOR_signaling_pathway | 3 | 52 | 0.3784 | 1 | |
| 56 | hsa04742_Taste_transduction | 3 | 52 | 0.3784 | 1 | |
| 57 | hsa00260_Glycine._serine_and_threonine_metabolism | 2 | 32 | 0.3948 | 1 | |
| 58 | hsa04370_VEGF_signaling_pathway | 4 | 76 | 0.4019 | 1 | |
| 59 | hsa00240_Pyrimidine_metabolism | 5 | 99 | 0.4086 | 1 | |
| 60 | hsa04260_Cardiac_muscle_contraction | 4 | 77 | 0.4115 | 1 | |
| 61 | hsa04974_Protein_digestion_and_absorption | 4 | 81 | 0.4496 | 1 | |
| 62 | hsa04722_Neurotrophin_signaling_pathway | 6 | 127 | 0.4506 | 1 | |
| 63 | hsa00051_Fructose_and_mannose_metabolism | 2 | 36 | 0.4537 | 1 | |
| 64 | hsa00270_Cysteine_and_methionine_metabolism | 2 | 36 | 0.4537 | 1 | |
| 65 | hsa03015_mRNA_surveillance_pathway | 4 | 83 | 0.4683 | 1 | |
| 66 | hsa04630_Jak.STAT_signaling_pathway | 7 | 155 | 0.4846 | 1 | |
| 67 | hsa04640_Hematopoietic_cell_lineage | 4 | 88 | 0.5141 | 1 | |
| 68 | hsa04210_Apoptosis | 4 | 89 | 0.523 | 1 | |
| 69 | hsa00230_Purine_metabolism | 7 | 162 | 0.5314 | 1 | |
| 70 | hsa02010_ABC_transporters | 2 | 44 | 0.5608 | 1 | |
| 71 | hsa04610_Complement_and_coagulation_cascades | 3 | 69 | 0.5633 | 1 | |
| 72 | hsa03050_Proteasome | 2 | 45 | 0.5731 | 1 | |
| 73 | hsa04144_Endocytosis | 8 | 203 | 0.631 | 1 | |
| 74 | hsa04145_Phagosome | 6 | 156 | 0.651 | 1 | |
| 75 | hsa04664_Fc_epsilon_RI_signaling_pathway | 3 | 79 | 0.6551 | 1 | |
| 76 | hsa00330_Arginine_and_proline_metabolism | 2 | 54 | 0.6723 | 1 | |
| 77 | hsa00562_Inositol_phosphate_metabolism | 2 | 57 | 0.701 | 1 | |
| 78 | hsa00590_Arachidonic_acid_metabolism | 2 | 59 | 0.7189 | 1 | |
| 79 | hsa04621_NOD.like_receptor_signaling_pathway | 2 | 59 | 0.7189 | 1 | |
| 80 | hsa00830_Retinol_metabolism | 2 | 64 | 0.7598 | 1 | |
| 81 | hsa04920_Adipocytokine_signaling_pathway | 2 | 68 | 0.7887 | 1 | |
| 82 | hsa04380_Osteoclast_differentiation | 4 | 128 | 0.7954 | 1 | |
| 83 | hsa04662_B_cell_receptor_signaling_pathway | 2 | 75 | 0.8319 | 1 | |
| 84 | hsa04612_Antigen_processing_and_presentation | 2 | 78 | 0.8479 | 1 | |
| 85 | hsa03013_RNA_transport | 3 | 152 | 0.9583 | 1 | |
| 86 | hsa04142_Lysosome | 2 | 121 | 0.9664 | 1 | |
| 87 | hsa04110_Cell_cycle | 2 | 128 | 0.974 | 1 | |
| 88 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 2 | 136 | 0.9807 | 1 | |
| 89 | hsa04120_Ubiquitin_mediated_proteolysis | 2 | 139 | 0.9827 | 1 | |
| 90 | hsa04740_Olfactory_transduction | 4 | 388 | 1 | 1 |