This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-107 | ACVR1 | 1.31 | 0 | -0.44 | 0.01143 | miRanda | -0.32 | 0 | NA | |
| 2 | hsa-miR-148b-3p | ACVR1 | 1.98 | 0 | -0.44 | 0.01143 | MirTarget | -0.38 | 0 | NA | |
| 3 | hsa-miR-186-5p | ACVR1 | 1.47 | 0 | -0.44 | 0.01143 | miRNAWalker2 validate | -0.37 | 0 | NA | |
| 4 | hsa-miR-30b-5p | ACVR1 | 0.8 | 0.00013 | -0.44 | 0.01143 | MirTarget | -0.33 | 0 | NA | |
| 5 | hsa-miR-30c-5p | ACVR1 | 0.78 | 0.00029 | -0.44 | 0.01143 | MirTarget; miRNATAP | -0.33 | 0 | NA | |
| 6 | hsa-miR-30e-5p | ACVR1 | 1.24 | 0 | -0.44 | 0.01143 | MirTarget | -0.41 | 0 | NA | |
| 7 | hsa-miR-181c-5p | ACVR1C | 1.59 | 0 | 0.21 | 0.68824 | miRNATAP | -0.41 | 6.0E-5 | NA | |
| 8 | hsa-let-7i-5p | ACVR2A | 0.41 | 0.00623 | -0.44 | 0.02646 | miRNATAP | -0.4 | 0 | NA | |
| 9 | hsa-miR-590-3p | BMP4 | 2.59 | 0 | -1.57 | 0.00013 | miRanda | -0.37 | 0 | NA | |
| 10 | hsa-miR-576-5p | BMP5 | 2.2 | 0 | -4.53 | 0 | mirMAP | -0.46 | 0.00072 | NA | |
| 11 | hsa-miR-590-3p | BMP5 | 2.59 | 0 | -4.53 | 0 | mirMAP | -0.38 | 0.00244 | NA | |
| 12 | hsa-miR-338-3p | BMP7 | 0.45 | 0.14458 | -1.78 | 0.02876 | miRanda | -0.41 | 0.00118 | NA | |
| 13 | hsa-miR-342-3p | BMP7 | 1.49 | 0 | -1.78 | 0.02876 | miRNAWalker2 validate; miRTarBase; miRanda | -0.56 | 4.0E-5 | NA | |
| 14 | hsa-miR-30e-3p | BMP8A | -0.22 | 0.23538 | 1.42 | 0 | MirTarget | -0.33 | 0 | NA | |
| 15 | hsa-miR-101-3p | BMPR1B | 0.52 | 0.00376 | 0.07 | 0.91404 | miRNATAP | -0.9 | 0 | NA | |
| 16 | hsa-miR-125a-5p | BMPR1B | 0.6 | 0.01023 | 0.07 | 0.91404 | MirTarget; miRNATAP | -0.81 | 0 | NA | |
| 17 | hsa-miR-182-5p | BMPR1B | 3.54 | 0 | 0.07 | 0.91404 | MirTarget | -0.35 | 1.0E-5 | NA | |
| 18 | hsa-miR-26b-5p | BMPR1B | 0.89 | 0 | 0.07 | 0.91404 | mirMAP | -0.53 | 0.00146 | NA | |
| 19 | hsa-miR-330-3p | BMPR1B | 0.88 | 0.00074 | 0.07 | 0.91404 | mirMAP | -0.65 | 0 | NA | |
| 20 | hsa-miR-429 | BMPR1B | 4.49 | 0 | 0.07 | 0.91404 | miRNATAP | -0.33 | 0 | NA | |
| 21 | hsa-miR-186-5p | BMPR2 | 1.47 | 0 | -0.59 | 4.0E-5 | mirMAP | -0.32 | 0 | NA | |
| 22 | hsa-miR-125a-5p | CDKN2B | 0.6 | 0.01023 | -0.11 | 0.83235 | miRanda | -0.31 | 0.00518 | NA | |
| 23 | hsa-miR-28-5p | CDKN2B | 0.23 | 0.07429 | -0.11 | 0.83235 | miRanda | -0.67 | 0.00119 | NA | |
| 24 | hsa-miR-582-5p | CDKN2B | 0.61 | 0.03299 | -0.11 | 0.83235 | miRNATAP | -0.44 | 0 | NA | |
| 25 | hsa-miR-671-5p | CDKN2B | 2.98 | 0 | -0.11 | 0.83235 | PITA | -0.43 | 0 | NA | |
| 26 | hsa-let-7a-5p | CHRD | 0.62 | 3.0E-5 | -2 | 0 | MirTarget; TargetScan; miRNATAP | -0.96 | 0 | NA | |
| 27 | hsa-let-7b-5p | CHRD | 0.6 | 0.0014 | -2 | 0 | MirTarget; miRNATAP | -0.71 | 0 | NA | |
| 28 | hsa-let-7d-5p | CHRD | 1.33 | 0 | -2 | 0 | MirTarget; miRNATAP | -0.72 | 0 | NA | |
| 29 | hsa-let-7f-5p | CHRD | 0.5 | 0.00356 | -2 | 0 | MirTarget; miRNATAP | -0.5 | 1.0E-5 | NA | |
| 30 | hsa-let-7g-3p | CHRD | 3.26 | 0 | -2 | 0 | MirTarget | -0.53 | 0 | NA | |
| 31 | hsa-let-7g-5p | CHRD | 1.2 | 0 | -2 | 0 | MirTarget; miRNATAP | -0.85 | 0 | NA | |
| 32 | hsa-miR-196b-5p | CHRD | 1.72 | 0 | -2 | 0 | miRNATAP | -0.41 | 0 | NA | |
| 33 | hsa-miR-455-3p | CHRD | 3.06 | 0 | -2 | 0 | miRNATAP | -0.3 | 0 | NA | |
| 34 | hsa-miR-98-5p | CHRD | 1.11 | 0 | -2 | 0 | MirTarget | -0.67 | 0 | NA | |
| 35 | hsa-let-7a-5p | DCN | 0.62 | 3.0E-5 | -3.45 | 0 | MirTarget | -1.3 | 0 | NA | |
| 36 | hsa-let-7b-5p | DCN | 0.6 | 0.0014 | -3.45 | 0 | MirTarget | -0.74 | 0 | NA | |
| 37 | hsa-let-7d-5p | DCN | 1.33 | 0 | -3.45 | 0 | MirTarget | -1.31 | 0 | NA | |
| 38 | hsa-let-7e-5p | DCN | 0.86 | 6.0E-5 | -3.45 | 0 | MirTarget | -0.61 | 0 | NA | |
| 39 | hsa-let-7f-5p | DCN | 0.5 | 0.00356 | -3.45 | 0 | MirTarget | -0.72 | 0 | NA | |
| 40 | hsa-let-7g-5p | DCN | 1.2 | 0 | -3.45 | 0 | MirTarget | -1.37 | 0 | NA | |
| 41 | hsa-miR-181a-5p | DCN | 2.3 | 0 | -3.45 | 0 | MirTarget | -0.51 | 0 | NA | |
| 42 | hsa-miR-181b-5p | DCN | 2.49 | 0 | -3.45 | 0 | MirTarget | -0.58 | 0 | NA | |
| 43 | hsa-miR-181c-5p | DCN | 1.59 | 0 | -3.45 | 0 | MirTarget | -0.5 | 0 | NA | |
| 44 | hsa-miR-181d-5p | DCN | 1.52 | 0 | -3.45 | 0 | MirTarget | -0.48 | 0 | NA | |
| 45 | hsa-miR-186-5p | DCN | 1.47 | 0 | -3.45 | 0 | mirMAP | -1.23 | 0 | NA | |
| 46 | hsa-miR-320a | DCN | 0.44 | 0.03902 | -3.45 | 0 | miRanda | -0.89 | 0 | NA | |
| 47 | hsa-miR-320b | DCN | 1.56 | 0 | -3.45 | 0 | miRanda | -0.87 | 0 | NA | |
| 48 | hsa-miR-374b-3p | DCN | 0.93 | 3.0E-5 | -3.45 | 0 | MirTarget | -0.6 | 0 | NA | |
| 49 | hsa-miR-421 | DCN | 1.18 | 1.0E-5 | -3.45 | 0 | miRanda | -0.71 | 0 | NA | |
| 50 | hsa-miR-491-5p | DCN | 0.62 | 0.04183 | -3.45 | 0 | miRanda | -0.48 | 0 | NA | |
| 51 | hsa-miR-501-5p | DCN | 1.94 | 0 | -3.45 | 0 | mirMAP | -0.49 | 0 | NA | |
| 52 | hsa-miR-590-3p | DCN | 2.59 | 0 | -3.45 | 0 | MirTarget; PITA; miRanda; mirMAP | -0.81 | 0 | NA | |
| 53 | hsa-miR-7-1-3p | DCN | 1.85 | 0 | -3.45 | 0 | mirMAP | -0.82 | 0 | NA | |
| 54 | hsa-miR-944 | DCN | 2.91 | 0 | -3.45 | 0 | mirMAP | -0.32 | 0 | NA | |
| 55 | hsa-miR-98-5p | DCN | 1.11 | 0 | -3.45 | 0 | MirTarget | -1.03 | 0 | NA | |
| 56 | hsa-let-7d-5p | GDF6 | 1.33 | 0 | -1.14 | 0.02791 | MirTarget; miRNATAP | -0.83 | 0 | NA | |
| 57 | hsa-let-7f-5p | GDF6 | 0.5 | 0.00356 | -1.14 | 0.02791 | MirTarget; miRNATAP | -0.5 | 0.00085 | NA | |
| 58 | hsa-let-7g-5p | GDF6 | 1.2 | 0 | -1.14 | 0.02791 | MirTarget; miRNATAP | -0.97 | 0 | NA | |
| 59 | hsa-miR-141-3p | GDF6 | 5.02 | 0 | -1.14 | 0.02791 | TargetScan | -0.34 | 0 | NA | |
| 60 | hsa-miR-141-5p | GDF6 | 3.81 | 0 | -1.14 | 0.02791 | mirMAP | -0.36 | 0 | NA | |
| 61 | hsa-miR-193a-3p | GDF6 | 1.6 | 0 | -1.14 | 0.02791 | miRanda | -0.31 | 0.0001 | NA | |
| 62 | hsa-miR-30d-3p | GDF6 | 0.98 | 4.0E-5 | -1.14 | 0.02791 | miRNATAP | -0.57 | 0 | NA | |
| 63 | hsa-miR-324-5p | GDF6 | 2.96 | 0 | -1.14 | 0.02791 | miRanda | -0.53 | 0 | NA | |
| 64 | hsa-miR-330-3p | GDF6 | 0.88 | 0.00074 | -1.14 | 0.02791 | PITA; miRNATAP | -0.43 | 1.0E-5 | NA | |
| 65 | hsa-miR-339-5p | GDF6 | 2.69 | 0 | -1.14 | 0.02791 | miRanda | -0.49 | 0 | NA | |
| 66 | hsa-miR-532-3p | GDF6 | 1.97 | 0 | -1.14 | 0.02791 | PITA; miRNATAP | -0.73 | 0 | NA | |
| 67 | hsa-miR-589-3p | GDF6 | 2.52 | 0 | -1.14 | 0.02791 | MirTarget | -0.45 | 0 | NA | |
| 68 | hsa-miR-590-3p | GDF6 | 2.59 | 0 | -1.14 | 0.02791 | miRanda; mirMAP | -0.55 | 0 | NA | |
| 69 | hsa-miR-98-5p | GDF6 | 1.11 | 0 | -1.14 | 0.02791 | MirTarget | -0.79 | 0 | NA | |
| 70 | hsa-miR-100-5p | ID1 | -1.48 | 2.0E-5 | -0.11 | 0.79783 | miRNAWalker2 validate | -0.39 | 0 | NA | |
| 71 | hsa-let-7a-3p | ID2 | 1.42 | 0 | -1.55 | 0 | miRNATAP | -0.36 | 0 | NA | |
| 72 | hsa-miR-27a-3p | ID2 | 1.67 | 0 | -1.55 | 0 | MirTarget; miRNATAP | -0.42 | 0 | NA | |
| 73 | hsa-miR-142-3p | ID4 | 2.75 | 0 | -2.19 | 0 | miRanda | -0.46 | 0 | NA | |
| 74 | hsa-miR-186-5p | ID4 | 1.47 | 0 | -2.19 | 0 | miRNAWalker2 validate; mirMAP | -0.31 | 0.00782 | NA | |
| 75 | hsa-miR-27a-3p | ID4 | 1.67 | 0 | -2.19 | 0 | miRNAWalker2 validate | -0.76 | 0 | NA | |
| 76 | hsa-miR-338-5p | ID4 | -0.58 | 0.04722 | -2.19 | 0 | MirTarget; PITA | -0.34 | 1.0E-5 | NA | |
| 77 | hsa-miR-339-5p | ID4 | 2.69 | 0 | -2.19 | 0 | miRanda | -0.3 | 0 | NA | |
| 78 | hsa-miR-342-3p | ID4 | 1.49 | 0 | -2.19 | 0 | MirTarget; PITA; miRanda | -0.42 | 0 | 24475217 | miR 342 regulates BRCA1 expression through modulation of ID4 in breast cancer; To investigate at functional level the role of miR-342 in the pathogenesis of breast cancer we focused our attention on its "in silico" predicted putative target gene ID4 a transcription factor of the helix-loop-helix protein family whose expression is inversely correlated with that of ER; We functionally validated the interaction between ID4 and miR-342 in a reporter Luciferase system; Based on these findings we hypothesized that regulation of ID4 mediated by miR-342 could be involved in the pathogenesis of breast cancer by downregulating BRCA1 expression; Overexpression of miR-342 in these cells reduced ID4 and increased BRCA1 expression supporting a possible role of this mechanism in breast cancer; In the ER-positive MCF7 and in the BRCA1-mutant HCC1937 cell lines miR-342 over-expression only reduced ID4 |
| 79 | hsa-miR-576-5p | ID4 | 2.2 | 0 | -2.19 | 0 | mirMAP | -0.35 | 4.0E-5 | NA | |
| 80 | hsa-miR-589-3p | ID4 | 2.52 | 0 | -2.19 | 0 | MirTarget | -0.31 | 0 | NA | |
| 81 | hsa-miR-590-3p | ID4 | 2.59 | 0 | -2.19 | 0 | miRNAWalker2 validate; miRanda | -0.43 | 0 | NA | |
| 82 | hsa-miR-590-5p | ID4 | 3.18 | 0 | -2.19 | 0 | miRanda | -0.33 | 0 | NA | |
| 83 | hsa-miR-629-3p | ID4 | 2.37 | 0 | -2.19 | 0 | MirTarget | -0.43 | 0 | NA | |
| 84 | hsa-miR-125a-5p | IFNG | 0.6 | 0.01023 | 0.73 | 0.24132 | miRanda | -0.8 | 0 | NA | |
| 85 | hsa-miR-429 | IFNG | 4.49 | 0 | 0.73 | 0.24132 | miRanda | -0.34 | 0 | NA | |
| 86 | hsa-miR-10b-5p | INHBA | -0.16 | 0.55501 | 1.44 | 0.01419 | miRNAWalker2 validate | -0.81 | 0 | NA | |
| 87 | hsa-miR-148b-3p | INHBA | 1.98 | 0 | 1.44 | 0.01419 | miRNAWalker2 validate | -0.82 | 0 | NA | |
| 88 | hsa-miR-181c-5p | INHBA | 1.59 | 0 | 1.44 | 0.01419 | mirMAP | -0.48 | 2.0E-5 | NA | |
| 89 | hsa-miR-200b-3p | INHBA | 3.78 | 0 | 1.44 | 0.01419 | TargetScan | -0.41 | 0 | NA | |
| 90 | hsa-miR-23b-3p | INHBA | -0.25 | 0.1502 | 1.44 | 0.01419 | mirMAP | -0.52 | 0.00183 | NA | |
| 91 | hsa-miR-26b-5p | INHBA | 0.89 | 0 | 1.44 | 0.01419 | mirMAP | -0.48 | 0.00141 | NA | |
| 92 | hsa-miR-30b-5p | INHBA | 0.8 | 0.00013 | 1.44 | 0.01419 | mirMAP | -0.91 | 0 | NA | |
| 93 | hsa-miR-30c-5p | INHBA | 0.78 | 0.00029 | 1.44 | 0.01419 | mirMAP | -1.13 | 0 | NA | |
| 94 | hsa-miR-30d-5p | INHBA | 0.68 | 0.00271 | 1.44 | 0.01419 | mirMAP | -1.29 | 0 | NA | |
| 95 | hsa-miR-30e-5p | INHBA | 1.24 | 0 | 1.44 | 0.01419 | mirMAP | -1.37 | 0 | NA | |
| 96 | hsa-miR-664a-3p | INHBA | 0.63 | 0.0052 | 1.44 | 0.01419 | mirMAP | -1.1 | 0 | NA | |
| 97 | hsa-let-7f-1-3p | INHBB | 1.55 | 0 | -1.64 | 0 | mirMAP | -0.34 | 0 | NA | |
| 98 | hsa-miR-19a-3p | INHBB | 3.42 | 0 | -1.64 | 0 | miRNATAP | -0.33 | 0 | NA | |
| 99 | hsa-miR-19b-3p | INHBB | 2.5 | 0 | -1.64 | 0 | miRNATAP | -0.4 | 0 | NA | |
| 100 | hsa-miR-26b-5p | INHBB | 0.89 | 0 | -1.64 | 0 | miRNATAP | -0.35 | 5.0E-5 | NA | |
| 101 | hsa-miR-32-3p | INHBB | 2.02 | 0 | -1.64 | 0 | mirMAP | -0.34 | 0 | NA | |
| 102 | hsa-miR-454-3p | INHBB | 2.47 | 0 | -1.64 | 0 | miRNATAP | -0.34 | 0 | NA | |
| 103 | hsa-miR-590-3p | INHBB | 2.59 | 0 | -1.64 | 0 | miRanda | -0.37 | 0 | NA | |
| 104 | hsa-miR-590-5p | INHBB | 3.18 | 0 | -1.64 | 0 | miRanda | -0.34 | 0 | NA | |
| 105 | hsa-miR-148a-5p | LEFTY2 | 1.16 | 0.00015 | -2.33 | 0.00054 | MirTarget | -0.32 | 0.00295 | NA | |
| 106 | hsa-miR-192-5p | LEFTY2 | 2.69 | 0 | -2.33 | 0.00054 | MirTarget | -0.87 | 0 | NA | |
| 107 | hsa-miR-215-5p | LEFTY2 | 2.31 | 0 | -2.33 | 0.00054 | MirTarget | -0.45 | 0 | NA | |
| 108 | hsa-miR-339-5p | LEFTY2 | 2.69 | 0 | -2.33 | 0.00054 | miRanda | -0.61 | 0 | NA | |
| 109 | hsa-miR-421 | LEFTY2 | 1.18 | 1.0E-5 | -2.33 | 0.00054 | miRanda | -0.74 | 0 | NA | |
| 110 | hsa-miR-625-5p | LEFTY2 | 1.43 | 0 | -2.33 | 0.00054 | MirTarget | -0.62 | 0 | NA | |
| 111 | hsa-miR-128-3p | LTBP1 | 1.64 | 0 | -1.1 | 0.00013 | MirTarget | -0.41 | 0 | NA | |
| 112 | hsa-miR-148b-3p | LTBP1 | 1.98 | 0 | -1.1 | 0.00013 | MirTarget | -0.49 | 0 | NA | |
| 113 | hsa-miR-7-1-3p | LTBP1 | 1.85 | 0 | -1.1 | 0.00013 | MirTarget | -0.31 | 0 | NA | |
| 114 | hsa-miR-92a-3p | LTBP1 | 2.06 | 0 | -1.1 | 0.00013 | miRNAWalker2 validate | -0.44 | 0 | NA | |
| 115 | hsa-let-7f-5p | MYC | 0.5 | 0.00356 | -1.77 | 0 | miRNAWalker2 validate | -0.3 | 0.00594 | NA | |
| 116 | hsa-let-7g-5p | MYC | 1.2 | 0 | -1.77 | 0 | miRNAWalker2 validate; miRTarBase | -0.65 | 0 | NA | |
| 117 | hsa-miR-34a-5p | MYC | 1.9 | 0 | -1.77 | 0 | miRNAWalker2 validate; miRTarBase | -0.36 | 0 | 25572695; 25686834; 21460242; 22159222; 23640973; 22830357; 22235332 | The c-Myc and CD44 were confirmed as direct targets of miR-34a in EJ cell apoptosis induced by PRE;miR 34a induces cellular senescence via modulation of telomerase activity in human hepatocellular carcinoma by targeting FoxM1/c Myc pathway;Myc mediated repression of microRNA 34a promotes high grade transformation of B cell lymphoma by dysregulation of FoxP1;MicroRNA 34a suppresses malignant transformation by targeting c Myc transcriptional complexes in human renal cell carcinoma; We investigated the functional effects of microRNA-34a miR-34a on c-Myc transcriptional complexes in renal cell carcinoma; miR-34a down-regulated expression of multiple oncogenes including c-Myc by targeting its 3' untranslated region which was revealed by luciferase reporter assays; Our results demonstrate that miR-34a suppresses assembly and function of the c-Myc complex that activates or elongates transcription indicating a novel role of miR-34a in the regulation of transcription by c-Myc;Among them miR-34a was also associated with poor prognosis in 2 independent series of leukemic and nodal MCL and in cooperation with high expression of the MYC oncogene;We report that miR-34a did not inhibit cell proliferation notwithstanding a marked down-regulation of c-MYC;MicroRNA 34a modulates c Myc transcriptional complexes to suppress malignancy in human prostate cancer cells; We studied the functional effects of miR-34a on c-Myc transcriptional complexes in PC-3 prostate cancer cells; miR-34a downregulated expression of c-Myc oncogene by targeting its 3' UTR as shown by luciferase reporter assays; This is the first report to document that miR-34a suppresses assembly and function of the c-Myc-Skp2-Miz1 complex that activates RhoA and the c-Myc-pTEFB complex that elongates transcription of various genes suggesting a novel role of miR-34a in the regulation of transcription by c-Myc complex |
| 118 | hsa-miR-423-5p | MYC | 0.96 | 0 | -1.77 | 0 | miRNAWalker2 validate | -0.59 | 0 | NA | |
| 119 | hsa-miR-98-5p | MYC | 1.11 | 0 | -1.77 | 0 | miRNAWalker2 validate; miRTarBase | -0.6 | 0 | NA | |
| 120 | hsa-miR-146b-5p | NODAL | 1.76 | 0 | -0.19 | 0.58962 | miRanda | -0.36 | 0 | NA | |
| 121 | hsa-miR-148b-3p | NOG | 1.98 | 0 | -1.05 | 0.08272 | MirTarget | -0.71 | 0 | NA | |
| 122 | hsa-miR-186-5p | NOG | 1.47 | 0 | -1.05 | 0.08272 | MirTarget; miRNATAP | -0.57 | 0.00017 | NA | |
| 123 | hsa-miR-200b-3p | NOG | 3.78 | 0 | -1.05 | 0.08272 | MirTarget; TargetScan | -0.31 | 0 | NA | |
| 124 | hsa-miR-200c-3p | NOG | 3.5 | 0 | -1.05 | 0.08272 | MirTarget; miRNATAP | -0.38 | 0 | NA | |
| 125 | hsa-miR-26b-3p | NOG | 2.18 | 0 | -1.05 | 0.08272 | MirTarget | -0.37 | 0.00098 | NA | |
| 126 | hsa-miR-582-3p | NOG | 0.01 | 0.97713 | -1.05 | 0.08272 | PITA; miRNATAP | -0.49 | 0 | NA | |
| 127 | hsa-miR-21-5p | PITX2 | 2.74 | 0 | -2.23 | 0 | miRNATAP | -0.41 | 6.0E-5 | NA | |
| 128 | hsa-miR-320a | PITX2 | 0.44 | 0.03902 | -2.23 | 0 | miRanda | -0.35 | 0.00105 | NA | |
| 129 | hsa-miR-324-5p | PITX2 | 2.96 | 0 | -2.23 | 0 | miRanda | -0.3 | 1.0E-5 | NA | |
| 130 | hsa-miR-590-5p | PITX2 | 3.18 | 0 | -2.23 | 0 | PITA; miRanda; miRNATAP | -0.35 | 0 | NA | |
| 131 | hsa-miR-132-3p | SMAD3 | 0.16 | 0.40819 | -0.15 | 0.56924 | mirMAP | -0.31 | 0 | NA | |
| 132 | hsa-miR-134-5p | SMAD6 | 0.14 | 0.65533 | -0.42 | 0.263 | MirTarget | -0.31 | 0 | NA | |
| 133 | hsa-miR-340-5p | SMAD6 | 0.3 | 0.15774 | -0.42 | 0.263 | mirMAP | -0.49 | 0 | NA | |
| 134 | hsa-miR-15b-5p | SMAD7 | 1.57 | 0 | -1.19 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.31 | 0 | NA | |
| 135 | hsa-miR-25-3p | SMAD7 | 1.36 | 0 | -1.19 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.32 | 0 | 23435373 | MicroRNA 25 functions as a potential tumor suppressor in colon cancer by targeting Smad7; Furthermore bioinformatic predictions and experimental validation were used to identify Smad7 as a direct target of miR-25; Functional reverse experiments indicated that the antitumor effects of miR-25 were probably mediated by its repression of Smad7; These results suggest that miR-25 may function as a tumor suppressor by targeting Smad7 in colon cancer |
| 136 | hsa-let-7a-3p | SMAD9 | 1.42 | 0 | -2.78 | 0 | mirMAP | -0.45 | 7.0E-5 | NA | |
| 137 | hsa-let-7b-3p | SMAD9 | 0.87 | 1.0E-5 | -2.78 | 0 | mirMAP | -0.33 | 0.00484 | NA | |
| 138 | hsa-let-7f-1-3p | SMAD9 | 1.55 | 0 | -2.78 | 0 | mirMAP | -0.56 | 0 | NA | |
| 139 | hsa-miR-103a-2-5p | SMAD9 | 2.48 | 0 | -2.78 | 0 | mirMAP | -0.41 | 0 | NA | |
| 140 | hsa-miR-142-5p | SMAD9 | 1.18 | 0.00182 | -2.78 | 0 | mirMAP | -0.4 | 0 | NA | |
| 141 | hsa-miR-148b-5p | SMAD9 | 2.18 | 0 | -2.78 | 0 | MirTarget | -0.36 | 1.0E-5 | NA | |
| 142 | hsa-miR-155-5p | SMAD9 | 1.2 | 0.00086 | -2.78 | 0 | mirMAP | -0.43 | 0 | NA | |
| 143 | hsa-miR-16-2-3p | SMAD9 | 2.32 | 0 | -2.78 | 0 | mirMAP | -0.61 | 0 | NA | |
| 144 | hsa-miR-17-5p | SMAD9 | 3.27 | 0 | -2.78 | 0 | mirMAP | -0.42 | 0 | NA | |
| 145 | hsa-miR-186-5p | SMAD9 | 1.47 | 0 | -2.78 | 0 | mirMAP | -0.58 | 0 | NA | |
| 146 | hsa-miR-193a-5p | SMAD9 | 0.56 | 0.02393 | -2.78 | 0 | MirTarget; TargetScan; miRNATAP | -0.36 | 0.00012 | NA | |
| 147 | hsa-miR-20a-5p | SMAD9 | 3.16 | 0 | -2.78 | 0 | mirMAP | -0.42 | 0 | NA | |
| 148 | hsa-miR-2355-5p | SMAD9 | 1.33 | 0 | -2.78 | 0 | MirTarget | -0.42 | 0 | NA | |
| 149 | hsa-miR-23a-3p | SMAD9 | 1 | 0 | -2.78 | 0 | mirMAP | -0.74 | 0 | NA | |
| 150 | hsa-miR-26a-2-3p | SMAD9 | 0.82 | 0.00017 | -2.78 | 0 | mirMAP | -0.49 | 0 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 26 | 190 | 3.494e-45 | 1.626e-41 |
| 2 | REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 24 | 207 | 1.837e-39 | 4.274e-36 |
| 3 | REGULATION OF PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 18 | 60 | 3.874e-37 | 6.008e-34 |
| 4 | POSITIVE REGULATION OF PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 15 | 48 | 3.969e-31 | 4.617e-28 |
| 5 | POSITIVE REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 17 | 100 | 2.671e-30 | 2.486e-27 |
| 6 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 26 | 689 | 3.494e-30 | 2.71e-27 |
| 7 | RESPONSE TO GROWTH FACTOR | 23 | 475 | 1.157e-28 | 7.689e-26 |
| 8 | RESPONSE TO BMP | 16 | 94 | 1.757e-28 | 9.082e-26 |
| 9 | CELLULAR RESPONSE TO BMP STIMULUS | 16 | 94 | 1.757e-28 | 9.082e-26 |
| 10 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 26 | 983 | 3.533e-26 | 1.644e-23 |
| 11 | SMAD PROTEIN SIGNAL TRANSDUCTION | 13 | 56 | 4.897e-25 | 2.071e-22 |
| 12 | RESPONSE TO ENDOGENOUS STIMULUS | 28 | 1450 | 6.313e-25 | 2.448e-22 |
| 13 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 29 | 1672 | 8.822e-25 | 3.158e-22 |
| 14 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 25 | 1008 | 2.832e-24 | 9.413e-22 |
| 15 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 28 | 1618 | 1.291e-23 | 4.005e-21 |
| 16 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 23 | 801 | 1.912e-23 | 5.56e-21 |
| 17 | REGULATION OF PROTEIN MODIFICATION PROCESS | 28 | 1710 | 5.891e-23 | 1.612e-20 |
| 18 | ACTIVIN RECEPTOR SIGNALING PATHWAY | 10 | 22 | 7.883e-23 | 2.038e-20 |
| 19 | REGULATION OF OSSIFICATION | 15 | 178 | 7.548e-22 | 1.848e-19 |
| 20 | REGULATION OF CELL PROLIFERATION | 26 | 1496 | 1.639e-21 | 3.813e-19 |
| 21 | POSITIVE REGULATION OF GENE EXPRESSION | 27 | 1733 | 2.557e-21 | 5.666e-19 |
| 22 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 27 | 1784 | 5.48e-21 | 1.159e-18 |
| 23 | REGULATION OF OSTEOBLAST DIFFERENTIATION | 13 | 112 | 8.269e-21 | 1.673e-18 |
| 24 | NEGATIVE REGULATION OF CELL PROLIFERATION | 20 | 643 | 9.934e-21 | 1.926e-18 |
| 25 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 27 | 1848 | 1.382e-20 | 2.571e-18 |
| 26 | TUBE DEVELOPMENT | 19 | 552 | 1.957e-20 | 3.502e-18 |
| 27 | REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 15 | 229 | 3.597e-20 | 6.199e-18 |
| 28 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 23 | 1142 | 5.778e-20 | 9.601e-18 |
| 29 | TISSUE DEVELOPMENT | 25 | 1518 | 6.39e-20 | 1.025e-17 |
| 30 | ORGAN MORPHOGENESIS | 21 | 841 | 6.706e-20 | 1.04e-17 |
| 31 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 22 | 1004 | 9.5e-20 | 1.426e-17 |
| 32 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 12 | 95 | 1.079e-19 | 1.569e-17 |
| 33 | NEGATIVE REGULATION OF CELL COMMUNICATION | 23 | 1192 | 1.508e-19 | 2.127e-17 |
| 34 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 26 | 1805 | 1.879e-19 | 2.571e-17 |
| 35 | SKELETAL SYSTEM DEVELOPMENT | 17 | 455 | 9.122e-19 | 1.213e-16 |
| 36 | REGULATION OF CELL DIFFERENTIATION | 24 | 1492 | 1.023e-18 | 1.286e-16 |
| 37 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 24 | 1492 | 1.023e-18 | 1.286e-16 |
| 38 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 22 | 1135 | 1.313e-18 | 1.608e-16 |
| 39 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 21 | 1036 | 4.802e-18 | 5.585e-16 |
| 40 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 21 | 1036 | 4.802e-18 | 5.585e-16 |
| 41 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 23 | 1395 | 5.011e-18 | 5.687e-16 |
| 42 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 19 | 771 | 1.009e-17 | 1.118e-15 |
| 43 | REGULATION OF CELL DEATH | 23 | 1472 | 1.647e-17 | 1.782e-15 |
| 44 | EPITHELIUM DEVELOPMENT | 20 | 945 | 1.875e-17 | 1.983e-15 |
| 45 | RESPONSE TO TRANSFORMING GROWTH FACTOR BETA | 12 | 144 | 1.933e-17 | 1.999e-15 |
| 46 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 25 | 1929 | 2.057e-17 | 2.081e-15 |
| 47 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 14 | 285 | 4.711e-17 | 4.664e-15 |
| 48 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 20 | 1021 | 8.385e-17 | 8.128e-15 |
| 49 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 17 | 609 | 1.206e-16 | 1.145e-14 |
| 50 | CELL DEVELOPMENT | 22 | 1426 | 1.662e-16 | 1.547e-14 |
| 51 | NEGATIVE REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 11 | 121 | 1.871e-16 | 1.707e-14 |
| 52 | PATTERN SPECIFICATION PROCESS | 15 | 418 | 3.043e-16 | 2.723e-14 |
| 53 | POSITIVE REGULATION OF OSSIFICATION | 10 | 84 | 3.119e-16 | 2.738e-14 |
| 54 | POSITIVE REGULATION OF CELL COMMUNICATION | 22 | 1532 | 7.503e-16 | 6.348e-14 |
| 55 | CONNECTIVE TISSUE DEVELOPMENT | 12 | 194 | 7.38e-16 | 6.348e-14 |
| 56 | REGULATION OF CELL DEVELOPMENT | 18 | 836 | 1.061e-15 | 8.813e-14 |
| 57 | NEGATIVE REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 10 | 102 | 2.353e-15 | 1.921e-13 |
| 58 | POSITIVE REGULATION OF BIOMINERAL TISSUE DEVELOPMENT | 8 | 38 | 2.865e-15 | 2.298e-13 |
| 59 | GASTRULATION | 11 | 155 | 3.041e-15 | 2.359e-13 |
| 60 | MESODERM MORPHOGENESIS | 9 | 66 | 3.042e-15 | 2.359e-13 |
| 61 | EMBRYO DEVELOPMENT | 18 | 894 | 3.401e-15 | 2.594e-13 |
| 62 | MESODERM DEVELOPMENT | 10 | 118 | 1.056e-14 | 7.922e-13 |
| 63 | TISSUE MORPHOGENESIS | 15 | 533 | 1.091e-14 | 8.06e-13 |
| 64 | REGULATION OF MAPK CASCADE | 16 | 660 | 1.118e-14 | 8.126e-13 |
| 65 | EMBRYONIC MORPHOGENESIS | 15 | 539 | 1.286e-14 | 9.203e-13 |
| 66 | POSITIVE REGULATION OF CELL PROLIFERATION | 17 | 814 | 1.45e-14 | 1.022e-12 |
| 67 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 17 | 823 | 1.736e-14 | 1.205e-12 |
| 68 | PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 6 | 13 | 4.45e-14 | 3.045e-12 |
| 69 | SKELETAL SYSTEM MORPHOGENESIS | 11 | 201 | 5.46e-14 | 3.682e-12 |
| 70 | POSITIVE REGULATION OF CELL DEATH | 15 | 605 | 6.929e-14 | 4.606e-12 |
| 71 | CARTILAGE DEVELOPMENT | 10 | 147 | 9.923e-14 | 6.503e-12 |
| 72 | UROGENITAL SYSTEM DEVELOPMENT | 12 | 299 | 1.327e-13 | 8.46e-12 |
| 73 | GLAND DEVELOPMENT | 13 | 395 | 1.326e-13 | 8.46e-12 |
| 74 | REGULATION OF TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 9 | 99 | 1.365e-13 | 8.467e-12 |
| 75 | REGULATION OF CELLULAR RESPONSE TO TRANSFORMING GROWTH FACTOR BETA STIMULUS | 9 | 99 | 1.365e-13 | 8.467e-12 |
| 76 | EMBRYONIC ORGAN DEVELOPMENT | 13 | 406 | 1.882e-13 | 1.152e-11 |
| 77 | REPRODUCTIVE SYSTEM DEVELOPMENT | 13 | 408 | 2.004e-13 | 1.211e-11 |
| 78 | GROWTH | 13 | 410 | 2.132e-13 | 1.272e-11 |
| 79 | REGULATION OF CARTILAGE DEVELOPMENT | 8 | 63 | 2.197e-13 | 1.294e-11 |
| 80 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 19 | 1360 | 3.158e-13 | 1.837e-11 |
| 81 | FORMATION OF PRIMARY GERM LAYER | 9 | 110 | 3.61e-13 | 2.074e-11 |
| 82 | REGULATION OF STEM CELL DIFFERENTIATION | 9 | 113 | 4.624e-13 | 2.624e-11 |
| 83 | EMBRYONIC SKELETAL SYSTEM DEVELOPMENT | 9 | 122 | 9.34e-13 | 5.219e-11 |
| 84 | REGULATION OF BIOMINERAL TISSUE DEVELOPMENT | 8 | 75 | 9.422e-13 | 5.219e-11 |
| 85 | HEART DEVELOPMENT | 13 | 466 | 1.082e-12 | 5.922e-11 |
| 86 | VASCULATURE DEVELOPMENT | 13 | 469 | 1.173e-12 | 6.348e-11 |
| 87 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 14 | 602 | 1.392e-12 | 7.446e-11 |
| 88 | EMBRYONIC ORGAN MORPHOGENESIS | 11 | 279 | 1.982e-12 | 1.048e-10 |
| 89 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 15 | 788 | 3.129e-12 | 1.618e-10 |
| 90 | CIRCULATORY SYSTEM DEVELOPMENT | 15 | 788 | 3.129e-12 | 1.618e-10 |
| 91 | MESONEPHROS DEVELOPMENT | 8 | 90 | 4.246e-12 | 2.171e-10 |
| 92 | DIGESTIVE SYSTEM DEVELOPMENT | 9 | 148 | 5.428e-12 | 2.745e-10 |
| 93 | EMBRYONIC SKELETAL SYSTEM MORPHOGENESIS | 8 | 93 | 5.557e-12 | 2.78e-10 |
| 94 | REGIONALIZATION | 11 | 311 | 6.43e-12 | 3.183e-10 |
| 95 | GLAND MORPHOGENESIS | 8 | 97 | 7.844e-12 | 3.802e-10 |
| 96 | POSITIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 9 | 154 | 7.78e-12 | 3.802e-10 |
| 97 | HEAD DEVELOPMENT | 14 | 709 | 1.251e-11 | 6e-10 |
| 98 | POSITIVE REGULATION OF OSTEOBLAST DIFFERENTIATION | 7 | 60 | 1.462e-11 | 6.94e-10 |
| 99 | REPRODUCTION | 17 | 1297 | 2.631e-11 | 1.236e-09 |
| 100 | NEGATIVE REGULATION OF CELLULAR RESPONSE TO TRANSFORMING GROWTH FACTOR BETA STIMULUS | 7 | 66 | 2.925e-11 | 1.347e-09 |
| 101 | NEGATIVE REGULATION OF TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 7 | 66 | 2.925e-11 | 1.347e-09 |
| 102 | REGULATION OF EPITHELIAL TO MESENCHYMAL TRANSITION | 7 | 67 | 3.261e-11 | 1.488e-09 |
| 103 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 11 | 363 | 3.394e-11 | 1.533e-09 |
| 104 | SENSORY ORGAN DEVELOPMENT | 12 | 493 | 4.664e-11 | 2.087e-09 |
| 105 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 15 | 957 | 4.931e-11 | 2.185e-09 |
| 106 | MESENCHYME DEVELOPMENT | 9 | 190 | 5.154e-11 | 2.262e-09 |
| 107 | KIDNEY EPITHELIUM DEVELOPMENT | 8 | 125 | 6.157e-11 | 2.677e-09 |
| 108 | REGULATION OF BMP SIGNALING PATHWAY | 7 | 77 | 8.901e-11 | 3.835e-09 |
| 109 | MORPHOGENESIS OF AN EPITHELIUM | 11 | 400 | 9.576e-11 | 4.062e-09 |
| 110 | NEGATIVE REGULATION OF OSTEOBLAST DIFFERENTIATION | 6 | 40 | 9.602e-11 | 4.062e-09 |
| 111 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 18 | 1656 | 1.172e-10 | 4.915e-09 |
| 112 | NEGATIVE REGULATION OF CELL DEVELOPMENT | 10 | 303 | 1.333e-10 | 5.537e-09 |
| 113 | HEART MORPHOGENESIS | 9 | 212 | 1.371e-10 | 5.547e-09 |
| 114 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 12 | 541 | 1.36e-10 | 5.547e-09 |
| 115 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 12 | 541 | 1.36e-10 | 5.547e-09 |
| 116 | POSITIVE REGULATION OF LOCOMOTION | 11 | 420 | 1.609e-10 | 6.456e-09 |
| 117 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 12 | 554 | 1.786e-10 | 7.105e-09 |
| 118 | CARDIAC SEPTUM DEVELOPMENT | 7 | 85 | 1.808e-10 | 7.128e-09 |
| 119 | LUNG MORPHOGENESIS | 6 | 45 | 2.024e-10 | 7.914e-09 |
| 120 | EMBRYONIC CRANIAL SKELETON MORPHOGENESIS | 6 | 46 | 2.325e-10 | 9.014e-09 |
| 121 | RESPONSE TO LIPID | 14 | 888 | 2.46e-10 | 9.459e-09 |
| 122 | TUBE MORPHOGENESIS | 10 | 323 | 2.487e-10 | 9.485e-09 |
| 123 | EYE DEVELOPMENT | 10 | 326 | 2.721e-10 | 1.029e-08 |
| 124 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 8 | 153 | 3.121e-10 | 1.171e-08 |
| 125 | REGULATION OF VASCULATURE DEVELOPMENT | 9 | 233 | 3.173e-10 | 1.181e-08 |
| 126 | CELL CYCLE ARREST | 8 | 154 | 3.288e-10 | 1.214e-08 |
| 127 | NEGATIVE REGULATION OF GROWTH | 9 | 236 | 3.554e-10 | 1.302e-08 |
| 128 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 14 | 917 | 3.747e-10 | 1.362e-08 |
| 129 | LOCOMOTION | 15 | 1114 | 4.128e-10 | 1.489e-08 |
| 130 | PROTEIN PHOSPHORYLATION | 14 | 944 | 5.473e-10 | 1.959e-08 |
| 131 | POSITIVE REGULATION OF CELL DEVELOPMENT | 11 | 472 | 5.537e-10 | 1.967e-08 |
| 132 | CRANIAL SKELETAL SYSTEM DEVELOPMENT | 6 | 55 | 7.109e-10 | 2.506e-08 |
| 133 | NEUROGENESIS | 16 | 1402 | 9.823e-10 | 3.436e-08 |
| 134 | POSITIVE REGULATION OF CARTILAGE DEVELOPMENT | 5 | 29 | 1.881e-09 | 6.531e-08 |
| 135 | SIGNAL TRANSDUCTION BY PROTEIN PHOSPHORYLATION | 10 | 404 | 2.168e-09 | 7.472e-08 |
| 136 | RESPIRATORY SYSTEM DEVELOPMENT | 8 | 197 | 2.323e-09 | 7.948e-08 |
| 137 | NEGATIVE REGULATION OF CELL DEATH | 13 | 872 | 2.536e-09 | 8.612e-08 |
| 138 | RHYTHMIC PROCESS | 9 | 298 | 2.769e-09 | 9.335e-08 |
| 139 | NEGATIVE REGULATION OF OSSIFICATION | 6 | 69 | 2.885e-09 | 9.659e-08 |
| 140 | NEGATIVE REGULATION OF PHOSPHORYLATION | 10 | 422 | 3.294e-09 | 1.095e-07 |
| 141 | RESPONSE TO HORMONE | 13 | 893 | 3.379e-09 | 1.115e-07 |
| 142 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 14 | 1087 | 3.402e-09 | 1.115e-07 |
| 143 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 7 | 129 | 3.455e-09 | 1.124e-07 |
| 144 | NEGATIVE REGULATION OF CELL CYCLE | 10 | 433 | 4.214e-09 | 1.362e-07 |
| 145 | MESENCHYMAL CELL DIFFERENTIATION | 7 | 134 | 4.507e-09 | 1.446e-07 |
| 146 | NEGATIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 12 | 740 | 4.766e-09 | 1.519e-07 |
| 147 | ARTERY DEVELOPMENT | 6 | 75 | 4.808e-09 | 1.522e-07 |
| 148 | POSITIVE REGULATION OF SMAD PROTEIN IMPORT INTO NUCLEUS | 4 | 12 | 4.854e-09 | 1.526e-07 |
| 149 | IMMUNE SYSTEM DEVELOPMENT | 11 | 582 | 4.962e-09 | 1.549e-07 |
| 150 | RESPONSE TO EXTERNAL STIMULUS | 17 | 1821 | 5.101e-09 | 1.582e-07 |
| 151 | REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 8 | 218 | 5.155e-09 | 1.589e-07 |
| 152 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 12 | 750 | 5.539e-09 | 1.696e-07 |
| 153 | REGULATION OF HOMEOSTATIC PROCESS | 10 | 447 | 5.711e-09 | 1.737e-07 |
| 154 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 14 | 1152 | 7.164e-09 | 2.151e-07 |
| 155 | NEGATIVE REGULATION OF VASCULATURE DEVELOPMENT | 6 | 80 | 7.128e-09 | 2.151e-07 |
| 156 | CARDIAC CHAMBER DEVELOPMENT | 7 | 144 | 7.444e-09 | 2.22e-07 |
| 157 | REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 9 | 337 | 8.083e-09 | 2.396e-07 |
| 158 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 11 | 616 | 8.936e-09 | 2.632e-07 |
| 159 | REGULATION OF CELL ADHESION | 11 | 629 | 1.109e-08 | 3.245e-07 |
| 160 | REGULATION OF ORGAN MORPHOGENESIS | 8 | 242 | 1.167e-08 | 3.395e-07 |
| 161 | OVULATION CYCLE PROCESS | 6 | 88 | 1.272e-08 | 3.676e-07 |
| 162 | BONE DEVELOPMENT | 7 | 156 | 1.298e-08 | 3.728e-07 |
| 163 | CELL DEATH | 13 | 1001 | 1.328e-08 | 3.791e-07 |
| 164 | OSSIFICATION | 8 | 251 | 1.552e-08 | 4.403e-07 |
| 165 | BLOOD VESSEL MORPHOGENESIS | 9 | 364 | 1.575e-08 | 4.442e-07 |
| 166 | PHOSPHORYLATION | 14 | 1228 | 1.616e-08 | 4.53e-07 |
| 167 | REGULATION OF SMAD PROTEIN IMPORT INTO NUCLEUS | 4 | 16 | 1.775e-08 | 4.946e-07 |
| 168 | NEGATIVE REGULATION OF IMMUNE SYSTEM PROCESS | 9 | 372 | 1.9e-08 | 5.264e-07 |
| 169 | REGULATION OF EPITHELIAL CELL MIGRATION | 7 | 166 | 1.996e-08 | 5.495e-07 |
| 170 | POSITIVE REGULATION OF CELL ADHESION | 9 | 376 | 2.084e-08 | 5.671e-07 |
| 171 | MORPHOGENESIS OF A BRANCHING STRUCTURE | 7 | 167 | 2.08e-08 | 5.671e-07 |
| 172 | REGULATION OF CHONDROCYTE DIFFERENTIATION | 5 | 46 | 2.122e-08 | 5.742e-07 |
| 173 | NEGATIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 8 | 262 | 2.166e-08 | 5.827e-07 |
| 174 | NEGATIVE REGULATION OF LOCOMOTION | 8 | 263 | 2.232e-08 | 5.967e-07 |
| 175 | NEGATIVE REGULATION OF GENE EXPRESSION | 15 | 1493 | 2.259e-08 | 6.007e-07 |
| 176 | SEX DIFFERENTIATION | 8 | 266 | 2.437e-08 | 6.439e-07 |
| 177 | POSITIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 7 | 171 | 2.45e-08 | 6.439e-07 |
| 178 | DIGESTIVE TRACT MORPHOGENESIS | 5 | 48 | 2.644e-08 | 6.912e-07 |
| 179 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 15 | 1517 | 2.798e-08 | 7.274e-07 |
| 180 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 12 | 872 | 2.959e-08 | 7.649e-07 |
| 181 | NEURON DIFFERENTIATION | 12 | 874 | 3.035e-08 | 7.801e-07 |
| 182 | POSITIVE REGULATION OF EPITHELIAL CELL MIGRATION | 6 | 103 | 3.289e-08 | 8.409e-07 |
| 183 | POSITIVE REGULATION OF PROTEIN IMPORT | 6 | 104 | 3.486e-08 | 8.863e-07 |
| 184 | ODONTOGENESIS | 6 | 105 | 3.692e-08 | 9.337e-07 |
| 185 | REGULATION OF PROTEIN IMPORT | 7 | 183 | 3.908e-08 | 9.83e-07 |
| 186 | REGULATION OF BINDING | 8 | 283 | 3.938e-08 | 9.852e-07 |
| 187 | REGULATION OF NEURON DIFFERENTIATION | 10 | 554 | 4.357e-08 | 1.084e-06 |
| 188 | VENTRICULAR SEPTUM DEVELOPMENT | 5 | 54 | 4.845e-08 | 1.199e-06 |
| 189 | STEM CELL DIFFERENTIATION | 7 | 190 | 5.058e-08 | 1.245e-06 |
| 190 | ANGIOGENESIS | 8 | 293 | 5.15e-08 | 1.261e-06 |
| 191 | OVULATION CYCLE | 6 | 113 | 5.735e-08 | 1.397e-06 |
| 192 | ANTERIOR POSTERIOR PATTERN SPECIFICATION | 7 | 194 | 5.835e-08 | 1.414e-06 |
| 193 | SPECIFICATION OF SYMMETRY | 6 | 117 | 7.062e-08 | 1.7e-06 |
| 194 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 14 | 1381 | 7.088e-08 | 1.7e-06 |
| 195 | REGULATION OF PROTEIN LOCALIZATION | 12 | 950 | 7.582e-08 | 1.809e-06 |
| 196 | IN UTERO EMBRYONIC DEVELOPMENT | 8 | 311 | 8.151e-08 | 1.935e-06 |
| 197 | MULTICELLULAR ORGANISM REPRODUCTION | 11 | 768 | 8.545e-08 | 2.018e-06 |
| 198 | REGULATION OF IMMUNE SYSTEM PROCESS | 14 | 1403 | 8.633e-08 | 2.019e-06 |
| 199 | POSITIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 6 | 121 | 8.632e-08 | 2.019e-06 |
| 200 | REGULATION OF HEMOPOIESIS | 8 | 314 | 8.775e-08 | 2.041e-06 |
| 201 | ENDOCRINE SYSTEM DEVELOPMENT | 6 | 123 | 9.518e-08 | 2.203e-06 |
| 202 | NEGATIVE REGULATION OF CYTOKINE PRODUCTION | 7 | 211 | 1.036e-07 | 2.387e-06 |
| 203 | REGULATION OF ACTIVIN RECEPTOR SIGNALING PATHWAY | 4 | 25 | 1.219e-07 | 2.781e-06 |
| 204 | DEVELOPMENT OF PRIMARY SEXUAL CHARACTERISTICS | 7 | 216 | 1.216e-07 | 2.781e-06 |
| 205 | REGULATION OF REPRODUCTIVE PROCESS | 6 | 129 | 1.264e-07 | 2.869e-06 |
| 206 | REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 7 | 220 | 1.378e-07 | 3.112e-06 |
| 207 | BRANCHING MORPHOGENESIS OF AN EPITHELIAL TUBE | 6 | 131 | 1.385e-07 | 3.113e-06 |
| 208 | NEGATIVE REGULATION OF GLIAL CELL DIFFERENTIATION | 4 | 26 | 1.439e-07 | 3.204e-06 |
| 209 | NEGATIVE REGULATION OF CARTILAGE DEVELOPMENT | 4 | 26 | 1.439e-07 | 3.204e-06 |
| 210 | REGULATION OF GROWTH | 10 | 633 | 1.512e-07 | 3.35e-06 |
| 211 | RESPONSE TO ABIOTIC STIMULUS | 12 | 1024 | 1.715e-07 | 3.782e-06 |
| 212 | GASTRULATION WITH MOUTH FORMING SECOND | 4 | 28 | 1.966e-07 | 4.314e-06 |
| 213 | REGULATION OF LEUKOCYTE DIFFERENTIATION | 7 | 232 | 1.977e-07 | 4.318e-06 |
| 214 | EPITHELIAL CELL DIFFERENTIATION | 9 | 495 | 2.176e-07 | 4.732e-06 |
| 215 | RESPONSE TO STEROID HORMONE | 9 | 497 | 2.251e-07 | 4.873e-06 |
| 216 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 13 | 1275 | 2.277e-07 | 4.906e-06 |
| 217 | REGULATION OF CELLULAR LOCALIZATION | 13 | 1277 | 2.319e-07 | 4.973e-06 |
| 218 | POSITIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 8 | 360 | 2.496e-07 | 5.327e-06 |
| 219 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 9 | 513 | 2.94e-07 | 6.247e-06 |
| 220 | BONE MORPHOGENESIS | 5 | 79 | 3.339e-07 | 7.062e-06 |
| 221 | SALIVARY GLAND DEVELOPMENT | 4 | 32 | 3.434e-07 | 7.23e-06 |
| 222 | POSITIVE REGULATION OF MYELOID CELL DIFFERENTIATION | 5 | 81 | 3.786e-07 | 7.935e-06 |
| 223 | EMBRYONIC DIGESTIVE TRACT DEVELOPMENT | 4 | 33 | 3.903e-07 | 8.107e-06 |
| 224 | PROTEIN LOCALIZATION TO NUCLEUS | 6 | 156 | 3.893e-07 | 8.107e-06 |
| 225 | POSITIVE REGULATION OF EPITHELIAL TO MESENCHYMAL TRANSITION | 4 | 34 | 4.417e-07 | 9.054e-06 |
| 226 | REGULATION OF MESENCHYMAL CELL PROLIFERATION | 4 | 34 | 4.417e-07 | 9.054e-06 |
| 227 | HEART VALVE DEVELOPMENT | 4 | 34 | 4.417e-07 | 9.054e-06 |
| 228 | POSITIVE REGULATION OF HEMOPOIESIS | 6 | 163 | 5.04e-07 | 1.029e-05 |
| 229 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 9 | 552 | 5.436e-07 | 1.1e-05 |
| 230 | REGULATION OF CELL MORPHOGENESIS | 9 | 552 | 5.436e-07 | 1.1e-05 |
| 231 | EPITHELIAL CELL PROLIFERATION | 5 | 89 | 6.068e-07 | 1.222e-05 |
| 232 | MUSCLE TISSUE DEVELOPMENT | 7 | 275 | 6.229e-07 | 1.239e-05 |
| 233 | APPENDAGE DEVELOPMENT | 6 | 169 | 6.232e-07 | 1.239e-05 |
| 234 | LIMB DEVELOPMENT | 6 | 169 | 6.232e-07 | 1.239e-05 |
| 235 | NEGATIVE REGULATION OF GLIOGENESIS | 4 | 37 | 6.266e-07 | 1.241e-05 |
| 236 | REGULATION OF CYTOKINE PRODUCTION | 9 | 563 | 6.409e-07 | 1.264e-05 |
| 237 | DORSAL VENTRAL PATTERN FORMATION | 5 | 91 | 6.781e-07 | 1.331e-05 |
| 238 | REGULATION OF TRANSFERASE ACTIVITY | 11 | 946 | 6.89e-07 | 1.341e-05 |
| 239 | ACTIVATION OF PROTEIN KINASE ACTIVITY | 7 | 279 | 6.863e-07 | 1.341e-05 |
| 240 | REGULATION OF CELL CYCLE | 11 | 949 | 7.109e-07 | 1.378e-05 |
| 241 | REGULATION OF CYTOKINE BIOSYNTHETIC PROCESS | 5 | 94 | 7.972e-07 | 1.539e-05 |
| 242 | NEGATIVE REGULATION OF PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 3 | 11 | 9.519e-07 | 1.83e-05 |
| 243 | REGULATION OF KINASE ACTIVITY | 10 | 776 | 9.811e-07 | 1.863e-05 |
| 244 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 5 | 98 | 9.81e-07 | 1.863e-05 |
| 245 | REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 5 | 98 | 9.81e-07 | 1.863e-05 |
| 246 | REGULATION OF MYELOID CELL DIFFERENTIATION | 6 | 183 | 9.929e-07 | 1.878e-05 |
| 247 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 5 | 99 | 1.032e-06 | 1.944e-05 |
| 248 | NEGATIVE REGULATION OF BMP SIGNALING PATHWAY | 4 | 42 | 1.055e-06 | 1.979e-05 |
| 249 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 8 | 437 | 1.079e-06 | 2.016e-05 |
| 250 | REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 5 | 100 | 1.085e-06 | 2.019e-05 |
| 251 | RESPONSE TO EXTRACELLULAR STIMULUS | 8 | 441 | 1.155e-06 | 2.142e-05 |
| 252 | NEGATIVE REGULATION OF STEM CELL DIFFERENTIATION | 4 | 43 | 1.162e-06 | 2.145e-05 |
| 253 | MYELOID CELL DIFFERENTIATION | 6 | 189 | 1.199e-06 | 2.204e-05 |
| 254 | REGULATION OF PROTEIN TARGETING | 7 | 307 | 1.301e-06 | 2.383e-05 |
| 255 | EXOCRINE SYSTEM DEVELOPMENT | 4 | 45 | 1.399e-06 | 2.542e-05 |
| 256 | ENDOCHONDRAL BONE MORPHOGENESIS | 4 | 45 | 1.399e-06 | 2.542e-05 |
| 257 | CARDIAC VENTRICLE DEVELOPMENT | 5 | 106 | 1.448e-06 | 2.622e-05 |
| 258 | REGULATION OF MYELOID LEUKOCYTE DIFFERENTIATION | 5 | 108 | 1.589e-06 | 2.865e-05 |
| 259 | NEGATIVE REGULATION OF OLIGODENDROCYTE DIFFERENTIATION | 3 | 13 | 1.646e-06 | 2.945e-05 |
| 260 | REGULATION OF CELL PROLIFERATION INVOLVED IN KIDNEY DEVELOPMENT | 3 | 13 | 1.646e-06 | 2.945e-05 |
| 261 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 6 | 200 | 1.666e-06 | 2.97e-05 |
| 262 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 8 | 465 | 1.717e-06 | 3.048e-05 |
| 263 | POSITIVE REGULATION OF MAPK CASCADE | 8 | 470 | 1.859e-06 | 3.275e-05 |
| 264 | WOUND HEALING | 8 | 470 | 1.859e-06 | 3.275e-05 |
| 265 | REGULATION OF TRANSPORT | 14 | 1804 | 1.865e-06 | 3.275e-05 |
| 266 | CELL MOTILITY | 10 | 835 | 1.903e-06 | 3.316e-05 |
| 267 | LOCALIZATION OF CELL | 10 | 835 | 1.903e-06 | 3.316e-05 |
| 268 | CARDIAC SEPTUM MORPHOGENESIS | 4 | 49 | 1.978e-06 | 3.435e-05 |
| 269 | REGULATION OF GLOMERULUS DEVELOPMENT | 3 | 14 | 2.092e-06 | 3.592e-05 |
| 270 | EMBRYONIC SKELETAL JOINT DEVELOPMENT | 3 | 14 | 2.092e-06 | 3.592e-05 |
| 271 | RESPONSE TO LAMINAR FLUID SHEAR STRESS | 3 | 14 | 2.092e-06 | 3.592e-05 |
| 272 | POSITIVE REGULATION OF STEM CELL DIFFERENTIATION | 4 | 50 | 2.148e-06 | 3.674e-05 |
| 273 | NEGATIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 5 | 115 | 2.167e-06 | 3.694e-05 |
| 274 | FEMALE SEX DIFFERENTIATION | 5 | 116 | 2.262e-06 | 3.841e-05 |
| 275 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 6 | 213 | 2.4e-06 | 4.061e-05 |
| 276 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 11 | 1079 | 2.502e-06 | 4.218e-05 |
| 277 | CELL CYCLE PROCESS | 11 | 1081 | 2.547e-06 | 4.279e-05 |
| 278 | POSITIVE REGULATION OF HOMEOSTATIC PROCESS | 6 | 216 | 2.602e-06 | 4.356e-05 |
| 279 | REGULATION OF MORPHOGENESIS OF A BRANCHING STRUCTURE | 4 | 53 | 2.72e-06 | 4.52e-05 |
| 280 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 4 | 53 | 2.72e-06 | 4.52e-05 |
| 281 | CELL PROLIFERATION | 9 | 672 | 2.768e-06 | 4.584e-05 |
| 282 | REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 5 | 122 | 2.9e-06 | 4.785e-05 |
| 283 | NEGATIVE REGULATION OF REPRODUCTIVE PROCESS | 4 | 54 | 2.934e-06 | 4.807e-05 |
| 284 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 10 | 876 | 2.925e-06 | 4.807e-05 |
| 285 | REGULATION OF KIDNEY DEVELOPMENT | 4 | 55 | 3.16e-06 | 5.158e-05 |
| 286 | REGULATION OF MONONUCLEAR CELL MIGRATION | 3 | 16 | 3.21e-06 | 5.205e-05 |
| 287 | NEGATIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 9 | 684 | 3.2e-06 | 5.205e-05 |
| 288 | EPITHELIAL TO MESENCHYMAL TRANSITION | 4 | 56 | 3.398e-06 | 5.481e-05 |
| 289 | MULTI ORGANISM REPRODUCTIVE PROCESS | 10 | 891 | 3.404e-06 | 5.481e-05 |
| 290 | CELL FATE COMMITMENT | 6 | 227 | 3.467e-06 | 5.562e-05 |
| 291 | FOREBRAIN DEVELOPMENT | 7 | 357 | 3.535e-06 | 5.652e-05 |
| 292 | POSITIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 8 | 514 | 3.609e-06 | 5.751e-05 |
| 293 | CELLULAR COMPONENT MORPHOGENESIS | 10 | 900 | 3.723e-06 | 5.913e-05 |
| 294 | EMBRYONIC PATTERN SPECIFICATION | 4 | 58 | 3.915e-06 | 6.196e-05 |
| 295 | POSITIVE REGULATION OF PROTEOLYSIS | 7 | 363 | 3.945e-06 | 6.222e-05 |
| 296 | REGULATION OF GLIAL CELL DIFFERENTIATION | 4 | 59 | 4.194e-06 | 6.593e-05 |
| 297 | REGULATION OF LYMPHOCYTE DIFFERENTIATION | 5 | 132 | 4.27e-06 | 6.689e-05 |
| 298 | POSITIVE REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 4 | 60 | 4.488e-06 | 6.984e-05 |
| 299 | CHONDROCYTE DIFFERENTIATION | 4 | 60 | 4.488e-06 | 6.984e-05 |
| 300 | OVARIAN FOLLICLE DEVELOPMENT | 4 | 61 | 4.796e-06 | 7.415e-05 |
| 301 | POSITIVE REGULATION OF STEM CELL PROLIFERATION | 4 | 61 | 4.796e-06 | 7.415e-05 |
| 302 | POSITIVE REGULATION OF CELL CELL ADHESION | 6 | 243 | 5.129e-06 | 7.902e-05 |
| 303 | POSITIVE REGULATION OF TRANSPORT | 10 | 936 | 5.275e-06 | 8.101e-05 |
| 304 | REGULATION OF CELL CELL ADHESION | 7 | 380 | 5.327e-06 | 8.154e-05 |
| 305 | REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 7 | 381 | 5.42e-06 | 8.268e-05 |
| 306 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 14 | 1977 | 5.52e-06 | 8.387e-05 |
| 307 | POSITIVE REGULATION OF CHONDROCYTE DIFFERENTIATION | 3 | 19 | 5.534e-06 | 8.387e-05 |
| 308 | CARDIAC MUSCLE TISSUE DEVELOPMENT | 5 | 140 | 5.695e-06 | 8.603e-05 |
| 309 | REGULATION OF PROTEIN SECRETION | 7 | 389 | 6.209e-06 | 9.35e-05 |
| 310 | CELLULAR PROCESS INVOLVED IN REPRODUCTION IN MULTICELLULAR ORGANISM | 6 | 252 | 6.318e-06 | 9.482e-05 |
| 311 | LENS DEVELOPMENT IN CAMERA TYPE EYE | 4 | 66 | 6.581e-06 | 9.846e-05 |
| 312 | REGULATION OF PEPTIDE TRANSPORT | 6 | 256 | 6.913e-06 | 0.0001031 |
| 313 | NEURON FATE COMMITMENT | 4 | 67 | 6.989e-06 | 0.0001039 |
| 314 | RESPONSE TO WOUNDING | 8 | 563 | 7.047e-06 | 0.0001044 |
| 315 | POSITIVE REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 6 | 258 | 7.228e-06 | 0.0001068 |
| 316 | POSITIVE REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 4 | 68 | 7.416e-06 | 0.0001092 |
| 317 | MALE SEX DIFFERENTIATION | 5 | 148 | 7.468e-06 | 0.0001096 |
| 318 | CHONDROCYTE DEVELOPMENT | 3 | 21 | 7.576e-06 | 0.0001108 |
| 319 | REGULATION OF HORMONE SECRETION | 6 | 262 | 7.891e-06 | 0.0001151 |
| 320 | REGULATION OF CELLULAR COMPONENT BIOGENESIS | 9 | 767 | 8.105e-06 | 0.0001178 |
| 321 | POSITIVE REGULATION OF CELLULAR COMPONENT BIOGENESIS | 7 | 406 | 8.209e-06 | 0.000119 |
| 322 | REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 5 | 152 | 8.503e-06 | 0.0001229 |
| 323 | NEGATIVE REGULATION OF CYTOKINE PRODUCTION INVOLVED IN IMMUNE RESPONSE | 3 | 22 | 8.761e-06 | 0.0001262 |
| 324 | ENDODERM DEVELOPMENT | 4 | 71 | 8.813e-06 | 0.0001266 |
| 325 | AMEBOIDAL TYPE CELL MIGRATION | 5 | 154 | 9.061e-06 | 0.0001293 |
| 326 | POSITIVE REGULATION OF PEPTIDASE ACTIVITY | 5 | 154 | 9.061e-06 | 0.0001293 |
| 327 | REGULATION OF CELL DIVISION | 6 | 272 | 9.77e-06 | 0.000139 |
| 328 | POSITIVE REGULATION OF ERYTHROCYTE DIFFERENTIATION | 3 | 23 | 1.006e-05 | 0.0001419 |
| 329 | PROSTATE GLAND MORPHOGENESIS | 3 | 23 | 1.006e-05 | 0.0001419 |
| 330 | NEGATIVE REGULATION OF EPITHELIAL TO MESENCHYMAL TRANSITION | 3 | 23 | 1.006e-05 | 0.0001419 |
| 331 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 13 | 1791 | 1.041e-05 | 0.0001459 |
| 332 | NEGATIVE REGULATION OF HORMONE SECRETION | 4 | 74 | 1.04e-05 | 0.0001459 |
| 333 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 12 | 1518 | 1.083e-05 | 0.0001514 |
| 334 | RESPONSE TO STEROL | 3 | 24 | 1.148e-05 | 0.00016 |
| 335 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 5 | 162 | 1.159e-05 | 0.0001609 |
| 336 | POSITIVE REGULATION OF CYTOPLASMIC TRANSPORT | 6 | 282 | 1.2e-05 | 0.0001661 |
| 337 | REGULATION OF TRANSFORMING GROWTH FACTOR BETA PRODUCTION | 3 | 25 | 1.303e-05 | 0.00018 |
| 338 | MESODERMAL CELL DIFFERENTIATION | 3 | 26 | 1.472e-05 | 0.0002026 |
| 339 | NEGATIVE REGULATION OF LEUKOCYTE DIFFERENTIATION | 4 | 82 | 1.564e-05 | 0.0002146 |
| 340 | POSITIVE REGULATION OF RESPONSE TO EXTERNAL STIMULUS | 6 | 296 | 1.579e-05 | 0.000216 |
| 341 | REGULATION OF ENDOTHELIAL CELL DIFFERENTIATION | 3 | 27 | 1.653e-05 | 0.0002243 |
| 342 | REGULATION OF ASTROCYTE DIFFERENTIATION | 3 | 27 | 1.653e-05 | 0.0002243 |
| 343 | POSITIVE REGULATION OF MESENCHYMAL CELL PROLIFERATION | 3 | 27 | 1.653e-05 | 0.0002243 |
| 344 | CELL CYCLE | 11 | 1316 | 1.669e-05 | 0.0002258 |
| 345 | NEGATIVE REGULATION OF TRANSPORT | 7 | 458 | 1.792e-05 | 0.0002417 |
| 346 | PALATE DEVELOPMENT | 4 | 85 | 1.803e-05 | 0.0002418 |
| 347 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 6 | 303 | 1.802e-05 | 0.0002418 |
| 348 | VENTRICULAR SEPTUM MORPHOGENESIS | 3 | 28 | 1.849e-05 | 0.0002473 |
| 349 | POSITIVE REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 4 | 86 | 1.888e-05 | 0.0002518 |
| 350 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 6 | 306 | 1.905e-05 | 0.0002532 |
| 351 | TAXIS | 7 | 464 | 1.948e-05 | 0.0002583 |
| 352 | TISSUE REMODELING | 4 | 87 | 1.977e-05 | 0.0002613 |
| 353 | MYELOID CELL HOMEOSTASIS | 4 | 88 | 2.068e-05 | 0.0002696 |
| 354 | NEGATIVE REGULATION OF PRODUCTION OF MOLECULAR MEDIATOR OF IMMUNE RESPONSE | 3 | 29 | 2.06e-05 | 0.0002696 |
| 355 | REGULATION OF HEART MORPHOGENESIS | 3 | 29 | 2.06e-05 | 0.0002696 |
| 356 | REGULATION OF OLIGODENDROCYTE DIFFERENTIATION | 3 | 29 | 2.06e-05 | 0.0002696 |
| 357 | REGULATION OF STEM CELL PROLIFERATION | 4 | 88 | 2.068e-05 | 0.0002696 |
| 358 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 7 | 470 | 2.116e-05 | 0.000275 |
| 359 | POSITIVE REGULATION OF IMMUNE SYSTEM PROCESS | 9 | 867 | 2.157e-05 | 0.0002796 |
| 360 | POSITIVE REGULATION OF DNA METABOLIC PROCESS | 5 | 185 | 2.199e-05 | 0.0002843 |
| 361 | AXIS SPECIFICATION | 4 | 90 | 2.26e-05 | 0.0002897 |
| 362 | ENDOTHELIUM DEVELOPMENT | 4 | 90 | 2.26e-05 | 0.0002897 |
| 363 | REGULATION OF GLIOGENESIS | 4 | 90 | 2.26e-05 | 0.0002897 |
| 364 | SMOOTH MUSCLE CELL DIFFERENTIATION | 3 | 30 | 2.286e-05 | 0.0002922 |
| 365 | REGULATION OF HORMONE LEVELS | 7 | 478 | 2.358e-05 | 0.0003006 |
| 366 | REGULATION OF MAP KINASE ACTIVITY | 6 | 319 | 2.407e-05 | 0.000306 |
| 367 | REGULATION OF CYTOPLASMIC TRANSPORT | 7 | 481 | 2.454e-05 | 0.0003112 |
| 368 | POSITIVE REGULATION OF KINASE ACTIVITY | 7 | 482 | 2.487e-05 | 0.0003145 |
| 369 | ENDOCARDIAL CUSHION DEVELOPMENT | 3 | 32 | 2.786e-05 | 0.0003494 |
| 370 | PATTERNING OF BLOOD VESSELS | 3 | 32 | 2.786e-05 | 0.0003494 |
| 371 | POSITIVE REGULATION OF EPIDERMIS DEVELOPMENT | 3 | 32 | 2.786e-05 | 0.0003494 |
| 372 | IMMUNE SYSTEM PROCESS | 13 | 1984 | 3.135e-05 | 0.0003921 |
| 373 | NEGATIVE REGULATION OF SECRETION | 5 | 200 | 3.196e-05 | 0.0003987 |
| 374 | RESPONSE TO FLUID SHEAR STRESS | 3 | 34 | 3.353e-05 | 0.0004159 |
| 375 | ORGAN FORMATION | 3 | 34 | 3.353e-05 | 0.0004159 |
| 376 | REGULATION OF SECRETION | 8 | 699 | 3.361e-05 | 0.0004159 |
| 377 | REGULATION OF DNA METABOLIC PROCESS | 6 | 340 | 3.442e-05 | 0.0004248 |
| 378 | REGULATION OF PRODUCTION OF MOLECULAR MEDIATOR OF IMMUNE RESPONSE | 4 | 101 | 3.56e-05 | 0.0004382 |
| 379 | REGULATION OF PROTEOLYSIS | 8 | 711 | 3.794e-05 | 0.0004657 |
| 380 | REGULATION OF MUSCLE TISSUE DEVELOPMENT | 4 | 103 | 3.845e-05 | 0.0004695 |
| 381 | REGULATION OF MUSCLE ORGAN DEVELOPMENT | 4 | 103 | 3.845e-05 | 0.0004695 |
| 382 | CARDIAC CHAMBER MORPHOGENESIS | 4 | 104 | 3.993e-05 | 0.0004789 |
| 383 | NEGATIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 5 | 209 | 3.944e-05 | 0.0004789 |
| 384 | REGULATION OF ERYTHROCYTE DIFFERENTIATION | 3 | 36 | 3.99e-05 | 0.0004789 |
| 385 | NEGATIVE REGULATION OF MUSCLE ORGAN DEVELOPMENT | 3 | 36 | 3.99e-05 | 0.0004789 |
| 386 | NEGATIVE REGULATION OF MUSCLE TISSUE DEVELOPMENT | 3 | 36 | 3.99e-05 | 0.0004789 |
| 387 | REGULATION OF PEPTIDE SECRETION | 5 | 209 | 3.944e-05 | 0.0004789 |
| 388 | HEAD MORPHOGENESIS | 3 | 36 | 3.99e-05 | 0.0004789 |
| 389 | REGULATION OF FAT CELL DIFFERENTIATION | 4 | 106 | 4.303e-05 | 0.0005133 |
| 390 | FAT CELL DIFFERENTIATION | 4 | 106 | 4.303e-05 | 0.0005133 |
| 391 | MULTI MULTICELLULAR ORGANISM PROCESS | 5 | 213 | 4.317e-05 | 0.0005138 |
| 392 | NEGATIVE REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 3 | 37 | 4.337e-05 | 0.0005147 |
| 393 | PLATELET DEGRANULATION | 4 | 107 | 4.464e-05 | 0.0005285 |
| 394 | NEGATIVE REGULATION OF PROTEIN SECRETION | 4 | 108 | 4.629e-05 | 0.0005467 |
| 395 | MESENCHYME MORPHOGENESIS | 3 | 38 | 4.702e-05 | 0.0005539 |
| 396 | RESPONSE TO ESTROGEN | 5 | 218 | 4.821e-05 | 0.0005665 |
| 397 | REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 6 | 365 | 5.11e-05 | 0.000599 |
| 398 | CELL CYCLE G1 S PHASE TRANSITION | 4 | 111 | 5.153e-05 | 0.0006009 |
| 399 | G1 S TRANSITION OF MITOTIC CELL CYCLE | 4 | 111 | 5.153e-05 | 0.0006009 |
| 400 | POSITIVE REGULATION OF INTRACELLULAR TRANSPORT | 6 | 370 | 5.511e-05 | 0.0006379 |
| 401 | POSITIVE REGULATION OF SECRETION | 6 | 370 | 5.511e-05 | 0.0006379 |
| 402 | NEGATIVE REGULATION OF LYMPHOCYTE DIFFERENTIATION | 3 | 40 | 5.493e-05 | 0.0006379 |
| 403 | REGULATION OF ENDOTHELIAL CELL MIGRATION | 4 | 114 | 5.719e-05 | 0.0006603 |
| 404 | NEGATIVE REGULATION OF FAT CELL DIFFERENTIATION | 3 | 41 | 5.919e-05 | 0.0006751 |
| 405 | POSITIVE REGULATION OF KIDNEY DEVELOPMENT | 3 | 41 | 5.919e-05 | 0.0006751 |
| 406 | PROSTATE GLAND DEVELOPMENT | 3 | 41 | 5.919e-05 | 0.0006751 |
| 407 | NEPHRON DEVELOPMENT | 4 | 115 | 5.917e-05 | 0.0006751 |
| 408 | LUNG ALVEOLUS DEVELOPMENT | 3 | 41 | 5.919e-05 | 0.0006751 |
| 409 | REGULATION OF CELL PROJECTION ORGANIZATION | 7 | 558 | 6.304e-05 | 0.0007165 |
| 410 | NEGATIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 4 | 117 | 6.329e-05 | 0.0007165 |
| 411 | MAMMARY GLAND DEVELOPMENT | 4 | 117 | 6.329e-05 | 0.0007165 |
| 412 | PITUITARY GLAND DEVELOPMENT | 3 | 42 | 6.366e-05 | 0.000719 |
| 413 | POSITIVE REGULATION OF CHEMOTAXIS | 4 | 120 | 6.985e-05 | 0.0007869 |
| 414 | BODY MORPHOGENESIS | 3 | 44 | 7.326e-05 | 0.0008234 |
| 415 | REGULATION OF PEPTIDASE ACTIVITY | 6 | 392 | 7.585e-05 | 0.0008505 |
| 416 | REGULATION OF ALCOHOL BIOSYNTHETIC PROCESS | 3 | 45 | 7.839e-05 | 0.0008768 |
| 417 | PEPTIDYL THREONINE MODIFICATION | 3 | 46 | 8.376e-05 | 0.0009323 |
| 418 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 3 | 46 | 8.376e-05 | 0.0009323 |
| 419 | BIOLOGICAL ADHESION | 9 | 1032 | 8.408e-05 | 0.0009337 |
| 420 | OSTEOBLAST DIFFERENTIATION | 4 | 126 | 8.443e-05 | 0.0009354 |
| 421 | REGULATION OF PHOSPHATASE ACTIVITY | 4 | 128 | 8.976e-05 | 0.0009897 |
| 422 | NEGATIVE REGULATION OF HEMOPOIESIS | 4 | 128 | 8.976e-05 | 0.0009897 |
| 423 | NUCLEAR IMPORT | 4 | 129 | 9.251e-05 | 0.001018 |
| 424 | REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 3 | 48 | 9.519e-05 | 0.001045 |
| 425 | POSITIVE REGULATION OF LEUKOCYTE DIFFERENTIATION | 4 | 131 | 9.819e-05 | 0.001075 |
| 426 | CELL CYCLE PHASE TRANSITION | 5 | 255 | 0.0001012 | 0.001101 |
| 427 | NEGATIVE REGULATION OF PEPTIDE SECRETION | 3 | 49 | 0.0001013 | 0.001101 |
| 428 | REGULATION OF STEROID BIOSYNTHETIC PROCESS | 3 | 49 | 0.0001013 | 0.001101 |
| 429 | FACE DEVELOPMENT | 3 | 50 | 0.0001076 | 0.001159 |
| 430 | RESPONSE TO PROGESTERONE | 3 | 50 | 0.0001076 | 0.001159 |
| 431 | ENDODERM FORMATION | 3 | 50 | 0.0001076 | 0.001159 |
| 432 | POSITIVE REGULATION OF MYELOID LEUKOCYTE DIFFERENTIATION | 3 | 50 | 0.0001076 | 0.001159 |
| 433 | CELL GROWTH | 4 | 135 | 0.0001103 | 0.001185 |
| 434 | NEGATIVE REGULATION OF CATALYTIC ACTIVITY | 8 | 829 | 0.0001116 | 0.001196 |
| 435 | NEGATIVE REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 4 | 136 | 0.0001135 | 0.001214 |
| 436 | ARTERY MORPHOGENESIS | 3 | 51 | 0.0001142 | 0.001218 |
| 437 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 7 | 616 | 0.000117 | 0.001246 |
| 438 | REGULATION OF INTRACELLULAR TRANSPORT | 7 | 621 | 0.000123 | 0.001307 |
| 439 | RESPONSE TO DRUG | 6 | 431 | 0.0001276 | 0.001353 |
| 440 | MESONEPHRIC TUBULE MORPHOGENESIS | 3 | 53 | 0.0001281 | 0.001355 |
| 441 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 6 | 437 | 0.0001376 | 0.001452 |
| 442 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 3 | 55 | 0.0001431 | 0.001506 |
| 443 | REGULATION OF CYTOKINE PRODUCTION INVOLVED IN IMMUNE RESPONSE | 3 | 57 | 0.0001592 | 0.001672 |
| 444 | NEURAL TUBE DEVELOPMENT | 4 | 149 | 0.0001614 | 0.001687 |
| 445 | REGULATION OF LEUKOCYTE MIGRATION | 4 | 149 | 0.0001614 | 0.001687 |
| 446 | PROTEOGLYCAN BIOSYNTHETIC PROCESS | 3 | 58 | 0.0001677 | 0.001745 |
| 447 | POSITIVE REGULATION OF CYTOKINE BIOSYNTHETIC PROCESS | 3 | 58 | 0.0001677 | 0.001745 |
| 448 | NEGATIVE REGULATION OF MULTI ORGANISM PROCESS | 4 | 151 | 0.0001699 | 0.001764 |
| 449 | CELLULAR RESPONSE TO LIPID | 6 | 457 | 0.0001755 | 0.001819 |
| 450 | REGULATION OF REMOVAL OF SUPEROXIDE RADICALS | 2 | 11 | 0.0001812 | 0.00187 |
| 451 | APOPTOTIC SIGNALING PATHWAY | 5 | 289 | 0.0001816 | 0.00187 |
| 452 | POSITIVE REGULATION OF HAIR CYCLE | 2 | 11 | 0.0001812 | 0.00187 |
| 453 | NEGATIVE REGULATION OF CELL DIVISION | 3 | 60 | 0.0001855 | 0.001897 |
| 454 | REGULATION OF MONOOXYGENASE ACTIVITY | 3 | 60 | 0.0001855 | 0.001897 |
| 455 | STEM CELL PROLIFERATION | 3 | 60 | 0.0001855 | 0.001897 |
| 456 | PROTEIN IMPORT | 4 | 155 | 0.0001878 | 0.001916 |
| 457 | LEUKOCYTE DIFFERENTIATION | 5 | 292 | 0.0001905 | 0.00194 |
| 458 | REGULATION OF DEPHOSPHORYLATION | 4 | 158 | 0.0002021 | 0.002053 |
| 459 | REGULATION OF MULTI ORGANISM PROCESS | 6 | 470 | 0.0002043 | 0.002068 |
| 460 | EMBRYONIC HEART TUBE MORPHOGENESIS | 3 | 62 | 0.0002045 | 0.002068 |
| 461 | REGULATION OF EPIDERMIS DEVELOPMENT | 3 | 63 | 0.0002144 | 0.00216 |
| 462 | NEGATIVE REGULATION OF CARDIAC MUSCLE TISSUE GROWTH | 2 | 12 | 0.0002172 | 0.00216 |
| 463 | REGULATION OF VITAMIN METABOLIC PROCESS | 2 | 12 | 0.0002172 | 0.00216 |
| 464 | POSITIVE REGULATION OF DNA DEPENDENT DNA REPLICATION | 2 | 12 | 0.0002172 | 0.00216 |
| 465 | REGULATION OF MACROPHAGE CYTOKINE PRODUCTION | 2 | 12 | 0.0002172 | 0.00216 |
| 466 | REGULATION OF CHEMOKINE BIOSYNTHETIC PROCESS | 2 | 12 | 0.0002172 | 0.00216 |
| 467 | TRACHEA MORPHOGENESIS | 2 | 12 | 0.0002172 | 0.00216 |
| 468 | NEGATIVE REGULATION OF HEART GROWTH | 2 | 12 | 0.0002172 | 0.00216 |
| 469 | EXTRACELLULAR STRUCTURE ORGANIZATION | 5 | 304 | 0.0002297 | 0.002278 |
| 470 | REGULATION OF CELL ACTIVATION | 6 | 484 | 0.0002394 | 0.00237 |
| 471 | REGULATION OF RESPONSE TO STRESS | 10 | 1468 | 0.0002447 | 0.002417 |
| 472 | REGULATION OF PROTEIN BINDING | 4 | 168 | 0.0002556 | 0.002486 |
| 473 | TYPE B PANCREATIC CELL DEVELOPMENT | 2 | 13 | 0.0002564 | 0.002486 |
| 474 | NEGATIVE REGULATION OF ASTROCYTE DIFFERENTIATION | 2 | 13 | 0.0002564 | 0.002486 |
| 475 | POSITIVE REGULATION OF STEROID BIOSYNTHETIC PROCESS | 2 | 13 | 0.0002564 | 0.002486 |
| 476 | BEHAVIORAL RESPONSE TO PAIN | 2 | 13 | 0.0002564 | 0.002486 |
| 477 | RESPONSE TO OXYGEN LEVELS | 5 | 311 | 0.0002551 | 0.002486 |
| 478 | MESENCHYMAL CELL PROLIFERATION | 2 | 13 | 0.0002564 | 0.002486 |
| 479 | MESODERMAL CELL FATE COMMITMENT | 2 | 13 | 0.0002564 | 0.002486 |
| 480 | REGULATION OF GONADOTROPIN SECRETION | 2 | 13 | 0.0002564 | 0.002486 |
| 481 | POSITIVE REGULATION OF ENDOTHELIAL CELL MIGRATION | 3 | 67 | 0.0002573 | 0.002489 |
| 482 | NEGATIVE REGULATION OF CELL GROWTH | 4 | 170 | 0.0002674 | 0.002581 |
| 483 | REGULATION OF CELL JUNCTION ASSEMBLY | 3 | 68 | 0.0002689 | 0.00259 |
| 484 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 2 | 14 | 0.0002988 | 0.002861 |
| 485 | HOMEOSTASIS OF NUMBER OF CELLS | 4 | 175 | 0.0002986 | 0.002861 |
| 486 | CRANIOFACIAL SUTURE MORPHOGENESIS | 2 | 14 | 0.0002988 | 0.002861 |
| 487 | REGULATION OF SYSTEM PROCESS | 6 | 507 | 0.0003071 | 0.002934 |
| 488 | ENDOTHELIAL CELL DIFFERENTIATION | 3 | 72 | 0.0003184 | 0.003035 |
| 489 | CELLULAR MACROMOLECULE LOCALIZATION | 9 | 1234 | 0.0003246 | 0.003088 |
| 490 | ERYTHROCYTE HOMEOSTASIS | 3 | 73 | 0.0003316 | 0.003142 |
| 491 | EMBRYONIC HEART TUBE DEVELOPMENT | 3 | 73 | 0.0003316 | 0.003142 |
| 492 | REGULATION OF CHEMOTAXIS | 4 | 180 | 0.0003323 | 0.003143 |
| 493 | REGULATION OF HAIR FOLLICLE DEVELOPMENT | 2 | 15 | 0.0003444 | 0.003218 |
| 494 | MORPHOGENESIS OF AN EPITHELIAL FOLD | 2 | 15 | 0.0003444 | 0.003218 |
| 495 | CHRONIC INFLAMMATORY RESPONSE | 2 | 15 | 0.0003444 | 0.003218 |
| 496 | POSITIVE REGULATION OF ENDOTHELIAL CELL APOPTOTIC PROCESS | 2 | 15 | 0.0003444 | 0.003218 |
| 497 | POSITIVE REGULATION OF MEMBRANE PROTEIN ECTODOMAIN PROTEOLYSIS | 2 | 15 | 0.0003444 | 0.003218 |
| 498 | VENOUS BLOOD VESSEL DEVELOPMENT | 2 | 15 | 0.0003444 | 0.003218 |
| 499 | REGULATION OF STEROID METABOLIC PROCESS | 3 | 74 | 0.0003451 | 0.003218 |
| 500 | RESPONSE TO KETONE | 4 | 182 | 0.0003465 | 0.003225 |
| 501 | DEVELOPMENTAL GROWTH | 5 | 333 | 0.0003494 | 0.003245 |
| 502 | ODONTOGENESIS OF DENTIN CONTAINING TOOTH | 3 | 75 | 0.0003591 | 0.003328 |
| 503 | CELL JUNCTION ORGANIZATION | 4 | 185 | 0.0003687 | 0.00341 |
| 504 | DIENCEPHALON DEVELOPMENT | 3 | 77 | 0.000388 | 0.003582 |
| 505 | MODULATION OF GROWTH OF SYMBIONT INVOLVED IN INTERACTION WITH HOST | 2 | 16 | 0.0003931 | 0.003615 |
| 506 | POSITIVE REGULATION OF TRANSFORMING GROWTH FACTOR BETA PRODUCTION | 2 | 16 | 0.0003931 | 0.003615 |
| 507 | RENAL TUBULE DEVELOPMENT | 3 | 78 | 0.000403 | 0.003699 |
| 508 | CELLULAR RESPONSE TO STRESS | 10 | 1565 | 0.00041 | 0.003756 |
| 509 | NEGATIVE REGULATION OF NEURON DIFFERENTIATION | 4 | 191 | 0.000416 | 0.003796 |
| 510 | RESPONSE TO NUTRIENT | 4 | 191 | 0.000416 | 0.003796 |
| 511 | NEGATIVE REGULATION OF KIDNEY DEVELOPMENT | 2 | 17 | 0.000445 | 0.004005 |
| 512 | EMBRYONIC DIGESTIVE TRACT MORPHOGENESIS | 2 | 17 | 0.000445 | 0.004005 |
| 513 | POSITIVE REGULATION OF EXTRACELLULAR MATRIX ORGANIZATION | 2 | 17 | 0.000445 | 0.004005 |
| 514 | NEGATIVE REGULATION OF TRANSFERASE ACTIVITY | 5 | 351 | 0.0004446 | 0.004005 |
| 515 | REGULATION OF STEM CELL POPULATION MAINTENANCE | 2 | 17 | 0.000445 | 0.004005 |
| 516 | NEGATIVE REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 2 | 17 | 0.000445 | 0.004005 |
| 517 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 2 | 17 | 0.000445 | 0.004005 |
| 518 | METANEPHROS DEVELOPMENT | 3 | 81 | 0.0004502 | 0.004037 |
| 519 | POSITIVE REGULATION OF PROTEIN KINASE B SIGNALING | 3 | 81 | 0.0004502 | 0.004037 |
| 520 | REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 4 | 197 | 0.0004676 | 0.004176 |
| 521 | KIDNEY MORPHOGENESIS | 3 | 82 | 0.0004668 | 0.004176 |
| 522 | PROTEOGLYCAN METABOLIC PROCESS | 3 | 83 | 0.0004836 | 0.004311 |
| 523 | NEGATIVE REGULATION OF MITOTIC CELL CYCLE | 4 | 199 | 0.0004857 | 0.004321 |
| 524 | REGULATION OF HETEROTYPIC CELL CELL ADHESION | 2 | 18 | 0.0005001 | 0.004412 |
| 525 | NEGATIVE REGULATION OF DEVELOPMENTAL GROWTH | 3 | 84 | 0.0005009 | 0.004412 |
| 526 | GLANDULAR EPITHELIAL CELL DEVELOPMENT | 2 | 18 | 0.0005001 | 0.004412 |
| 527 | PROTEIN LOCALIZATION TO ORGANELLE | 6 | 556 | 0.0005016 | 0.004412 |
| 528 | KIDNEY MESENCHYME DEVELOPMENT | 2 | 18 | 0.0005001 | 0.004412 |
| 529 | PHARYNGEAL SYSTEM DEVELOPMENT | 2 | 18 | 0.0005001 | 0.004412 |
| 530 | RESPONSE TO ALCOHOL | 5 | 362 | 0.0005116 | 0.004491 |
| 531 | POSITIVE REGULATION OF DNA REPLICATION | 3 | 86 | 0.0005366 | 0.004702 |
| 532 | NEURON PROJECTION GUIDANCE | 4 | 205 | 0.0005432 | 0.004751 |
| 533 | CELL MORPHOGENESIS INVOLVED IN NEURON DIFFERENTIATION | 5 | 368 | 0.0005512 | 0.004812 |
| 534 | ENTEROENDOCRINE CELL DIFFERENTIATION | 2 | 19 | 0.0005583 | 0.004855 |
| 535 | ATRIOVENTRICULAR VALVE DEVELOPMENT | 2 | 19 | 0.0005583 | 0.004855 |
| 536 | POSITIVE REGULATION OF CYTOKINE PRODUCTION | 5 | 370 | 0.000565 | 0.004904 |
| 537 | GERM CELL DEVELOPMENT | 4 | 209 | 0.0005841 | 0.005061 |
| 538 | RESPONSE TO MECHANICAL STIMULUS | 4 | 210 | 0.0005946 | 0.005143 |
| 539 | POSITIVE REGULATION OF PROTEIN SECRETION | 4 | 211 | 0.0006053 | 0.005225 |
| 540 | REGULATION OF OXIDOREDUCTASE ACTIVITY | 3 | 90 | 0.0006129 | 0.005281 |
| 541 | NEGATIVE REGULATION OF CELL JUNCTION ASSEMBLY | 2 | 20 | 0.0006196 | 0.0053 |
| 542 | TRACHEA DEVELOPMENT | 2 | 20 | 0.0006196 | 0.0053 |
| 543 | REGULATION OF MACROPHAGE DIFFERENTIATION | 2 | 20 | 0.0006196 | 0.0053 |
| 544 | NEGATIVE REGULATION OF CHONDROCYTE DIFFERENTIATION | 2 | 20 | 0.0006196 | 0.0053 |
| 545 | NEPHRON EPITHELIUM DEVELOPMENT | 3 | 93 | 0.0006744 | 0.005747 |
| 546 | REGULATION OF DNA BINDING | 3 | 93 | 0.0006744 | 0.005747 |
| 547 | RESPONSE TO DIETARY EXCESS | 2 | 21 | 0.000684 | 0.005776 |
| 548 | NEGATIVE REGULATION OF ORGAN GROWTH | 2 | 21 | 0.000684 | 0.005776 |
| 549 | CELL AGGREGATION | 2 | 21 | 0.000684 | 0.005776 |
| 550 | REGULATION OF MEMBRANE PROTEIN ECTODOMAIN PROTEOLYSIS | 2 | 21 | 0.000684 | 0.005776 |
| 551 | CARTILAGE CONDENSATION | 2 | 21 | 0.000684 | 0.005776 |
| 552 | NEURAL TUBE FORMATION | 3 | 94 | 0.0006958 | 0.005865 |
| 553 | GAMETE GENERATION | 6 | 595 | 0.0007163 | 0.006027 |
| 554 | REGULATION OF TRANSCRIPTION FACTOR IMPORT INTO NUCLEUS | 3 | 95 | 0.0007176 | 0.006027 |
| 555 | REGULATION OF CELL GROWTH | 5 | 391 | 0.000725 | 0.006078 |
| 556 | MYELOID LEUKOCYTE DIFFERENTIATION | 3 | 96 | 0.0007398 | 0.00618 |
| 557 | CARDIOCYTE DIFFERENTIATION | 3 | 96 | 0.0007398 | 0.00618 |
| 558 | REGULATION OF HAIR CYCLE | 2 | 22 | 0.0007515 | 0.006256 |
| 559 | REGULATION OF B CELL DIFFERENTIATION | 2 | 22 | 0.0007515 | 0.006256 |
| 560 | REGULATED EXOCYTOSIS | 4 | 224 | 0.0007571 | 0.00629 |
| 561 | HEMATOPOIETIC PROGENITOR CELL DIFFERENTIATION | 3 | 98 | 0.0007855 | 0.006515 |
| 562 | TELENCEPHALON DEVELOPMENT | 4 | 228 | 0.0008087 | 0.006696 |
| 563 | POSITIVE REGULATION OF ALCOHOL BIOSYNTHETIC PROCESS | 2 | 23 | 0.0008221 | 0.006723 |
| 564 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL METABOLIC PROCESS | 2 | 23 | 0.0008221 | 0.006723 |
| 565 | NEURON PROJECTION MORPHOGENESIS | 5 | 402 | 0.0008213 | 0.006723 |
| 566 | REGULATION OF METANEPHROS DEVELOPMENT | 2 | 23 | 0.0008221 | 0.006723 |
| 567 | REGULATION OF SUPEROXIDE METABOLIC PROCESS | 2 | 23 | 0.0008221 | 0.006723 |
| 568 | POSITIVE REGULATION OF COLLAGEN METABOLIC PROCESS | 2 | 23 | 0.0008221 | 0.006723 |
| 569 | POSITIVE REGULATION OF STEROID METABOLIC PROCESS | 2 | 23 | 0.0008221 | 0.006723 |
| 570 | CAMERA TYPE EYE MORPHOGENESIS | 3 | 101 | 0.0008574 | 0.006999 |
| 571 | POSITIVE REGULATION OF NEURON PROJECTION DEVELOPMENT | 4 | 232 | 0.0008628 | 0.007031 |
| 572 | RESPONSE TO NITROGEN COMPOUND | 7 | 859 | 0.0008761 | 0.007127 |
| 573 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 5 | 408 | 0.0008778 | 0.007128 |
| 574 | NEGATIVE REGULATION OF IMMUNE EFFECTOR PROCESS | 3 | 102 | 0.0008822 | 0.007152 |
| 575 | POSITIVE REGULATION OF CELLULAR RESPONSE TO TRANSFORMING GROWTH FACTOR BETA STIMULUS | 2 | 24 | 0.0008958 | 0.007174 |
| 576 | POSITIVE REGULATION OF OSTEOCLAST DIFFERENTIATION | 2 | 24 | 0.0008958 | 0.007174 |
| 577 | POSITIVE REGULATION OF EPITHELIAL CELL APOPTOTIC PROCESS | 2 | 24 | 0.0008958 | 0.007174 |
| 578 | POSITIVE REGULATION OF TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 2 | 24 | 0.0008958 | 0.007174 |
| 579 | NEGATIVE REGULATION OF MYOBLAST DIFFERENTIATION | 2 | 24 | 0.0008958 | 0.007174 |
| 580 | ENDOTHELIAL CELL PROLIFERATION | 2 | 24 | 0.0008958 | 0.007174 |
| 581 | CARDIAC EPITHELIAL TO MESENCHYMAL TRANSITION | 2 | 24 | 0.0008958 | 0.007174 |
| 582 | POSITIVE REGULATION OF GROWTH | 4 | 238 | 0.0009487 | 0.007585 |
| 583 | SENSORY ORGAN MORPHOGENESIS | 4 | 239 | 0.0009636 | 0.007691 |
| 584 | REGULATION OF FIBROBLAST GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 2 | 25 | 0.0009726 | 0.007736 |
| 585 | EPITHELIAL CELL APOPTOTIC PROCESS | 2 | 25 | 0.0009726 | 0.007736 |
| 586 | REGULATION OF T CELL DIFFERENTIATION | 3 | 107 | 0.001013 | 0.008047 |
| 587 | POSITIVE REGULATION OF B CELL MEDIATED IMMUNITY | 2 | 26 | 0.001052 | 0.0083 |
| 588 | REGULATION OF MESONEPHROS DEVELOPMENT | 2 | 26 | 0.001052 | 0.0083 |
| 589 | NEGATIVE REGULATION OF EMBRYONIC DEVELOPMENT | 2 | 26 | 0.001052 | 0.0083 |
| 590 | POSITIVE REGULATION OF IMMUNOGLOBULIN MEDIATED IMMUNE RESPONSE | 2 | 26 | 0.001052 | 0.0083 |
| 591 | POSITIVE REGULATION OF LEUKOCYTE MIGRATION | 3 | 109 | 0.001069 | 0.008417 |
| 592 | MUSCLE STRUCTURE DEVELOPMENT | 5 | 432 | 0.001133 | 0.008893 |
| 593 | POSITIVE REGULATION OF FILOPODIUM ASSEMBLY | 2 | 27 | 0.001135 | 0.008893 |
| 594 | LIPID STORAGE | 2 | 27 | 0.001135 | 0.008893 |
| 595 | NEGATIVE REGULATION OF KINASE ACTIVITY | 4 | 250 | 0.001138 | 0.008902 |
| 596 | REGULATION OF ORGANELLE ORGANIZATION | 8 | 1178 | 0.00118 | 0.009211 |
| 597 | CELL FATE COMMITMENT INVOLVED IN FORMATION OF PRIMARY GERM LAYER | 2 | 28 | 0.001221 | 0.009471 |
| 598 | NEGATIVE REGULATION OF CYTOKINE BIOSYNTHETIC PROCESS | 2 | 28 | 0.001221 | 0.009471 |
| 599 | REGULATION OF EMBRYONIC DEVELOPMENT | 3 | 114 | 0.001217 | 0.009471 |
| 600 | POSITIVE REGULATION OF PHOSPHATASE ACTIVITY | 2 | 28 | 0.001221 | 0.009471 |
| 601 | LEUKOCYTE CELL CELL ADHESION | 4 | 255 | 0.001225 | 0.009481 |
| 602 | PROTEIN LOCALIZATION | 10 | 1805 | 0.001255 | 0.009699 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR BINDING | 16 | 50 | 1.969e-33 | 1.829e-30 |
| 2 | CYTOKINE RECEPTOR BINDING | 18 | 271 | 1.885e-24 | 8.756e-22 |
| 3 | TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE ACTIVITY | 8 | 17 | 1.463e-18 | 4.531e-16 |
| 4 | GROWTH FACTOR ACTIVITY | 12 | 160 | 7.05e-17 | 1.31e-14 |
| 5 | CYTOKINE ACTIVITY | 13 | 219 | 6.425e-17 | 1.31e-14 |
| 6 | RECEPTOR SIGNALING PROTEIN ACTIVITY | 12 | 172 | 1.707e-16 | 2.643e-14 |
| 7 | RECEPTOR SERINE THREONINE KINASE BINDING | 7 | 15 | 2.584e-16 | 3.429e-14 |
| 8 | CYTOKINE BINDING | 9 | 92 | 6.91e-14 | 8.024e-12 |
| 9 | RECEPTOR BINDING | 20 | 1476 | 9.553e-14 | 9.861e-12 |
| 10 | TRANSFORMING GROWTH FACTOR BETA BINDING | 6 | 16 | 2.068e-13 | 1.921e-11 |
| 11 | GROWTH FACTOR BINDING | 9 | 123 | 1.006e-12 | 8.5e-11 |
| 12 | TRANSMEMBRANE RECEPTOR PROTEIN KINASE ACTIVITY | 8 | 81 | 1.782e-12 | 1.38e-10 |
| 13 | RECEPTOR SIGNALING PROTEIN SERINE THREONINE KINASE ACTIVITY | 8 | 92 | 5.085e-12 | 3.634e-10 |
| 14 | SMAD BINDING | 7 | 72 | 5.488e-11 | 3.642e-09 |
| 15 | GLYCOSAMINOGLYCAN BINDING | 8 | 205 | 3.179e-09 | 1.969e-07 |
| 16 | PROTEIN DIMERIZATION ACTIVITY | 14 | 1149 | 6.929e-09 | 4.023e-07 |
| 17 | IDENTICAL PROTEIN BINDING | 14 | 1209 | 1.326e-08 | 7.245e-07 |
| 18 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 9 | 445 | 8.825e-08 | 4.554e-06 |
| 19 | COLLAGEN BINDING | 5 | 65 | 1.247e-07 | 6.097e-06 |
| 20 | HEPARIN BINDING | 6 | 157 | 4.042e-07 | 1.878e-05 |
| 21 | I SMAD BINDING | 3 | 11 | 9.519e-07 | 4.211e-05 |
| 22 | SIGNAL TRANSDUCER ACTIVITY | 14 | 1731 | 1.136e-06 | 4.797e-05 |
| 23 | ACTIVIN BINDING | 3 | 12 | 1.268e-06 | 5.12e-05 |
| 24 | PROTEIN KINASE ACTIVITY | 9 | 640 | 1.855e-06 | 7.179e-05 |
| 25 | SULFUR COMPOUND BINDING | 6 | 234 | 4.129e-06 | 0.0001534 |
| 26 | MACROMOLECULAR COMPLEX BINDING | 12 | 1399 | 4.686e-06 | 0.0001674 |
| 27 | PROTEIN HOMODIMERIZATION ACTIVITY | 9 | 722 | 4.97e-06 | 0.000171 |
| 28 | PROTEIN COMPLEX BINDING | 10 | 935 | 5.226e-06 | 0.0001734 |
| 29 | KINASE ACTIVITY | 9 | 842 | 1.71e-05 | 0.0005479 |
| 30 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 9 | 992 | 6.197e-05 | 0.001919 |
| 31 | EXTRACELLULAR MATRIX BINDING | 3 | 51 | 0.0001142 | 0.003421 |
| 32 | CO SMAD BINDING | 2 | 12 | 0.0002172 | 0.006307 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | EXTRACELLULAR SPACE | 19 | 1376 | 3.892e-13 | 2.273e-10 |
| 2 | EXTRACELLULAR MATRIX | 10 | 426 | 3.605e-09 | 1.053e-06 |
| 3 | PLATELET ALPHA GRANULE | 5 | 75 | 2.571e-07 | 5.005e-05 |
| 4 | RECEPTOR COMPLEX | 7 | 327 | 1.979e-06 | 0.0002889 |
| 5 | PLATELET ALPHA GRANULE LUMEN | 4 | 55 | 3.16e-06 | 0.000369 |
| 6 | PLASMA MEMBRANE RECEPTOR COMPLEX | 5 | 175 | 1.683e-05 | 0.001504 |
| 7 | SECRETORY GRANULE LUMEN | 4 | 85 | 1.803e-05 | 0.001504 |
| 8 | PROTEIN KINASE COMPLEX | 4 | 90 | 2.26e-05 | 0.00165 |
| 9 | PLASMA MEMBRANE PROTEIN COMPLEX | 7 | 510 | 3.567e-05 | 0.002315 |
| 10 | VESICLE LUMEN | 4 | 106 | 4.303e-05 | 0.002513 |
| 11 | CELL SURFACE | 8 | 757 | 5.91e-05 | 0.003138 |
| 12 | EXTRACELLULAR MATRIX COMPONENT | 4 | 125 | 8.186e-05 | 0.003984 |
| 13 | TRANSCRIPTION FACTOR COMPLEX | 5 | 298 | 0.0002094 | 0.009407 |
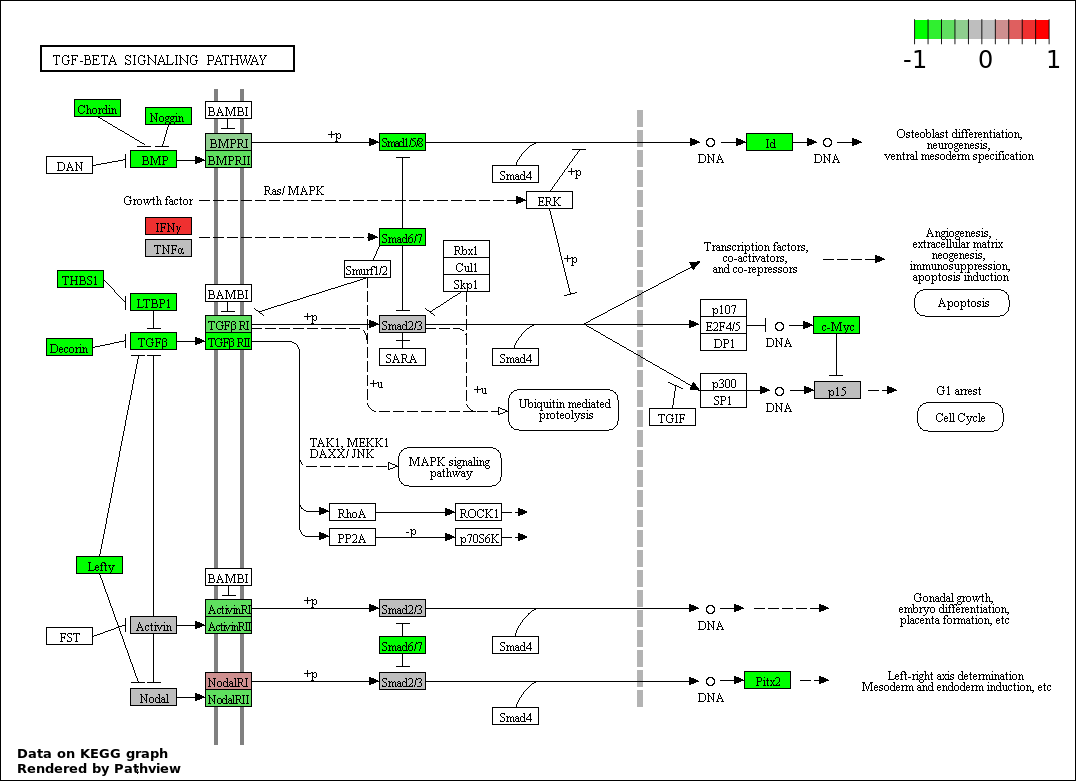
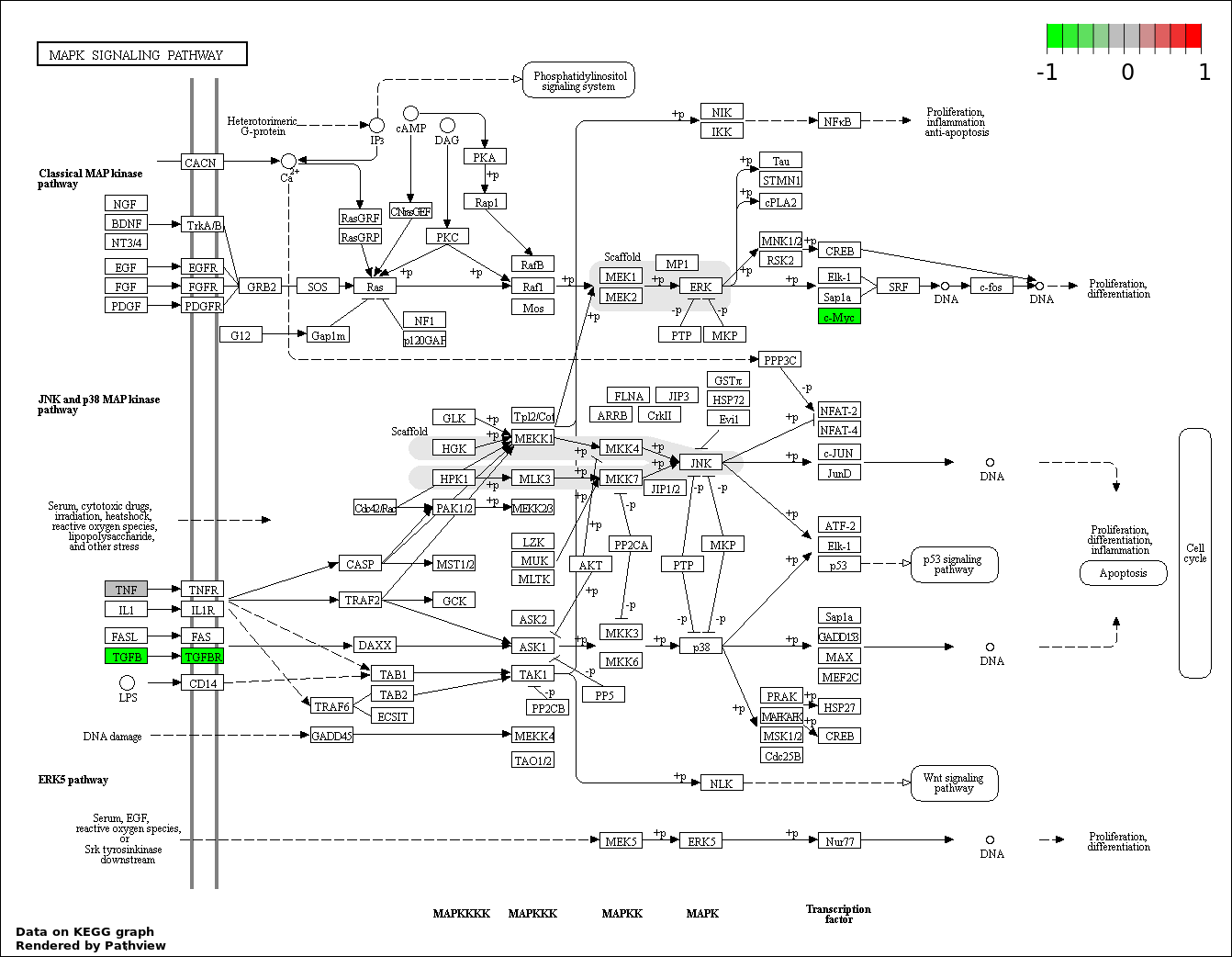
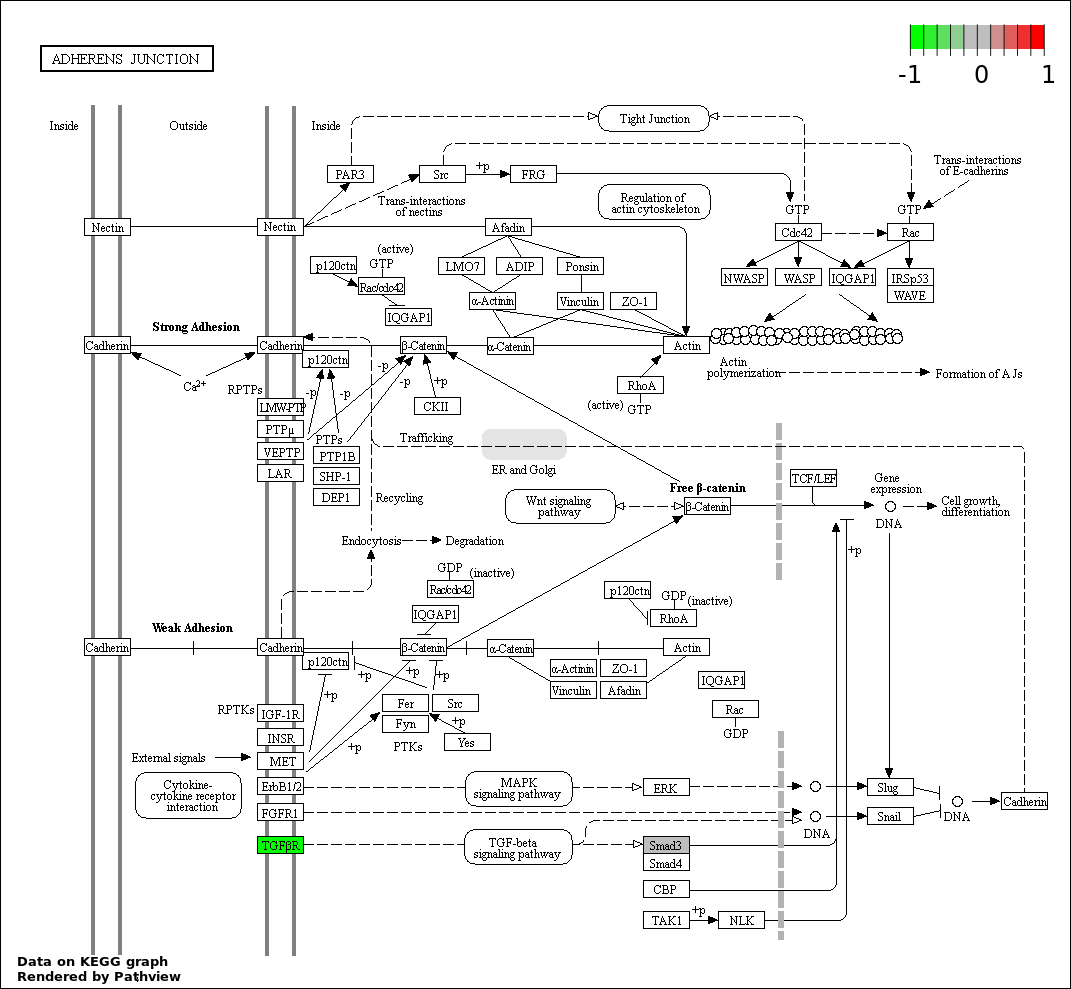
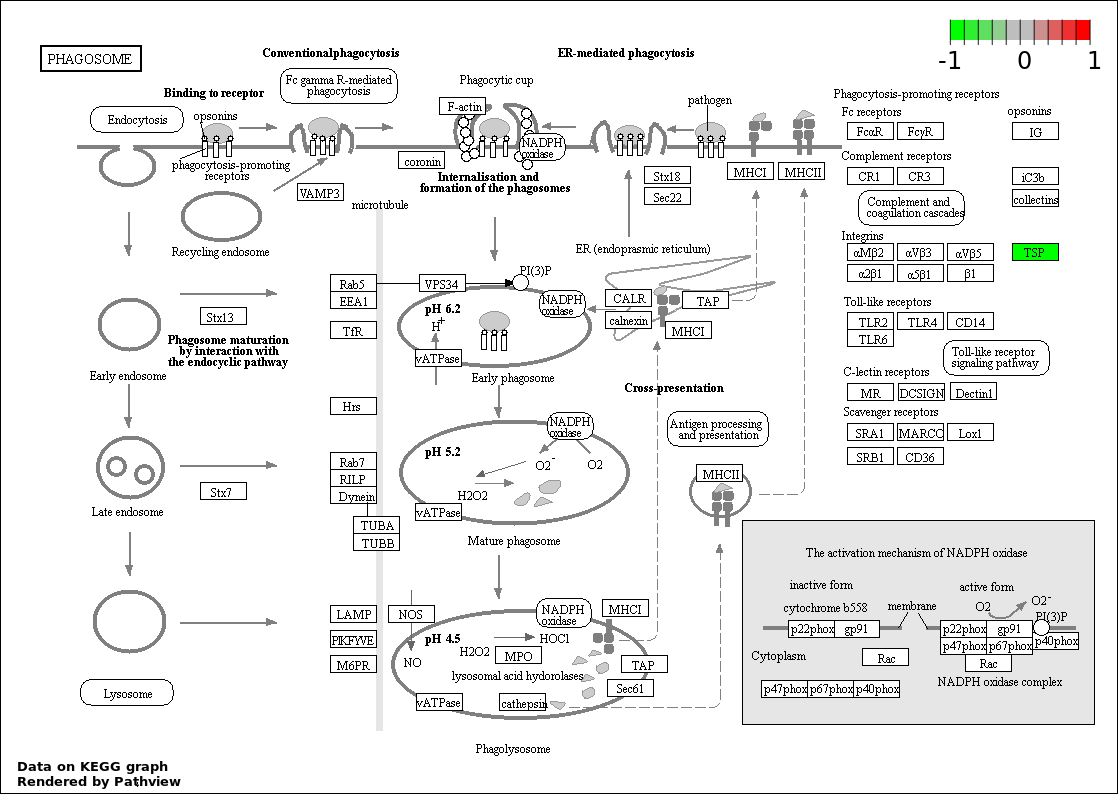
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04350_TGF.beta_signaling_pathway | 37 | 85 | 1.707e-92 | 3.022e-90 | |
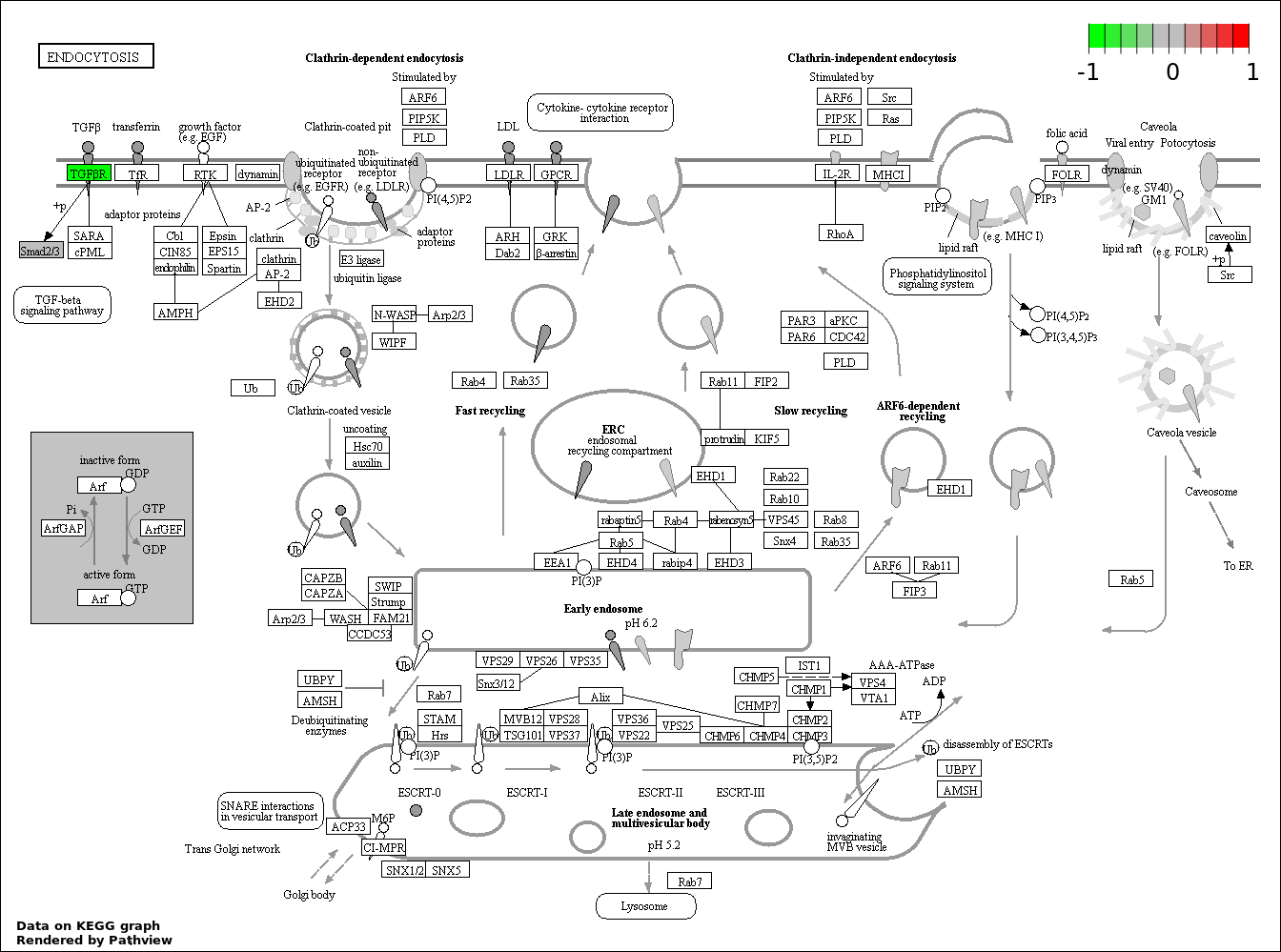
| 2 | hsa04144_Endocytosis | 7 | 203 | 7.959e-08 | 7.044e-06 | |
| 3 | hsa04340_Hedgehog_signaling_pathway | 4 | 56 | 3.398e-06 | 0.00013 | |
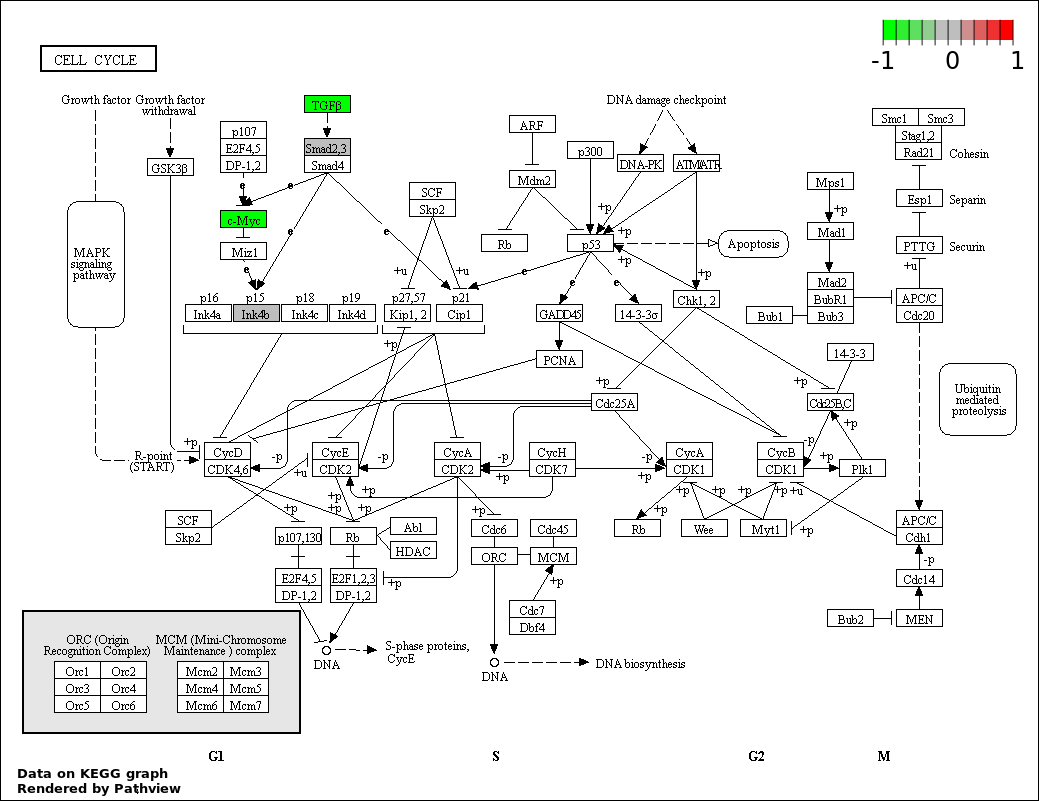
| 4 | hsa04110_Cell_cycle | 5 | 128 | 3.672e-06 | 0.00013 | |
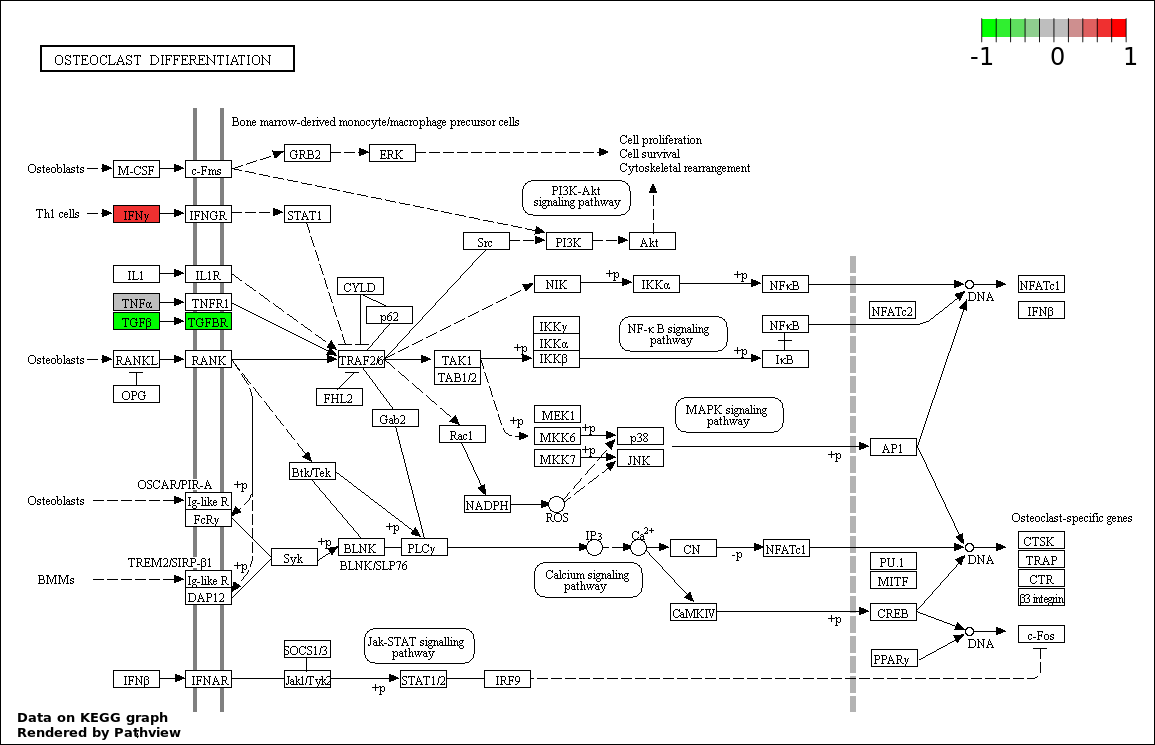
| 5 | hsa04380_Osteoclast_differentiation | 5 | 128 | 3.672e-06 | 0.00013 | |
| 6 | hsa04010_MAPK_signaling_pathway | 6 | 268 | 8.979e-06 | 0.0002649 | |
| 7 | hsa04520_Adherens_junction | 3 | 73 | 0.0003316 | 0.008384 | |
| 8 | hsa04512_ECM.receptor_interaction | 3 | 85 | 0.0005186 | 0.01147 | |
| 9 | hsa04145_Phagosome | 3 | 156 | 0.002978 | 0.05856 | |
| 10 | hsa04510_Focal_adhesion | 3 | 200 | 0.005961 | 0.1055 | |
| 11 | hsa04612_Antigen_processing_and_presentation | 2 | 78 | 0.009155 | 0.1473 | |
| 12 | hsa04660_T_cell_receptor_signaling_pathway | 2 | 108 | 0.01701 | 0.251 | |
| 13 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 2 | 136 | 0.02617 | 0.3564 | |
| 14 | hsa04310_Wnt_signaling_pathway | 2 | 151 | 0.03174 | 0.3929 | |
| 15 | hsa04630_Jak.STAT_signaling_pathway | 2 | 155 | 0.03329 | 0.3929 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | RP11-166D19.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-374b-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 17 | DCN | Sponge network | -3.855 | 0 | -3.453 | 0 | 0.84 |
| 2 | HAND2-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-374b-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-944;hsa-miR-98-5p | 20 | DCN | Sponge network | -5.605 | 0 | -3.453 | 0 | 0.745 |
| 3 | MAGI2-AS3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-98-5p | 12 | DCN | Sponge network | -2.414 | 0 | -3.453 | 0 | 0.738 |
| 4 | MIR143HG |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-374b-3p;hsa-miR-491-5p;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-98-5p | 19 | DCN | Sponge network | -4.237 | 0 | -3.453 | 0 | 0.728 |
| 5 | RP11-166D19.1 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-940 | 29 | THBS1 | Sponge network | -3.855 | 0 | -2.128 | 0 | 0.727 |
| 6 | DNM3OS |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-7-1-3p | 10 | DCN | Sponge network | -2.298 | 1.0E-5 | -3.453 | 0 | 0.715 |
| 7 | RP11-554A11.4 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-421;hsa-miR-491-5p | 12 | DCN | Sponge network | -3.989 | 0 | -3.453 | 0 | 0.705 |
| 8 | LINC00702 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-374b-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 19 | DCN | Sponge network | -2.704 | 0 | -3.453 | 0 | 0.699 |
| 9 | LINC00702 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-940 | 29 | THBS1 | Sponge network | -2.704 | 0 | -2.128 | 0 | 0.679 |
| 10 | RP11-166D19.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-96-5p | 13 | THBS2 | Sponge network | -3.855 | 0 | -0.819 | 0.127 | 0.678 |
| 11 | RP11-536O18.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p | 10 | DCN | Sponge network | -5.37 | 4.0E-5 | -3.453 | 0 | 0.675 |
| 12 | MAGI2-AS3 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-33a-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-98-5p | 21 | THBS1 | Sponge network | -2.414 | 0 | -2.128 | 0 | 0.668 |
| 13 | ADAMTS9-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-374b-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-944;hsa-miR-98-5p | 20 | DCN | Sponge network | -7.614 | 0 | -3.453 | 0 | 0.634 |
| 14 | C20orf166-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-491-5p | 13 | DCN | Sponge network | -6.333 | 0 | -3.453 | 0 | 0.621 |
| 15 | DNM3OS |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-940 | 21 | THBS1 | Sponge network | -2.298 | 1.0E-5 | -2.128 | 0 | 0.61 |
| 16 | DNM3OS |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 13 | THBS2 | Sponge network | -2.298 | 1.0E-5 | -0.819 | 0.127 | 0.607 |
| 17 | GAS6-AS2 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-421;hsa-miR-7-1-3p | 11 | DCN | Sponge network | -2.655 | 0 | -3.453 | 0 | 0.602 |
| 18 | MIR143HG |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-33a-3p;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-940;hsa-miR-98-5p | 29 | THBS1 | Sponge network | -4.237 | 0 | -2.128 | 0 | 0.601 |
| 19 | AC093627.8 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-20a-5p | 10 | THBS1 | Sponge network | -5.744 | 0 | -2.128 | 0 | 0.588 |
| 20 | CTC-296K1.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-374b-3p | 12 | DCN | Sponge network | -5.677 | 0 | -3.453 | 0 | 0.587 |
| 21 | RP11-720L2.4 | hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-491-5p;hsa-miR-940 | 15 | THBS1 | Sponge network | -3.686 | 2.0E-5 | -2.128 | 0 | 0.585 |
| 22 | RP5-1172A22.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b | 12 | DCN | Sponge network | -3.373 | 0.00572 | -3.453 | 0 | 0.58 |
| 23 | ACTA2-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-374b-3p;hsa-miR-491-5p;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 13 | DCN | Sponge network | -3.838 | 0 | -3.453 | 0 | 0.574 |
| 24 | AC093627.8 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p | 10 | DCN | Sponge network | -5.744 | 0 | -3.453 | 0 | 0.571 |
| 25 | RP11-401O9.4 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-590-5p;hsa-miR-671-5p | 14 | THBS1 | Sponge network | -0.359 | 0.78702 | -2.128 | 0 | 0.568 |
| 26 | HAND2-AS1 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-940;hsa-miR-98-5p | 29 | THBS1 | Sponge network | -5.605 | 0 | -2.128 | 0 | 0.565 |
| 27 | RP11-536O18.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p | 11 | THBS1 | Sponge network | -5.37 | 4.0E-5 | -2.128 | 0 | 0.563 |
| 28 | RP11-20J15.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-491-5p;hsa-miR-501-5p | 12 | DCN | Sponge network | -5.104 | 8.0E-5 | -3.453 | 0 | 0.551 |
| 29 | PDZRN3-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-374b-3p;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 12 | DCN | Sponge network | -5.049 | 1.0E-5 | -3.453 | 0 | 0.541 |
| 30 | GAS6-AS2 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-421;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-940 | 21 | THBS1 | Sponge network | -2.655 | 0 | -2.128 | 0 | 0.539 |
| 31 | RP11-753H16.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-320a;hsa-miR-320b | 12 | DCN | Sponge network | -5.702 | 0 | -3.453 | 0 | 0.539 |
| 32 | RP11-750H9.5 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-671-5p;hsa-miR-940 | 10 | THBS1 | Sponge network | -1.207 | 0.04802 | -2.128 | 0 | 0.533 |
| 33 | RP11-867G23.10 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p | 12 | THBS1 | Sponge network | -5.684 | 0 | -2.128 | 0 | 0.533 |
| 34 | RP11-693J15.4 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-374b-3p;hsa-miR-491-5p;hsa-miR-501-5p;hsa-miR-7-1-3p | 12 | DCN | Sponge network | -3.319 | 0.00281 | -3.453 | 0 | 0.529 |
| 35 | RP11-887P2.5 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-374b-3p;hsa-miR-421;hsa-miR-7-1-3p;hsa-miR-944 | 10 | DCN | Sponge network | -6.751 | 0 | -3.453 | 0 | 0.523 |
| 36 | RP11-166D19.1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-590-5p | 15 | TGFB2 | Sponge network | -3.855 | 0 | -1.199 | 0.00331 | 0.52 |
| 37 | RP11-175K6.1 | hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-590-5p;hsa-miR-671-5p;hsa-miR-98-5p | 12 | THBS1 | Sponge network | -2.386 | 0 | -2.128 | 0 | 0.517 |
| 38 | FENDRR |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-374b-3p;hsa-miR-421;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-98-5p | 15 | DCN | Sponge network | -4.793 | 0 | -3.453 | 0 | 0.516 |
| 39 | LINC00702 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-582-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-598-3p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 15 | THBS2 | Sponge network | -2.704 | 0 | -0.819 | 0.127 | 0.511 |
| 40 | RP5-1172A22.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p | 14 | THBS1 | Sponge network | -3.373 | 0.00572 | -2.128 | 0 | 0.505 |
| 41 | LINC00473 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-7-1-3p | 11 | DCN | Sponge network | -5.53 | 0 | -3.453 | 0 | 0.505 |
| 42 | RP11-1049A21.2 | hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-664a-3p | 10 | THBS1 | Sponge network | -4.749 | 0.00016 | -2.128 | 0 | 0.505 |
| 43 | MIR143HG |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 12 | THBS2 | Sponge network | -4.237 | 0 | -0.819 | 0.127 | 0.503 |
| 44 | RP11-554A11.4 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-491-5p | 14 | THBS1 | Sponge network | -3.989 | 0 | -2.128 | 0 | 0.5 |
| 45 | AC002480.3 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-182-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-33a-3p;hsa-miR-664a-3p;hsa-miR-98-5p | 11 | THBS1 | Sponge network | -1.522 | 0.03484 | -2.128 | 0 | 0.498 |
| 46 | HAND2-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 13 | THBS2 | Sponge network | -5.605 | 0 | -0.819 | 0.127 | 0.497 |
| 47 | AC002480.5 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-671-5p | 10 | THBS1 | Sponge network | -2.128 | 0.02032 | -2.128 | 0 | 0.488 |
| 48 | MAGI2-AS3 |
hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | THBS2 | Sponge network | -2.414 | 0 | -0.819 | 0.127 | 0.487 |
| 49 | RP11-389G6.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-944 | 16 | DCN | Sponge network | -7.573 | 0 | -3.453 | 0 | 0.483 |
| 50 | NR2F1-AS1 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-33a-3p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-98-5p | 16 | THBS1 | Sponge network | -1.881 | 0 | -2.128 | 0 | 0.481 |
| 51 | LINC00473 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 19 | THBS1 | Sponge network | -5.53 | 0 | -2.128 | 0 | 0.48 |
| 52 | LINC00702 |
hsa-miR-10b-5p;hsa-miR-148b-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-664a-3p | 10 | INHBA | Sponge network | -2.704 | 0 | 1.441 | 0.01419 | 0.477 |
| 53 | RP11-20J15.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-491-5p;hsa-miR-940 | 17 | THBS1 | Sponge network | -5.104 | 8.0E-5 | -2.128 | 0 | 0.475 |
| 54 | LINC00327 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-98-5p | 12 | DCN | Sponge network | -1.951 | 0.01135 | -3.453 | 0 | 0.475 |
| 55 | RP11-805I24.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-320b;hsa-miR-421;hsa-miR-491-5p;hsa-miR-7-1-3p | 11 | DCN | Sponge network | -5.815 | 0 | -3.453 | 0 | 0.474 |
| 56 | RP11-145A3.1 | hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-491-5p;hsa-miR-98-5p | 10 | THBS1 | Sponge network | 1.041 | 0.23373 | -2.128 | 0 | 0.471 |
| 57 | CTD-2589M5.4 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-320a;hsa-miR-320b | 10 | DCN | Sponge network | -7.181 | 0 | -3.453 | 0 | 0.47 |
| 58 | MAGI2-AS3 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-454-3p | 11 | TGFB2 | Sponge network | -2.414 | 0 | -1.199 | 0.00331 | 0.468 |
| 59 | LINC00565 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-664a-3p | 14 | THBS1 | Sponge network | -1.493 | 0.05998 | -2.128 | 0 | 0.466 |
| 60 | RP11-753H16.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-671-5p | 13 | THBS1 | Sponge network | -5.702 | 0 | -2.128 | 0 | 0.466 |
| 61 | RP11-401O9.3 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-671-5p;hsa-miR-92a-3p;hsa-miR-940 | 14 | THBS1 | Sponge network | -0.262 | 0.83673 | -2.128 | 0 | 0.465 |
| 62 | CTC-296K1.4 | hsa-let-7b-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-576-5p;hsa-miR-590-5p | 10 | THBS1 | Sponge network | -6.559 | 0 | -2.128 | 0 | 0.465 |
| 63 | ADAMTS9-AS1 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-940;hsa-miR-98-5p | 30 | THBS1 | Sponge network | -7.614 | 0 | -2.128 | 0 | 0.46 |
| 64 | ACTA2-AS1 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-940 | 23 | THBS1 | Sponge network | -3.838 | 0 | -2.128 | 0 | 0.457 |
| 65 | LINC00702 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-454-3p;hsa-miR-590-5p | 14 | TGFB2 | Sponge network | -2.704 | 0 | -1.199 | 0.00331 | 0.457 |
| 66 | ACTA2-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-590-3p;hsa-miR-598-3p;hsa-miR-7-1-3p;hsa-miR-96-5p | 10 | THBS2 | Sponge network | -3.838 | 0 | -0.819 | 0.127 | 0.455 |
| 67 | C20orf166-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-491-5p;hsa-miR-664a-3p;hsa-miR-671-5p | 18 | THBS1 | Sponge network | -6.333 | 0 | -2.128 | 0 | 0.455 |
| 68 | CTC-296K1.3 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-671-5p;hsa-miR-940 | 17 | THBS1 | Sponge network | -5.677 | 0 | -2.128 | 0 | 0.453 |
| 69 | RP11-243M5.1 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-940 | 14 | THBS1 | Sponge network | -8.824 | 0 | -2.128 | 0 | 0.451 |
| 70 | RP11-693J15.4 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-491-5p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-940 | 21 | THBS1 | Sponge network | -3.319 | 0.00281 | -2.128 | 0 | 0.445 |
| 71 | LINC00163 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-664a-3p;hsa-miR-671-5p | 15 | THBS1 | Sponge network | -4.312 | 0.00014 | -2.128 | 0 | 0.445 |
| 72 | LINC00694 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-590-3p | 11 | THBS1 | Sponge network | -4.485 | 1.0E-5 | -2.128 | 0 | 0.442 |
| 73 | PDZRN3-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-671-5p;hsa-miR-7-1-3p | 18 | THBS1 | Sponge network | -5.049 | 1.0E-5 | -2.128 | 0 | 0.44 |
| 74 | RP11-62F24.2 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-671-5p;hsa-miR-7-1-3p | 10 | THBS1 | Sponge network | -4.034 | 0.00034 | -2.128 | 0 | 0.439 |
| 75 | AC104654.2 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-7-1-3p;hsa-miR-944 | 11 | DCN | Sponge network | -3.012 | 0.00731 | -3.453 | 0 | 0.438 |
| 76 | LINC00327 |
hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-32-3p;hsa-miR-582-3p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | THBS2 | Sponge network | -1.951 | 0.01135 | -0.819 | 0.127 | 0.436 |
| 77 | CTD-2298J14.2 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-421;hsa-miR-664a-3p;hsa-miR-98-5p | 11 | THBS1 | Sponge network | -5.822 | 0 | -2.128 | 0 | 0.435 |
| 78 | RP11-693J15.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-5p;hsa-miR-7-1-3p | 11 | THBS2 | Sponge network | -3.319 | 0.00281 | -0.819 | 0.127 | 0.432 |
| 79 | RP11-180N14.1 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-576-5p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-98-5p | 19 | THBS1 | Sponge network | -4.46 | 0 | -2.128 | 0 | 0.43 |
| 80 | LINC00163 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p | 11 | DCN | Sponge network | -4.312 | 0.00014 | -3.453 | 0 | 0.423 |
| 81 | AC104654.2 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-940 | 15 | THBS1 | Sponge network | -3.012 | 0.00731 | -2.128 | 0 | 0.42 |
| 82 | TBX5-AS1 | hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 17 | THBS1 | Sponge network | -2.557 | 2.0E-5 | -2.128 | 0 | 0.42 |
| 83 | DNM3OS |
hsa-miR-107;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-33a-3p;hsa-miR-454-3p;hsa-miR-590-5p | 10 | TGFB2 | Sponge network | -2.298 | 1.0E-5 | -1.199 | 0.00331 | 0.418 |
| 84 | LINC00327 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-98-5p | 23 | THBS1 | Sponge network | -1.951 | 0.01135 | -2.128 | 0 | 0.412 |
| 85 | PDZRN3-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 12 | THBS2 | Sponge network | -5.049 | 1.0E-5 | -0.819 | 0.127 | 0.405 |
| 86 | C20orf166-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-93-5p | 10 | THBS2 | Sponge network | -6.333 | 0 | -0.819 | 0.127 | 0.404 |
| 87 | BVES-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-98-5p | 14 | DCN | Sponge network | -4.161 | 1.0E-5 | -3.453 | 0 | 0.403 |
| 88 | ZFHX4-AS1 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-940 | 17 | THBS1 | Sponge network | -2.966 | 0.00743 | -2.128 | 0 | 0.401 |
| 89 | RP11-359E10.1 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-20a-5p;hsa-miR-576-5p;hsa-miR-671-5p | 12 | THBS1 | Sponge network | -1.216 | 0.0438 | -2.128 | 0 | 0.391 |
| 90 | C4A-AS1 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-590-5p;hsa-miR-671-5p;hsa-miR-940 | 15 | THBS1 | Sponge network | -1.76 | 0.00265 | -2.128 | 0 | 0.388 |
| 91 | RP11-498E2.9 | hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p | 11 | THBS1 | Sponge network | -5.841 | 0 | -2.128 | 0 | 0.387 |
| 92 | RP11-753H16.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-93-5p | 10 | THBS2 | Sponge network | -5.702 | 0 | -0.819 | 0.127 | 0.387 |
| 93 | MAGI2-AS3 |
hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-23a-3p;hsa-miR-590-3p;hsa-miR-93-5p | 10 | SMAD9 | Sponge network | -2.414 | 0 | -2.785 | 0 | 0.386 |
| 94 | MIR143HG |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-590-5p | 14 | TGFB2 | Sponge network | -4.237 | 0 | -1.199 | 0.00331 | 0.386 |
| 95 | RP11-887P2.5 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-940 | 21 | THBS1 | Sponge network | -6.751 | 0 | -2.128 | 0 | 0.385 |
| 96 | RP1-65J11.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-421;hsa-miR-7-1-3p;hsa-miR-944 | 10 | DCN | Sponge network | -5.989 | 0 | -3.453 | 0 | 0.378 |
| 97 | APCDD1L-AS1 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-664a-3p;hsa-miR-940 | 13 | THBS1 | Sponge network | -2.022 | 0.00702 | -2.128 | 0 | 0.369 |
| 98 | ADAMTS9-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 13 | THBS2 | Sponge network | -7.614 | 0 | -0.819 | 0.127 | 0.368 |
| 99 | RP11-384P7.7 |
hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 11 | SMAD9 | Sponge network | -3.649 | 3.0E-5 | -2.785 | 0 | 0.362 |
| 100 | EMX2OS |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-26a-2-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-455-5p;hsa-miR-590-3p;hsa-miR-629-3p | 17 | SMAD9 | Sponge network | -2.206 | 0.03389 | -2.785 | 0 | 0.359 |
| 101 | LINC00473 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | THBS2 | Sponge network | -5.53 | 0 | -0.819 | 0.127 | 0.358 |
| 102 | RP11-88I18.3 |
hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-455-5p;hsa-miR-629-3p | 11 | SMAD9 | Sponge network | -7.88 | 0 | -2.785 | 0 | 0.358 |
| 103 | RP11-456K23.1 | hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-32-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-92a-3p | 14 | THBS1 | Sponge network | -1.962 | 1.0E-5 | -2.128 | 0 | 0.357 |
| 104 | HAND2-AS1 |
hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-193a-3p;hsa-miR-30d-3p;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-339-5p;hsa-miR-532-3p;hsa-miR-590-3p;hsa-miR-98-5p | 13 | GDF6 | Sponge network | -5.605 | 0 | -1.135 | 0.02791 | 0.352 |
| 105 | RP11-161D15.1 | hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-664a-3p;hsa-miR-671-5p;hsa-miR-98-5p | 12 | THBS1 | Sponge network | -6.672 | 0 | -2.128 | 0 | 0.348 |
| 106 | BVES-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-576-5p;hsa-miR-664a-3p;hsa-miR-98-5p | 20 | THBS1 | Sponge network | -4.161 | 1.0E-5 | -2.128 | 0 | 0.345 |
| 107 | FENDRR |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-940;hsa-miR-98-5p | 21 | THBS1 | Sponge network | -4.793 | 0 | -2.128 | 0 | 0.343 |
| 108 | AF131217.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-193a-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-26a-2-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 18 | SMAD9 | Sponge network | -5.31 | 0 | -2.785 | 0 | 0.342 |
| 109 | MIR497HG |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-590-5p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-940 | 21 | THBS1 | Sponge network | -3.802 | 0 | -2.128 | 0 | 0.336 |
| 110 | LINC00702 |
hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-30d-3p;hsa-miR-339-5p;hsa-miR-532-3p;hsa-miR-589-3p;hsa-miR-590-3p | 10 | GDF6 | Sponge network | -2.704 | 0 | -1.135 | 0.02791 | 0.335 |
| 111 | PWAR6 | hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-155-5p;hsa-miR-16-2-3p;hsa-miR-193a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-590-3p | 10 | SMAD9 | Sponge network | -2.542 | 0 | -2.785 | 0 | 0.335 |
| 112 | RP11-81H14.2 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-576-5p;hsa-miR-664a-3p;hsa-miR-940 | 17 | THBS1 | Sponge network | -2.322 | 0.00014 | -2.128 | 0 | 0.334 |
| 113 | AF131217.1 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-186-5p;hsa-miR-320b;hsa-miR-421;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-98-5p | 15 | DCN | Sponge network | -5.31 | 0 | -3.453 | 0 | 0.332 |
| 114 | FENDRR |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-193a-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-26a-2-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-455-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 19 | SMAD9 | Sponge network | -4.793 | 0 | -2.785 | 0 | 0.33 |
| 115 | CTD-2554C21.2 | hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-155-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-629-3p;hsa-miR-93-5p | 12 | SMAD9 | Sponge network | -2.822 | 0.00245 | -2.785 | 0 | 0.329 |
| 116 | PART1 |
hsa-miR-142-3p;hsa-miR-186-5p;hsa-miR-27a-3p;hsa-miR-338-5p;hsa-miR-342-3p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p | 10 | ID4 | Sponge network | -3.06 | 7.0E-5 | -2.192 | 0 | 0.328 |
| 117 | PGM5-AS1 |
hsa-let-7a-3p;hsa-let-7d-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-7-1-3p;hsa-miR-940 | 13 | THBS1 | Sponge network | -11.072 | 0 | -2.128 | 0 | 0.328 |
| 118 | ACTA2-AS1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-454-3p | 12 | TGFB2 | Sponge network | -3.838 | 0 | -1.199 | 0.00331 | 0.327 |
| 119 | MIR497HG |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-26a-2-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-629-3p;hsa-miR-93-5p | 16 | SMAD9 | Sponge network | -3.802 | 0 | -2.785 | 0 | 0.327 |
| 120 | PGM5-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p | 11 | SMAD9 | Sponge network | -11.072 | 0 | -2.785 | 0 | 0.325 |
| 121 | RP11-384P7.7 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-98-5p | 10 | DCN | Sponge network | -3.649 | 3.0E-5 | -3.453 | 0 | 0.325 |
| 122 | RP11-439E19.10 | hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-193a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p | 10 | SMAD9 | Sponge network | -0.548 | 0.31987 | -2.785 | 0 | 0.321 |
| 123 | PDZRN3-AS1 |
hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-23a-3p;hsa-miR-26a-2-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p;hsa-miR-629-3p | 12 | SMAD9 | Sponge network | -5.049 | 1.0E-5 | -2.785 | 0 | 0.319 |
| 124 | RP11-166D19.1 |
hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-30d-3p;hsa-miR-330-3p;hsa-miR-339-5p;hsa-miR-532-3p;hsa-miR-589-3p;hsa-miR-590-3p | 11 | GDF6 | Sponge network | -3.855 | 0 | -1.135 | 0.02791 | 0.318 |
| 125 | RP11-120J1.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p;hsa-miR-629-3p | 14 | SMAD9 | Sponge network | -6.067 | 0 | -2.785 | 0 | 0.317 |
| 126 | RP11-283G6.5 | hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-940 | 10 | THBS1 | Sponge network | -1.071 | 0.39761 | -2.128 | 0 | 0.317 |
| 127 | RP11-401P9.4 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-93-5p | 11 | SMAD9 | Sponge network | -2.738 | 0 | -2.785 | 0 | 0.316 |
| 128 | RP11-13K12.5 |
hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-193a-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-629-3p;hsa-miR-93-5p | 12 | SMAD9 | Sponge network | -4.086 | 0 | -2.785 | 0 | 0.313 |
| 129 | C20orf166-AS1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-330-5p;hsa-miR-429;hsa-miR-454-3p | 10 | TGFB2 | Sponge network | -6.333 | 0 | -1.199 | 0.00331 | 0.313 |
| 130 | RP11-428G5.5 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-940 | 14 | THBS1 | Sponge network | -1.039 | 0.39301 | -2.128 | 0 | 0.311 |
| 131 | HAND2-AS1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-429 | 14 | TGFB2 | Sponge network | -5.605 | 0 | -1.199 | 0.00331 | 0.309 |
| 132 | RP11-389G6.3 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-590-3p | 18 | THBS1 | Sponge network | -7.573 | 0 | -2.128 | 0 | 0.309 |
| 133 | MEG9 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-671-5p | 11 | THBS1 | Sponge network | -3.37 | 3.0E-5 | -2.128 | 0 | 0.301 |
| 134 | RP5-887A10.1 | hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-16-1-3p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-671-5p;hsa-miR-92a-3p;hsa-miR-940 | 14 | THBS1 | Sponge network | -3.078 | 0.0212 | -2.128 | 0 | 0.299 |
| 135 | RP11-6O2.3 |
hsa-let-7a-3p;hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-629-3p | 12 | SMAD9 | Sponge network | -4.533 | 0 | -2.785 | 0 | 0.298 |
| 136 | CTC-296K1.3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-23a-3p;hsa-miR-29a-5p | 11 | SMAD9 | Sponge network | -5.677 | 0 | -2.785 | 0 | 0.294 |
| 137 | RP11-389G6.3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-142-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p;hsa-miR-93-5p | 15 | SMAD9 | Sponge network | -7.573 | 0 | -2.785 | 0 | 0.29 |
| 138 | GATA6-AS1 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-671-5p | 14 | THBS1 | Sponge network | -3.855 | 0 | -2.128 | 0 | 0.288 |
| 139 | MIR143HG |
hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-30d-3p;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-339-5p;hsa-miR-532-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-98-5p | 13 | GDF6 | Sponge network | -4.237 | 0 | -1.135 | 0.02791 | 0.286 |
| 140 | ZFHX4-AS1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-590-5p | 11 | TGFB2 | Sponge network | -2.966 | 0.00743 | -1.199 | 0.00331 | 0.286 |
| 141 | ADAMTS9-AS1 |
hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-193a-3p;hsa-miR-30d-3p;hsa-miR-324-5p;hsa-miR-339-5p;hsa-miR-532-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-98-5p | 13 | GDF6 | Sponge network | -7.614 | 0 | -1.135 | 0.02791 | 0.283 |
| 142 | RP11-389G6.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-93-5p | 10 | THBS2 | Sponge network | -7.573 | 0 | -0.819 | 0.127 | 0.282 |
| 143 | ADAMTS9-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-455-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 17 | SMAD9 | Sponge network | -7.614 | 0 | -2.785 | 0 | 0.282 |
| 144 | ACTA2-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-455-5p;hsa-miR-590-3p | 13 | SMAD9 | Sponge network | -3.838 | 0 | -2.785 | 0 | 0.281 |
| 145 | DIO3OS |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-671-5p;hsa-miR-940 | 16 | THBS1 | Sponge network | -3.619 | 0 | -2.128 | 0 | 0.279 |
| 146 | HAND2-AS1 |
hsa-miR-10b-5p;hsa-miR-148b-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-664a-3p | 10 | INHBA | Sponge network | -5.605 | 0 | 1.441 | 0.01419 | 0.279 |
| 147 | LINC01018 | hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-629-3p | 13 | SMAD9 | Sponge network | -1.607 | 0.09425 | -2.785 | 0 | 0.277 |
| 148 | MIR143HG |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 15 | SMAD9 | Sponge network | -4.237 | 0 | -2.785 | 0 | 0.274 |
| 149 | HAR1A | hsa-let-7a-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-33a-3p;hsa-miR-940 | 10 | THBS1 | Sponge network | -2.085 | 0 | -2.128 | 0 | 0.274 |
| 150 | RP11-678G14.3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-148b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-2355-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p | 11 | SMAD9 | Sponge network | -4.141 | 0.0002 | -2.785 | 0 | 0.273 |
| 151 | RP11-496N12.6 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-671-5p | 11 | THBS1 | Sponge network | -5.574 | 0 | -2.128 | 0 | 0.27 |
| 152 | BZRAP1-AS1 | hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-671-5p;hsa-miR-940 | 12 | THBS1 | Sponge network | -2.343 | 0 | -2.128 | 0 | 0.27 |
| 153 | RP1-65J11.1 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-940 | 16 | THBS1 | Sponge network | -5.989 | 0 | -2.128 | 0 | 0.269 |
| 154 | RP11-384P7.7 |
hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-7-1-3p;hsa-miR-940;hsa-miR-98-5p | 19 | THBS1 | Sponge network | -3.649 | 3.0E-5 | -2.128 | 0 | 0.269 |
| 155 | RP11-887P2.5 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-142-5p;hsa-miR-148b-5p;hsa-miR-155-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-193a-5p;hsa-miR-20a-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-93-5p | 16 | SMAD9 | Sponge network | -6.751 | 0 | -2.785 | 0 | 0.264 |
| 156 | MIR143HG |
hsa-miR-10b-5p;hsa-miR-148b-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-664a-3p | 10 | INHBA | Sponge network | -4.237 | 0 | 1.441 | 0.01419 | 0.263 |
| 157 | RP11-416N2.4 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-148b-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p;hsa-miR-93-5p | 12 | SMAD9 | Sponge network | -2.355 | 0.00054 | -2.785 | 0 | 0.263 |
| 158 | FENDRR |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 13 | THBS2 | Sponge network | -4.793 | 0 | -0.819 | 0.127 | 0.261 |
| 159 | RP11-805I24.3 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-491-5p;hsa-miR-576-5p;hsa-miR-7-1-3p;hsa-miR-940 | 20 | THBS1 | Sponge network | -5.815 | 0 | -2.128 | 0 | 0.259 |
| 160 | RP11-88I18.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-20a-5p | 13 | THBS1 | Sponge network | -7.88 | 0 | -2.128 | 0 | 0.259 |
| 161 | RP11-887P2.5 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-32-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | THBS2 | Sponge network | -6.751 | 0 | -0.819 | 0.127 | 0.258 |
| 162 | RP11-120J1.1 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-590-3p | 16 | THBS1 | Sponge network | -6.067 | 0 | -2.128 | 0 | 0.254 |
| 163 | RP1-65J11.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-103a-2-5p;hsa-miR-142-5p;hsa-miR-148b-5p;hsa-miR-155-5p;hsa-miR-186-5p;hsa-miR-193a-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-455-5p | 13 | SMAD9 | Sponge network | -5.989 | 0 | -2.785 | 0 | 0.252 |
| 164 | RP11-401P9.4 |
hsa-let-7a-3p;hsa-let-7a-5p;hsa-let-7d-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-590-5p | 12 | THBS1 | Sponge network | -2.738 | 0 | -2.128 | 0 | 0.251 |