This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-106a-5p | TGFBR2 | 1.39 | 6.0E-5 | -1.92 | 0 | miRTarBase; miRNATAP | -0.31 | 0 | 22912877 | MiR-106a inhibits the expression of transforming growth factor-β receptor 2 TGFBR2 leading to increased CRC cell migration and invasion |
| 2 | hsa-miR-106b-5p | TGFBR2 | 1.47 | 0 | -1.92 | 0 | miRNATAP | -0.49 | 0 | NA | |
| 3 | hsa-miR-107 | TGFBR2 | 0.66 | 0 | -1.92 | 0 | miRanda; miRNATAP | -0.25 | 0.00092 | NA | |
| 4 | hsa-miR-130a-3p | TGFBR2 | 0.88 | 0.00016 | -1.92 | 0 | miRNATAP | -0.13 | 0.00147 | NA | |
| 5 | hsa-miR-130b-3p | TGFBR2 | 1.83 | 0 | -1.92 | 0 | miRNAWalker2 validate; miRNATAP | -0.42 | 0 | 25024357 | Follow-up experiments showed two miRNAs miR-9-5p and miR-130b-3p in this module had increased expression while their target gene TGFBR2 had decreased expression in a cohort of human NSCLC |
| 6 | hsa-miR-142-3p | TGFBR2 | 3.98 | 0 | -1.92 | 0 | miRanda | -0.19 | 0 | NA | |
| 7 | hsa-miR-17-5p | TGFBR2 | 2.07 | 0 | -1.92 | 0 | miRNAWalker2 validate; miRTarBase; TargetScan; miRNATAP | -0.46 | 0 | 25011053; 27120811 | We demonstrate that miR-17 overexpression interferes with the TGFβ-EMT axis and hinders RCC sphere formation; and validated TGFBR2 as a direct and biologically relevant target during this process;MiR-17-5p was found to bind to the 3'UTR of TGFBR2 mRNA and further validation of this specific binding was performed through a reporter assay; An inverse correlation between miR-17-5p and TGFBR2 protein was observed in gastric cancer tissues; Cell studies revealed that miR-17-5p negatively regulated TGFBR2 expression by directly binding to the 3'UTR of TGFBR2 mRNA thereby promoting cell growth and migration; The results of our study suggest a novel regulatory network in gastric cancer mediated by miR-17-5p and TGFBR2 and may indicate that TGFBR2 could serve as a new therapeutic target in gastric cancer |
| 8 | hsa-miR-186-5p | TGFBR2 | 0.85 | 0 | -1.92 | 0 | miRNATAP | -0.38 | 0 | NA | |
| 9 | hsa-miR-18a-5p | TGFBR2 | 1.37 | 1.0E-5 | -1.92 | 0 | miRNAWalker2 validate | -0.39 | 0 | NA | |
| 10 | hsa-miR-19a-3p | TGFBR2 | 2.12 | 0 | -1.92 | 0 | miRNAWalker2 validate; miRNATAP | -0.32 | 0 | NA | |
| 11 | hsa-miR-19b-3p | TGFBR2 | 2.11 | 0 | -1.92 | 0 | miRNAWalker2 validate; miRNATAP | -0.33 | 0 | NA | |
| 12 | hsa-miR-20a-5p | TGFBR2 | 2.65 | 0 | -1.92 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.38 | 0 | NA | |
| 13 | hsa-miR-20b-5p | TGFBR2 | 1.36 | 0.00261 | -1.92 | 0 | miRNATAP | -0.19 | 0 | NA | |
| 14 | hsa-miR-21-5p | TGFBR2 | 4.38 | 0 | -1.92 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.29 | 0 | 24037531 | Androgen receptor and microRNA 21 axis downregulates transforming growth factor beta receptor II TGFBR2 expression in prostate cancer; Our results revealed that miR-21 suppresses TGFBR2 levels by binding to its 3'-UTR and AR signaling further potentiates this effect in both untransformed and transformed human prostate epithelial cells as well as in human prostate cancers; Manipulation of androgen signaling or the expression levels of AR or miR-21 negatively altered TGFBR2 expression in untransformed and transformed human prostate epithelial cells human prostate cancer xenografts and mouse prostate glands; Together these results suggest that the AR and miR-21 axis exerts its oncogenic effects in prostate tumors by downregulating TGFBR2 hence inhibiting the tumor-suppressive activity of TGFβ pathway |
| 15 | hsa-miR-28-5p | TGFBR2 | 1.2 | 0 | -1.92 | 0 | miRanda | -0.25 | 0.00025 | NA | |
| 16 | hsa-miR-301a-3p | TGFBR2 | 2.7 | 0 | -1.92 | 0 | miRNATAP | -0.31 | 0 | 25551793 | MicroRNA 301a promotes migration and invasion by targeting TGFBR2 in human colorectal cancer; TGFBR2 was identified to be the downstream target of miR-301a; Knockdown of TGFBR2 in cells treated by miR-301a inhibitor elevated the previously abrogated migration and invasion; Our data indicated that miR-301a correlated with the metastatic and invasive ability in human colorectal cancers and miR-301a exerted its role as oncogene by targeting TGFBR2 |
| 17 | hsa-miR-320b | TGFBR2 | 0.23 | 0.37882 | -1.92 | 0 | miRanda; miRNATAP | -0.17 | 1.0E-5 | NA | |
| 18 | hsa-miR-335-3p | TGFBR2 | 1.51 | 0 | -1.92 | 0 | mirMAP | -0.31 | 0 | NA | |
| 19 | hsa-miR-429 | TGFBR2 | 2.38 | 0 | -1.92 | 0 | miRNATAP | -0.18 | 0 | NA | |
| 20 | hsa-miR-454-3p | TGFBR2 | 1.49 | 0 | -1.92 | 0 | miRNATAP | -0.39 | 0 | NA | |
| 21 | hsa-miR-590-3p | TGFBR2 | 0.84 | 0.00129 | -1.92 | 0 | miRanda | -0.3 | 0 | NA | |
| 22 | hsa-miR-590-5p | TGFBR2 | 2.07 | 0 | -1.92 | 0 | miRNAWalker2 validate; miRTarBase; PITA; miRanda; miRNATAP | -0.43 | 0 | NA | |
| 23 | hsa-miR-9-5p | TGFBR2 | 4.99 | 0 | -1.92 | 0 | miRNATAP | -0.17 | 0 | 25024357 | Follow-up experiments showed two miRNAs miR-9-5p and miR-130b-3p in this module had increased expression while their target gene TGFBR2 had decreased expression in a cohort of human NSCLC |
| 24 | hsa-miR-92a-3p | TGFBR2 | -0.14 | 0.49341 | -1.92 | 0 | miRNAWalker2 validate | -0.24 | 0 | NA | |
| 25 | hsa-miR-93-5p | TGFBR2 | 1.51 | 0 | -1.92 | 0 | miRNAWalker2 validate; miRNATAP | -0.47 | 0 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CELL CYCLE | 38 | 1316 | 5.116e-18 | 1.19e-14 |
| 2 | REGULATION OF CELL DEATH | 40 | 1472 | 4.239e-18 | 1.19e-14 |
| 3 | REGULATION OF CELL CYCLE | 32 | 949 | 5.397e-17 | 8.371e-14 |
| 4 | DNA METABOLIC PROCESS | 29 | 758 | 7.886e-17 | 9.173e-14 |
| 5 | CELLULAR RESPONSE TO STRESS | 37 | 1565 | 9.71e-15 | 9.036e-12 |
| 6 | NEGATIVE REGULATION OF CELL CYCLE | 21 | 433 | 2.654e-14 | 2.058e-11 |
| 7 | CELL CYCLE PHASE TRANSITION | 17 | 255 | 5.709e-14 | 3.795e-11 |
| 8 | CELL CYCLE PROCESS | 30 | 1081 | 1.092e-13 | 6.354e-11 |
| 9 | CELLULAR RESPONSE TO DNA DAMAGE STIMULUS | 25 | 720 | 1.355e-13 | 7.003e-11 |
| 10 | CELL CYCLE CHECKPOINT | 15 | 194 | 2.21e-13 | 1.028e-10 |
| 11 | CELL DEATH | 28 | 1001 | 7.185e-13 | 3.039e-10 |
| 12 | MITOTIC CELL CYCLE | 24 | 766 | 4.013e-12 | 1.53e-09 |
| 13 | REGULATION OF PROTEIN MODIFICATION PROCESS | 35 | 1710 | 4.276e-12 | 1.53e-09 |
| 14 | REGULATION OF DNA METABOLIC PROCESS | 17 | 340 | 5.955e-12 | 1.847e-09 |
| 15 | SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 12 | 127 | 5.871e-12 | 1.847e-09 |
| 16 | REGULATION OF TRANSFERASE ACTIVITY | 26 | 946 | 8.328e-12 | 2.422e-09 |
| 17 | NEGATIVE REGULATION OF CELL CYCLE PROCESS | 14 | 214 | 1.419e-11 | 3.884e-09 |
| 18 | DNA REPAIR | 19 | 480 | 1.788e-11 | 4.379e-09 |
| 19 | MITOTIC CELL CYCLE CHECKPOINT | 12 | 139 | 1.722e-11 | 4.379e-09 |
| 20 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 35 | 1805 | 1.973e-11 | 4.525e-09 |
| 21 | REGULATION OF CELL CYCLE ARREST | 11 | 108 | 2.042e-11 | 4.525e-09 |
| 22 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 33 | 1618 | 2.357e-11 | 4.985e-09 |
| 23 | REGULATION OF CELL CYCLE PROCESS | 20 | 558 | 2.95e-11 | 5.967e-09 |
| 24 | POSITIVE REGULATION OF DNA METABOLIC PROCESS | 13 | 185 | 3.238e-11 | 6.277e-09 |
| 25 | NEGATIVE REGULATION OF MITOTIC CELL CYCLE | 13 | 199 | 8.085e-11 | 1.505e-08 |
| 26 | SIGNAL TRANSDUCTION IN RESPONSE TO DNA DAMAGE | 10 | 96 | 1.39e-10 | 2.487e-08 |
| 27 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 24 | 917 | 1.663e-10 | 2.764e-08 |
| 28 | RESPONSE TO ALCOHOL | 16 | 362 | 1.635e-10 | 2.764e-08 |
| 29 | RESPONSE TO ABIOTIC STIMULUS | 25 | 1024 | 2.776e-10 | 4.454e-08 |
| 30 | CELL CYCLE G2 M PHASE TRANSITION | 11 | 138 | 2.966e-10 | 4.601e-08 |
| 31 | MISMATCH REPAIR | 7 | 31 | 3.155e-10 | 4.736e-08 |
| 32 | NEGATIVE REGULATION OF CELL CYCLE PHASE TRANSITION | 11 | 146 | 5.438e-10 | 7.668e-08 |
| 33 | DNA INTEGRITY CHECKPOINT | 11 | 146 | 5.438e-10 | 7.668e-08 |
| 34 | CELL CYCLE G1 S PHASE TRANSITION | 10 | 111 | 5.926e-10 | 7.878e-08 |
| 35 | G1 S TRANSITION OF MITOTIC CELL CYCLE | 10 | 111 | 5.926e-10 | 7.878e-08 |
| 36 | POSITIVE REGULATION OF GENE EXPRESSION | 32 | 1733 | 6.552e-10 | 8.468e-08 |
| 37 | RESPONSE TO ENDOGENOUS STIMULUS | 29 | 1450 | 8.433e-10 | 1.061e-07 |
| 38 | DNA BIOSYNTHETIC PROCESS | 10 | 121 | 1.389e-09 | 1.701e-07 |
| 39 | REGULATION OF KINASE ACTIVITY | 21 | 776 | 1.527e-09 | 1.796e-07 |
| 40 | REGENERATION | 11 | 161 | 1.544e-09 | 1.796e-07 |
| 41 | RESPONSE TO STEROID HORMONE | 17 | 497 | 2.122e-09 | 2.408e-07 |
| 42 | REGULATION OF CELL CYCLE PHASE TRANSITION | 14 | 321 | 2.943e-09 | 3.26e-07 |
| 43 | MITOTIC DNA INTEGRITY CHECKPOINT | 9 | 100 | 4.49e-09 | 4.858e-07 |
| 44 | CELL DIVISION | 16 | 460 | 5.208e-09 | 5.507e-07 |
| 45 | CHROMOSOME ORGANIZATION | 23 | 1009 | 6.015e-09 | 6.155e-07 |
| 46 | POSITIVE REGULATION OF CELL DEATH | 18 | 605 | 6.085e-09 | 6.155e-07 |
| 47 | REGULATION OF MITOTIC CELL CYCLE | 16 | 468 | 6.647e-09 | 6.581e-07 |
| 48 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 16 | 470 | 7.06e-09 | 6.844e-07 |
| 49 | APOPTOTIC SIGNALING PATHWAY | 13 | 289 | 7.762e-09 | 7.203e-07 |
| 50 | REGULATION OF CELLULAR RESPONSE TO STRESS | 19 | 691 | 7.895e-09 | 7.203e-07 |
| 51 | REGULATION OF CELL PROLIFERATION | 28 | 1496 | 7.691e-09 | 7.203e-07 |
| 52 | RESPONSE TO INORGANIC SUBSTANCE | 16 | 479 | 9.225e-09 | 8.255e-07 |
| 53 | ORGAN REGENERATION | 8 | 83 | 1.947e-08 | 1.709e-06 |
| 54 | DNA REPLICATION | 11 | 208 | 2.255e-08 | 1.943e-06 |
| 55 | POSITIVE REGULATION OF CELL CYCLE ARREST | 8 | 85 | 2.354e-08 | 1.991e-06 |
| 56 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 23 | 1087 | 2.408e-08 | 2.001e-06 |
| 57 | RESPONSE TO METAL ION | 13 | 333 | 4.152e-08 | 3.389e-06 |
| 58 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 28 | 1656 | 6.816e-08 | 5.468e-06 |
| 59 | NEGATIVE REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 8 | 98 | 7.252e-08 | 5.719e-06 |
| 60 | REGULATION OF CATABOLIC PROCESS | 18 | 731 | 1.085e-07 | 8.412e-06 |
| 61 | RESPONSE TO ESTRADIOL | 9 | 146 | 1.234e-07 | 9.411e-06 |
| 62 | POSITIVE REGULATION OF CELL CYCLE PROCESS | 11 | 247 | 1.302e-07 | 9.774e-06 |
| 63 | G1 DNA DAMAGE CHECKPOINT | 7 | 73 | 1.615e-07 | 1.193e-05 |
| 64 | TISSUE DEVELOPMENT | 26 | 1518 | 1.755e-07 | 1.276e-05 |
| 65 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 29 | 1848 | 1.858e-07 | 1.33e-05 |
| 66 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 6 | 46 | 1.994e-07 | 1.406e-05 |
| 67 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 30 | 1977 | 2.228e-07 | 1.547e-05 |
| 68 | REGULATION OF DNA REPLICATION | 9 | 161 | 2.844e-07 | 1.946e-05 |
| 69 | REGULATION OF SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 9 | 162 | 2.997e-07 | 2.021e-05 |
| 70 | NEGATIVE REGULATION OF CELL DEATH | 19 | 872 | 3.061e-07 | 2.035e-05 |
| 71 | POSITIVE REGULATION OF CELL CYCLE | 12 | 332 | 3.256e-07 | 2.134e-05 |
| 72 | REGULATION OF TRANSCRIPTION INVOLVED IN G1 S TRANSITION OF MITOTIC CELL CYCLE | 5 | 27 | 3.522e-07 | 2.276e-05 |
| 73 | RESPONSE TO ESTROGEN | 10 | 218 | 3.805e-07 | 2.423e-05 |
| 74 | RESPONSE TO GROWTH FACTOR | 14 | 475 | 3.853e-07 | 2.423e-05 |
| 75 | RESPONSE TO LIPID | 19 | 888 | 4.038e-07 | 2.505e-05 |
| 76 | RESPONSE TO HORMONE | 19 | 893 | 4.397e-07 | 2.68e-05 |
| 77 | REGULATION OF ORGANELLE ORGANIZATION | 22 | 1178 | 4.435e-07 | 2.68e-05 |
| 78 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 25 | 1492 | 4.824e-07 | 2.878e-05 |
| 79 | RESPONSE TO RADIATION | 13 | 413 | 4.947e-07 | 2.913e-05 |
| 80 | POSTREPLICATION REPAIR | 6 | 54 | 5.303e-07 | 3.085e-05 |
| 81 | NEGATIVE REGULATION OF CELL COMMUNICATION | 22 | 1192 | 5.415e-07 | 3.111e-05 |
| 82 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 15 | 573 | 6.458e-07 | 3.664e-05 |
| 83 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 25 | 1517 | 6.568e-07 | 3.682e-05 |
| 84 | RESPONSE TO DRUG | 13 | 431 | 7.993e-07 | 4.427e-05 |
| 85 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 4 | 14 | 8.618e-07 | 4.717e-05 |
| 86 | NUCLEUS ORGANIZATION | 8 | 136 | 9.112e-07 | 4.93e-05 |
| 87 | CELL PROLIFERATION | 16 | 672 | 9.276e-07 | 4.961e-05 |
| 88 | RESPONSE TO TOXIC SUBSTANCE | 10 | 241 | 9.521e-07 | 5.034e-05 |
| 89 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 7 | 95 | 9.907e-07 | 5.179e-05 |
| 90 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 27 | 1784 | 1.122e-06 | 5.8e-05 |
| 91 | TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 9 | 190 | 1.145e-06 | 5.852e-05 |
| 92 | SOMATIC DIVERSIFICATION OF IMMUNE RECEPTORS VIA SOMATIC MUTATION | 4 | 15 | 1.17e-06 | 5.918e-05 |
| 93 | NEURON APOPTOTIC PROCESS | 5 | 35 | 1.367e-06 | 6.84e-05 |
| 94 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 15 | 616 | 1.591e-06 | 7.873e-05 |
| 95 | REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 8 | 147 | 1.64e-06 | 8.033e-05 |
| 96 | NEGATIVE REGULATION OF GENE EXPRESSION | 24 | 1493 | 1.792e-06 | 8.685e-05 |
| 97 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 20 | 1079 | 1.81e-06 | 8.685e-05 |
| 98 | POSITIVE REGULATION OF NEURON DEATH | 6 | 67 | 1.933e-06 | 9.177e-05 |
| 99 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 8 | 153 | 2.215e-06 | 0.0001041 |
| 100 | NUCLEOTIDE EXCISION REPAIR DNA INCISION | 5 | 39 | 2.382e-06 | 0.0001108 |
| 101 | POSITIVE REGULATION OF CELL PROLIFERATION | 17 | 814 | 2.461e-06 | 0.0001134 |
| 102 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 19 | 1004 | 2.518e-06 | 0.0001149 |
| 103 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 19 | 1008 | 2.669e-06 | 0.0001206 |
| 104 | TRANSCRIPTION COUPLED NUCLEOTIDE EXCISION REPAIR | 6 | 73 | 3.209e-06 | 0.000142 |
| 105 | ERROR FREE TRANSLESION SYNTHESIS | 4 | 19 | 3.266e-06 | 0.000142 |
| 106 | DNA RECOMBINATION | 9 | 215 | 3.175e-06 | 0.000142 |
| 107 | ERROR PRONE TRANSLESION SYNTHESIS | 4 | 19 | 3.266e-06 | 0.000142 |
| 108 | PHOSPHORYLATION | 21 | 1228 | 3.44e-06 | 0.0001482 |
| 109 | DNA SYNTHESIS INVOLVED IN DNA REPAIR | 6 | 74 | 3.477e-06 | 0.0001484 |
| 110 | RESPONSE TO LIGHT STIMULUS | 10 | 280 | 3.661e-06 | 0.0001549 |
| 111 | REGULATION OF MAPK CASCADE | 15 | 660 | 3.701e-06 | 0.0001552 |
| 112 | ORGANELLE FISSION | 13 | 496 | 3.769e-06 | 0.0001566 |
| 113 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 25 | 1672 | 3.837e-06 | 0.000158 |
| 114 | EPITHELIUM DEVELOPMENT | 18 | 945 | 4.382e-06 | 0.0001775 |
| 115 | INTRACELLULAR SIGNAL TRANSDUCTION | 24 | 1572 | 4.388e-06 | 0.0001775 |
| 116 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 20 | 1152 | 4.876e-06 | 0.0001956 |
| 117 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 27 | 1929 | 4.969e-06 | 0.0001976 |
| 118 | RESPONSE TO NITROGEN COMPOUND | 17 | 859 | 5.046e-06 | 0.000199 |
| 119 | GENERATION OF PRECURSOR METABOLITES AND ENERGY | 10 | 292 | 5.307e-06 | 0.0002075 |
| 120 | MITOTIC NUCLEAR DIVISION | 11 | 361 | 5.379e-06 | 0.0002086 |
| 121 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 11 | 363 | 5.668e-06 | 0.000218 |
| 122 | MEMBRANE DISASSEMBLY | 5 | 47 | 6.126e-06 | 0.0002245 |
| 123 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 22 | 1381 | 6.078e-06 | 0.0002245 |
| 124 | NUCLEAR ENVELOPE ORGANIZATION | 6 | 81 | 5.902e-06 | 0.0002245 |
| 125 | NUCLEAR ENVELOPE DISASSEMBLY | 5 | 47 | 6.126e-06 | 0.0002245 |
| 126 | NEURON DEATH | 5 | 47 | 6.126e-06 | 0.0002245 |
| 127 | POSITIVE REGULATION OF NEURON APOPTOTIC PROCESS | 5 | 47 | 6.126e-06 | 0.0002245 |
| 128 | RESPONSE TO UV | 7 | 126 | 6.574e-06 | 0.000239 |
| 129 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 14 | 616 | 8.02e-06 | 0.0002893 |
| 130 | POSITIVE REGULATION OF DNA REPLICATION | 6 | 86 | 8.36e-06 | 0.0002992 |
| 131 | NUCLEOTIDE EXCISION REPAIR DNA GAP FILLING | 4 | 24 | 8.765e-06 | 0.000309 |
| 132 | HEAD DEVELOPMENT | 15 | 709 | 8.751e-06 | 0.000309 |
| 133 | REGULATION OF PROTEOLYSIS | 15 | 711 | 9.049e-06 | 0.0003166 |
| 134 | RESPONSE TO OXYGEN LEVELS | 10 | 311 | 9.22e-06 | 0.0003202 |
| 135 | REGULATION OF GENERATION OF PRECURSOR METABOLITES AND ENERGY | 6 | 89 | 1.02e-05 | 0.0003514 |
| 136 | EPITHELIAL CELL APOPTOTIC PROCESS | 4 | 25 | 1.039e-05 | 0.0003555 |
| 137 | HEART DEVELOPMENT | 12 | 466 | 1.088e-05 | 0.0003696 |
| 138 | RESPONSE TO REACTIVE OXYGEN SPECIES | 8 | 191 | 1.14e-05 | 0.0003843 |
| 139 | VENTRICULAR SYSTEM DEVELOPMENT | 4 | 26 | 1.223e-05 | 0.0004093 |
| 140 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 19 | 1135 | 1.457e-05 | 0.0004841 |
| 141 | CELLULAR RESPONSE TO OXYGEN LEVELS | 7 | 143 | 1.507e-05 | 0.0004971 |
| 142 | RESPONSE TO TRANSFORMING GROWTH FACTOR BETA | 7 | 144 | 1.576e-05 | 0.0005165 |
| 143 | REGULATION OF RESPONSE TO STRESS | 22 | 1468 | 1.591e-05 | 0.0005178 |
| 144 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 21 | 1360 | 1.645e-05 | 0.0005316 |
| 145 | AGING | 9 | 264 | 1.66e-05 | 0.0005328 |
| 146 | REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 6 | 97 | 1.672e-05 | 0.000533 |
| 147 | PROTEIN PHOSPHORYLATION | 17 | 944 | 1.726e-05 | 0.0005464 |
| 148 | REGULATION OF CELL CYCLE G2 M PHASE TRANSITION | 5 | 59 | 1.896e-05 | 0.0005962 |
| 149 | SOMATIC DIVERSIFICATION OF IMMUNOGLOBULINS | 4 | 29 | 1.918e-05 | 0.0005988 |
| 150 | POSITIVE REGULATION OF CHROMOSOME ORGANIZATION | 7 | 150 | 2.054e-05 | 0.000637 |
| 151 | NEGATIVE REGULATION OF INTRACELLULAR STEROID HORMONE RECEPTOR SIGNALING PATHWAY | 4 | 30 | 2.203e-05 | 0.0006789 |
| 152 | INTRINSIC APOPTOTIC SIGNALING PATHWAY | 7 | 152 | 2.237e-05 | 0.0006848 |
| 153 | NEGATIVE REGULATION OF PHOSPHORYLATION | 11 | 422 | 2.313e-05 | 0.0007033 |
| 154 | REGULATION OF MUSCLE TISSUE DEVELOPMENT | 6 | 103 | 2.356e-05 | 0.0007072 |
| 155 | REGULATION OF MUSCLE ORGAN DEVELOPMENT | 6 | 103 | 2.356e-05 | 0.0007072 |
| 156 | MACROMOLECULAR COMPLEX ASSEMBLY | 21 | 1398 | 2.481e-05 | 0.0007316 |
| 157 | REGULATION OF CHROMOSOME ORGANIZATION | 9 | 278 | 2.496e-05 | 0.0007316 |
| 158 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 16 | 872 | 2.5e-05 | 0.0007316 |
| 159 | CELLULAR RESPONSE TO REACTIVE OXYGEN SPECIES | 6 | 104 | 2.489e-05 | 0.0007316 |
| 160 | REGULATION OF PROTEIN MODIFICATION BY SMALL PROTEIN CONJUGATION OR REMOVAL | 9 | 280 | 2.641e-05 | 0.000768 |
| 161 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 22 | 1518 | 2.667e-05 | 0.0007695 |
| 162 | RESPONSE TO OXIDATIVE STRESS | 10 | 352 | 2.679e-05 | 0.0007695 |
| 163 | CELLULAR RESPONSE TO STEROID HORMONE STIMULUS | 8 | 218 | 2.956e-05 | 0.0008437 |
| 164 | FOREBRAIN DEVELOPMENT | 10 | 357 | 3.02e-05 | 0.0008569 |
| 165 | G2 DNA DAMAGE CHECKPOINT | 4 | 33 | 3.248e-05 | 0.0009159 |
| 166 | MICROTUBULE BASED PROCESS | 12 | 522 | 3.332e-05 | 0.000934 |
| 167 | EMBRYO DEVELOPMENT | 16 | 894 | 3.378e-05 | 0.0009412 |
| 168 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 15 | 801 | 3.636e-05 | 0.001007 |
| 169 | NUCLEOTIDE EXCISION REPAIR | 6 | 113 | 3.98e-05 | 0.001096 |
| 170 | TELENCEPHALON DEVELOPMENT | 8 | 228 | 4.065e-05 | 0.001113 |
| 171 | REGULATION OF GLYCOGEN METABOLIC PROCESS | 4 | 35 | 4.12e-05 | 0.001121 |
| 172 | RHYTHMIC PROCESS | 9 | 298 | 4.293e-05 | 0.001161 |
| 173 | REGULATION OF HISTONE PHOSPHORYLATION | 3 | 13 | 4.57e-05 | 0.001229 |
| 174 | DNA DAMAGE RESPONSE DETECTION OF DNA DAMAGE | 4 | 36 | 4.616e-05 | 0.001234 |
| 175 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 12 | 541 | 4.709e-05 | 0.001245 |
| 176 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 12 | 541 | 4.709e-05 | 0.001245 |
| 177 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 17 | 1036 | 5.575e-05 | 0.001451 |
| 178 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 11 | 465 | 5.583e-05 | 0.001451 |
| 179 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 17 | 1036 | 5.575e-05 | 0.001451 |
| 180 | RESPONSE TO CORTICOSTEROID | 7 | 176 | 5.708e-05 | 0.001469 |
| 181 | TUBE DEVELOPMENT | 12 | 552 | 5.713e-05 | 0.001469 |
| 182 | RESPONSE TO KETONE | 7 | 182 | 7.052e-05 | 0.001803 |
| 183 | CELLULAR RESPONSE TO NITRIC OXIDE | 3 | 15 | 7.211e-05 | 0.001834 |
| 184 | REGULATION OF MAP KINASE ACTIVITY | 9 | 319 | 7.254e-05 | 0.001834 |
| 185 | TRANSLESION SYNTHESIS | 4 | 41 | 7.768e-05 | 0.001954 |
| 186 | REGULATION OF NEURON DEATH | 8 | 252 | 8.211e-05 | 0.002054 |
| 187 | SOMATIC DIVERSIFICATION OF IMMUNE RECEPTORS | 4 | 42 | 8.549e-05 | 0.002116 |
| 188 | REGULATION OF HEART GROWTH | 4 | 42 | 8.549e-05 | 0.002116 |
| 189 | SOMATIC RECOMBINATION OF IMMUNOGLOBULIN GENES INVOLVED IN IMMUNE RESPONSE | 3 | 16 | 8.839e-05 | 0.00212 |
| 190 | POSITIVE REGULATION OF HISTONE H3 K4 METHYLATION | 3 | 16 | 8.839e-05 | 0.00212 |
| 191 | NUCLEIC ACID PHOSPHODIESTER BOND HYDROLYSIS | 8 | 254 | 8.675e-05 | 0.00212 |
| 192 | REGULATION OF FIBROBLAST PROLIFERATION | 5 | 81 | 8.81e-05 | 0.00212 |
| 193 | ISOTYPE SWITCHING | 3 | 16 | 8.839e-05 | 0.00212 |
| 194 | SOMATIC DIVERSIFICATION OF IMMUNOGLOBULINS INVOLVED IN IMMUNE RESPONSE | 3 | 16 | 8.839e-05 | 0.00212 |
| 195 | REGULATION OF POLYSACCHARIDE METABOLIC PROCESS | 4 | 43 | 9.386e-05 | 0.00224 |
| 196 | GROWTH | 10 | 410 | 9.603e-05 | 0.00228 |
| 197 | REGULATION OF NEURON APOPTOTIC PROCESS | 7 | 192 | 9.86e-05 | 0.002329 |
| 198 | CELL DEVELOPMENT | 20 | 1426 | 0.0001035 | 0.002431 |
| 199 | NEGATIVE REGULATION OF CELL AGING | 3 | 17 | 0.0001069 | 0.0025 |
| 200 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 23 | 1791 | 0.0001105 | 0.002565 |
| 201 | POSITIVE REGULATION OF CHROMATIN MODIFICATION | 5 | 85 | 0.0001108 | 0.002565 |
| 202 | RESPONSE TO ETHANOL | 6 | 136 | 0.0001117 | 0.002572 |
| 203 | PROTEIN LOCALIZATION TO CHROMOSOME | 4 | 45 | 0.0001124 | 0.002575 |
| 204 | REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 7 | 197 | 0.0001157 | 0.002635 |
| 205 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 13 | 689 | 0.0001161 | 0.002635 |
| 206 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 14 | 788 | 0.0001179 | 0.00265 |
| 207 | CIRCULATORY SYSTEM DEVELOPMENT | 14 | 788 | 0.0001179 | 0.00265 |
| 208 | REGULATION OF HYDROLASE ACTIVITY | 19 | 1327 | 0.0001208 | 0.002702 |
| 209 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 7 | 200 | 0.0001271 | 0.00283 |
| 210 | MICROTUBULE CYTOSKELETON ORGANIZATION | 9 | 348 | 0.0001403 | 0.003109 |
| 211 | REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 4 | 48 | 0.0001449 | 0.003164 |
| 212 | IMMUNOGLOBULIN PRODUCTION | 4 | 48 | 0.0001449 | 0.003164 |
| 213 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 12 | 609 | 0.0001444 | 0.003164 |
| 214 | NEGATIVE REGULATION OF TRANSFERASE ACTIVITY | 9 | 351 | 0.0001496 | 0.003253 |
| 215 | CELLULAR RESPONSE TO REACTIVE NITROGEN SPECIES | 3 | 19 | 0.0001511 | 0.00327 |
| 216 | RESPONSE TO IONIZING RADIATION | 6 | 145 | 0.0001587 | 0.003418 |
| 217 | REGULATION OF CELLULAR AMIDE METABOLIC PROCESS | 9 | 354 | 0.0001595 | 0.003419 |
| 218 | PROTEIN COMPLEX BIOGENESIS | 17 | 1132 | 0.0001637 | 0.003477 |
| 219 | PROTEIN COMPLEX ASSEMBLY | 17 | 1132 | 0.0001637 | 0.003477 |
| 220 | PEPTIDYL SERINE MODIFICATION | 6 | 148 | 0.0001774 | 0.003735 |
| 221 | RESPONSE TO TRANSITION METAL NANOPARTICLE | 6 | 148 | 0.0001774 | 0.003735 |
| 222 | POSITIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 9 | 360 | 0.0001808 | 0.003789 |
| 223 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 13 | 724 | 0.0001891 | 0.003946 |
| 224 | POSTTRANSCRIPTIONAL REGULATION OF GENE EXPRESSION | 10 | 448 | 0.0001975 | 0.004102 |
| 225 | NEGATIVE REGULATION OF CATALYTIC ACTIVITY | 14 | 829 | 0.0001996 | 0.004128 |
| 226 | REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 6 | 152 | 0.000205 | 0.00418 |
| 227 | REGULATION OF CHROMATIN ORGANIZATION | 6 | 152 | 0.000205 | 0.00418 |
| 228 | SOMATIC RECOMBINATION OF IMMUNOGLOBULIN GENE SEGMENTS | 3 | 21 | 0.0002057 | 0.00418 |
| 229 | RESPONSE TO NITRIC OXIDE | 3 | 21 | 0.0002057 | 0.00418 |
| 230 | ENERGY DERIVATION BY OXIDATION OF ORGANIC COMPOUNDS | 7 | 217 | 0.0002101 | 0.00425 |
| 231 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 4 | 53 | 0.0002134 | 0.004299 |
| 232 | REGULATION OF CELL DEVELOPMENT | 14 | 836 | 0.0002176 | 0.004364 |
| 233 | DNA DEPENDENT DNA REPLICATION | 5 | 99 | 0.0002272 | 0.004518 |
| 234 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 5 | 99 | 0.0002272 | 0.004518 |
| 235 | PEPTIDYL AMINO ACID MODIFICATION | 14 | 841 | 0.0002313 | 0.00456 |
| 236 | ORGAN MORPHOGENESIS | 14 | 841 | 0.0002313 | 0.00456 |
| 237 | CELLULAR RESPONSE TO INORGANIC SUBSTANCE | 6 | 156 | 0.000236 | 0.004633 |
| 238 | LIMBIC SYSTEM DEVELOPMENT | 5 | 100 | 0.0002381 | 0.004656 |
| 239 | CELLULAR RESPONSE TO HORMONE STIMULUS | 11 | 552 | 0.0002513 | 0.004892 |
| 240 | HORMONE MEDIATED SIGNALING PATHWAY | 6 | 158 | 0.0002528 | 0.004901 |
| 241 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 11 | 554 | 0.0002592 | 0.005004 |
| 242 | REGULATION OF JNK CASCADE | 6 | 159 | 0.0002615 | 0.005028 |
| 243 | POSITIVE REGULATION OF CELL COMMUNICATION | 20 | 1532 | 0.00027 | 0.005169 |
| 244 | REPRODUCTION | 18 | 1297 | 0.0002735 | 0.005215 |
| 245 | APOPTOTIC MITOCHONDRIAL CHANGES | 4 | 57 | 0.000283 | 0.005375 |
| 246 | REGULATION OF NUCLEAR DIVISION | 6 | 163 | 0.0002989 | 0.005653 |
| 247 | CELLULAR GLUCAN METABOLIC PROCESS | 4 | 58 | 0.0003027 | 0.005679 |
| 248 | GLUCAN METABOLIC PROCESS | 4 | 58 | 0.0003027 | 0.005679 |
| 249 | IMMUNOGLOBULIN PRODUCTION INVOLVED IN IMMUNOGLOBULIN MEDIATED IMMUNE RESPONSE | 3 | 24 | 0.0003093 | 0.005711 |
| 250 | INTERSPECIES INTERACTION BETWEEN ORGANISMS | 12 | 662 | 0.0003105 | 0.005711 |
| 251 | SYMBIOSIS ENCOMPASSING MUTUALISM THROUGH PARASITISM | 12 | 662 | 0.0003105 | 0.005711 |
| 252 | NEGATIVE REGULATION OF ORGANELLE ORGANIZATION | 9 | 387 | 0.0003084 | 0.005711 |
| 253 | REGULATION OF PROTEIN PHOSPHATASE TYPE 2A ACTIVITY | 3 | 24 | 0.0003093 | 0.005711 |
| 254 | REGULATION OF INTRACELLULAR STEROID HORMONE RECEPTOR SIGNALING PATHWAY | 4 | 59 | 0.0003233 | 0.005923 |
| 255 | REGULATION OF WNT SIGNALING PATHWAY | 8 | 310 | 0.0003375 | 0.006159 |
| 256 | REGULATION OF PROTEIN BINDING | 6 | 168 | 0.0003513 | 0.006288 |
| 257 | SPINDLE CHECKPOINT | 3 | 25 | 0.0003501 | 0.006288 |
| 258 | INTRACELLULAR RECEPTOR SIGNALING PATHWAY | 6 | 168 | 0.0003513 | 0.006288 |
| 259 | HISTONE PHOSPHORYLATION | 3 | 25 | 0.0003501 | 0.006288 |
| 260 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 15 | 983 | 0.0003474 | 0.006288 |
| 261 | POSITIVE REGULATION OF KINASE ACTIVITY | 10 | 482 | 0.0003534 | 0.006301 |
| 262 | RESPONSE TO HYDROGEN PEROXIDE | 5 | 109 | 0.0003552 | 0.006309 |
| 263 | POSITIVE REGULATION OF CATABOLIC PROCESS | 9 | 395 | 0.0003581 | 0.006335 |
| 264 | CELLULAR RESPONSE TO HYDROGEN PEROXIDE | 4 | 61 | 0.0003676 | 0.006454 |
| 265 | POSITIVE REGULATION OF STEM CELL PROLIFERATION | 4 | 61 | 0.0003676 | 0.006454 |
| 266 | NUCLEOSIDE MONOPHOSPHATE METABOLIC PROCESS | 7 | 239 | 0.0003773 | 0.0066 |
| 267 | POSITIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 6 | 171 | 0.0003861 | 0.006729 |
| 268 | NEGATIVE REGULATION OF STRIATED MUSCLE CELL DIFFERENTIATION | 3 | 26 | 0.0003941 | 0.006843 |
| 269 | RESPONSE TO AMINO ACID | 5 | 112 | 0.0004026 | 0.006963 |
| 270 | SINGLE ORGANISM BIOSYNTHETIC PROCESS | 18 | 1340 | 0.0004052 | 0.006984 |
| 271 | SENSORY ORGAN DEVELOPMENT | 10 | 493 | 0.0004219 | 0.007245 |
| 272 | REGULATION OF HISTONE H3 K4 METHYLATION | 3 | 27 | 0.0004416 | 0.007555 |
| 273 | PRODUCTION OF MOLECULAR MEDIATOR OF IMMUNE RESPONSE | 4 | 65 | 0.0004688 | 0.007991 |
| 274 | RESPONSE TO COPPER ION | 3 | 28 | 0.0004926 | 0.008366 |
| 275 | CELL AGING | 4 | 67 | 0.0005263 | 0.008905 |
| 276 | REGULATION OF CELL DIFFERENTIATION | 19 | 1492 | 0.0005332 | 0.008989 |
| 277 | DEVELOPMENTAL GROWTH | 8 | 333 | 0.0005422 | 0.009108 |
| 278 | REGULATION OF CARDIAC MUSCLE CELL PROLIFERATION | 3 | 29 | 0.0005473 | 0.009127 |
| 279 | NUCLEOTIDE EXCISION REPAIR PREINCISION COMPLEX ASSEMBLY | 3 | 29 | 0.0005473 | 0.009127 |
| 280 | CELLULAR RESPONSE TO OXIDATIVE STRESS | 6 | 184 | 0.0005691 | 0.009457 |
| 281 | MEIOTIC CELL CYCLE | 6 | 186 | 0.0006024 | 0.009975 |
| 282 | DNA STRAND ELONGATION | 3 | 30 | 0.0006056 | 0.009993 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | KINASE BINDING | 19 | 606 | 9.211e-10 | 8.557e-07 |
| 2 | MISMATCHED DNA BINDING | 5 | 12 | 3.692e-09 | 1.715e-06 |
| 3 | ENZYME BINDING | 30 | 1737 | 1.239e-08 | 3.836e-06 |
| 4 | CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 6 | 34 | 3.022e-08 | 7.018e-06 |
| 5 | STEROID HORMONE RECEPTOR BINDING | 7 | 81 | 3.323e-07 | 6.173e-05 |
| 6 | IDENTICAL PROTEIN BINDING | 22 | 1209 | 6.873e-07 | 0.0001064 |
| 7 | ANDROGEN RECEPTOR BINDING | 5 | 39 | 2.382e-06 | 0.0002766 |
| 8 | MACROMOLECULAR COMPLEX BINDING | 23 | 1399 | 2.107e-06 | 0.0002766 |
| 9 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 12 | 445 | 6.848e-06 | 0.0007069 |
| 10 | PROTEIN COMPLEX BINDING | 17 | 935 | 1.526e-05 | 0.001402 |
| 11 | CYCLIN DEPENDENT PROTEIN SERINE THREONINE KINASE REGULATOR ACTIVITY | 4 | 28 | 1.66e-05 | 0.001402 |
| 12 | DAMAGED DNA BINDING | 5 | 63 | 2.616e-05 | 0.002025 |
| 13 | TAU PROTEIN KINASE ACTIVITY | 3 | 12 | 3.529e-05 | 0.002522 |
| 14 | HORMONE RECEPTOR BINDING | 7 | 168 | 4.249e-05 | 0.002819 |
| 15 | RECEPTOR BINDING | 21 | 1476 | 5.492e-05 | 0.003364 |
| 16 | RNA BINDING | 22 | 1598 | 5.795e-05 | 0.003364 |
| 17 | KINASE ACTIVITY | 15 | 842 | 6.407e-05 | 0.003501 |
| 18 | ADENYL NUCLEOTIDE BINDING | 21 | 1514 | 7.91e-05 | 0.004082 |
| 19 | RNA POLYMERASE II CARBOXY TERMINAL DOMAIN KINASE ACTIVITY | 3 | 16 | 8.839e-05 | 0.004322 |
| 20 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 16 | 992 | 0.0001152 | 0.005353 |
| 21 | CYCLIN BINDING | 3 | 19 | 0.0001511 | 0.006381 |
| 22 | HISTONE KINASE ACTIVITY | 3 | 19 | 0.0001511 | 0.006381 |
| 23 | TRANSCRIPTION FACTOR BINDING | 11 | 524 | 0.0001604 | 0.006479 |
| 24 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR BINDING | 4 | 50 | 0.00017 | 0.00658 |
| 25 | PROTEIN DIMERIZATION ACTIVITY | 17 | 1149 | 0.0001954 | 0.006722 |
| 26 | PROTEIN HOMODIMERIZATION ACTIVITY | 13 | 722 | 0.0001841 | 0.006722 |
| 27 | RIBONUCLEOTIDE BINDING | 23 | 1860 | 0.0001939 | 0.006722 |
| 28 | PROTEIN PHOSPHATASE TYPE 2A REGULATOR ACTIVITY | 3 | 21 | 0.0002057 | 0.006826 |
| 29 | PROTEIN KINASE ACTIVITY | 12 | 640 | 0.0002283 | 0.007314 |
| 30 | TRANSFERASE ACTIVITY TRANSFERRING NITROGENOUS GROUPS | 3 | 23 | 0.0002717 | 0.008415 |
| 31 | DNA SECONDARY STRUCTURE BINDING | 3 | 24 | 0.0003093 | 0.009269 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CHROMOSOME | 25 | 880 | 1.113e-11 | 6.499e-09 |
| 2 | NUCLEAR CHROMOSOME | 16 | 523 | 3.148e-08 | 9.193e-06 |
| 3 | CHROMOSOMAL REGION | 12 | 330 | 3.053e-07 | 5.943e-05 |
| 4 | PROTEIN KINASE COMPLEX | 7 | 90 | 6.853e-07 | 1e-04 |
| 5 | MISMATCH REPAIR COMPLEX | 4 | 14 | 8.618e-07 | 0.0001007 |
| 6 | MICROTUBULE CYTOSKELETON | 20 | 1068 | 1.547e-06 | 0.0001291 |
| 7 | CONDENSED CHROMOSOME | 9 | 195 | 1.42e-06 | 0.0001291 |
| 8 | CENTROSOME | 13 | 487 | 3.088e-06 | 0.0002254 |
| 9 | MICROTUBULE ORGANIZING CENTER | 14 | 623 | 9.112e-06 | 0.0005913 |
| 10 | CONDENSED CHROMOSOME OUTER KINETOCHORE | 3 | 12 | 3.529e-05 | 0.002061 |
| 11 | TRANSCRIPTION FACTOR COMPLEX | 9 | 298 | 4.293e-05 | 0.002279 |
| 12 | TRANSFERASE COMPLEX TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 8 | 237 | 5.343e-05 | 0.002565 |
| 13 | CATALYTIC COMPLEX | 17 | 1038 | 5.71e-05 | 0.002565 |
| 14 | PRONUCLEUS | 3 | 15 | 7.211e-05 | 0.003008 |
| 15 | MITOCHONDRIAL MATRIX | 10 | 412 | 9.995e-05 | 0.003648 |
| 16 | DNA REPAIR COMPLEX | 4 | 43 | 9.386e-05 | 0.003648 |
| 17 | CONDENSED NUCLEAR CHROMOSOME | 5 | 85 | 0.0001108 | 0.003807 |
| 18 | CYTOSKELETON | 24 | 1967 | 0.0001676 | 0.005439 |
| 19 | MITOCHONDRIAL PART | 15 | 953 | 0.0002498 | 0.007678 |
| 20 | CHROMOSOME TELOMERIC REGION | 6 | 162 | 0.0002892 | 0.008444 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
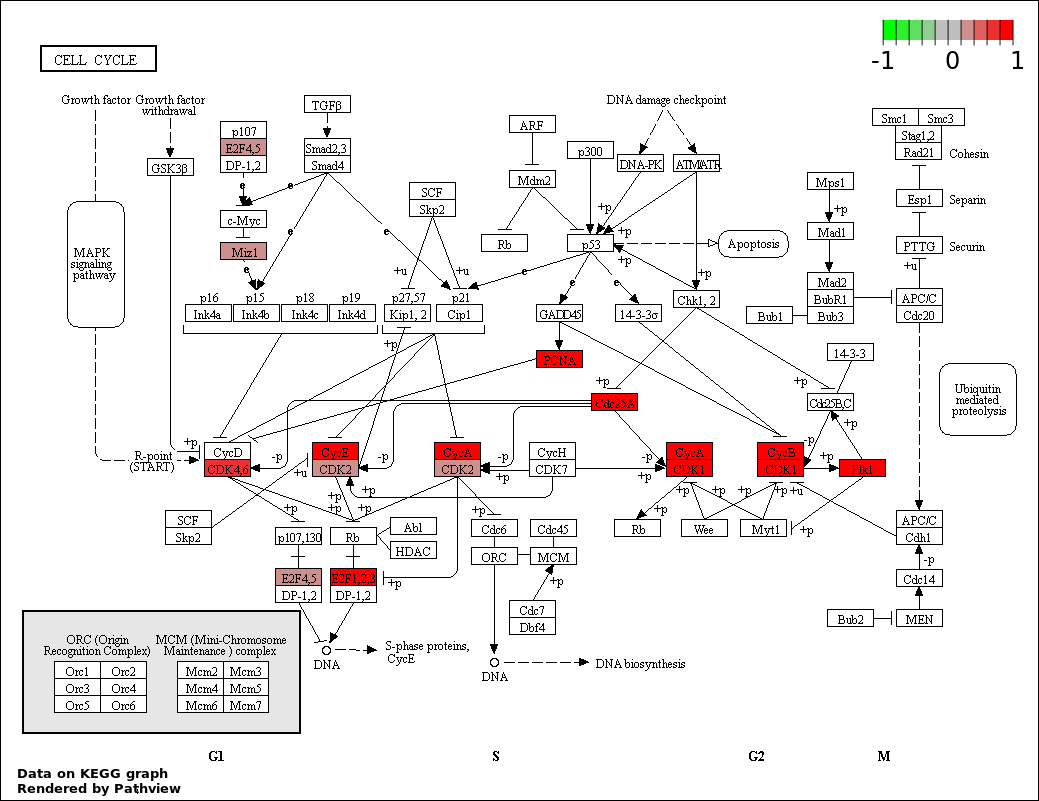
| 1 | hsa04110_Cell_cycle | 14 | 128 | 1.164e-14 | 2.06e-12 | |
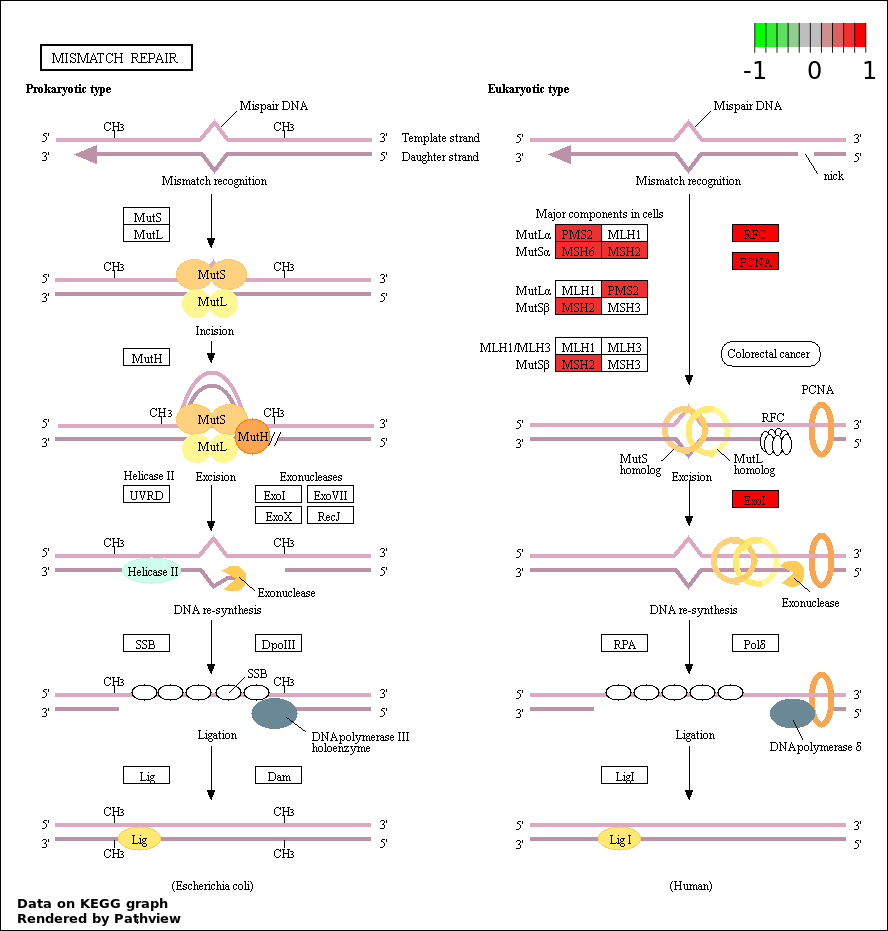
| 2 | hsa03430_Mismatch_repair | 7 | 23 | 3.051e-11 | 2.7e-09 | |
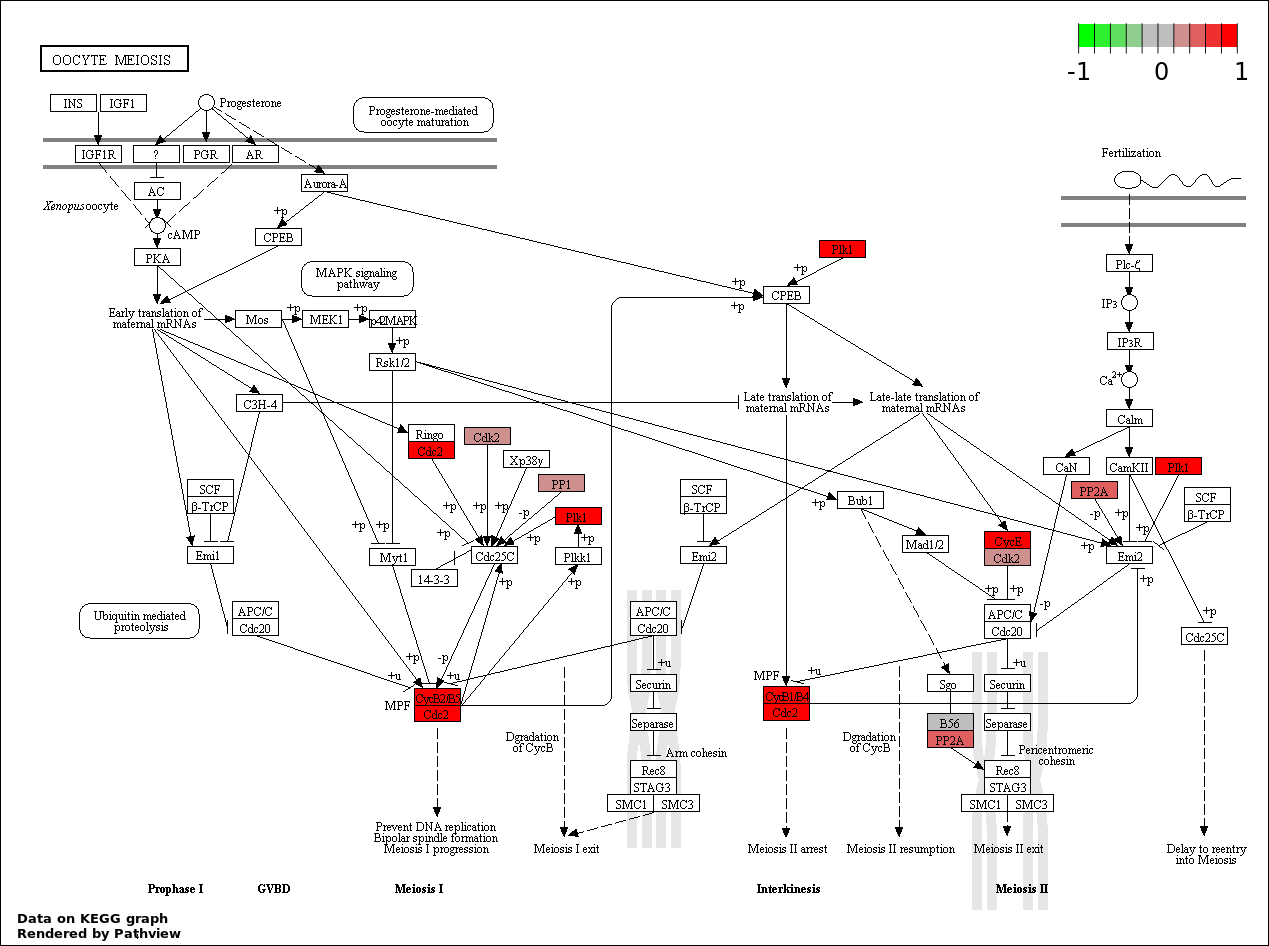
| 3 | hsa04114_Oocyte_meiosis | 10 | 114 | 7.717e-10 | 4.553e-08 | |
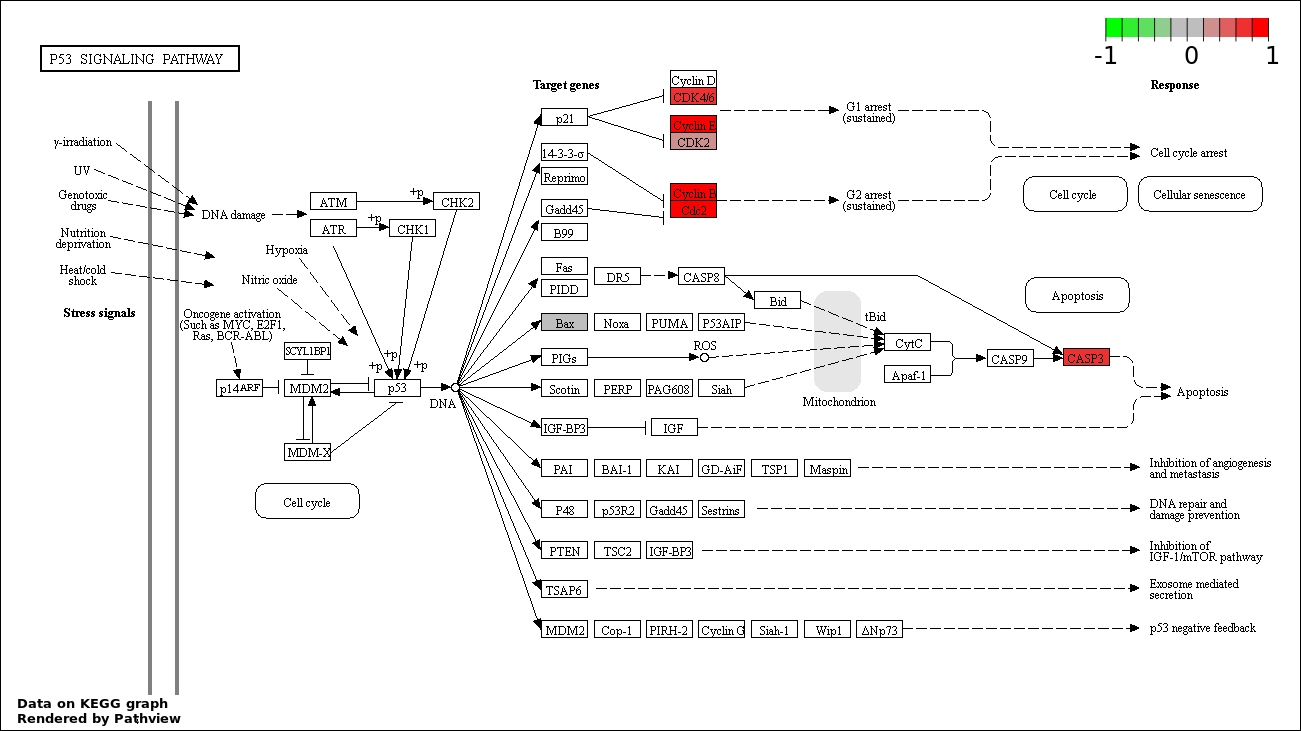
| 4 | hsa04115_p53_signaling_pathway | 8 | 69 | 4.401e-09 | 1.948e-07 | |
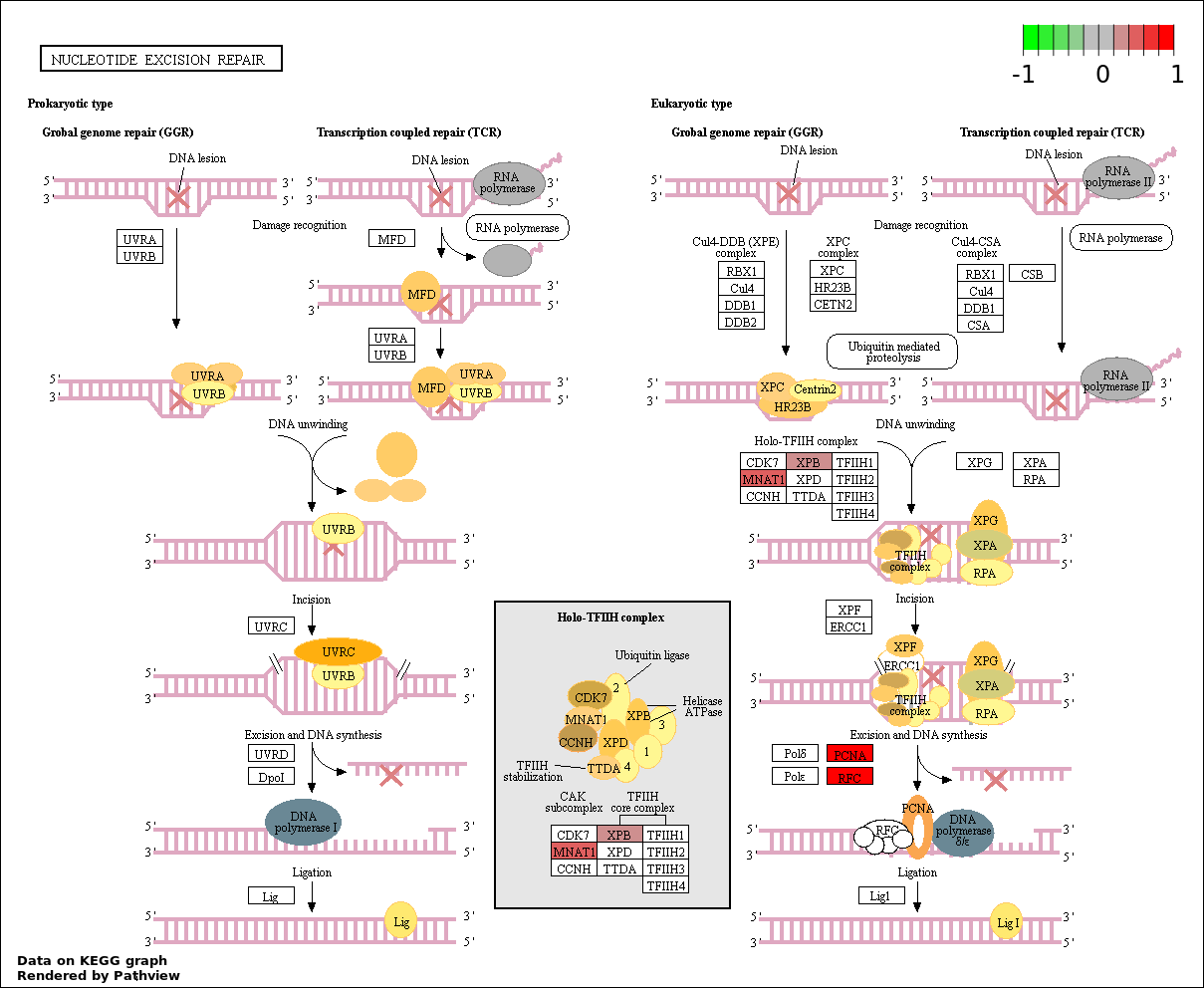
| 5 | hsa03420_Nucleotide_excision_repair | 5 | 45 | 4.923e-06 | 0.0001743 | |
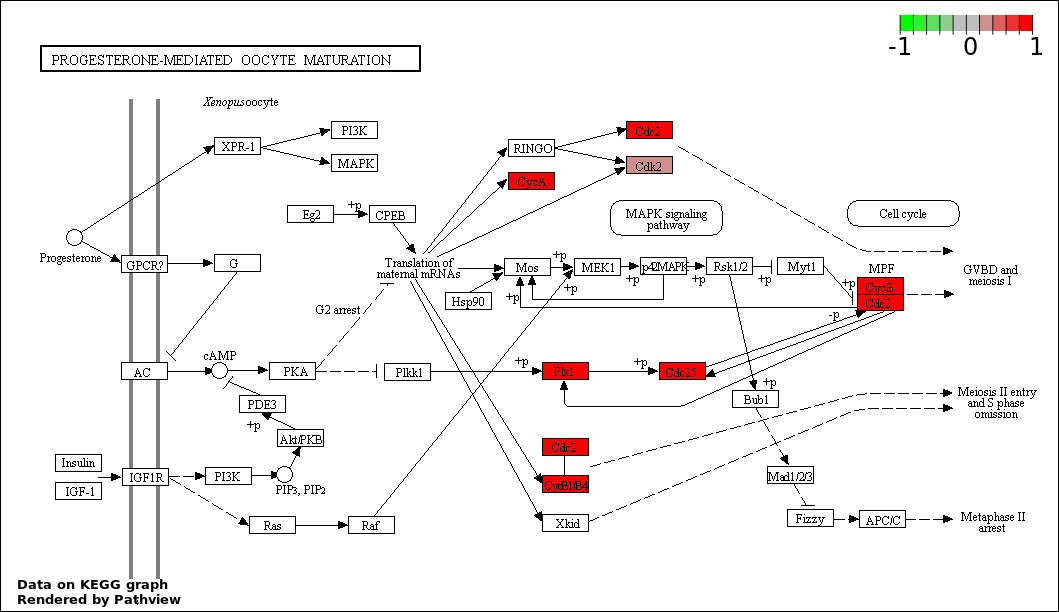
| 6 | hsa04914_Progesterone.mediated_oocyte_maturation | 6 | 87 | 8.939e-06 | 0.0002637 | |
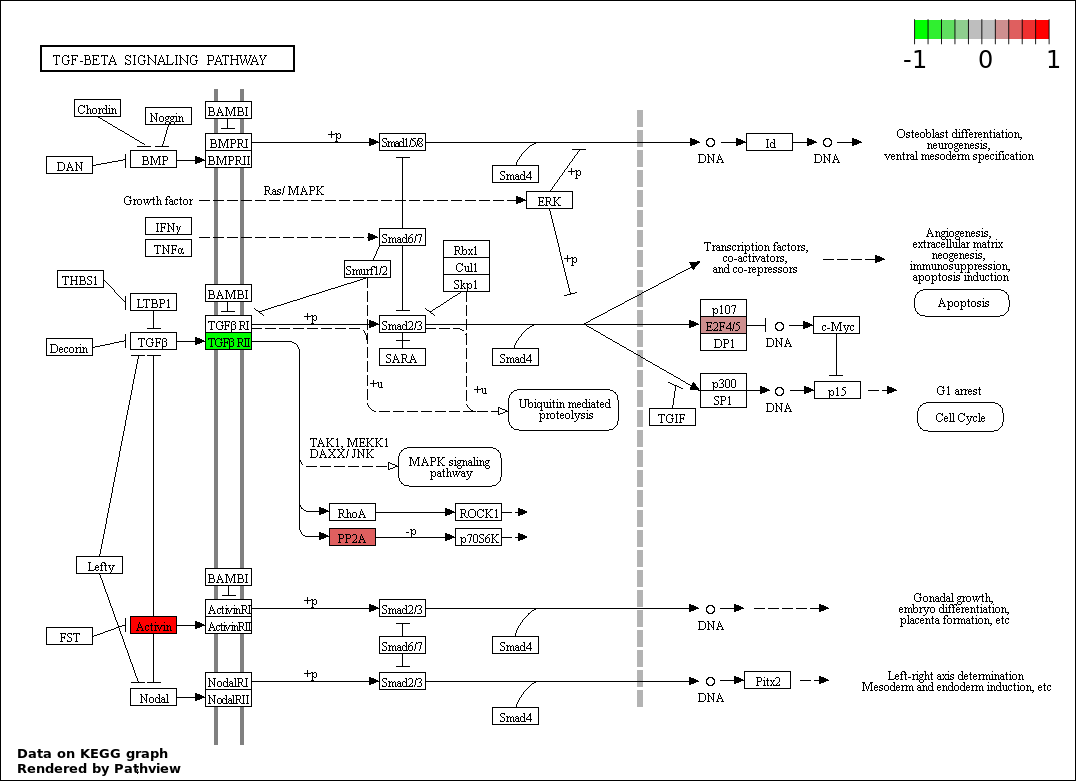
| 7 | hsa04350_TGF.beta_signaling_pathway | 5 | 85 | 0.0001108 | 0.002802 | |
| 8 | hsa04530_Tight_junction | 5 | 133 | 0.0008797 | 0.01946 | |
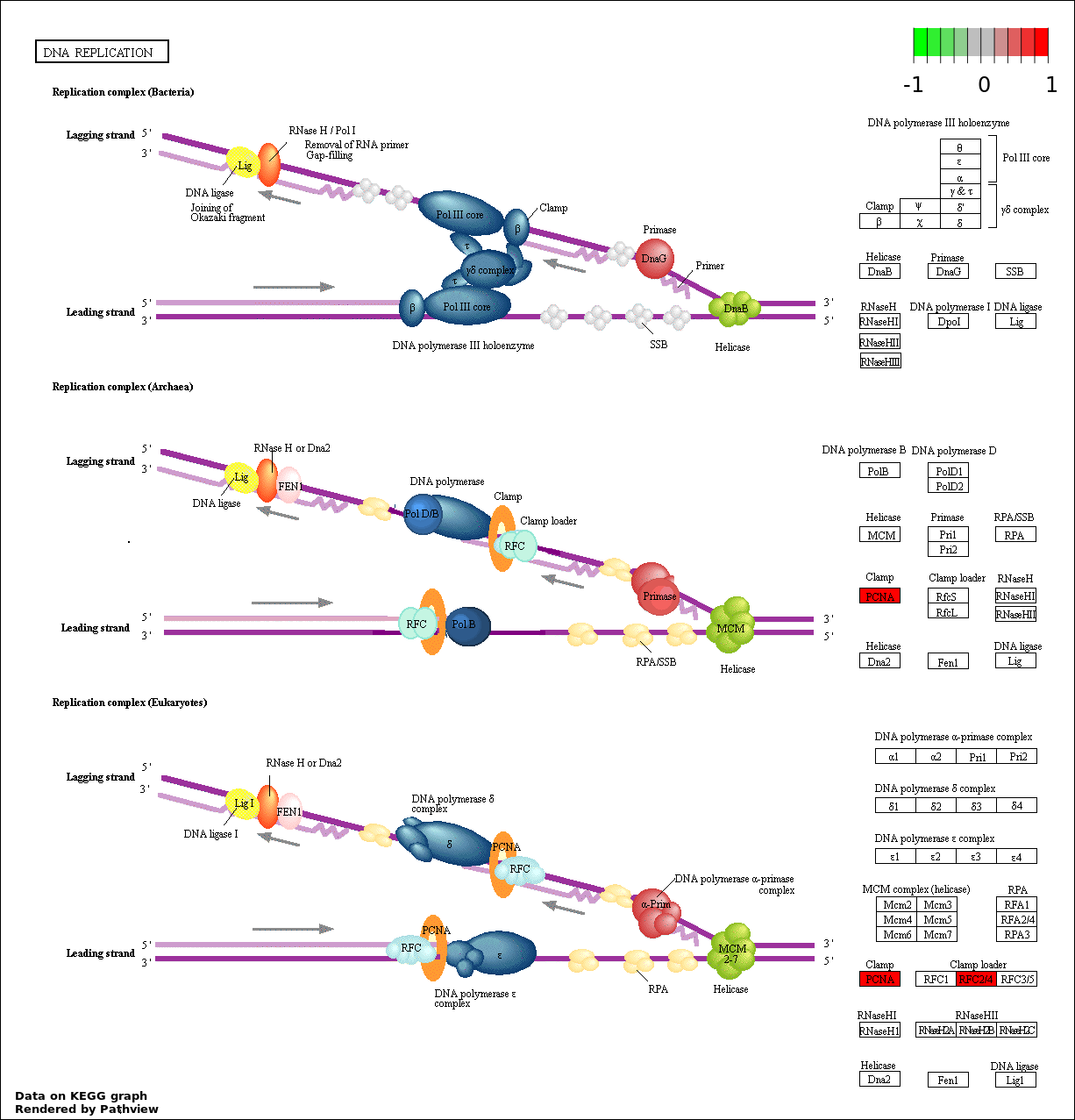
| 9 | hsa03030_DNA_replication | 3 | 36 | 0.00104 | 0.02045 | |
| 10 | hsa04210_Apoptosis | 4 | 89 | 0.001527 | 0.02703 | |
| 11 | hsa04730_Long.term_depression | 3 | 70 | 0.006954 | 0.1119 | |
| 12 | hsa04310_Wnt_signaling_pathway | 4 | 151 | 0.01002 | 0.1478 | |
| 13 | hsa03015_mRNA_surveillance_pathway | 3 | 83 | 0.01108 | 0.1509 | |
| 14 | hsa00250_Alanine._aspartate_and_glutamate_metabolism | 2 | 32 | 0.01358 | 0.1717 | |
| 15 | hsa03410_Base_excision_repair | 2 | 34 | 0.01525 | 0.18 | |
| 16 | hsa04010_MAPK_signaling_pathway | 5 | 268 | 0.0169 | 0.187 | |
| 17 | hsa04916_Melanogenesis | 3 | 101 | 0.01872 | 0.1949 | |
| 18 | hsa04330_Notch_signaling_pathway | 2 | 47 | 0.02806 | 0.2614 | |
| 19 | hsa00480_Glutathione_metabolism | 2 | 50 | 0.03145 | 0.2784 | |
| 20 | hsa00983_Drug_metabolism_._other_enzymes | 2 | 52 | 0.03381 | 0.2849 | |
| 21 | hsa04910_Insulin_signaling_pathway | 3 | 138 | 0.04159 | 0.3347 | |
| 22 | hsa04720_Long.term_potentiation | 2 | 70 | 0.05776 | 0.4445 | |
| 23 | hsa04974_Protein_digestion_and_absorption | 2 | 81 | 0.07454 | 0.5498 | |
| 24 | hsa04510_Focal_adhesion | 3 | 200 | 0.1002 | 0.6396 | |
| 25 | hsa04144_Endocytosis | 3 | 203 | 0.1036 | 0.6396 | |
| 26 | hsa00240_Pyrimidine_metabolism | 2 | 99 | 0.1048 | 0.6396 | |
| 27 | hsa04810_Regulation_of_actin_cytoskeleton | 3 | 214 | 0.1164 | 0.6869 | |
| 28 | hsa04660_T_cell_receptor_signaling_pathway | 2 | 108 | 0.121 | 0.6907 | |
| 29 | hsa04722_Neurotrophin_signaling_pathway | 2 | 127 | 0.1569 | 0.8027 | |
| 30 | hsa03040_Spliceosome | 2 | 128 | 0.1588 | 0.8027 | |
| 31 | hsa04360_Axon_guidance | 2 | 130 | 0.1627 | 0.8027 | |
| 32 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 2 | 136 | 0.1745 | 0.8247 | |
| 33 | hsa04120_Ubiquitin_mediated_proteolysis | 2 | 139 | 0.1805 | 0.8247 | |
| 34 | hsa03013_RNA_transport | 2 | 152 | 0.2066 | 0.8706 | |
| 35 | hsa00230_Purine_metabolism | 2 | 162 | 0.2269 | 0.9128 | |
| 36 | hsa04141_Protein_processing_in_endoplasmic_reticulum | 2 | 168 | 0.2392 | 0.9409 | |
| 37 | hsa04062_Chemokine_signaling_pathway | 2 | 189 | 0.2824 | 1 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | RP11-1024P17.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-9-5p | 20 | TGFBR2 | Sponge network | -2.062 | 0 | -1.915 | 0 | 0.762 |
| 2 | LINC00968 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-93-5p | 20 | TGFBR2 | Sponge network | -4.19 | 0 | -1.915 | 0 | 0.609 |
| 3 | RP11-389C8.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-93-5p | 18 | TGFBR2 | Sponge network | -2.039 | 0 | -1.915 | 0 | 0.579 |
| 4 | TBX5-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-92a-3p | 19 | TGFBR2 | Sponge network | -2.108 | 0 | -1.915 | 0 | 0.563 |
| 5 | AC109642.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 19 | TGFBR2 | Sponge network | -2.791 | 0 | -1.915 | 0 | 0.551 |
| 6 | LINC00702 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 22 | TGFBR2 | Sponge network | -2.856 | 0 | -1.915 | 0 | 0.534 |
| 7 | MAGI2-AS3 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 21 | TGFBR2 | Sponge network | -1.892 | 0 | -1.915 | 0 | 0.522 |
| 8 | RP11-720L2.4 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -2.305 | 0 | -1.915 | 0 | 0.509 |
| 9 | FENDRR | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-93-5p | 15 | TGFBR2 | Sponge network | -4.222 | 0 | -1.915 | 0 | 0.506 |
| 10 | LINC00961 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-9-5p | 12 | TGFBR2 | Sponge network | -2.724 | 0 | -1.915 | 0 | 0.497 |
| 11 | LINC00472 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -2.952 | 0 | -1.915 | 0 | 0.479 |
| 12 | AC011899.9 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 14 | TGFBR2 | Sponge network | -2.611 | 0 | -1.915 | 0 | 0.448 |
| 13 | RP11-1008C21.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-92a-3p;hsa-miR-93-5p | 16 | TGFBR2 | Sponge network | -1.249 | 0 | -1.915 | 0 | 0.446 |
| 14 | RP11-378A13.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-93-5p | 17 | TGFBR2 | Sponge network | -1.713 | 0 | -1.915 | 0 | 0.446 |
| 15 | RP11-399O19.9 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-93-5p | 15 | TGFBR2 | Sponge network | -0.873 | 0.00072 | -1.915 | 0 | 0.445 |
| 16 | AF131215.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-9-5p | 12 | TGFBR2 | Sponge network | -2.09 | 0 | -1.915 | 0 | 0.435 |
| 17 | AC079630.4 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-93-5p | 16 | TGFBR2 | Sponge network | -3.758 | 0 | -1.915 | 0 | 0.433 |
| 18 | CTD-2013N24.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-93-5p | 19 | TGFBR2 | Sponge network | -1.745 | 0 | -1.915 | 0 | 0.42 |
| 19 | RP11-354E11.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-9-5p | 17 | TGFBR2 | Sponge network | -2.138 | 0 | -1.915 | 0 | 0.418 |
| 20 | RP11-401P9.4 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -3.04 | 0 | -1.915 | 0 | 0.413 |
| 21 | RP11-476D10.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-590-3p | 12 | TGFBR2 | Sponge network | -4.519 | 0 | -1.915 | 0 | 0.41 |
| 22 | WDFY3-AS2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-590-5p | 16 | TGFBR2 | Sponge network | -1.297 | 0 | -1.915 | 0 | 0.407 |
| 23 | RP11-532F6.3 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-9-5p | 12 | TGFBR2 | Sponge network | -2.028 | 0 | -1.915 | 0 | 0.391 |
| 24 | RP11-352D13.6 | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-93-5p | 10 | TGFBR2 | Sponge network | -4.634 | 0 | -1.915 | 0 | 0.39 |
| 25 | MIR22HG | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -1.704 | 0 | -1.915 | 0 | 0.39 |
| 26 | MIR497HG | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-93-5p | 18 | TGFBR2 | Sponge network | -2.142 | 0 | -1.915 | 0 | 0.387 |
| 27 | RP11-293M10.6 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-590-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -1.199 | 0.00063 | -1.915 | 0 | 0.386 |
| 28 | RP5-839B4.8 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -5.037 | 0 | -1.915 | 0 | 0.384 |
| 29 | RP11-1008C21.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-590-3p | 14 | TGFBR2 | Sponge network | -1.826 | 3.0E-5 | -1.915 | 0 | 0.382 |
| 30 | AC007743.1 | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-320b;hsa-miR-429;hsa-miR-590-5p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -2.595 | 0 | -1.915 | 0 | 0.375 |
| 31 | HHIP-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-590-3p | 14 | TGFBR2 | Sponge network | -2.807 | 0 | -1.915 | 0 | 0.374 |
| 32 | RP11-456K23.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-93-5p | 19 | TGFBR2 | Sponge network | -1.488 | 0 | -1.915 | 0 | 0.372 |
| 33 | GAS6-AS2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-5p;hsa-miR-92a-3p | 18 | TGFBR2 | Sponge network | -1.761 | 0 | -1.915 | 0 | 0.371 |
| 34 | LINC00261 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p | 15 | TGFBR2 | Sponge network | -2.566 | 0.00025 | -1.915 | 0 | 0.367 |
| 35 | RP11-10C24.3 | hsa-miR-106a-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-5p | 11 | TGFBR2 | Sponge network | -1.013 | 0 | -1.915 | 0 | 0.361 |
| 36 | RP1-78O14.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p | 12 | TGFBR2 | Sponge network | -4.409 | 0 | -1.915 | 0 | 0.358 |
| 37 | RP11-77A13.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -6.738 | 0 | -1.915 | 0 | 0.35 |
| 38 | RP11-365O16.3 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -2.765 | 0.00017 | -1.915 | 0 | 0.349 |
| 39 | LINC00920 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p | 10 | TGFBR2 | Sponge network | -0.998 | 0.0011 | -1.915 | 0 | 0.347 |
| 40 | RP11-88I21.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -8.789 | 0 | -1.915 | 0 | 0.346 |
| 41 | RP5-1042I8.7 | hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-92a-3p;hsa-miR-93-5p | 10 | TGFBR2 | Sponge network | -0.733 | 0.00018 | -1.915 | 0 | 0.345 |
| 42 | RP11-517P14.2 | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.795 | 6.0E-5 | -1.915 | 0 | 0.344 |
| 43 | LIPE-AS1 | hsa-miR-106a-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-5p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.734 | 0.00039 | -1.915 | 0 | 0.343 |
| 44 | RP11-680F8.3 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -1.628 | 0 | -1.915 | 0 | 0.34 |
| 45 | CYP1B1-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20b-5p;hsa-miR-28-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -1.073 | 0.00045 | -1.915 | 0 | 0.34 |
| 46 | LL22NC03-86G7.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-9-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -1.177 | 3.0E-5 | -1.915 | 0 | 0.32 |
| 47 | LINC00443 | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p | 11 | TGFBR2 | Sponge network | -3.704 | 0.0003 | -1.915 | 0 | 0.311 |
| 48 | BZRAP1-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-335-3p | 10 | TGFBR2 | Sponge network | -0.785 | 0.00723 | -1.915 | 0 | 0.309 |
| 49 | LINC00092 | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-590-3p | 10 | TGFBR2 | Sponge network | -2.383 | 0 | -1.915 | 0 | 0.308 |
| 50 | AC004947.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 12 | TGFBR2 | Sponge network | -3.94 | 0 | -1.915 | 0 | 0.306 |
| 51 | RP11-1223D19.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -0.862 | 0.05389 | -1.915 | 0 | 0.303 |
| 52 | RBPMS-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130a-3p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-93-5p | 15 | TGFBR2 | Sponge network | -1.548 | 1.0E-5 | -1.915 | 0 | 0.301 |
| 53 | RP11-452C13.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p | 10 | TGFBR2 | Sponge network | -2.725 | 1.0E-5 | -1.915 | 0 | 0.3 |
| 54 | CASC2 | hsa-miR-106a-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 15 | TGFBR2 | Sponge network | -1.086 | 0 | -1.915 | 0 | 0.299 |
| 55 | PSMG3-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20b-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-590-3p | 10 | TGFBR2 | Sponge network | -0.522 | 0.09584 | -1.915 | 0 | 0.294 |
| 56 | RP11-166D19.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-92a-3p | 18 | TGFBR2 | Sponge network | -0.582 | 0.05253 | -1.915 | 0 | 0.294 |
| 57 | AF131215.9 | hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 10 | TGFBR2 | Sponge network | -1.808 | 0 | -1.915 | 0 | 0.292 |
| 58 | RP11-462G12.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -1.071 | 0.01175 | -1.915 | 0 | 0.291 |
| 59 | RP11-400K9.4 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -1.193 | 0.00359 | -1.915 | 0 | 0.291 |
| 60 | RP11-716O23.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-92a-3p | 11 | TGFBR2 | Sponge network | -5.465 | 0 | -1.915 | 0 | 0.286 |
| 61 | HLA-F-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -0.495 | 0.12126 | -1.915 | 0 | 0.283 |
| 62 | TPRG1-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-9-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.947 | 0.04384 | -1.915 | 0 | 0.282 |
| 63 | BAIAP2-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.182 | 0.51705 | -1.915 | 0 | 0.272 |
| 64 | CTD-2008P7.9 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130a-3p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-9-5p | 15 | TGFBR2 | Sponge network | -1.912 | 1.0E-5 | -1.915 | 0 | 0.27 |
| 65 | LINC01010 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.54 | 0.27666 | -1.915 | 0 | 0.267 |
| 66 | CTD-2003C8.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-454-3p | 11 | TGFBR2 | Sponge network | -3.403 | 0 | -1.915 | 0 | 0.266 |
| 67 | BDNF-AS | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -0.568 | 0.02011 | -1.915 | 0 | 0.263 |
| 68 | SAP30L-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-590-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 10 | TGFBR2 | Sponge network | 0.631 | 0.02291 | -1.915 | 0 | 0.263 |
| 69 | RP4-639F20.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-320b | 10 | TGFBR2 | Sponge network | -1.312 | 0 | -1.915 | 0 | 0.263 |
| 70 | CTC-523E23.4 | hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-9-5p | 10 | TGFBR2 | Sponge network | -1.636 | 0.00051 | -1.915 | 0 | 0.261 |
| 71 | LINC00922 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-9-5p;hsa-miR-92a-3p | 11 | TGFBR2 | Sponge network | -0.842 | 0.11239 | -1.915 | 0 | 0.26 |
| 72 | LINC00619 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p | 10 | TGFBR2 | Sponge network | -2.307 | 0.02217 | -1.915 | 0 | 0.253 |
| 73 | PCED1B-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-92a-3p;hsa-miR-93-5p | 14 | TGFBR2 | Sponge network | -0.672 | 0.02084 | -1.915 | 0 | 0.252 |
| 74 | AC093495.4 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-9-5p;hsa-miR-93-5p | 13 | TGFBR2 | Sponge network | -1.583 | 0.00599 | -1.915 | 0 | 0.25 |