microRNA information: hsa-miR-148b-3p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-148b-3p | miRbase |
| Accession: | MIMAT0000759 | miRbase |
| Precursor name: | hsa-mir-148b | miRbase |
| Precursor accession: | MI0000811 | miRbase |
| Symbol: | MIR148B | HGNC |
| RefSeq ID: | NR_029894 | GenBank |
| Sequence: | UCAGUGCAUCACAGAACUUUGU |
Reported expression in cancers: hsa-miR-148b-3p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-148b-3p | bladder cancer | upregulation | "Identification and validation of reference genes f ......" | 25738263 | qPCR; RNA-Seq |
| hsa-miR-148b-3p | breast cancer | upregulation | "Selected candidates identified in the initial scre ......" | 22927033 | qPCR |
| hsa-miR-148b-3p | breast cancer | downregulation | "Among them miR-148b down-regulated in aggressive b ......" | 23233531 | |
| hsa-miR-148b-3p | breast cancer | deregulation | "The objective of this study is to investigate the ......" | 25232827 | qPCR |
| hsa-miR-148b-3p | chronic myeloid leukemia | downregulation | "Downregulated microRNA 148b in circulating PBMCs i ......" | 25187697 | |
| hsa-miR-148b-3p | colorectal cancer | downregulation | "The purpose of this study was to elucidate the mol ......" | 22020560 | qPCR; in situ hybridization |
| hsa-miR-148b-3p | gastric cancer | downregulation | "MicroRNA 148b is frequently down regulated in gast ......" | 21205300 | qPCR; in situ hybridization |
| hsa-miR-148b-3p | gastric cancer | downregulation | "Using Limma package in R a total of 27 differentia ......" | 26692821 | |
| hsa-miR-148b-3p | liver cancer | downregulation | "MiR 148b suppresses cell proliferation and invasio ......" | 25627001 | |
| hsa-miR-148b-3p | liver cancer | downregulation | "Profiles of tissue microRNAs; miR 148b and miR 25 ......" | 26209296 | qPCR |
| hsa-miR-148b-3p | lung cancer | downregulation | "miR-148b functions as a tumor suppressor in non-sm ......" | 26664144 | |
| hsa-miR-148b-3p | lung cancer | downregulation | "Western Blot and luciferase assay were performed t ......" | 26759383 | |
| hsa-miR-148b-3p | lung squamous cell cancer | downregulation | "Serum miR 152 miR 148a miR 148b and miR 21 as nove ......" | 25501703 | qPCR |
| hsa-miR-148b-3p | lung squamous cell cancer | downregulation | "MicroRNA 148b is down regulated in non small cell ......" | 25755777 | qPCR |
| hsa-miR-148b-3p | lung squamous cell cancer | downregulation | "Down regulated MicroRNA 148b expression as predict ......" | 26377406 | qPCR |
| hsa-miR-148b-3p | lymphoma | upregulation | "Among the differentially expressed miRNAs miR-148b ......" | 22843616 | qPCR |
| hsa-miR-148b-3p | pancreatic cancer | downregulation | "miR 148b functions as a tumor suppressor in pancre ......" | 23171948 | |
| hsa-miR-148b-3p | prostate cancer | deregulation | "Upregulation of miR-122 miR-335 miR-184 miR-193 mi ......" | 23781281 | |
| hsa-miR-148b-3p | thyroid cancer | upregulation | "High expression levels of miR-146-5p miR-199a-5p m ......" | 27586203 |
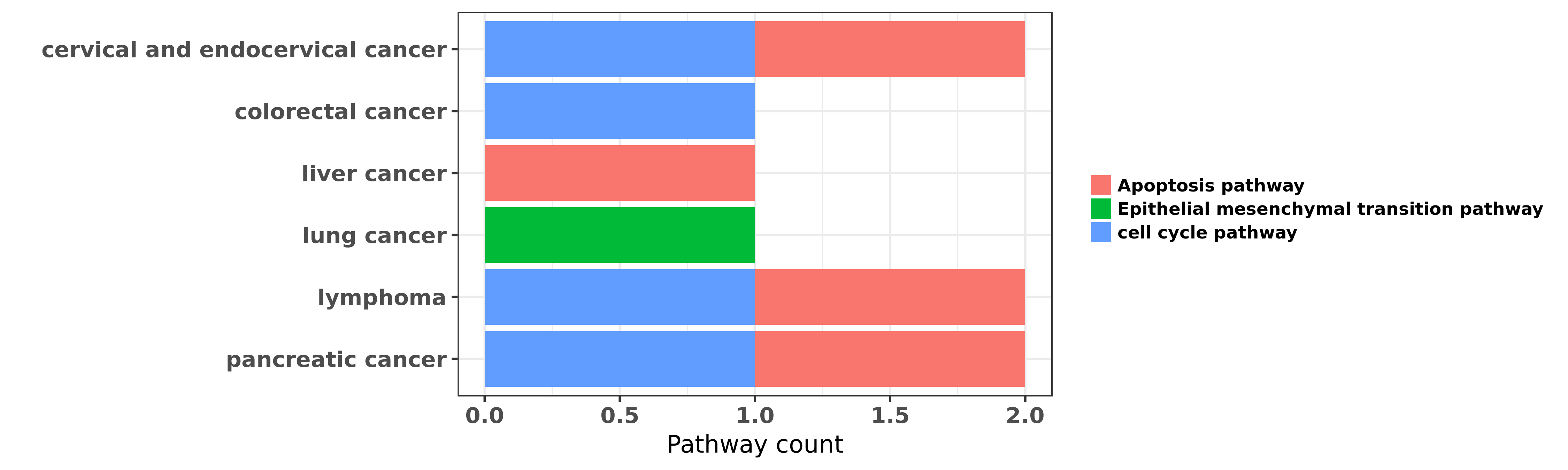
Reported cancer pathway affected by hsa-miR-148b-3p
| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-148b-3p | cervical and endocervical cancer | cell cycle pathway; Apoptosis pathway | "MicroRNA 148b Acts as a Tumor Suppressor in Cervic ......" | 27505047 | Colony formation |
| hsa-miR-148b-3p | colorectal cancer | cell cycle pathway | "Altered p53 regulation of miR 148b and p55PIK cont ......" | 24632606 | |
| hsa-miR-148b-3p | liver cancer | Apoptosis pathway | "MiR 148b suppresses cell proliferation and invasio ......" | 25627001 | Luciferase |
| hsa-miR-148b-3p | lung cancer | Epithelial mesenchymal transition pathway | "Restoration of BRG1 inhibits proliferation and met ......" | 26664144 | MTT assay; Western blot; Luciferase |
| hsa-miR-148b-3p | lymphoma | Apoptosis pathway; cell cycle pathway | "MicroRNA 148b enhances the radiosensitivity of non ......" | 22843616 | |
| hsa-miR-148b-3p | pancreatic cancer | cell cycle pathway; Apoptosis pathway | "miR 148b functions as a tumor suppressor in pancre ......" | 23171948 | RNAi |
| hsa-miR-148b-3p | pancreatic cancer | Apoptosis pathway | "MicroRNA 148b and microRNA 152 reactivate tumor su ......" | 24448385 | Luciferase |
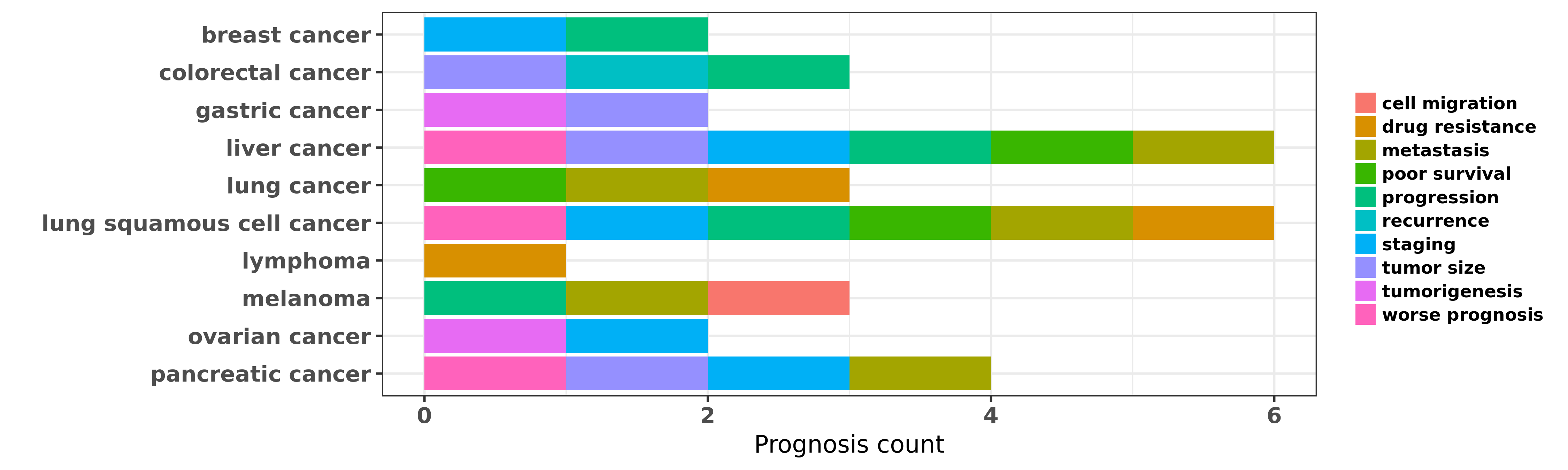
Reported cancer prognosis affected by hsa-miR-148b-3p
| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-148b-3p | breast cancer | progression | "miR148b is a major coordinator of breast cancer pr ......" | 23233531 | |
| hsa-miR-148b-3p | breast cancer | staging | "Further we validated our previously reported circu ......" | 24194846 | |
| hsa-miR-148b-3p | colorectal cancer | tumor size | "MicroRNA 148b suppresses cell growth by targeting ......" | 22020560 | Western blot; Luciferase |
| hsa-miR-148b-3p | colorectal cancer | recurrence | "TaqMan® Human MicroRNA Array Set v2.0 was used to ......" | 23280316 | |
| hsa-miR-148b-3p | colorectal cancer | progression | "Altered p53 regulation of miR 148b and p55PIK cont ......" | 24632606 | |
| hsa-miR-148b-3p | gastric cancer | tumorigenesis; tumor size | "MicroRNA 148b is frequently down regulated in gast ......" | 21205300 | MTT assay; Western blot; Luciferase |
| hsa-miR-148b-3p | liver cancer | metastasis | "Identification of metastasis related microRNAs of ......" | 19912688 | |
| hsa-miR-148b-3p | liver cancer | worse prognosis; staging; poor survival; progression | "MicroRNA 148b expression is decreased in hepatocel ......" | 24805877 | |
| hsa-miR-148b-3p | liver cancer | metastasis; worse prognosis; tumor size | "MiR 148b suppresses cell proliferation and invasio ......" | 25627001 | Luciferase |
| hsa-miR-148b-3p | liver cancer | metastasis | "In the present study we observed that overexpressi ......" | 25997710 | |
| hsa-miR-148b-3p | liver cancer | poor survival; staging; metastasis; worse prognosis | "Downregulation of miR 148b as biomarker for early ......" | 26206498 | |
| hsa-miR-148b-3p | liver cancer | poor survival; staging; metastasis | "Profiles of tissue microRNAs; miR 148b and miR 25 ......" | 26209296 | |
| hsa-miR-148b-3p | liver cancer | metastasis | "MicroRNA 148b targets Rho associated protein kinas ......" | 26530325 | Western blot; MTT assay; Luciferase |
| hsa-miR-148b-3p | lung cancer | metastasis | "Restoration of BRG1 inhibits proliferation and met ......" | 26664144 | MTT assay; Western blot; Luciferase |
| hsa-miR-148b-3p | lung cancer | drug resistance; poor survival | "miR-148b has been reported to be implicated regula ......" | 26759383 | Western blot; Luciferase |
| hsa-miR-148b-3p | lung squamous cell cancer | progression | "miR 148b functions as a tumor suppressor in non sm ......" | 25232379 | Luciferase |
| hsa-miR-148b-3p | lung squamous cell cancer | poor survival; worse prognosis; staging; metastasis | "MicroRNA 148b is down regulated in non small cell ......" | 25755777 | |
| hsa-miR-148b-3p | lung squamous cell cancer | poor survival; staging; metastasis | "Down regulated MicroRNA 148b expression as predict ......" | 26377406 | |
| hsa-miR-148b-3p | lung squamous cell cancer | drug resistance; staging; poor survival | "MicroRNA 148b is a potential prognostic biomarker ......" | 27083571 | |
| hsa-miR-148b-3p | lung squamous cell cancer | staging | "The aim of this study was to determine whether cir ......" | 27461630 | |
| hsa-miR-148b-3p | lymphoma | drug resistance | "MicroRNA 148b enhances the radiosensitivity of non ......" | 22843616 | |
| hsa-miR-148b-3p | melanoma | progression; cell migration; metastasis | "miR 214 coordinates melanoma progression by upregu ......" | 23667173 | |
| hsa-miR-148b-3p | melanoma | metastasis | "miR 214 and miR 148b Targeting Inhibits Disseminat ......" | 27328731 | |
| hsa-miR-148b-3p | ovarian cancer | staging; tumorigenesis | "Increased expression of miR 148b in ovarian carcin ......" | 22344713 | |
| hsa-miR-148b-3p | pancreatic cancer | staging; metastasis; worse prognosis; tumor size | "miR 148b functions as a tumor suppressor in pancre ......" | 23171948 | RNAi |
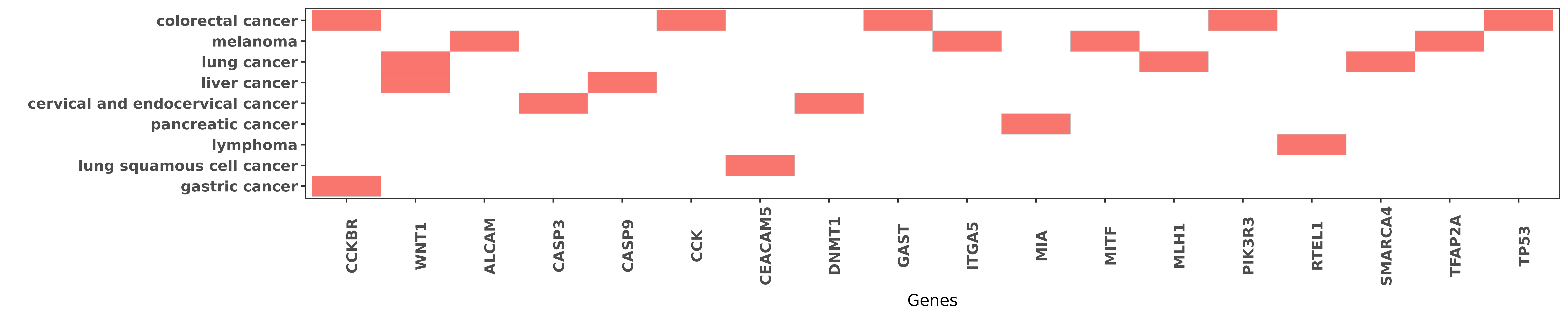
Reported gene related to hsa-miR-148b-3p
| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-miR-148b-3p | melanoma | ALCAM | "miR 214 coordinates melanoma progression by upregu ......" | 23667173 |
| hsa-miR-148b-3p | melanoma | ALCAM | "Notably transendothelial migration in vitro and ex ......" | 27328731 |
| hsa-miR-148b-3p | colorectal cancer | CCKBR | "MicroRNA 148b suppresses cell growth by targeting ......" | 22020560 |
| hsa-miR-148b-3p | gastric cancer | CCKBR | "Using a luciferase activity assay and western blot ......" | 21205300 |
| hsa-miR-148b-3p | liver cancer | WNT1 | "Data from the dual-luciferase reporter gene assay ......" | 25627001 |
| hsa-miR-148b-3p | lung cancer | WNT1 | "Additionally we suggested that miR-148b suppressed ......" | 26664144 |
| hsa-miR-148b-3p | cervical and endocervical cancer | CASP3 | "MicroRNA 148b Acts as a Tumor Suppressor in Cervic ......" | 27505047 |
| hsa-miR-148b-3p | liver cancer | CASP9 | "Further miR-148b induced cells apoptosis by activa ......" | 25627001 |
| hsa-miR-148b-3p | colorectal cancer | CCK | "MicroRNA 148b suppresses cell growth by targeting ......" | 22020560 |
| hsa-miR-148b-3p | lung squamous cell cancer | CEACAM5 | "miR 148b functions as a tumor suppressor in non sm ......" | 25232379 |
| hsa-miR-148b-3p | cervical and endocervical cancer | DNMT1 | "Forty-eight hours after transfection the mRNA expr ......" | 27505047 |
| hsa-miR-148b-3p | colorectal cancer | GAST | "Then we used siRNA radioimmunoassay and ELISA to d ......" | 22020560 |
| hsa-miR-148b-3p | melanoma | ITGA5 | "Notably transendothelial migration in vitro and ex ......" | 27328731 |
| hsa-miR-148b-3p | pancreatic cancer | MIA | "Moreover the introduced miR-148b and -152 could in ......" | 24448385 |
| hsa-miR-148b-3p | melanoma | MITF | "miR 148 regulates Mitf in melanoma cells; Furtherm ......" | 20644734 |
| hsa-miR-148b-3p | lung cancer | MLH1 | "MutL homologue 1 MLH1 is the direct target of miR- ......" | 26759383 |
| hsa-miR-148b-3p | colorectal cancer | PIK3R3 | "Altered p53 regulation of miR 148b and p55PIK cont ......" | 24632606 |
| hsa-miR-148b-3p | lymphoma | RTEL1 | "Taken together these results demonstrate that miR- ......" | 22843616 |
| hsa-miR-148b-3p | lung cancer | SMARCA4 | "Restoration of BRG1 inhibits proliferation and met ......" | 26664144 |
| hsa-miR-148b-3p | melanoma | TFAP2A | "miR 214 coordinates melanoma progression by upregu ......" | 23667173 |
| hsa-miR-148b-3p | colorectal cancer | TP53 | "Altered p53 regulation of miR 148b and p55PIK cont ......" | 24632606 |
Expression profile in cancer corhorts:

Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-148b-3p | ACACB | 9 cancers: BLCA; BRCA; COAD; KIRC; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.678; TCGA BRCA -0.751; TCGA COAD -0.333; TCGA KIRC -0.547; TCGA OV -0.184; TCGA PRAD -0.402; TCGA SARC -0.314; TCGA STAD -0.647; TCGA UCEC -0.469 |
hsa-miR-148b-3p | ADAM33 | 16 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -1.593; TCGA BRCA -1.432; TCGA CESC -1.042; TCGA COAD -1.7; TCGA ESCA -1.111; TCGA HNSC -0.496; TCGA KIRC -1.253; TCGA KIRP -0.445; TCGA LIHC -0.422; TCGA LUAD -0.437; TCGA LUSC -0.665; TCGA OV -0.744; TCGA PAAD -1.089; TCGA PRAD -0.651; TCGA STAD -1.453; TCGA UCEC -1.015 |
hsa-miR-148b-3p | ADAMTS4 | 9 cancers: BLCA; CESC; COAD; LUSC; OV; PAAD; PRAD; THCA; UCEC | miRNAWalker2 validate | TCGA BLCA -0.818; TCGA CESC -0.307; TCGA COAD -0.433; TCGA LUSC -0.476; TCGA OV -0.423; TCGA PAAD -0.574; TCGA PRAD -0.489; TCGA THCA -0.521; TCGA UCEC -0.234 |
hsa-miR-148b-3p | ADAMTS6 | 11 cancers: BLCA; COAD; ESCA; KIRC; KIRP; LGG; LUSC; PAAD; SARC; THCA; STAD | miRNAWalker2 validate | TCGA BLCA -0.8; TCGA COAD -0.261; TCGA ESCA -0.332; TCGA KIRC -0.686; TCGA KIRP -0.557; TCGA LGG -0.605; TCGA LUSC -0.532; TCGA PAAD -0.54; TCGA SARC -0.341; TCGA THCA -0.572; TCGA STAD -0.309 |
hsa-miR-148b-3p | AKAP11 | 10 cancers: BLCA; CESC; ESCA; LIHC; LUSC; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.163; TCGA CESC -0.201; TCGA ESCA -0.141; TCGA LIHC -0.242; TCGA LUSC -0.106; TCGA OV -0.226; TCGA PRAD -0.215; TCGA SARC -0.142; TCGA STAD -0.198; TCGA UCEC -0.243 |
hsa-miR-148b-3p | APBB1 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LUAD; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.864; TCGA BRCA -0.307; TCGA CESC -0.294; TCGA COAD -0.87; TCGA ESCA -0.452; TCGA KIRC -0.121; TCGA KIRP -0.176; TCGA LUAD -0.2; TCGA OV -0.343; TCGA PRAD -0.463; TCGA SARC -0.352; TCGA STAD -0.718; TCGA UCEC -0.611 |
hsa-miR-148b-3p | APBB2 | 13 cancers: BLCA; CESC; HNSC; KIRC; KIRP; LGG; LIHC; LUAD; LUSC; OV; PAAD; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.35; TCGA CESC -0.261; TCGA HNSC -0.159; TCGA KIRC -0.268; TCGA KIRP -0.385; TCGA LGG -0.155; TCGA LIHC -0.314; TCGA LUAD -0.216; TCGA LUSC -0.198; TCGA OV -0.258; TCGA PAAD -0.231; TCGA STAD -0.24; TCGA UCEC -0.203 |
hsa-miR-148b-3p | CCL19 | 10 cancers: BLCA; BRCA; COAD; ESCA; LUAD; PAAD; PRAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -1.496; TCGA BRCA -0.698; TCGA COAD -1.592; TCGA ESCA -0.879; TCGA LUAD -0.318; TCGA PAAD -1.486; TCGA PRAD -0.626; TCGA THCA -2.133; TCGA STAD -0.685; TCGA UCEC -0.455 |
hsa-miR-148b-3p | CPA3 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; LIHC; LUAD; OV; PAAD; PRAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -1.486; TCGA BRCA -0.72; TCGA CESC -0.619; TCGA COAD -0.975; TCGA ESCA -0.466; TCGA KIRP -0.89; TCGA LIHC -0.668; TCGA LUAD -0.834; TCGA OV -0.482; TCGA PAAD -0.908; TCGA PRAD -0.406; TCGA SARC -0.911; TCGA THCA -2.421; TCGA STAD -0.607; TCGA UCEC -1.069 |
hsa-miR-148b-3p | DLG2 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; OV; SARC; STAD; UCEC | miRNAWalker2 validate; MirTarget | TCGA BLCA -0.773; TCGA BRCA -0.469; TCGA CESC -0.971; TCGA COAD -1.335; TCGA ESCA -0.947; TCGA KIRC -0.619; TCGA KIRP -0.345; TCGA LIHC -0.405; TCGA LUAD -0.486; TCGA OV -0.506; TCGA SARC -0.602; TCGA STAD -1.302; TCGA UCEC -0.641 |
hsa-miR-148b-3p | DNAJB4 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; PAAD; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.574; TCGA BRCA -0.227; TCGA CESC -0.41; TCGA COAD -0.254; TCGA ESCA -0.447; TCGA HNSC -0.228; TCGA LIHC -0.327; TCGA LUAD -0.189; TCGA PAAD -0.313; TCGA PRAD -0.593; TCGA SARC -0.281; TCGA STAD -0.572; TCGA UCEC -0.482 |
hsa-miR-148b-3p | DPT | 16 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -2.387; TCGA BRCA -1.072; TCGA CESC -1.156; TCGA COAD -1.573; TCGA ESCA -1.012; TCGA HNSC -0.542; TCGA KIRC -1.529; TCGA KIRP -0.59; TCGA LIHC -0.574; TCGA LUAD -0.601; TCGA LUSC -0.463; TCGA OV -1.009; TCGA PAAD -0.669; TCGA PRAD -1.03; TCGA STAD -1.258; TCGA UCEC -1.508 |
hsa-miR-148b-3p | EYA4 | 11 cancers: BLCA; BRCA; COAD; HNSC; KIRC; KIRP; LUAD; PAAD; PRAD; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.847; TCGA BRCA -0.481; TCGA COAD -0.614; TCGA HNSC -0.431; TCGA KIRC -1.444; TCGA KIRP -1.124; TCGA LUAD -0.388; TCGA PAAD -0.855; TCGA PRAD -0.876; TCGA STAD -0.422; TCGA UCEC -0.734 |
hsa-miR-148b-3p | GFRA1 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LIHC; LUAD; OV; PAAD; PRAD; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -1.013; TCGA BRCA -0.476; TCGA CESC -1.165; TCGA COAD -1.388; TCGA ESCA -0.827; TCGA HNSC -0.394; TCGA LGG -0.737; TCGA LIHC -0.755; TCGA LUAD -0.313; TCGA OV -0.973; TCGA PAAD -0.413; TCGA PRAD -0.7; TCGA STAD -1.33; TCGA UCEC -1.15 |
hsa-miR-148b-3p | GLI3 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; LIHC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.521; TCGA BRCA -0.19; TCGA CESC -1.117; TCGA COAD -1.228; TCGA ESCA -1.15; TCGA LIHC -0.436; TCGA LUAD -0.606; TCGA LUSC -0.284; TCGA OV -0.4; TCGA PAAD -0.714; TCGA PRAD -0.537; TCGA STAD -0.848; TCGA UCEC -0.664 |
hsa-miR-148b-3p | GNAI2 | 13 cancers: BLCA; BRCA; COAD; ESCA; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.146; TCGA BRCA -0.086; TCGA COAD -0.233; TCGA ESCA -0.136; TCGA LIHC -0.11; TCGA LUAD -0.276; TCGA LUSC -0.17; TCGA OV -0.093; TCGA PAAD -0.127; TCGA PRAD -0.283; TCGA THCA -0.33; TCGA STAD -0.112; TCGA UCEC -0.097 |
hsa-miR-148b-3p | HEY2 | 11 cancers: BLCA; BRCA; CESC; COAD; LGG; LIHC; LUAD; LUSC; SARC; THCA; STAD | miRNAWalker2 validate | TCGA BLCA -0.296; TCGA BRCA -0.374; TCGA CESC -0.698; TCGA COAD -0.329; TCGA LGG -0.19; TCGA LIHC -0.443; TCGA LUAD -0.205; TCGA LUSC -0.282; TCGA SARC -0.869; TCGA THCA -0.69; TCGA STAD -0.275 |
hsa-miR-148b-3p | IGSF1 | 9 cancers: BLCA; BRCA; CESC; COAD; HNSC; PRAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.392; TCGA BRCA -0.66; TCGA CESC -0.676; TCGA COAD -0.54; TCGA HNSC -0.475; TCGA PRAD -1.35; TCGA THCA -0.963; TCGA STAD -0.763; TCGA UCEC -0.323 |
hsa-miR-148b-3p | IGSF10 | 10 cancers: BLCA; BRCA; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; STAD | miRNAWalker2 validate | TCGA BLCA -1.393; TCGA BRCA -1.057; TCGA COAD -0.579; TCGA ESCA -0.651; TCGA KIRC -0.752; TCGA KIRP -0.622; TCGA LIHC -0.911; TCGA LUAD -0.352; TCGA PAAD -0.417; TCGA STAD -0.83 |
hsa-miR-148b-3p | INHBA | 14 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; SARC; THCA; UCEC | miRNAWalker2 validate | TCGA BLCA -0.815; TCGA CESC -0.667; TCGA COAD -0.734; TCGA ESCA -0.805; TCGA HNSC -0.542; TCGA KIRP -0.457; TCGA LIHC -0.478; TCGA LUAD -0.308; TCGA LUSC -0.985; TCGA OV -0.64; TCGA PAAD -1.21; TCGA SARC -0.712; TCGA THCA -1.379; TCGA UCEC -0.311 |
hsa-miR-148b-3p | ITGA5 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; LUAD; LUSC; OV; PAAD; PRAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -1.091; TCGA BRCA -0.122; TCGA CESC -0.363; TCGA COAD -0.676; TCGA ESCA -0.482; TCGA LUAD -0.243; TCGA LUSC -0.588; TCGA OV -0.2; TCGA PAAD -0.702; TCGA PRAD -0.588; TCGA SARC -0.288; TCGA THCA -0.184; TCGA STAD -0.517; TCGA UCEC -0.278 |
hsa-miR-148b-3p | KLF6 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.276; TCGA BRCA -0.327; TCGA CESC -0.353; TCGA COAD -0.169; TCGA ESCA -0.314; TCGA HNSC -0.34; TCGA LIHC -0.532; TCGA LUAD -0.246; TCGA LUSC -0.284; TCGA PAAD -0.567; TCGA PRAD -0.295; TCGA STAD -0.182; TCGA UCEC -0.414 |
hsa-miR-148b-3p | LCA5 | 13 cancers: BLCA; BRCA; CESC; COAD; KIRP; LIHC; LUAD; PAAD; PRAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.182; TCGA BRCA -0.277; TCGA CESC -0.444; TCGA COAD -0.443; TCGA KIRP -0.295; TCGA LIHC -0.278; TCGA LUAD -0.285; TCGA PAAD -0.224; TCGA PRAD -0.332; TCGA SARC -0.311; TCGA THCA -0.338; TCGA STAD -0.397; TCGA UCEC -0.424 |
hsa-miR-148b-3p | LOXL1 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD | miRNAWalker2 validate | TCGA BLCA -0.416; TCGA BRCA -0.187; TCGA CESC -0.407; TCGA COAD -0.516; TCGA ESCA -0.316; TCGA KIRC -0.576; TCGA KIRP -0.352; TCGA LUAD -0.283; TCGA LUSC -0.377; TCGA OV -0.301; TCGA PAAD -0.638; TCGA PRAD -0.268; TCGA THCA -1.001; TCGA STAD -0.246 |
hsa-miR-148b-3p | LPL | 9 cancers: BLCA; BRCA; COAD; HNSC; KIRC; LGG; OV; THCA; STAD | miRNAWalker2 validate | TCGA BLCA -0.573; TCGA BRCA -0.913; TCGA COAD -0.528; TCGA HNSC -0.428; TCGA KIRC -0.276; TCGA LGG -0.273; TCGA OV -0.328; TCGA THCA -0.646; TCGA STAD -0.322 |
hsa-miR-148b-3p | MARCH2 | 9 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LUAD; PAAD; PRAD; STAD | miRNAWalker2 validate; MirTarget; miRNATAP | TCGA BLCA -0.217; TCGA BRCA -0.168; TCGA COAD -0.366; TCGA ESCA -0.259; TCGA HNSC -0.225; TCGA LUAD -0.135; TCGA PAAD -0.148; TCGA PRAD -0.075; TCGA STAD -0.349 |
hsa-miR-148b-3p | MECP2 | 12 cancers: BLCA; BRCA; COAD; ESCA; KIRC; KIRP; LUAD; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.13; TCGA BRCA -0.097; TCGA COAD -0.135; TCGA ESCA -0.139; TCGA KIRC -0.229; TCGA KIRP -0.205; TCGA LUAD -0.081; TCGA OV -0.124; TCGA PRAD -0.096; TCGA SARC -0.128; TCGA STAD -0.246; TCGA UCEC -0.206 |
hsa-miR-148b-3p | MKX | 12 cancers: BLCA; BRCA; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; SARC; THCA; STAD | miRNAWalker2 validate | TCGA BLCA -1.568; TCGA BRCA -0.33; TCGA ESCA -0.426; TCGA KIRC -0.737; TCGA KIRP -0.361; TCGA LIHC -0.516; TCGA LUAD -0.288; TCGA PAAD -0.662; TCGA PRAD -0.589; TCGA SARC -0.485; TCGA THCA -1.066; TCGA STAD -1.201 |
hsa-miR-148b-3p | MTMR10 | 10 cancers: BLCA; BRCA; CESC; HNSC; KIRC; KIRP; LIHC; PRAD; STAD; UCEC | miRNAWalker2 validate; MirTarget | TCGA BLCA -0.104; TCGA BRCA -0.129; TCGA CESC -0.125; TCGA HNSC -0.129; TCGA KIRC -0.226; TCGA KIRP -0.16; TCGA LIHC -0.197; TCGA PRAD -0.081; TCGA STAD -0.241; TCGA UCEC -0.186 |
hsa-miR-148b-3p | MYOM1 | 11 cancers: BLCA; BRCA; COAD; ESCA; HNSC; KIRC; KIRP; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -1.37; TCGA BRCA -0.89; TCGA COAD -1.203; TCGA ESCA -0.6; TCGA HNSC -1.151; TCGA KIRC -0.686; TCGA KIRP -0.807; TCGA PRAD -0.852; TCGA SARC -1.174; TCGA STAD -1.155; TCGA UCEC -0.689 |
hsa-miR-148b-3p | NCS1 | 12 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.659; TCGA CESC -0.537; TCGA COAD -0.586; TCGA ESCA -1.034; TCGA HNSC -0.159; TCGA LUAD -0.502; TCGA LUSC -0.341; TCGA OV -0.306; TCGA PRAD -0.596; TCGA SARC -0.561; TCGA STAD -1.004; TCGA UCEC -0.355 |
hsa-miR-148b-3p | NDN | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LGG; LUAD; LUSC; PRAD; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.688; TCGA BRCA -0.739; TCGA CESC -0.646; TCGA COAD -0.861; TCGA ESCA -0.689; TCGA HNSC -0.506; TCGA KIRP -0.362; TCGA LGG -0.224; TCGA LUAD -0.278; TCGA LUSC -0.282; TCGA PRAD -0.462; TCGA STAD -0.856; TCGA UCEC -0.872 |
hsa-miR-148b-3p | NIN | 10 cancers: BLCA; CESC; COAD; ESCA; LUAD; LUSC; PAAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.193; TCGA CESC -0.24; TCGA COAD -0.189; TCGA ESCA -0.544; TCGA LUAD -0.176; TCGA LUSC -0.202; TCGA PAAD -0.189; TCGA THCA -0.291; TCGA STAD -0.364; TCGA UCEC -0.084 |
hsa-miR-148b-3p | NOX4 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; THCA; UCEC | miRNAWalker2 validate | TCGA BLCA -0.452; TCGA CESC -0.52; TCGA COAD -1.018; TCGA ESCA -0.615; TCGA HNSC -0.259; TCGA LUAD -0.317; TCGA LUSC -0.473; TCGA PAAD -1.291; TCGA THCA -1.381; TCGA UCEC -0.155 |
hsa-miR-148b-3p | NR3C2 | 11 cancers: BLCA; BRCA; KIRC; KIRP; LIHC; LUAD; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.923; TCGA BRCA -0.589; TCGA KIRC -0.868; TCGA KIRP -0.613; TCGA LIHC -0.549; TCGA LUAD -0.323; TCGA OV -0.252; TCGA PRAD -0.174; TCGA SARC -0.302; TCGA STAD -0.51; TCGA UCEC -0.502 |
hsa-miR-148b-3p | NTN1 | 13 cancers: BLCA; BRCA; COAD; ESCA; KIRC; KIRP; LGG; LUAD; OV; PRAD; SARC; THCA; STAD | miRNAWalker2 validate; mirMAP | TCGA BLCA -1.184; TCGA BRCA -0.18; TCGA COAD -0.803; TCGA ESCA -0.814; TCGA KIRC -0.682; TCGA KIRP -0.839; TCGA LGG -0.188; TCGA LUAD -0.433; TCGA OV -0.394; TCGA PRAD -0.984; TCGA SARC -0.87; TCGA THCA -0.209; TCGA STAD -1.139 |
hsa-miR-148b-3p | PIK3CA | 9 cancers: BLCA; CESC; COAD; ESCA; LUAD; PAAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.097; TCGA CESC -0.244; TCGA COAD -0.118; TCGA ESCA -0.271; TCGA LUAD -0.093; TCGA PAAD -0.326; TCGA THCA -0.341; TCGA STAD -0.152; TCGA UCEC -0.121 |
hsa-miR-148b-3p | PLXDC2 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PAAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.769; TCGA BRCA -0.085; TCGA CESC -0.568; TCGA COAD -0.651; TCGA ESCA -0.874; TCGA HNSC -0.497; TCGA LIHC -0.392; TCGA LUAD -0.423; TCGA LUSC -0.327; TCGA PAAD -0.83; TCGA THCA -0.56; TCGA STAD -0.408; TCGA UCEC -0.174 |
hsa-miR-148b-3p | PPP2R3A | 12 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; PAAD; PRAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.357; TCGA CESC -0.55; TCGA COAD -0.342; TCGA ESCA -0.644; TCGA HNSC -0.438; TCGA LUAD -0.27; TCGA PAAD -0.464; TCGA PRAD -0.082; TCGA SARC -0.198; TCGA THCA -0.374; TCGA STAD -0.608; TCGA UCEC -0.288 |
hsa-miR-148b-3p | PRNP | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PRAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate; MirTarget | TCGA BLCA -0.567; TCGA BRCA -0.331; TCGA CESC -0.632; TCGA COAD -0.372; TCGA ESCA -0.719; TCGA HNSC -0.334; TCGA LUAD -0.368; TCGA LUSC -0.372; TCGA OV -0.142; TCGA PRAD -0.408; TCGA SARC -0.151; TCGA THCA -0.342; TCGA STAD -0.592; TCGA UCEC -0.6 |
hsa-miR-148b-3p | PTGER2 | 10 cancers: BLCA; BRCA; CESC; KIRP; LUAD; OV; PAAD; PRAD; THCA; UCEC | miRNAWalker2 validate | TCGA BLCA -0.434; TCGA BRCA -0.339; TCGA CESC -0.368; TCGA KIRP -0.818; TCGA LUAD -0.338; TCGA OV -0.395; TCGA PAAD -0.531; TCGA PRAD -0.761; TCGA THCA -0.955; TCGA UCEC -0.44 |
hsa-miR-148b-3p | RGMA | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LGG; LUAD; PAAD; PRAD; THCA; STAD; UCEC | miRNAWalker2 validate; miRNATAP | TCGA BLCA -1.497; TCGA BRCA -0.64; TCGA CESC -1.017; TCGA COAD -1.565; TCGA ESCA -1.477; TCGA KIRC -0.756; TCGA KIRP -0.914; TCGA LGG -0.119; TCGA LUAD -0.361; TCGA PAAD -0.943; TCGA PRAD -0.622; TCGA THCA -0.292; TCGA STAD -1.572; TCGA UCEC -0.678 |
hsa-miR-148b-3p | RNF150 | 12 cancers: BLCA; BRCA; COAD; ESCA; KIRC; KIRP; LIHC; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.92; TCGA BRCA -0.517; TCGA COAD -1.379; TCGA ESCA -1.212; TCGA KIRC -1.176; TCGA KIRP -0.995; TCGA LIHC -0.496; TCGA OV -0.197; TCGA PRAD -0.169; TCGA SARC -0.868; TCGA STAD -1.348; TCGA UCEC -0.288 |
hsa-miR-148b-3p | RTN4 | 14 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.119; TCGA BRCA -0.062; TCGA CESC -0.207; TCGA ESCA -0.276; TCGA HNSC -0.407; TCGA LIHC -0.424; TCGA LUAD -0.093; TCGA LUSC -0.276; TCGA OV -0.091; TCGA PAAD -0.17; TCGA PRAD -0.134; TCGA THCA -0.321; TCGA STAD -0.312; TCGA UCEC -0.184 |
hsa-miR-148b-3p | SESTD1 | 11 cancers: BLCA; COAD; ESCA; KIRP; OV; PAAD; PRAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate; MirTarget | TCGA BLCA -0.201; TCGA COAD -0.107; TCGA ESCA -0.204; TCGA KIRP -0.178; TCGA OV -0.431; TCGA PAAD -0.174; TCGA PRAD -0.151; TCGA SARC -0.293; TCGA THCA -0.459; TCGA STAD -0.329; TCGA UCEC -0.118 |
hsa-miR-148b-3p | SLC22A3 | 9 cancers: BLCA; BRCA; HNSC; LGG; PAAD; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -1.682; TCGA BRCA -0.867; TCGA HNSC -0.935; TCGA LGG -0.218; TCGA PAAD -1.014; TCGA PRAD -0.998; TCGA SARC -1.026; TCGA STAD -0.863; TCGA UCEC -1.377 |
hsa-miR-148b-3p | SNAI2 | 16 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.512; TCGA BRCA -0.36; TCGA CESC -1.204; TCGA COAD -0.347; TCGA ESCA -0.88; TCGA HNSC -0.281; TCGA KIRC -0.345; TCGA LIHC -0.544; TCGA LUAD -0.557; TCGA LUSC -0.48; TCGA OV -0.51; TCGA PAAD -0.948; TCGA PRAD -0.808; TCGA THCA -0.396; TCGA STAD -0.149; TCGA UCEC -0.725 |
hsa-miR-148b-3p | ST6GALNAC5 | 11 cancers: BLCA; CESC; COAD; ESCA; KIRC; KIRP; LUAD; PRAD; SARC; THCA; STAD | miRNAWalker2 validate | TCGA BLCA -1.063; TCGA CESC -0.453; TCGA COAD -1.012; TCGA ESCA -0.487; TCGA KIRC -0.62; TCGA KIRP -1.11; TCGA LUAD -0.711; TCGA PRAD -0.351; TCGA SARC -1.136; TCGA THCA -4.126; TCGA STAD -0.688 |
hsa-miR-148b-3p | STEAP4 | 12 cancers: BLCA; BRCA; COAD; HNSC; LIHC; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | miRNAWalker2 validate; MirTarget | TCGA BLCA -0.866; TCGA BRCA -0.529; TCGA COAD -0.732; TCGA HNSC -0.446; TCGA LIHC -0.922; TCGA LUAD -0.276; TCGA OV -0.379; TCGA PAAD -0.525; TCGA PRAD -0.38; TCGA THCA -0.777; TCGA STAD -0.831; TCGA UCEC -0.481 |
hsa-miR-148b-3p | SYNE1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; LUAD; OV; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.793; TCGA BRCA -0.291; TCGA CESC -0.264; TCGA COAD -0.621; TCGA ESCA -0.406; TCGA KIRP -0.3; TCGA LUAD -0.331; TCGA OV -0.188; TCGA PRAD -0.357; TCGA SARC -0.521; TCGA STAD -0.756; TCGA UCEC -0.739 |
hsa-miR-148b-3p | SYNE2 | 10 cancers: BLCA; BRCA; ESCA; KIRC; KIRP; LGG; LIHC; LUSC; PAAD; SARC | miRNAWalker2 validate | TCGA BLCA -0.18; TCGA BRCA -0.071; TCGA ESCA -0.212; TCGA KIRC -0.325; TCGA KIRP -0.3; TCGA LGG -0.177; TCGA LIHC -0.33; TCGA LUSC -0.135; TCGA PAAD -0.215; TCGA SARC -0.433 |
hsa-miR-148b-3p | TACC1 | 11 cancers: BLCA; BRCA; CESC; KIRC; LIHC; LUAD; PRAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.371; TCGA BRCA -0.379; TCGA CESC -0.379; TCGA KIRC -0.268; TCGA LIHC -0.392; TCGA LUAD -0.153; TCGA PRAD -0.395; TCGA SARC -0.499; TCGA THCA -0.116; TCGA STAD -0.603; TCGA UCEC -0.594 |
hsa-miR-148b-3p | TMEM47 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.376; TCGA BRCA -0.403; TCGA CESC -0.759; TCGA COAD -0.745; TCGA ESCA -0.72; TCGA HNSC -0.257; TCGA KIRC -0.238; TCGA KIRP -0.516; TCGA LIHC -0.881; TCGA LUAD -0.324; TCGA PAAD -0.437; TCGA PRAD -0.311; TCGA SARC -0.746; TCGA STAD -0.849; TCGA UCEC -0.701 |
hsa-miR-148b-3p | TXNIP | 11 cancers: BLCA; BRCA; ESCA; KIRC; KIRP; LGG; LUAD; PAAD; PRAD; STAD; UCEC | miRNAWalker2 validate; MirTarget | TCGA BLCA -0.247; TCGA BRCA -0.475; TCGA ESCA -0.305; TCGA KIRC -0.169; TCGA KIRP -0.312; TCGA LGG -0.111; TCGA LUAD -0.359; TCGA PAAD -0.219; TCGA PRAD -0.115; TCGA STAD -0.642; TCGA UCEC -0.439 |
hsa-miR-148b-3p | YPEL4 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; LGG; LUAD; LUSC; OV; PAAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.297; TCGA BRCA -0.714; TCGA CESC -0.377; TCGA COAD -0.791; TCGA ESCA -0.507; TCGA KIRP -0.508; TCGA LGG -0.431; TCGA LUAD -0.181; TCGA LUSC -0.227; TCGA OV -0.317; TCGA PAAD -0.441; TCGA THCA -1.2; TCGA STAD -0.527; TCGA UCEC -0.631 |
hsa-miR-148b-3p | ZDHHC15 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LUAD; PRAD; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BLCA -0.932; TCGA BRCA -0.338; TCGA CESC -1.003; TCGA COAD -1.198; TCGA ESCA -0.595; TCGA KIRC -1.103; TCGA KIRP -0.47; TCGA LUAD -0.712; TCGA PRAD -0.607; TCGA SARC -0.56; TCGA STAD -0.992; TCGA UCEC -0.743 |
hsa-miR-148b-3p | RBM24 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -1.537; TCGA BRCA -0.474; TCGA CESC -0.848; TCGA COAD -0.779; TCGA ESCA -0.685; TCGA HNSC -1.355; TCGA LIHC -0.417; TCGA LUAD -0.517; TCGA PRAD -0.879; TCGA SARC -1.544; TCGA STAD -1.386; TCGA UCEC -0.728 |
hsa-miR-148b-3p | TCF4 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; LGG; LIHC; LUAD; OV; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.652; TCGA BRCA -0.31; TCGA CESC -0.565; TCGA COAD -0.609; TCGA ESCA -0.398; TCGA LGG -0.242; TCGA LIHC -0.217; TCGA LUAD -0.322; TCGA OV -0.283; TCGA PAAD -0.459; TCGA PRAD -0.261; TCGA STAD -0.414; TCGA UCEC -0.41 |
hsa-miR-148b-3p | ROBO1 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; LGG; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.419; TCGA BRCA -0.14; TCGA CESC -0.339; TCGA COAD -0.528; TCGA ESCA -0.296; TCGA KIRP -0.431; TCGA LGG -0.209; TCGA LUAD -0.165; TCGA OV -0.276; TCGA PAAD -0.611; TCGA PRAD -0.564; TCGA THCA -0.303; TCGA STAD -0.405; TCGA UCEC -0.173 |
hsa-miR-148b-3p | BAG3 | 11 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LUSC; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.316; TCGA BRCA -0.071; TCGA CESC -0.421; TCGA ESCA -0.273; TCGA HNSC -0.228; TCGA LUSC -0.224; TCGA PAAD -0.41; TCGA PRAD -0.185; TCGA THCA -0.632; TCGA STAD -0.43; TCGA UCEC -0.105 |
hsa-miR-148b-3p | KLF4 | 10 cancers: BLCA; BRCA; CESC; HNSC; KIRC; KIRP; LIHC; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.433; TCGA BRCA -0.572; TCGA CESC -0.356; TCGA HNSC -0.334; TCGA KIRC -0.22; TCGA KIRP -0.267; TCGA LIHC -0.385; TCGA PRAD -0.263; TCGA THCA -0.223; TCGA UCEC -0.388 |
hsa-miR-148b-3p | FAM168B | 9 cancers: BLCA; BRCA; CESC; KIRC; OV; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.246; TCGA BRCA -0.072; TCGA CESC -0.2; TCGA KIRC -0.091; TCGA OV -0.197; TCGA PRAD -0.142; TCGA SARC -0.26; TCGA STAD -0.212; TCGA UCEC -0.183 |
hsa-miR-148b-3p | GREM2 | 16 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUSC; OV; PAAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -1.083; TCGA BRCA -1.084; TCGA CESC -0.652; TCGA COAD -1.883; TCGA ESCA -1.228; TCGA HNSC -0.638; TCGA KIRC -1.306; TCGA KIRP -0.927; TCGA LIHC -0.991; TCGA LUSC -0.442; TCGA OV -0.637; TCGA PAAD -0.903; TCGA SARC -1.171; TCGA THCA -0.636; TCGA STAD -1.386; TCGA UCEC -0.779 |
hsa-miR-148b-3p | TGFB2 | 16 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.765; TCGA BRCA -0.423; TCGA CESC -0.652; TCGA COAD -0.557; TCGA ESCA -0.468; TCGA KIRC -0.36; TCGA KIRP -0.67; TCGA LIHC -0.308; TCGA LUAD -0.598; TCGA LUSC -0.355; TCGA OV -0.302; TCGA PAAD -0.904; TCGA PRAD -0.473; TCGA THCA -0.819; TCGA STAD -0.257; TCGA UCEC -0.405 |
hsa-miR-148b-3p | RAB34 | 10 cancers: BLCA; BRCA; COAD; ESCA; KIRP; PAAD; PRAD; SARC; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.202; TCGA BRCA -0.082; TCGA COAD -0.817; TCGA ESCA -0.353; TCGA KIRP -0.301; TCGA PAAD -0.806; TCGA PRAD -0.659; TCGA SARC -0.252; TCGA THCA -0.501; TCGA STAD -0.344 |
hsa-miR-148b-3p | DNER | 9 cancers: BLCA; BRCA; COAD; KIRP; LGG; OV; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.495; TCGA BRCA -0.477; TCGA COAD -1.101; TCGA KIRP -1.138; TCGA LGG -0.161; TCGA OV -0.535; TCGA PRAD -0.349; TCGA STAD -0.766; TCGA UCEC -0.556 |
hsa-miR-148b-3p | PTPN14 | 10 cancers: BLCA; BRCA; ESCA; KIRP; LUAD; PAAD; PRAD; SARC; THCA; STAD | MirTarget | TCGA BLCA -0.099; TCGA BRCA -0.262; TCGA ESCA -0.44; TCGA KIRP -0.362; TCGA LUAD -0.166; TCGA PAAD -0.71; TCGA PRAD -0.173; TCGA SARC -0.373; TCGA THCA -0.445; TCGA STAD -0.369 |
hsa-miR-148b-3p | CHD9 | 10 cancers: BLCA; CESC; ESCA; LGG; LIHC; LUAD; LUSC; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.203; TCGA CESC -0.305; TCGA ESCA -0.355; TCGA LGG -0.168; TCGA LIHC -0.359; TCGA LUAD -0.13; TCGA LUSC -0.244; TCGA PRAD -0.118; TCGA STAD -0.257; TCGA UCEC -0.222 |
hsa-miR-148b-3p | ADAM23 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.762; TCGA BRCA -0.433; TCGA CESC -1.202; TCGA COAD -1.135; TCGA ESCA -1.419; TCGA KIRC -0.731; TCGA KIRP -0.527; TCGA LUAD -0.426; TCGA OV -0.455; TCGA PAAD -0.546; TCGA PRAD -0.62; TCGA THCA -1.112; TCGA STAD -0.62; TCGA UCEC -0.337 |
hsa-miR-148b-3p | CYTH3 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.378; TCGA BRCA -0.281; TCGA CESC -0.35; TCGA COAD -0.345; TCGA ESCA -0.266; TCGA HNSC -0.295; TCGA KIRC -0.345; TCGA KIRP -0.257; TCGA LIHC -0.312; TCGA LUAD -0.477; TCGA LUSC -0.412; TCGA OV -0.231; TCGA PAAD -0.473; TCGA STAD -0.329; TCGA UCEC -0.235 |
hsa-miR-148b-3p | PRICKLE2 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PAAD; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.786; TCGA BRCA -0.368; TCGA CESC -0.429; TCGA COAD -0.704; TCGA ESCA -0.744; TCGA HNSC -0.167; TCGA LIHC -0.464; TCGA LUAD -0.346; TCGA LUSC -0.415; TCGA OV -0.198; TCGA PAAD -0.293; TCGA PRAD -0.759; TCGA SARC -0.473; TCGA STAD -1.116; TCGA UCEC -0.566 |
hsa-miR-148b-3p | ABCA1 | 9 cancers: BLCA; BRCA; COAD; ESCA; LGG; LUAD; LUSC; THCA; STAD | MirTarget | TCGA BLCA -0.192; TCGA BRCA -0.134; TCGA COAD -0.248; TCGA ESCA -0.48; TCGA LGG -0.119; TCGA LUAD -0.291; TCGA LUSC -0.324; TCGA THCA -0.162; TCGA STAD -0.21 |
hsa-miR-148b-3p | SSBP2 | 10 cancers: BLCA; CESC; COAD; ESCA; KIRP; LUAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.369; TCGA CESC -0.472; TCGA COAD -0.439; TCGA ESCA -0.531; TCGA KIRP -0.258; TCGA LUAD -0.256; TCGA PRAD -0.26; TCGA SARC -0.223; TCGA STAD -0.761; TCGA UCEC -0.232 |
hsa-miR-148b-3p | ERRFI1 | 10 cancers: BLCA; BRCA; CESC; KIRP; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.156; TCGA BRCA -0.205; TCGA CESC -0.255; TCGA KIRP -0.68; TCGA LUAD -0.245; TCGA LUSC -0.355; TCGA OV -0.265; TCGA PRAD -0.279; TCGA STAD -0.347; TCGA UCEC -0.161 |
hsa-miR-148b-3p | SCN9A | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; OV; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.389; TCGA BRCA -0.661; TCGA CESC -0.599; TCGA COAD -0.821; TCGA ESCA -0.977; TCGA KIRP -0.624; TCGA OV -0.537; TCGA PRAD -0.524; TCGA THCA -1.078; TCGA STAD -0.748 |
hsa-miR-148b-3p | ARRDC3 | 9 cancers: BLCA; BRCA; CESC; ESCA; KIRP; LUAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.191; TCGA BRCA -0.259; TCGA CESC -0.216; TCGA ESCA -0.322; TCGA KIRP -0.272; TCGA LUAD -0.222; TCGA PRAD -0.392; TCGA STAD -0.285; TCGA UCEC -0.128 |
hsa-miR-148b-3p | ADCY2 | 12 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LGG; LUAD; OV; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -1.245; TCGA BRCA -0.406; TCGA COAD -1.244; TCGA ESCA -1.351; TCGA HNSC -0.894; TCGA LGG -0.237; TCGA LUAD -0.334; TCGA OV -0.992; TCGA PRAD -0.459; TCGA SARC -1.193; TCGA STAD -1.136; TCGA UCEC -0.959 |
hsa-miR-148b-3p | PPARGC1A | 10 cancers: BLCA; BRCA; HNSC; KIRC; LIHC; OV; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -1.418; TCGA BRCA -0.553; TCGA HNSC -0.656; TCGA KIRC -0.674; TCGA LIHC -0.521; TCGA OV -1.216; TCGA PRAD -0.924; TCGA SARC -1.258; TCGA STAD -0.497; TCGA UCEC -0.402 |
hsa-miR-148b-3p | ATP1A2 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LIHC; LUAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -1.954; TCGA BRCA -1.437; TCGA CESC -0.949; TCGA COAD -2.372; TCGA ESCA -1.774; TCGA HNSC -1.704; TCGA KIRC -0.772; TCGA LIHC -0.619; TCGA LUAD -0.483; TCGA PRAD -1.116; TCGA SARC -0.803; TCGA THCA -0.294; TCGA STAD -2.218; TCGA UCEC -0.775 |
hsa-miR-148b-3p | DNAJC18 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LUAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.328; TCGA BRCA -0.208; TCGA CESC -0.175; TCGA COAD -0.165; TCGA ESCA -0.425; TCGA KIRC -0.248; TCGA LUAD -0.1; TCGA PRAD -0.316; TCGA SARC -0.13; TCGA STAD -0.681; TCGA UCEC -0.35 |
hsa-miR-148b-3p | SLC24A3 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LGG; LIHC; LUSC; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -1.275; TCGA BRCA -0.289; TCGA CESC -1.035; TCGA COAD -0.995; TCGA ESCA -1.021; TCGA HNSC -1.178; TCGA KIRC -0.761; TCGA KIRP -0.976; TCGA LGG -0.47; TCGA LIHC -0.68; TCGA LUSC -0.767; TCGA PRAD -0.848; TCGA SARC -1.564; TCGA STAD -0.517; TCGA UCEC -1.029 |
hsa-miR-148b-3p | ATP2B4 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.622; TCGA BRCA -0.131; TCGA CESC -0.387; TCGA COAD -0.657; TCGA ESCA -0.821; TCGA KIRP -0.127; TCGA LUAD -0.13; TCGA LUSC -0.188; TCGA PAAD -0.402; TCGA PRAD -0.822; TCGA SARC -0.576; TCGA THCA -0.482; TCGA STAD -0.896; TCGA UCEC -0.272 |
hsa-miR-148b-3p | PTEN | 9 cancers: BLCA; ESCA; HNSC; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.169; TCGA ESCA -0.109; TCGA HNSC -0.118; TCGA LIHC -0.179; TCGA LUAD -0.131; TCGA PAAD -0.164; TCGA PRAD -0.123; TCGA STAD -0.195; TCGA UCEC -0.217 |
hsa-miR-148b-3p | FAM43A | 12 cancers: BLCA; BRCA; CESC; HNSC; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; THCA; UCEC | MirTarget | TCGA BLCA -0.516; TCGA BRCA -0.466; TCGA CESC -0.395; TCGA HNSC -0.322; TCGA KIRC -0.408; TCGA KIRP -0.508; TCGA LIHC -0.476; TCGA LUAD -0.159; TCGA PAAD -0.256; TCGA PRAD -0.168; TCGA THCA -1.138; TCGA UCEC -0.29 |
hsa-miR-148b-3p | RNF38 | 11 cancers: BLCA; CESC; ESCA; KIRC; KIRP; LGG; LUAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.142; TCGA CESC -0.227; TCGA ESCA -0.143; TCGA KIRC -0.191; TCGA KIRP -0.104; TCGA LGG -0.096; TCGA LUAD -0.147; TCGA PRAD -0.075; TCGA SARC -0.416; TCGA STAD -0.332; TCGA UCEC -0.206 |
hsa-miR-148b-3p | TNRC6C | 10 cancers: BLCA; BRCA; COAD; KIRC; KIRP; LGG; LIHC; OV; STAD; UCEC | MirTarget | TCGA BLCA -0.231; TCGA BRCA -0.087; TCGA COAD -0.238; TCGA KIRC -0.179; TCGA KIRP -0.265; TCGA LGG -0.154; TCGA LIHC -0.147; TCGA OV -0.219; TCGA STAD -0.335; TCGA UCEC -0.223 |
hsa-miR-148b-3p | PQLC3 | 10 cancers: BLCA; COAD; KIRP; LIHC; LUAD; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.209; TCGA COAD -0.165; TCGA KIRP -0.176; TCGA LIHC -0.169; TCGA LUAD -0.111; TCGA PAAD -0.566; TCGA PRAD -0.105; TCGA THCA -0.24; TCGA STAD -0.174; TCGA UCEC -0.124 |
hsa-miR-148b-3p | CCDC6 | 9 cancers: BLCA; CESC; HNSC; LUAD; PAAD; PRAD; SARC; THCA; STAD | MirTarget | TCGA BLCA -0.173; TCGA CESC -0.128; TCGA HNSC -0.093; TCGA LUAD -0.173; TCGA PAAD -0.272; TCGA PRAD -0.082; TCGA SARC -0.429; TCGA THCA -0.165; TCGA STAD -0.114 |
hsa-miR-148b-3p | BACH2 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LGG; LIHC; LUAD; OV; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -1.061; TCGA BRCA -0.514; TCGA CESC -0.471; TCGA COAD -0.854; TCGA ESCA -0.638; TCGA KIRC -0.418; TCGA KIRP -0.59; TCGA LGG -0.149; TCGA LIHC -0.355; TCGA LUAD -0.302; TCGA OV -0.427; TCGA PRAD -0.252; TCGA THCA -0.948; TCGA STAD -0.441; TCGA UCEC -0.627 |
hsa-miR-148b-3p | CHST1 | 10 cancers: BLCA; CESC; COAD; ESCA; KIRC; KIRP; LGG; LUSC; PAAD; THCA | MirTarget | TCGA BLCA -0.469; TCGA CESC -0.617; TCGA COAD -0.502; TCGA ESCA -0.362; TCGA KIRC -0.299; TCGA KIRP -0.668; TCGA LGG -0.163; TCGA LUSC -0.316; TCGA PAAD -0.521; TCGA THCA -0.299 |
hsa-miR-148b-3p | PLEKHM3 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; LIHC; OV; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.107; TCGA BRCA -0.224; TCGA CESC -0.218; TCGA COAD -0.156; TCGA ESCA -0.183; TCGA LIHC -0.23; TCGA OV -0.113; TCGA PRAD -0.122; TCGA SARC -0.131; TCGA STAD -0.355; TCGA UCEC -0.241 |
hsa-miR-148b-3p | ADAMTS15 | 12 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LIHC; LUAD; LUSC; OV; PAAD; PRAD; STAD | MirTarget | TCGA BLCA -1.466; TCGA BRCA -0.554; TCGA CESC -0.774; TCGA ESCA -0.926; TCGA KIRC -0.466; TCGA LIHC -0.55; TCGA LUAD -0.327; TCGA LUSC -0.829; TCGA OV -0.3; TCGA PAAD -0.836; TCGA PRAD -0.273; TCGA STAD -0.325 |
hsa-miR-148b-3p | GAP43 | 10 cancers: BLCA; BRCA; COAD; ESCA; LUAD; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -1.024; TCGA BRCA -0.359; TCGA COAD -1.992; TCGA ESCA -0.822; TCGA LUAD -0.407; TCGA PAAD -0.6; TCGA PRAD -0.638; TCGA THCA -3.455; TCGA STAD -1.272; TCGA UCEC -0.513 |
hsa-miR-148b-3p | SLC8A1 | 12 cancers: BLCA; CESC; COAD; ESCA; KIRP; LIHC; LUAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.717; TCGA CESC -0.367; TCGA COAD -0.576; TCGA ESCA -0.357; TCGA KIRP -0.333; TCGA LIHC -0.309; TCGA LUAD -0.207; TCGA PRAD -0.891; TCGA SARC -0.704; TCGA THCA -0.645; TCGA STAD -0.673; TCGA UCEC -0.593 |
hsa-miR-148b-3p | PBXIP1 | 10 cancers: BLCA; BRCA; COAD; KIRC; KIRP; LIHC; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.186; TCGA BRCA -0.168; TCGA COAD -0.349; TCGA KIRC -0.156; TCGA KIRP -0.179; TCGA LIHC -0.161; TCGA PRAD -0.222; TCGA SARC -0.484; TCGA STAD -0.414; TCGA UCEC -0.218 |
hsa-miR-148b-3p | SYNC | 12 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LUAD; LUSC; OV; PAAD; PRAD; SARC; STAD | MirTarget | TCGA BLCA -0.566; TCGA BRCA -0.21; TCGA COAD -1.174; TCGA ESCA -1.038; TCGA HNSC -0.792; TCGA LUAD -0.411; TCGA LUSC -0.693; TCGA OV -0.26; TCGA PAAD -0.826; TCGA PRAD -0.563; TCGA SARC -0.664; TCGA STAD -0.393 |
hsa-miR-148b-3p | SACS | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; LUAD; LUSC; PAAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.743; TCGA BRCA -0.141; TCGA CESC -0.361; TCGA COAD -0.225; TCGA ESCA -0.268; TCGA LUAD -0.13; TCGA LUSC -0.181; TCGA PAAD -0.279; TCGA PRAD -0.127; TCGA SARC -0.198; TCGA STAD -0.368; TCGA UCEC -0.155 |
hsa-miR-148b-3p | STK38L | 10 cancers: BLCA; CESC; ESCA; KIRC; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.319; TCGA CESC -0.362; TCGA ESCA -0.225; TCGA KIRC -0.173; TCGA PAAD -0.36; TCGA PRAD -0.203; TCGA SARC -0.396; TCGA THCA -0.083; TCGA STAD -0.207; TCGA UCEC -0.178 |
hsa-miR-148b-3p | EGR3 | 14 cancers: BLCA; BRCA; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -1.1; TCGA BRCA -0.758; TCGA COAD -0.442; TCGA ESCA -0.549; TCGA HNSC -0.426; TCGA KIRC -0.552; TCGA KIRP -0.459; TCGA LIHC -0.628; TCGA LUAD -0.297; TCGA LUSC -0.511; TCGA OV -0.28; TCGA PRAD -0.602; TCGA STAD -0.375; TCGA UCEC -0.23 |
hsa-miR-148b-3p | NOG | 12 cancers: BLCA; BRCA; ESCA; HNSC; KIRC; KIRP; LGG; LUSC; OV; PAAD; PRAD; STAD | MirTarget | TCGA BLCA -0.708; TCGA BRCA -0.582; TCGA ESCA -0.629; TCGA HNSC -0.755; TCGA KIRC -0.669; TCGA KIRP -1.082; TCGA LGG -0.868; TCGA LUSC -0.402; TCGA OV -0.592; TCGA PAAD -0.675; TCGA PRAD -0.718; TCGA STAD -0.737 |
hsa-miR-148b-3p | FOXF1 | 13 cancers: BLCA; BRCA; COAD; ESCA; KIRP; LIHC; LUAD; OV; PAAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.892; TCGA BRCA -0.202; TCGA COAD -0.594; TCGA ESCA -0.37; TCGA KIRP -0.436; TCGA LIHC -0.593; TCGA LUAD -0.253; TCGA OV -0.225; TCGA PAAD -0.508; TCGA PRAD -0.884; TCGA SARC -0.577; TCGA STAD -0.832; TCGA UCEC -0.226 |
hsa-miR-148b-3p | PANX1 | 9 cancers: BLCA; ESCA; HNSC; KIRP; LIHC; LUAD; LUSC; PAAD; THCA | MirTarget | TCGA BLCA -0.22; TCGA ESCA -0.15; TCGA HNSC -0.24; TCGA KIRP -0.119; TCGA LIHC -0.37; TCGA LUAD -0.089; TCGA LUSC -0.178; TCGA PAAD -0.39; TCGA THCA -0.117 |
hsa-miR-148b-3p | PEA15 | 12 cancers: BLCA; CESC; COAD; ESCA; HNSC; LGG; LUAD; LUSC; PAAD; SARC; THCA; STAD | MirTarget | TCGA BLCA -0.133; TCGA CESC -0.161; TCGA COAD -0.289; TCGA ESCA -0.294; TCGA HNSC -0.168; TCGA LGG -0.268; TCGA LUAD -0.21; TCGA LUSC -0.193; TCGA PAAD -0.321; TCGA SARC -0.14; TCGA THCA -0.41; TCGA STAD -0.228 |
hsa-miR-148b-3p | ACVR1 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.378; TCGA BRCA -0.102; TCGA CESC -0.32; TCGA COAD -0.142; TCGA ESCA -0.284; TCGA HNSC -0.183; TCGA LUAD -0.14; TCGA LUSC -0.265; TCGA PAAD -0.312; TCGA PRAD -0.181; TCGA SARC -0.266; TCGA THCA -0.504; TCGA STAD -0.131; TCGA UCEC -0.078 |
hsa-miR-148b-3p | ANK2 | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -1.438; TCGA BRCA -0.717; TCGA CESC -0.781; TCGA COAD -1.28; TCGA ESCA -0.832; TCGA HNSC -0.343; TCGA PRAD -0.843; TCGA SARC -0.279; TCGA STAD -1.188; TCGA UCEC -0.497 |
hsa-miR-148b-3p | GLIPR2 | 10 cancers: BLCA; BRCA; COAD; ESCA; LUAD; PAAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.852; TCGA BRCA -0.246; TCGA COAD -0.48; TCGA ESCA -0.441; TCGA LUAD -0.463; TCGA PAAD -0.657; TCGA SARC -0.284; TCGA THCA -0.377; TCGA STAD -0.408; TCGA UCEC -0.347 |
hsa-miR-148b-3p | JMY | 9 cancers: BLCA; CESC; ESCA; KIRC; KIRP; LGG; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.112; TCGA CESC -0.297; TCGA ESCA -0.353; TCGA KIRC -0.415; TCGA KIRP -0.222; TCGA LGG -0.083; TCGA SARC -0.186; TCGA STAD -0.395; TCGA UCEC -0.238 |
hsa-miR-148b-3p | MEOX2 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; OV; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -1.696; TCGA BRCA -0.946; TCGA CESC -0.932; TCGA COAD -1.576; TCGA ESCA -0.892; TCGA KIRC -0.415; TCGA KIRP -0.427; TCGA LIHC -0.354; TCGA LUAD -0.56; TCGA OV -0.632; TCGA PAAD -1.058; TCGA PRAD -0.594; TCGA STAD -1.19; TCGA UCEC -1.232 |
hsa-miR-148b-3p | TRAK2 | 9 cancers: BLCA; ESCA; KIRC; LUAD; PAAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.333; TCGA ESCA -0.163; TCGA KIRC -0.182; TCGA LUAD -0.112; TCGA PAAD -0.162; TCGA SARC -0.197; TCGA THCA -0.263; TCGA STAD -0.264; TCGA UCEC -0.127 |
hsa-miR-148b-3p | DMPK | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; OV; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.407; TCGA BRCA -0.27; TCGA CESC -0.32; TCGA COAD -0.35; TCGA ESCA -0.488; TCGA HNSC -0.461; TCGA KIRC -0.497; TCGA KIRP -0.305; TCGA LUAD -0.133; TCGA LUSC -0.31; TCGA OV -0.204; TCGA PRAD -0.378; TCGA SARC -0.606; TCGA STAD -1.038; TCGA UCEC -0.304 |
hsa-miR-148b-3p | F3 | 9 cancers: BLCA; BRCA; HNSC; KIRC; KIRP; LIHC; LUSC; PAAD; PRAD | MirTarget | TCGA BLCA -0.795; TCGA BRCA -0.662; TCGA HNSC -0.332; TCGA KIRC -0.862; TCGA KIRP -1.027; TCGA LIHC -0.362; TCGA LUSC -0.372; TCGA PAAD -0.698; TCGA PRAD -0.857 |
hsa-miR-148b-3p | GPCPD1 | 10 cancers: BLCA; BRCA; ESCA; KIRC; KIRP; LGG; LIHC; SARC; THCA; STAD | MirTarget | TCGA BLCA -0.196; TCGA BRCA -0.151; TCGA ESCA -0.288; TCGA KIRC -0.111; TCGA KIRP -0.193; TCGA LGG -0.078; TCGA LIHC -0.197; TCGA SARC -0.115; TCGA THCA -0.118; TCGA STAD -0.294 |
hsa-miR-148b-3p | ZMAT3 | 10 cancers: BLCA; BRCA; CESC; ESCA; LIHC; OV; PAAD; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.273; TCGA BRCA -0.083; TCGA CESC -0.357; TCGA ESCA -0.148; TCGA LIHC -0.409; TCGA OV -0.161; TCGA PAAD -0.27; TCGA PRAD -0.139; TCGA THCA -0.884; TCGA STAD -0.228 |
hsa-miR-148b-3p | MMP13 | 11 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; PAAD; PRAD; THCA | MirTarget | TCGA BLCA -0.449; TCGA CESC -1.635; TCGA COAD -0.656; TCGA ESCA -1.486; TCGA HNSC -0.794; TCGA LUAD -1.007; TCGA LUSC -1.414; TCGA OV -0.972; TCGA PAAD -1.377; TCGA PRAD -0.485; TCGA THCA -3.32 |
hsa-miR-148b-3p | TEAD1 | 13 cancers: BLCA; BRCA; COAD; ESCA; LGG; LIHC; LUAD; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.418; TCGA BRCA -0.139; TCGA COAD -0.175; TCGA ESCA -0.213; TCGA LGG -0.119; TCGA LIHC -0.151; TCGA LUAD -0.249; TCGA PAAD -0.171; TCGA PRAD -0.373; TCGA SARC -0.381; TCGA THCA -0.421; TCGA STAD -0.62; TCGA UCEC -0.215 |
hsa-miR-148b-3p | ADPRH | 9 cancers: BLCA; BRCA; COAD; LUAD; PAAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.366; TCGA BRCA -0.09; TCGA COAD -0.338; TCGA LUAD -0.252; TCGA PAAD -0.303; TCGA PRAD -0.183; TCGA SARC -0.323; TCGA STAD -0.239; TCGA UCEC -0.237 |
hsa-miR-148b-3p | S1PR1 | 11 cancers: BLCA; BRCA; CESC; COAD; KIRP; LIHC; LUAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.598; TCGA BRCA -0.605; TCGA CESC -0.406; TCGA COAD -0.506; TCGA KIRP -0.249; TCGA LIHC -0.435; TCGA LUAD -0.201; TCGA PRAD -0.207; TCGA THCA -0.277; TCGA STAD -0.481; TCGA UCEC -0.573 |
hsa-miR-148b-3p | CFL2 | 11 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LIHC; LUAD; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.951; TCGA BRCA -0.273; TCGA COAD -0.635; TCGA ESCA -0.534; TCGA HNSC -0.403; TCGA LIHC -0.236; TCGA LUAD -0.152; TCGA PRAD -0.567; TCGA SARC -0.745; TCGA STAD -1.145; TCGA UCEC -0.45 |
hsa-miR-148b-3p | MTMR9 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRC; LGG; LIHC; OV; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.164; TCGA BRCA -0.07; TCGA CESC -0.287; TCGA COAD -0.185; TCGA HNSC -0.084; TCGA KIRC -0.129; TCGA LGG -0.061; TCGA LIHC -0.18; TCGA OV -0.129; TCGA SARC -0.174; TCGA STAD -0.27; TCGA UCEC -0.21 |
hsa-miR-148b-3p | GADD45A | 14 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; LIHC; LUAD; LUSC; PAAD; PRAD; SARC; THCA; UCEC | MirTarget | TCGA BLCA -0.242; TCGA BRCA -0.231; TCGA CESC -0.526; TCGA ESCA -0.215; TCGA HNSC -0.298; TCGA KIRC -0.218; TCGA LIHC -0.533; TCGA LUAD -0.34; TCGA LUSC -0.367; TCGA PAAD -0.274; TCGA PRAD -0.246; TCGA SARC -0.544; TCGA THCA -0.46; TCGA UCEC -0.212 |
hsa-miR-148b-3p | MIER1 | 9 cancers: BLCA; CESC; LIHC; LUAD; PAAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.085; TCGA CESC -0.138; TCGA LIHC -0.213; TCGA LUAD -0.15; TCGA PAAD -0.101; TCGA SARC -0.21; TCGA THCA -0.051; TCGA STAD -0.135; TCGA UCEC -0.158 |
hsa-miR-148b-3p | LTBP1 | 15 cancers: BLCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.492; TCGA CESC -0.376; TCGA COAD -0.861; TCGA ESCA -0.871; TCGA HNSC -0.243; TCGA KIRC -0.594; TCGA KIRP -0.718; TCGA LUAD -0.271; TCGA LUSC -0.225; TCGA PAAD -0.896; TCGA PRAD -0.509; TCGA SARC -0.464; TCGA THCA -1.188; TCGA STAD -0.703; TCGA UCEC -0.239 |
hsa-miR-148b-3p | SCARA3 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.866; TCGA BRCA -0.329; TCGA CESC -0.894; TCGA COAD -0.856; TCGA ESCA -0.753; TCGA KIRC -0.678; TCGA KIRP -0.816; TCGA PAAD -0.697; TCGA PRAD -0.668; TCGA SARC -0.494; TCGA THCA -1.356; TCGA STAD -0.865; TCGA UCEC -0.397 |
hsa-miR-148b-3p | PDE5A | 16 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; OV; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.772; TCGA BRCA -0.38; TCGA CESC -0.342; TCGA COAD -0.49; TCGA ESCA -0.63; TCGA KIRC -0.318; TCGA KIRP -0.247; TCGA LIHC -0.328; TCGA LUAD -0.36; TCGA OV -0.208; TCGA PAAD -0.326; TCGA PRAD -0.85; TCGA SARC -0.815; TCGA THCA -1.725; TCGA STAD -0.838; TCGA UCEC -0.503 |
hsa-miR-148b-3p | FLYWCH1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; LGG; LIHC; LUSC; OV; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.178; TCGA BRCA -0.196; TCGA CESC -0.17; TCGA COAD -0.173; TCGA ESCA -0.36; TCGA LGG -0.119; TCGA LIHC -0.162; TCGA LUSC -0.275; TCGA OV -0.21; TCGA SARC -0.219; TCGA STAD -0.218; TCGA UCEC -0.128 |
hsa-miR-148b-3p | NDST1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; OV; STAD; UCEC | mirMAP | TCGA BLCA -0.176; TCGA BRCA -0.166; TCGA CESC -0.388; TCGA COAD -0.224; TCGA ESCA -0.364; TCGA HNSC -0.202; TCGA KIRC -0.141; TCGA LUAD -0.143; TCGA LUSC -0.292; TCGA OV -0.26; TCGA STAD -0.162; TCGA UCEC -0.095 |
hsa-miR-148b-3p | PITPNM3 | 16 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; PRAD; SARC; THCA; STAD | mirMAP | TCGA BLCA -0.492; TCGA BRCA -0.182; TCGA CESC -0.378; TCGA ESCA -0.675; TCGA HNSC -0.23; TCGA KIRC -0.733; TCGA KIRP -0.711; TCGA LIHC -0.74; TCGA LUAD -0.324; TCGA LUSC -0.278; TCGA OV -0.24; TCGA PAAD -0.351; TCGA PRAD -0.348; TCGA SARC -0.768; TCGA THCA -0.116; TCGA STAD -0.893 |
hsa-miR-148b-3p | TTC28 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; LGG; OV; PAAD; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.205; TCGA BRCA -0.372; TCGA CESC -0.362; TCGA COAD -0.636; TCGA ESCA -0.365; TCGA LGG -0.073; TCGA OV -0.224; TCGA PAAD -0.271; TCGA PRAD -0.425; TCGA SARC -0.238; TCGA STAD -0.633; TCGA UCEC -0.143 |
hsa-miR-148b-3p | SNPH | 10 cancers: BLCA; BRCA; COAD; HNSC; KIRC; LUSC; PRAD; SARC; STAD; UCEC | mirMAP; miRNATAP | TCGA BLCA -0.289; TCGA BRCA -0.189; TCGA COAD -0.386; TCGA HNSC -0.471; TCGA KIRC -0.183; TCGA LUSC -0.25; TCGA PRAD -0.201; TCGA SARC -0.259; TCGA STAD -0.372; TCGA UCEC -0.437 |
hsa-miR-148b-3p | XYLT1 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LIHC; LUSC; OV; PAAD; PRAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.631; TCGA BRCA -0.192; TCGA CESC -0.42; TCGA COAD -0.32; TCGA ESCA -0.336; TCGA KIRC -0.25; TCGA LIHC -0.24; TCGA LUSC -0.396; TCGA OV -0.522; TCGA PAAD -0.353; TCGA PRAD -0.304; TCGA SARC -0.287; TCGA THCA -0.144; TCGA STAD -0.286; TCGA UCEC -0.732 |
hsa-miR-148b-3p | MPP2 | 10 cancers: BLCA; COAD; ESCA; KIRC; KIRP; LUAD; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.328; TCGA COAD -0.708; TCGA ESCA -0.584; TCGA KIRC -1.162; TCGA KIRP -0.513; TCGA LUAD -0.299; TCGA PRAD -0.924; TCGA SARC -0.531; TCGA STAD -0.734; TCGA UCEC -0.413 |
hsa-miR-148b-3p | SRF | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; KIRP; LUSC; PAAD; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.365; TCGA BRCA -0.181; TCGA CESC -0.198; TCGA COAD -0.167; TCGA HNSC -0.161; TCGA KIRP -0.204; TCGA LUSC -0.142; TCGA PAAD -0.114; TCGA PRAD -0.32; TCGA SARC -0.514; TCGA STAD -0.401; TCGA UCEC -0.257 |
hsa-miR-148b-3p | CHD5 | 10 cancers: BLCA; COAD; ESCA; KIRC; KIRP; PRAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.639; TCGA COAD -0.633; TCGA ESCA -0.705; TCGA KIRC -1.864; TCGA KIRP -1.482; TCGA PRAD -0.792; TCGA SARC -0.827; TCGA THCA -0.689; TCGA STAD -0.922; TCGA UCEC -0.401 |
hsa-miR-148b-3p | PDLIM2 | 16 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LGG; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.598; TCGA BRCA -0.26; TCGA CESC -0.485; TCGA COAD -0.39; TCGA ESCA -0.388; TCGA HNSC -0.562; TCGA LGG -0.103; TCGA LIHC -0.305; TCGA LUAD -0.327; TCGA LUSC -0.334; TCGA OV -0.271; TCGA PAAD -0.286; TCGA PRAD -0.224; TCGA THCA -0.322; TCGA STAD -0.173; TCGA UCEC -0.243 |
hsa-miR-148b-3p | SEPT7 | 12 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.187; TCGA CESC -0.195; TCGA COAD -0.125; TCGA ESCA -0.221; TCGA HNSC -0.097; TCGA LUAD -0.132; TCGA LUSC -0.152; TCGA PAAD -0.095; TCGA PRAD -0.116; TCGA THCA -0.078; TCGA STAD -0.263; TCGA UCEC -0.274 |
hsa-miR-148b-3p | ZBTB46 | 9 cancers: BLCA; BRCA; COAD; KIRC; KIRP; LUAD; LUSC; OV; UCEC | mirMAP | TCGA BLCA -0.243; TCGA BRCA -0.145; TCGA COAD -0.285; TCGA KIRC -0.323; TCGA KIRP -0.227; TCGA LUAD -0.242; TCGA LUSC -0.317; TCGA OV -0.197; TCGA UCEC -0.433 |
hsa-miR-148b-3p | ADAM19 | 10 cancers: BLCA; COAD; ESCA; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD | mirMAP | TCGA BLCA -1.03; TCGA COAD -0.462; TCGA ESCA -0.507; TCGA LUAD -0.257; TCGA LUSC -0.53; TCGA PAAD -0.789; TCGA PRAD -0.473; TCGA SARC -0.472; TCGA THCA -0.899; TCGA STAD -0.278 |
hsa-miR-148b-3p | SCRN1 | 11 cancers: BLCA; COAD; ESCA; KIRC; KIRP; LIHC; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.375; TCGA COAD -0.444; TCGA ESCA -0.219; TCGA KIRC -0.49; TCGA KIRP -0.302; TCGA LIHC -0.455; TCGA LUSC -0.199; TCGA PRAD -0.509; TCGA THCA -0.492; TCGA STAD -0.304; TCGA UCEC -0.171 |
hsa-miR-148b-3p | ENTPD1 | 10 cancers: BLCA; COAD; KIRP; LUAD; PAAD; PRAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.485; TCGA COAD -0.368; TCGA KIRP -0.136; TCGA LUAD -0.181; TCGA PAAD -0.336; TCGA PRAD -0.109; TCGA SARC -0.456; TCGA THCA -0.515; TCGA STAD -0.343; TCGA UCEC -0.306 |
hsa-miR-148b-3p | KIAA1644 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -1.877; TCGA BRCA -0.232; TCGA CESC -0.956; TCGA COAD -1.392; TCGA ESCA -1.204; TCGA HNSC -0.573; TCGA LIHC -0.717; TCGA LUAD -0.351; TCGA LUSC -0.808; TCGA OV -0.902; TCGA PAAD -0.332; TCGA PRAD -1.026; TCGA THCA -1.047; TCGA STAD -1.246; TCGA UCEC -1.142 |
hsa-miR-148b-3p | WNT9A | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; LUAD; LUSC; OV; STAD; UCEC | mirMAP | TCGA BLCA -0.797; TCGA BRCA -0.217; TCGA CESC -0.879; TCGA COAD -1.038; TCGA ESCA -1.441; TCGA LUAD -0.194; TCGA LUSC -0.636; TCGA OV -0.26; TCGA STAD -1.127; TCGA UCEC -0.255 |
hsa-miR-148b-3p | SETD7 | 11 cancers: BLCA; CESC; ESCA; HNSC; LIHC; OV; PAAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.387; TCGA CESC -0.206; TCGA ESCA -0.139; TCGA HNSC -0.125; TCGA LIHC -0.287; TCGA OV -0.145; TCGA PAAD -0.202; TCGA SARC -0.156; TCGA THCA -0.499; TCGA STAD -0.394; TCGA UCEC -0.463 |
hsa-miR-148b-3p | GFOD1 | 13 cancers: BLCA; BRCA; COAD; ESCA; HNSC; KIRP; LUAD; LUSC; OV; PRAD; SARC; THCA; STAD | mirMAP | TCGA BLCA -0.673; TCGA BRCA -0.181; TCGA COAD -0.222; TCGA ESCA -0.511; TCGA HNSC -0.327; TCGA KIRP -0.28; TCGA LUAD -0.156; TCGA LUSC -0.344; TCGA OV -0.249; TCGA PRAD -0.344; TCGA SARC -0.334; TCGA THCA -0.127; TCGA STAD -0.67 |
hsa-miR-148b-3p | RNF217 | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; LIHC; PAAD; PRAD; SARC; STAD | mirMAP | TCGA BLCA -0.726; TCGA BRCA -0.297; TCGA CESC -0.44; TCGA COAD -0.601; TCGA ESCA -0.931; TCGA LIHC -0.489; TCGA PAAD -0.58; TCGA PRAD -0.168; TCGA SARC -0.311; TCGA STAD -0.61 |
hsa-miR-148b-3p | PLXDC1 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LUAD; LUSC; OV; PAAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.39; TCGA BRCA -0.271; TCGA CESC -0.396; TCGA COAD -0.404; TCGA HNSC -0.273; TCGA LUAD -0.213; TCGA LUSC -0.479; TCGA OV -0.352; TCGA PAAD -0.333; TCGA THCA -0.542; TCGA STAD -0.197; TCGA UCEC -0.495 |
hsa-miR-148b-3p | ADCY9 | 9 cancers: BLCA; COAD; ESCA; LUAD; LUSC; OV; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.649; TCGA COAD -0.191; TCGA ESCA -0.341; TCGA LUAD -0.223; TCGA LUSC -0.184; TCGA OV -0.22; TCGA SARC -0.346; TCGA STAD -0.357; TCGA UCEC -0.404 |
hsa-miR-148b-3p | KIF26B | 9 cancers: BLCA; CESC; COAD; KIRC; KIRP; LUSC; OV; PAAD; THCA | mirMAP | TCGA BLCA -0.288; TCGA CESC -0.346; TCGA COAD -0.762; TCGA KIRC -0.497; TCGA KIRP -0.283; TCGA LUSC -0.514; TCGA OV -0.572; TCGA PAAD -1.062; TCGA THCA -0.349 |
hsa-miR-148b-3p | FYCO1 | 12 cancers: BLCA; BRCA; COAD; ESCA; HNSC; KIRC; LIHC; LUAD; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.378; TCGA BRCA -0.079; TCGA COAD -0.22; TCGA ESCA -0.302; TCGA HNSC -0.273; TCGA KIRC -0.156; TCGA LIHC -0.191; TCGA LUAD -0.201; TCGA PRAD -0.208; TCGA SARC -0.611; TCGA STAD -0.594; TCGA UCEC -0.287 |
hsa-miR-148b-3p | FOXK1 | 9 cancers: BLCA; BRCA; CESC; KIRC; OV; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.136; TCGA BRCA -0.079; TCGA CESC -0.337; TCGA KIRC -0.211; TCGA OV -0.311; TCGA SARC -0.159; TCGA THCA -0.25; TCGA STAD -0.113; TCGA UCEC -0.201 |
hsa-miR-148b-3p | LONRF2 | 9 cancers: BLCA; CESC; COAD; KIRC; KIRP; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.756; TCGA CESC -0.745; TCGA COAD -2.034; TCGA KIRC -0.753; TCGA KIRP -1.186; TCGA SARC -1.505; TCGA THCA -0.598; TCGA STAD -1.315; TCGA UCEC -0.501 |
hsa-miR-148b-3p | SYNPO | 16 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PAAD; PRAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.7; TCGA BRCA -0.505; TCGA CESC -0.392; TCGA COAD -0.443; TCGA ESCA -0.531; TCGA HNSC -0.249; TCGA LIHC -0.195; TCGA LUAD -0.484; TCGA LUSC -0.516; TCGA OV -0.181; TCGA PAAD -0.275; TCGA PRAD -0.416; TCGA SARC -0.331; TCGA THCA -0.744; TCGA STAD -0.448; TCGA UCEC -0.502 |
hsa-miR-148b-3p | STARD5 | 10 cancers: BLCA; BRCA; CESC; ESCA; LIHC; LUSC; PAAD; PRAD; SARC; UCEC | mirMAP | TCGA BLCA -0.159; TCGA BRCA -0.122; TCGA CESC -0.363; TCGA ESCA -0.303; TCGA LIHC -0.585; TCGA LUSC -0.207; TCGA PAAD -0.197; TCGA PRAD -0.168; TCGA SARC -0.306; TCGA UCEC -0.211 |
hsa-miR-148b-3p | PPP1R3B | 9 cancers: BLCA; BRCA; HNSC; KIRP; LIHC; PAAD; PRAD; SARC; UCEC | mirMAP | TCGA BLCA -0.372; TCGA BRCA -0.189; TCGA HNSC -0.303; TCGA KIRP -0.186; TCGA LIHC -0.366; TCGA PAAD -0.333; TCGA PRAD -0.229; TCGA SARC -0.901; TCGA UCEC -0.226 |
hsa-miR-148b-3p | ZHX3 | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.404; TCGA BRCA -0.156; TCGA CESC -0.318; TCGA COAD -0.181; TCGA ESCA -0.301; TCGA HNSC -0.33; TCGA KIRC -0.246; TCGA KIRP -0.249; TCGA LIHC -0.332; TCGA LUAD -0.114; TCGA LUSC -0.245; TCGA OV -0.235; TCGA SARC -0.163; TCGA STAD -0.442; TCGA UCEC -0.276 |
hsa-miR-148b-3p | ARL10 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; LUAD; LUSC; PRAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.544; TCGA BRCA -0.243; TCGA CESC -0.71; TCGA COAD -0.667; TCGA ESCA -0.711; TCGA LUAD -0.286; TCGA LUSC -0.248; TCGA PRAD -0.403; TCGA SARC -0.229; TCGA THCA -0.458; TCGA STAD -0.473; TCGA UCEC -0.307 |
hsa-miR-148b-3p | LYNX1 | 17 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; OV; PAAD; PRAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.858; TCGA BRCA -0.378; TCGA CESC -0.621; TCGA COAD -0.952; TCGA ESCA -1.057; TCGA HNSC -1.206; TCGA KIRC -0.617; TCGA KIRP -0.399; TCGA LUAD -0.474; TCGA LUSC -0.861; TCGA OV -0.973; TCGA PAAD -0.34; TCGA PRAD -0.514; TCGA SARC -0.811; TCGA THCA -0.952; TCGA STAD -0.901; TCGA UCEC -0.583 |
hsa-miR-148b-3p | KCTD2 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.101; TCGA BRCA -0.07; TCGA CESC -0.096; TCGA COAD -0.151; TCGA ESCA -0.121; TCGA HNSC -0.178; TCGA KIRC -0.096; TCGA KIRP -0.163; TCGA LIHC -0.106; TCGA LUAD -0.206; TCGA PRAD -0.087; TCGA STAD -0.181; TCGA UCEC -0.178 |
hsa-miR-148b-3p | MXRA7 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUAD; LUSC; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BLCA -0.722; TCGA BRCA -0.293; TCGA CESC -0.461; TCGA COAD -0.57; TCGA ESCA -0.717; TCGA HNSC -0.138; TCGA KIRP -0.312; TCGA LUAD -0.525; TCGA LUSC -0.185; TCGA PRAD -0.575; TCGA SARC -0.425; TCGA STAD -0.772; TCGA UCEC -0.429 |
hsa-miR-148b-3p | LRRC55 | 10 cancers: BLCA; CESC; COAD; KIRC; LIHC; LUAD; PAAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.215; TCGA CESC -0.467; TCGA COAD -0.959; TCGA KIRC -0.354; TCGA LIHC -0.647; TCGA LUAD -0.258; TCGA PAAD -1.178; TCGA THCA -0.576; TCGA STAD -0.527; TCGA UCEC -0.272 |
hsa-miR-148b-3p | BCL9L | 10 cancers: BLCA; CESC; COAD; KIRP; LGG; LUAD; LUSC; PAAD; SARC; THCA | mirMAP | TCGA BLCA -0.172; TCGA CESC -0.163; TCGA COAD -0.118; TCGA KIRP -0.157; TCGA LGG -0.161; TCGA LUAD -0.198; TCGA LUSC -0.15; TCGA PAAD -0.394; TCGA SARC -0.202; TCGA THCA -0.415 |
hsa-miR-148b-3p | TECPR2 | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LUSC; SARC; THCA; STAD | mirMAP | TCGA BLCA -0.218; TCGA BRCA -0.088; TCGA CESC -0.228; TCGA COAD -0.113; TCGA ESCA -0.287; TCGA KIRC -0.223; TCGA LUSC -0.11; TCGA SARC -0.17; TCGA THCA -0.072; TCGA STAD -0.284 |
hsa-miR-148b-3p | ADARB1 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LUAD; LUSC; PAAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.829; TCGA BRCA -0.199; TCGA CESC -0.226; TCGA COAD -0.331; TCGA ESCA -0.45; TCGA HNSC -0.155; TCGA KIRC -0.252; TCGA KIRP -0.466; TCGA LUAD -0.363; TCGA LUSC -0.396; TCGA PAAD -0.19; TCGA THCA -0.293; TCGA STAD -0.563; TCGA UCEC -0.399 |
hsa-miR-148b-3p | MYO5A | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.384; TCGA CESC -0.57; TCGA COAD -0.471; TCGA ESCA -0.693; TCGA HNSC -0.261; TCGA LUAD -0.285; TCGA LUSC -0.367; TCGA THCA -0.307; TCGA STAD -0.266; TCGA UCEC -0.142 |
hsa-miR-148b-3p | BMPR2 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; LIHC; LUAD; PAAD; PRAD; SARC; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.217; TCGA BRCA -0.068; TCGA CESC -0.196; TCGA COAD -0.221; TCGA ESCA -0.264; TCGA KIRP -0.127; TCGA LIHC -0.275; TCGA LUAD -0.221; TCGA PAAD -0.292; TCGA PRAD -0.16; TCGA SARC -0.153; TCGA THCA -0.135; TCGA STAD -0.223; TCGA UCEC -0.264 |
hsa-miR-148b-3p | ATXN1L | 10 cancers: BLCA; CESC; COAD; ESCA; LIHC; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.115; TCGA CESC -0.276; TCGA COAD -0.094; TCGA ESCA -0.177; TCGA LIHC -0.18; TCGA LUSC -0.124; TCGA OV -0.116; TCGA PAAD -0.113; TCGA STAD -0.107; TCGA UCEC -0.111 |
hsa-miR-148b-3p | PALM2 | 10 cancers: BLCA; COAD; ESCA; HNSC; KIRC; LIHC; LUSC; OV; PRAD; SARC | mirMAP | TCGA BLCA -0.447; TCGA COAD -0.398; TCGA ESCA -0.515; TCGA HNSC -0.254; TCGA KIRC -0.306; TCGA LIHC -0.56; TCGA LUSC -0.534; TCGA OV -0.258; TCGA PRAD -0.132; TCGA SARC -0.356 |
hsa-miR-148b-3p | PTPRD | 11 cancers: BLCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; OV; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -1.383; TCGA CESC -1.009; TCGA COAD -0.645; TCGA ESCA -0.856; TCGA HNSC -0.773; TCGA LIHC -1.024; TCGA LUAD -0.513; TCGA OV -0.785; TCGA THCA -0.67; TCGA STAD -0.455; TCGA UCEC -0.728 |
hsa-miR-148b-3p | MAFB | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; OV; STAD; UCEC | miRNATAP | TCGA BLCA -0.524; TCGA BRCA -0.237; TCGA CESC -0.468; TCGA COAD -0.574; TCGA ESCA -0.92; TCGA HNSC -0.583; TCGA LUAD -0.347; TCGA LUSC -0.274; TCGA OV -0.2; TCGA STAD -0.265; TCGA UCEC -0.118 |
hsa-miR-148b-3p | NCAM1 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LGG; LIHC; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -1.992; TCGA BRCA -0.764; TCGA CESC -1.008; TCGA COAD -0.888; TCGA ESCA -0.948; TCGA HNSC -1.092; TCGA KIRP -0.878; TCGA LGG -0.336; TCGA LIHC -0.833; TCGA PRAD -1.118; TCGA SARC -0.629; TCGA STAD -1.167; TCGA UCEC -1.118 |
hsa-miR-148b-3p | CCDC85A | 11 cancers: BLCA; BRCA; CESC; COAD; KIRP; LIHC; LUAD; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.274; TCGA BRCA -0.516; TCGA CESC -0.53; TCGA COAD -0.678; TCGA KIRP -0.419; TCGA LIHC -0.396; TCGA LUAD -0.395; TCGA PRAD -0.566; TCGA SARC -0.35; TCGA STAD -0.541; TCGA UCEC -0.835 |
hsa-miR-148b-3p | IL17RD | 15 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LGG; LIHC; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.585; TCGA BRCA -0.423; TCGA CESC -0.568; TCGA COAD -0.468; TCGA ESCA -0.475; TCGA KIRC -0.24; TCGA LGG -0.138; TCGA LIHC -0.324; TCGA LUAD -0.301; TCGA OV -0.269; TCGA PAAD -0.58; TCGA PRAD -0.334; TCGA THCA -1.557; TCGA STAD -0.731; TCGA UCEC -0.312 |
hsa-miR-148b-3p | PPFIA2 | 10 cancers: BLCA; BRCA; ESCA; KIRC; KIRP; LUAD; PAAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -1.038; TCGA BRCA -0.315; TCGA ESCA -0.548; TCGA KIRC -0.603; TCGA KIRP -0.694; TCGA LUAD -0.346; TCGA PAAD -0.967; TCGA SARC -1.286; TCGA STAD -0.579; TCGA UCEC -0.561 |
hsa-miR-148b-3p | PDK4 | 10 cancers: BLCA; BRCA; CESC; COAD; HNSC; LIHC; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.819; TCGA BRCA -0.994; TCGA CESC -0.449; TCGA COAD -0.644; TCGA HNSC -0.888; TCGA LIHC -0.91; TCGA PRAD -0.532; TCGA SARC -0.365; TCGA STAD -1.237; TCGA UCEC -0.664 |
hsa-miR-148b-3p | ATXN1 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUAD; LUSC; PAAD; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.613; TCGA BRCA -0.073; TCGA CESC -0.288; TCGA COAD -0.21; TCGA ESCA -0.315; TCGA HNSC -0.151; TCGA LUAD -0.194; TCGA LUSC -0.169; TCGA PAAD -0.335; TCGA PRAD -0.343; TCGA SARC -0.126; TCGA STAD -0.329; TCGA UCEC -0.19 |
hsa-miR-148b-3p | LAMC2 | 10 cancers: BRCA; ESCA; KIRC; KIRP; LIHC; LUAD; LUSC; PAAD; PRAD; THCA | miRNAWalker2 validate | TCGA BRCA -0.468; TCGA ESCA -0.617; TCGA KIRC -0.667; TCGA KIRP -0.666; TCGA LIHC -0.826; TCGA LUAD -0.531; TCGA LUSC -1.008; TCGA PAAD -1.648; TCGA PRAD -0.672; TCGA THCA -0.929 |
hsa-miR-148b-3p | NLGN4X | 10 cancers: BRCA; CESC; COAD; ESCA; KIRP; OV; PRAD; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BRCA -0.513; TCGA CESC -0.895; TCGA COAD -0.792; TCGA ESCA -0.474; TCGA KIRP -1.041; TCGA OV -0.876; TCGA PRAD -0.335; TCGA THCA -0.431; TCGA STAD -0.601; TCGA UCEC -0.363 |
hsa-miR-148b-3p | PKP1 | 9 cancers: BRCA; CESC; COAD; ESCA; HNSC; KIRC; PAAD; PRAD; THCA | miRNAWalker2 validate | TCGA BRCA -0.905; TCGA CESC -1.249; TCGA COAD -0.462; TCGA ESCA -2.201; TCGA HNSC -0.456; TCGA KIRC -0.485; TCGA PAAD -1.318; TCGA PRAD -0.389; TCGA THCA -1.351 |
hsa-miR-148b-3p | TOM1L2 | 13 cancers: BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LGG; LUAD; OV; SARC; STAD; UCEC | miRNAWalker2 validate | TCGA BRCA -0.189; TCGA CESC -0.506; TCGA COAD -0.146; TCGA ESCA -0.336; TCGA HNSC -0.366; TCGA KIRC -0.284; TCGA KIRP -0.144; TCGA LGG -0.069; TCGA LUAD -0.122; TCGA OV -0.124; TCGA SARC -0.422; TCGA STAD -0.407; TCGA UCEC -0.237 |
hsa-miR-148b-3p | YPEL1 | 11 cancers: BRCA; COAD; KIRC; KIRP; LGG; LIHC; OV; SARC; THCA; STAD; UCEC | miRNAWalker2 validate | TCGA BRCA -0.087; TCGA COAD -0.222; TCGA KIRC -0.139; TCGA KIRP -0.267; TCGA LGG -0.153; TCGA LIHC -0.443; TCGA OV -0.262; TCGA SARC -0.382; TCGA THCA -0.136; TCGA STAD -0.301; TCGA UCEC -0.091 |
hsa-miR-148b-3p | MTSS1L | 15 cancers: BRCA; CESC; COAD; ESCA; KIRC; KIRP; LGG; LUAD; LUSC; OV; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BRCA -0.103; TCGA CESC -0.464; TCGA COAD -0.253; TCGA ESCA -0.491; TCGA KIRC -0.332; TCGA KIRP -0.299; TCGA LGG -0.295; TCGA LUAD -0.196; TCGA LUSC -0.282; TCGA OV -0.211; TCGA PRAD -0.188; TCGA SARC -0.478; TCGA THCA -0.414; TCGA STAD -0.478; TCGA UCEC -0.125 |
hsa-miR-148b-3p | MPPED2 | 9 cancers: BRCA; CESC; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; STAD | MirTarget | TCGA BRCA -0.518; TCGA CESC -1.094; TCGA COAD -0.932; TCGA ESCA -0.947; TCGA KIRC -1.353; TCGA KIRP -1.046; TCGA LIHC -0.316; TCGA LUAD -0.336; TCGA STAD -1.016 |
hsa-miR-148b-3p | EPN2 | 12 cancers: BRCA; CESC; ESCA; HNSC; KIRC; KIRP; LGG; LUAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BRCA -0.066; TCGA CESC -0.327; TCGA ESCA -0.327; TCGA HNSC -0.172; TCGA KIRC -0.133; TCGA KIRP -0.193; TCGA LGG -0.175; TCGA LUAD -0.079; TCGA SARC -0.385; TCGA THCA -0.117; TCGA STAD -0.322; TCGA UCEC -0.168 |
hsa-miR-148b-3p | MUM1L1 | 10 cancers: BRCA; CESC; COAD; ESCA; HNSC; LUAD; OV; PAAD; PRAD; UCEC | MirTarget | TCGA BRCA -0.45; TCGA CESC -0.988; TCGA COAD -0.827; TCGA ESCA -0.701; TCGA HNSC -1.245; TCGA LUAD -0.378; TCGA OV -0.333; TCGA PAAD -1.576; TCGA PRAD -0.426; TCGA UCEC -1.276 |
hsa-miR-148b-3p | DOCK6 | 9 cancers: BRCA; CESC; HNSC; KIRC; KIRP; LUSC; OV; THCA; UCEC | MirTarget | TCGA BRCA -0.265; TCGA CESC -0.339; TCGA HNSC -0.204; TCGA KIRC -0.315; TCGA KIRP -0.295; TCGA LUSC -0.264; TCGA OV -0.22; TCGA THCA -0.084; TCGA UCEC -0.148 |
hsa-miR-148b-3p | KIAA1217 | 9 cancers: BRCA; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; THCA | MirTarget; miRNATAP | TCGA BRCA -0.091; TCGA ESCA -0.205; TCGA KIRC -0.171; TCGA KIRP -0.139; TCGA LIHC -0.146; TCGA LUAD -0.205; TCGA PAAD -0.869; TCGA PRAD -0.083; TCGA THCA -0.612 |
hsa-miR-148b-3p | PIK3IP1 | 11 cancers: BRCA; CESC; COAD; ESCA; LGG; LIHC; LUAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BRCA -0.301; TCGA CESC -0.261; TCGA COAD -0.325; TCGA ESCA -0.215; TCGA LGG -0.092; TCGA LIHC -0.257; TCGA LUAD -0.182; TCGA SARC -0.28; TCGA THCA -0.087; TCGA STAD -0.266; TCGA UCEC -0.329 |
hsa-miR-148b-3p | BMP3 | 9 cancers: BRCA; COAD; ESCA; KIRC; KIRP; LUAD; PAAD; PRAD; STAD | MirTarget | TCGA BRCA -0.914; TCGA COAD -1.875; TCGA ESCA -1.468; TCGA KIRC -0.514; TCGA KIRP -0.683; TCGA LUAD -0.331; TCGA PAAD -0.735; TCGA PRAD -1.025; TCGA STAD -1.973 |
hsa-miR-148b-3p | MDFIC | 13 cancers: BRCA; CESC; COAD; ESCA; KIRP; LIHC; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BRCA -0.22; TCGA CESC -0.275; TCGA COAD -0.467; TCGA ESCA -0.547; TCGA KIRP -0.213; TCGA LIHC -0.479; TCGA LUAD -0.402; TCGA OV -0.38; TCGA PAAD -0.778; TCGA PRAD -0.355; TCGA THCA -0.651; TCGA STAD -0.459; TCGA UCEC -0.596 |
hsa-miR-148b-3p | EFNB2 | 11 cancers: BRCA; CESC; HNSC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; THCA; UCEC | MirTarget | TCGA BRCA -0.299; TCGA CESC -0.364; TCGA HNSC -0.389; TCGA KIRP -0.688; TCGA LIHC -0.202; TCGA LUAD -0.347; TCGA LUSC -0.322; TCGA OV -0.486; TCGA PAAD -0.502; TCGA THCA -0.125; TCGA UCEC -0.326 |
hsa-miR-148b-3p | USP31 | 9 cancers: BRCA; CESC; ESCA; LUAD; LUSC; PRAD; SARC; THCA; UCEC | MirTarget | TCGA BRCA -0.067; TCGA CESC -0.389; TCGA ESCA -0.439; TCGA LUAD -0.11; TCGA LUSC -0.289; TCGA PRAD -0.102; TCGA SARC -0.276; TCGA THCA -0.413; TCGA UCEC -0.269 |
hsa-miR-148b-3p | SIK1 | 10 cancers: BRCA; ESCA; KIRC; KIRP; LIHC; LUSC; PAAD; PRAD; THCA; UCEC | MirTarget | TCGA BRCA -0.35; TCGA ESCA -0.226; TCGA KIRC -0.669; TCGA KIRP -0.773; TCGA LIHC -0.366; TCGA LUSC -0.21; TCGA PAAD -0.283; TCGA PRAD -0.474; TCGA THCA -0.786; TCGA UCEC -0.277 |
hsa-miR-148b-3p | BHLHE41 | 9 cancers: BRCA; COAD; ESCA; LUAD; LUSC; PAAD; PRAD; THCA; STAD | MirTarget | TCGA BRCA -0.497; TCGA COAD -0.554; TCGA ESCA -0.591; TCGA LUAD -0.578; TCGA LUSC -0.538; TCGA PAAD -0.39; TCGA PRAD -0.394; TCGA THCA -0.656; TCGA STAD -0.321 |
hsa-miR-148b-3p | CDH23 | 12 cancers: BRCA; COAD; ESCA; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BRCA -0.841; TCGA COAD -0.798; TCGA ESCA -0.688; TCGA LIHC -0.611; TCGA LUAD -0.479; TCGA LUSC -0.359; TCGA OV -0.292; TCGA PAAD -0.384; TCGA PRAD -0.734; TCGA THCA -0.487; TCGA STAD -0.494; TCGA UCEC -0.457 |
hsa-miR-148b-3p | SLC25A26 | 11 cancers: BRCA; CESC; HNSC; KIRC; LGG; LIHC; OV; PRAD; SARC; STAD; UCEC | mirMAP | TCGA BRCA -0.14; TCGA CESC -0.13; TCGA HNSC -0.191; TCGA KIRC -0.246; TCGA LGG -0.1; TCGA LIHC -0.116; TCGA OV -0.106; TCGA PRAD -0.076; TCGA SARC -0.107; TCGA STAD -0.373; TCGA UCEC -0.089 |
hsa-miR-148b-3p | FAM160B2 | 9 cancers: BRCA; CESC; ESCA; KIRC; LGG; LUSC; OV; SARC; STAD | mirMAP | TCGA BRCA -0.202; TCGA CESC -0.161; TCGA ESCA -0.132; TCGA KIRC -0.121; TCGA LGG -0.1; TCGA LUSC -0.324; TCGA OV -0.108; TCGA SARC -0.19; TCGA STAD -0.129 |
hsa-miR-148b-3p | IGF2 | 11 cancers: BRCA; HNSC; KIRC; KIRP; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BRCA -0.895; TCGA HNSC -0.815; TCGA KIRC -1.719; TCGA KIRP -0.558; TCGA LUAD -0.391; TCGA OV -0.783; TCGA PAAD -1.002; TCGA PRAD -0.358; TCGA THCA -0.959; TCGA STAD -0.337; TCGA UCEC -0.425 |
hsa-miR-148b-3p | AR | 9 cancers: BRCA; CESC; COAD; ESCA; LIHC; PAAD; SARC; STAD; UCEC | mirMAP | TCGA BRCA -0.268; TCGA CESC -0.514; TCGA COAD -0.987; TCGA ESCA -0.632; TCGA LIHC -0.987; TCGA PAAD -0.873; TCGA SARC -1.016; TCGA STAD -1.177; TCGA UCEC -0.368 |
hsa-miR-148b-3p | SRRM3 | 13 cancers: BRCA; CESC; COAD; ESCA; HNSC; KIRC; LGG; LUAD; LUSC; OV; PRAD; THCA; UCEC | mirMAP | TCGA BRCA -0.357; TCGA CESC -0.957; TCGA COAD -0.378; TCGA ESCA -0.911; TCGA HNSC -0.678; TCGA KIRC -0.414; TCGA LGG -0.774; TCGA LUAD -0.25; TCGA LUSC -0.631; TCGA OV -0.36; TCGA PRAD -0.481; TCGA THCA -0.347; TCGA UCEC -0.372 |
hsa-miR-148b-3p | SEPT5 | 10 cancers: BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUSC; OV; STAD; UCEC | mirMAP | TCGA BRCA -0.107; TCGA CESC -0.484; TCGA COAD -0.381; TCGA ESCA -0.73; TCGA HNSC -0.904; TCGA KIRP -0.394; TCGA LUSC -0.26; TCGA OV -0.203; TCGA STAD -0.627; TCGA UCEC -0.353 |
hsa-miR-148b-3p | NFAT5 | 13 cancers: BRCA; CESC; ESCA; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BRCA -0.132; TCGA CESC -0.13; TCGA ESCA -0.261; TCGA KIRC -0.302; TCGA KIRP -0.337; TCGA LIHC -0.21; TCGA LUAD -0.113; TCGA LUSC -0.323; TCGA OV -0.19; TCGA PAAD -0.17; TCGA PRAD -0.112; TCGA STAD -0.208; TCGA UCEC -0.169 |
hsa-miR-148b-3p | ARL8B | 11 cancers: CESC; ESCA; HNSC; KIRC; LUAD; LUSC; PAAD; SARC; THCA; STAD; UCEC | miRNAWalker2 validate; MirTarget | TCGA CESC -0.259; TCGA ESCA -0.257; TCGA HNSC -0.193; TCGA KIRC -0.155; TCGA LUAD -0.084; TCGA LUSC -0.164; TCGA PAAD -0.089; TCGA SARC -0.166; TCGA THCA -0.235; TCGA STAD -0.141; TCGA UCEC -0.119 |
hsa-miR-148b-3p | MATN3 | 9 cancers: CESC; COAD; KIRC; LUAD; LUSC; PAAD; PRAD; THCA; UCEC | miRNAWalker2 validate; MirTarget | TCGA CESC -0.562; TCGA COAD -1.013; TCGA KIRC -0.441; TCGA LUAD -0.368; TCGA LUSC -0.496; TCGA PAAD -1.442; TCGA PRAD -0.269; TCGA THCA -1.191; TCGA UCEC -0.19 |
hsa-miR-148b-3p | KCNH1 | 9 cancers: CESC; COAD; ESCA; KIRC; KIRP; PRAD; THCA; STAD; UCEC | MirTarget | TCGA CESC -0.518; TCGA COAD -0.382; TCGA ESCA -0.838; TCGA KIRC -0.73; TCGA KIRP -0.844; TCGA PRAD -0.496; TCGA THCA -0.697; TCGA STAD -0.309; TCGA UCEC -0.332 |
hsa-miR-148b-3p | RAI14 | 10 cancers: CESC; COAD; ESCA; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA CESC -0.279; TCGA COAD -0.415; TCGA ESCA -0.419; TCGA LUAD -0.194; TCGA OV -0.272; TCGA PAAD -0.495; TCGA PRAD -0.109; TCGA THCA -0.531; TCGA STAD -0.266; TCGA UCEC -0.257 |
hsa-miR-148b-3p | ADAMTS19 | 9 cancers: CESC; COAD; KIRC; KIRP; LGG; OV; PRAD; STAD; UCEC | MirTarget | TCGA CESC -0.65; TCGA COAD -0.505; TCGA KIRC -0.542; TCGA KIRP -0.681; TCGA LGG -0.411; TCGA OV -0.447; TCGA PRAD -0.641; TCGA STAD -0.496; TCGA UCEC -0.444 |
hsa-miR-148b-3p | CTTNBP2NL | 9 cancers: CESC; ESCA; LIHC; LUAD; PAAD; PRAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA CESC -0.314; TCGA ESCA -0.199; TCGA LIHC -0.156; TCGA LUAD -0.203; TCGA PAAD -0.345; TCGA PRAD -0.088; TCGA THCA -0.342; TCGA STAD -0.139; TCGA UCEC -0.139 |
hsa-miR-148b-3p | MTMR3 | 11 cancers: CESC; COAD; ESCA; HNSC; KIRC; KIRP; LGG; LUAD; SARC; STAD; UCEC | MirTarget; mirMAP | TCGA CESC -0.163; TCGA COAD -0.104; TCGA ESCA -0.193; TCGA HNSC -0.158; TCGA KIRC -0.201; TCGA KIRP -0.156; TCGA LGG -0.106; TCGA LUAD -0.177; TCGA SARC -0.09; TCGA STAD -0.107; TCGA UCEC -0.074 |
hsa-miR-148b-3p | EPB41L1 | 9 cancers: CESC; COAD; KIRC; LUAD; PAAD; PRAD; SARC; THCA; UCEC | MirTarget | TCGA CESC -0.234; TCGA COAD -0.244; TCGA KIRC -0.197; TCGA LUAD -0.17; TCGA PAAD -0.304; TCGA PRAD -0.086; TCGA SARC -0.191; TCGA THCA -0.279; TCGA UCEC -0.2 |
hsa-miR-148b-3p | TANC2 | 9 cancers: CESC; COAD; ESCA; KIRP; LGG; LUAD; LUSC; THCA; UCEC | mirMAP | TCGA CESC -0.299; TCGA COAD -0.329; TCGA ESCA -0.414; TCGA KIRP -0.398; TCGA LGG -0.074; TCGA LUAD -0.172; TCGA LUSC -0.311; TCGA THCA -0.673; TCGA UCEC -0.209 |
hsa-miR-148b-3p | NOS1 | 9 cancers: COAD; ESCA; HNSC; KIRC; KIRP; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA COAD -1.582; TCGA ESCA -1.681; TCGA HNSC -0.651; TCGA KIRC -0.59; TCGA KIRP -0.87; TCGA PAAD -0.625; TCGA PRAD -1.323; TCGA STAD -1.685; TCGA UCEC -0.363 |
Enriched cancer pathways of putative targets