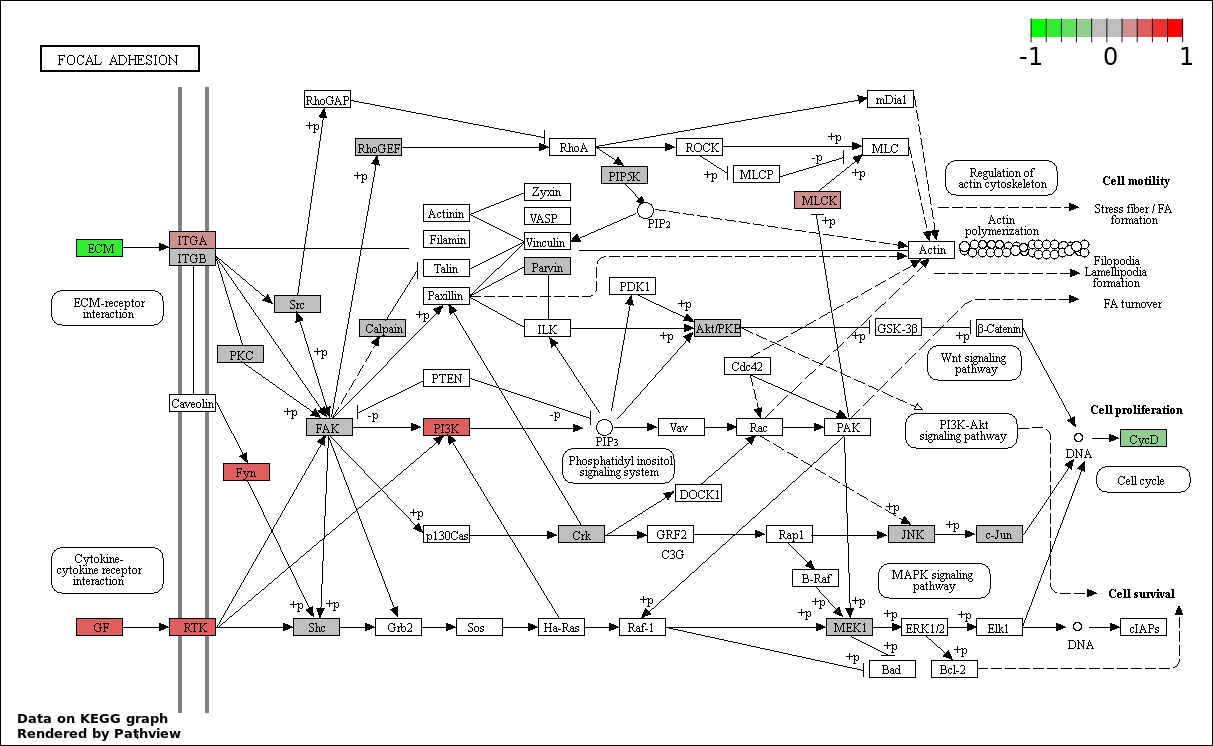
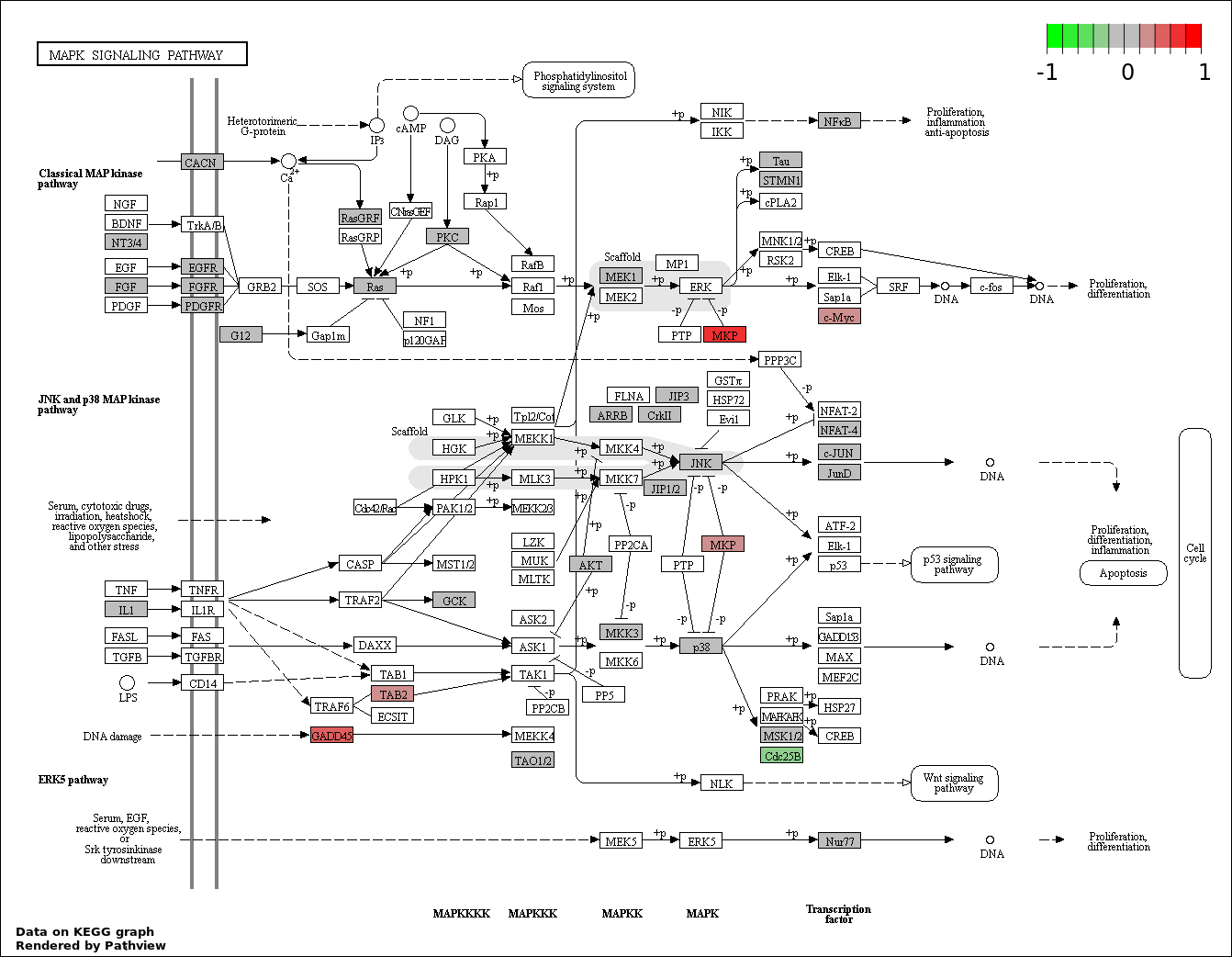
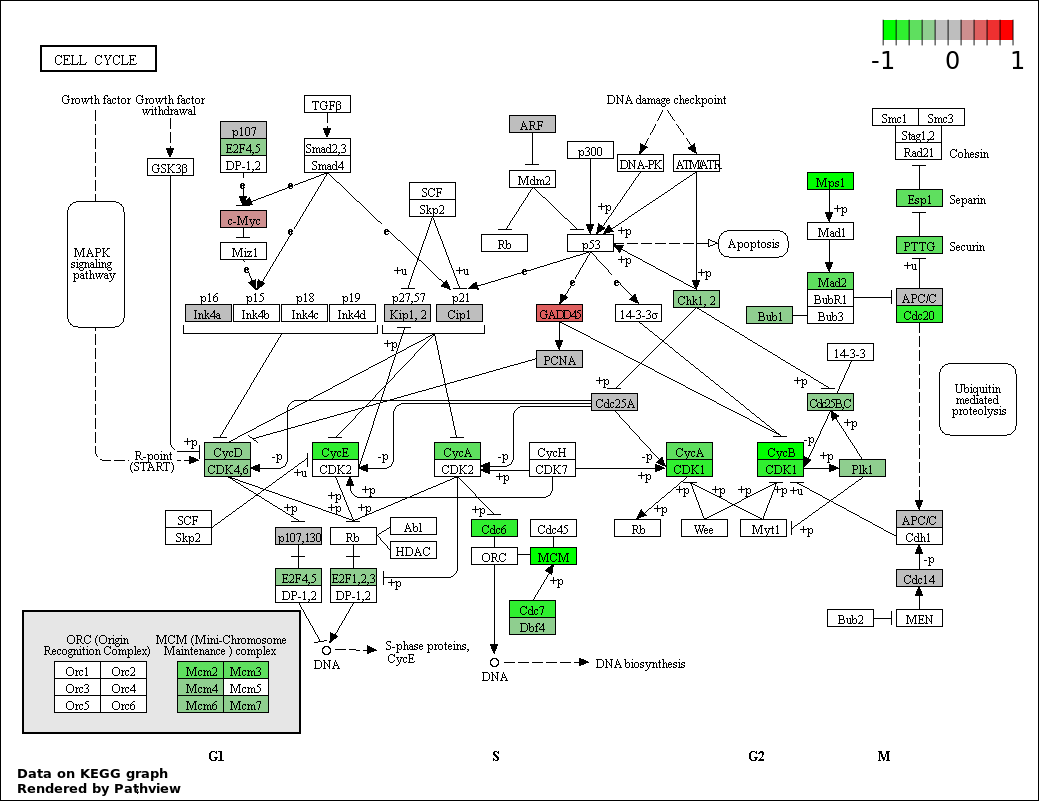
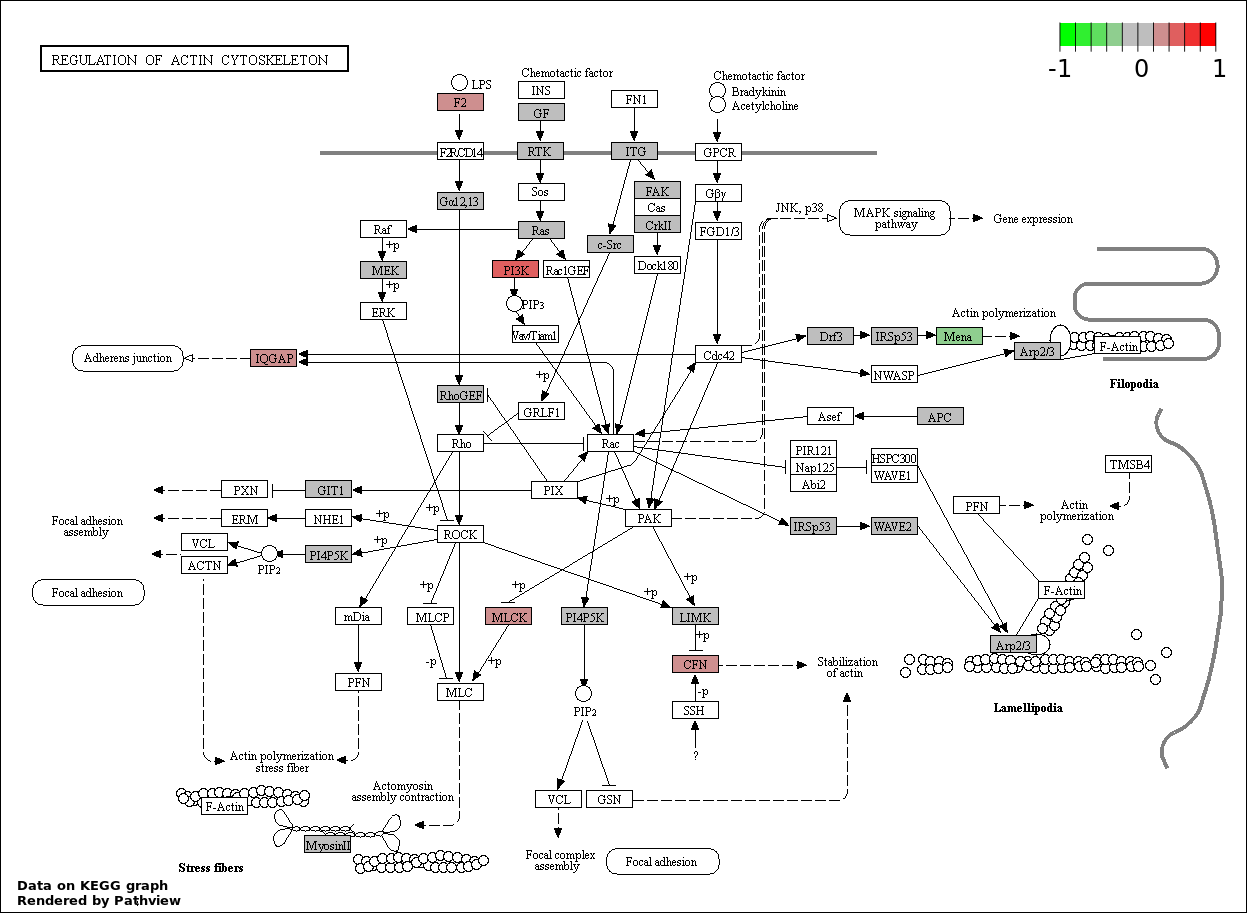
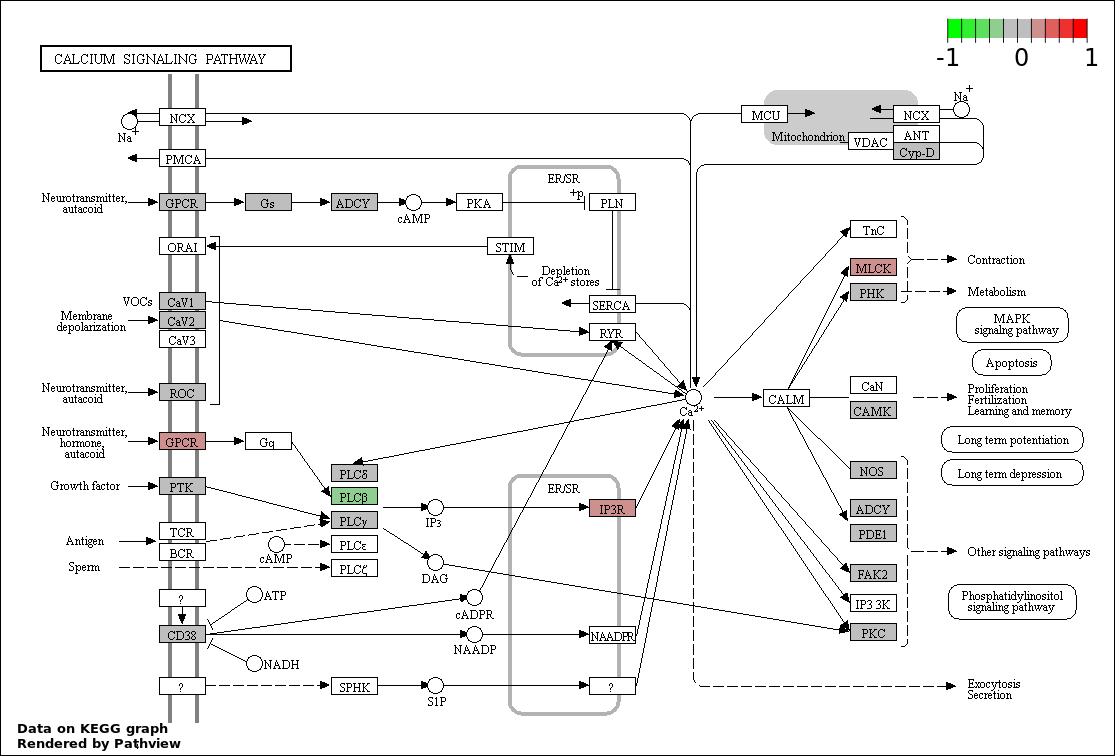
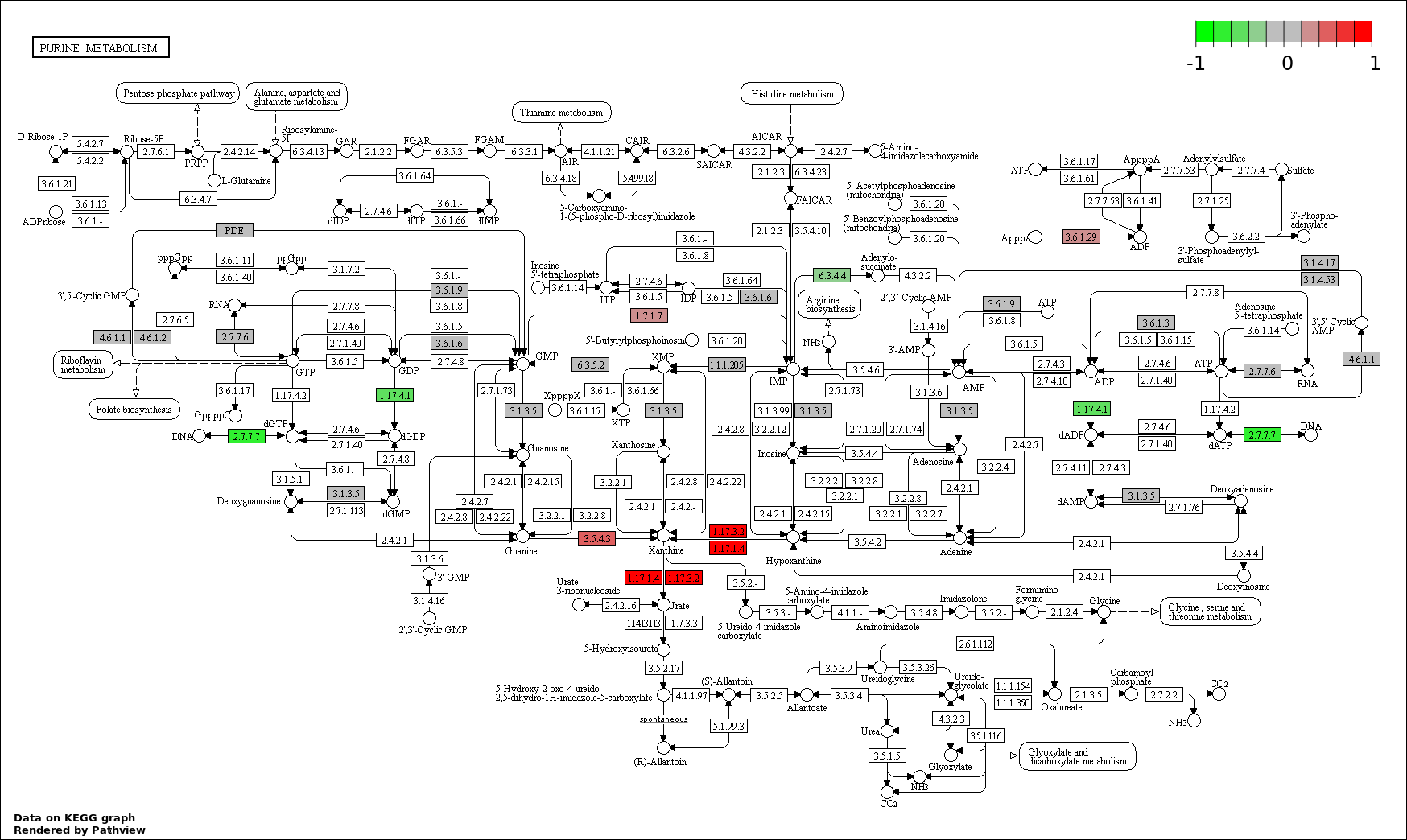
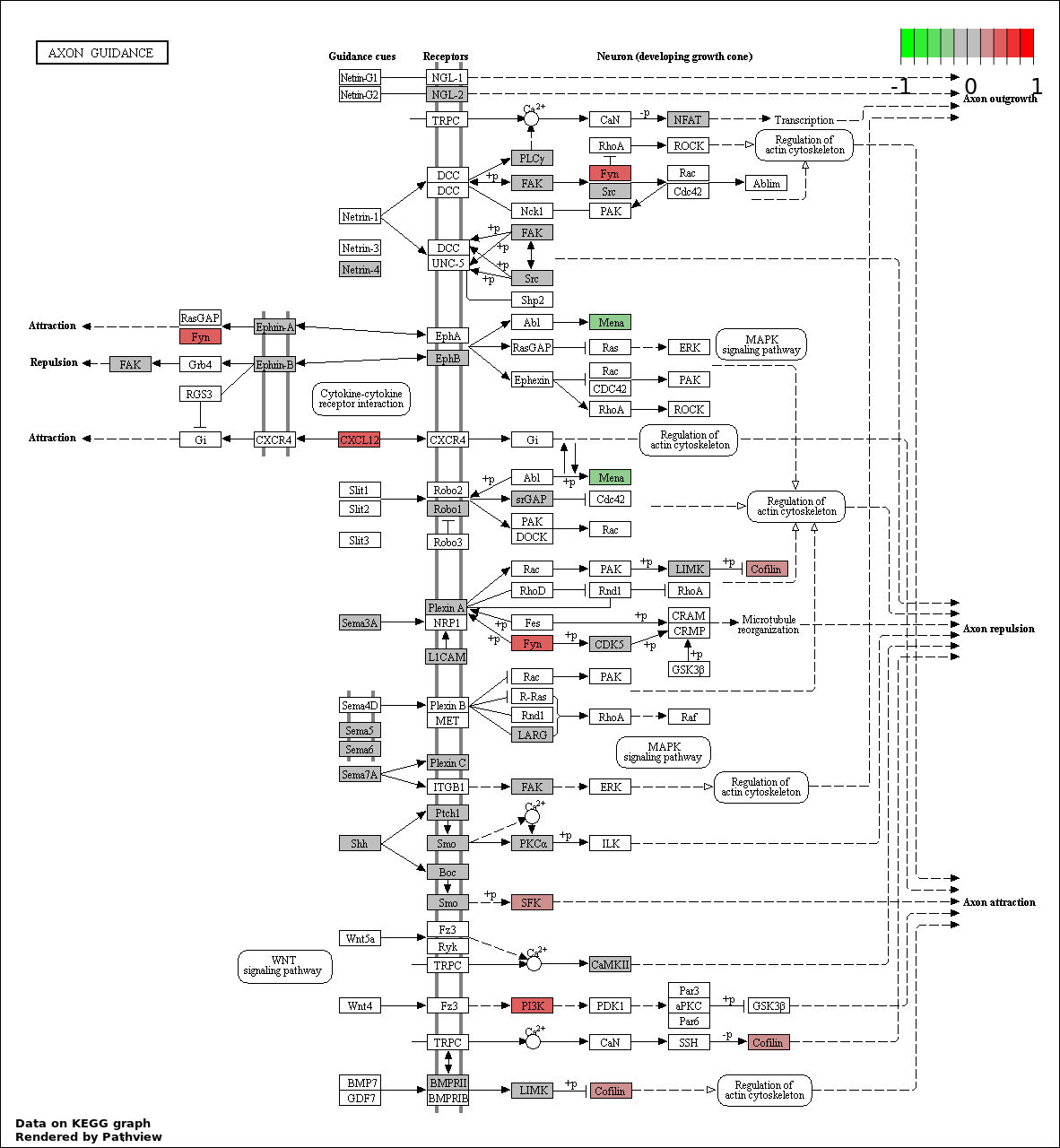
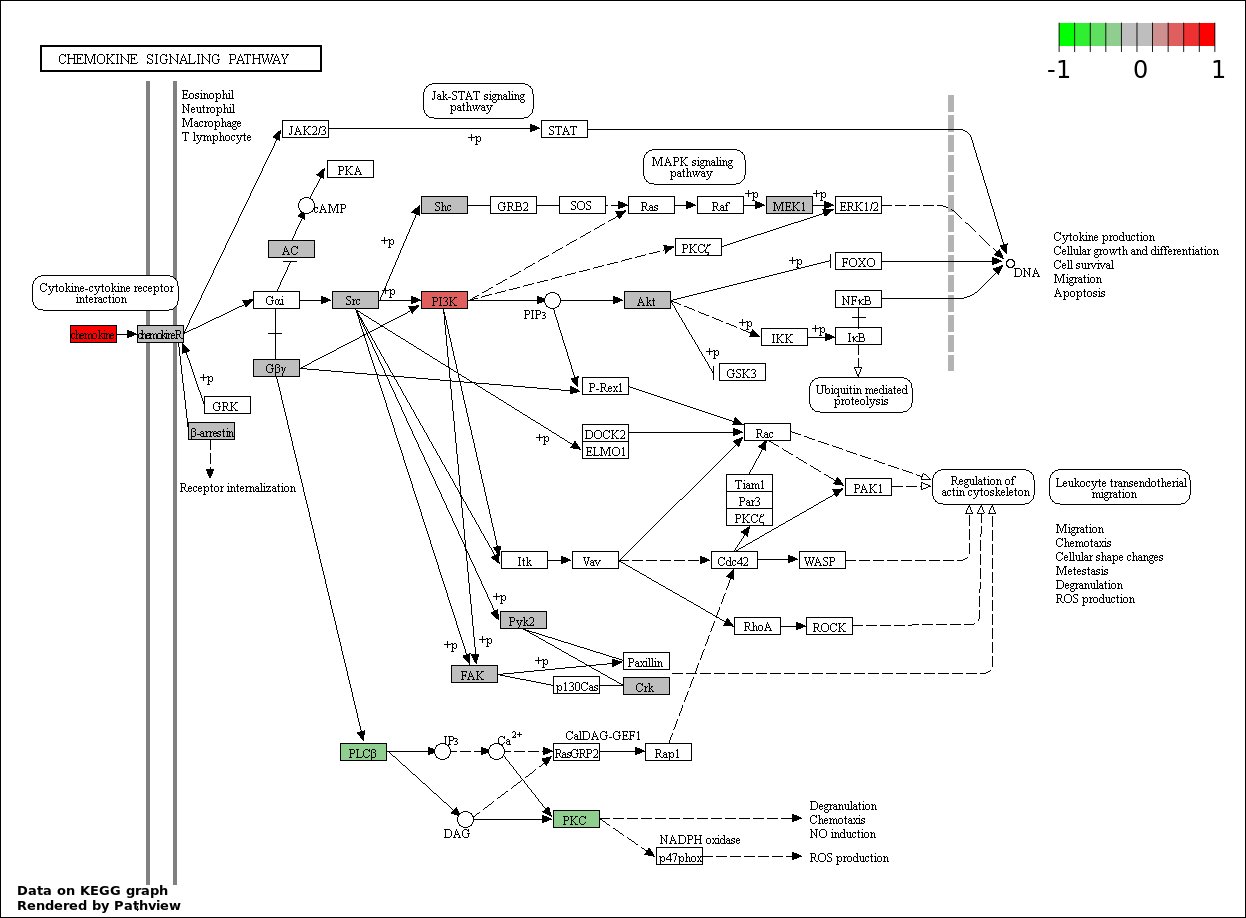
This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-let-7a-5p | FAM171A2 | 0.06 | 0.9566 | 0.1 | 0.82435 | miRNAWalker2 validate | -0.38 | 0.04942 | ||
| 2 | hsa-let-7a-5p | HOXC4 | 0.06 | 0.9566 | -0.06 | 0.90675 | miRNAWalker2 validate | -0.4 | 0.026 | ||
| 3 | hsa-let-7a-5p | HSF2 | 0.06 | 0.9566 | -0.22 | 0.66591 | miRNAWalker2 validate | -0.92 | 0.00536 | ||
| 4 | hsa-let-7a-5p | REPS1 | 0.06 | 0.9566 | 0.18 | 0.80922 | miRNAWalker2 validate | -0.87 | 0.02632 | ||
| 5 | hsa-let-7a-5p | SKA3 | 0.06 | 0.9566 | -0.2 | 0.72348 | miRNAWalker2 validate | -0.74 | 0.012 | ||
| 6 | hsa-let-7a-5p | SNRPC | 0.06 | 0.9566 | -0.09 | 0.90563 | miRNAWalker2 validate | -0.73 | 0.007 | ||
| 7 | hsa-let-7b-5p | AATF | 0.02 | 0.98368 | -0.22 | 0.78826 | miRNAWalker2 validate | -0.36 | 0 | ||
| 8 | hsa-let-7b-5p | ACACA | 0.02 | 0.98368 | -0.18 | 0.7702 | miRNAWalker2 validate | -0.35 | 1.0E-5 | ||
| 9 | hsa-let-7b-5p | ALG3 | 0.02 | 0.98368 | -0.18 | 0.82449 | miRNAWalker2 validate | -0.36 | 0.00017 | ||
| 10 | hsa-let-7b-5p | ASPSCR1 | 0.02 | 0.98368 | -0.11 | 0.8742 | miRNAWalker2 validate | -0.58 | 0.00632 | ||
| 11 | hsa-let-7b-5p | ATAD3B | 0.02 | 0.98368 | -0.01 | 0.98938 | miRNAWalker2 validate | -0.09 | 0.04214 | ||
| 12 | hsa-let-7b-5p | ATP6V1F | 0.02 | 0.98368 | -0.1 | 0.90016 | miRNAWalker2 validate | -0.18 | 0.01116 | ||
| 13 | hsa-let-7b-5p | AURKA | 0.02 | 0.98368 | -0.39 | 0.56777 | miRNAWalker2 validate | -0.58 | 0.00184 | ||
| 14 | hsa-let-7b-5p | BIRC5 | 0.02 | 0.98368 | -0.52 | 0.35824 | miRNAWalker2 validate | -0.43 | 0.00183 | ||
| 15 | hsa-let-7b-5p | CCNA2 | 0.02 | 0.98368 | -0.45 | 0.39878 | miRNAWalker2 validate; miRTarBase | -0.45 | 0.00615 | ||
| 16 | hsa-let-7b-5p | CCNB1 | 0.02 | 0.98368 | -0.85 | 0.15986 | miRNAWalker2 validate | -0.74 | 0.001 | ||
| 17 | hsa-let-7b-5p | CCNB2 | 0.02 | 0.98368 | -0.71 | 0.25562 | miRNAWalker2 validate | -0.71 | 0.00351 | ||
| 18 | hsa-let-7b-5p | CCNF | 0.02 | 0.98368 | -0.14 | 0.78436 | miRNAWalker2 validate; MirTarget | -0.12 | 0.01767 | ||
| 19 | hsa-let-7b-5p | CDC25A | 0.02 | 0.98368 | -0.17 | 0.70355 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.14 | 0.04849 | ||
| 20 | hsa-let-7b-5p | CIZ1 | 0.02 | 0.98368 | -0.14 | 0.80332 | miRNAWalker2 validate | -0.1 | 0.03881 | ||
| 21 | hsa-let-7b-5p | CKAP2 | 0.02 | 0.98368 | -0.15 | 0.72955 | miRNAWalker2 validate | -0.23 | 0.01508 | ||
| 22 | hsa-let-7b-5p | CKS2 | 0.02 | 0.98368 | -0.39 | 0.63253 | miRNAWalker2 validate | -0.61 | 0.00094 | ||
| 23 | hsa-let-7b-5p | DDX21 | 0.02 | 0.98368 | 0.06 | 0.93742 | miRNAWalker2 validate | -0.22 | 0.03681 | ||
| 24 | hsa-let-7b-5p | DDX41 | 0.02 | 0.98368 | -0.1 | 0.88826 | miRNAWalker2 validate | -0.22 | 0.00618 | ||
| 25 | hsa-let-7b-5p | DDX49 | 0.02 | 0.98368 | -0.17 | 0.77652 | miRNAWalker2 validate | -0.13 | 0.04053 | ||
| 26 | hsa-let-7b-5p | DHX57 | 0.02 | 0.98368 | -0.07 | 0.8799 | miRNAWalker2 validate | -0.11 | 0.038 | ||
| 27 | hsa-let-7b-5p | DPF2 | 0.02 | 0.98368 | -0.11 | 0.87328 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.2 | 0.00176 | ||
| 28 | hsa-let-7b-5p | E2F5 | 0.02 | 0.98368 | -0.38 | 0.46904 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.41 | 0.00468 | ||
| 29 | hsa-let-7b-5p | FAM49B | 0.02 | 0.98368 | -0.26 | 0.62777 | miRNAWalker2 validate | -0.41 | 4.0E-5 | ||
| 30 | hsa-let-7b-5p | FOXK1 | 0.02 | 0.98368 | -0.07 | 0.88647 | miRNAWalker2 validate | -0.12 | 0.03407 | ||
| 31 | hsa-let-7b-5p | GPATCH4 | 0.02 | 0.98368 | -0.15 | 0.83977 | miRNAWalker2 validate | -0.29 | 0.00596 | ||
| 32 | hsa-let-7b-5p | GRPEL2 | 0.02 | 0.98368 | -0.07 | 0.9018 | miRNAWalker2 validate; MirTarget | -0.17 | 0.01287 | ||
| 33 | hsa-let-7b-5p | HIST1H1C | 0.02 | 0.98368 | -0.09 | 0.91755 | miRNAWalker2 validate | -0.47 | 0.03675 | ||
| 34 | hsa-let-7b-5p | HSF2 | 0.02 | 0.98368 | -0.22 | 0.66591 | miRNAWalker2 validate | -0.34 | 0.00027 | ||
| 35 | hsa-let-7b-5p | IGF2BP1 | 0.02 | 0.98368 | -0.26 | 0.60749 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.33 | 0.00092 | ||
| 36 | hsa-let-7b-5p | IGF2BP3 | 0.02 | 0.98368 | -0.31 | 0.57427 | miRNAWalker2 validate; MirTarget | -0.65 | 0.00246 | ||
| 37 | hsa-let-7b-5p | IMPDH2 | 0.02 | 0.98368 | -0.18 | 0.8283 | miRNAWalker2 validate | -0.39 | 0.00189 | ||
| 38 | hsa-let-7b-5p | KIF2A | 0.02 | 0.98368 | -0.28 | 0.65454 | miRNAWalker2 validate | -0.44 | 0.00595 | ||
| 39 | hsa-let-7b-5p | KIFC1 | 0.02 | 0.98368 | -0.21 | 0.72576 | miRNAWalker2 validate | -0.16 | 0.04412 | ||
| 40 | hsa-let-7b-5p | LARP1 | 0.02 | 0.98368 | -0.05 | 0.93886 | miRNAWalker2 validate | -0.17 | 0.01736 | ||
| 41 | hsa-let-7b-5p | LIN28B | 0.02 | 0.98368 | -0.74 | 0.20501 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -1.19 | 0.00091 | ||
| 42 | hsa-let-7b-5p | MCM4 | 0.02 | 0.98368 | -0.29 | 0.63472 | miRNAWalker2 validate | -0.47 | 0.00032 | ||
| 43 | hsa-let-7b-5p | MCM7 | 0.02 | 0.98368 | -0.34 | 0.62445 | miRNAWalker2 validate | -0.26 | 0.01099 | ||
| 44 | hsa-let-7b-5p | MIPEP | 0.02 | 0.98368 | 0.06 | 0.9247 | miRNAWalker2 validate | -0.2 | 0.02658 | ||
| 45 | hsa-let-7b-5p | MSI1 | 0.02 | 0.98368 | 0.08 | 0.86414 | miRNAWalker2 validate | -0.14 | 0.01223 | ||
| 46 | hsa-let-7b-5p | MYC | 0.02 | 0.98368 | 0.33 | 0.7029 | miRNAWalker2 validate | -0.52 | 0.0321 | ||
| 47 | hsa-let-7b-5p | NCAPD2 | 0.02 | 0.98368 | -0.41 | 0.4535 | miRNAWalker2 validate | -0.3 | 0.01038 | ||
| 48 | hsa-let-7b-5p | NCAPG2 | 0.02 | 0.98368 | -0.38 | 0.49946 | miRNAWalker2 validate | -0.23 | 0.04548 | ||
| 49 | hsa-let-7b-5p | NFATC3 | 0.02 | 0.98368 | 0.01 | 0.98354 | miRNAWalker2 validate | -0.14 | 0.04252 | ||
| 50 | hsa-let-7b-5p | NLE1 | 0.02 | 0.98368 | -0.05 | 0.91455 | miRNAWalker2 validate | -0.23 | 1.0E-5 | ||
| 51 | hsa-let-7b-5p | NUP155 | 0.02 | 0.98368 | -0.06 | 0.92061 | miRNAWalker2 validate | -0.37 | 0.00031 | ||
| 52 | hsa-let-7b-5p | PIGU | 0.02 | 0.98368 | -0.23 | 0.72389 | miRNAWalker2 validate | -0.22 | 0.00402 | ||
| 53 | hsa-let-7b-5p | POP1 | 0.02 | 0.98368 | 0.01 | 0.98549 | miRNAWalker2 validate | -0.16 | 0.03985 | ||
| 54 | hsa-let-7b-5p | PRIM1 | 0.02 | 0.98368 | -0.36 | 0.56869 | miRNAWalker2 validate | -0.56 | 4.0E-5 | ||
| 55 | hsa-let-7b-5p | PTTG1 | 0.02 | 0.98368 | -0.49 | 0.52313 | miRNAWalker2 validate | -0.46 | 0.00477 | ||
| 56 | hsa-let-7b-5p | PUS1 | 0.02 | 0.98368 | -0.13 | 0.83634 | miRNAWalker2 validate | -0.19 | 0.01751 | ||
| 57 | hsa-let-7b-5p | RPIA | 0.02 | 0.98368 | -0.24 | 0.71413 | miRNAWalker2 validate | -0.2 | 0.03095 | ||
| 58 | hsa-let-7b-5p | RRM2 | 0.02 | 0.98368 | -0.59 | 0.43494 | miRNAWalker2 validate; miRNATAP | -0.61 | 0.00957 | ||
| 59 | hsa-let-7b-5p | SCAMP3 | 0.02 | 0.98368 | -0.36 | 0.64555 | miRNAWalker2 validate | -0.22 | 0.03353 | ||
| 60 | hsa-let-7b-5p | SCML2 | 0.02 | 0.98368 | -0.16 | 0.78606 | miRNAWalker2 validate | -0.31 | 0.00111 | ||
| 61 | hsa-let-7b-5p | SLC1A4 | 0.02 | 0.98368 | -0.3 | 0.63058 | miRNAWalker2 validate | -0.27 | 0.01083 | ||
| 62 | hsa-let-7b-5p | STIP1 | 0.02 | 0.98368 | -0.29 | 0.70874 | miRNAWalker2 validate | -0.37 | 0.00677 | ||
| 63 | hsa-let-7b-5p | TAB2 | 0.02 | 0.98368 | 0.2 | 0.76431 | miRNAWalker2 validate | -0.28 | 0.0247 | ||
| 64 | hsa-let-7b-5p | TPD52L2 | 0.02 | 0.98368 | -0.26 | 0.72043 | miRNAWalker2 validate | -0.27 | 0.00029 | ||
| 65 | hsa-let-7b-5p | TYMS | 0.02 | 0.98368 | -0.69 | 0.35822 | miRNAWalker2 validate | -0.44 | 0.02928 | ||
| 66 | hsa-let-7b-5p | UBAP2L | 0.02 | 0.98368 | -0.26 | 0.6878 | miRNAWalker2 validate | -0.26 | 0.00118 | ||
| 67 | hsa-let-7b-5p | UCK2 | 0.02 | 0.98368 | -0.11 | 0.87036 | miRNAWalker2 validate | -0.31 | 0.00119 | ||
| 68 | hsa-let-7b-5p | USP54 | 0.02 | 0.98368 | -0.01 | 0.98913 | miRNAWalker2 validate | -0.45 | 0.00849 | ||
| 69 | hsa-let-7b-5p | WDR4 | 0.02 | 0.98368 | -0.09 | 0.86538 | miRNAWalker2 validate | -0.24 | 0.0013 | ||
| 70 | hsa-let-7b-5p | XPO5 | 0.02 | 0.98368 | -0.28 | 0.6557 | miRNAWalker2 validate | -0.35 | 0.0005 | ||
| 71 | hsa-let-7c-5p | CCNB2 | 0.17 | 0.86763 | -0.71 | 0.25562 | miRNAWalker2 validate | -1.26 | 0 | ||
| 72 | hsa-let-7c-5p | CCNF | 0.17 | 0.86763 | -0.14 | 0.78436 | miRNAWalker2 validate; MirTarget | -0.23 | 3.0E-5 | ||
| 73 | hsa-let-7c-5p | DNAJC6 | 0.17 | 0.86763 | -0.07 | 0.86401 | miRNAWalker2 validate | -0.17 | 0.01897 | ||
| 74 | hsa-let-7c-5p | FANCI | 0.17 | 0.86763 | -0.39 | 0.452 | miRNAWalker2 validate | -0.55 | 0.00012 | ||
| 75 | hsa-let-7c-5p | MYC | 0.17 | 0.86763 | 0.33 | 0.7029 | miRNAWalker2 validate; miRTarBase | -0.6 | 0.02772 | ||
| 76 | hsa-let-7c-5p | PTK2 | 0.17 | 0.86763 | -0.09 | 0.90224 | miRNAWalker2 validate | -0.35 | 0.0053 | ||
| 77 | hsa-let-7c-5p | TMEM165 | 0.17 | 0.86763 | -0.1 | 0.88176 | miRNAWalker2 validate | -0.28 | 0.00423 | ||
| 78 | hsa-let-7c-5p | TRIM71 | 0.17 | 0.86763 | -0.45 | 0.45563 | miRNAWalker2 validate; MirTarget | -0.85 | 2.0E-5 | ||
| 79 | hsa-let-7d-5p | LIN28B | 0.02 | 0.98157 | -0.74 | 0.20501 | miRNAWalker2 validate; MirTarget; miRNATAP | -1.17 | 0.00074 | ||
| 80 | hsa-let-7e-5p | AGMAT | -0.32 | 0.66172 | 0.17 | 0.81353 | miRNAWalker2 validate | -0.22 | 0.00354 | ||
| 81 | hsa-let-7e-5p | CCDC106 | -0.32 | 0.66172 | 0.01 | 0.98038 | miRNAWalker2 validate | -0.14 | 0.02525 | ||
| 82 | hsa-let-7e-5p | IRS2 | -0.32 | 0.66172 | 0.5 | 0.59412 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.21 | 0.03182 | ||
| 83 | hsa-let-7e-5p | MYCN | -0.32 | 0.66172 | -0.03 | 0.94965 | miRNAWalker2 validate; miRNATAP | -0.06 | 0.04093 | ||
| 84 | hsa-let-7e-5p | NUP155 | -0.32 | 0.66172 | -0.06 | 0.92061 | miRNAWalker2 validate | -0.19 | 0.00729 | ||
| 85 | hsa-let-7e-5p | TXNRD1 | -0.32 | 0.66172 | 0.01 | 0.9939 | miRNAWalker2 validate | -0.26 | 0.04355 | ||
| 86 | hsa-let-7e-5p | UBAP2L | -0.32 | 0.66172 | -0.26 | 0.6878 | miRNAWalker2 validate | -0.12 | 0.0301 | ||
| 87 | hsa-let-7f-5p | CCNB2 | 0.18 | 0.86357 | -0.71 | 0.25562 | miRNAWalker2 validate | -0.69 | 0.03654 | ||
| 88 | hsa-let-7f-5p | HMGA2 | 0.18 | 0.86357 | -0.01 | 0.9752 | miRNAWalker2 validate; MirTarget | -0.05 | 0.03568 | ||
| 89 | hsa-let-7f-5p | IGF2BP1 | 0.18 | 0.86357 | -0.26 | 0.60749 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.39 | 0.00325 | ||
| 90 | hsa-let-7f-5p | SNRPC | 0.18 | 0.86357 | -0.09 | 0.90563 | miRNAWalker2 validate | -0.31 | 0.00227 | ||
| 91 | hsa-let-7f-5p | ZNF280B | 0.18 | 0.86357 | -0.06 | 0.86397 | miRNAWalker2 validate | -0.14 | 0.01079 | ||
| 92 | hsa-miR-100-5p | ALG3 | 0.39 | 0.64595 | -0.18 | 0.82449 | miRNAWalker2 validate | -0.15 | 0.00731 | ||
| 93 | hsa-miR-100-5p | ATP1B3 | 0.39 | 0.64595 | -0.03 | 0.96135 | miRNAWalker2 validate | -0.15 | 0.00013 | ||
| 94 | hsa-miR-100-5p | DDX21 | 0.39 | 0.64595 | 0.06 | 0.93742 | miRNAWalker2 validate | -0.13 | 0.02652 | ||
| 95 | hsa-miR-100-5p | H2AFY | 0.39 | 0.64595 | -0.02 | 0.97274 | miRNAWalker2 validate | -0.21 | 0 | ||
| 96 | hsa-miR-100-5p | IARS | 0.39 | 0.64595 | -0.32 | 0.71092 | miRNAWalker2 validate | -0.31 | 0 | ||
| 97 | hsa-miR-100-5p | LIN28B | 0.39 | 0.64595 | -0.74 | 0.20501 | miRNAWalker2 validate | -0.44 | 0.03778 | ||
| 98 | hsa-miR-100-5p | PLK1 | 0.39 | 0.64595 | -0.36 | 0.48024 | miRNAWalker2 validate; miRTarBase | -0.15 | 0.00489 | 23842624; 23842624; 23842624 | In addition we found that upregulation of miR-100 could inhibit growth and increase apoptosis of HCC cells by downregulating polo-like kinase 1 plk1;In HCC tissues miR-100 expression was inversely correlated with the expression of plk1 protein r = -0.418; P = 0.029;Therefore downregulation of miR-100 was correlated with progressive pathological feature and poor prognosis in HCC patients and miR-100 could function as a tumor suppressor by targeting plk1 |
| 99 | hsa-miR-100-5p | RAD51C | 0.39 | 0.64595 | -0.21 | 0.73039 | miRNAWalker2 validate | -0.21 | 0.00015 | ||
| 100 | hsa-miR-100-5p | RCN2 | 0.39 | 0.64595 | -0.3 | 0.69009 | miRNAWalker2 validate | -0.15 | 0.03374 | ||
| 101 | hsa-miR-100-5p | RRM2 | 0.39 | 0.64595 | -0.59 | 0.43494 | miRNAWalker2 validate | -0.5 | 0.00018 | ||
| 102 | hsa-miR-100-5p | VPS45 | 0.39 | 0.64595 | -0.24 | 0.73838 | miRNAWalker2 validate | -0.13 | 0.0149 | ||
| 103 | hsa-miR-100-5p | WDR4 | 0.39 | 0.64595 | -0.09 | 0.86538 | miRNAWalker2 validate | -0.16 | 0.00021 | ||
| 104 | hsa-miR-101-3p | BIRC5 | 0.46 | 0.57468 | -0.52 | 0.35824 | miRNAWalker2 validate | -0.27 | 0.00406 | ||
| 105 | hsa-miR-101-3p | CDC7 | 0.46 | 0.57468 | -0.6 | 0.27879 | miRNAWalker2 validate | -0.24 | 0.04603 | ||
| 106 | hsa-miR-101-3p | FZD6 | 0.46 | 0.57468 | -0.07 | 0.91012 | miRNAWalker2 validate; MirTarget | -0.52 | 0.00135 | ||
| 107 | hsa-miR-101-3p | KIF2C | 0.46 | 0.57468 | -0.39 | 0.48428 | miRNAWalker2 validate | -0.2 | 0.02263 | ||
| 108 | hsa-miR-101-3p | LMNB1 | 0.46 | 0.57468 | -0.41 | 0.49217 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.29 | 0.00328 | ||
| 109 | hsa-miR-101-3p | MSH2 | 0.46 | 0.57468 | -0.32 | 0.61258 | miRNAWalker2 validate | -0.32 | 0.00495 | ||
| 110 | hsa-miR-101-3p | PLAG1 | 0.46 | 0.57468 | -0.23 | 0.6194 | miRNAWalker2 validate | -0.34 | 0.00254 | ||
| 111 | hsa-miR-101-3p | STMN1 | 0.46 | 0.57468 | -0.17 | 0.74564 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.11 | 0.00767 | ||
| 112 | hsa-miR-101-3p | SUB1 | 0.46 | 0.57468 | -0.07 | 0.93647 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.18 | 0.001 | ||
| 113 | hsa-miR-103a-3p | GPR180 | -0.14 | 0.87827 | -0.05 | 0.89036 | miRNAWalker2 validate | -0.23 | 0.00595 | ||
| 114 | hsa-miR-103a-3p | PHKA1 | -0.14 | 0.87827 | -0.04 | 0.92803 | miRNAWalker2 validate | -0.21 | 0.02293 | ||
| 115 | hsa-miR-103a-3p | RAD51 | -0.14 | 0.87827 | -0.09 | 0.80562 | miRNAWalker2 validate | -0.09 | 0.03199 | ||
| 116 | hsa-miR-106b-5p | DAP3 | -0.05 | 0.95955 | -0.19 | 0.83402 | miRNAWalker2 validate | -0.2 | 0.03816 | ||
| 117 | hsa-miR-106b-5p | ZNF706 | -0.05 | 0.95955 | -0.07 | 0.9149 | miRNAWalker2 validate | -0.27 | 0.01262 | ||
| 118 | hsa-miR-107 | MYB | -0.11 | 0.89701 | -0.04 | 0.90582 | miRNAWalker2 validate; miRTarBase; PITA; miRanda; miRNATAP | -0.08 | 0.0233 | ||
| 119 | hsa-miR-10a-5p | ABCG2 | -0.05 | 0.95447 | 0.42 | 0.53864 | miRNAWalker2 validate | -0.4 | 0.04299 | ||
| 120 | hsa-miR-10a-5p | AGER | -0.05 | 0.95447 | -0 | 0.99946 | miRNAWalker2 validate | -0.09 | 0.00502 | ||
| 121 | hsa-miR-10a-5p | CHRNA5 | -0.05 | 0.95447 | -0.03 | 0.93738 | miRNAWalker2 validate | -0.11 | 0.01081 | ||
| 122 | hsa-miR-10a-5p | E2F1 | -0.05 | 0.95447 | -0.08 | 0.88089 | miRNAWalker2 validate | -0.12 | 0.00466 | ||
| 123 | hsa-miR-10a-5p | EXTL3 | -0.05 | 0.95447 | -0.1 | 0.84831 | miRNAWalker2 validate; mirMAP | -0.08 | 0.04589 | ||
| 124 | hsa-miR-10a-5p | FASN | -0.05 | 0.95447 | -0.01 | 0.97924 | miRNAWalker2 validate | -0.09 | 0.04934 | ||
| 125 | hsa-miR-10a-5p | PTPRT | -0.05 | 0.95447 | 0.06 | 0.89967 | miRNAWalker2 validate; mirMAP | -0.13 | 0.01138 | ||
| 126 | hsa-miR-10a-5p | SLC41A3 | -0.05 | 0.95447 | -0.09 | 0.8902 | miRNAWalker2 validate | -0.13 | 0.02943 | ||
| 127 | hsa-miR-10b-5p | ADRA1B | -0.35 | 0.69442 | 0.15 | 0.70639 | miRNAWalker2 validate | -0.07 | 0.00229 | ||
| 128 | hsa-miR-10b-5p | APLNR | -0.35 | 0.69442 | 0 | 0.99494 | miRNAWalker2 validate | -0.16 | 0.02271 | ||
| 129 | hsa-miR-10b-5p | FAM13A | -0.35 | 0.69442 | 0.39 | 0.5079 | miRNAWalker2 validate | -0.16 | 0.00417 | ||
| 130 | hsa-miR-10b-5p | NR4A3 | -0.35 | 0.69442 | 0.28 | 0.51115 | miRNAWalker2 validate | -0.07 | 0.01705 | ||
| 131 | hsa-miR-10b-5p | NRP2 | -0.35 | 0.69442 | 0.04 | 0.92545 | miRNAWalker2 validate; miRNATAP | -0.05 | 0.00185 | ||
| 132 | hsa-miR-10b-5p | PTPRT | -0.35 | 0.69442 | 0.06 | 0.89967 | miRNAWalker2 validate; mirMAP | -0.07 | 0.02488 | ||
| 133 | hsa-miR-10b-5p | SLC2A3 | -0.35 | 0.69442 | 0.34 | 0.57207 | miRNAWalker2 validate | -0.22 | 0.01081 | ||
| 134 | hsa-miR-122-5p | AACS | 0.33 | 0.76417 | -0.53 | 0.36212 | miRNAWalker2 validate; miRTarBase | -0.43 | 0.00162 | ||
| 135 | hsa-miR-122-5p | ALDOA | 0.33 | 0.76417 | -0.46 | 0.61124 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.67 | 4.0E-5 | ||
| 136 | hsa-miR-122-5p | ANK2 | 0.33 | 0.76417 | -0.12 | 0.78484 | miRNAWalker2 validate; miRTarBase | -0.31 | 0.00167 | ||
| 137 | hsa-miR-122-5p | AP3M2 | 0.33 | 0.76417 | -0.25 | 0.66584 | miRNAWalker2 validate; miRTarBase | -0.25 | 0.01463 | ||
| 138 | hsa-miR-122-5p | CD83 | 0.33 | 0.76417 | 0.11 | 0.85702 | miRNAWalker2 validate | -0.25 | 0.03136 | ||
| 139 | hsa-miR-122-5p | CDK4 | 0.33 | 0.76417 | -0.31 | 0.69218 | miRNAWalker2 validate | -0.34 | 0.00305 | ||
| 140 | hsa-miR-122-5p | CENPF | 0.33 | 0.76417 | -0.35 | 0.45736 | miRNAWalker2 validate | -0.37 | 0.00454 | ||
| 141 | hsa-miR-122-5p | CLSPN | 0.33 | 0.76417 | 0 | 0.98724 | miRNAWalker2 validate | -0.12 | 9.0E-5 | ||
| 142 | hsa-miR-122-5p | DSTYK | 0.33 | 0.76417 | -0.12 | 0.81931 | miRNAWalker2 validate; miRTarBase | -0.14 | 0.04097 | ||
| 143 | hsa-miR-122-5p | FAM118A | 0.33 | 0.76417 | -0.26 | 0.61976 | miRNAWalker2 validate | -0.38 | 0.00957 | ||
| 144 | hsa-miR-122-5p | GALNT10 | 0.33 | 0.76417 | 0.02 | 0.97535 | miRNAWalker2 validate; miRTarBase | -0.31 | 0 | ||
| 145 | hsa-miR-122-5p | HMOX1 | 0.33 | 0.76417 | 0.19 | 0.81377 | miRNAWalker2 validate | -0.6 | 0.00059 | ||
| 146 | hsa-miR-122-5p | LCA5 | 0.33 | 0.76417 | -0.03 | 0.93386 | miRNAWalker2 validate | -0.1 | 0.00199 | ||
| 147 | hsa-miR-122-5p | LMNB2 | 0.33 | 0.76417 | -0.33 | 0.56202 | miRNAWalker2 validate | -0.39 | 1.0E-5 | ||
| 148 | hsa-miR-122-5p | NMNAT2 | 0.33 | 0.76417 | 0 | 0.99068 | miRNAWalker2 validate | -0.07 | 0.04107 | ||
| 149 | hsa-miR-122-5p | PHKA1 | 0.33 | 0.76417 | -0.04 | 0.92803 | miRNAWalker2 validate | -0.33 | 0.00061 | ||
| 150 | hsa-miR-122-5p | PHPT1 | 0.33 | 0.76417 | -0.32 | 0.69151 | miRNAWalker2 validate | -0.31 | 0.00132 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CELL CYCLE PROCESS | 238 | 1081 | 3.184e-23 | 1.482e-19 |
| 2 | CELL CYCLE | 271 | 1316 | 6.905e-22 | 1.606e-18 |
| 3 | MITOTIC CELL CYCLE | 181 | 766 | 3.095e-21 | 4.8e-18 |
| 4 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 308 | 1672 | 3.253e-17 | 3.784e-14 |
| 5 | CELL DIVISION | 118 | 460 | 5.236e-17 | 4.06e-14 |
| 6 | REGULATION OF CELL DIFFERENTIATION | 281 | 1492 | 5.1e-17 | 4.06e-14 |
| 7 | POSITIVE REGULATION OF CELL CYCLE | 94 | 332 | 1.032e-16 | 6.858e-14 |
| 8 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 191 | 917 | 4.116e-16 | 2.394e-13 |
| 9 | REGULATION OF CELL CYCLE | 195 | 949 | 8.426e-16 | 4.356e-13 |
| 10 | RESPONSE TO ENDOGENOUS STIMULUS | 270 | 1450 | 1.002e-15 | 4.665e-13 |
| 11 | SMALL MOLECULE METABOLIC PROCESS | 314 | 1767 | 2.657e-15 | 1.124e-12 |
| 12 | REGULATION OF KINASE ACTIVITY | 166 | 776 | 3.299e-15 | 1.279e-12 |
| 13 | MITOTIC NUCLEAR DIVISION | 96 | 361 | 3.976e-15 | 1.321e-12 |
| 14 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 292 | 1618 | 3.959e-15 | 1.321e-12 |
| 15 | SISTER CHROMATID SEGREGATION | 59 | 176 | 1.73e-14 | 5.365e-12 |
| 16 | CELL PROLIFERATION | 147 | 672 | 2.05e-14 | 5.612e-12 |
| 17 | REGULATION OF CELL DEVELOPMENT | 173 | 836 | 2.049e-14 | 5.612e-12 |
| 18 | ORGANIC ACID METABOLIC PROCESS | 190 | 953 | 4.289e-14 | 1.109e-11 |
| 19 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 238 | 1275 | 4.708e-14 | 1.153e-11 |
| 20 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 311 | 1791 | 8.302e-14 | 1.84e-11 |
| 21 | POSITIVE REGULATION OF CELL CYCLE PROCESS | 72 | 247 | 8.175e-14 | 1.84e-11 |
| 22 | REGULATION OF TRANSFERASE ACTIVITY | 187 | 946 | 1.579e-13 | 3.34e-11 |
| 23 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 251 | 1381 | 1.974e-13 | 3.675e-11 |
| 24 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 267 | 1492 | 1.946e-13 | 3.675e-11 |
| 25 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 198 | 1021 | 1.967e-13 | 3.675e-11 |
| 26 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 162 | 788 | 2.58e-13 | 4.437e-11 |
| 27 | CIRCULATORY SYSTEM DEVELOPMENT | 162 | 788 | 2.58e-13 | 4.437e-11 |
| 28 | REGULATION OF CELL PROLIFERATION | 267 | 1496 | 2.67e-13 | 4.437e-11 |
| 29 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 252 | 1395 | 3.525e-13 | 5.656e-11 |
| 30 | RESPONSE TO HORMONE | 177 | 893 | 5.764e-13 | 8.94e-11 |
| 31 | NEUROGENESIS | 252 | 1402 | 6.202e-13 | 9.309e-11 |
| 32 | EMBRYO DEVELOPMENT | 176 | 894 | 1.241e-12 | 1.732e-10 |
| 33 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 116 | 513 | 1.265e-12 | 1.732e-10 |
| 34 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 213 | 1142 | 1.254e-12 | 1.732e-10 |
| 35 | ORGANELLE FISSION | 113 | 496 | 1.486e-12 | 1.975e-10 |
| 36 | CHROMOSOME SEGREGATION | 74 | 272 | 1.681e-12 | 2.115e-10 |
| 37 | RESPONSE TO DRUG | 102 | 431 | 1.648e-12 | 2.115e-10 |
| 38 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 185 | 957 | 1.73e-12 | 2.119e-10 |
| 39 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 172 | 872 | 1.919e-12 | 2.289e-10 |
| 40 | VASCULATURE DEVELOPMENT | 108 | 469 | 2.299e-12 | 2.674e-10 |
| 41 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 266 | 1518 | 2.477e-12 | 2.811e-10 |
| 42 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 196 | 1036 | 2.838e-12 | 3.071e-10 |
| 43 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 196 | 1036 | 2.838e-12 | 3.071e-10 |
| 44 | HEAD DEVELOPMENT | 146 | 709 | 3.604e-12 | 3.812e-10 |
| 45 | RESPONSE TO NITROGEN COMPOUND | 169 | 859 | 3.779e-12 | 3.907e-10 |
| 46 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 311 | 1848 | 4.514e-12 | 4.566e-10 |
| 47 | REGULATION OF MITOTIC CELL CYCLE | 107 | 468 | 4.66e-12 | 4.614e-10 |
| 48 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 155 | 771 | 5.486e-12 | 5.318e-10 |
| 49 | POSITIVE REGULATION OF KINASE ACTIVITY | 109 | 482 | 6.068e-12 | 5.763e-10 |
| 50 | BLOOD VESSEL MORPHOGENESIS | 89 | 364 | 6.686e-12 | 6.194e-10 |
| 51 | REGULATION OF PROTEIN MODIFICATION PROCESS | 291 | 1710 | 6.789e-12 | 6.194e-10 |
| 52 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 130 | 616 | 9.916e-12 | 8.873e-10 |
| 53 | REGULATION OF NEURON DIFFERENTIATION | 120 | 554 | 1.079e-11 | 9.47e-10 |
| 54 | TISSUE DEVELOPMENT | 263 | 1518 | 1.178e-11 | 1.015e-09 |
| 55 | NUCLEAR CHROMOSOME SEGREGATION | 64 | 228 | 1.216e-11 | 1.029e-09 |
| 56 | REGULATION OF CELL CYCLE PROCESS | 120 | 558 | 1.768e-11 | 1.469e-09 |
| 57 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 74 | 285 | 1.957e-11 | 1.598e-09 |
| 58 | SINGLE ORGANISM CATABOLIC PROCESS | 181 | 957 | 2.109e-11 | 1.692e-09 |
| 59 | SISTER CHROMATID COHESION | 40 | 111 | 2.175e-11 | 1.716e-09 |
| 60 | CELL DEVELOPMENT | 248 | 1426 | 3.313e-11 | 2.569e-09 |
| 61 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 184 | 983 | 3.524e-11 | 2.688e-09 |
| 62 | RESPONSE TO EXTRACELLULAR STIMULUS | 100 | 441 | 3.872e-11 | 2.906e-09 |
| 63 | REGULATION OF CELL DEATH | 254 | 1472 | 4.249e-11 | 3.138e-09 |
| 64 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 205 | 1135 | 7.051e-11 | 5.126e-09 |
| 65 | RESPONSE TO ALCOHOL | 86 | 362 | 7.501e-11 | 5.369e-09 |
| 66 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 103 | 465 | 8.331e-11 | 5.701e-09 |
| 67 | TUBE DEVELOPMENT | 117 | 552 | 8.289e-11 | 5.701e-09 |
| 68 | REGULATION OF CYTOSKELETON ORGANIZATION | 109 | 502 | 8.304e-11 | 5.701e-09 |
| 69 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 155 | 799 | 9.22e-11 | 6.217e-09 |
| 70 | LOCOMOTION | 201 | 1114 | 1.204e-10 | 8.001e-09 |
| 71 | NEGATIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 137 | 684 | 1.278e-10 | 8.374e-09 |
| 72 | NEGATIVE REGULATION OF GENE EXPRESSION | 254 | 1493 | 1.833e-10 | 1.185e-08 |
| 73 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 297 | 1805 | 1.962e-10 | 1.251e-08 |
| 74 | POSITIVE REGULATION OF MITOTIC CELL CYCLE | 41 | 123 | 2.031e-10 | 1.277e-08 |
| 75 | REGULATION OF ORGANELLE ORGANIZATION | 209 | 1178 | 2.325e-10 | 1.442e-08 |
| 76 | INTRACELLULAR SIGNAL TRANSDUCTION | 264 | 1572 | 2.958e-10 | 1.811e-08 |
| 77 | CELLULAR RESPONSE TO STRESS | 263 | 1565 | 2.999e-10 | 1.812e-08 |
| 78 | CELL MOTILITY | 158 | 835 | 4.098e-10 | 2.414e-08 |
| 79 | LOCALIZATION OF CELL | 158 | 835 | 4.098e-10 | 2.414e-08 |
| 80 | REGULATION OF GROWTH | 127 | 633 | 5.503e-10 | 3.201e-08 |
| 81 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 274 | 1656 | 6.22e-10 | 3.573e-08 |
| 82 | RESPONSE TO LIPID | 165 | 888 | 6.783e-10 | 3.849e-08 |
| 83 | ORGAN MORPHOGENESIS | 158 | 841 | 7.017e-10 | 3.934e-08 |
| 84 | LIPID METABOLIC PROCESS | 204 | 1158 | 7.217e-10 | 3.998e-08 |
| 85 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 101 | 470 | 7.487e-10 | 4.099e-08 |
| 86 | NEGATIVE REGULATION OF CELL CYCLE | 95 | 433 | 7.691e-10 | 4.161e-08 |
| 87 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 317 | 1977 | 8.597e-10 | 4.546e-08 |
| 88 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 155 | 823 | 8.557e-10 | 4.546e-08 |
| 89 | TELENCEPHALON DEVELOPMENT | 60 | 228 | 8.756e-10 | 4.578e-08 |
| 90 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 144 | 750 | 9.126e-10 | 4.666e-08 |
| 91 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 182 | 1008 | 9.122e-10 | 4.666e-08 |
| 92 | POSITIVE REGULATION OF CELL PROLIFERATION | 153 | 814 | 1.275e-09 | 6.451e-08 |
| 93 | EMBRYONIC MORPHOGENESIS | 111 | 539 | 1.472e-09 | 7.362e-08 |
| 94 | OXIDATION REDUCTION PROCESS | 165 | 898 | 1.587e-09 | 7.858e-08 |
| 95 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 90 | 408 | 1.615e-09 | 7.912e-08 |
| 96 | NEURON PROJECTION MORPHOGENESIS | 89 | 402 | 1.644e-09 | 7.97e-08 |
| 97 | REGULATION OF MICROTUBULE BASED PROCESS | 62 | 243 | 1.744e-09 | 8.366e-08 |
| 98 | MICROTUBULE CYTOSKELETON ORGANIZATION | 80 | 348 | 1.779e-09 | 8.448e-08 |
| 99 | RESPONSE TO EXTERNAL STIMULUS | 294 | 1821 | 1.906e-09 | 8.96e-08 |
| 100 | POSITIVE REGULATION OF GENE EXPRESSION | 282 | 1733 | 1.93e-09 | 8.982e-08 |
| 101 | CELL CYCLE CHECKPOINT | 53 | 194 | 1.996e-09 | 9.196e-08 |
| 102 | REGULATION OF TRANSPORT | 291 | 1804 | 2.561e-09 | 1.168e-07 |
| 103 | NEGATIVE REGULATION OF CELL DEATH | 160 | 872 | 3.141e-09 | 1.419e-07 |
| 104 | POSITIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 45 | 154 | 3.323e-09 | 1.455e-07 |
| 105 | CELLULAR COMPONENT MORPHOGENESIS | 164 | 900 | 3.302e-09 | 1.455e-07 |
| 106 | ANGIOGENESIS | 70 | 293 | 3.347e-09 | 1.455e-07 |
| 107 | POSITIVE REGULATION OF CELL CYCLE PHASE TRANSITION | 27 | 68 | 3.295e-09 | 1.455e-07 |
| 108 | TAXIS | 98 | 464 | 3.474e-09 | 1.497e-07 |
| 109 | RESPIRATORY SYSTEM DEVELOPMENT | 53 | 197 | 3.595e-09 | 1.521e-07 |
| 110 | TUBE MORPHOGENESIS | 75 | 323 | 3.576e-09 | 1.521e-07 |
| 111 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 112 | 554 | 3.73e-09 | 1.55e-07 |
| 112 | CHEMICAL HOMEOSTASIS | 160 | 874 | 3.709e-09 | 1.55e-07 |
| 113 | HOMEOSTATIC PROCESS | 225 | 1337 | 5.877e-09 | 2.42e-07 |
| 114 | EMBRYONIC ORGAN DEVELOPMENT | 88 | 406 | 6.007e-09 | 2.452e-07 |
| 115 | REGULATION OF CELL CYCLE PHASE TRANSITION | 74 | 321 | 6.317e-09 | 2.556e-07 |
| 116 | ANION TRANSMEMBRANE TRANSPORT | 62 | 251 | 6.745e-09 | 2.705e-07 |
| 117 | RESPONSE TO STEROID HORMONE | 102 | 497 | 8.233e-09 | 3.274e-07 |
| 118 | PALLIUM DEVELOPMENT | 44 | 153 | 8.472e-09 | 3.341e-07 |
| 119 | RESPONSE TO ABIOTIC STIMULUS | 180 | 1024 | 9.133e-09 | 3.571e-07 |
| 120 | CELL MORPHOGENESIS INVOLVED IN NEURON DIFFERENTIATION | 81 | 368 | 1.165e-08 | 4.517e-07 |
| 121 | REGULATION OF CELL DIVISION | 65 | 272 | 1.192e-08 | 4.583e-07 |
| 122 | ANION TRANSPORT | 103 | 507 | 1.215e-08 | 4.632e-07 |
| 123 | CELL CYCLE PHASE TRANSITION | 62 | 255 | 1.285e-08 | 4.862e-07 |
| 124 | FOREBRAIN DEVELOPMENT | 79 | 357 | 1.374e-08 | 5.155e-07 |
| 125 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 147 | 801 | 1.385e-08 | 5.155e-07 |
| 126 | MICROTUBULE BASED PROCESS | 105 | 522 | 1.515e-08 | 5.594e-07 |
| 127 | SMALL MOLECULE CATABOLIC PROCESS | 74 | 328 | 1.671e-08 | 6.122e-07 |
| 128 | MITOTIC SISTER CHROMATID SEGREGATION | 31 | 91 | 1.782e-08 | 6.478e-07 |
| 129 | ION HOMEOSTASIS | 113 | 576 | 1.821e-08 | 6.567e-07 |
| 130 | NEGATIVE REGULATION OF PHOSPHORYLATION | 89 | 422 | 1.948e-08 | 6.974e-07 |
| 131 | REGULATION OF CELL PROJECTION ORGANIZATION | 110 | 558 | 2.151e-08 | 7.642e-07 |
| 132 | REGULATION OF CELL MORPHOGENESIS | 109 | 552 | 2.273e-08 | 8.011e-07 |
| 133 | CHROMOSOME ORGANIZATION | 176 | 1009 | 2.376e-08 | 8.314e-07 |
| 134 | TISSUE MORPHOGENESIS | 106 | 533 | 2.413e-08 | 8.38e-07 |
| 135 | REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 75 | 337 | 2.469e-08 | 8.51e-07 |
| 136 | RESPONSE TO NUTRIENT | 50 | 191 | 2.603e-08 | 8.906e-07 |
| 137 | BIOLOGICAL ADHESION | 179 | 1032 | 2.72e-08 | 9.24e-07 |
| 138 | CELLULAR AMINO ACID METABOLIC PROCESS | 74 | 332 | 2.857e-08 | 9.633e-07 |
| 139 | POSITIVE REGULATION OF LOCOMOTION | 88 | 420 | 3.216e-08 | 1.069e-06 |
| 140 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 246 | 1517 | 3.218e-08 | 1.069e-06 |
| 141 | NEURON DIFFERENTIATION | 156 | 874 | 3.362e-08 | 1.109e-06 |
| 142 | NEGATIVE REGULATION OF CELL DEVELOPMENT | 69 | 303 | 3.415e-08 | 1.119e-06 |
| 143 | REPRODUCTIVE SYSTEM DEVELOPMENT | 86 | 408 | 3.458e-08 | 1.125e-06 |
| 144 | RESPONSE TO OXYGEN LEVELS | 70 | 311 | 4.463e-08 | 1.442e-06 |
| 145 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 174 | 1004 | 4.556e-08 | 1.462e-06 |
| 146 | REGULATION OF MAPK CASCADE | 124 | 660 | 5e-08 | 1.594e-06 |
| 147 | REGULATION OF LIPID METABOLIC PROCESS | 65 | 282 | 5.214e-08 | 1.65e-06 |
| 148 | NEGATIVE REGULATION OF ORGANELLE ORGANIZATION | 82 | 387 | 5.646e-08 | 1.775e-06 |
| 149 | REGULATION OF DEVELOPMENTAL GROWTH | 66 | 289 | 6.004e-08 | 1.839e-06 |
| 150 | MORPHOGENESIS OF AN EPITHELIUM | 84 | 400 | 5.92e-08 | 1.839e-06 |
| 151 | POSITIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 66 | 289 | 6.004e-08 | 1.839e-06 |
| 152 | REGULATION OF BINDING | 65 | 283 | 6.008e-08 | 1.839e-06 |
| 153 | MONOCARBOXYLIC ACID METABOLIC PROCESS | 100 | 503 | 6.106e-08 | 1.857e-06 |
| 154 | REGULATION OF ANATOMICAL STRUCTURE SIZE | 95 | 472 | 7.107e-08 | 2.147e-06 |
| 155 | DEVELOPMENTAL GROWTH | 73 | 333 | 7.218e-08 | 2.167e-06 |
| 156 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 280 | 1784 | 7.515e-08 | 2.242e-06 |
| 157 | ION TRANSPORT | 209 | 1262 | 7.667e-08 | 2.272e-06 |
| 158 | NEURON PROJECTION DEVELOPMENT | 106 | 545 | 7.943e-08 | 2.339e-06 |
| 159 | CYTOSKELETON ORGANIZATION | 149 | 838 | 8.821e-08 | 2.581e-06 |
| 160 | REGULATION OF CELL GROWTH | 82 | 391 | 9.014e-08 | 2.621e-06 |
| 161 | SKELETAL SYSTEM DEVELOPMENT | 92 | 455 | 9.141e-08 | 2.642e-06 |
| 162 | REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 31 | 97 | 9.549e-08 | 2.743e-06 |
| 163 | NEGATIVE REGULATION OF MITOTIC CELL CYCLE | 50 | 199 | 1.068e-07 | 3.049e-06 |
| 164 | NEURON PROJECTION GUIDANCE | 51 | 205 | 1.123e-07 | 3.186e-06 |
| 165 | ALPHA AMINO ACID METABOLIC PROCESS | 55 | 229 | 1.293e-07 | 3.625e-06 |
| 166 | CELLULAR CHEMICAL HOMEOSTASIS | 109 | 570 | 1.288e-07 | 3.625e-06 |
| 167 | PROTEIN PHOSPHORYLATION | 163 | 944 | 1.577e-07 | 4.394e-06 |
| 168 | DNA INTEGRITY CHECKPOINT | 40 | 146 | 1.687e-07 | 4.672e-06 |
| 169 | NEGATIVE REGULATION OF CELL CYCLE PROCESS | 52 | 214 | 1.901e-07 | 5.233e-06 |
| 170 | CEREBRAL CORTEX DEVELOPMENT | 32 | 105 | 2.053e-07 | 5.619e-06 |
| 171 | NEGATIVE REGULATION OF CYTOSKELETON ORGANIZATION | 53 | 221 | 2.286e-07 | 6.22e-06 |
| 172 | NEGATIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 133 | 740 | 2.365e-07 | 6.399e-06 |
| 173 | CELL CYCLE G1 S PHASE TRANSITION | 33 | 111 | 2.545e-07 | 6.807e-06 |
| 174 | G1 S TRANSITION OF MITOTIC CELL CYCLE | 33 | 111 | 2.545e-07 | 6.807e-06 |
| 175 | POSITIVE REGULATION OF CELL DIVISION | 37 | 132 | 2.585e-07 | 6.872e-06 |
| 176 | RESPONSE TO ACID CHEMICAL | 69 | 319 | 2.784e-07 | 7.319e-06 |
| 177 | REGULATION OF MAP KINASE ACTIVITY | 69 | 319 | 2.784e-07 | 7.319e-06 |
| 178 | CELLULAR RESPONSE TO HORMONE STIMULUS | 105 | 552 | 2.859e-07 | 7.473e-06 |
| 179 | PLACENTA DEVELOPMENT | 38 | 138 | 2.953e-07 | 7.675e-06 |
| 180 | REGULATION OF CELL CYCLE G2 M PHASE TRANSITION | 22 | 59 | 3.431e-07 | 8.87e-06 |
| 181 | CELLULAR LIPID METABOLIC PROCESS | 157 | 913 | 3.487e-07 | 8.964e-06 |
| 182 | MITOTIC CELL CYCLE CHECKPOINT | 38 | 139 | 3.61e-07 | 9.229e-06 |
| 183 | GROWTH | 83 | 410 | 3.655e-07 | 9.294e-06 |
| 184 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 112 | 602 | 3.714e-07 | 9.391e-06 |
| 185 | RESPONSE TO PEPTIDE | 82 | 404 | 3.831e-07 | 9.636e-06 |
| 186 | EMBRYONIC ORGAN MORPHOGENESIS | 62 | 279 | 4.139e-07 | 1.035e-05 |
| 187 | REGULATION OF CELLULAR LOCALIZATION | 207 | 1277 | 4.48e-07 | 1.115e-05 |
| 188 | DNA REPLICATION | 50 | 208 | 4.603e-07 | 1.139e-05 |
| 189 | MUSCLE STRUCTURE DEVELOPMENT | 86 | 432 | 4.78e-07 | 1.177e-05 |
| 190 | IN UTERO EMBRYONIC DEVELOPMENT | 67 | 311 | 4.807e-07 | 1.177e-05 |
| 191 | ORGANIC ANION TRANSPORT | 79 | 387 | 4.896e-07 | 1.193e-05 |
| 192 | UROGENITAL SYSTEM DEVELOPMENT | 65 | 299 | 5.055e-07 | 1.225e-05 |
| 193 | TRANSMEMBRANE TRANSPORT | 182 | 1098 | 5.26e-07 | 1.268e-05 |
| 194 | RESPONSE TO INORGANIC SUBSTANCE | 93 | 479 | 5.343e-07 | 1.282e-05 |
| 195 | MITOTIC DNA INTEGRITY CHECKPOINT | 30 | 100 | 7.101e-07 | 1.694e-05 |
| 196 | CELLULAR RESPONSE TO OXYGEN LEVELS | 38 | 143 | 7.857e-07 | 1.865e-05 |
| 197 | REGULATION OF EXTENT OF CELL GROWTH | 30 | 101 | 8.981e-07 | 2.121e-05 |
| 198 | POSITIVE REGULATION OF VASCULATURE DEVELOPMENT | 36 | 133 | 9.415e-07 | 2.208e-05 |
| 199 | CELLULAR RESPONSE TO NITROGEN COMPOUND | 96 | 505 | 9.443e-07 | 2.208e-05 |
| 200 | POSITIVE REGULATION OF MAP KINASE ACTIVITY | 49 | 207 | 9.681e-07 | 2.252e-05 |
| 201 | ION TRANSMEMBRANE TRANSPORT | 142 | 822 | 9.84e-07 | 2.278e-05 |
| 202 | DRUG TRANSPORT | 13 | 25 | 9.946e-07 | 2.291e-05 |
| 203 | CELLULAR HOMEOSTASIS | 121 | 676 | 1.029e-06 | 2.358e-05 |
| 204 | MORPHOGENESIS OF A BRANCHING STRUCTURE | 42 | 167 | 1.052e-06 | 2.4e-05 |
| 205 | GLAND DEVELOPMENT | 79 | 395 | 1.143e-06 | 2.595e-05 |
| 206 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 111 | 609 | 1.181e-06 | 2.667e-05 |
| 207 | POSITIVE REGULATION OF CELL COMMUNICATION | 239 | 1532 | 1.207e-06 | 2.712e-05 |
| 208 | NEGATIVE REGULATION OF CELL COMMUNICATION | 193 | 1192 | 1.215e-06 | 2.718e-05 |
| 209 | SINGLE ORGANISM BIOSYNTHETIC PROCESS | 213 | 1340 | 1.283e-06 | 2.856e-05 |
| 210 | ORGANIC HYDROXY COMPOUND METABOLIC PROCESS | 92 | 482 | 1.323e-06 | 2.93e-05 |
| 211 | NEURON DEVELOPMENT | 122 | 687 | 1.406e-06 | 3.101e-05 |
| 212 | ORGANIC ACID TRANSMEMBRANE TRANSPORT | 30 | 103 | 1.419e-06 | 3.114e-05 |
| 213 | AGING | 58 | 264 | 1.426e-06 | 3.116e-05 |
| 214 | REGULATION OF ENDOTHELIAL CELL MIGRATION | 32 | 114 | 1.572e-06 | 3.418e-05 |
| 215 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 122 | 689 | 1.635e-06 | 3.54e-05 |
| 216 | REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 38 | 147 | 1.643e-06 | 3.54e-05 |
| 217 | REGULATION OF HORMONE LEVELS | 91 | 478 | 1.689e-06 | 3.622e-05 |
| 218 | DRUG TRANSMEMBRANE TRANSPORT | 11 | 19 | 1.698e-06 | 3.625e-05 |
| 219 | CARBOXYLIC ACID CATABOLIC PROCESS | 48 | 205 | 1.732e-06 | 3.664e-05 |
| 220 | ORGANIC ACID CATABOLIC PROCESS | 48 | 205 | 1.732e-06 | 3.664e-05 |
| 221 | EPITHELIUM DEVELOPMENT | 158 | 945 | 1.812e-06 | 3.764e-05 |
| 222 | RESPONSE TO CORTICOSTEROID | 43 | 176 | 1.807e-06 | 3.764e-05 |
| 223 | CELL PROJECTION ORGANIZATION | 152 | 902 | 1.808e-06 | 3.764e-05 |
| 224 | VITAMIN METABOLIC PROCESS | 33 | 120 | 1.795e-06 | 3.764e-05 |
| 225 | BRANCHING MORPHOGENESIS OF AN EPITHELIAL TUBE | 35 | 131 | 1.841e-06 | 3.807e-05 |
| 226 | HEART DEVELOPMENT | 89 | 466 | 1.909e-06 | 3.929e-05 |
| 227 | POSITIVE REGULATION OF POTASSIUM ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 10 | 16 | 1.944e-06 | 3.984e-05 |
| 228 | REGULATION OF SYSTEM PROCESS | 95 | 507 | 2.057e-06 | 4.198e-05 |
| 229 | ORGANIC ACID TRANSPORT | 57 | 261 | 2.135e-06 | 4.338e-05 |
| 230 | SKELETAL SYSTEM MORPHOGENESIS | 47 | 201 | 2.3e-06 | 4.654e-05 |
| 231 | RESPONSE TO WOUNDING | 103 | 563 | 2.396e-06 | 4.825e-05 |
| 232 | REGULATION OF MICROTUBULE POLYMERIZATION OR DEPOLYMERIZATION | 43 | 178 | 2.483e-06 | 4.979e-05 |
| 233 | ARTERY DEVELOPMENT | 24 | 75 | 2.542e-06 | 5.076e-05 |
| 234 | PHOSPHORYLATION | 196 | 1228 | 2.653e-06 | 5.275e-05 |
| 235 | TRANSITION METAL ION HOMEOSTASIS | 30 | 106 | 2.737e-06 | 5.418e-05 |
| 236 | POSITIVE REGULATION OF MAPK CASCADE | 89 | 470 | 2.749e-06 | 5.421e-05 |
| 237 | CELLULAR RESPONSE TO LIPID | 87 | 457 | 2.84e-06 | 5.576e-05 |
| 238 | RESPONSE TO OXIDATIVE STRESS | 71 | 352 | 2.86e-06 | 5.591e-05 |
| 239 | CARBOXYLIC ACID BIOSYNTHETIC PROCESS | 58 | 270 | 3.027e-06 | 5.844e-05 |
| 240 | NEGATIVE REGULATION OF NEURON DIFFERENTIATION | 45 | 191 | 3.019e-06 | 5.844e-05 |
| 241 | ORGANIC ACID BIOSYNTHETIC PROCESS | 58 | 270 | 3.027e-06 | 5.844e-05 |
| 242 | REGULATION OF AXONOGENESIS | 41 | 168 | 3.202e-06 | 6.157e-05 |
| 243 | POSITIVE REGULATION OF CELL DEVELOPMENT | 89 | 472 | 3.29e-06 | 6.3e-05 |
| 244 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 34 | 129 | 3.572e-06 | 6.811e-05 |
| 245 | RESPONSE TO ESTRADIOL | 37 | 146 | 3.714e-06 | 6.996e-05 |
| 246 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 99 | 541 | 3.701e-06 | 6.996e-05 |
| 247 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 99 | 541 | 3.701e-06 | 6.996e-05 |
| 248 | APPENDAGE DEVELOPMENT | 41 | 169 | 3.756e-06 | 7.019e-05 |
| 249 | LIMB DEVELOPMENT | 41 | 169 | 3.756e-06 | 7.019e-05 |
| 250 | REGULATION OF PROTEIN LOCALIZATION | 157 | 950 | 3.871e-06 | 7.204e-05 |
| 251 | POSITIVE REGULATION OF ENDOTHELIAL CELL MIGRATION | 22 | 67 | 4.089e-06 | 7.556e-05 |
| 252 | RESPONSE TO INSULIN | 47 | 205 | 4.092e-06 | 7.556e-05 |
| 253 | REGULATION OF CELL CYCLE ARREST | 30 | 108 | 4.163e-06 | 7.656e-05 |
| 254 | KIDNEY MORPHOGENESIS | 25 | 82 | 4.242e-06 | 7.771e-05 |
| 255 | ORGANONITROGEN COMPOUND CATABOLIC PROCESS | 69 | 343 | 4.322e-06 | 7.886e-05 |
| 256 | CELL PART MORPHOGENESIS | 112 | 633 | 4.503e-06 | 8.184e-05 |
| 257 | RESPONSE TO KETONE | 43 | 182 | 4.591e-06 | 8.311e-05 |
| 258 | REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 49 | 218 | 4.655e-06 | 8.396e-05 |
| 259 | SMALL MOLECULE BIOSYNTHETIC PROCESS | 84 | 443 | 4.918e-06 | 8.835e-05 |
| 260 | REGULATION OF DNA BINDING | 27 | 93 | 5.015e-06 | 8.974e-05 |
| 261 | REGULATION OF TRANSCRIPTION REGULATORY REGION DNA BINDING | 15 | 36 | 5.077e-06 | 9.051e-05 |
| 262 | RESPONSE TO HYDROGEN PEROXIDE | 30 | 109 | 5.107e-06 | 9.07e-05 |
| 263 | NEGATIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 56 | 262 | 5.216e-06 | 9.228e-05 |
| 264 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 212 | 1360 | 5.331e-06 | 9.397e-05 |
| 265 | RESPONSE TO FATTY ACID | 25 | 83 | 5.401e-06 | 9.484e-05 |
| 266 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 92 | 498 | 5.433e-06 | 9.503e-05 |
| 267 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 287 | 1929 | 5.662e-06 | 9.867e-05 |
| 268 | CELLULAR RESPONSE TO ALCOHOL | 31 | 115 | 5.729e-06 | 9.947e-05 |
| 269 | REGULATION OF EPITHELIAL CELL MIGRATION | 40 | 166 | 5.854e-06 | 0.0001013 |
| 270 | REGULATION OF BLOOD CIRCULATION | 61 | 295 | 6.194e-06 | 0.0001067 |
| 271 | REPRODUCTION | 203 | 1297 | 6.544e-06 | 0.0001124 |
| 272 | REGULATION OF HYDROLASE ACTIVITY | 207 | 1327 | 6.611e-06 | 0.0001131 |
| 273 | REGULATION OF HOMEOSTATIC PROCESS | 84 | 447 | 7.037e-06 | 0.0001195 |
| 274 | MITOTIC SPINDLE ORGANIZATION | 22 | 69 | 7.034e-06 | 0.0001195 |
| 275 | LIPID MODIFICATION | 47 | 210 | 8.141e-06 | 0.0001374 |
| 276 | NEURON RECOGNITION | 14 | 33 | 8.149e-06 | 0.0001374 |
| 277 | REGULATION OF CHEMOTAXIS | 42 | 180 | 8.209e-06 | 0.0001379 |
| 278 | DNA REPLICATION INITIATION | 13 | 29 | 8.245e-06 | 0.000138 |
| 279 | POSITIVE REGULATION OF CELL CYCLE G2 M PHASE TRANSITION | 10 | 18 | 8.501e-06 | 0.0001413 |
| 280 | AMINO ACID BETAINE METABOLIC PROCESS | 10 | 18 | 8.501e-06 | 0.0001413 |
| 281 | CELLULAR AMINO ACID CATABOLIC PROCESS | 30 | 112 | 9.242e-06 | 0.0001525 |
| 282 | EAR MORPHOGENESIS | 30 | 112 | 9.242e-06 | 0.0001525 |
| 283 | ORGANONITROGEN COMPOUND METABOLIC PROCESS | 268 | 1796 | 9.577e-06 | 0.0001575 |
| 284 | AMINO ACID IMPORT | 9 | 15 | 1.049e-05 | 0.0001718 |
| 285 | L AMINO ACID IMPORT | 8 | 12 | 1.115e-05 | 0.000182 |
| 286 | REGULATION OF DNA METABOLIC PROCESS | 67 | 340 | 1.205e-05 | 0.000196 |
| 287 | COFACTOR METABOLIC PROCESS | 66 | 334 | 1.271e-05 | 0.0002055 |
| 288 | INNER EAR MORPHOGENESIS | 26 | 92 | 1.272e-05 | 0.0002055 |
| 289 | POSITIVE REGULATION OF HYDROLASE ACTIVITY | 148 | 905 | 1.296e-05 | 0.0002086 |
| 290 | CATABOLIC PROCESS | 264 | 1773 | 1.32e-05 | 0.0002118 |
| 291 | POSITIVE REGULATION OF EPITHELIAL CELL MIGRATION | 28 | 103 | 1.342e-05 | 0.0002146 |
| 292 | AMEBOIDAL TYPE CELL MIGRATION | 37 | 154 | 1.382e-05 | 0.0002202 |
| 293 | REGULATION OF CELL SIZE | 40 | 172 | 1.449e-05 | 0.0002291 |
| 294 | MESENCHYME DEVELOPMENT | 43 | 190 | 1.45e-05 | 0.0002291 |
| 295 | REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 27 | 98 | 1.457e-05 | 0.0002291 |
| 296 | REGULATION OF VASCULATURE DEVELOPMENT | 50 | 233 | 1.454e-05 | 0.0002291 |
| 297 | WOUND HEALING | 86 | 470 | 1.581e-05 | 0.0002477 |
| 298 | REGULATION OF LIPID KINASE ACTIVITY | 17 | 48 | 1.625e-05 | 0.0002538 |
| 299 | DEVELOPMENTAL GROWTH INVOLVED IN MORPHOGENESIS | 28 | 104 | 1.634e-05 | 0.0002543 |
| 300 | IMMUNE SYSTEM DEVELOPMENT | 102 | 582 | 1.79e-05 | 0.000277 |
| 301 | NEGATIVE REGULATION OF CELL SUBSTRATE ADHESION | 18 | 53 | 1.792e-05 | 0.000277 |
| 302 | SINGLE ORGANISM BEHAVIOR | 73 | 384 | 1.829e-05 | 0.0002819 |
| 303 | REGULATION OF ORGAN GROWTH | 22 | 73 | 1.925e-05 | 0.0002956 |
| 304 | POSITIVE REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 21 | 68 | 1.957e-05 | 0.0002996 |
| 305 | RESPONSE TO ORGANOPHOSPHORUS | 34 | 139 | 2.032e-05 | 0.00031 |
| 306 | CELL DEATH | 160 | 1001 | 2.083e-05 | 0.0003168 |
| 307 | VASCULAR PROCESS IN CIRCULATORY SYSTEM | 38 | 163 | 2.211e-05 | 0.0003351 |
| 308 | REGULATION OF SMOOTH MUSCLE CELL MIGRATION | 17 | 49 | 2.219e-05 | 0.0003352 |
| 309 | MESENCHYMAL CELL DIFFERENTIATION | 33 | 134 | 2.302e-05 | 0.0003467 |
| 310 | MUSCLE TISSUE DEVELOPMENT | 56 | 275 | 2.315e-05 | 0.000347 |
| 311 | BEHAVIOR | 92 | 516 | 2.32e-05 | 0.000347 |
| 312 | NEGATIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 30 | 117 | 2.331e-05 | 0.0003477 |
| 313 | REGULATION OF GLUCOSE METABOLIC PROCESS | 28 | 106 | 2.396e-05 | 0.0003562 |
| 314 | NEGATIVE REGULATION OF KINASE ACTIVITY | 52 | 250 | 2.453e-05 | 0.0003623 |
| 315 | NEGATIVE REGULATION OF CELL CYCLE PHASE TRANSITION | 35 | 146 | 2.452e-05 | 0.0003623 |
| 316 | NEGATIVE REGULATION OF LOCOMOTION | 54 | 263 | 2.529e-05 | 0.0003724 |
| 317 | RESPONSE TO PURINE CONTAINING COMPOUND | 37 | 158 | 2.537e-05 | 0.0003724 |
| 318 | NEGATIVE REGULATION OF FATTY ACID BIOSYNTHETIC PROCESS | 8 | 13 | 2.599e-05 | 0.0003803 |
| 319 | ORGANELLE LOCALIZATION | 77 | 415 | 2.638e-05 | 0.0003848 |
| 320 | SPROUTING ANGIOGENESIS | 16 | 45 | 2.714e-05 | 0.0003947 |
| 321 | EAR DEVELOPMENT | 43 | 195 | 2.832e-05 | 0.0004105 |
| 322 | AMINO ACID TRANSPORT | 31 | 124 | 2.938e-05 | 0.0004246 |
| 323 | POSITIVE REGULATION OF BMP SIGNALING PATHWAY | 13 | 32 | 3.006e-05 | 0.000433 |
| 324 | STEM CELL PROLIFERATION | 19 | 60 | 3.255e-05 | 0.000466 |
| 325 | GLUTAMINE FAMILY AMINO ACID METABOLIC PROCESS | 20 | 65 | 3.254e-05 | 0.000466 |
| 326 | REGULATION OF PEPTIDE SECRETION | 45 | 209 | 3.489e-05 | 0.000498 |
| 327 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 169 | 1079 | 3.819e-05 | 0.0005434 |
| 328 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 170 | 1087 | 3.93e-05 | 0.0005576 |
| 329 | RESPONSE TO MECHANICAL STIMULUS | 45 | 210 | 3.946e-05 | 0.000558 |
| 330 | CIRCULATORY SYSTEM PROCESS | 69 | 366 | 4.033e-05 | 0.0005687 |
| 331 | RESPONSE TO VITAMIN | 26 | 98 | 4.254e-05 | 0.0005981 |
| 332 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 33 | 138 | 4.354e-05 | 0.0006103 |
| 333 | RESPONSE TO CAMP | 27 | 104 | 4.627e-05 | 0.0006465 |
| 334 | EMBRYONIC SKELETAL SYSTEM MORPHOGENESIS | 25 | 93 | 4.677e-05 | 0.0006515 |
| 335 | REGULATION OF PEPTIDE TRANSPORT | 52 | 256 | 4.775e-05 | 0.0006632 |
| 336 | CELLULAR TRANSITION METAL ION HOMEOSTASIS | 22 | 77 | 4.804e-05 | 0.0006652 |
| 337 | SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 31 | 127 | 4.822e-05 | 0.0006658 |
| 338 | STEROID METABOLIC PROCESS | 49 | 237 | 4.872e-05 | 0.0006688 |
| 339 | MUSCLE CELL DIFFERENTIATION | 49 | 237 | 4.872e-05 | 0.0006688 |
| 340 | PEPTIDYL AMINO ACID MODIFICATION | 136 | 841 | 4.986e-05 | 0.0006823 |
| 341 | WATER SOLUBLE VITAMIN METABOLIC PROCESS | 24 | 88 | 5.109e-05 | 0.0006963 |
| 342 | HISTONE PHOSPHORYLATION | 11 | 25 | 5.118e-05 | 0.0006963 |
| 343 | REGULATION OF NUCLEAR DIVISION | 37 | 163 | 5.196e-05 | 0.0007049 |
| 344 | AMINO ACID TRANSMEMBRANE TRANSPORT | 20 | 67 | 5.284e-05 | 0.0007147 |
| 345 | EMBRYONIC SKELETAL SYSTEM DEVELOPMENT | 30 | 122 | 5.471e-05 | 0.0007359 |
| 346 | POSITIVE REGULATION OF CELL DEATH | 103 | 605 | 5.472e-05 | 0.0007359 |
| 347 | EMBRYONIC PLACENTA DEVELOPMENT | 23 | 83 | 5.541e-05 | 0.0007431 |
| 348 | REGULATION OF MEMBRANE POTENTIAL | 65 | 343 | 5.661e-05 | 0.0007569 |
| 349 | COENZYME METABOLIC PROCESS | 53 | 265 | 6.254e-05 | 0.0008338 |
| 350 | NEGATIVE REGULATION OF DEVELOPMENTAL GROWTH | 23 | 84 | 6.795e-05 | 0.0009033 |
| 351 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 140 | 876 | 6.858e-05 | 0.0009067 |
| 352 | ESTABLISHMENT OR MAINTENANCE OF CELL POLARITY | 33 | 141 | 6.859e-05 | 0.0009067 |
| 353 | ACTIVATION OF PROTEIN KINASE ACTIVITY | 55 | 279 | 6.916e-05 | 0.0009116 |
| 354 | REGULATION OF GTPASE ACTIVITY | 112 | 673 | 6.997e-05 | 0.0009197 |
| 355 | CARBOHYDRATE DERIVATIVE METABOLIC PROCESS | 163 | 1047 | 7.191e-05 | 0.0009426 |
| 356 | POSITIVE REGULATION OF TRANSPORT | 148 | 936 | 7.277e-05 | 0.0009511 |
| 357 | SMOOTH MUSCLE TISSUE DEVELOPMENT | 9 | 18 | 7.326e-05 | 0.0009549 |
| 358 | DICARBOXYLIC ACID METABOLIC PROCESS | 26 | 101 | 7.402e-05 | 0.000962 |
| 359 | ORGANOPHOSPHATE METABOLIC PROCESS | 141 | 885 | 7.501e-05 | 0.0009722 |
| 360 | REGULATION OF AXON GUIDANCE | 14 | 39 | 7.597e-05 | 0.0009819 |
| 361 | REGULATION OF STEROID METABOLIC PROCESS | 21 | 74 | 7.89e-05 | 0.001017 |
| 362 | TEMPERATURE HOMEOSTASIS | 11 | 26 | 7.935e-05 | 0.00102 |
| 363 | CELLULAR RESPONSE TO PEPTIDE | 54 | 274 | 8.07e-05 | 0.001034 |
| 364 | POSITIVE REGULATION OF CELL CYCLE ARREST | 23 | 85 | 8.296e-05 | 0.00106 |
| 365 | POSITIVE REGULATION OF DNA METABOLIC PROCESS | 40 | 185 | 8.387e-05 | 0.001069 |
| 366 | NEGATIVE REGULATION OF GROWTH | 48 | 236 | 8.845e-05 | 0.001125 |
| 367 | REGULATION OF HORMONE SECRETION | 52 | 262 | 8.972e-05 | 0.001137 |
| 368 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 78 | 437 | 9.071e-05 | 0.001144 |
| 369 | REGULATION OF DNA REPLICATION | 36 | 161 | 9.054e-05 | 0.001144 |
| 370 | HINDBRAIN DEVELOPMENT | 32 | 137 | 9.132e-05 | 0.001148 |
| 371 | REGULATION OF NEURON APOPTOTIC PROCESS | 41 | 192 | 9.226e-05 | 0.001157 |
| 372 | POSITIVE REGULATION OF DNA REPLICATION | 23 | 86 | 0.0001008 | 0.00125 |
| 373 | RESPONSE TO ESTROGEN | 45 | 218 | 0.0001008 | 0.00125 |
| 374 | CELLULAR RESPONSE TO STEROID HORMONE STIMULUS | 45 | 218 | 0.0001008 | 0.00125 |
| 375 | CELLULAR COMPONENT DISASSEMBLY | 89 | 515 | 0.0001002 | 0.00125 |
| 376 | NEGATIVE REGULATION OF CELL PROLIFERATION | 107 | 643 | 0.000101 | 0.00125 |
| 377 | DIVALENT INORGANIC CATION HOMEOSTASIS | 64 | 343 | 0.0001029 | 0.00127 |
| 378 | NEGATIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 30 | 126 | 0.0001032 | 0.001271 |
| 379 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 14 | 40 | 0.0001044 | 0.001274 |
| 380 | CELLULAR RESPONSE TO STEROL | 8 | 15 | 0.0001046 | 0.001274 |
| 381 | RESPONSE TO CADMIUM ION | 14 | 40 | 0.0001044 | 0.001274 |
| 382 | DEVELOPMENTAL MATURATION | 41 | 193 | 0.0001041 | 0.001274 |
| 383 | CHROMOSOME CONDENSATION | 12 | 31 | 0.0001064 | 0.001286 |
| 384 | NEGATIVE REGULATION OF CELL PROJECTION ORGANIZATION | 33 | 144 | 0.000106 | 0.001286 |
| 385 | REGULATION OF PLATELET ACTIVATION | 12 | 31 | 0.0001064 | 0.001286 |
| 386 | POSITIVE REGULATION OF GROWTH | 48 | 238 | 0.0001097 | 0.001322 |
| 387 | METANEPHROS DEVELOPMENT | 22 | 81 | 0.0001105 | 0.001325 |
| 388 | OSSIFICATION | 50 | 251 | 0.0001103 | 0.001325 |
| 389 | CENTROSOME CYCLE | 15 | 45 | 0.0001128 | 0.001346 |
| 390 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 94 | 552 | 0.0001128 | 0.001346 |
| 391 | NEGATIVE REGULATION OF CELL DIVISION | 18 | 60 | 0.0001148 | 0.001366 |
| 392 | CONNECTIVE TISSUE DEVELOPMENT | 41 | 194 | 0.0001173 | 0.001393 |
| 393 | POSITIVE REGULATION OF HEART GROWTH | 11 | 27 | 0.0001198 | 0.001418 |
| 394 | REGULATION OF NEURON DEATH | 50 | 252 | 0.0001222 | 0.001443 |
| 395 | RESPONSE TO INCREASED OXYGEN LEVELS | 10 | 23 | 0.0001277 | 0.001493 |
| 396 | SEMAPHORIN PLEXIN SIGNALING PATHWAY | 13 | 36 | 0.0001273 | 0.001493 |
| 397 | RESPONSE TO HYPEROXIA | 10 | 23 | 0.0001277 | 0.001493 |
| 398 | POSITIVE REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 13 | 36 | 0.0001273 | 0.001493 |
| 399 | DICARBOXYLIC ACID TRANSPORT | 20 | 71 | 0.0001295 | 0.001511 |
| 400 | NEGATIVE REGULATION OF PROTEIN COMPLEX DISASSEMBLY | 37 | 170 | 0.0001317 | 0.001532 |
| 401 | CELLULAR AMINO ACID BIOSYNTHETIC PROCESS | 24 | 93 | 0.0001322 | 0.001534 |
| 402 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 58 | 306 | 0.0001375 | 0.001591 |
| 403 | KINETOCHORE ORGANIZATION | 7 | 12 | 0.0001383 | 0.001597 |
| 404 | DNA DEPENDENT DNA REPLICATION | 25 | 99 | 0.0001408 | 0.001598 |
| 405 | RESPONSE TO MITOCHONDRIAL DEPOLARISATION | 31 | 134 | 0.0001407 | 0.001598 |
| 406 | CELLULAR RESPONSE TO INSULIN STIMULUS | 33 | 146 | 0.0001402 | 0.001598 |
| 407 | REGULATION OF LIPID BIOSYNTHETIC PROCESS | 30 | 128 | 0.0001397 | 0.001598 |
| 408 | MITOPHAGY IN RESPONSE TO MITOCHONDRIAL DEPOLARIZATION | 31 | 134 | 0.0001407 | 0.001598 |
| 409 | LIPID PHOSPHORYLATION | 25 | 99 | 0.0001408 | 0.001598 |
| 410 | MACROMITOPHAGY | 31 | 134 | 0.0001407 | 0.001598 |
| 411 | AORTA DEVELOPMENT | 14 | 41 | 0.0001415 | 0.001598 |
| 412 | REGULATION OF POTASSIUM ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 14 | 41 | 0.0001415 | 0.001598 |
| 413 | CHROMOSOME LOCALIZATION | 18 | 61 | 0.0001452 | 0.001636 |
| 414 | DEVELOPMENTAL CELL GROWTH | 21 | 77 | 0.0001472 | 0.001655 |
| 415 | RESPONSE TO XENOBIOTIC STIMULUS | 26 | 105 | 0.0001478 | 0.001657 |
| 416 | NEGATIVE REGULATION OF DNA BINDING | 15 | 46 | 0.0001494 | 0.001671 |
| 417 | TUBE FORMATION | 30 | 129 | 0.000162 | 0.001807 |
| 418 | REGULATION OF CATABOLIC PROCESS | 118 | 731 | 0.0001663 | 0.001851 |
| 419 | REGULATION OF GLUCOSE TRANSPORT | 25 | 100 | 0.0001671 | 0.001856 |
| 420 | RESPONSE TO REACTIVE OXYGEN SPECIES | 40 | 191 | 0.0001731 | 0.001917 |
| 421 | GLYCEROLIPID METABOLIC PROCESS | 65 | 356 | 0.0001745 | 0.001924 |
| 422 | NEGATIVE REGULATION OF CELL ADHESION | 45 | 223 | 0.0001744 | 0.001924 |
| 423 | REGULATION OF FATTY ACID OXIDATION | 11 | 28 | 0.0001765 | 0.001942 |
| 424 | REGULATION OF PROTEIN COMPLEX DISASSEMBLY | 44 | 217 | 0.0001829 | 0.002007 |
| 425 | POSITIVE REGULATION OF CELL GROWTH | 33 | 148 | 0.0001841 | 0.002011 |
| 426 | RESPONSE TO TRANSITION METAL NANOPARTICLE | 33 | 148 | 0.0001841 | 0.002011 |
| 427 | PLATELET ACTIVATION | 32 | 142 | 0.0001861 | 0.002028 |
| 428 | METAPHASE PLATE CONGRESSION | 14 | 42 | 0.0001896 | 0.002061 |
| 429 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 57 | 303 | 0.0001919 | 0.002081 |
| 430 | REGULATION OF CELLULAR COMPONENT SIZE | 62 | 337 | 0.0001963 | 0.002109 |
| 431 | HIPPOCAMPUS DEVELOPMENT | 20 | 73 | 0.0001962 | 0.002109 |
| 432 | G1 DNA DAMAGE CHECKPOINT | 20 | 73 | 0.0001962 | 0.002109 |
| 433 | LIPID BIOSYNTHETIC PROCESS | 91 | 539 | 0.0001956 | 0.002109 |
| 434 | CYTOKINESIS | 22 | 84 | 0.0001968 | 0.00211 |
| 435 | RESPONSE TO VITAMIN A | 9 | 20 | 0.0002033 | 0.00217 |
| 436 | AXONAL FASCICULATION | 9 | 20 | 0.0002033 | 0.00217 |
| 437 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 174 | 1152 | 0.0002052 | 0.002185 |
| 438 | G2 DNA DAMAGE CHECKPOINT | 12 | 33 | 0.0002138 | 0.002271 |
| 439 | REGULATION OF PROTEIN BINDING | 36 | 168 | 0.0002229 | 0.002363 |
| 440 | CARDIOCYTE DIFFERENTIATION | 24 | 96 | 0.0002237 | 0.002365 |
| 441 | PALATE DEVELOPMENT | 22 | 85 | 0.0002366 | 0.002491 |
| 442 | REGULATION OF CHROMOSOME SEGREGATION | 22 | 85 | 0.0002366 | 0.002491 |
| 443 | POSITIVE REGULATION OF POTASSIUM ION TRANSPORT | 13 | 38 | 0.0002383 | 0.002497 |
| 444 | REGULATION OF INTRACELLULAR TRANSPORT | 102 | 621 | 0.0002378 | 0.002497 |
| 445 | CARDIAC MUSCLE CELL DIFFERENTIATION | 20 | 74 | 0.0002397 | 0.002506 |
| 446 | L AMINO ACID TRANSPORT | 17 | 58 | 0.0002408 | 0.002506 |
| 447 | NEGATIVE REGULATION OF WOUND HEALING | 17 | 58 | 0.0002408 | 0.002506 |
| 448 | GLIOGENESIS | 37 | 175 | 0.0002438 | 0.002532 |
| 449 | RESPONSE TO METAL ION | 61 | 333 | 0.0002481 | 0.002571 |
| 450 | SENSORY ORGAN DEVELOPMENT | 84 | 493 | 0.0002499 | 0.002584 |
| 451 | CENTROMERE COMPLEX ASSEMBLY | 15 | 48 | 0.0002543 | 0.002619 |
| 452 | NEGATIVE REGULATION OF FATTY ACID METABOLIC PROCESS | 11 | 29 | 0.0002545 | 0.002619 |
| 453 | NUCLEOSIDE PHOSPHATE CATABOLIC PROCESS | 19 | 69 | 0.0002615 | 0.002686 |
| 454 | NEGATIVE REGULATION OF LIPID METABOLIC PROCESS | 21 | 80 | 0.0002633 | 0.002699 |
| 455 | CARNITINE METABOLIC PROCESS | 7 | 13 | 0.0002693 | 0.002742 |
| 456 | SEMAPHORIN PLEXIN SIGNALING PATHWAY INVOLVED IN NEURON PROJECTION GUIDANCE | 7 | 13 | 0.0002693 | 0.002742 |
| 457 | VENTRICULAR CARDIAC MUSCLE CELL DEVELOPMENT | 7 | 13 | 0.0002693 | 0.002742 |
| 458 | REGULATION OF CATION TRANSMEMBRANE TRANSPORT | 42 | 208 | 0.0002777 | 0.002821 |
| 459 | PROTEIN COMPLEX SUBUNIT ORGANIZATION | 222 | 1527 | 0.0002815 | 0.002854 |
| 460 | POSITIVE REGULATION OF RESPONSE TO DNA DAMAGE STIMULUS | 18 | 64 | 0.0002826 | 0.002859 |
| 461 | NEPHRON DEVELOPMENT | 27 | 115 | 0.0002855 | 0.002881 |
| 462 | CELL DIFFERENTIATION INVOLVED IN EMBRYONIC PLACENTA DEVELOPMENT | 10 | 25 | 0.0002926 | 0.002947 |
| 463 | RESPONSE TO FLUID SHEAR STRESS | 12 | 34 | 0.0002955 | 0.002969 |
| 464 | METANEPHRIC NEPHRON MORPHOGENESIS | 9 | 21 | 0.0003189 | 0.003167 |
| 465 | NEGATIVE REGULATION OF PLATELET ACTIVATION | 8 | 17 | 0.0003182 | 0.003167 |
| 466 | REGULATION OF PLATELET AGGREGATION | 8 | 17 | 0.0003182 | 0.003167 |
| 467 | REGULATION OF SECRETION | 112 | 699 | 0.0003219 | 0.003167 |
| 468 | POSITIVE REGULATION OF PROTEIN IMPORT | 25 | 104 | 0.0003215 | 0.003167 |
| 469 | STEM CELL DIFFERENTIATION | 39 | 190 | 0.0003187 | 0.003167 |
| 470 | NEGATIVE REGULATION OF NEURON DEATH | 36 | 171 | 0.0003201 | 0.003167 |
| 471 | REGULATION OF ORGAN MORPHOGENESIS | 47 | 242 | 0.0003203 | 0.003167 |
| 472 | CELLULAR RESPONSE TO REACTIVE OXYGEN SPECIES | 25 | 104 | 0.0003215 | 0.003167 |
| 473 | REGULATION OF TRANSMEMBRANE TRANSPORT | 74 | 426 | 0.0003171 | 0.003167 |
| 474 | NEURON MIGRATION | 26 | 110 | 0.0003278 | 0.003204 |
| 475 | REGULATION OF STEROID BIOSYNTHETIC PROCESS | 15 | 49 | 0.0003273 | 0.003204 |
| 476 | CARDIAC CELL DEVELOPMENT | 15 | 49 | 0.0003273 | 0.003204 |
| 477 | NEGATIVE REGULATION OF LIPID BIOSYNTHETIC PROCESS | 14 | 44 | 0.000329 | 0.003209 |
| 478 | CELLULAR RESPONSE TO OXIDATIVE STRESS | 38 | 184 | 0.0003325 | 0.003237 |
| 479 | REGULATION OF OSSIFICATION | 37 | 178 | 0.0003464 | 0.003365 |
| 480 | NEGATIVE REGULATION OF AXONOGENESIS | 18 | 65 | 0.0003486 | 0.003379 |
| 481 | REGULATION OF CALCIUM MEDIATED SIGNALING | 20 | 76 | 0.0003525 | 0.003402 |
| 482 | NEGATIVE REGULATION OF TRANSFERASE ACTIVITY | 63 | 351 | 0.000352 | 0.003402 |
| 483 | REGULATION OF CARBOHYDRATE METABOLIC PROCESS | 36 | 172 | 0.00036 | 0.003461 |
| 484 | DNA STRAND ELONGATION | 11 | 30 | 0.0003596 | 0.003461 |
| 485 | CENTRAL NERVOUS SYSTEM NEURON DIFFERENTIATION | 35 | 166 | 0.0003734 | 0.003582 |
| 486 | REGULATION OF TRANSPORTER ACTIVITY | 40 | 198 | 0.0003789 | 0.003628 |
| 487 | REGULATION OF RECEPTOR ACTIVITY | 27 | 117 | 0.0003833 | 0.003656 |
| 488 | REGIONALIZATION | 57 | 311 | 0.0003834 | 0.003656 |
| 489 | PURINE CONTAINING COMPOUND METABOLIC PROCESS | 69 | 394 | 0.0003926 | 0.003736 |
| 490 | ESTABLISHMENT OF CELL POLARITY | 22 | 88 | 0.0004013 | 0.003811 |
| 491 | RHYTHMIC PROCESS | 55 | 298 | 0.0004026 | 0.003815 |
| 492 | CELLULAR RESPONSE TO CAMP | 15 | 50 | 0.0004175 | 0.003948 |
| 493 | METENCEPHALON DEVELOPMENT | 24 | 100 | 0.0004302 | 0.004052 |
| 494 | LIMBIC SYSTEM DEVELOPMENT | 24 | 100 | 0.0004302 | 0.004052 |
| 495 | POSITIVE REGULATION OF NEURON PROJECTION DEVELOPMENT | 45 | 232 | 0.0004366 | 0.004104 |
| 496 | PHOSPHATIDYLINOSITOL METABOLIC PROCESS | 39 | 193 | 0.0004427 | 0.004153 |
| 497 | CELLULAR RESPONSE TO DNA DAMAGE STIMULUS | 114 | 720 | 0.0004526 | 0.004237 |
| 498 | CELLULAR RESPONSE TO HYDROGEN PEROXIDE | 17 | 61 | 0.0004647 | 0.004333 |
| 499 | POSITIVE REGULATION OF STEM CELL PROLIFERATION | 17 | 61 | 0.0004647 | 0.004333 |
| 500 | CARBOHYDRATE METABOLIC PROCESS | 106 | 662 | 0.0004713 | 0.004386 |
| 501 | EPITHELIAL CELL PROLIFERATION | 22 | 89 | 0.0004751 | 0.004412 |
| 502 | BIOTIN METABOLIC PROCESS | 7 | 14 | 0.0004842 | 0.004435 |
| 503 | DNA CONFORMATION CHANGE | 51 | 273 | 0.0004795 | 0.004435 |
| 504 | INSULIN LIKE GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 7 | 14 | 0.0004842 | 0.004435 |
| 505 | REGULATION OF CHOLESTEROL METABOLIC PROCESS | 9 | 22 | 0.0004839 | 0.004435 |
| 506 | CAMP CATABOLIC PROCESS | 7 | 14 | 0.0004842 | 0.004435 |
| 507 | AORTA MORPHOGENESIS | 9 | 22 | 0.0004839 | 0.004435 |
| 508 | HISTONE H3 DEACETYLATION | 9 | 22 | 0.0004839 | 0.004435 |
| 509 | RESPONSE TO ALKALOID | 30 | 137 | 0.0004883 | 0.004446 |
| 510 | PATTERN SPECIFICATION PROCESS | 72 | 418 | 0.0004864 | 0.004446 |
| 511 | ACTIVATION OF MAPK ACTIVITY | 30 | 137 | 0.0004883 | 0.004446 |
| 512 | MITOTIC CYTOKINESIS | 11 | 31 | 0.0004988 | 0.004533 |
| 513 | CENTROSOME LOCALIZATION | 8 | 18 | 0.0005142 | 0.004664 |
| 514 | POSITIVE REGULATION OF MITOTIC NUCLEAR DIVISION | 15 | 51 | 0.0005282 | 0.004763 |
| 515 | ARTERY MORPHOGENESIS | 15 | 51 | 0.0005282 | 0.004763 |
| 516 | REGULATION OF BLOOD VESSEL ENDOTHELIAL CELL MIGRATION | 15 | 51 | 0.0005282 | 0.004763 |
| 517 | RESPONSE TO TOXIC SUBSTANCE | 46 | 241 | 0.0005476 | 0.004928 |
| 518 | IMMUNE SYSTEM PROCESS | 278 | 1984 | 0.0005542 | 0.004978 |
| 519 | CELL CYCLE G2 M PHASE TRANSITION | 30 | 138 | 0.0005552 | 0.004978 |
| 520 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 114 | 724 | 0.000558 | 0.004993 |
| 521 | FOREBRAIN CELL MIGRATION | 17 | 62 | 0.0005714 | 0.005093 |
| 522 | POSITIVE REGULATION OF NUCLEAR DIVISION | 17 | 62 | 0.0005714 | 0.005093 |
| 523 | REGULATION OF CELL ADHESION | 101 | 629 | 0.0005744 | 0.00511 |
| 524 | MOTOR NEURON AXON GUIDANCE | 10 | 27 | 0.0006059 | 0.005349 |
| 525 | POSITIVE REGULATION OF MESENCHYMAL CELL PROLIFERATION | 10 | 27 | 0.0006059 | 0.005349 |
| 526 | POSITIVE REGULATION OF POTASSIUM ION TRANSMEMBRANE TRANSPORT | 10 | 27 | 0.0006059 | 0.005349 |
| 527 | REGULATION OF TRANSCRIPTION INVOLVED IN G1 S TRANSITION OF MITOTIC CELL CYCLE | 10 | 27 | 0.0006059 | 0.005349 |
| 528 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 99 | 616 | 0.0006284 | 0.005538 |
| 529 | HORMONE MEDIATED SIGNALING PATHWAY | 33 | 158 | 0.0006437 | 0.005662 |
| 530 | SULFUR COMPOUND METABOLIC PROCESS | 63 | 359 | 0.0006491 | 0.005698 |
| 531 | POSITIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 27 | 121 | 0.0006711 | 0.005881 |
| 532 | NEGATIVE REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 11 | 32 | 0.0006804 | 0.00594 |
| 533 | METANEPHRIC NEPHRON DEVELOPMENT | 11 | 32 | 0.0006804 | 0.00594 |
| 534 | MEMBRANE DISASSEMBLY | 14 | 47 | 0.0006973 | 0.006035 |
| 535 | POSITIVE REGULATION OF CATABOLIC PROCESS | 68 | 395 | 0.0006972 | 0.006035 |
| 536 | POSITIVE REGULATION OF OXIDOREDUCTASE ACTIVITY | 14 | 47 | 0.0006973 | 0.006035 |
| 537 | NUCLEAR ENVELOPE DISASSEMBLY | 14 | 47 | 0.0006973 | 0.006035 |
| 538 | HETEROCHROMATIN ORGANIZATION | 6 | 11 | 0.000699 | 0.006035 |
| 539 | CORPUS CALLOSUM DEVELOPMENT | 6 | 11 | 0.000699 | 0.006035 |
| 540 | REGULATION OF SYNAPTIC PLASTICITY | 30 | 140 | 0.0007139 | 0.006151 |
| 541 | REGULATION OF HEART GROWTH | 13 | 42 | 0.0007169 | 0.006166 |
| 542 | METAL ION TRANSPORT | 94 | 582 | 0.0007224 | 0.006181 |
| 543 | REGULATION OF JNK CASCADE | 33 | 159 | 0.0007227 | 0.006181 |
| 544 | REGULATED EXOCYTOSIS | 43 | 224 | 0.0007214 | 0.006181 |
| 545 | REGULATION OF SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 51 | 278 | 0.0007378 | 0.006299 |
| 546 | MONOCARBOXYLIC ACID BIOSYNTHETIC PROCESS | 35 | 172 | 0.000741 | 0.006314 |
| 547 | CELL ACTIVATION | 92 | 568 | 0.0007458 | 0.006344 |
| 548 | ORGANELLE DISASSEMBLY | 37 | 185 | 0.0007483 | 0.006354 |
| 549 | POSITIVE REGULATION OF LIPID METABOLIC PROCESS | 28 | 128 | 0.0007523 | 0.006376 |
| 550 | CARTILAGE DEVELOPMENT | 31 | 147 | 0.0007801 | 0.00659 |
| 551 | MICROTUBULE BASED MOVEMENT | 40 | 205 | 0.0007803 | 0.00659 |
| 552 | NEGATIVE REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 23 | 98 | 0.0007831 | 0.006601 |
| 553 | LIPID HOMEOSTASIS | 25 | 110 | 0.0007859 | 0.006612 |
| 554 | VENTRICULAR CARDIAC MUSCLE CELL DIFFERENTIATION | 8 | 19 | 0.0007975 | 0.006698 |
| 555 | MESENCHYMAL TO EPITHELIAL TRANSITION | 7 | 15 | 0.0008164 | 0.00682 |
| 556 | NEGATIVE REGULATION OF SMOOTH MUSCLE CONTRACTION | 7 | 15 | 0.0008164 | 0.00682 |
| 557 | FRUCTOSE METABOLIC PROCESS | 7 | 15 | 0.0008164 | 0.00682 |
| 558 | NEURON PROJECTION EXTENSION | 15 | 53 | 0.0008259 | 0.006842 |
| 559 | POSITIVE REGULATION OF FIBROBLAST PROLIFERATION | 15 | 53 | 0.0008259 | 0.006842 |
| 560 | STRIATED MUSCLE CELL DIFFERENTIATION | 35 | 173 | 0.0008263 | 0.006842 |
| 561 | REGULATION OF CELLULAR KETONE METABOLIC PROCESS | 35 | 173 | 0.0008263 | 0.006842 |
| 562 | MESONEPHRIC TUBULE MORPHOGENESIS | 15 | 53 | 0.0008259 | 0.006842 |
| 563 | ADULT BEHAVIOR | 29 | 135 | 0.0008325 | 0.00688 |
| 564 | METANEPHROS MORPHOGENESIS | 10 | 28 | 0.0008444 | 0.006966 |
| 565 | DNA GEOMETRIC CHANGE | 20 | 81 | 0.0008553 | 0.007043 |
| 566 | NEPHRON EPITHELIUM DEVELOPMENT | 22 | 93 | 0.0009006 | 0.007391 |
| 567 | POSITIVE REGULATION OF BLOOD CIRCULATION | 22 | 93 | 0.0009006 | 0.007391 |
| 568 | CEREBRAL CORTEX CELL MIGRATION | 13 | 43 | 0.0009187 | 0.007526 |
| 569 | REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 42 | 220 | 0.0009258 | 0.00757 |
| 570 | POSITIVE REGULATION OF ORGAN GROWTH | 12 | 38 | 0.0009339 | 0.007624 |
| 571 | VASCULOGENESIS | 16 | 59 | 0.0009358 | 0.007625 |
| 572 | RESPONSE TO ETHANOL | 29 | 136 | 0.000942 | 0.007662 |
| 573 | NEUROTRANSMITTER TRANSPORT | 32 | 155 | 0.000947 | 0.00769 |
| 574 | CARBOHYDRATE DERIVATIVE BIOSYNTHETIC PROCESS | 95 | 595 | 0.0009922 | 0.008043 |
| 575 | REGULATION OF CELLULAR RESPONSE TO STRESS | 108 | 691 | 0.001 | 0.008095 |
| 576 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 33 | 162 | 0.001013 | 0.008185 |
| 577 | REGULATION OF HEART CONTRACTION | 42 | 221 | 0.001016 | 0.008195 |
| 578 | STEROID CATABOLIC PROCESS | 9 | 24 | 0.001023 | 0.008236 |
| 579 | CATION TRANSPORT | 122 | 796 | 0.001032 | 0.008292 |
| 580 | REGULATION OF BLOOD PRESSURE | 34 | 169 | 0.001076 | 0.008636 |
| 581 | CELL MATURATION | 28 | 131 | 0.001099 | 0.008774 |
| 582 | PROTEIN STABILIZATION | 28 | 131 | 0.001099 | 0.008774 |
| 583 | NEGATIVE REGULATION OF BINDING | 28 | 131 | 0.001099 | 0.008774 |
| 584 | MONOSACCHARIDE METABOLIC PROCESS | 39 | 202 | 0.001109 | 0.008822 |
| 585 | EPITHELIAL CELL DIFFERENTIATION | 81 | 495 | 0.001109 | 0.008822 |
| 586 | STEROID HORMONE MEDIATED SIGNALING PATHWAY | 27 | 125 | 0.001132 | 0.00899 |
| 587 | REGULATION OF GLUCOSE IMPORT | 16 | 60 | 0.001139 | 0.009032 |
| 588 | ALPHA AMINO ACID BIOSYNTHETIC PROCESS | 19 | 77 | 0.001149 | 0.009092 |
| 589 | RESPONSE TO ARSENIC CONTAINING SUBSTANCE | 10 | 29 | 0.001155 | 0.009092 |
| 590 | CEREBRAL CORTEX RADIALLY ORIENTED CELL MIGRATION | 10 | 29 | 0.001155 | 0.009092 |
| 591 | NEUROBLAST PROLIFERATION | 10 | 29 | 0.001155 | 0.009092 |
| 592 | BRANCHING INVOLVED IN URETERIC BUD MORPHOGENESIS | 13 | 44 | 0.001166 | 0.009165 |
| 593 | CEREBRAL CORTEX RADIAL GLIA GUIDED MIGRATION | 8 | 20 | 0.001194 | 0.009226 |
| 594 | BILE ACID BIOSYNTHETIC PROCESS | 8 | 20 | 0.001194 | 0.009226 |
| 595 | TELENCEPHALON GLIAL CELL MIGRATION | 8 | 20 | 0.001194 | 0.009226 |
| 596 | MIDDLE EAR MORPHOGENESIS | 8 | 20 | 0.001194 | 0.009226 |
| 597 | ORGANOPHOSPHATE CATABOLIC PROCESS | 25 | 113 | 0.001185 | 0.009226 |
| 598 | TRACHEA DEVELOPMENT | 8 | 20 | 0.001194 | 0.009226 |
| 599 | REGULATION OF PROTEIN IMPORT | 36 | 183 | 0.001193 | 0.009226 |
| 600 | RESPONSE TO LEAD ION | 8 | 20 | 0.001194 | 0.009226 |
| 601 | RESPONSE TO LEPTIN | 8 | 20 | 0.001194 | 0.009226 |
| 602 | DEVELOPMENT OF PRIMARY SEXUAL CHARACTERISTICS | 41 | 216 | 0.001187 | 0.009226 |
| 603 | LUNG EPITHELIUM DEVELOPMENT | 11 | 34 | 0.001209 | 0.00927 |
| 604 | REGULATION OF MESENCHYMAL CELL PROLIFERATION | 11 | 34 | 0.001209 | 0.00927 |
| 605 | CAMP METABOLIC PROCESS | 11 | 34 | 0.001209 | 0.00927 |
| 606 | NEGATIVE CHEMOTAXIS | 12 | 39 | 0.001207 | 0.00927 |
| 607 | CYTOSKELETON DEPENDENT CYTOKINESIS | 12 | 39 | 0.001207 | 0.00927 |
| 608 | ALPHA AMINO ACID CATABOLIC PROCESS | 22 | 95 | 0.001216 | 0.009304 |
| 609 | REGULATION OF ACTIN FILAMENT BASED PROCESS | 55 | 312 | 0.001239 | 0.009465 |
| 610 | REGULATION OF NEUROTRANSMITTER LEVELS | 37 | 190 | 0.001247 | 0.009509 |
| 611 | CARTILAGE MORPHOGENESIS | 6 | 12 | 0.00126 | 0.009516 |
| 612 | CELL SUBSTRATE ADHESION | 33 | 164 | 0.001259 | 0.009516 |
| 613 | CELLULAR RESPONSE TO CHOLESTEROL | 6 | 12 | 0.00126 | 0.009516 |
| 614 | CRANIAL SKELETAL SYSTEM DEVELOPMENT | 15 | 55 | 0.001255 | 0.009516 |
| 615 | ASPARTATE FAMILY AMINO ACID METABOLIC PROCESS | 15 | 55 | 0.001255 | 0.009516 |
| 616 | REGULATION OF ION TRANSPORT | 94 | 592 | 0.001255 | 0.009516 |
| 617 | CELLULAR RESPONSE TO EXTERNAL STIMULUS | 48 | 264 | 0.001262 | 0.009519 |
| 618 | REGULATION OF WOUND HEALING | 27 | 126 | 0.001283 | 0.009662 |
| 619 | REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 38 | 197 | 0.001296 | 0.009745 |
| 620 | GENERATION OF PRECURSOR METABOLITES AND ENERGY | 52 | 292 | 0.001316 | 0.009873 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | RIBONUCLEOTIDE BINDING | 329 | 1860 | 9.843e-16 | 4.572e-13 |
| 2 | MACROMOLECULAR COMPLEX BINDING | 263 | 1399 | 7.485e-16 | 4.572e-13 |
| 3 | ADENYL NUCLEOTIDE BINDING | 277 | 1514 | 3.971e-15 | 1.23e-12 |
| 4 | PROTEIN COMPLEX BINDING | 185 | 935 | 1.966e-13 | 4.565e-11 |
| 5 | ENZYME BINDING | 297 | 1737 | 2.237e-12 | 4.156e-10 |
| 6 | KINASE BINDING | 130 | 606 | 2.993e-12 | 4.634e-10 |
| 7 | IDENTICAL PROTEIN BINDING | 221 | 1209 | 3.588e-12 | 4.762e-10 |
| 8 | HYDROLASE ACTIVITY ACTING ON ACID ANHYDRIDES | 162 | 820 | 7.67e-12 | 8.907e-10 |
| 9 | CYTOSKELETAL PROTEIN BINDING | 158 | 819 | 9.288e-11 | 9.587e-09 |
| 10 | COFACTOR BINDING | 68 | 263 | 1.552e-10 | 1.442e-08 |
| 11 | ACTIVE TRANSMEMBRANE TRANSPORTER ACTIVITY | 82 | 356 | 1.012e-09 | 8.546e-08 |
| 12 | ANION TRANSMEMBRANE TRANSPORTER ACTIVITY | 70 | 293 | 3.347e-09 | 2.591e-07 |
| 13 | TUBULIN BINDING | 66 | 273 | 5.63e-09 | 4.023e-07 |
| 14 | TRANSITION METAL ION BINDING | 233 | 1400 | 8.293e-09 | 5.503e-07 |
| 15 | KINASE ACTIVITY | 152 | 842 | 2.39e-08 | 1.48e-06 |
| 16 | TRANSMEMBRANE TRANSPORTER ACTIVITY | 173 | 997 | 4.571e-08 | 2.654e-06 |
| 17 | ORGANIC ANION TRANSMEMBRANE TRANSPORTER ACTIVITY | 47 | 178 | 5.069e-08 | 2.77e-06 |
| 18 | MICROTUBULE BINDING | 51 | 201 | 5.699e-08 | 2.941e-06 |
| 19 | PROTEIN KINASE ACTIVITY | 119 | 640 | 1.66e-07 | 8.114e-06 |
| 20 | ZINC ION BINDING | 192 | 1155 | 2.049e-07 | 9.52e-06 |
| 21 | ORGANIC ACID TRANSMEMBRANE TRANSPORTER ACTIVITY | 39 | 142 | 2.232e-07 | 9.723e-06 |
| 22 | OXIDOREDUCTASE ACTIVITY | 130 | 719 | 2.303e-07 | 9.723e-06 |
| 23 | TRANSPORTER ACTIVITY | 208 | 1276 | 2.764e-07 | 1.117e-05 |
| 24 | PROTEIN HOMODIMERIZATION ACTIVITY | 130 | 722 | 2.921e-07 | 1.131e-05 |
| 25 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 167 | 992 | 5.902e-07 | 2.193e-05 |
| 26 | ATPASE ACTIVITY | 84 | 427 | 1.077e-06 | 3.847e-05 |
| 27 | COENZYME BINDING | 44 | 179 | 1.154e-06 | 3.972e-05 |
| 28 | PROTEIN DIMERIZATION ACTIVITY | 187 | 1149 | 1.273e-06 | 4.222e-05 |
| 29 | RECEPTOR BINDING | 231 | 1476 | 1.455e-06 | 4.662e-05 |
| 30 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 68 | 328 | 1.697e-06 | 5.254e-05 |
| 31 | ACTIVE ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 42 | 171 | 2.046e-06 | 6.132e-05 |
| 32 | PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 23 | 70 | 2.455e-06 | 7.128e-05 |
| 33 | SECONDARY ACTIVE TRANSMEMBRANE TRANSPORTER ACTIVITY | 52 | 233 | 3.014e-06 | 8.486e-05 |
| 34 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION REGULATORY REGION SEQUENCE SPECIFIC BINDING | 65 | 315 | 3.359e-06 | 9.166e-05 |
| 35 | KINASE REGULATOR ACTIVITY | 44 | 186 | 3.453e-06 | 9.166e-05 |
| 36 | ANION CATION SYMPORTER ACTIVITY | 19 | 53 | 4.251e-06 | 0.0001073 |
| 37 | MICROTUBULE MOTOR ACTIVITY | 24 | 77 | 4.273e-06 | 0.0001073 |
| 38 | SYMPORTER ACTIVITY | 36 | 142 | 4.948e-06 | 0.000121 |
| 39 | MOTOR ACTIVITY | 34 | 131 | 5.161e-06 | 0.0001229 |
| 40 | OXIDOREDUCTASE ACTIVITY ACTING ON PAIRED DONORS WITH INCORPORATION OR REDUCTION OF MOLECULAR OXYGEN NAD P H AS ONE DONOR AND INCORPORATION OF ONE ATOM OF OXYGEN | 15 | 37 | 7.617e-06 | 0.0001769 |
| 41 | RNA POLYMERASE II TRANSCRIPTION FACTOR ACTIVITY SEQUENCE SPECIFIC DNA BINDING | 110 | 629 | 9.376e-06 | 0.0002124 |
| 42 | ORGANIC ACID SODIUM SYMPORTER ACTIVITY | 13 | 30 | 1.299e-05 | 0.0002874 |
| 43 | DOUBLE STRANDED DNA BINDING | 128 | 764 | 1.573e-05 | 0.0003398 |
| 44 | TRANSITION METAL ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 15 | 39 | 1.628e-05 | 0.0003436 |
| 45 | ATPASE ACTIVITY COUPLED | 62 | 313 | 2.126e-05 | 0.000439 |
| 46 | AMINO ACID TRANSMEMBRANE TRANSPORTER ACTIVITY | 23 | 79 | 2.34e-05 | 0.0004725 |
| 47 | STEROID HORMONE RECEPTOR ACTIVITY | 19 | 59 | 2.496e-05 | 0.0004933 |
| 48 | NUCLEIC ACID BINDING TRANSCRIPTION FACTOR ACTIVITY | 186 | 1199 | 2.746e-05 | 0.0005315 |
| 49 | REGULATORY REGION NUCLEIC ACID BINDING | 134 | 818 | 3.052e-05 | 0.0005787 |
| 50 | METAL ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 77 | 417 | 3.129e-05 | 0.0005813 |
| 51 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 81 | 445 | 3.338e-05 | 0.000608 |
| 52 | SOLUTE SODIUM SYMPORTER ACTIVITY | 17 | 51 | 4.009e-05 | 0.0007162 |
| 53 | TRANSCRIPTION FACTOR BINDING | 92 | 524 | 4.231e-05 | 0.0007417 |
| 54 | SOLUTE CATION SYMPORTER ACTIVITY | 26 | 99 | 5.135e-05 | 0.0008834 |
| 55 | TRANSCRIPTION FACTOR ACTIVITY DIRECT LIGAND REGULATED SEQUENCE SPECIFIC DNA BINDING | 16 | 48 | 6.719e-05 | 0.001135 |
| 56 | SEQUENCE SPECIFIC DNA BINDING | 161 | 1037 | 9.159e-05 | 0.001519 |
| 57 | HISTONE KINASE ACTIVITY | 9 | 19 | 0.0001248 | 0.002033 |
| 58 | SODIUM AMINO ACID SYMPORTER ACTIVITY | 7 | 12 | 0.0001383 | 0.002215 |
| 59 | GROWTH FACTOR BINDING | 29 | 123 | 0.0001604 | 0.002525 |
| 60 | CATION TRANSMEMBRANE TRANSPORTER ACTIVITY | 103 | 622 | 0.0001631 | 0.002526 |
| 61 | KINASE INHIBITOR ACTIVITY | 23 | 89 | 0.0001768 | 0.002649 |
| 62 | CYCLIN DEPENDENT PROTEIN SERINE THREONINE KINASE REGULATOR ACTIVITY | 11 | 28 | 0.0001765 | 0.002649 |
| 63 | MODIFIED AMINO ACID TRANSMEMBRANE TRANSPORTER ACTIVITY | 8 | 16 | 0.0001877 | 0.002683 |
| 64 | CATION AMINO ACID SYMPORTER ACTIVITY | 8 | 16 | 0.0001877 | 0.002683 |
| 65 | SODIUM ION TRANSMEMBRANE TRANSPORTER ACTIVITY | 31 | 136 | 0.0001873 | 0.002683 |
| 66 | EPHRIN RECEPTOR BINDING | 10 | 24 | 0.000196 | 0.002759 |
| 67 | DICARBOXYLIC ACID TRANSMEMBRANE TRANSPORTER ACTIVITY | 12 | 33 | 0.0002138 | 0.002964 |
| 68 | INORGANIC CATION TRANSMEMBRANE TRANSPORTER ACTIVITY | 89 | 527 | 0.0002264 | 0.003093 |
| 69 | MOLECULAR FUNCTION REGULATOR | 200 | 1353 | 0.0002338 | 0.003136 |
| 70 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 45 | 226 | 0.0002391 | 0.003136 |
| 71 | FLAVIN ADENINE DINUCLEOTIDE BINDING | 20 | 74 | 0.0002397 | 0.003136 |
| 72 | PROTEIN TYROSINE KINASE ACTIVITY | 37 | 176 | 0.0002745 | 0.003541 |
| 73 | TRANSMEMBRANE RECEPTOR PROTEIN KINASE ACTIVITY | 21 | 81 | 0.0003168 | 0.003899 |
| 74 | CORE PROMOTER BINDING | 33 | 152 | 0.0003102 | 0.003899 |
| 75 | NAD DEPENDENT PROTEIN DEACETYLASE ACTIVITY | 8 | 17 | 0.0003182 | 0.003899 |
| 76 | DRUG TRANSPORTER ACTIVITY | 9 | 21 | 0.0003189 | 0.003899 |
| 77 | HISTONE DEACETYLASE BINDING | 25 | 105 | 0.0003758 | 0.004476 |
| 78 | INTEGRIN BINDING | 25 | 105 | 0.0003758 | 0.004476 |
| 79 | CARBON OXYGEN LYASE ACTIVITY | 19 | 71 | 0.0003897 | 0.004583 |
| 80 | TRANSCRIPTIONAL REPRESSOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION REGULATORY REGION SEQUENCE SPECIFIC BINDING | 35 | 168 | 0.000472 | 0.005482 |
| 81 | DNA DEPENDENT ATPASE ACTIVITY | 20 | 79 | 0.0006076 | 0.006883 |
| 82 | CHEMOREPELLENT ACTIVITY | 10 | 27 | 0.0006059 | 0.006883 |
| 83 | ENZYME REGULATOR ACTIVITY | 145 | 959 | 0.0006533 | 0.007312 |
| 84 | ATPASE ACTIVITY COUPLED TO MOVEMENT OF SUBSTANCES | 27 | 121 | 0.0006711 | 0.007422 |
| 85 | TRANSCRIPTION COACTIVATOR BINDING | 6 | 11 | 0.000699 | 0.007464 |
| 86 | ANION TRANSMEMBRANE TRANSPORTING ATPASE ACTIVITY | 6 | 11 | 0.000699 | 0.007464 |
| 87 | SEMAPHORIN RECEPTOR ACTIVITY | 6 | 11 | 0.000699 | 0.007464 |
| 88 | TRANSFERASE ACTIVITY TRANSFERRING NITROGENOUS GROUPS | 9 | 23 | 0.0007129 | 0.007526 |
| 89 | LYASE ACTIVITY | 36 | 179 | 0.0007869 | 0.008214 |
| 90 | NEUROTRANSMITTER SODIUM SYMPORTER ACTIVITY | 8 | 19 | 0.0007975 | 0.008232 |
| 91 | X3 5 CYCLIC AMP PHOSPHODIESTERASE ACTIVITY | 7 | 15 | 0.0008164 | 0.008243 |
| 92 | APOLIPOPROTEIN BINDING | 7 | 15 | 0.0008164 | 0.008243 |
| 93 | DNA HELICASE ACTIVITY | 15 | 53 | 0.0008259 | 0.00825 |
| 94 | PROTEIN DEACETYLASE ACTIVITY | 13 | 43 | 0.0009187 | 0.008984 |
| 95 | TRANSCRIPTIONAL REPRESSOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 24 | 105 | 0.0009096 | 0.008984 |
| 96 | OXIDOREDUCTASE ACTIVITY ACTING ON PAIRED DONORS WITH INCORPORATION OR REDUCTION OF MOLECULAR OXYGEN | 32 | 155 | 0.000947 | 0.009079 |
| 97 | CHROMATIN BINDING | 73 | 435 | 0.0009479 | 0.009079 |
| 98 | ACTIN BINDING | 67 | 393 | 0.0009882 | 0.009368 |
| 99 | L AMINO ACID TRANSMEMBRANE TRANSPORTER ACTIVITY | 15 | 54 | 0.001021 | 0.009585 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CHROMOSOME | 169 | 880 | 2.903e-11 | 1.316e-08 |
| 2 | NEURON PART | 224 | 1265 | 5.951e-11 | 1.316e-08 |
| 3 | NEURON PROJECTION | 176 | 942 | 1.127e-10 | 1.316e-08 |
| 4 | CONDENSED CHROMOSOME | 56 | 195 | 8.796e-11 | 1.316e-08 |
| 5 | SPINDLE | 73 | 289 | 1.063e-10 | 1.316e-08 |
| 6 | CHROMOSOME CENTROMERIC REGION | 51 | 174 | 2.708e-10 | 2.636e-08 |
| 7 | CELL PROJECTION | 292 | 1786 | 5.736e-10 | 4.785e-08 |
| 8 | MICROTUBULE CYTOSKELETON | 187 | 1068 | 6.284e-09 | 4.587e-07 |
| 9 | CHROMOSOMAL REGION | 74 | 330 | 2.189e-08 | 1.42e-06 |
| 10 | CYTOSKELETON | 307 | 1967 | 2.815e-08 | 1.644e-06 |
| 11 | SPINDLE POLE | 37 | 126 | 6.98e-08 | 3.136e-06 |
| 12 | CELL PROJECTION PART | 165 | 946 | 6.674e-08 | 3.136e-06 |
| 13 | SOMATODENDRITIC COMPARTMENT | 122 | 650 | 6.792e-08 | 3.136e-06 |
| 14 | INTRINSIC COMPONENT OF PLASMA MEMBRANE | 261 | 1649 | 1.057e-07 | 4.115e-06 |
| 15 | MICROTUBULE | 84 | 405 | 1.048e-07 | 4.115e-06 |
| 16 | MICROTUBULE ASSOCIATED COMPLEX | 40 | 145 | 1.379e-07 | 5.033e-06 |
| 17 | KINETOCHORE | 35 | 120 | 1.876e-07 | 6.445e-06 |
| 18 | MEMBRANE REGION | 189 | 1134 | 2.131e-07 | 6.914e-06 |
| 19 | PLASMA MEMBRANE REGION | 160 | 929 | 2.43e-07 | 7.095e-06 |
| 20 | DENDRITE | 90 | 451 | 2.329e-07 | 7.095e-06 |
| 21 | CONDENSED CHROMOSOME CENTROMERIC REGION | 31 | 102 | 3.388e-07 | 9.423e-06 |
| 22 | SYNAPSE | 134 | 754 | 4.088e-07 | 1.085e-05 |
| 23 | SPINDLE MICROTUBULE | 21 | 58 | 1.113e-06 | 2.825e-05 |
| 24 | CYTOSKELETAL PART | 223 | 1436 | 3.921e-06 | 9.16e-05 |
| 25 | SYNAPSE PART | 109 | 610 | 3.805e-06 | 9.16e-05 |
| 26 | APICAL PART OF CELL | 71 | 361 | 7.197e-06 | 0.0001617 |
| 27 | KINESIN COMPLEX | 19 | 55 | 7.952e-06 | 0.000172 |
| 28 | PROTEINACEOUS EXTRACELLULAR MATRIX | 70 | 356 | 8.394e-06 | 0.0001751 |
| 29 | AXON | 79 | 418 | 1.053e-05 | 0.0002121 |
| 30 | CONDENSED CHROMOSOME OUTER KINETOCHORE | 8 | 12 | 1.115e-05 | 0.000217 |
| 31 | PERINUCLEAR REGION OF CYTOPLASM | 111 | 642 | 1.438e-05 | 0.0002708 |
| 32 | MIDBODY | 33 | 132 | 1.651e-05 | 0.0003012 |
| 33 | APICAL PLASMA MEMBRANE | 59 | 292 | 1.809e-05 | 0.0003202 |
| 34 | SPINDLE MIDZONE | 12 | 27 | 2.053e-05 | 0.0003527 |
| 35 | EXTRACELLULAR MATRIX | 79 | 426 | 2.125e-05 | 0.0003547 |
| 36 | MITOTIC SPINDLE | 18 | 55 | 3.174e-05 | 0.0005149 |
| 37 | NUCLEAR CHROMOSOME | 92 | 523 | 3.931e-05 | 0.0006204 |
| 38 | MICROTUBULE ORGANIZING CENTER | 104 | 623 | 0.000111 | 0.001706 |
| 39 | EXCITATORY SYNAPSE | 41 | 197 | 0.0001666 | 0.002495 |
| 40 | POSTSYNAPSE | 68 | 378 | 0.0001965 | 0.002798 |
| 41 | MITOCHONDRION | 237 | 1633 | 0.0001939 | 0.002798 |
| 42 | PHOSPHATIDYLINOSITOL 3 KINASE COMPLEX | 9 | 20 | 0.0002033 | 0.002827 |
| 43 | CONDENSED NUCLEAR CHROMOSOME | 22 | 85 | 0.0002366 | 0.003213 |
| 44 | CENTROSOME | 83 | 487 | 0.0002688 | 0.003567 |
| 45 | AXON PART | 43 | 219 | 0.0004428 | 0.005747 |
| 46 | CHROMATIN | 75 | 441 | 0.000551 | 0.006847 |
| 47 | MEMBRANE MICRODOMAIN | 53 | 288 | 0.000545 | 0.006847 |
| 48 | MCM COMPLEX | 6 | 11 | 0.000699 | 0.007963 |
| 49 | CELL JUNCTION | 170 | 1151 | 0.0007091 | 0.007963 |
| 50 | SEMAPHORIN RECEPTOR COMPLEX | 6 | 11 | 0.000699 | 0.007963 |
| 51 | CELL BODY | 82 | 494 | 0.0006724 | 0.007963 |
| 52 | ANCHORED COMPONENT OF MEMBRANE | 32 | 152 | 0.0006681 | 0.007963 |
| 53 | BASOLATERAL PLASMA MEMBRANE | 41 | 211 | 0.0007369 | 0.00812 |
| 54 | PLASMA MEMBRANE RAFT | 21 | 86 | 0.0007515 | 0.008127 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04510_Focal_adhesion | 47 | 387 | 0.4166 | 1 | |
| 2 | hsa04010_MAPK_signaling_pathway | 46 | 387 | 0.4793 | 1 | |
| 3 | hsa04110_Cell_cycle | 40 | 387 | 0.8221 | 1 | |
| 4 | hsa04810_Regulation_of_actin_cytoskeleton | 40 | 387 | 0.8221 | 1 | |
| 5 | hsa04020_Calcium_signaling_pathway | 30 | 387 | 0.996 | 1 | |
| 6 | hsa00230_Purine_metabolism | 29 | 387 | 0.9977 | 1 | |
| 7 | hsa04360_Axon_guidance | 29 | 387 | 0.9977 | 1 | |
| 8 | hsa04012_ErbB_signaling_pathway | 27 | 387 | 0.9993 | 1 | |
| 9 | hsa04062_Chemokine_signaling_pathway | 26 | 387 | 0.9996 | 1 | |
| 10 | hsa04115_p53_signaling_pathway | 25 | 387 | 0.9998 | 1 | |
| 11 | hsa04144_Endocytosis | 24 | 387 | 0.9999 | 1 | |
| 12 | hsa00010_Glycolysis_._Gluconeogenesis | 13 | 387 | 1 | 1 | |
| 13 | hsa00020_Citrate_cycle_.TCA_cycle. | 5 | 387 | 1 | 1 | |
| 14 | hsa00030_Pentose_phosphate_pathway | 7 | 387 | 1 | 1 | |
| 15 | hsa00040_Pentose_and_glucuronate_interconversions | 5 | 387 | 1 | 1 | |
| 16 | hsa00051_Fructose_and_mannose_metabolism | 9 | 387 | 1 | 1 | |
| 17 | hsa00052_Galactose_metabolism | 4 | 387 | 1 | 1 | |
| 18 | hsa00053_Ascorbate_and_aldarate_metabolism | 5 | 387 | 1 | 1 | |
| 19 | hsa00061_Fatty_acid_biosynthesis | 3 | 387 | 1 | 1 | |
| 20 | hsa00071_Fatty_acid_metabolism | 17 | 387 | 1 | 1 | |
| 21 | hsa00100_Steroid_biosynthesis | 2 | 387 | 1 | 1 | |
| 22 | hsa00120_Primary_bile_acid_biosynthesis | 8 | 387 | 1 | 1 | |
| 23 | hsa00140_Steroid_hormone_biosynthesis | 14 | 387 | 1 | 1 | |
| 24 | hsa00190_Oxidative_phosphorylation | 7 | 387 | 1 | 1 | |
| 25 | hsa00232_Caffeine_metabolism | 2 | 387 | 1 | 1 | |
| 26 | hsa00240_Pyrimidine_metabolism | 13 | 387 | 1 | 1 | |
| 27 | hsa00250_Alanine._aspartate_and_glutamate_metabolism | 11 | 387 | 1 | 1 | |
| 28 | hsa00260_Glycine._serine_and_threonine_metabolism | 11 | 387 | 1 | 1 | |
| 29 | hsa00270_Cysteine_and_methionine_metabolism | 8 | 387 | 1 | 1 | |
| 30 | hsa00280_Valine._leucine_and_isoleucine_degradation | 14 | 387 | 1 | 1 | |
| 31 | hsa00290_Valine._leucine_and_isoleucine_biosynthesis | 3 | 387 | 1 | 1 | |
| 32 | hsa00300_Lysine_biosynthesis | 2 | 387 | 1 | 1 | |
| 33 | hsa00310_Lysine_degradation | 12 | 387 | 1 | 1 | |
| 34 | hsa00330_Arginine_and_proline_metabolism | 14 | 387 | 1 | 1 | |
| 35 | hsa00340_Histidine_metabolism | 8 | 387 | 1 | 1 | |
| 36 | hsa00350_Tyrosine_metabolism | 11 | 387 | 1 | 1 | |
| 37 | hsa00360_Phenylalanine_metabolism | 4 | 387 | 1 | 1 | |
| 38 | hsa00380_Tryptophan_metabolism | 11 | 387 | 1 | 1 | |
| 39 | hsa00400_Phenylalanine._tyrosine_and_tryptophan_biosynthesis | 3 | 387 | 1 | 1 | |
| 40 | hsa00410_beta.Alanine_metabolism | 5 | 387 | 1 | 1 | |
| 41 | hsa00430_Taurine_and_hypotaurine_metabolism | 2 | 387 | 1 | 1 | |
| 42 | hsa00450_Selenocompound_metabolism | 3 | 387 | 1 | 1 | |
| 43 | hsa00471_D.Glutamine_and_D.glutamate_metabolism | 2 | 387 | 1 | 1 | |
| 44 | hsa00480_Glutathione_metabolism | 7 | 387 | 1 | 1 | |
| 45 | hsa00500_Starch_and_sucrose_metabolism | 8 | 387 | 1 | 1 | |
| 46 | hsa00510_N.Glycan_biosynthesis | 7 | 387 | 1 | 1 | |
| 47 | hsa00511_Other_glycan_degradation | 3 | 387 | 1 | 1 | |
| 48 | hsa00512_Mucin_type_O.Glycan_biosynthesis | 4 | 387 | 1 | 1 | |
| 49 | hsa00514_Other_types_of_O.glycan_biosynthesis | 5 | 387 | 1 | 1 | |
| 50 | hsa00520_Amino_sugar_and_nucleotide_sugar_metabolism | 7 | 387 | 1 | 1 | |
| 51 | hsa00532_Glycosaminoglycan_biosynthesis_._chondroitin_sulfate | 2 | 387 | 1 | 1 | |
| 52 | hsa00533_Glycosaminoglycan_biosynthesis_._keratan_sulfate | 5 | 387 | 1 | 1 | |
| 53 | hsa00561_Glycerolipid_metabolism | 10 | 387 | 1 | 1 | |
| 54 | hsa00562_Inositol_phosphate_metabolism | 11 | 387 | 1 | 1 | |
| 55 | hsa00563_Glycosylphosphatidylinositol.GPI..anchor_biosynthesis | 5 | 387 | 1 | 1 | |
| 56 | hsa00564_Glycerophospholipid_metabolism | 9 | 387 | 1 | 1 | |
| 57 | hsa00565_Ether_lipid_metabolism | 3 | 387 | 1 | 1 | |
| 58 | hsa00590_Arachidonic_acid_metabolism | 7 | 387 | 1 | 1 | |
| 59 | hsa00591_Linoleic_acid_metabolism | 5 | 387 | 1 | 1 | |
| 60 | hsa00592_alpha.Linolenic_acid_metabolism | 2 | 387 | 1 | 1 | |
| 61 | hsa00600_Sphingolipid_metabolism | 8 | 387 | 1 | 1 | |
| 62 | hsa00601_Glycosphingolipid_biosynthesis_._lacto_and_neolacto_series | 6 | 387 | 1 | 1 | |
| 63 | hsa00604_Glycosphingolipid_biosynthesis_._ganglio_series | 4 | 387 | 1 | 1 | |
| 64 | hsa00620_Pyruvate_metabolism | 8 | 387 | 1 | 1 | |
| 65 | hsa00630_Glyoxylate_and_dicarboxylate_metabolism | 3 | 387 | 1 | 1 | |
| 66 | hsa00640_Propanoate_metabolism | 9 | 387 | 1 | 1 | |
| 67 | hsa00650_Butanoate_metabolism | 8 | 387 | 1 | 1 | |
| 68 | hsa00670_One_carbon_pool_by_folate | 5 | 387 | 1 | 1 | |
| 69 | hsa00740_Riboflavin_metabolism | 2 | 387 | 1 | 1 | |
| 70 | hsa00750_Vitamin_B6_metabolism | 2 | 387 | 1 | 1 | |
| 71 | hsa00760_Nicotinate_and_nicotinamide_metabolism | 6 | 387 | 1 | 1 | |
| 72 | hsa00770_Pantothenate_and_CoA_biosynthesis | 5 | 387 | 1 | 1 | |
| 73 | hsa00830_Retinol_metabolism | 15 | 387 | 1 | 1 | |
| 74 | hsa00860_Porphyrin_and_chlorophyll_metabolism | 7 | 387 | 1 | 1 | |
| 75 | hsa00900_Terpenoid_backbone_biosynthesis | 2 | 387 | 1 | 1 | |
| 76 | hsa00910_Nitrogen_metabolism | 8 | 387 | 1 | 1 | |
| 77 | hsa00920_Sulfur_metabolism | 2 | 387 | 1 | 1 | |
| 78 | hsa00970_Aminoacyl.tRNA_biosynthesis | 8 | 387 | 1 | 1 | |
| 79 | hsa00980_Metabolism_of_xenobiotics_by_cytochrome_P450 | 15 | 387 | 1 | 1 | |
| 80 | hsa00982_Drug_metabolism_._cytochrome_P450 | 17 | 387 | 1 | 1 | |
| 81 | hsa00983_Drug_metabolism_._other_enzymes | 11 | 387 | 1 | 1 | |
| 82 | hsa01040_Biosynthesis_of_unsaturated_fatty_acids | 3 | 387 | 1 | 1 | |
| 83 | hsa02010_ABC_transporters | 17 | 387 | 1 | 1 | |
| 84 | hsa03008_Ribosome_biogenesis_in_eukaryotes | 7 | 387 | 1 | 1 | |
| 85 | hsa03010_Ribosome | 2 | 387 | 1 | 1 | |
| 86 | hsa03013_RNA_transport | 11 | 387 | 1 | 1 | |
| 87 | hsa03015_mRNA_surveillance_pathway | 9 | 387 | 1 | 1 | |
| 88 | hsa03018_RNA_degradation | 4 | 387 | 1 | 1 | |
| 89 | hsa03020_RNA_polymerase | 2 | 387 | 1 | 1 | |
| 90 | hsa03022_Basal_transcription_factors | 2 | 387 | 1 | 1 | |
| 91 | hsa03030_DNA_replication | 9 | 387 | 1 | 1 | |
| 92 | hsa03040_Spliceosome | 6 | 387 | 1 | 1 | |
| 93 | hsa03050_Proteasome | 2 | 387 | 1 | 1 | |
| 94 | hsa03320_PPAR_signaling_pathway | 11 | 387 | 1 | 1 | |
| 95 | hsa03410_Base_excision_repair | 6 | 387 | 1 | 1 | |
| 96 | hsa03420_Nucleotide_excision_repair | 2 | 387 | 1 | 1 | |
| 97 | hsa03430_Mismatch_repair | 4 | 387 | 1 | 1 | |
| 98 | hsa03440_Homologous_recombination | 5 | 387 | 1 | 1 | |
| 99 | hsa03450_Non.homologous_end.joining | 2 | 387 | 1 | 1 | |
| 100 | hsa04070_Phosphatidylinositol_signaling_system | 18 | 387 | 1 | 1 | |
| 101 | hsa04114_Oocyte_meiosis | 23 | 387 | 1 | 1 | |
| 102 | hsa04120_Ubiquitin_mediated_proteolysis | 10 | 387 | 1 | 1 | |
| 103 | hsa04130_SNARE_interactions_in_vesicular_transport | 4 | 387 | 1 | 1 | |
| 104 | hsa04140_Regulation_of_autophagy | 2 | 387 | 1 | 1 | |
| 105 | hsa04141_Protein_processing_in_endoplasmic_reticulum | 11 | 387 | 1 | 1 | |
| 106 | hsa04142_Lysosome | 15 | 387 | 1 | 1 | |
| 107 | hsa04145_Phagosome | 11 | 387 | 1 | 1 | |
| 108 | hsa04146_Peroxisome | 18 | 387 | 1 | 1 | |
| 109 | hsa04150_mTOR_signaling_pathway | 9 | 387 | 1 | 1 | |
| 110 | hsa04210_Apoptosis | 13 | 387 | 1 | 1 | |
| 111 | hsa04260_Cardiac_muscle_contraction | 5 | 387 | 1 | 1 | |
| 112 | hsa04270_Vascular_smooth_muscle_contraction | 19 | 387 | 1 | 1 | |
| 113 | hsa04310_Wnt_signaling_pathway | 23 | 387 | 1 | 1 | |
| 114 | hsa04320_Dorso.ventral_axis_formation | 4 | 387 | 1 | 1 | |
| 115 | hsa04330_Notch_signaling_pathway | 6 | 387 | 1 | 1 | |
| 116 | hsa04340_Hedgehog_signaling_pathway | 9 | 387 | 1 | 1 | |
| 117 | hsa04350_TGF.beta_signaling_pathway | 13 | 387 | 1 | 1 | |
| 118 | hsa04370_VEGF_signaling_pathway | 18 | 387 | 1 | 1 | |
| 119 | hsa04380_Osteoclast_differentiation | 22 | 387 | 1 | 1 | |
| 120 | hsa04512_ECM.receptor_interaction | 19 | 387 | 1 | 1 | |
| 121 | hsa04514_Cell_adhesion_molecules_.CAMs. | 23 | 387 | 1 | 1 | |
| 122 | hsa04520_Adherens_junction | 13 | 387 | 1 | 1 | |
| 123 | hsa04530_Tight_junction | 23 | 387 | 1 | 1 | |
| 124 | hsa04540_Gap_junction | 17 | 387 | 1 | 1 | |
| 125 | hsa04610_Complement_and_coagulation_cascades | 15 | 387 | 1 | 1 | |
| 126 | hsa04612_Antigen_processing_and_presentation | 4 | 387 | 1 | 1 | |
| 127 | hsa04614_Renin.angiotensin_system | 4 | 387 | 1 | 1 | |
| 128 | hsa04620_Toll.like_receptor_signaling_pathway | 14 | 387 | 1 | 1 | |
| 129 | hsa04621_NOD.like_receptor_signaling_pathway | 3 | 387 | 1 | 1 | |
| 130 | hsa04622_RIG.I.like_receptor_signaling_pathway | 4 | 387 | 1 | 1 | |
| 131 | hsa04623_Cytosolic_DNA.sensing_pathway | 3 | 387 | 1 | 1 | |
| 132 | hsa04630_Jak.STAT_signaling_pathway | 19 | 387 | 1 | 1 | |
| 133 | hsa04640_Hematopoietic_cell_lineage | 15 | 387 | 1 | 1 | |
| 134 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 18 | 387 | 1 | 1 | |
| 135 | hsa04660_T_cell_receptor_signaling_pathway | 19 | 387 | 1 | 1 | |
| 136 | hsa04662_B_cell_receptor_signaling_pathway | 13 | 387 | 1 | 1 | |
| 137 | hsa04664_Fc_epsilon_RI_signaling_pathway | 18 | 387 | 1 | 1 | |
| 138 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 23 | 387 | 1 | 1 | |
| 139 | hsa04670_Leukocyte_transendothelial_migration | 22 | 387 | 1 | 1 | |
| 140 | hsa04672_Intestinal_immune_network_for_IgA_production | 4 | 387 | 1 | 1 | |
| 141 | hsa04710_Circadian_rhythm_._mammal | 6 | 387 | 1 | 1 | |
| 142 | hsa04720_Long.term_potentiation | 9 | 387 | 1 | 1 | |
| 143 | hsa04722_Neurotrophin_signaling_pathway | 22 | 387 | 1 | 1 | |
| 144 | hsa04730_Long.term_depression | 13 | 387 | 1 | 1 | |
| 145 | hsa04740_Olfactory_transduction | 3 | 388 | 1 | 1 | |
| 146 | hsa04742_Taste_transduction | 3 | 387 | 1 | 1 | |
| 147 | hsa04910_Insulin_signaling_pathway | 22 | 387 | 1 | 1 | |
| 148 | hsa04912_GnRH_signaling_pathway | 20 | 387 | 1 | 1 | |
| 149 | hsa04914_Progesterone.mediated_oocyte_maturation | 23 | 387 | 1 | 1 | |
| 150 | hsa04916_Melanogenesis | 18 | 387 | 1 | 1 | |
| 151 | hsa04920_Adipocytokine_signaling_pathway | 12 | 387 | 1 | 1 | |
| 152 | hsa04960_Aldosterone.regulated_sodium_reabsorption | 16 | 387 | 1 | 1 | |
| 153 | hsa04962_Vasopressin.regulated_water_reabsorption | 5 | 387 | 1 | 1 | |
| 154 | hsa04964_Proximal_tubule_bicarbonate_reclamation | 6 | 387 | 1 | 1 | |
| 155 | hsa04966_Collecting_duct_acid_secretion | 5 | 387 | 1 | 1 | |
| 156 | hsa04970_Salivary_secretion | 15 | 387 | 1 | 1 | |
| 157 | hsa04971_Gastric_acid_secretion | 12 | 387 | 1 | 1 | |
| 158 | hsa04972_Pancreatic_secretion | 13 | 387 | 1 | 1 | |
| 159 | hsa04973_Carbohydrate_digestion_and_absorption | 12 | 387 | 1 | 1 | |
| 160 | hsa04974_Protein_digestion_and_absorption | 14 | 387 | 1 | 1 | |
| 161 | hsa04975_Fat_digestion_and_absorption | 7 | 387 | 1 | 1 | |
| 162 | hsa04976_Bile_secretion | 20 | 387 | 1 | 1 | |
| 163 | hsa04977_Vitamin_digestion_and_absorption | 4 | 387 | 1 | 1 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | RP11-1007O24.3 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-7-5p | 14 | MDGA1 | Sponge network | 0.069 | 0.8938 | 0.083 | 0.8765 | 0.783 |
| 2 | Z83844.1 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.019 | 0.96538 | 0.083 | 0.8765 | 0.753 |
| 3 | RP3-470B24.5 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-24-3p | 10 | MDGA1 | Sponge network | 0.074 | 0.87619 | 0.083 | 0.8765 | 0.737 |
| 4 | LINC00599 | hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-376b-3p | 10 | MDGA1 | Sponge network | 0.08 | 0.87396 | 0.083 | 0.8765 | 0.715 |
| 5 | AC096559.1 |
hsa-miR-130a-3p;hsa-miR-142-5p;hsa-miR-143-3p;hsa-miR-330-3p;hsa-miR-519c-5p;hsa-miR-520a-5p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-522-3p;hsa-miR-525-5p | 10 | CENPP | Sponge network | -0.021 | 0.95521 | -0.025 | 0.95334 | 0.713 |
| 6 | AC090286.4 | hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 10 | MDGA1 | Sponge network | 0.075 | 0.84624 | 0.083 | 0.8765 | 0.71 |
| 7 | RP11-314O13.1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-301a-3p | 13 | MDGA1 | Sponge network | 0.063 | 0.89816 | 0.083 | 0.8765 | 0.71 |
| 8 | RP11-983P16.4 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-7-5p | 11 | MDGA1 | Sponge network | 0.1 | 0.8545 | 0.083 | 0.8765 | 0.701 |
| 9 | RP5-1126H10.2 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-326 | 13 | MDGA1 | Sponge network | 0.01 | 0.98441 | 0.083 | 0.8765 | 0.7 |
| 10 | LA16c-313D11.9 |
hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-24-3p;hsa-miR-25-3p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-324-3p;hsa-miR-424-5p | 10 | GPRIN2 | Sponge network | 0.053 | 0.91948 | -0.003 | 0.99411 | 0.693 |
| 11 | C10orf71-AS1 | hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-362-5p | 12 | MDGA1 | Sponge network | 0.012 | 0.97275 | 0.083 | 0.8765 | 0.692 |
| 12 | MEF2C-AS1 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-7-5p | 12 | MDGA1 | Sponge network | 0.048 | 0.91154 | 0.083 | 0.8765 | 0.692 |
| 13 | LINC00967 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-331-3p;hsa-miR-362-5p;hsa-miR-7-5p | 16 | MDGA1 | Sponge network | 0.044 | 0.91085 | 0.083 | 0.8765 | 0.692 |
| 14 | LINC01015 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-7-5p | 15 | MDGA1 | Sponge network | 0.054 | 0.88526 | 0.083 | 0.8765 | 0.689 |
| 15 | HOXA-AS3 |
hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-7-5p | 14 | MDGA1 | Sponge network | 0.072 | 0.86414 | 0.083 | 0.8765 | 0.686 |
| 16 | AC007389.3 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 11 | MDGA1 | Sponge network | 0.105 | 0.82727 | 0.083 | 0.8765 | 0.681 |
| 17 | RP11-351J23.1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 10 | MDGA1 | Sponge network | 0.048 | 0.89961 | 0.083 | 0.8765 | 0.68 |
| 18 | SDCBP2-AS1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-362-5p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-7-5p | 19 | MDGA1 | Sponge network | 0.03 | 0.94851 | 0.083 | 0.8765 | 0.678 |
| 19 | RP1-197B17.3 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-301a-3p;hsa-miR-326;hsa-miR-362-5p;hsa-miR-7-5p | 17 | MDGA1 | Sponge network | 0.04 | 0.9392 | 0.083 | 0.8765 | 0.677 |
| 20 | RP11-626H12.3 | hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.089 | 0.82141 | 0.083 | 0.8765 | 0.674 |
| 21 | RP11-168P8.3 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-301a-3p | 10 | MDGA1 | Sponge network | 0 | 0.99975 | 0.083 | 0.8765 | 0.669 |
| 22 | WWC2-AS2 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p | 11 | MDGA1 | Sponge network | 0.045 | 0.92359 | 0.083 | 0.8765 | 0.669 |
| 23 | ADORA2A-AS1 |
hsa-miR-106a-5p;hsa-miR-1323;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-512-3p;hsa-miR-515-3p;hsa-miR-516b-3p;hsa-miR-517a-3p;hsa-miR-517c-3p;hsa-miR-519b-3p;hsa-miR-519c-3p;hsa-miR-519e-3p;hsa-miR-520c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-877-5p;hsa-miR-92a-3p | 19 | NFIA | Sponge network | 0.296 | 0.58272 | 0.294 | 0.66575 | 0.668 |
| 24 | BCDIN3D-AS1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.02 | 0.96325 | 0.083 | 0.8765 | 0.666 |
| 25 | RP11-141J13.5 |
hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 11 | MDGA1 | Sponge network | 0.065 | 0.88633 | 0.083 | 0.8765 | 0.665 |
| 26 | CTD-2377D24.6 | hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.087 | 0.85124 | 0.083 | 0.8765 | 0.664 |
| 27 | AC005523.3 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-27a-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 12 | MDGA1 | Sponge network | 0.053 | 0.91591 | 0.083 | 0.8765 | 0.661 |
| 28 | AC096559.1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 12 | MDGA1 | Sponge network | -0.021 | 0.95521 | 0.083 | 0.8765 | 0.654 |
| 29 | PLAC4 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p;hsa-miR-7-5p | 12 | MDGA1 | Sponge network | 0.081 | 0.84504 | 0.083 | 0.8765 | 0.652 |
| 30 | DNAJB8-AS1 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-362-5p | 10 | MDGA1 | Sponge network | 0.093 | 0.78509 | 0.083 | 0.8765 | 0.652 |
| 31 | LINC00929 | hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.014 | 0.97083 | 0.083 | 0.8765 | 0.65 |
| 32 | RP1-197B17.3 |
hsa-miR-145-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-7-5p | 10 | ESR1 | Sponge network | 0.04 | 0.9392 | 0.121 | 0.80686 | 0.644 |
| 33 | MAP3K14-AS1 |
hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-515-5p | 10 | ESR1 | Sponge network | 0.031 | 0.93669 | 0.121 | 0.80686 | 0.644 |
| 34 | RP11-150O12.3 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-331-3p | 13 | MDGA1 | Sponge network | -0.014 | 0.97424 | 0.083 | 0.8765 | 0.64 |
| 35 | ADORA2A-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p | 11 | PPARA | Sponge network | 0.296 | 0.58272 | 0.224 | 0.72425 | 0.64 |
| 36 | RP11-793H13.3 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 13 | MDGA1 | Sponge network | 0.005 | 0.99113 | 0.083 | 0.8765 | 0.639 |
| 37 | LINC00184 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-7-5p | 12 | MDGA1 | Sponge network | 0.039 | 0.91915 | 0.083 | 0.8765 | 0.639 |
| 38 | AC106801.1 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 13 | MDGA1 | Sponge network | 0.104 | 0.83294 | 0.083 | 0.8765 | 0.639 |
| 39 | RP11-63A11.1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-362-5p | 14 | MDGA1 | Sponge network | 0.071 | 0.84677 | 0.083 | 0.8765 | 0.636 |
| 40 | RP11-251P6.1 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-223-3p;hsa-miR-362-5p;hsa-miR-376a-3p;hsa-miR-376b-3p | 11 | MDGA1 | Sponge network | 0.059 | 0.90381 | 0.083 | 0.8765 | 0.636 |
| 41 | RP11-166A12.1 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-326;hsa-miR-376a-3p;hsa-miR-376b-3p | 13 | MDGA1 | Sponge network | 0.088 | 0.84584 | 0.083 | 0.8765 | 0.635 |
| 42 | RP11-453A12.1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.096 | 0.81164 | 0.083 | 0.8765 | 0.635 |
| 43 | POLR2J4 |
hsa-miR-145-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-324-5p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-522-3p | 11 | ESR1 | Sponge network | 0.083 | 0.85528 | 0.121 | 0.80686 | 0.633 |
| 44 | PVT1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 14 | MDGA1 | Sponge network | 0.026 | 0.95534 | 0.083 | 0.8765 | 0.633 |
| 45 | KIF25-AS1 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 11 | MDGA1 | Sponge network | 0.023 | 0.948 | 0.083 | 0.8765 | 0.633 |
| 46 | LINC00208 |
hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-7-5p | 12 | ESR1 | Sponge network | 0.105 | 0.81216 | 0.121 | 0.80686 | 0.627 |
| 47 | LINC00886 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-377-3p;hsa-miR-382-5p;hsa-miR-493-5p | 17 | KLF12 | Sponge network | 0.196 | 0.7299 | 0.171 | 0.73281 | 0.626 |
| 48 | RBM26-AS1 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-188-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-376a-3p;hsa-miR-429;hsa-miR-491-5p | 13 | KLF12 | Sponge network | 0.109 | 0.80346 | 0.171 | 0.73281 | 0.623 |
| 49 | LA16c-313D11.9 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p | 11 | MDGA1 | Sponge network | 0.053 | 0.91948 | 0.083 | 0.8765 | 0.622 |
| 50 | RP11-798K3.2 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-302c-3p | 14 | HLF | Sponge network | 0.217 | 0.77181 | 0.706 | 0.36208 | 0.621 |
| 51 | LINC00642 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-326;hsa-miR-362-5p | 12 | MDGA1 | Sponge network | 0.033 | 0.93191 | 0.083 | 0.8765 | 0.619 |
| 52 | RP11-426A6.5 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-331-3p;hsa-miR-362-5p | 13 | MDGA1 | Sponge network | 0.072 | 0.89604 | 0.083 | 0.8765 | 0.619 |
| 53 | LBX2-AS1 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-491-5p | 13 | HLF | Sponge network | 0.007 | 0.99154 | 0.706 | 0.36208 | 0.617 |
| 54 | PART1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-7-5p | 16 | MDGA1 | Sponge network | -0.062 | 0.88185 | 0.083 | 0.8765 | 0.616 |
| 55 | RP3-400B16.1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 13 | MDGA1 | Sponge network | 0.061 | 0.87432 | 0.083 | 0.8765 | 0.616 |
| 56 | ADORA2A-AS1 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-223-3p;hsa-miR-23a-3p | 11 | KLF12 | Sponge network | 0.296 | 0.58272 | 0.171 | 0.73281 | 0.614 |
| 57 | RP11-280O1.2 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 13 | MDGA1 | Sponge network | 0.028 | 0.94098 | 0.083 | 0.8765 | 0.611 |
| 58 | RP11-319E16.1 | hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-223-3p;hsa-miR-24-3p | 10 | MDGA1 | Sponge network | 0.002 | 0.99707 | 0.083 | 0.8765 | 0.61 |
| 59 | AC007255.8 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-7-5p | 14 | MDGA1 | Sponge network | 0.075 | 0.85163 | 0.083 | 0.8765 | 0.61 |
| 60 | RP1-187B23.1 |
hsa-miR-17-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-515-5p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e | 10 | ESR1 | Sponge network | 0.075 | 0.87147 | 0.121 | 0.80686 | 0.607 |
| 61 | CTD-3247F14.2 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-301a-3p;hsa-miR-7-5p | 12 | MDGA1 | Sponge network | 0.104 | 0.78681 | 0.083 | 0.8765 | 0.606 |
| 62 | NAPA-AS1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-376b-3p | 13 | MDGA1 | Sponge network | 0.065 | 0.89644 | 0.083 | 0.8765 | 0.605 |
| 63 | NTRK3-AS1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p | 11 | MDGA1 | Sponge network | 0.031 | 0.95002 | 0.083 | 0.8765 | 0.604 |
| 64 | LINC00652 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-223-3p;hsa-miR-24-3p | 12 | MDGA1 | Sponge network | 0.024 | 0.96591 | 0.083 | 0.8765 | 0.602 |
| 65 | LINC00208 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-7-5p | 16 | MDGA1 | Sponge network | 0.105 | 0.81216 | 0.083 | 0.8765 | 0.601 |
| 66 | LINC00886 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-302c-3p;hsa-miR-377-3p | 20 | HLF | Sponge network | 0.196 | 0.7299 | 0.706 | 0.36208 | 0.6 |
| 67 | RP11-798K3.2 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p | 11 | PPARA | Sponge network | 0.217 | 0.77181 | 0.224 | 0.72425 | 0.599 |
| 68 | RP11-416N2.4 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 11 | MDGA1 | Sponge network | 0.011 | 0.97606 | 0.083 | 0.8765 | 0.598 |
| 69 | RP11-847H18.2 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 14 | MDGA1 | Sponge network | 0.112 | 0.81863 | 0.083 | 0.8765 | 0.598 |
| 70 | MIR205HG |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 12 | MDGA1 | Sponge network | 0.025 | 0.94278 | 0.083 | 0.8765 | 0.598 |
| 71 | RP11-805I24.3 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 12 | MDGA1 | Sponge network | -0.009 | 0.9868 | 0.083 | 0.8765 | 0.595 |
| 72 | LINC00642 |
hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-25-3p;hsa-miR-27a-3p;hsa-miR-324-3p;hsa-miR-342-3p;hsa-miR-424-5p;hsa-miR-503-5p | 10 | GPRIN2 | Sponge network | 0.033 | 0.93191 | -0.003 | 0.99411 | 0.594 |
| 73 | RP11-304C12.3 |
hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-362-5p;hsa-miR-7-5p | 13 | MDGA1 | Sponge network | 0.058 | 0.89219 | 0.083 | 0.8765 | 0.592 |
| 74 | ZNF503-AS2 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p | 11 | MDGA1 | Sponge network | -0.049 | 0.92396 | 0.083 | 0.8765 | 0.589 |
| 75 | RP11-63B19.1 | hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-362-5p;hsa-miR-7-5p | 10 | MDGA1 | Sponge network | 0.078 | 0.86406 | 0.083 | 0.8765 | 0.588 |
| 76 | LINC00238 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p | 10 | TTC9 | Sponge network | 0.096 | 0.75184 | 0.309 | 0.51342 | 0.587 |
| 77 | LINC00208 |
hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-25-3p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-324-3p;hsa-miR-424-5p;hsa-miR-503-5p;hsa-miR-93-5p | 15 | GPRIN2 | Sponge network | 0.105 | 0.81216 | -0.003 | 0.99411 | 0.586 |
| 78 | AC003090.1 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-376c-3p;hsa-miR-382-5p | 13 | KLF12 | Sponge network | -0.005 | 0.99146 | 0.171 | 0.73281 | 0.585 |
| 79 | AC109828.1 | hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-25-3p;hsa-miR-27a-3p;hsa-miR-424-5p;hsa-miR-503-5p;hsa-miR-93-5p | 12 | GPRIN2 | Sponge network | 0.009 | 0.98192 | -0.003 | 0.99411 | 0.584 |
| 80 | LBX2-AS1 |
hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-491-5p | 10 | KLF12 | Sponge network | 0.007 | 0.99154 | 0.171 | 0.73281 | 0.582 |
| 81 | RP11-8L2.1 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 11 | MDGA1 | Sponge network | -0.005 | 0.9887 | 0.083 | 0.8765 | 0.576 |
| 82 | LINC00700 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 12 | MDGA1 | Sponge network | 0.09 | 0.78132 | 0.083 | 0.8765 | 0.576 |
| 83 | AC012487.2 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 11 | MDGA1 | Sponge network | 0.064 | 0.89114 | 0.083 | 0.8765 | 0.573 |
| 84 | AC068138.1 |
hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 14 | MDGA1 | Sponge network | 0.06 | 0.8855 | 0.083 | 0.8765 | 0.571 |
| 85 | LINC00886 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-331-3p | 10 | SLC1A2 | Sponge network | 0.196 | 0.7299 | 0.329 | 0.56983 | 0.571 |
| 86 | SMIM2-AS1 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-125a-5p;hsa-miR-1323;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-224-5p;hsa-miR-375;hsa-miR-485-5p;hsa-miR-519c-5p;hsa-miR-7-5p | 15 | SLC4A4 | Sponge network | 0.194 | 0.64908 | 0.236 | 0.64507 | 0.57 |
| 87 | PRKAG2-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-212-3p;hsa-miR-223-3p;hsa-miR-299-3p;hsa-miR-301a-3p;hsa-miR-485-5p | 10 | AR | Sponge network | 0.284 | 0.58395 | 0.661 | 0.388 | 0.57 |
| 88 | LINC00559 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-331-3p;hsa-miR-362-5p | 17 | MDGA1 | Sponge network | 0.04 | 0.91022 | 0.083 | 0.8765 | 0.57 |
| 89 | RP11-620J15.2 | hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-362-5p;hsa-miR-7-5p | 10 | MDGA1 | Sponge network | -0.017 | 0.96861 | 0.083 | 0.8765 | 0.57 |
| 90 | RP11-449P15.2 |
hsa-miR-17-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18b-5p;hsa-miR-324-5p;hsa-miR-515-5p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-522-3p | 10 | ESR1 | Sponge network | 0.046 | 0.92254 | 0.121 | 0.80686 | 0.569 |
| 91 | RP11-375O18.2 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 10 | MDGA1 | Sponge network | 0.036 | 0.92297 | 0.083 | 0.8765 | 0.569 |
| 92 | RP3-333B15.4 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-7-5p | 14 | MDGA1 | Sponge network | 0.032 | 0.93813 | 0.083 | 0.8765 | 0.568 |
| 93 | RP11-65L19.4 |
hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-324-5p;hsa-miR-520d-3p | 10 | ESR1 | Sponge network | 0.056 | 0.92382 | 0.121 | 0.80686 | 0.567 |
| 94 | RP11-545I5.3 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-376c-3p;hsa-miR-377-3p;hsa-miR-382-5p;hsa-miR-429;hsa-miR-491-5p | 18 | KLF12 | Sponge network | 0.103 | 0.76843 | 0.171 | 0.73281 | 0.564 |
| 95 | AC034220.3 |
hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-324-5p | 10 | ESR1 | Sponge network | 0.029 | 0.95595 | 0.121 | 0.80686 | 0.563 |
| 96 | AC012487.2 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-218-5p | 11 | HLF | Sponge network | 0.064 | 0.89114 | 0.706 | 0.36208 | 0.562 |
| 97 | MIR205HG |
hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-324-3p;hsa-miR-93-5p | 10 | GPRIN2 | Sponge network | 0.025 | 0.94278 | -0.003 | 0.99411 | 0.562 |
| 98 | OSER1-AS1 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-218-5p | 11 | HLF | Sponge network | 0.081 | 0.86429 | 0.706 | 0.36208 | 0.562 |
| 99 | RP11-320N7.2 |
hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-218-5p | 10 | HLF | Sponge network | 0.397 | 0.4155 | 0.706 | 0.36208 | 0.561 |
| 100 | POLR2J4 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p | 12 | MDGA1 | Sponge network | 0.083 | 0.85528 | 0.083 | 0.8765 | 0.56 |
| 101 | XXyac-YX155B6.5 | hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-362-5p | 10 | MDGA1 | Sponge network | 0.025 | 0.94233 | 0.083 | 0.8765 | 0.559 |
| 102 | MTUS2-AS1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-7-5p | 12 | MDGA1 | Sponge network | 0.038 | 0.93568 | 0.083 | 0.8765 | 0.558 |
| 103 | CTD-2201E18.3 |
hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-515-5p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-7-5p | 10 | ESR1 | Sponge network | 0.121 | 0.85821 | 0.121 | 0.80686 | 0.557 |
| 104 | RP11-1006G14.1 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p | 14 | MDGA1 | Sponge network | 0.059 | 0.8921 | 0.083 | 0.8765 | 0.557 |
| 105 | RP4-781K5.4 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p;hsa-miR-376a-3p;hsa-miR-376b-3p | 12 | MDGA1 | Sponge network | 0.005 | 0.98885 | 0.083 | 0.8765 | 0.554 |
| 106 | RP11-397A16.3 | hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-24-3p;hsa-miR-27a-3p | 11 | MDGA1 | Sponge network | 0.023 | 0.94586 | 0.083 | 0.8765 | 0.553 |
| 107 | ADORA2A-AS1 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-212-3p;hsa-miR-223-3p;hsa-miR-877-5p | 12 | AR | Sponge network | 0.296 | 0.58272 | 0.661 | 0.388 | 0.553 |
| 108 | RP11-453A12.1 |
hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-25-3p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-324-3p | 12 | GPRIN2 | Sponge network | 0.096 | 0.81164 | -0.003 | 0.99411 | 0.552 |
| 109 | SMIM2-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-302c-3p;hsa-miR-485-5p;hsa-miR-877-5p | 12 | AR | Sponge network | 0.194 | 0.64908 | 0.661 | 0.388 | 0.552 |
| 110 | PLAC4 |
hsa-miR-106b-5p;hsa-miR-18b-5p;hsa-miR-24-3p;hsa-miR-25-3p;hsa-miR-27a-3p;hsa-miR-324-3p;hsa-miR-342-3p;hsa-miR-424-5p;hsa-miR-503-5p;hsa-miR-93-5p | 10 | GPRIN2 | Sponge network | 0.081 | 0.84504 | -0.003 | 0.99411 | 0.548 |
| 111 | LINC00261 | hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181b-5p;hsa-miR-20b-5p;hsa-miR-212-3p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-877-5p | 10 | AR | Sponge network | 0.464 | 0.56742 | 0.661 | 0.388 | 0.548 |
| 112 | RP11-304C12.3 |
hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-301a-3p;hsa-miR-7-5p | 10 | ESR1 | Sponge network | 0.058 | 0.89219 | 0.121 | 0.80686 | 0.547 |
| 113 | MAP3K14-AS1 |
hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-324-3p;hsa-miR-342-3p;hsa-miR-424-5p;hsa-miR-503-5p | 13 | GPRIN2 | Sponge network | 0.031 | 0.93669 | -0.003 | 0.99411 | 0.545 |
| 114 | RP11-798K3.2 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-20b-5p;hsa-miR-212-3p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-302c-3p | 13 | AR | Sponge network | 0.217 | 0.77181 | 0.661 | 0.388 | 0.543 |
| 115 | CTD-2366F13.1 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-218-5p | 12 | HLF | Sponge network | 0.092 | 0.81374 | 0.706 | 0.36208 | 0.542 |
| 116 | RP13-616I3.1 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-301a-3p | 10 | MDGA1 | Sponge network | 0.149 | 0.75223 | 0.083 | 0.8765 | 0.542 |
| 117 | LINC00460 | hsa-let-7b-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | -0.043 | 0.92258 | 0.083 | 0.8765 | 0.539 |
| 118 | RP11-407B7.1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-7-5p | 12 | MDGA1 | Sponge network | 0.009 | 0.97909 | 0.083 | 0.8765 | 0.537 |
| 119 | HOXA-AS2 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-218-5p;hsa-miR-27a-3p;hsa-miR-377-3p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e | 10 | CNTNAP2 | Sponge network | -0.046 | 0.91915 | -0.008 | 0.98698 | 0.537 |
| 120 | ADORA2A-AS1 |
hsa-miR-106a-5p;hsa-miR-1323;hsa-miR-17-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-519b-3p;hsa-miR-519d-3p;hsa-miR-92a-3p | 10 | RORA | Sponge network | 0.296 | 0.58272 | 0.521 | 0.41408 | 0.536 |
| 121 | THAP7-AS1 |
hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-324-5p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-7-5p;hsa-miR-9-5p | 12 | ESR1 | Sponge network | -0.058 | 0.89073 | 0.121 | 0.80686 | 0.535 |
| 122 | AP000253.1 |
hsa-miR-132-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-376c-3p | 10 | KLF12 | Sponge network | 0.27 | 0.48487 | 0.171 | 0.73281 | 0.533 |
| 123 | TRAM2-AS1 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-369-3p;hsa-miR-375 | 12 | KLF12 | Sponge network | 0.066 | 0.90989 | 0.171 | 0.73281 | 0.531 |
| 124 | TMEM254-AS1 |
hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-331-3p | 12 | MDGA1 | Sponge network | 0.03 | 0.94824 | 0.083 | 0.8765 | 0.53 |
| 125 | RP11-613D13.8 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-331-3p;hsa-miR-7-5p | 15 | MDGA1 | Sponge network | -0.059 | 0.88272 | 0.083 | 0.8765 | 0.53 |
| 126 | ADORA2A-AS1 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-200c-3p | 11 | HLF | Sponge network | 0.296 | 0.58272 | 0.706 | 0.36208 | 0.529 |
| 127 | RP11-24D15.1 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-301a-3p | 10 | MDGA1 | Sponge network | 0.11 | 0.75766 | 0.083 | 0.8765 | 0.527 |
| 128 | RP11-177H13.2 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 11 | MDGA1 | Sponge network | -0.017 | 0.96779 | 0.083 | 0.8765 | 0.527 |
| 129 | SMIM2-AS1 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-188-5p;hsa-miR-199b-5p;hsa-miR-301a-3p;hsa-miR-375;hsa-miR-376c-3p;hsa-miR-491-5p | 10 | KLF12 | Sponge network | 0.194 | 0.64908 | 0.171 | 0.73281 | 0.525 |
| 130 | ALDH1L1-AS2 |
hsa-miR-145-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-515-5p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-7-5p;hsa-miR-9-5p | 12 | ESR1 | Sponge network | 0.044 | 0.93158 | 0.121 | 0.80686 | 0.524 |
| 131 | ZBED5-AS1 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-363-3p;hsa-miR-367-3p;hsa-miR-376c-3p | 11 | SLC1A2 | Sponge network | 0.166 | 0.80245 | 0.329 | 0.56983 | 0.523 |
| 132 | SMIM2-AS1 |
hsa-miR-106a-5p;hsa-miR-1323;hsa-miR-142-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-512-3p;hsa-miR-516b-3p;hsa-miR-519b-3p;hsa-miR-520c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-877-5p | 14 | NFIA | Sponge network | 0.194 | 0.64908 | 0.294 | 0.66575 | 0.52 |
| 133 | AP000696.2 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 11 | MDGA1 | Sponge network | 0.064 | 0.8886 | 0.083 | 0.8765 | 0.518 |
| 134 | RP4-794H19.1 |
hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-320a;hsa-miR-424-5p;hsa-miR-491-5p | 11 | SCN2A | Sponge network | 0.028 | 0.9507 | -0.073 | 0.8182 | 0.517 |
| 135 | RP11-319G6.1 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 11 | MDGA1 | Sponge network | 0.019 | 0.97302 | 0.083 | 0.8765 | 0.517 |
| 136 | RP11-588H23.3 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p | 13 | MDGA1 | Sponge network | 0.009 | 0.98308 | 0.083 | 0.8765 | 0.516 |
| 137 | LINC00208 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-221-3p;hsa-miR-222-3p;hsa-miR-27a-3p | 10 | STX1B | Sponge network | 0.105 | 0.81216 | 0.074 | 0.87501 | 0.515 |
| 138 | RP11-736K20.6 |
hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p | 11 | MDGA1 | Sponge network | 0.069 | 0.87221 | 0.083 | 0.8765 | 0.514 |
| 139 | LINC00886 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-299-3p;hsa-miR-301a-3p;hsa-miR-302c-3p;hsa-miR-493-5p;hsa-miR-877-5p | 17 | AR | Sponge network | 0.196 | 0.7299 | 0.661 | 0.388 | 0.513 |
| 140 | RP11-798K3.2 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-223-3p;hsa-miR-375 | 11 | SLC4A4 | Sponge network | 0.217 | 0.77181 | 0.236 | 0.64507 | 0.509 |
| 141 | TRAM2-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-375;hsa-miR-494-3p | 11 | SLC4A4 | Sponge network | 0.066 | 0.90989 | 0.236 | 0.64507 | 0.508 |
| 142 | RP11-6N17.4 |
hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-377-3p | 12 | HLF | Sponge network | 0.195 | 0.73464 | 0.706 | 0.36208 | 0.507 |
| 143 | TMEM254-AS1 |
hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-320a;hsa-miR-491-5p | 10 | SCN2A | Sponge network | 0.03 | 0.94824 | -0.073 | 0.8182 | 0.505 |
| 144 | LINC00559 |
hsa-miR-106b-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-24-3p;hsa-miR-25-3p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-335-5p;hsa-miR-342-3p;hsa-miR-424-5p;hsa-miR-93-5p | 12 | GPRIN2 | Sponge network | 0.04 | 0.91022 | -0.003 | 0.99411 | 0.505 |
| 145 | ZBED5-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-302c-3p | 13 | AR | Sponge network | 0.166 | 0.80245 | 0.661 | 0.388 | 0.5 |
| 146 | PTOV1-AS1 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-92a-3p | 11 | PRKCA | Sponge network | -0.074 | 0.84862 | -0.093 | 0.86855 | 0.499 |
| 147 | RP11-34A14.3 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-223-3p;hsa-miR-24-3p | 10 | MDGA1 | Sponge network | 0.044 | 0.91589 | 0.083 | 0.8765 | 0.499 |
| 148 | SMIM2-AS1 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-302c-3p;hsa-miR-491-5p | 17 | HLF | Sponge network | 0.194 | 0.64908 | 0.706 | 0.36208 | 0.498 |
| 149 | POLR2J4 |
hsa-miR-1323;hsa-miR-142-5p;hsa-miR-143-3p;hsa-miR-520a-5p;hsa-miR-520d-3p;hsa-miR-520d-5p;hsa-miR-520e;hsa-miR-522-3p;hsa-miR-524-5p;hsa-miR-525-5p | 10 | CENPP | Sponge network | 0.083 | 0.85528 | -0.025 | 0.95334 | 0.498 |
| 150 | MAP3K14-AS1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 13 | MDGA1 | Sponge network | 0.031 | 0.93669 | 0.083 | 0.8765 | 0.497 |
| 151 | RP11-244O19.1 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-331-3p;hsa-miR-362-5p | 12 | MDGA1 | Sponge network | -0.01 | 0.98445 | 0.083 | 0.8765 | 0.497 |
| 152 | RP11-798K3.2 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-375 | 12 | KLF12 | Sponge network | 0.217 | 0.77181 | 0.171 | 0.73281 | 0.497 |
| 153 | AC078842.4 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.062 | 0.84089 | 0.083 | 0.8765 | 0.493 |
| 154 | LINC00242 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-301a-3p;hsa-miR-7-5p | 11 | MDGA1 | Sponge network | 0.057 | 0.88871 | 0.083 | 0.8765 | 0.493 |
| 155 | RP11-280O1.2 |
hsa-miR-17-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-342-3p;hsa-miR-424-5p;hsa-miR-93-5p | 11 | GPRIN2 | Sponge network | 0.028 | 0.94098 | -0.003 | 0.99411 | 0.492 |
| 156 | HOXA-AS2 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-331-3p | 12 | MDGA1 | Sponge network | -0.046 | 0.91915 | 0.083 | 0.8765 | 0.492 |
| 157 | CCDC13-AS1 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.035 | 0.92276 | 0.083 | 0.8765 | 0.492 |
| 158 | THAP7-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-512-3p;hsa-miR-515-3p;hsa-miR-516b-3p;hsa-miR-517c-3p;hsa-miR-519b-3p;hsa-miR-519c-3p;hsa-miR-519e-3p;hsa-miR-520c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-9-5p;hsa-miR-92a-3p | 20 | NFIA | Sponge network | -0.058 | 0.89073 | 0.294 | 0.66575 | 0.491 |
| 159 | STK4-AS1 | hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-7-5p | 11 | MDGA1 | Sponge network | -0.01 | 0.98497 | 0.083 | 0.8765 | 0.491 |
| 160 | ZBED5-AS1 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-302c-3p;hsa-miR-491-5p | 20 | HLF | Sponge network | 0.166 | 0.80245 | 0.706 | 0.36208 | 0.49 |
| 161 | RP11-276H19.2 |
hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-192-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-30a-5p;hsa-miR-30e-5p | 12 | NEBL | Sponge network | -0.077 | 0.83993 | -0.126 | 0.74294 | 0.489 |
| 162 | LINC00886 |
hsa-miR-106a-5p;hsa-miR-145-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-23a-3p | 10 | GFRA1 | Sponge network | 0.196 | 0.7299 | 0.161 | 0.75691 | 0.489 |
| 163 | LINC00910 | hsa-let-7b-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-331-3p | 10 | MDGA1 | Sponge network | -0.017 | 0.95793 | 0.083 | 0.8765 | 0.489 |
| 164 | AC012507.3 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 12 | MDGA1 | Sponge network | -0.021 | 0.96252 | 0.083 | 0.8765 | 0.488 |
| 165 | MTUS2-AS1 |
hsa-miR-145-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-515-5p;hsa-miR-522-3p;hsa-miR-7-5p | 10 | ESR1 | Sponge network | 0.038 | 0.93568 | 0.121 | 0.80686 | 0.487 |
| 166 | WEE2-AS1 |
hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-424-5p;hsa-miR-491-5p | 10 | SCN2A | Sponge network | -0.042 | 0.91109 | -0.073 | 0.8182 | 0.484 |
| 167 | RP11-847H18.2 |
hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-324-3p;hsa-miR-424-5p;hsa-miR-93-5p | 11 | GPRIN2 | Sponge network | 0.112 | 0.81863 | -0.003 | 0.99411 | 0.483 |
| 168 | LINC00559 |
hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-424-5p;hsa-miR-491-5p | 10 | SCN2A | Sponge network | 0.04 | 0.91022 | -0.073 | 0.8182 | 0.482 |
| 169 | MAP3K14-AS1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-222-3p;hsa-miR-27a-3p;hsa-miR-342-3p;hsa-miR-342-5p | 11 | STX1B | Sponge network | 0.031 | 0.93669 | 0.074 | 0.87501 | 0.481 |
| 170 | AP000253.1 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-141-3p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-299-3p;hsa-miR-485-5p;hsa-miR-877-5p | 13 | AR | Sponge network | 0.27 | 0.48487 | 0.661 | 0.388 | 0.481 |
| 171 | AP000253.1 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-500a-3p | 14 | HLF | Sponge network | 0.27 | 0.48487 | 0.706 | 0.36208 | 0.48 |
| 172 | DICER1-AS1 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-376a-3p;hsa-miR-491-5p | 11 | KLF12 | Sponge network | 0.089 | 0.85424 | 0.171 | 0.73281 | 0.479 |
| 173 | RP5-968P14.2 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-377-3p;hsa-miR-491-5p | 12 | HLF | Sponge network | 0.045 | 0.87863 | 0.706 | 0.36208 | 0.479 |
| 174 | RP11-6N17.4 |
hsa-let-7e-5p;hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 10 | SLC4A4 | Sponge network | 0.195 | 0.73464 | 0.236 | 0.64507 | 0.477 |
| 175 | LINC00886 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-223-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-376b-3p | 13 | SLC4A4 | Sponge network | 0.196 | 0.7299 | 0.236 | 0.64507 | 0.477 |
| 176 | ZBED5-AS1 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-369-3p;hsa-miR-376c-3p;hsa-miR-491-5p | 13 | KLF12 | Sponge network | 0.166 | 0.80245 | 0.171 | 0.73281 | 0.474 |
| 177 | AP000253.1 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p | 10 | PPARA | Sponge network | 0.27 | 0.48487 | 0.224 | 0.72425 | 0.47 |
| 178 | RP11-448A19.1 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p | 10 | MDGA1 | Sponge network | 0.064 | 0.88097 | 0.083 | 0.8765 | 0.468 |
| 179 | ALDH1L1-AS2 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p | 10 | PRKCA | Sponge network | 0.044 | 0.93158 | -0.093 | 0.86855 | 0.468 |
| 180 | DPH6-AS1 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-301a-3p;hsa-miR-376c-3p | 11 | KLF12 | Sponge network | 0.046 | 0.88414 | 0.171 | 0.73281 | 0.468 |
| 181 | RP4-647C14.3 | hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-376b-3p | 10 | MDGA1 | Sponge network | 0.063 | 0.88387 | 0.083 | 0.8765 | 0.467 |
| 182 | RP11-300A12.2 | hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.108 | 0.76999 | 0.083 | 0.8765 | 0.466 |
| 183 | RP11-458F8.4 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-376a-3p;hsa-miR-491-5p | 10 | KLF12 | Sponge network | 0.135 | 0.76373 | 0.171 | 0.73281 | 0.466 |
| 184 | PTOV1-AS1 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-376c-3p;hsa-miR-382-5p | 13 | KLF12 | Sponge network | -0.074 | 0.84862 | 0.171 | 0.73281 | 0.465 |
| 185 | UBAC2-AS1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-331-3p | 12 | MDGA1 | Sponge network | 0 | 0.9996 | 0.083 | 0.8765 | 0.465 |
| 186 | RP11-696N14.1 | hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | -0.016 | 0.9557 | 0.083 | 0.8765 | 0.464 |
| 187 | RP11-46C24.7 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199a-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-301a-3p;hsa-miR-369-3p;hsa-miR-375 | 11 | KLF12 | Sponge network | -0.039 | 0.92939 | 0.171 | 0.73281 | 0.463 |
| 188 | AC008703.1 | hsa-let-7b-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | -0.032 | 0.9323 | 0.083 | 0.8765 | 0.462 |
| 189 | RP11-401P9.5 |
hsa-miR-106a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-494-3p | 10 | SLC1A2 | Sponge network | 0.033 | 0.92794 | 0.329 | 0.56983 | 0.462 |
| 190 | RP11-328N19.1 |
hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30a-5p | 10 | NEBL | Sponge network | -0.044 | 0.89095 | -0.126 | 0.74294 | 0.462 |
| 191 | ALDH1L1-AS2 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-7-5p | 14 | MDGA1 | Sponge network | 0.044 | 0.93158 | 0.083 | 0.8765 | 0.46 |
| 192 | LINC00526 | hsa-let-7e-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-218-5p | 12 | HLF | Sponge network | -0.04 | 0.94499 | 0.706 | 0.36208 | 0.459 |
| 193 | SPTY2D1-AS1 |
hsa-miR-106b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-25-3p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-324-3p;hsa-miR-93-5p | 10 | GPRIN2 | Sponge network | -0.012 | 0.97196 | -0.003 | 0.99411 | 0.458 |
| 194 | ALDH1L1-AS2 |
hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-377-3p;hsa-miR-491-5p | 14 | HLF | Sponge network | 0.044 | 0.93158 | 0.706 | 0.36208 | 0.457 |
| 195 | RP11-545I5.3 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-877-5p | 11 | AR | Sponge network | 0.103 | 0.76843 | 0.661 | 0.388 | 0.455 |
| 196 | TRAM2-AS1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-200c-3p;hsa-miR-218-5p | 13 | HLF | Sponge network | 0.066 | 0.90989 | 0.706 | 0.36208 | 0.454 |
| 197 | ATP6V0E2-AS1 |
hsa-miR-106a-5p;hsa-miR-139-5p;hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-424-5p;hsa-miR-491-5p | 13 | SCN2A | Sponge network | -0.095 | 0.8128 | -0.073 | 0.8182 | 0.454 |
| 198 | RBM26-AS1 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-429;hsa-miR-491-5p;hsa-miR-500a-3p | 21 | HLF | Sponge network | 0.109 | 0.80346 | 0.706 | 0.36208 | 0.452 |
| 199 | DPH6-AS1 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-218-5p | 10 | HLF | Sponge network | 0.046 | 0.88414 | 0.706 | 0.36208 | 0.452 |
| 200 | KB-431C1.4 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-miR-101-3p;hsa-miR-139-5p;hsa-miR-195-5p;hsa-miR-26b-5p;hsa-miR-27a-3p;hsa-miR-27b-3p | 10 | ZBTB10 | Sponge network | -0.017 | 0.97318 | 0.057 | 0.91732 | 0.452 |
| 201 | AFAP1-AS1 |
hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-144-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-16-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30e-5p | 12 | NEBL | Sponge network | -0.118 | 0.77941 | -0.126 | 0.74294 | 0.451 |
| 202 | LBX2-AS1 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-23a-3p | 10 | GFRA1 | Sponge network | 0.007 | 0.99154 | 0.161 | 0.75691 | 0.449 |
| 203 | RP11-545I5.3 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-342-3p;hsa-miR-376b-3p;hsa-miR-429 | 13 | PPARA | Sponge network | 0.103 | 0.76843 | 0.224 | 0.72425 | 0.449 |
| 204 | ADORA2A-AS1 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-1323;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-519c-3p;hsa-miR-519d-3p | 13 | SLC4A4 | Sponge network | 0.296 | 0.58272 | 0.236 | 0.64507 | 0.442 |
| 205 | RP11-492E3.2 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 11 | MDGA1 | Sponge network | 0.082 | 0.89032 | 0.083 | 0.8765 | 0.442 |
| 206 | RP11-873E20.1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-424-5p | 10 | SCN2A | Sponge network | 0.019 | 0.95215 | -0.073 | 0.8182 | 0.442 |
| 207 | TPTEP1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-197-3p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-7-5p | 11 | MDGA1 | Sponge network | 0.044 | 0.90203 | 0.083 | 0.8765 | 0.441 |
| 208 | LINC00238 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-224-5p;hsa-miR-342-3p | 12 | DENND5B | Sponge network | 0.096 | 0.75184 | 0.008 | 0.98644 | 0.44 |
| 209 | ZBED5-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-363-3p;hsa-miR-369-3p;hsa-miR-9-5p | 13 | PPARA | Sponge network | 0.166 | 0.80245 | 0.224 | 0.72425 | 0.438 |
| 210 | PTOV1-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-363-3p | 12 | PPARA | Sponge network | -0.074 | 0.84862 | 0.224 | 0.72425 | 0.438 |
| 211 | PTOV1-AS1 |
hsa-let-7d-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p | 17 | HLF | Sponge network | -0.074 | 0.84862 | 0.706 | 0.36208 | 0.436 |
| 212 | THAP7-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-519c-3p;hsa-miR-7-5p | 11 | SLC4A4 | Sponge network | -0.058 | 0.89073 | 0.236 | 0.64507 | 0.436 |
| 213 | ALDH1L1-AS2 |
hsa-let-7e-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-519c-3p;hsa-miR-7-5p | 11 | SLC4A4 | Sponge network | 0.044 | 0.93158 | 0.236 | 0.64507 | 0.434 |
| 214 | THAP7-AS1 |
hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-7-5p | 10 | MDGA1 | Sponge network | -0.058 | 0.89073 | 0.083 | 0.8765 | 0.433 |
| 215 | TRAM2-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-223-3p;hsa-miR-299-3p | 11 | AR | Sponge network | 0.066 | 0.90989 | 0.661 | 0.388 | 0.432 |
| 216 | AP000253.1 |
hsa-miR-106a-5p;hsa-miR-1323;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-224-5p;hsa-miR-483-3p;hsa-miR-485-5p | 12 | SLC4A4 | Sponge network | 0.27 | 0.48487 | 0.236 | 0.64507 | 0.432 |
| 217 | ZBTB11-AS1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-24-3p;hsa-miR-331-3p | 11 | MDGA1 | Sponge network | -0.049 | 0.90441 | 0.083 | 0.8765 | 0.432 |
| 218 | LINC00886 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-222-3p;hsa-miR-27a-3p;hsa-miR-342-5p | 10 | STX1B | Sponge network | 0.196 | 0.7299 | 0.074 | 0.87501 | 0.431 |
| 219 | PRKAG2-AS1 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-331-3p | 10 | SLC1A2 | Sponge network | 0.284 | 0.58395 | 0.329 | 0.56983 | 0.43 |
| 220 | RP11-874J12.4 |
hsa-miR-126-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-192-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30a-5p | 11 | NEBL | Sponge network | -0.062 | 0.84559 | -0.126 | 0.74294 | 0.429 |
| 221 | AC016700.5 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-214-3p;hsa-miR-223-3p;hsa-miR-7-5p | 10 | MDGA1 | Sponge network | -0.006 | 0.98822 | 0.083 | 0.8765 | 0.429 |
| 222 | CTB-131B5.5 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-376c-3p;hsa-miR-377-3p;hsa-miR-382-5p | 12 | KLF12 | Sponge network | 0.216 | 0.59133 | 0.171 | 0.73281 | 0.428 |
| 223 | RP11-829H16.3 | hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-223-3p;hsa-miR-24-3p | 10 | MDGA1 | Sponge network | -0.019 | 0.94705 | 0.083 | 0.8765 | 0.428 |
| 224 | RP11-27N21.3 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 12 | MDGA1 | Sponge network | -0.047 | 0.90233 | 0.083 | 0.8765 | 0.425 |
| 225 | RP11-522B15.3 | hsa-miR-1323;hsa-miR-512-3p;hsa-miR-519b-3p;hsa-miR-519c-3p;hsa-miR-520c-3p;hsa-miR-520d-3p;hsa-miR-520d-5p;hsa-miR-520e;hsa-miR-524-5p;hsa-miR-9-5p | 10 | NFIA | Sponge network | 0.031 | 0.93373 | 0.294 | 0.66575 | 0.423 |
| 226 | ALDH1L1-AS2 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-376c-3p;hsa-miR-377-3p;hsa-miR-491-5p | 13 | KLF12 | Sponge network | 0.044 | 0.93158 | 0.171 | 0.73281 | 0.42 |
| 227 | THAP7-AS1 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-369-3p;hsa-miR-376a-3p;hsa-miR-376c-3p;hsa-miR-377-3p;hsa-miR-491-5p;hsa-miR-493-5p | 14 | KLF12 | Sponge network | -0.058 | 0.89073 | 0.171 | 0.73281 | 0.418 |
| 228 | AC003090.1 |
hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-218-5p | 10 | HLF | Sponge network | -0.005 | 0.99146 | 0.706 | 0.36208 | 0.418 |
| 229 | SNHG16 |
hsa-miR-181a-5p;hsa-miR-192-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p | 10 | SLC7A11 | Sponge network | -0.137 | 0.87069 | -0.564 | 0.28432 | 0.418 |
| 230 | RP11-874J12.4 |
hsa-miR-122-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-181a-5p;hsa-miR-192-5p;hsa-miR-215-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27a-3p;hsa-miR-27b-3p | 10 | SLC7A11 | Sponge network | -0.062 | 0.84559 | -0.564 | 0.28432 | 0.416 |
| 231 | AC016700.5 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-20b-5p | 12 | HLF | Sponge network | -0.006 | 0.98822 | 0.706 | 0.36208 | 0.415 |
| 232 | LINC00886 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 18 | MDGA1 | Sponge network | 0.196 | 0.7299 | 0.083 | 0.8765 | 0.414 |
| 233 | USP46-AS1 | hsa-let-7b-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-199a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p | 10 | HLF | Sponge network | -0.106 | 0.76346 | 0.706 | 0.36208 | 0.413 |
| 234 | RP11-6N17.4 |
hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-342-3p;hsa-miR-369-3p;hsa-miR-376b-3p | 10 | PPARA | Sponge network | 0.195 | 0.73464 | 0.224 | 0.72425 | 0.413 |
| 235 | RP11-401P9.5 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-377-3p | 19 | HLF | Sponge network | 0.033 | 0.92794 | 0.706 | 0.36208 | 0.411 |
| 236 | PTOV1-AS1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-92a-3p | 12 | DENND5B | Sponge network | -0.074 | 0.84862 | 0.008 | 0.98644 | 0.41 |
| 237 | RP11-61I13.3 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-223-3p | 10 | MDGA1 | Sponge network | 0.004 | 0.98985 | 0.083 | 0.8765 | 0.41 |
| 238 | RP11-1006G14.1 |
hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-301a-3p | 10 | KLF12 | Sponge network | 0.059 | 0.8921 | 0.171 | 0.73281 | 0.408 |
| 239 | RP11-390E23.6 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-376c-3p;hsa-miR-494-3p | 10 | SLC1A2 | Sponge network | 0.037 | 0.93955 | 0.329 | 0.56983 | 0.408 |
| 240 | DICER1-AS1 |
hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-491-5p | 12 | HLF | Sponge network | 0.089 | 0.85424 | 0.706 | 0.36208 | 0.408 |
| 241 | THAP7-AS1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-377-3p;hsa-miR-491-5p | 15 | HLF | Sponge network | -0.058 | 0.89073 | 0.706 | 0.36208 | 0.408 |
| 242 | CTD-3193O13.1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p | 10 | MDGA1 | Sponge network | 0.047 | 0.90548 | 0.083 | 0.8765 | 0.407 |
| 243 | RP5-991G20.1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 13 | MDGA1 | Sponge network | 0.039 | 0.93103 | 0.083 | 0.8765 | 0.405 |
| 244 | RP11-734K2.4 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-362-5p;hsa-miR-376b-3p;hsa-miR-7-5p | 16 | MDGA1 | Sponge network | 0.094 | 0.83958 | 0.083 | 0.8765 | 0.404 |
| 245 | RP11-545I5.3 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-517c-3p;hsa-miR-519b-3p;hsa-miR-520d-3p;hsa-miR-877-5p;hsa-miR-92a-3p | 11 | NFIA | Sponge network | 0.103 | 0.76843 | 0.294 | 0.66575 | 0.404 |
| 246 | AP000472.2 | hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-20a-5p | 10 | HLF | Sponge network | 0.172 | 0.66101 | 0.706 | 0.36208 | 0.404 |
| 247 | RP11-736K20.6 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p | 10 | PPARA | Sponge network | 0.069 | 0.87221 | 0.224 | 0.72425 | 0.403 |
| 248 | LINC00886 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-375 | 10 | PRKCA | Sponge network | 0.196 | 0.7299 | -0.093 | 0.86855 | 0.401 |
| 249 | AC079354.3 | hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-362-5p | 10 | MDGA1 | Sponge network | 0.005 | 0.9876 | 0.083 | 0.8765 | 0.399 |
| 250 | LINC00238 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p | 15 | HLF | Sponge network | 0.096 | 0.75184 | 0.706 | 0.36208 | 0.397 |
| 251 | RP11-379H18.1 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-21-5p;hsa-miR-212-3p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-375 | 10 | KLF12 | Sponge network | 0.109 | 0.81097 | 0.171 | 0.73281 | 0.397 |
| 252 | LINC00930 | hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-515-5p;hsa-miR-516b-3p;hsa-miR-517c-3p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520d-5p;hsa-miR-520e | 11 | NFIA | Sponge network | -0.004 | 0.99459 | 0.294 | 0.66575 | 0.397 |
| 253 | THAP7-AS1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-410-3p;hsa-miR-519b-3p;hsa-miR-9-5p;hsa-miR-92a-3p | 10 | RORA | Sponge network | -0.058 | 0.89073 | 0.521 | 0.41408 | 0.396 |
| 254 | HOXA-AS2 |
hsa-let-7e-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-375;hsa-miR-519c-3p;hsa-miR-520d-5p | 11 | SLC4A4 | Sponge network | -0.046 | 0.91915 | 0.236 | 0.64507 | 0.395 |
| 255 | RP11-379H18.1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p | 12 | HLF | Sponge network | 0.109 | 0.81097 | 0.706 | 0.36208 | 0.395 |
| 256 | ZFAS1 | hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-144-3p;hsa-miR-15a-5p;hsa-miR-195-5p;hsa-miR-26b-5p;hsa-miR-29a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-30a-5p;hsa-miR-497-5p | 11 | PLAG1 | Sponge network | -0.014 | 0.98731 | -0.227 | 0.6194 | 0.394 |
| 257 | LINC00565 |
hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-362-5p;hsa-miR-376a-3p;hsa-miR-376b-3p | 12 | MDGA1 | Sponge network | -0.031 | 0.93622 | 0.083 | 0.8765 | 0.393 |
| 258 | RP11-545I5.3 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-376c-3p;hsa-miR-429;hsa-miR-494-3p;hsa-miR-92a-3p | 12 | SLC1A2 | Sponge network | 0.103 | 0.76843 | 0.329 | 0.56983 | 0.393 |
| 259 | SACS-AS1 | hsa-let-7b-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-24-3p | 10 | MDGA1 | Sponge network | -0.002 | 0.99513 | 0.083 | 0.8765 | 0.392 |
| 260 | RBM26-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-429;hsa-miR-877-5p | 10 | NFIA | Sponge network | 0.109 | 0.80346 | 0.294 | 0.66575 | 0.39 |
| 261 | RBM26-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-429;hsa-miR-877-5p | 13 | AR | Sponge network | 0.109 | 0.80346 | 0.661 | 0.388 | 0.389 |
| 262 | AC068138.1 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-301a-3p;hsa-miR-369-3p;hsa-miR-376a-3p;hsa-miR-382-5p | 15 | KLF12 | Sponge network | 0.06 | 0.8855 | 0.171 | 0.73281 | 0.389 |
| 263 | ZNF503-AS2 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p | 18 | HLF | Sponge network | -0.049 | 0.92396 | 0.706 | 0.36208 | 0.389 |
| 264 | LINC00238 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p | 11 | AR | Sponge network | 0.096 | 0.75184 | 0.661 | 0.388 | 0.389 |
| 265 | DICER1-AS1 |
hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-331-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 12 | MDGA1 | Sponge network | 0.089 | 0.85424 | 0.083 | 0.8765 | 0.388 |
| 266 | RP11-280O1.2 |
hsa-miR-126-5p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-19b-3p;hsa-miR-320a;hsa-miR-424-5p | 10 | ZFHX4 | Sponge network | 0.028 | 0.94098 | 0.088 | 0.8617 | 0.388 |
| 267 | RP11-545I5.3 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-377-3p;hsa-miR-429;hsa-miR-491-5p | 17 | HLF | Sponge network | 0.103 | 0.76843 | 0.706 | 0.36208 | 0.386 |
| 268 | RP11-6F2.5 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p | 17 | HLF | Sponge network | 0.253 | 0.44204 | 0.706 | 0.36208 | 0.386 |
| 269 | PSMG3-AS1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-214-3p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-7-5p | 12 | MDGA1 | Sponge network | -0.033 | 0.95037 | 0.083 | 0.8765 | 0.386 |
| 270 | TMEM254-AS1 |
hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-491-5p | 12 | HLF | Sponge network | 0.03 | 0.94824 | 0.706 | 0.36208 | 0.386 |
| 271 | ZBED5-AS1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p | 11 | DENND5B | Sponge network | 0.166 | 0.80245 | 0.008 | 0.98644 | 0.385 |
| 272 | RP11-873E20.1 |
hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 13 | MDGA1 | Sponge network | 0.019 | 0.95215 | 0.083 | 0.8765 | 0.384 |
| 273 | THAP7-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-363-3p;hsa-miR-369-3p;hsa-miR-376b-3p;hsa-miR-9-5p | 15 | PPARA | Sponge network | -0.058 | 0.89073 | 0.224 | 0.72425 | 0.383 |
| 274 | RP11-46C24.7 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-500a-3p | 14 | HLF | Sponge network | -0.039 | 0.92939 | 0.706 | 0.36208 | 0.383 |
| 275 | PTPRG-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-494-3p | 10 | SLC4A4 | Sponge network | 0.009 | 0.97688 | 0.236 | 0.64507 | 0.382 |
| 276 | LAMTOR5-AS1 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p | 10 | PRKCA | Sponge network | 0.011 | 0.9823 | -0.093 | 0.86855 | 0.382 |
| 277 | RP11-545I5.3 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-429;hsa-miR-494-3p | 16 | SLC4A4 | Sponge network | 0.103 | 0.76843 | 0.236 | 0.64507 | 0.382 |
| 278 | RP11-274B18.4 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 10 | MDGA1 | Sponge network | 0.09 | 0.79671 | 0.083 | 0.8765 | 0.381 |
| 279 | RP11-350G8.5 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p | 10 | MDGA1 | Sponge network | 0.059 | 0.84655 | 0.083 | 0.8765 | 0.381 |
| 280 | PTOV1-AS1 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-363-3p;hsa-miR-376c-3p;hsa-miR-92a-3p;hsa-miR-96-5p | 13 | SLC1A2 | Sponge network | -0.074 | 0.84862 | 0.329 | 0.56983 | 0.381 |
| 281 | RP11-736K20.6 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-376c-3p | 12 | KLF12 | Sponge network | 0.069 | 0.87221 | 0.171 | 0.73281 | 0.38 |
| 282 | RAD21-AS1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 11 | MDGA1 | Sponge network | 0.081 | 0.81401 | 0.083 | 0.8765 | 0.38 |
| 283 | RP11-1081M5.2 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-214-3p;hsa-miR-223-3p;hsa-miR-24-3p | 12 | MDGA1 | Sponge network | 0.065 | 0.84871 | 0.083 | 0.8765 | 0.38 |
| 284 | LINC00853 |
hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-377-3p;hsa-miR-382-5p | 10 | KLF12 | Sponge network | 0.101 | 0.80817 | 0.171 | 0.73281 | 0.379 |
| 285 | RP11-400K9.4 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-491-5p | 12 | KLF12 | Sponge network | 0.242 | 0.49109 | 0.171 | 0.73281 | 0.378 |
| 286 | AC091729.9 | hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-377-3p;hsa-miR-491-5p | 11 | HLF | Sponge network | 0.138 | 0.78179 | 0.706 | 0.36208 | 0.376 |
| 287 | ZNF503-AS2 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-494-3p | 10 | SLC4A4 | Sponge network | -0.049 | 0.92396 | 0.236 | 0.64507 | 0.375 |
| 288 | RP11-27N21.3 |
hsa-miR-139-5p;hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-424-5p | 10 | SCN2A | Sponge network | -0.047 | 0.90233 | -0.073 | 0.8182 | 0.375 |
| 289 | AC007255.8 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-494-3p;hsa-miR-7-5p | 13 | SLC4A4 | Sponge network | 0.075 | 0.85163 | 0.236 | 0.64507 | 0.375 |
| 290 | LINC00884 |
hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-342-3p;hsa-miR-376b-3p | 11 | PPARA | Sponge network | 0.137 | 0.76561 | 0.224 | 0.72425 | 0.374 |
| 291 | AC005523.3 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-376b-3p | 10 | PPARA | Sponge network | 0.053 | 0.91591 | 0.224 | 0.72425 | 0.374 |
| 292 | RP11-6N17.4 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-369-3p;hsa-miR-376a-3p;hsa-miR-376c-3p;hsa-miR-377-3p;hsa-miR-493-5p | 12 | KLF12 | Sponge network | 0.195 | 0.73464 | 0.171 | 0.73281 | 0.372 |
| 293 | RP11-390E23.6 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-299-3p;hsa-miR-301a-3p | 11 | AR | Sponge network | 0.037 | 0.93955 | 0.661 | 0.388 | 0.372 |
| 294 | AC016700.5 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-494-3p;hsa-miR-7-5p | 10 | SLC4A4 | Sponge network | -0.006 | 0.98822 | 0.236 | 0.64507 | 0.372 |
| 295 | RP11-552M11.8 | hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-223-3p;hsa-miR-7-5p | 10 | MDGA1 | Sponge network | -0.009 | 0.97886 | 0.083 | 0.8765 | 0.371 |
| 296 | ZNF503-AS2 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-382-5p | 10 | KLF12 | Sponge network | -0.049 | 0.92396 | 0.171 | 0.73281 | 0.371 |
| 297 | ADORA2A-AS1 |
hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-522-3p | 11 | ESR1 | Sponge network | 0.296 | 0.58272 | 0.121 | 0.80686 | 0.37 |
| 298 | TRAM2-AS1 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-369-3p;hsa-miR-9-5p | 12 | PPARA | Sponge network | 0.066 | 0.90989 | 0.224 | 0.72425 | 0.369 |
| 299 | ADORA2A-AS1 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-92a-3p | 10 | PRKCA | Sponge network | 0.296 | 0.58272 | -0.093 | 0.86855 | 0.369 |
| 300 | ALDH1L1-AS2 |
hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-9-5p | 10 | PPARA | Sponge network | 0.044 | 0.93158 | 0.224 | 0.72425 | 0.369 |
| 301 | LINC00174 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-382-5p;hsa-miR-493-5p | 11 | KLF12 | Sponge network | 0.123 | 0.81603 | 0.171 | 0.73281 | 0.368 |
| 302 | RP11-798K3.2 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-375 | 10 | PRKCA | Sponge network | 0.217 | 0.77181 | -0.093 | 0.86855 | 0.368 |
| 303 | RP11-1081M5.2 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199a-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-377-3p;hsa-miR-382-5p | 13 | KLF12 | Sponge network | 0.065 | 0.84871 | 0.171 | 0.73281 | 0.368 |
| 304 | RP11-847H18.2 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-222-3p;hsa-miR-27a-3p;hsa-miR-342-5p | 10 | STX1B | Sponge network | 0.112 | 0.81863 | 0.074 | 0.87501 | 0.366 |
| 305 | AC005682.5 | hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-192-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-30a-5p;hsa-miR-32-5p | 11 | NEBL | Sponge network | -0.124 | 0.71336 | -0.126 | 0.74294 | 0.365 |
| 306 | THAP7-AS1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-493-5p | 14 | AR | Sponge network | -0.058 | 0.89073 | 0.661 | 0.388 | 0.365 |
| 307 | ALDH1L1-AS2 |
hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-424-5p;hsa-miR-491-5p | 10 | SCN2A | Sponge network | 0.044 | 0.93158 | -0.073 | 0.8182 | 0.365 |
| 308 | ALDH1L1-AS2 |
hsa-miR-142-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-512-3p;hsa-miR-515-5p;hsa-miR-519b-3p;hsa-miR-519c-3p;hsa-miR-520c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-9-5p | 11 | NFIA | Sponge network | 0.044 | 0.93158 | 0.294 | 0.66575 | 0.364 |
| 309 | ALDH1L1-AS2 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-218-5p;hsa-miR-27a-3p;hsa-miR-377-3p;hsa-miR-519c-3p;hsa-miR-520c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-526b-5p | 12 | CNTNAP2 | Sponge network | 0.044 | 0.93158 | -0.008 | 0.98698 | 0.363 |
| 310 | RP11-401P9.5 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-494-3p | 14 | SLC4A4 | Sponge network | 0.033 | 0.92794 | 0.236 | 0.64507 | 0.363 |
| 311 | LINC00667 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-363-3p;hsa-miR-429 | 10 | SLC1A2 | Sponge network | -0.166 | 0.75617 | 0.329 | 0.56983 | 0.363 |
| 312 | RP11-16P6.1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 12 | MDGA1 | Sponge network | -0.003 | 0.99304 | 0.083 | 0.8765 | 0.362 |
| 313 | RBM26-AS1 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-429 | 12 | SLC4A4 | Sponge network | 0.109 | 0.80346 | 0.236 | 0.64507 | 0.362 |
| 314 | RP11-379H18.1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-212-3p;hsa-miR-223-3p;hsa-miR-877-5p | 15 | AR | Sponge network | 0.109 | 0.81097 | 0.661 | 0.388 | 0.361 |
| 315 | LINC00511 |
hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-30c-5p;hsa-miR-30e-5p;hsa-miR-335-5p | 11 | NEBL | Sponge network | -0.067 | 0.87356 | -0.126 | 0.74294 | 0.36 |
| 316 | ZBED5-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-363-3p | 10 | TTC9 | Sponge network | 0.166 | 0.80245 | 0.309 | 0.51342 | 0.359 |
| 317 | RP11-545I5.3 |
hsa-miR-106a-5p;hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-23a-3p;hsa-miR-376c-3p | 10 | GFRA1 | Sponge network | 0.103 | 0.76843 | 0.161 | 0.75691 | 0.359 |
| 318 | RP11-736K20.5 | hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-223-3p;hsa-miR-301a-3p | 10 | AR | Sponge network | 0.162 | 0.72458 | 0.661 | 0.388 | 0.358 |
| 319 | HOXA-AS2 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-375 | 10 | PRKCA | Sponge network | -0.046 | 0.91915 | -0.093 | 0.86855 | 0.358 |
| 320 | LINC00886 |
hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-301a-3p | 10 | ESR1 | Sponge network | 0.196 | 0.7299 | 0.121 | 0.80686 | 0.357 |
| 321 | ATP6V0E2-AS1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-331-3p | 13 | MDGA1 | Sponge network | -0.095 | 0.8128 | 0.083 | 0.8765 | 0.356 |
| 322 | ZBED5-AS1 |
hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-301a-3p;hsa-miR-9-5p | 10 | ESR1 | Sponge network | 0.166 | 0.80245 | 0.121 | 0.80686 | 0.355 |
| 323 | ADORA2A-AS1 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-212-3p;hsa-miR-214-3p;hsa-miR-23a-3p | 10 | GFRA1 | Sponge network | 0.296 | 0.58272 | 0.161 | 0.75691 | 0.355 |
| 324 | SMIM2-AS1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-199b-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-224-5p;hsa-miR-342-3p | 10 | DENND5B | Sponge network | 0.194 | 0.64908 | 0.008 | 0.98644 | 0.354 |
| 325 | RP11-400K9.4 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-491-5p | 17 | HLF | Sponge network | 0.242 | 0.49109 | 0.706 | 0.36208 | 0.354 |
| 326 | RP11-492E3.2 |
hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-144-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-32-5p | 12 | NEBL | Sponge network | 0.082 | 0.89032 | -0.126 | 0.74294 | 0.353 |
| 327 | LINC00667 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-429 | 12 | KLF12 | Sponge network | -0.166 | 0.75617 | 0.171 | 0.73281 | 0.352 |
| 328 | RP11-736K20.6 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p | 13 | AR | Sponge network | 0.069 | 0.87221 | 0.661 | 0.388 | 0.35 |
| 329 | RP11-401P9.5 |
hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-301a-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 15 | MDGA1 | Sponge network | 0.033 | 0.92794 | 0.083 | 0.8765 | 0.35 |
| 330 | RP11-620J15.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-miR-195-5p;hsa-miR-26b-5p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-30a-5p;hsa-miR-424-5p;hsa-miR-497-5p | 11 | ZBTB10 | Sponge network | -0.266 | 0.60278 | 0.057 | 0.91732 | 0.349 |
| 331 | LINC00559 |
hsa-miR-130a-3p;hsa-miR-143-3p;hsa-miR-145-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-217;hsa-miR-28-5p | 10 | SNX27 | Sponge network | 0.04 | 0.91022 | -0.226 | 0.73456 | 0.349 |
| 332 | PTOV1-AS1 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p | 14 | AR | Sponge network | -0.074 | 0.84862 | 0.661 | 0.388 | 0.349 |
| 333 | CTD-2619J13.17 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 12 | MDGA1 | Sponge network | 0.034 | 0.93595 | 0.083 | 0.8765 | 0.348 |
| 334 | ENTPD1-AS1 |
hsa-miR-106a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-512-3p;hsa-miR-515-3p;hsa-miR-515-5p;hsa-miR-516b-3p;hsa-miR-519c-3p;hsa-miR-519e-3p;hsa-miR-520d-3p;hsa-miR-520d-5p;hsa-miR-520e;hsa-miR-92a-3p | 14 | NFIA | Sponge network | 0.012 | 0.9694 | 0.294 | 0.66575 | 0.348 |
| 335 | RP11-95D17.1 |
hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p | 10 | NEBL | Sponge network | -0.032 | 0.94444 | -0.126 | 0.74294 | 0.347 |
| 336 | LINC00667 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p | 10 | DENND5B | Sponge network | -0.166 | 0.75617 | 0.008 | 0.98644 | 0.347 |
| 337 | RP11-401P9.5 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-301a-3p | 11 | AR | Sponge network | 0.033 | 0.92794 | 0.661 | 0.388 | 0.346 |
| 338 | RP11-401P9.5 |
hsa-miR-106a-5p;hsa-miR-145-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-23a-3p | 10 | GFRA1 | Sponge network | 0.033 | 0.92794 | 0.161 | 0.75691 | 0.344 |
| 339 | RP1-228H13.5 |
hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-144-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-30a-5p;hsa-miR-30e-5p | 11 | NEBL | Sponge network | -0.086 | 0.84203 | -0.126 | 0.74294 | 0.344 |
| 340 | RP11-401P9.5 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-342-3p | 11 | DENND5B | Sponge network | 0.033 | 0.92794 | 0.008 | 0.98644 | 0.344 |
| 341 | RP5-968P14.2 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p | 10 | PPARA | Sponge network | 0.045 | 0.87863 | 0.224 | 0.72425 | 0.343 |
| 342 | AC003090.1 |
hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-331-3p | 11 | MDGA1 | Sponge network | -0.005 | 0.99146 | 0.083 | 0.8765 | 0.343 |
| 343 | RP11-1006G14.1 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p | 11 | PPARA | Sponge network | 0.059 | 0.8921 | 0.224 | 0.72425 | 0.342 |
| 344 | SNHG6 |
hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-144-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-192-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30a-5p | 12 | NEBL | Sponge network | -0.227 | 0.81331 | -0.126 | 0.74294 | 0.342 |
| 345 | LINC00667 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-429 | 21 | HLF | Sponge network | -0.166 | 0.75617 | 0.706 | 0.36208 | 0.342 |
| 346 | LINC00667 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-429 | 12 | SLC4A4 | Sponge network | -0.166 | 0.75617 | 0.236 | 0.64507 | 0.342 |
| 347 | LINC00494 |
hsa-miR-126-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-195-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30c-5p;hsa-miR-335-5p | 10 | NEBL | Sponge network | -0.103 | 0.79947 | -0.126 | 0.74294 | 0.341 |
| 348 | LINC00886 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-223-3p | 10 | DENND5B | Sponge network | 0.196 | 0.7299 | 0.008 | 0.98644 | 0.341 |
| 349 | RP5-1157M23.2 |
hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30a-5p | 12 | NEBL | Sponge network | -0.074 | 0.87445 | -0.126 | 0.74294 | 0.34 |
| 350 | AC005523.3 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-491-5p | 13 | HLF | Sponge network | 0.053 | 0.91591 | 0.706 | 0.36208 | 0.34 |
| 351 | DCTN1-AS1 | hsa-let-7b-5p;hsa-let-7d-5p;hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p | 12 | HLF | Sponge network | 0.007 | 0.97862 | 0.706 | 0.36208 | 0.338 |
| 352 | HOXA-AS2 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-375;hsa-miR-376c-3p;hsa-miR-377-3p;hsa-miR-382-5p;hsa-miR-491-5p | 14 | KLF12 | Sponge network | -0.046 | 0.91915 | 0.171 | 0.73281 | 0.337 |
| 353 | LINC00339 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-218-5p | 10 | HLF | Sponge network | 0 | 0.99996 | 0.706 | 0.36208 | 0.337 |
| 354 | LINC00886 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-376b-3p | 12 | PPARA | Sponge network | 0.196 | 0.7299 | 0.224 | 0.72425 | 0.337 |
| 355 | DANCR |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-miR-101-3p;hsa-miR-133b;hsa-miR-195-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30a-5p | 10 | ZBTB10 | Sponge network | -0.092 | 0.90786 | 0.057 | 0.91732 | 0.336 |
| 356 | SNHG6 |
hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-15a-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30a-5p | 10 | PLAG1 | Sponge network | -0.227 | 0.81331 | -0.227 | 0.6194 | 0.336 |
| 357 | LINC00559 |
hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-150-5p;hsa-miR-195-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-27a-3p;hsa-miR-342-3p;hsa-miR-424-5p;hsa-miR-491-5p;hsa-miR-497-5p | 11 | SLC22A23 | Sponge network | 0.04 | 0.91022 | -0.03 | 0.95804 | 0.336 |
| 358 | LAMTOR5-AS1 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-377-3p;hsa-miR-429;hsa-miR-491-5p | 16 | HLF | Sponge network | 0.011 | 0.9823 | 0.706 | 0.36208 | 0.335 |
| 359 | RP11-873E20.1 |
hsa-miR-130a-3p;hsa-miR-143-3p;hsa-miR-145-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-193a-3p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-217;hsa-miR-376a-3p | 10 | SNX27 | Sponge network | 0.019 | 0.95215 | -0.226 | 0.73456 | 0.335 |
| 360 | RP11-399O19.9 |
hsa-let-7d-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-20b-5p | 14 | HLF | Sponge network | 0.032 | 0.9362 | 0.706 | 0.36208 | 0.335 |
| 361 | RBM26-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-320a;hsa-miR-491-5p | 12 | SCN2A | Sponge network | 0.109 | 0.80346 | -0.073 | 0.8182 | 0.335 |
| 362 | LINC00668 |
hsa-miR-126-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-30a-5p;hsa-miR-30e-5p | 10 | NEBL | Sponge network | -0.186 | 0.70101 | -0.126 | 0.74294 | 0.335 |
| 363 | LAMTOR5-AS1 |
hsa-let-7e-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-429;hsa-miR-494-3p | 13 | SLC4A4 | Sponge network | 0.011 | 0.9823 | 0.236 | 0.64507 | 0.335 |
| 364 | RP11-390E23.6 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-375;hsa-miR-494-3p | 10 | SLC4A4 | Sponge network | 0.037 | 0.93955 | 0.236 | 0.64507 | 0.333 |
| 365 | ST7-AS1 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-369-3p;hsa-miR-376c-3p;hsa-miR-491-5p | 11 | KLF12 | Sponge network | -0.009 | 0.98084 | 0.171 | 0.73281 | 0.33 |
| 366 | RP11-6F2.5 |
hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 11 | MDGA1 | Sponge network | 0.253 | 0.44204 | 0.083 | 0.8765 | 0.329 |
| 367 | AC007255.8 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200c-3p;hsa-miR-20b-5p | 16 | HLF | Sponge network | 0.075 | 0.85163 | 0.706 | 0.36208 | 0.329 |
| 368 | AC068138.1 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-301a-3p | 10 | AR | Sponge network | 0.06 | 0.8855 | 0.661 | 0.388 | 0.327 |
| 369 | RP11-873E20.1 |
hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-424-5p | 10 | GPRIN2 | Sponge network | 0.019 | 0.95215 | -0.003 | 0.99411 | 0.326 |
| 370 | RGMB-AS1 | hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-500a-3p | 10 | HLF | Sponge network | 0.168 | 0.64302 | 0.706 | 0.36208 | 0.326 |
| 371 | RP11-401P9.5 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-301a-3p;hsa-miR-369-3p;hsa-miR-376a-3p;hsa-miR-377-3p | 14 | KLF12 | Sponge network | 0.033 | 0.92794 | 0.171 | 0.73281 | 0.325 |
| 372 | RP11-798K3.2 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-223-3p | 10 | DENND5B | Sponge network | 0.217 | 0.77181 | 0.008 | 0.98644 | 0.324 |
| 373 | LINC00174 |
hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-299-3p;hsa-miR-493-5p;hsa-miR-877-5p | 11 | AR | Sponge network | 0.123 | 0.81603 | 0.661 | 0.388 | 0.324 |
| 374 | ALDH1L1-AS2 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-23a-3p;hsa-miR-514a-3p;hsa-miR-520e;hsa-miR-9-5p | 10 | SLC22A3 | Sponge network | 0.044 | 0.93158 | 0.276 | 0.68622 | 0.323 |
| 375 | AC007255.8 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-494-3p | 10 | SLC1A2 | Sponge network | 0.075 | 0.85163 | 0.329 | 0.56983 | 0.322 |
| 376 | LINC00094 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-369-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-376c-3p;hsa-miR-377-3p;hsa-miR-429;hsa-miR-491-5p | 12 | KLF12 | Sponge network | -0.246 | 0.62122 | 0.171 | 0.73281 | 0.32 |
| 377 | LINC00559 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-375 | 10 | PRKCA | Sponge network | 0.04 | 0.91022 | -0.093 | 0.86855 | 0.32 |
| 378 | LINC00238 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-342-3p | 10 | PPARA | Sponge network | 0.096 | 0.75184 | 0.224 | 0.72425 | 0.318 |
| 379 | RP11-347C12.10 | hsa-let-7d-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-302c-3p | 10 | HLF | Sponge network | 0.039 | 0.90964 | 0.706 | 0.36208 | 0.317 |
| 380 | RP11-375I20.6 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-145-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-23a-3p | 11 | GFRA1 | Sponge network | 0.009 | 0.9807 | 0.161 | 0.75691 | 0.316 |
| 381 | SDCBP2-AS1 |
hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-342-3p;hsa-miR-369-3p;hsa-miR-376b-3p | 13 | PPARA | Sponge network | 0.03 | 0.94851 | 0.224 | 0.72425 | 0.316 |
| 382 | CTB-131B5.5 |
hsa-miR-106a-5p;hsa-miR-1323;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-483-3p;hsa-miR-519d-3p;hsa-miR-524-5p | 14 | SLC4A4 | Sponge network | 0.216 | 0.59133 | 0.236 | 0.64507 | 0.316 |
| 383 | RP11-6F2.5 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-376a-3p | 10 | KLF12 | Sponge network | 0.253 | 0.44204 | 0.171 | 0.73281 | 0.315 |
| 384 | RP11-400K9.4 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-494-3p;hsa-miR-7-5p | 13 | SLC4A4 | Sponge network | 0.242 | 0.49109 | 0.236 | 0.64507 | 0.314 |
| 385 | RP11-6N17.4 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p | 10 | PRKCA | Sponge network | 0.195 | 0.73464 | -0.093 | 0.86855 | 0.313 |
| 386 | LINC00208 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-301a-3p | 16 | AR | Sponge network | 0.105 | 0.81216 | 0.661 | 0.388 | 0.31 |
| 387 | RP11-46C24.7 |
hsa-let-7e-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-375 | 10 | SLC4A4 | Sponge network | -0.039 | 0.92939 | 0.236 | 0.64507 | 0.31 |
| 388 | PART1 |
hsa-miR-142-3p;hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-424-5p;hsa-miR-491-5p | 10 | SCN2A | Sponge network | -0.062 | 0.88185 | -0.073 | 0.8182 | 0.309 |
| 389 | TOB1-AS1 |
hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200b-3p;hsa-miR-200c-3p | 11 | HLF | Sponge network | 0.01 | 0.98241 | 0.706 | 0.36208 | 0.309 |
| 390 | AC007255.8 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-301a-3p | 14 | AR | Sponge network | 0.075 | 0.85163 | 0.661 | 0.388 | 0.308 |
| 391 | WWC2-AS2 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-491-5p | 12 | HLF | Sponge network | 0.045 | 0.92359 | 0.706 | 0.36208 | 0.308 |
| 392 | RP11-448A19.1 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-376c-3p;hsa-miR-377-3p;hsa-miR-493-5p | 11 | KLF12 | Sponge network | 0.064 | 0.88097 | 0.171 | 0.73281 | 0.307 |
| 393 | LINC00667 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-373-3p;hsa-miR-429 | 15 | AR | Sponge network | -0.166 | 0.75617 | 0.661 | 0.388 | 0.306 |
| 394 | CTB-131B5.5 |
hsa-miR-106a-5p;hsa-miR-1323;hsa-miR-17-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-519b-3p;hsa-miR-519d-3p;hsa-miR-524-5p | 10 | RORA | Sponge network | 0.216 | 0.59133 | 0.521 | 0.41408 | 0.306 |
| 395 | RP11-257I14.1 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-493-5p | 13 | AR | Sponge network | -0.027 | 0.91947 | 0.661 | 0.388 | 0.305 |
| 396 | AC114730.3 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p | 13 | MDGA1 | Sponge network | 0.051 | 0.88172 | 0.083 | 0.8765 | 0.304 |
| 397 | RAD21-AS1 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-377-3p | 12 | HLF | Sponge network | 0.081 | 0.81401 | 0.706 | 0.36208 | 0.303 |
| 398 | RP11-399O19.9 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p | 11 | AR | Sponge network | 0.032 | 0.9362 | 0.661 | 0.388 | 0.303 |
| 399 | RP11-448A19.1 |
hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-493-5p | 10 | AR | Sponge network | 0.064 | 0.88097 | 0.661 | 0.388 | 0.303 |
| 400 | SDCBP2-AS1 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-301a-3p;hsa-miR-362-5p;hsa-miR-369-3p;hsa-miR-376a-3p;hsa-miR-376c-3p;hsa-miR-377-3p;hsa-miR-382-5p;hsa-miR-493-5p | 17 | KLF12 | Sponge network | 0.03 | 0.94851 | 0.171 | 0.73281 | 0.302 |
| 401 | RP11-379H18.1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-376b-3p | 12 | PPARA | Sponge network | 0.109 | 0.81097 | 0.224 | 0.72425 | 0.302 |
| 402 | LINC00174 |
hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-376b-3p | 11 | PPARA | Sponge network | 0.123 | 0.81603 | 0.224 | 0.72425 | 0.301 |
| 403 | RBM26-AS1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-222-3p;hsa-miR-27a-3p;hsa-miR-342-5p;hsa-miR-429 | 11 | STX1B | Sponge network | 0.109 | 0.80346 | 0.074 | 0.87501 | 0.301 |
| 404 | LAMTOR5-AS1 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-369-3p;hsa-miR-376a-3p;hsa-miR-377-3p;hsa-miR-382-5p;hsa-miR-429;hsa-miR-491-5p | 15 | KLF12 | Sponge network | 0.011 | 0.9823 | 0.171 | 0.73281 | 0.301 |
| 405 | RP11-314O13.1 |
hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p | 10 | PPARA | Sponge network | 0.063 | 0.89816 | 0.224 | 0.72425 | 0.301 |
| 406 | RP11-6F2.5 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-301a-3p | 13 | AR | Sponge network | 0.253 | 0.44204 | 0.661 | 0.388 | 0.301 |
| 407 | RP1-187B23.1 |
hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p | 10 | PPARA | Sponge network | 0.075 | 0.87147 | 0.224 | 0.72425 | 0.3 |
| 408 | RP11-798K3.2 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p | 12 | MDGA1 | Sponge network | 0.217 | 0.77181 | 0.083 | 0.8765 | 0.299 |
| 409 | KB-431C1.4 |
hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-144-3p;hsa-miR-15a-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-335-5p | 10 | NEBL | Sponge network | -0.017 | 0.97318 | -0.126 | 0.74294 | 0.299 |
| 410 | LINC00853 |
hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-377-3p | 10 | HLF | Sponge network | 0.101 | 0.80817 | 0.706 | 0.36208 | 0.298 |
| 411 | ENTPD1-AS1 |
hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-515-5p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-522-3p;hsa-miR-7-5p | 10 | ESR1 | Sponge network | 0.012 | 0.9694 | 0.121 | 0.80686 | 0.297 |
| 412 | WEE2-AS1 |
hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 13 | MDGA1 | Sponge network | -0.042 | 0.91109 | 0.083 | 0.8765 | 0.295 |
| 413 | PTOV1-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-363-3p;hsa-miR-92a-3p | 12 | TTC9 | Sponge network | -0.074 | 0.84862 | 0.309 | 0.51342 | 0.295 |
| 414 | LINC00884 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-483-3p;hsa-miR-485-5p;hsa-miR-494-3p | 11 | SLC4A4 | Sponge network | 0.137 | 0.76561 | 0.236 | 0.64507 | 0.294 |
| 415 | THAP7-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-363-3p;hsa-miR-520e;hsa-miR-92a-3p | 11 | TTC9 | Sponge network | -0.058 | 0.89073 | 0.309 | 0.51342 | 0.294 |
| 416 | MAP3K14-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-342-3p | 12 | PPARA | Sponge network | 0.031 | 0.93669 | 0.224 | 0.72425 | 0.293 |
| 417 | RP11-320N7.2 |
hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 11 | MDGA1 | Sponge network | 0.397 | 0.4155 | 0.083 | 0.8765 | 0.291 |
| 418 | RP11-390E23.6 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-491-5p | 12 | HLF | Sponge network | 0.037 | 0.93955 | 0.706 | 0.36208 | 0.291 |
| 419 | MAP3K14-AS1 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-377-3p;hsa-miR-491-5p | 19 | HLF | Sponge network | 0.031 | 0.93669 | 0.706 | 0.36208 | 0.289 |
| 420 | LINC00521 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-516b-3p;hsa-miR-520d-5p | 10 | NFIA | Sponge network | 0.041 | 0.9118 | 0.294 | 0.66575 | 0.288 |
| 421 | TMEM254-AS1 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-375 | 11 | PRKCA | Sponge network | 0.03 | 0.94824 | -0.093 | 0.86855 | 0.288 |
| 422 | RP11-350G8.5 |
hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-302c-3p | 11 | HLF | Sponge network | 0.059 | 0.84655 | 0.706 | 0.36208 | 0.288 |
| 423 | LINC00884 |
hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-299-3p;hsa-miR-485-5p | 10 | AR | Sponge network | 0.137 | 0.76561 | 0.661 | 0.388 | 0.286 |
| 424 | RP11-1007O24.3 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-218-5p | 16 | HLF | Sponge network | 0.069 | 0.8938 | 0.706 | 0.36208 | 0.286 |
| 425 | RP11-6F2.5 |
hsa-miR-106a-5p;hsa-miR-146a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-494-3p | 10 | SLC1A2 | Sponge network | 0.253 | 0.44204 | 0.329 | 0.56983 | 0.286 |
| 426 | RP11-545I5.3 |
hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-301a-3p;hsa-miR-520d-3p | 10 | ESR1 | Sponge network | 0.103 | 0.76843 | 0.121 | 0.80686 | 0.286 |
| 427 | KRBOX1-AS1 |
hsa-miR-130a-3p;hsa-miR-143-3p;hsa-miR-145-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-28-5p;hsa-miR-320a | 10 | SNX27 | Sponge network | 0.012 | 0.9741 | -0.226 | 0.73456 | 0.285 |
| 428 | RP11-545I5.3 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-150-5p;hsa-miR-17-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-23a-3p;hsa-miR-342-3p;hsa-miR-429;hsa-miR-494-3p | 10 | SFXN1 | Sponge network | 0.103 | 0.76843 | 0.212 | 0.75913 | 0.285 |
| 429 | RP11-1006G14.1 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-301a-3p | 11 | AR | Sponge network | 0.059 | 0.8921 | 0.661 | 0.388 | 0.284 |
| 430 | RP11-390E23.6 |
hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-369-3p;hsa-miR-375;hsa-miR-376c-3p;hsa-miR-382-5p;hsa-miR-491-5p | 14 | KLF12 | Sponge network | 0.037 | 0.93955 | 0.171 | 0.73281 | 0.284 |
| 431 | LINC00339 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-miR-101-3p;hsa-miR-195-5p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-424-5p;hsa-miR-497-5p | 10 | ZBTB10 | Sponge network | 0 | 0.99996 | 0.057 | 0.91732 | 0.283 |
| 432 | RP11-862G15.2 | hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-429;hsa-miR-515-5p;hsa-miR-519c-3p;hsa-miR-520d-3p;hsa-miR-520e | 11 | NFIA | Sponge network | 0.019 | 0.96899 | 0.294 | 0.66575 | 0.281 |
| 433 | RP11-1006G14.1 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-500a-3p | 15 | HLF | Sponge network | 0.059 | 0.8921 | 0.706 | 0.36208 | 0.28 |
| 434 | AC068138.1 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-145-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-23a-3p | 11 | GFRA1 | Sponge network | 0.06 | 0.8855 | 0.161 | 0.75691 | 0.28 |
| 435 | LINC00884 |
hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-197-3p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 12 | MDGA1 | Sponge network | 0.137 | 0.76561 | 0.083 | 0.8765 | 0.278 |
| 436 | RP11-375I20.6 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-369-3p | 10 | KLF12 | Sponge network | 0.009 | 0.9807 | 0.171 | 0.73281 | 0.277 |
| 437 | ST7-AS1 |
hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-491-5p | 10 | HLF | Sponge network | -0.009 | 0.98084 | 0.706 | 0.36208 | 0.276 |
| 438 | RP11-1081M5.2 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-19a-3p;hsa-miR-200c-3p;hsa-miR-377-3p;hsa-miR-500a-3p | 16 | HLF | Sponge network | 0.065 | 0.84871 | 0.706 | 0.36208 | 0.276 |
| 439 | WEE2-AS1 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-146b-5p;hsa-miR-150-5p;hsa-miR-17-5p;hsa-miR-342-3p;hsa-miR-369-3p;hsa-miR-514a-3p;hsa-miR-520a-5p;hsa-miR-525-5p | 10 | KCNB1 | Sponge network | -0.042 | 0.91109 | 0.378 | 0.44855 | 0.275 |
| 440 | POLR2J4 |
hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p | 10 | PPARA | Sponge network | 0.083 | 0.85528 | 0.224 | 0.72425 | 0.275 |
| 441 | CTD-3193O13.1 |
hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-218-5p;hsa-miR-491-5p | 10 | HLF | Sponge network | 0.047 | 0.90548 | 0.706 | 0.36208 | 0.275 |
| 442 | RP11-46C24.7 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-375 | 11 | PRKCA | Sponge network | -0.039 | 0.92939 | -0.093 | 0.86855 | 0.274 |
| 443 | AP000253.1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199b-5p;hsa-miR-20b-5p;hsa-miR-223-3p;hsa-miR-224-5p | 10 | DENND5B | Sponge network | 0.27 | 0.48487 | 0.008 | 0.98644 | 0.273 |
| 444 | RP11-736K20.6 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-218-5p | 16 | HLF | Sponge network | 0.069 | 0.87221 | 0.706 | 0.36208 | 0.272 |
| 445 | RP11-390E23.6 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-369-3p | 12 | PPARA | Sponge network | 0.037 | 0.93955 | 0.224 | 0.72425 | 0.271 |
| 446 | KB-1460A1.5 |
hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-144-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p | 10 | NEBL | Sponge network | -0.055 | 0.92334 | -0.126 | 0.74294 | 0.271 |
| 447 | BCDIN3D-AS1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-196a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-500a-3p | 12 | HLF | Sponge network | 0.02 | 0.96325 | 0.706 | 0.36208 | 0.27 |
| 448 | DLX6-AS1 |
hsa-miR-122-5p;hsa-miR-126-5p;hsa-miR-130a-3p;hsa-miR-192-5p;hsa-miR-215-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30e-5p | 12 | SLC7A11 | Sponge network | -0.076 | 0.80689 | -0.564 | 0.28432 | 0.27 |
| 449 | MAP3K14-AS1 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-223-3p | 14 | AR | Sponge network | 0.031 | 0.93669 | 0.661 | 0.388 | 0.27 |
| 450 | RP11-401P9.5 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-342-3p;hsa-miR-369-3p;hsa-miR-376b-3p | 13 | PPARA | Sponge network | 0.033 | 0.92794 | 0.224 | 0.72425 | 0.27 |
| 451 | HOXA-AS2 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-377-3p;hsa-miR-491-5p | 13 | HLF | Sponge network | -0.046 | 0.91915 | 0.706 | 0.36208 | 0.27 |
| 452 | HEXA-AS1 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-196a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-491-5p | 13 | HLF | Sponge network | -0.001 | 0.99633 | 0.706 | 0.36208 | 0.27 |
| 453 | ATP6V0E2-AS1 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-375;hsa-miR-429;hsa-miR-494-3p | 12 | SLC4A4 | Sponge network | -0.095 | 0.8128 | 0.236 | 0.64507 | 0.269 |
| 454 | MAP3K14-AS1 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-377-3p;hsa-miR-491-5p | 13 | KLF12 | Sponge network | 0.031 | 0.93669 | 0.171 | 0.73281 | 0.269 |
| 455 | ST7-AS1 |
hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-369-3p | 10 | PPARA | Sponge network | -0.009 | 0.98084 | 0.224 | 0.72425 | 0.269 |
| 456 | CTD-3074O7.5 |
hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-377-3p;hsa-miR-491-5p | 10 | KLF12 | Sponge network | -0.027 | 0.95238 | 0.171 | 0.73281 | 0.266 |
| 457 | RP5-991G20.1 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-491-5p | 11 | KLF12 | Sponge network | 0.039 | 0.93103 | 0.171 | 0.73281 | 0.266 |
| 458 | POLR2J4 |
hsa-miR-1323;hsa-miR-142-5p;hsa-miR-155-5p;hsa-miR-21-5p;hsa-miR-223-3p;hsa-miR-512-3p;hsa-miR-515-3p;hsa-miR-516b-3p;hsa-miR-519c-3p;hsa-miR-520c-3p;hsa-miR-520d-3p;hsa-miR-520d-5p;hsa-miR-520e;hsa-miR-524-5p;hsa-miR-618 | 15 | NFIA | Sponge network | 0.083 | 0.85528 | 0.294 | 0.66575 | 0.264 |
| 459 | AC068138.1 |
hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-218-5p | 12 | HLF | Sponge network | 0.06 | 0.8855 | 0.706 | 0.36208 | 0.263 |
| 460 | SMIM2-AS1 |
hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-188-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-7-5p | 11 | ESR1 | Sponge network | 0.194 | 0.64908 | 0.121 | 0.80686 | 0.262 |
| 461 | RFPL1S |
hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20b-5p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-335-5p;hsa-miR-93-5p | 11 | GPRIN2 | Sponge network | 0.026 | 0.93109 | -0.003 | 0.99411 | 0.262 |
| 462 | PTPRG-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-342-3p;hsa-miR-369-3p;hsa-miR-376b-3p | 13 | PPARA | Sponge network | 0.009 | 0.97688 | 0.224 | 0.72425 | 0.261 |
| 463 | RP1-167G20.1 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-218-5p | 11 | HLF | Sponge network | 0.038 | 0.90491 | 0.706 | 0.36208 | 0.261 |
| 464 | LINC00174 |
hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-375;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-494-3p | 10 | SLC4A4 | Sponge network | 0.123 | 0.81603 | 0.236 | 0.64507 | 0.26 |
| 465 | SNHG16 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-miR-101-3p;hsa-miR-195-5p;hsa-miR-26b-5p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-30a-5p;hsa-miR-497-5p;hsa-miR-98-5p | 12 | ZBTB10 | Sponge network | -0.137 | 0.87069 | 0.057 | 0.91732 | 0.26 |
| 466 | CTD-2619J13.17 |
hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-376a-3p | 10 | KLF12 | Sponge network | 0.034 | 0.93595 | 0.171 | 0.73281 | 0.259 |
| 467 | CTB-131B5.5 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199b-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-377-3p | 13 | HLF | Sponge network | 0.216 | 0.59133 | 0.706 | 0.36208 | 0.259 |
| 468 | RP11-620J15.3 |
hsa-miR-126-5p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30a-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30e-5p | 11 | NEBL | Sponge network | -0.266 | 0.60278 | -0.126 | 0.74294 | 0.259 |
| 469 | LINC00094 |
hsa-let-7b-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-218-5p;hsa-miR-377-3p;hsa-miR-429;hsa-miR-491-5p | 12 | HLF | Sponge network | -0.246 | 0.62122 | 0.706 | 0.36208 | 0.258 |
| 470 | RP11-736K20.6 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-23a-3p;hsa-miR-376c-3p | 11 | GFRA1 | Sponge network | 0.069 | 0.87221 | 0.161 | 0.75691 | 0.258 |
| 471 | RP11-6F2.5 |
hsa-miR-106a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-494-3p | 12 | SLC4A4 | Sponge network | 0.253 | 0.44204 | 0.236 | 0.64507 | 0.258 |
| 472 | RP11-375I20.6 |
hsa-miR-106a-5p;hsa-miR-132-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-20b-5p;hsa-miR-223-3p | 11 | AR | Sponge network | 0.009 | 0.9807 | 0.661 | 0.388 | 0.258 |
| 473 | SPTY2D1-AS1 |
hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-148b-3p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30a-5p | 11 | NEBL | Sponge network | -0.012 | 0.97196 | -0.126 | 0.74294 | 0.258 |
| 474 | RP11-375I20.6 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-342-3p;hsa-miR-369-3p | 12 | PPARA | Sponge network | 0.009 | 0.9807 | 0.224 | 0.72425 | 0.258 |
| 475 | RP11-734K2.4 |
hsa-miR-145-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-188-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-222-3p;hsa-miR-7-5p | 10 | ESR1 | Sponge network | 0.094 | 0.83958 | 0.121 | 0.80686 | 0.258 |
| 476 | OSTM1-AS1 |
hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27b-3p;hsa-miR-30a-5p;hsa-miR-30e-5p;hsa-miR-335-5p | 10 | NEBL | Sponge network | -0.029 | 0.92722 | -0.126 | 0.74294 | 0.257 |
| 477 | KB-431C1.4 |
hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-139-5p;hsa-miR-144-3p;hsa-miR-15a-5p;hsa-miR-195-5p;hsa-miR-26a-5p;hsa-miR-26b-5p;hsa-miR-27a-3p;hsa-miR-27b-3p;hsa-miR-29a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p | 13 | PLAG1 | Sponge network | -0.017 | 0.97318 | -0.227 | 0.6194 | 0.257 |
| 478 | RP11-448A19.1 |
hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199b-5p;hsa-miR-218-5p;hsa-miR-377-3p | 10 | HLF | Sponge network | 0.064 | 0.88097 | 0.706 | 0.36208 | 0.257 |
| 479 | LINC00667 |
hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181d-5p;hsa-miR-19b-3p;hsa-miR-363-3p;hsa-miR-375;hsa-miR-424-5p;hsa-miR-429 | 10 | ZFHX4 | Sponge network | -0.166 | 0.75617 | 0.088 | 0.8617 | 0.256 |
| 480 | PART1 |
hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-150-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-183-5p;hsa-miR-200c-3p;hsa-miR-214-3p;hsa-miR-218-5p | 10 | PRKCA | Sponge network | -0.062 | 0.88185 | -0.093 | 0.86855 | 0.256 |
| 481 | LINC00667 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-363-3p;hsa-miR-429 | 12 | PPARA | Sponge network | -0.166 | 0.75617 | 0.224 | 0.72425 | 0.256 |
| 482 | LINC00648 |
hsa-miR-125b-5p;hsa-miR-126-5p;hsa-miR-144-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-195-5p;hsa-miR-26b-5p;hsa-miR-30a-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30e-5p | 11 | NEBL | Sponge network | -0.197 | 0.54083 | -0.126 | 0.74294 | 0.255 |
| 483 | LAMTOR5-AS1 |
hsa-let-7b-5p;hsa-let-7e-5p;hsa-miR-10a-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-182-5p;hsa-miR-199b-5p;hsa-miR-214-3p;hsa-miR-218-5p;hsa-miR-223-3p;hsa-miR-24-3p;hsa-miR-27a-3p;hsa-miR-376a-3p;hsa-miR-376b-3p | 16 | MDGA1 | Sponge network | 0.011 | 0.9823 | 0.083 | 0.8765 | 0.255 |
| 484 | LINC00667 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-199a-5p;hsa-miR-199b-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-363-3p | 10 | TTC9 | Sponge network | -0.166 | 0.75617 | 0.309 | 0.51342 | 0.253 |
| 485 | RP11-290H9.4 | hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-196a-5p;hsa-miR-199b-5p;hsa-miR-200c-3p;hsa-miR-20b-5p;hsa-miR-218-5p;hsa-miR-491-5p | 12 | HLF | Sponge network | 0.088 | 0.83935 | 0.706 | 0.36208 | 0.253 |
| 486 | SDCBP2-AS1 |
hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-223-3p;hsa-miR-301a-3p;hsa-miR-493-5p | 10 | AR | Sponge network | 0.03 | 0.94851 | 0.661 | 0.388 | 0.253 |
| 487 | ATP6V0E2-AS1 |
hsa-miR-142-5p;hsa-miR-145-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-19b-3p;hsa-miR-375;hsa-miR-424-5p;hsa-miR-429 | 10 | ZFHX4 | Sponge network | -0.095 | 0.8128 | 0.088 | 0.8617 | 0.253 |
| 488 | RP11-375I20.6 |
hsa-let-7e-5p;hsa-miR-106a-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-199a-5p;hsa-miR-20b-5p;hsa-miR-218-5p | 12 | HLF | Sponge network | 0.009 | 0.9807 | 0.706 | 0.36208 | 0.253 |
| 489 | WEE2-AS1 |
hsa-let-7e-5p;hsa-miR-125a-5p;hsa-miR-142-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200c-3p;hsa-miR-223-3p;hsa-miR-376a-3p;hsa-miR-376b-3p;hsa-miR-494-3p | 12 | SLC4A4 | Sponge network | -0.042 | 0.91109 | 0.236 | 0.64507 | 0.252 |
| 490 | RP11-166A12.1 |
hsa-miR-106a-5p;hsa-miR-141-3p;hsa-miR-142-5p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-19a-3p;hsa-miR-223-3p;hsa-miR-299-3p;hsa-miR-301a-3p | 13 | AR | Sponge network | 0.088 | 0.84584 | 0.661 | 0.388 | 0.252 |
| 491 | RBM26-AS1 |
hsa-miR-106a-5p;hsa-miR-142-3p;hsa-miR-146a-5p;hsa-miR-155-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-18a-5p;hsa-miR-18b-5p;hsa-miR-21-5p;hsa-miR-376b-3p;hsa-miR-429 | 12 | PPARA | Sponge network | 0.109 | 0.80346 | 0.224 | 0.72425 | 0.252 |
| 492 | CYP1B1-AS1 | hsa-miR-106a-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-512-3p;hsa-miR-515-5p;hsa-miR-516b-3p;hsa-miR-519c-3p;hsa-miR-519e-3p;hsa-miR-520d-3p;hsa-miR-520e;hsa-miR-618 | 12 | NFIA | Sponge network | 0.011 | 0.97204 | 0.294 | 0.66575 | 0.252 |