microRNA information: hsa-miR-194-5p
| Section | ID | link |
|---|---|---|
| Mature name: | hsa-miR-194-5p | miRbase |
| Accession: | MIMAT0000460 | miRbase |
| Precursor name: | hsa-mir-194-1 | miRbase |
| Precursor accession: | MI0000488 | miRbase |
| Symbol: | MIR194-1 | HGNC |
| RefSeq ID: | NR_029711 | GenBank |
| Sequence: | UGUAACAGCAACUCCAUGUGGA |
Reported expression in cancers: hsa-miR-194-5p
| miRNA | cancer | regulation | reporting | PUBMED | method |
|---|---|---|---|---|---|
| hsa-miR-194-5p | bladder cancer | downregulation | "In the present study we investigated the roles and ......" | 27133066 | |
| hsa-miR-194-5p | breast cancer | deregulation | "We identified seven miRNAs that were differentiall ......" | 27409424 | |
| hsa-miR-194-5p | colorectal cancer | downregulation | "MiR 194 commonly repressed in colorectal cancer su ......" | 25602366 | Microarray |
| hsa-miR-194-5p | gastric cancer | upregulation | "We performed miRNA microarray and quantitative rev ......" | 23385731 | Reverse transcription PCR; Microarray; qPCR |
| hsa-miR-194-5p | kidney renal cell cancer | downregulation | "miR 192 miR 194 and miR 215: a convergent microRNA ......" | 23715501 | |
| hsa-miR-194-5p | kidney renal cell cancer | downregulation | "miR-194 was reported to be downregulated in severa ......" | 26860079 | |
| hsa-miR-194-5p | liver cancer | upregulation | "Up-regulation of miR-15a miR-16 miR-27b miR-30b mi ......" | 24314246 | qPCR |
| hsa-miR-194-5p | lung squamous cell cancer | downregulation | "Earlier studies have demonstrated that miR-194 was ......" | 26909612 | |
| hsa-miR-194-5p | melanoma | upregulation | "miR-194 is a tumor-suppressor gene in multiple tum ......" | 27573550 | |
| hsa-miR-194-5p | ovarian cancer | upregulation | "In this study we explored the role of miR-194 in o ......" | 27486333 | qPCR |
| hsa-miR-194-5p | prostate cancer | upregulation | "A trend of increased expression >40% of miR-135b a ......" | 18949015 | qPCR |
Reported cancer pathway affected by hsa-miR-194-5p

| miRNA | cancer | pathway | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-194-5p | bladder cancer | cell cycle pathway | "Long noncoding RNA H19 contributes to gallbladder ......" | 26803515 | |
| hsa-miR-194-5p | breast cancer | Apoptosis pathway | "We have performed miRNA microarray profiling befor ......" | 22829924 | Luciferase |
| hsa-miR-194-5p | gastric cancer | cell cycle pathway | "Inverse association between miR 194 expression and ......" | 21845495 | Western blot |
| hsa-miR-194-5p | gastric cancer | Epithelial mesenchymal transition pathway | "We hypothesized that miR-194 may control Forkhead ......" | 24748184 | |
| hsa-miR-194-5p | gastric cancer | cell cycle pathway; Apoptosis pathway | "miR 194 targets RBX1 gene to modulate proliferatio ......" | 25412959 | Flow cytometry; Transwell assay; Wound Healing Assay; Western blot; Luciferase |
| hsa-miR-194-5p | liver cancer | Epithelial mesenchymal transition pathway | "miR 194 is a marker of hepatic epithelial cells an ......" | 20979124 | |
| hsa-miR-194-5p | liver cancer | Apoptosis pathway | "In this study we aimed to analyze the role of micr ......" | 26722431 | |
| hsa-miR-194-5p | lung squamous cell cancer | Apoptosis pathway | "miR 194 inhibits the proliferation invasion migrat ......" | 26909612 | |
| hsa-miR-194-5p | sarcoma | Apoptosis pathway | "Studies have shown that miR-194 functions as a tum ......" | 25096247 | Western blot; Luciferase |
Reported cancer prognosis affected by hsa-miR-194-5p

| miRNA | cancer | prognosis | reporting | PUBMED | functional study |
|---|---|---|---|---|---|
| hsa-miR-194-5p | bladder cancer | progression | "MiR 194 inhibits cell proliferation and invasion v ......" | 27133066 | |
| hsa-miR-194-5p | breast cancer | cell migration | "We have performed miRNA microarray profiling befor ......" | 22829924 | Luciferase |
| hsa-miR-194-5p | breast cancer | recurrence | "We identified seven miRNAs that were differentiall ......" | 27409424 | |
| hsa-miR-194-5p | colorectal cancer | staging; differentiation; tumor size; poor survival | "MiR 194 commonly repressed in colorectal cancer su ......" | 25602366 | Luciferase |
| hsa-miR-194-5p | colorectal cancer | worse prognosis | "Circulating levels of the miRNAs miR 194 and miR 2 ......" | 26318304 | |
| hsa-miR-194-5p | colorectal cancer | poor survival | "The expression levels of miR-206 miR-219 miR-192 m ......" | 27666868 | |
| hsa-miR-194-5p | colorectal cancer | staging; poor survival | "Significant inverse correlation between the expres ......" | 27747302 | |
| hsa-miR-194-5p | endometrial cancer | metastasis | "Furthermore we discovered that the expression of B ......" | 21851624 | |
| hsa-miR-194-5p | endometrial cancer | staging; worse prognosis | "Prognostic significance of miR 194 in endometrial ......" | 24040515 | |
| hsa-miR-194-5p | esophageal cancer | drug resistance; staging | "Initially a microarray-based approach was performe ......" | 23649428 | |
| hsa-miR-194-5p | gastric cancer | staging; tumor size; progression | "Inverse association between miR 194 expression and ......" | 21845495 | Western blot |
| hsa-miR-194-5p | gastric cancer | staging | "We performed miRNA microarray and quantitative rev ......" | 23385731 | |
| hsa-miR-194-5p | gastric cancer | cell migration; progression | "We hypothesized that miR-194 may control Forkhead ......" | 24748184 | |
| hsa-miR-194-5p | gastric cancer | poor survival; metastasis; tumor size | "miR 194 targets RBX1 gene to modulate proliferatio ......" | 25412959 | Flow cytometry; Transwell assay; Wound Healing Assay; Western blot; Luciferase |
| hsa-miR-194-5p | kidney renal cell cancer | progression | "miR 192 miR 194 and miR 215: a convergent microRNA ......" | 23715501 | Luciferase |
| hsa-miR-194-5p | kidney renal cell cancer | staging; poor survival; tumor size; worse prognosis; progression | "miR-194 was reported to be downregulated in severa ......" | 26860079 | |
| hsa-miR-194-5p | liver cancer | metastasis; differentiation | "miR 194 is a marker of hepatic epithelial cells an ......" | 20979124 | |
| hsa-miR-194-5p | liver cancer | staging | "Gα12gep oncogene inhibits FOXO1 in hepatocellular ......" | 24631529 | |
| hsa-miR-194-5p | liver cancer | cell migration; metastasis | "NF κB signaling relieves negative regulation by m ......" | 26221053 | Luciferase |
| hsa-miR-194-5p | liver cancer | poor survival; staging; metastasis; tumor size; progression | "In this study we aimed to analyze the role of micr ......" | 26722431 | |
| hsa-miR-194-5p | lung cancer | progression | "We succeeded in establishing the MTA1 knockdown NS ......" | 22576802 | RNAi |
| hsa-miR-194-5p | lung squamous cell cancer | metastasis | "miR 194 suppresses metastasis of non small cell lu ......" | 23584484 | |
| hsa-miR-194-5p | lung squamous cell cancer | metastasis; drug resistance; poor survival | "miR 194 inhibits the proliferation invasion migrat ......" | 26909612 | |
| hsa-miR-194-5p | lymphoma | poor survival | "Functional studies indicated that CDKN1A/p21 and S ......" | 22102710 | |
| hsa-miR-194-5p | melanoma | metastasis; staging | "miR 194 is a negative regulator of GEF H1 pathway ......" | 27573550 | |
| hsa-miR-194-5p | ovarian cancer | staging | "The reactivation of p53 was detected by Western bl ......" | 22340095 | Western blot |
| hsa-miR-194-5p | pancreatic cancer | worse prognosis; poor survival | "The aim of this study was to investigate microRNAs ......" | 25906450 | |
| hsa-miR-194-5p | prostate cancer | recurrence; progression; metastasis; worse prognosis | "MicroRNAs in the serum of patients who had experie ......" | 23846169 | |
| hsa-miR-194-5p | sarcoma | metastasis; poor survival | "Studies have shown that miR-194 functions as a tum ......" | 25096247 | Western blot; Luciferase |
Reported gene related to hsa-miR-194-5p
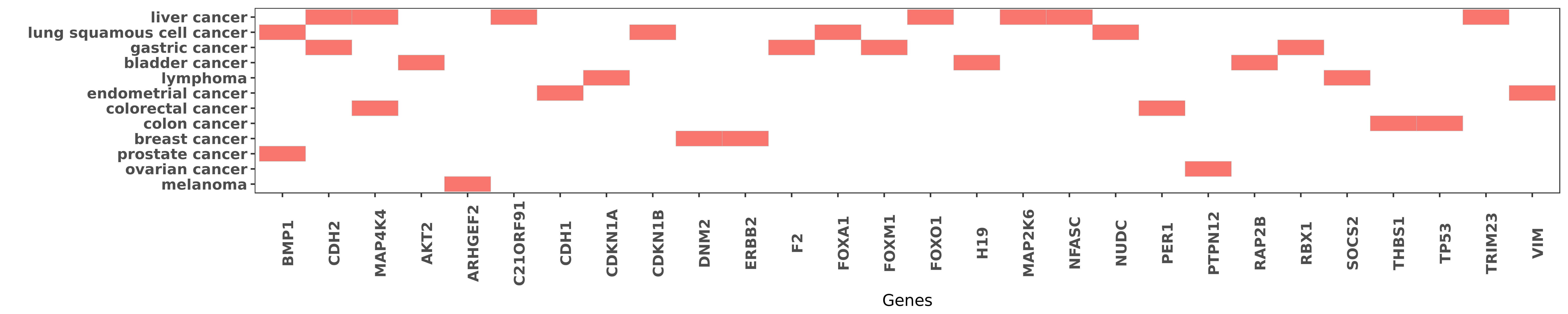
| miRNA | cancer | gene | reporting | PUBMED |
|---|---|---|---|---|
| hsa-miR-194-5p | lung squamous cell cancer | BMP1 | "miR 194 suppresses metastasis of non small cell lu ......" | 23584484 |
| hsa-miR-194-5p | prostate cancer | BMP1 | "Bone morphogenetic protein 1 BMP1 was shown to be ......" | 26820911 |
| hsa-miR-194-5p | gastric cancer | CDH2 | "The results of real-time PCR and western blot high ......" | 21845495 |
| hsa-miR-194-5p | liver cancer | CDH2 | "The overexpression of miR-194 in liver mesenchymal ......" | 20979124 |
| hsa-miR-194-5p | colorectal cancer | MAP4K4 | "In addition using dual-luciferase reporter gene as ......" | 25602366 |
| hsa-miR-194-5p | liver cancer | MAP4K4 | "Mitogen-activated protein kinase 4 MAP4K4 a potent ......" | 26722431 |
| hsa-miR-194-5p | bladder cancer | AKT2 | "Long noncoding RNA H19 contributes to gallbladder ......" | 26803515 |
| hsa-miR-194-5p | melanoma | ARHGEF2 | "miR 194 is a negative regulator of GEF H1 pathway ......" | 27573550 |
| hsa-miR-194-5p | liver cancer | C21ORF91 | "Transcripts encoding tripartite motif containing 2 ......" | 26221053 |
| hsa-miR-194-5p | endometrial cancer | CDH1 | "Ectopic expression of miR-194 in EC cells induced ......" | 21851624 |
| hsa-miR-194-5p | lymphoma | CDKN1A | "miR-20a/b and miR-194 target CDKN1A and SOCS2 in f ......" | 22102710 |
| hsa-miR-194-5p | lung squamous cell cancer | CDKN1B | "miR 194 suppresses metastasis of non small cell lu ......" | 23584484 |
| hsa-miR-194-5p | breast cancer | DNM2 | "Increased miR-194 expression markedly reduced leve ......" | 22829924 |
| hsa-miR-194-5p | breast cancer | ERBB2 | "Forced expression of miR-194 in breast cancer cell ......" | 22829924 |
| hsa-miR-194-5p | gastric cancer | F2 | "Though there was no significant difference between ......" | 21845495 |
| hsa-miR-194-5p | lung squamous cell cancer | FOXA1 | "miR 194 inhibits the proliferation invasion migrat ......" | 26909612 |
| hsa-miR-194-5p | gastric cancer | FOXM1 | "We hypothesized that miR-194 may control Forkhead ......" | 24748184 |
| hsa-miR-194-5p | liver cancer | FOXO1 | "Gα12gep oncogene inhibits FOXO1 in hepatocellular ......" | 24631529 |
| hsa-miR-194-5p | bladder cancer | H19 | "Long noncoding RNA H19 contributes to gallbladder ......" | 26803515 |
| hsa-miR-194-5p | liver cancer | MAP2K6 | "Mitogen-activated protein kinase 4 MAP4K4 a potent ......" | 26722431 |
| hsa-miR-194-5p | liver cancer | NFASC | "NF κB signaling relieves negative regulation by m ......" | 26221053 |
| hsa-miR-194-5p | lung squamous cell cancer | NUDC | "Human nuclear distribution C hNUDC was predicted t ......" | 27035759 |
| hsa-miR-194-5p | colorectal cancer | PER1 | "Significant inverse correlation between the expres ......" | 27747302 |
| hsa-miR-194-5p | ovarian cancer | PTPN12 | "Meanwhile bioinformatics tools were used to identi ......" | 27486333 |
| hsa-miR-194-5p | bladder cancer | RAP2B | "MiR 194 inhibits cell proliferation and invasion v ......" | 27133066 |
| hsa-miR-194-5p | gastric cancer | RBX1 | "miR 194 targets RBX1 gene to modulate proliferatio ......" | 25412959 |
| hsa-miR-194-5p | lymphoma | SOCS2 | "Functional studies indicated that CDKN1A/p21 and S ......" | 22102710 |
| hsa-miR-194-5p | colon cancer | THBS1 | "p53 responsive miR 194 inhibits thrombospondin 1 a ......" | 22028325 |
| hsa-miR-194-5p | colon cancer | TP53 | "p53 responsive miR 194 inhibits thrombospondin 1 a ......" | 22028325 |
| hsa-miR-194-5p | liver cancer | TRIM23 | "Transcripts encoding tripartite motif containing 2 ......" | 26221053 |
| hsa-miR-194-5p | endometrial cancer | VIM | "Ectopic expression of miR-194 in EC cells induced ......" | 21851624 |
Expression profile in cancer corhorts:

Putative target regulations
| miRNA | Gene | Cancer | Interaction | Correlation beta |
|---|---|---|---|---|
hsa-miR-194-5p | HBEGF | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; LIHC; PRAD; THCA; UCEC | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | TCGA BLCA -0.242; TCGA BRCA -0.134; TCGA CESC -0.24; TCGA ESCA -0.064; TCGA HNSC -0.281; TCGA KIRC -0.078; TCGA LIHC -0.259; TCGA PRAD -0.398; TCGA THCA -0.592; TCGA UCEC -0.084 |
hsa-miR-194-5p | IGF1R | 9 cancers: BLCA; CESC; ESCA; HNSC; LIHC; PAAD; SARC; STAD; UCEC | miRNAWalker2 validate; miRTarBase; miRNATAP | TCGA BLCA -0.189; TCGA CESC -0.188; TCGA ESCA -0.196; TCGA HNSC -0.091; TCGA LIHC -0.583; TCGA PAAD -0.213; TCGA SARC -0.244; TCGA STAD -0.139; TCGA UCEC -0.113 |
hsa-miR-194-5p | ITGA9 | 9 cancers: BLCA; BRCA; COAD; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | miRNAWalker2 validate; miRTarBase | TCGA BLCA -0.239; TCGA BRCA -0.122; TCGA COAD -0.307; TCGA LIHC -0.293; TCGA LUAD -0.159; TCGA PAAD -0.151; TCGA PRAD -0.607; TCGA STAD -0.305; TCGA UCEC -0.454 |
hsa-miR-194-5p | PTPN12 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LGG; LIHC; LUAD; THCA | miRNAWalker2 validate; miRTarBase | TCGA BLCA -0.121; TCGA BRCA -0.098; TCGA CESC -0.057; TCGA COAD -0.062; TCGA ESCA -0.057; TCGA HNSC -0.178; TCGA KIRP -0.078; TCGA LGG -0.071; TCGA LIHC -0.104; TCGA LUAD -0.058; TCGA THCA -0.136 |
hsa-miR-194-5p | PJA2 | 10 cancers: BLCA; BRCA; COAD; ESCA; HNSC; OV; PRAD; SARC; STAD; UCEC | MirTarget | TCGA BLCA -0.239; TCGA BRCA -0.192; TCGA COAD -0.147; TCGA ESCA -0.076; TCGA HNSC -0.081; TCGA OV -0.054; TCGA PRAD -0.197; TCGA SARC -0.131; TCGA STAD -0.206; TCGA UCEC -0.067 |
hsa-miR-194-5p | EHBP1 | 9 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.226; TCGA BRCA -0.121; TCGA CESC -0.071; TCGA COAD -0.165; TCGA ESCA -0.086; TCGA HNSC -0.129; TCGA THCA -0.183; TCGA STAD -0.197; TCGA UCEC -0.087 |
hsa-miR-194-5p | KLF12 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.321; TCGA BRCA -0.084; TCGA CESC -0.109; TCGA COAD -0.42; TCGA ESCA -0.121; TCGA LUAD -0.07; TCGA OV -0.107; TCGA PAAD -0.251; TCGA PRAD -0.189; TCGA THCA -0.284; TCGA STAD -0.258; TCGA UCEC -0.231 |
hsa-miR-194-5p | LRCH2 | 12 cancers: BLCA; BRCA; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.512; TCGA BRCA -0.347; TCGA COAD -0.751; TCGA ESCA -0.094; TCGA KIRC -0.183; TCGA KIRP -0.303; TCGA LIHC -0.36; TCGA LUAD -0.078; TCGA PAAD -0.147; TCGA PRAD -0.621; TCGA STAD -0.356; TCGA UCEC -0.387 |
hsa-miR-194-5p | BRMS1L | 10 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LUAD; PAAD; STAD; UCEC | MirTarget | TCGA BLCA -0.188; TCGA BRCA -0.121; TCGA CESC -0.141; TCGA COAD -0.081; TCGA ESCA -0.126; TCGA KIRC -0.055; TCGA LUAD -0.089; TCGA PAAD -0.073; TCGA STAD -0.112; TCGA UCEC -0.066 |
hsa-miR-194-5p | DISC1 | 9 cancers: BLCA; BRCA; COAD; KIRP; LIHC; OV; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.086; TCGA BRCA -0.134; TCGA COAD -0.227; TCGA KIRP -0.07; TCGA LIHC -0.346; TCGA OV -0.084; TCGA THCA -0.157; TCGA STAD -0.06; TCGA UCEC -0.131 |
hsa-miR-194-5p | BNIP2 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.065; TCGA BRCA -0.073; TCGA CESC -0.116; TCGA COAD -0.129; TCGA ESCA -0.078; TCGA KIRP -0.086; TCGA PAAD -0.109; TCGA PRAD -0.065; TCGA SARC -0.057; TCGA THCA -0.093; TCGA STAD -0.149; TCGA UCEC -0.1 |
hsa-miR-194-5p | ZFHX4 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; KIRP; LUAD; OV; PAAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.729; TCGA BRCA -0.357; TCGA CESC -0.154; TCGA COAD -1.046; TCGA ESCA -0.306; TCGA KIRC -0.392; TCGA KIRP -0.297; TCGA LUAD -0.107; TCGA OV -0.214; TCGA PAAD -0.259; TCGA THCA -0.3; TCGA STAD -0.364; TCGA UCEC -0.196 |
hsa-miR-194-5p | PITPNM2 | 10 cancers: BLCA; CESC; COAD; ESCA; HNSC; PAAD; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.288; TCGA CESC -0.205; TCGA COAD -0.108; TCGA ESCA -0.115; TCGA HNSC -0.211; TCGA PAAD -0.189; TCGA PRAD -0.183; TCGA SARC -0.121; TCGA STAD -0.214; TCGA UCEC -0.158 |
hsa-miR-194-5p | SLC7A5 | 11 cancers: BLCA; CESC; ESCA; HNSC; KIRC; LIHC; LUSC; PAAD; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.174; TCGA CESC -0.217; TCGA ESCA -0.273; TCGA HNSC -0.444; TCGA KIRC -0.249; TCGA LIHC -0.256; TCGA LUSC -0.289; TCGA PAAD -0.248; TCGA PRAD -0.237; TCGA THCA -0.911; TCGA STAD -0.125 |
hsa-miR-194-5p | TRPS1 | 11 cancers: BLCA; CESC; COAD; ESCA; KIRC; LIHC; LUSC; PAAD; PRAD; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.465; TCGA CESC -0.159; TCGA COAD -0.598; TCGA ESCA -0.26; TCGA KIRC -0.092; TCGA LIHC -0.195; TCGA LUSC -0.154; TCGA PAAD -0.3; TCGA PRAD -0.143; TCGA STAD -0.41; TCGA UCEC -0.102 |
hsa-miR-194-5p | ERG | 11 cancers: BLCA; BRCA; CESC; COAD; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.203; TCGA BRCA -0.327; TCGA CESC -0.213; TCGA COAD -0.452; TCGA LUAD -0.084; TCGA OV -0.09; TCGA PAAD -0.277; TCGA PRAD -0.344; TCGA THCA -0.112; TCGA STAD -0.144; TCGA UCEC -0.24 |
hsa-miR-194-5p | PHLDA1 | 9 cancers: BLCA; CESC; HNSC; KIRP; LGG; LUSC; OV; PRAD; STAD | MirTarget | TCGA BLCA -0.178; TCGA CESC -0.176; TCGA HNSC -0.181; TCGA KIRP -0.219; TCGA LGG -0.14; TCGA LUSC -0.231; TCGA OV -0.175; TCGA PRAD -0.347; TCGA STAD -0.12 |
hsa-miR-194-5p | MAP2 | 13 cancers: BLCA; BRCA; COAD; ESCA; HNSC; KIRC; LIHC; LUAD; OV; PAAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.278; TCGA BRCA -0.199; TCGA COAD -0.693; TCGA ESCA -0.163; TCGA HNSC -0.308; TCGA KIRC -0.129; TCGA LIHC -0.162; TCGA LUAD -0.129; TCGA OV -0.274; TCGA PAAD -0.155; TCGA THCA -0.387; TCGA STAD -0.361; TCGA UCEC -0.155 |
hsa-miR-194-5p | OSTM1 | 14 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PAAD; PRAD; SARC; THCA; STAD | MirTarget | TCGA BLCA -0.24; TCGA BRCA -0.153; TCGA COAD -0.144; TCGA ESCA -0.064; TCGA HNSC -0.137; TCGA LIHC -0.144; TCGA LUAD -0.051; TCGA LUSC -0.087; TCGA OV -0.06; TCGA PAAD -0.142; TCGA PRAD -0.158; TCGA SARC -0.208; TCGA THCA -0.152; TCGA STAD -0.164 |
hsa-miR-194-5p | LRRFIP1 | 10 cancers: BLCA; BRCA; HNSC; KIRC; KIRP; LUSC; PRAD; SARC; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.119; TCGA BRCA -0.089; TCGA HNSC -0.264; TCGA KIRC -0.126; TCGA KIRP -0.054; TCGA LUSC -0.113; TCGA PRAD -0.13; TCGA SARC -0.176; TCGA THCA -0.072; TCGA STAD -0.051 |
hsa-miR-194-5p | TMEM43 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; LIHC; LUAD; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.092; TCGA BRCA -0.115; TCGA CESC -0.116; TCGA COAD -0.106; TCGA ESCA -0.082; TCGA KIRP -0.106; TCGA LIHC -0.202; TCGA LUAD -0.09; TCGA PAAD -0.092; TCGA PRAD -0.104; TCGA SARC -0.104; TCGA THCA -0.163; TCGA STAD -0.176; TCGA UCEC -0.128 |
hsa-miR-194-5p | PARVA | 11 cancers: BLCA; BRCA; COAD; HNSC; LIHC; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.274; TCGA BRCA -0.175; TCGA COAD -0.162; TCGA HNSC -0.079; TCGA LIHC -0.25; TCGA PAAD -0.076; TCGA PRAD -0.284; TCGA SARC -0.188; TCGA THCA -0.184; TCGA STAD -0.239; TCGA UCEC -0.101 |
hsa-miR-194-5p | OSBPL8 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; LIHC; LUSC; OV; PAAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.108; TCGA BRCA -0.149; TCGA CESC -0.07; TCGA COAD -0.111; TCGA ESCA -0.062; TCGA LIHC -0.089; TCGA LUSC -0.061; TCGA OV -0.072; TCGA PAAD -0.121; TCGA THCA -0.145; TCGA STAD -0.096; TCGA UCEC -0.109 |
hsa-miR-194-5p | CDK14 | 11 cancers: BLCA; BRCA; COAD; ESCA; LUAD; LUSC; OV; PAAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.336; TCGA BRCA -0.168; TCGA COAD -0.609; TCGA ESCA -0.195; TCGA LUAD -0.085; TCGA LUSC -0.222; TCGA OV -0.166; TCGA PAAD -0.333; TCGA THCA -0.138; TCGA STAD -0.255; TCGA UCEC -0.125 |
hsa-miR-194-5p | ANGPTL1 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; OV; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.723; TCGA BRCA -0.555; TCGA CESC -0.152; TCGA COAD -1.491; TCGA ESCA -0.105; TCGA KIRC -0.357; TCGA OV -0.359; TCGA PAAD -0.433; TCGA PRAD -0.784; TCGA STAD -0.771; TCGA UCEC -0.367 |
hsa-miR-194-5p | CLIP4 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.461; TCGA BRCA -0.212; TCGA CESC -0.346; TCGA COAD -0.786; TCGA ESCA -0.353; TCGA HNSC -0.259; TCGA KIRC -0.145; TCGA LIHC -0.392; TCGA LUAD -0.083; TCGA PAAD -0.222; TCGA PRAD -0.458; TCGA STAD -0.443; TCGA UCEC -0.127 |
hsa-miR-194-5p | ASAP1 | 10 cancers: BLCA; CESC; COAD; ESCA; LIHC; PAAD; PRAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.108; TCGA CESC -0.131; TCGA COAD -0.149; TCGA ESCA -0.095; TCGA LIHC -0.36; TCGA PAAD -0.226; TCGA PRAD -0.174; TCGA THCA -0.289; TCGA STAD -0.101; TCGA UCEC -0.115 |
hsa-miR-194-5p | BNC1 | 10 cancers: BLCA; BRCA; CESC; ESCA; HNSC; KIRC; LUSC; PRAD; THCA; STAD | MirTarget | TCGA BLCA -1.139; TCGA BRCA -0.433; TCGA CESC -1.306; TCGA ESCA -1.116; TCGA HNSC -0.586; TCGA KIRC -0.234; TCGA LUSC -0.636; TCGA PRAD -0.519; TCGA THCA -0.833; TCGA STAD -0.528 |
hsa-miR-194-5p | SAMD8 | 9 cancers: BLCA; BRCA; CESC; ESCA; KIRC; LIHC; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.124; TCGA BRCA -0.107; TCGA CESC -0.08; TCGA ESCA -0.076; TCGA KIRC -0.062; TCGA LIHC -0.122; TCGA PRAD -0.138; TCGA THCA -0.12; TCGA STAD -0.108 |
hsa-miR-194-5p | PPFIBP1 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUSC; PAAD; PRAD; STAD | MirTarget | TCGA BLCA -0.211; TCGA BRCA -0.215; TCGA CESC -0.102; TCGA COAD -0.197; TCGA ESCA -0.093; TCGA HNSC -0.215; TCGA LIHC -0.137; TCGA LUSC -0.163; TCGA PAAD -0.106; TCGA PRAD -0.105; TCGA STAD -0.121 |
hsa-miR-194-5p | ATP8B4 | 10 cancers: BLCA; BRCA; CESC; COAD; LIHC; LUAD; PAAD; SARC; THCA; STAD | MirTarget | TCGA BLCA -0.416; TCGA BRCA -0.255; TCGA CESC -0.119; TCGA COAD -0.301; TCGA LIHC -0.136; TCGA LUAD -0.072; TCGA PAAD -0.277; TCGA SARC -0.544; TCGA THCA -0.361; TCGA STAD -0.083 |
hsa-miR-194-5p | DMD | 9 cancers: BLCA; BRCA; COAD; KIRC; LUAD; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.515; TCGA BRCA -0.366; TCGA COAD -0.378; TCGA KIRC -0.063; TCGA LUAD -0.078; TCGA PRAD -0.615; TCGA SARC -0.553; TCGA STAD -0.417; TCGA UCEC -0.387 |
hsa-miR-194-5p | TEAD1 | 9 cancers: BLCA; BRCA; COAD; LUAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.181; TCGA BRCA -0.168; TCGA COAD -0.16; TCGA LUAD -0.081; TCGA PRAD -0.346; TCGA SARC -0.164; TCGA THCA -0.185; TCGA STAD -0.268; TCGA UCEC -0.097 |
hsa-miR-194-5p | MAP1B | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; LIHC; LUAD; PAAD; PRAD; SARC; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.574; TCGA BRCA -0.226; TCGA CESC -0.176; TCGA COAD -0.611; TCGA ESCA -0.163; TCGA LIHC -0.445; TCGA LUAD -0.149; TCGA PAAD -0.187; TCGA PRAD -0.626; TCGA SARC -0.351; TCGA THCA -0.544; TCGA STAD -0.426; TCGA UCEC -0.33 |
hsa-miR-194-5p | PPP3CA | 11 cancers: BLCA; BRCA; CESC; COAD; HNSC; LIHC; LUAD; LUSC; OV; THCA; UCEC | MirTarget | TCGA BLCA -0.171; TCGA BRCA -0.132; TCGA CESC -0.06; TCGA COAD -0.07; TCGA HNSC -0.227; TCGA LIHC -0.095; TCGA LUAD -0.07; TCGA LUSC -0.18; TCGA OV -0.14; TCGA THCA -0.142; TCGA UCEC -0.106 |
hsa-miR-194-5p | FLI1 | 11 cancers: BLCA; BRCA; CESC; COAD; LIHC; LUAD; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.266; TCGA BRCA -0.189; TCGA CESC -0.118; TCGA COAD -0.415; TCGA LIHC -0.214; TCGA LUAD -0.067; TCGA PAAD -0.346; TCGA PRAD -0.151; TCGA THCA -0.221; TCGA STAD -0.099; TCGA UCEC -0.124 |
hsa-miR-194-5p | TMEM108 | 12 cancers: BLCA; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; THCA; STAD; UCEC | MirTarget | TCGA BLCA -0.468; TCGA COAD -0.43; TCGA ESCA -0.193; TCGA KIRC -0.107; TCGA KIRP -0.225; TCGA LIHC -0.38; TCGA LUAD -0.095; TCGA PAAD -0.148; TCGA PRAD -0.58; TCGA THCA -0.241; TCGA STAD -0.364; TCGA UCEC -0.364 |
hsa-miR-194-5p | EXOC5 | 9 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LUSC; SARC; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.11; TCGA BRCA -0.103; TCGA CESC -0.077; TCGA ESCA -0.086; TCGA HNSC -0.124; TCGA LUSC -0.08; TCGA SARC -0.09; TCGA THCA -0.069; TCGA STAD -0.083 |
hsa-miR-194-5p | TRIM23 | 9 cancers: BLCA; BRCA; COAD; ESCA; LUAD; PAAD; PRAD; STAD; UCEC | MirTarget | TCGA BLCA -0.135; TCGA BRCA -0.152; TCGA COAD -0.141; TCGA ESCA -0.064; TCGA LUAD -0.079; TCGA PAAD -0.101; TCGA PRAD -0.079; TCGA STAD -0.166; TCGA UCEC -0.098 |
hsa-miR-194-5p | GYG1 | 11 cancers: BLCA; CESC; COAD; ESCA; KIRC; LIHC; PAAD; PRAD; SARC; THCA; STAD | MirTarget; miRNATAP | TCGA BLCA -0.096; TCGA CESC -0.058; TCGA COAD -0.094; TCGA ESCA -0.058; TCGA KIRC -0.088; TCGA LIHC -0.253; TCGA PAAD -0.077; TCGA PRAD -0.212; TCGA SARC -0.289; TCGA THCA -0.212; TCGA STAD -0.219 |
hsa-miR-194-5p | ITPKB | 13 cancers: BLCA; CESC; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; THCA; STAD; UCEC | MirTarget; miRNATAP | TCGA BLCA -0.156; TCGA CESC -0.077; TCGA COAD -0.254; TCGA ESCA -0.16; TCGA KIRC -0.11; TCGA KIRP -0.068; TCGA LIHC -0.271; TCGA LUAD -0.051; TCGA PAAD -0.152; TCGA PRAD -0.071; TCGA THCA -0.117; TCGA STAD -0.358; TCGA UCEC -0.248 |
hsa-miR-194-5p | DCBLD2 | 11 cancers: BLCA; BRCA; COAD; ESCA; KIRC; KIRP; LUAD; PAAD; PRAD; THCA; STAD | MirTarget | TCGA BLCA -0.304; TCGA BRCA -0.202; TCGA COAD -0.437; TCGA ESCA -0.352; TCGA KIRC -0.206; TCGA KIRP -0.208; TCGA LUAD -0.199; TCGA PAAD -0.338; TCGA PRAD -0.392; TCGA THCA -0.328; TCGA STAD -0.292 |
hsa-miR-194-5p | CYP26B1 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LIHC; OV; PAAD; PRAD; UCEC | mirMAP | TCGA BLCA -0.339; TCGA BRCA -0.17; TCGA CESC -0.245; TCGA COAD -0.346; TCGA ESCA -0.139; TCGA HNSC -0.538; TCGA KIRC -0.107; TCGA LIHC -0.572; TCGA OV -0.177; TCGA PAAD -0.286; TCGA PRAD -0.459; TCGA UCEC -0.179 |
hsa-miR-194-5p | GAS7 | 13 cancers: BLCA; BRCA; COAD; ESCA; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.584; TCGA BRCA -0.219; TCGA COAD -0.613; TCGA ESCA -0.128; TCGA LIHC -0.513; TCGA LUAD -0.081; TCGA LUSC -0.172; TCGA OV -0.176; TCGA PAAD -0.303; TCGA PRAD -0.345; TCGA THCA -0.594; TCGA STAD -0.346; TCGA UCEC -0.301 |
hsa-miR-194-5p | CYTH3 | 13 cancers: BLCA; BRCA; COAD; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; STAD; UCEC | mirMAP | TCGA BLCA -0.238; TCGA BRCA -0.15; TCGA COAD -0.162; TCGA HNSC -0.227; TCGA KIRC -0.092; TCGA KIRP -0.091; TCGA LIHC -0.285; TCGA LUAD -0.165; TCGA LUSC -0.127; TCGA OV -0.101; TCGA PAAD -0.15; TCGA STAD -0.079; TCGA UCEC -0.175 |
hsa-miR-194-5p | RBMS2 | 10 cancers: BLCA; BRCA; COAD; HNSC; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.159; TCGA BRCA -0.153; TCGA COAD -0.161; TCGA HNSC -0.11; TCGA LIHC -0.336; TCGA LUAD -0.068; TCGA PAAD -0.129; TCGA PRAD -0.187; TCGA STAD -0.064; TCGA UCEC -0.079 |
hsa-miR-194-5p | SH3PXD2A | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRP; LUSC; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.297; TCGA BRCA -0.087; TCGA CESC -0.112; TCGA COAD -0.119; TCGA ESCA -0.109; TCGA HNSC -0.184; TCGA KIRP -0.104; TCGA LUSC -0.169; TCGA PAAD -0.14; TCGA PRAD -0.337; TCGA THCA -0.249; TCGA STAD -0.176; TCGA UCEC -0.169 |
hsa-miR-194-5p | ALDH1L2 | 10 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LIHC; PAAD; PRAD; THCA; STAD | mirMAP | TCGA BLCA -0.622; TCGA BRCA -0.201; TCGA COAD -0.336; TCGA ESCA -0.099; TCGA HNSC -0.195; TCGA LIHC -0.538; TCGA PAAD -0.308; TCGA PRAD -0.534; TCGA THCA -0.345; TCGA STAD -0.145 |
hsa-miR-194-5p | KIAA1644 | 14 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; OV; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.902; TCGA BRCA -0.27; TCGA COAD -0.888; TCGA ESCA -0.273; TCGA HNSC -0.648; TCGA LIHC -0.636; TCGA LUAD -0.095; TCGA LUSC -0.455; TCGA OV -0.373; TCGA PAAD -0.211; TCGA PRAD -0.963; TCGA THCA -0.697; TCGA STAD -0.49; TCGA UCEC -0.454 |
hsa-miR-194-5p | NTRK3 | 9 cancers: BLCA; COAD; KIRP; OV; PAAD; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.357; TCGA COAD -0.874; TCGA KIRP -0.157; TCGA OV -0.266; TCGA PAAD -0.238; TCGA PRAD -0.588; TCGA THCA -0.256; TCGA STAD -0.458; TCGA UCEC -0.492 |
hsa-miR-194-5p | ENAH | 14 cancers: BLCA; CESC; ESCA; HNSC; LGG; LIHC; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD; UCEC | mirMAP; miRNATAP | TCGA BLCA -0.167; TCGA CESC -0.183; TCGA ESCA -0.142; TCGA HNSC -0.276; TCGA LGG -0.106; TCGA LIHC -0.215; TCGA LUAD -0.093; TCGA LUSC -0.221; TCGA PAAD -0.108; TCGA PRAD -0.099; TCGA SARC -0.219; TCGA THCA -0.327; TCGA STAD -0.149; TCGA UCEC -0.112 |
hsa-miR-194-5p | ADAMTS5 | 13 cancers: BLCA; BRCA; COAD; KIRP; LGG; LIHC; LUAD; LUSC; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BLCA -0.257; TCGA BRCA -0.424; TCGA COAD -0.28; TCGA KIRP -0.344; TCGA LGG -0.153; TCGA LIHC -0.292; TCGA LUAD -0.087; TCGA LUSC -0.333; TCGA OV -0.335; TCGA PAAD -0.221; TCGA PRAD -0.66; TCGA STAD -0.105; TCGA UCEC -0.464 |
hsa-miR-194-5p | PLXDC1 | 12 cancers: BLCA; BRCA; COAD; HNSC; KIRP; LIHC; LUAD; LUSC; PAAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.217; TCGA BRCA -0.234; TCGA COAD -0.268; TCGA HNSC -0.141; TCGA KIRP -0.113; TCGA LIHC -0.644; TCGA LUAD -0.053; TCGA LUSC -0.139; TCGA PAAD -0.207; TCGA THCA -0.515; TCGA STAD -0.063; TCGA UCEC -0.215 |
hsa-miR-194-5p | RNF152 | 11 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUSC; PRAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.195; TCGA BRCA -0.123; TCGA CESC -0.124; TCGA COAD -0.225; TCGA ESCA -0.15; TCGA HNSC -0.296; TCGA LUSC -0.21; TCGA PRAD -0.461; TCGA THCA -0.241; TCGA STAD -0.074; TCGA UCEC -0.158 |
hsa-miR-194-5p | MYO5A | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PAAD; THCA; STAD; UCEC | mirMAP | TCGA BLCA -0.319; TCGA BRCA -0.067; TCGA CESC -0.339; TCGA COAD -0.486; TCGA ESCA -0.288; TCGA HNSC -0.383; TCGA LIHC -0.272; TCGA LUAD -0.109; TCGA LUSC -0.198; TCGA PAAD -0.315; TCGA THCA -0.249; TCGA STAD -0.228; TCGA UCEC -0.114 |
hsa-miR-194-5p | ZNF532 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRP; LIHC; LUAD; LUSC; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.229; TCGA BRCA -0.075; TCGA CESC -0.206; TCGA COAD -0.424; TCGA ESCA -0.172; TCGA KIRP -0.13; TCGA LIHC -0.451; TCGA LUAD -0.056; TCGA LUSC -0.102; TCGA PAAD -0.231; TCGA PRAD -0.173; TCGA STAD -0.242; TCGA UCEC -0.162 |
hsa-miR-194-5p | SAMD4A | 13 cancers: BLCA; BRCA; COAD; HNSC; KIRC; LUAD; LUSC; PAAD; PRAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.458; TCGA BRCA -0.234; TCGA COAD -0.405; TCGA HNSC -0.133; TCGA KIRC -0.051; TCGA LUAD -0.177; TCGA LUSC -0.304; TCGA PAAD -0.232; TCGA PRAD -0.356; TCGA SARC -0.153; TCGA THCA -0.456; TCGA STAD -0.268; TCGA UCEC -0.204 |
hsa-miR-194-5p | SNX29 | 10 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LIHC; LUAD; PAAD; PRAD; STAD | miRNATAP | TCGA BLCA -0.157; TCGA BRCA -0.139; TCGA COAD -0.266; TCGA ESCA -0.118; TCGA HNSC -0.166; TCGA LIHC -0.242; TCGA LUAD -0.056; TCGA PAAD -0.192; TCGA PRAD -0.058; TCGA STAD -0.225 |
hsa-miR-194-5p | DAAM1 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; LUSC; OV; PAAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.206; TCGA BRCA -0.189; TCGA CESC -0.171; TCGA COAD -0.09; TCGA ESCA -0.111; TCGA HNSC -0.439; TCGA KIRC -0.109; TCGA LUAD -0.05; TCGA LUSC -0.226; TCGA OV -0.082; TCGA PAAD -0.09; TCGA SARC -0.205; TCGA STAD -0.221; TCGA UCEC -0.133 |
hsa-miR-194-5p | NRP1 | 12 cancers: BLCA; BRCA; CESC; COAD; HNSC; LIHC; LUAD; LUSC; PAAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.274; TCGA BRCA -0.212; TCGA CESC -0.086; TCGA COAD -0.418; TCGA HNSC -0.137; TCGA LIHC -0.269; TCGA LUAD -0.124; TCGA LUSC -0.261; TCGA PAAD -0.233; TCGA THCA -0.222; TCGA STAD -0.122; TCGA UCEC -0.099 |
hsa-miR-194-5p | PTPRD | 11 cancers: BLCA; BRCA; ESCA; HNSC; LIHC; LUAD; OV; PAAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.57; TCGA BRCA -0.27; TCGA ESCA -0.307; TCGA HNSC -0.257; TCGA LIHC -0.227; TCGA LUAD -0.106; TCGA OV -0.316; TCGA PAAD -0.276; TCGA THCA -0.503; TCGA STAD -0.223; TCGA UCEC -0.382 |
hsa-miR-194-5p | CEP170 | 11 cancers: BLCA; CESC; COAD; ESCA; LIHC; LUAD; OV; PAAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.152; TCGA CESC -0.11; TCGA COAD -0.323; TCGA ESCA -0.051; TCGA LIHC -0.194; TCGA LUAD -0.097; TCGA OV -0.116; TCGA PAAD -0.198; TCGA THCA -0.14; TCGA STAD -0.105; TCGA UCEC -0.11 |
hsa-miR-194-5p | LHX6 | 10 cancers: BLCA; BRCA; CESC; COAD; LIHC; LUAD; PAAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.097; TCGA BRCA -0.176; TCGA CESC -0.163; TCGA COAD -0.418; TCGA LIHC -0.301; TCGA LUAD -0.096; TCGA PAAD -0.204; TCGA THCA -0.422; TCGA STAD -0.136; TCGA UCEC -0.17 |
hsa-miR-194-5p | PER3 | 9 cancers: BLCA; BRCA; COAD; LUAD; LUSC; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.219; TCGA BRCA -0.164; TCGA COAD -0.178; TCGA LUAD -0.131; TCGA LUSC -0.164; TCGA PRAD -0.269; TCGA SARC -0.228; TCGA STAD -0.22; TCGA UCEC -0.151 |
hsa-miR-194-5p | EYA4 | 10 cancers: BLCA; BRCA; COAD; ESCA; KIRC; LIHC; OV; PAAD; PRAD; STAD | miRNATAP | TCGA BLCA -0.485; TCGA BRCA -0.549; TCGA COAD -0.693; TCGA ESCA -0.228; TCGA KIRC -0.39; TCGA LIHC -0.295; TCGA OV -0.302; TCGA PAAD -0.367; TCGA PRAD -0.887; TCGA STAD -0.217 |
hsa-miR-194-5p | MEF2C | 11 cancers: BLCA; BRCA; CESC; COAD; LIHC; LUAD; PAAD; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.286; TCGA BRCA -0.24; TCGA CESC -0.067; TCGA COAD -0.392; TCGA LIHC -0.2; TCGA LUAD -0.126; TCGA PAAD -0.271; TCGA PRAD -0.419; TCGA SARC -0.202; TCGA STAD -0.23; TCGA UCEC -0.205 |
hsa-miR-194-5p | CHD9 | 9 cancers: BLCA; BRCA; CESC; ESCA; HNSC; LUSC; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.132; TCGA BRCA -0.206; TCGA CESC -0.059; TCGA ESCA -0.076; TCGA HNSC -0.175; TCGA LUSC -0.082; TCGA PRAD -0.12; TCGA STAD -0.099; TCGA UCEC -0.064 |
hsa-miR-194-5p | KLF7 | 14 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; LUSC; PRAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.283; TCGA BRCA -0.197; TCGA CESC -0.152; TCGA COAD -0.206; TCGA ESCA -0.253; TCGA HNSC -0.413; TCGA LIHC -0.399; TCGA LUAD -0.053; TCGA LUSC -0.243; TCGA PRAD -0.251; TCGA SARC -0.271; TCGA THCA -0.223; TCGA STAD -0.16; TCGA UCEC -0.141 |
hsa-miR-194-5p | BICD2 | 11 cancers: BLCA; CESC; COAD; ESCA; HNSC; LIHC; LUAD; PAAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.164; TCGA CESC -0.38; TCGA COAD -0.125; TCGA ESCA -0.296; TCGA HNSC -0.242; TCGA LIHC -0.124; TCGA LUAD -0.082; TCGA PAAD -0.242; TCGA THCA -0.07; TCGA STAD -0.224; TCGA UCEC -0.089 |
hsa-miR-194-5p | FOXO1 | 10 cancers: BLCA; BRCA; COAD; KIRC; LUSC; OV; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.178; TCGA BRCA -0.306; TCGA COAD -0.157; TCGA KIRC -0.09; TCGA LUSC -0.106; TCGA OV -0.165; TCGA PAAD -0.224; TCGA PRAD -0.304; TCGA STAD -0.114; TCGA UCEC -0.147 |
hsa-miR-194-5p | FOXP2 | 9 cancers: BLCA; BRCA; COAD; ESCA; OV; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.644; TCGA BRCA -0.517; TCGA COAD -0.881; TCGA ESCA -0.484; TCGA OV -0.337; TCGA PAAD -0.207; TCGA PRAD -0.728; TCGA STAD -0.632; TCGA UCEC -0.294 |
hsa-miR-194-5p | PAFAH1B1 | 10 cancers: BLCA; BRCA; COAD; HNSC; LIHC; LUSC; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.099; TCGA BRCA -0.117; TCGA COAD -0.079; TCGA HNSC -0.11; TCGA LIHC -0.098; TCGA LUSC -0.079; TCGA PAAD -0.059; TCGA PRAD -0.112; TCGA STAD -0.078; TCGA UCEC -0.052 |
hsa-miR-194-5p | GPM6B | 12 cancers: BLCA; BRCA; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.381; TCGA BRCA -0.22; TCGA COAD -0.538; TCGA ESCA -0.24; TCGA KIRC -0.464; TCGA KIRP -0.186; TCGA LIHC -0.513; TCGA LUAD -0.076; TCGA PAAD -0.329; TCGA PRAD -0.236; TCGA STAD -0.561; TCGA UCEC -0.188 |
hsa-miR-194-5p | MAF | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; LUSC; PAAD; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.441; TCGA BRCA -0.214; TCGA CESC -0.36; TCGA COAD -0.411; TCGA ESCA -0.308; TCGA HNSC -0.257; TCGA LUSC -0.276; TCGA PAAD -0.22; TCGA PRAD -0.125; TCGA THCA -0.085; TCGA STAD -0.176; TCGA UCEC -0.223 |
hsa-miR-194-5p | PKD2 | 14 cancers: BLCA; BRCA; COAD; ESCA; KIRC; LIHC; LUAD; OV; PAAD; PRAD; SARC; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.181; TCGA BRCA -0.311; TCGA COAD -0.425; TCGA ESCA -0.077; TCGA KIRC -0.064; TCGA LIHC -0.352; TCGA LUAD -0.085; TCGA OV -0.104; TCGA PAAD -0.175; TCGA PRAD -0.125; TCGA SARC -0.173; TCGA THCA -0.088; TCGA STAD -0.302; TCGA UCEC -0.208 |
hsa-miR-194-5p | EGR2 | 12 cancers: BLCA; BRCA; CESC; COAD; ESCA; KIRC; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | miRNATAP | TCGA BLCA -0.46; TCGA BRCA -0.259; TCGA CESC -0.121; TCGA COAD -0.53; TCGA ESCA -0.225; TCGA KIRC -0.183; TCGA LIHC -0.45; TCGA LUAD -0.061; TCGA PAAD -0.283; TCGA PRAD -0.346; TCGA STAD -0.125; TCGA UCEC -0.202 |
hsa-miR-194-5p | ZEB1 | 11 cancers: BLCA; BRCA; COAD; LGG; LUAD; OV; PAAD; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.465; TCGA BRCA -0.324; TCGA COAD -0.564; TCGA LGG -0.124; TCGA LUAD -0.059; TCGA OV -0.163; TCGA PAAD -0.195; TCGA PRAD -0.433; TCGA THCA -0.2; TCGA STAD -0.356; TCGA UCEC -0.293 |
hsa-miR-194-5p | MYH10 | 13 cancers: BLCA; BRCA; CESC; COAD; ESCA; HNSC; KIRC; LUAD; OV; PAAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.13; TCGA BRCA -0.081; TCGA CESC -0.139; TCGA COAD -0.513; TCGA ESCA -0.287; TCGA HNSC -0.403; TCGA KIRC -0.183; TCGA LUAD -0.096; TCGA OV -0.099; TCGA PAAD -0.228; TCGA THCA -0.441; TCGA STAD -0.256; TCGA UCEC -0.135 |
hsa-miR-194-5p | MAP3K3 | 13 cancers: BLCA; BRCA; CESC; COAD; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; THCA; STAD; UCEC | miRNATAP | TCGA BLCA -0.191; TCGA BRCA -0.098; TCGA CESC -0.069; TCGA COAD -0.278; TCGA KIRC -0.066; TCGA KIRP -0.092; TCGA LIHC -0.076; TCGA LUAD -0.094; TCGA PAAD -0.201; TCGA PRAD -0.075; TCGA THCA -0.151; TCGA STAD -0.211; TCGA UCEC -0.139 |
hsa-miR-194-5p | LIMK2 | 9 cancers: BLCA; CESC; ESCA; HNSC; KIRC; LIHC; LUSC; PRAD; SARC | miRNATAP | TCGA BLCA -0.068; TCGA CESC -0.144; TCGA ESCA -0.071; TCGA HNSC -0.269; TCGA KIRC -0.078; TCGA LIHC -0.262; TCGA LUSC -0.245; TCGA PRAD -0.153; TCGA SARC -0.09 |
hsa-miR-194-5p | PPP2R2C | 9 cancers: BLCA; CESC; ESCA; HNSC; LIHC; LUSC; PRAD; THCA; STAD | miRNATAP | TCGA BLCA -0.361; TCGA CESC -0.291; TCGA ESCA -0.724; TCGA HNSC -0.907; TCGA LIHC -0.956; TCGA LUSC -0.382; TCGA PRAD -0.155; TCGA THCA -0.402; TCGA STAD -0.504 |
hsa-miR-194-5p | MBNL2 | 9 cancers: BLCA; BRCA; CESC; HNSC; OV; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.095; TCGA BRCA -0.192; TCGA CESC -0.081; TCGA HNSC -0.14; TCGA OV -0.095; TCGA PRAD -0.517; TCGA SARC -0.178; TCGA STAD -0.104; TCGA UCEC -0.125 |
hsa-miR-194-5p | GLIS3 | 10 cancers: BLCA; BRCA; COAD; KIRC; LIHC; LUAD; LUSC; PAAD; PRAD; SARC | miRNATAP | TCGA BLCA -0.217; TCGA BRCA -0.164; TCGA COAD -0.471; TCGA KIRC -0.056; TCGA LIHC -0.454; TCGA LUAD -0.058; TCGA LUSC -0.292; TCGA PAAD -0.127; TCGA PRAD -0.468; TCGA SARC -0.507 |
hsa-miR-194-5p | SLC2A3 | 10 cancers: BLCA; BRCA; CESC; COAD; KIRP; LIHC; PAAD; PRAD; THCA; STAD | miRNATAP | TCGA BLCA -0.41; TCGA BRCA -0.135; TCGA CESC -0.077; TCGA COAD -0.405; TCGA KIRP -0.111; TCGA LIHC -0.304; TCGA PAAD -0.269; TCGA PRAD -0.267; TCGA THCA -0.467; TCGA STAD -0.093 |
hsa-miR-194-5p | ZNF25 | 11 cancers: BLCA; BRCA; COAD; ESCA; KIRP; LIHC; LUAD; PAAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.173; TCGA BRCA -0.143; TCGA COAD -0.261; TCGA ESCA -0.064; TCGA KIRP -0.06; TCGA LIHC -0.125; TCGA LUAD -0.068; TCGA PAAD -0.118; TCGA SARC -0.115; TCGA STAD -0.264; TCGA UCEC -0.199 |
hsa-miR-194-5p | PRICKLE2 | 13 cancers: BLCA; BRCA; COAD; ESCA; HNSC; LIHC; LUAD; OV; PAAD; PRAD; SARC; STAD; UCEC | miRNATAP | TCGA BLCA -0.333; TCGA BRCA -0.291; TCGA COAD -0.509; TCGA ESCA -0.125; TCGA HNSC -0.107; TCGA LIHC -0.256; TCGA LUAD -0.1; TCGA OV -0.092; TCGA PAAD -0.25; TCGA PRAD -0.669; TCGA SARC -0.188; TCGA STAD -0.48; TCGA UCEC -0.325 |
hsa-miR-194-5p | PTPN13 | 14 cancers: BRCA; CESC; COAD; ESCA; HNSC; KIRC; KIRP; LIHC; LUAD; LUSC; OV; PAAD; STAD; UCEC | miRNAWalker2 validate; miRTarBase | TCGA BRCA -0.212; TCGA CESC -0.315; TCGA COAD -0.347; TCGA ESCA -0.498; TCGA HNSC -0.127; TCGA KIRC -0.136; TCGA KIRP -0.132; TCGA LIHC -0.775; TCGA LUAD -0.127; TCGA LUSC -0.208; TCGA OV -0.308; TCGA PAAD -0.207; TCGA STAD -0.447; TCGA UCEC -0.181 |
hsa-miR-194-5p | ZFHX3 | 9 cancers: BRCA; COAD; KIRC; KIRP; OV; PRAD; SARC; STAD; UCEC | MirTarget; miRNATAP | TCGA BRCA -0.157; TCGA COAD -0.113; TCGA KIRC -0.101; TCGA KIRP -0.058; TCGA OV -0.068; TCGA PRAD -0.135; TCGA SARC -0.115; TCGA STAD -0.156; TCGA UCEC -0.096 |
hsa-miR-194-5p | SLC1A3 | 10 cancers: BRCA; CESC; COAD; ESCA; LUAD; LUSC; PAAD; PRAD; THCA; STAD | MirTarget | TCGA BRCA -0.265; TCGA CESC -0.319; TCGA COAD -0.538; TCGA ESCA -0.231; TCGA LUAD -0.059; TCGA LUSC -0.258; TCGA PAAD -0.452; TCGA PRAD -0.231; TCGA THCA -0.408; TCGA STAD -0.092 |
hsa-miR-194-5p | ATP6V1E1 | 9 cancers: BRCA; CESC; ESCA; HNSC; LIHC; OV; SARC; THCA; STAD | MirTarget | TCGA BRCA -0.058; TCGA CESC -0.062; TCGA ESCA -0.068; TCGA HNSC -0.064; TCGA LIHC -0.08; TCGA OV -0.071; TCGA SARC -0.091; TCGA THCA -0.089; TCGA STAD -0.078 |
hsa-miR-194-5p | FZD6 | 11 cancers: BRCA; CESC; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; SARC; STAD; UCEC | MirTarget | TCGA BRCA -0.157; TCGA CESC -0.251; TCGA ESCA -0.253; TCGA KIRC -0.196; TCGA KIRP -0.094; TCGA LIHC -0.265; TCGA LUAD -0.071; TCGA PAAD -0.113; TCGA SARC -0.482; TCGA STAD -0.115; TCGA UCEC -0.083 |
hsa-miR-194-5p | WTIP | 12 cancers: BRCA; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; OV; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA BRCA -0.109; TCGA COAD -0.391; TCGA ESCA -0.123; TCGA KIRC -0.15; TCGA KIRP -0.117; TCGA LIHC -0.519; TCGA LUAD -0.109; TCGA OV -0.166; TCGA PAAD -0.215; TCGA PRAD -0.137; TCGA STAD -0.183; TCGA UCEC -0.214 |
hsa-miR-194-5p | RAB6B | 9 cancers: BRCA; COAD; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; STAD | miRNATAP | TCGA BRCA -0.114; TCGA COAD -0.364; TCGA ESCA -0.122; TCGA KIRC -0.194; TCGA KIRP -0.058; TCGA LIHC -0.449; TCGA LUAD -0.058; TCGA PAAD -0.128; TCGA STAD -0.289 |
hsa-miR-194-5p | CADM1 | 9 cancers: BRCA; COAD; KIRC; KIRP; LUAD; OV; PAAD; STAD; UCEC | miRNATAP | TCGA BRCA -0.126; TCGA COAD -0.453; TCGA KIRC -0.168; TCGA KIRP -0.057; TCGA LUAD -0.139; TCGA OV -0.105; TCGA PAAD -0.222; TCGA STAD -0.28; TCGA UCEC -0.101 |
hsa-miR-194-5p | GRHL3 | 9 cancers: CESC; ESCA; HNSC; KIRC; KIRP; LGG; LUAD; LUSC; THCA | MirTarget | TCGA CESC -0.452; TCGA ESCA -0.45; TCGA HNSC -0.315; TCGA KIRC -0.62; TCGA KIRP -0.471; TCGA LGG -0.326; TCGA LUAD -0.098; TCGA LUSC -0.463; TCGA THCA -1.263 |
hsa-miR-194-5p | MTSS1 | 9 cancers: CESC; ESCA; HNSC; KIRP; LGG; LUSC; PAAD; STAD; UCEC | MirTarget; miRNATAP | TCGA CESC -0.555; TCGA ESCA -0.388; TCGA HNSC -0.254; TCGA KIRP -0.143; TCGA LGG -0.108; TCGA LUSC -0.223; TCGA PAAD -0.094; TCGA STAD -0.22; TCGA UCEC -0.169 |
hsa-miR-194-5p | TUSC3 | 9 cancers: COAD; ESCA; KIRC; LIHC; OV; PAAD; THCA; STAD; UCEC | MirTarget | TCGA COAD -0.39; TCGA ESCA -0.43; TCGA KIRC -0.118; TCGA LIHC -0.681; TCGA OV -0.115; TCGA PAAD -0.226; TCGA THCA -0.735; TCGA STAD -0.317; TCGA UCEC -0.315 |
hsa-miR-194-5p | MPP2 | 10 cancers: COAD; ESCA; KIRC; KIRP; LIHC; LUAD; PAAD; PRAD; STAD; UCEC | mirMAP | TCGA COAD -0.561; TCGA ESCA -0.187; TCGA KIRC -0.488; TCGA KIRP -0.136; TCGA LIHC -0.497; TCGA LUAD -0.09; TCGA PAAD -0.195; TCGA PRAD -0.713; TCGA STAD -0.41; TCGA UCEC -0.234 |
Enriched cancer pathways of putative targets