This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-122-5p | ADAM17 | -1.24 | 0 | 0.08 | 0.57225 | miRNAWalker2 validate; miRTarBase | -0.19 | 0 | 19296470 | We further showed that one of the miR-122 targets ADAM17 a disintegrin and metalloprotease 17 is involved in metastasis; Silencing of ADAM17 resulted in a dramatic reduction of in vitro migration invasion in vivo tumorigenesis angiogenesis and local invasion in the livers of nude mice which is similar to that which occurs with the restoration of miR-122; Our study suggests that miR-122 a tumor suppressor microRNA affecting hepatocellular carcinoma intrahepatic metastasis by angiogenesis suppression exerts some of its action via regulation of ADAM17; Using the concomitant down-regulation of its targets including ADAM17 a rational therapeutic strategy based on miR-122 may prove to be beneficial for patients with hepatocellular carcinoma |
| 2 | hsa-miR-145-5p | APH1A | -1.48 | 0 | 0.33 | 1.0E-5 | miRNAWalker2 validate | -0.1 | 0 | NA | |
| 3 | hsa-miR-1976 | APH1A | -0.43 | 0.00325 | 0.33 | 1.0E-5 | MirTarget | -0.13 | 0 | NA | |
| 4 | hsa-miR-2355-5p | APH1A | -0.23 | 0.15791 | 0.33 | 1.0E-5 | MirTarget | -0.1 | 0 | NA | |
| 5 | hsa-miR-342-3p | APH1A | -0.32 | 0.04498 | 0.33 | 1.0E-5 | miRanda | -0.13 | 0 | NA | |
| 6 | hsa-miR-582-5p | APH1A | -0.68 | 0.00104 | 0.33 | 1.0E-5 | PITA | -0.12 | 0 | NA | |
| 7 | hsa-miR-324-3p | CREBBP | 0.26 | 0.05061 | -0.2 | 0.01387 | miRNAWalker2 validate | -0.12 | 3.0E-5 | NA | |
| 8 | hsa-miR-330-5p | CREBBP | 0.44 | 0.00533 | -0.2 | 0.01387 | miRanda | -0.11 | 1.0E-5 | NA | |
| 9 | hsa-miR-769-5p | CREBBP | 0.22 | 0.03334 | -0.2 | 0.01387 | miRNAWalker2 validate | -0.13 | 0.00077 | NA | |
| 10 | hsa-miR-146b-5p | CTBP2 | 0.42 | 0.04574 | -0.39 | 0.02251 | miRanda | -0.18 | 1.0E-5 | NA | |
| 11 | hsa-miR-616-5p | CTBP2 | 0.15 | 0.40284 | -0.39 | 0.02251 | mirMAP | -0.31 | 0 | NA | |
| 12 | hsa-miR-618 | CTBP2 | 0.14 | 0.51715 | -0.39 | 0.02251 | MirTarget; PITA | -0.33 | 0 | NA | |
| 13 | hsa-miR-103a-3p | DLL1 | 0.77 | 0 | -0.55 | 0.01603 | miRNATAP | -0.51 | 0 | NA | |
| 14 | hsa-miR-107 | DLL1 | 0.24 | 0.01708 | -0.55 | 0.01603 | PITA; miRanda; miRNATAP | -0.62 | 0 | NA | |
| 15 | hsa-miR-130b-3p | DLL1 | 0.69 | 0.00011 | -0.55 | 0.01603 | miRNATAP | -0.21 | 0.00085 | NA | |
| 16 | hsa-miR-146b-5p | DLL1 | 0.42 | 0.04574 | -0.55 | 0.01603 | miRanda | -0.18 | 0.00088 | NA | |
| 17 | hsa-miR-15a-5p | DLL1 | 0.35 | 0.00077 | -0.55 | 0.01603 | miRNATAP | -0.37 | 0.00062 | NA | |
| 18 | hsa-miR-15b-5p | DLL1 | 0.23 | 0.08248 | -0.55 | 0.01603 | miRNATAP | -0.35 | 3.0E-5 | NA | |
| 19 | hsa-miR-301a-3p | DLL1 | 0.84 | 0 | -0.55 | 0.01603 | miRNATAP | -0.26 | 4.0E-5 | NA | |
| 20 | hsa-miR-423-3p | DLL1 | 0.3 | 0.00067 | -0.55 | 0.01603 | PITA | -0.61 | 0 | NA | |
| 21 | hsa-miR-454-3p | DLL1 | 0.67 | 0 | -0.55 | 0.01603 | miRNATAP | -0.21 | 0.00766 | NA | |
| 22 | hsa-miR-589-5p | DLL1 | 1.19 | 0 | -0.55 | 0.01603 | miRNATAP | -0.63 | 0 | NA | |
| 23 | hsa-miR-29a-5p | DLL4 | -0.11 | 0.34962 | 1.07 | 0 | miRNATAP | -0.21 | 0.00016 | NA | |
| 24 | hsa-miR-30c-5p | DLL4 | -0.43 | 0.00016 | 1.07 | 0 | miRNATAP | -0.17 | 0.00359 | NA | |
| 25 | hsa-miR-424-5p | DLL4 | -2.63 | 0 | 1.07 | 0 | miRNATAP | -0.13 | 0.0002 | NA | |
| 26 | hsa-miR-188-5p | DTX1 | 1.12 | 0 | -2.8 | 0 | PITA | -0.32 | 0.0009 | NA | |
| 27 | hsa-miR-421 | DTX1 | 0.94 | 0 | -2.8 | 0 | PITA; miRanda; miRNATAP | -0.35 | 0.00027 | NA | |
| 28 | hsa-let-7a-5p | DTX2 | -0.33 | 0.00046 | 0.39 | 0 | TargetScan; miRNATAP | -0.23 | 0 | NA | |
| 29 | hsa-let-7b-5p | DTX2 | -0.96 | 0 | 0.39 | 0 | miRNATAP | -0.12 | 1.0E-5 | NA | |
| 30 | hsa-let-7g-5p | DTX2 | -0.46 | 2.0E-5 | 0.39 | 0 | miRNATAP | -0.2 | 0 | NA | |
| 31 | hsa-miR-107 | DTX3 | 0.24 | 0.01708 | 0.08 | 0.67413 | miRanda | -0.29 | 0.00171 | NA | |
| 32 | hsa-miR-148a-5p | DTX3 | -0.77 | 0 | 0.08 | 0.67413 | mirMAP | -0.35 | 0 | NA | |
| 33 | hsa-miR-339-5p | DTX3L | 0.28 | 0.03557 | -0.19 | 0.02216 | miRanda | -0.1 | 0.00089 | NA | |
| 34 | hsa-miR-340-3p | DTX3L | -0.33 | 0.0033 | -0.19 | 0.02216 | MirTarget | -0.11 | 0.00292 | NA | |
| 35 | hsa-miR-140-3p | DTX4 | 0.55 | 0 | -0.32 | 0.16271 | miRNATAP | -0.43 | 0.0011 | NA | |
| 36 | hsa-miR-204-5p | DTX4 | -0.54 | 0.03309 | -0.32 | 0.16271 | miRNATAP | -0.29 | 0 | NA | |
| 37 | hsa-miR-29a-5p | DTX4 | -0.11 | 0.34962 | -0.32 | 0.16271 | mirMAP | -0.46 | 0 | NA | |
| 38 | hsa-miR-29b-3p | DTX4 | -0.35 | 0.01214 | -0.32 | 0.16271 | miRNATAP | -0.26 | 0.00109 | NA | |
| 39 | hsa-miR-374a-5p | DTX4 | 0.02 | 0.86978 | -0.32 | 0.16271 | mirMAP | -0.8 | 0 | NA | |
| 40 | hsa-miR-374b-5p | DTX4 | -0.31 | 0.00301 | -0.32 | 0.16271 | mirMAP | -0.62 | 0 | NA | |
| 41 | hsa-miR-548b-3p | DTX4 | -0.02 | 0.93082 | -0.32 | 0.16271 | PITA; mirMAP; miRNATAP | -0.19 | 9.0E-5 | NA | |
| 42 | hsa-miR-10a-5p | DVL1 | -1.48 | 0 | 0.1 | 0.35133 | miRNAWalker2 validate | -0.12 | 0 | NA | |
| 43 | hsa-miR-27a-3p | DVL2 | -0.37 | 0.00876 | 0.93 | 0 | miRNATAP | -0.1 | 0.00527 | NA | |
| 44 | hsa-miR-27b-3p | DVL2 | -0.82 | 0 | 0.93 | 0 | miRNATAP | -0.22 | 0 | NA | |
| 45 | hsa-miR-125b-5p | DVL3 | -1.36 | 0 | 0.61 | 0 | miRNATAP | -0.13 | 0 | NA | |
| 46 | hsa-miR-26b-5p | DVL3 | -1.11 | 0 | 0.61 | 0 | miRNAWalker2 validate | -0.11 | 0.00053 | NA | |
| 47 | hsa-miR-29b-2-5p | DVL3 | -0.22 | 0.08431 | 0.61 | 0 | MirTarget | -0.1 | 0.00053 | NA | |
| 48 | hsa-miR-339-5p | EP300 | 0.28 | 0.03557 | -0.2 | 0.07049 | miRanda | -0.16 | 0.0001 | NA | |
| 49 | hsa-miR-342-3p | EP300 | -0.32 | 0.04498 | -0.2 | 0.07049 | MirTarget; PITA; miRanda; miRNATAP | -0.11 | 0.00085 | NA | |
| 50 | hsa-miR-30a-5p | HDAC1 | -0.63 | 0.00011 | 0.31 | 0.00015 | miRNAWalker2 validate | -0.15 | 0 | NA | |
| 51 | hsa-let-7b-3p | HDAC2 | -1.22 | 0 | 0.16 | 0.06327 | mirMAP | -0.12 | 1.0E-5 | NA | |
| 52 | hsa-miR-125b-2-3p | HDAC2 | -1.66 | 0 | 0.16 | 0.06327 | mirMAP | -0.1 | 0 | NA | |
| 53 | hsa-miR-16-1-3p | JAG1 | 0.39 | 0.00112 | 0.28 | 0.09623 | MirTarget | -0.26 | 0.00018 | NA | |
| 54 | hsa-miR-186-5p | JAG1 | -0.06 | 0.53529 | 0.28 | 0.09623 | MirTarget; miRNATAP | -0.3 | 0.00055 | NA | |
| 55 | hsa-miR-26b-3p | JAG1 | -1.26 | 0 | 0.28 | 0.09623 | miRNATAP | -0.27 | 5.0E-5 | NA | |
| 56 | hsa-miR-26b-5p | JAG1 | -1.11 | 0 | 0.28 | 0.09623 | MirTarget; miRNATAP | -0.2 | 0.00313 | NA | |
| 57 | hsa-miR-34a-5p | JAG1 | 1.04 | 0 | 0.28 | 0.09623 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.19 | 0.00096 | NA | |
| 58 | hsa-miR-361-3p | JAG1 | 0.28 | 0.00549 | 0.28 | 0.09623 | miRNATAP | -0.24 | 0.00533 | NA | |
| 59 | hsa-miR-590-5p | JAG1 | -0.1 | 0.31003 | 0.28 | 0.09623 | MirTarget; PITA; miRanda; miRNATAP | -0.31 | 0.00018 | NA | |
| 60 | hsa-miR-140-5p | JAG2 | -0.22 | 0.01407 | 1.64 | 0 | miRanda | -0.37 | 0.00048 | NA | |
| 61 | hsa-miR-3607-3p | JAG2 | -2.16 | 0 | 1.64 | 0 | miRNATAP | -0.24 | 0 | NA | |
| 62 | hsa-miR-505-3p | JAG2 | -1.2 | 0 | 1.64 | 0 | mirMAP | -0.33 | 0 | NA | |
| 63 | hsa-miR-342-3p | KAT2A | -0.32 | 0.04498 | 1.46 | 0 | MirTarget; miRanda | -0.12 | 0.00265 | NA | |
| 64 | hsa-miR-106a-5p | KAT2B | -0.46 | 0.00972 | -0.94 | 0 | MirTarget | -0.18 | 1.0E-5 | NA | |
| 65 | hsa-miR-106b-5p | KAT2B | 0.65 | 0 | -0.94 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.52 | 0 | NA | |
| 66 | hsa-miR-125a-3p | KAT2B | -0.84 | 4.0E-5 | -0.94 | 0 | miRanda | -0.15 | 4.0E-5 | NA | |
| 67 | hsa-miR-17-5p | KAT2B | 0.7 | 2.0E-5 | -0.94 | 0 | MirTarget; TargetScan | -0.4 | 0 | 23095762 | miR 17 5p targets the p300/CBP associated factor and modulates androgen receptor transcriptional activity in cultured prostate cancer cells; Targeting of PCAF by miR-17-5p was evaluated using the luciferase reporter assay; Expression of PCAF in PCa cells was associated with the downregulation of miR-17-5p; Targeting of the 3'-untranslated region of PCAF mRNA by miR-17-5p caused translational suppression and RNA degradation and consequently modulation of AR transcriptional activity in PCa cells; PCAF is upregulated in cultured PCa cells and upregulation of PCAF is associated with the downregulation of miR-17-5p; Targeting of PCAF by miR-17-5p modulates AR transcriptional activity and cell growth in cultured PCa cells |
| 68 | hsa-miR-181a-5p | KAT2B | 0.25 | 0.05519 | -0.94 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.43 | 0 | NA | |
| 69 | hsa-miR-181b-5p | KAT2B | 0.49 | 0.00105 | -0.94 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.46 | 0 | NA | |
| 70 | hsa-miR-181c-5p | KAT2B | -0.01 | 0.96913 | -0.94 | 0 | MirTarget | -0.27 | 0 | NA | |
| 71 | hsa-miR-181d-5p | KAT2B | 0.16 | 0.36381 | -0.94 | 0 | MirTarget | -0.28 | 0 | NA | |
| 72 | hsa-miR-19a-3p | KAT2B | 1.02 | 0 | -0.94 | 0 | miRNAWalker2 validate | -0.26 | 0 | NA | |
| 73 | hsa-miR-19b-3p | KAT2B | 0.6 | 0.00017 | -0.94 | 0 | miRNAWalker2 validate | -0.3 | 0 | NA | |
| 74 | hsa-miR-20a-3p | KAT2B | -0.32 | 0.04679 | -0.94 | 0 | MirTarget | -0.22 | 0 | NA | |
| 75 | hsa-miR-20a-5p | KAT2B | 0.85 | 0 | -0.94 | 0 | MirTarget | -0.32 | 0 | NA | |
| 76 | hsa-miR-20b-5p | KAT2B | 0.46 | 0.02859 | -0.94 | 0 | MirTarget | -0.11 | 0.00122 | NA | |
| 77 | hsa-miR-25-3p | KAT2B | 0.63 | 0 | -0.94 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.26 | 0.00023 | NA | |
| 78 | hsa-miR-330-3p | KAT2B | -0.33 | 0.03161 | -0.94 | 0 | MirTarget | -0.27 | 0 | NA | |
| 79 | hsa-miR-338-5p | KAT2B | -0.22 | 0.25239 | -0.94 | 0 | MirTarget; miRNATAP | -0.15 | 7.0E-5 | NA | |
| 80 | hsa-miR-342-3p | KAT2B | -0.32 | 0.04498 | -0.94 | 0 | miRanda | -0.18 | 7.0E-5 | NA | |
| 81 | hsa-miR-363-3p | KAT2B | -0.17 | 0.37519 | -0.94 | 0 | MirTarget | -0.1 | 0.00633 | NA | |
| 82 | hsa-miR-429 | KAT2B | -1.4 | 7.0E-5 | -0.94 | 0 | miRanda | -0.13 | 0 | NA | |
| 83 | hsa-miR-590-5p | KAT2B | -0.1 | 0.31003 | -0.94 | 0 | miRanda | -0.26 | 0.0004 | NA | |
| 84 | hsa-miR-92a-3p | KAT2B | 0.21 | 0.13429 | -0.94 | 0 | miRNAWalker2 validate; MirTarget | -0.35 | 0 | NA | |
| 85 | hsa-miR-92b-3p | KAT2B | 0.22 | 0.29619 | -0.94 | 0 | MirTarget | -0.23 | 0 | NA | |
| 86 | hsa-miR-93-5p | KAT2B | 1.4 | 0 | -0.94 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.38 | 0 | NA | |
| 87 | hsa-miR-107 | LFNG | 0.24 | 0.01708 | 0.26 | 0.14017 | miRanda | -0.36 | 3.0E-5 | NA | |
| 88 | hsa-miR-365a-3p | LFNG | 0.16 | 0.15325 | 0.26 | 0.14017 | mirMAP | -0.45 | 0 | NA | |
| 89 | hsa-miR-219a-5p | MFNG | 0.18 | 0.32269 | -0.41 | 0.0014 | MirTarget | -0.14 | 0.00012 | NA | |
| 90 | hsa-miR-28-5p | MFNG | -0.43 | 0 | -0.41 | 0.0014 | miRanda | -0.25 | 0.00032 | NA | |
| 91 | hsa-miR-29a-3p | NCOR2 | -0.86 | 0 | 0.53 | 0 | miRNATAP | -0.15 | 0 | NA | |
| 92 | hsa-miR-29b-3p | NCOR2 | -0.35 | 0.01214 | 0.53 | 0 | miRNATAP | -0.13 | 1.0E-5 | NA | |
| 93 | hsa-miR-29c-3p | NCOR2 | -1.44 | 0 | 0.53 | 0 | miRNATAP | -0.16 | 0 | NA | |
| 94 | hsa-miR-3607-3p | NCOR2 | -2.16 | 0 | 0.53 | 0 | miRNATAP | -0.14 | 0 | NA | |
| 95 | hsa-miR-101-3p | NOTCH1 | -1.48 | 0 | -0.03 | 0.79556 | MirTarget | -0.13 | 0.00575 | NA | |
| 96 | hsa-miR-34a-5p | NOTCH1 | 1.04 | 0 | -0.03 | 0.79556 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.24 | 0 | 22438124; 23140286; 24349627; 24565525; 25783790; 20351093; 27082152; 23642368; 22347519; 23430952; 23902763; 21743299; 22992310; 23145211; 25623761; 23085450 | Delta tocotrienol suppresses Notch 1 pathway by upregulating miR 34a in nonsmall cell lung cancer cells; In our study using miRNA microarray we observed that downregulation of the Notch-1 pathway by delta-tocotrienol correlated with upregulation of miR-34a in nonsmall cell lung cancer cells NSCLC;We found that Re-expression forced expression of miR-34a inhibits cell growth and induces apoptosis with concomitant down-regulation of Notch-1 signaling pathway one of the target of miR-34a; Moreover treatment of PC cells with a natural compound genistein led to the up-regulation of miR-34a resulting in the down-regulation of Notch-1 which was correlated with inhibition of cell growth and induction of apoptosis;Most importantly BR-DIM intervention in PCa patients prior to radical prostatectomy showed reexpression of miR-34a which was consistent with decreased expression of AR PSA and Notch-1 in PCa tissue specimens;In addition intracellular restoration of miR-34a inhibited breast cancer cell migration via targeting Notch-1 signaling;MicroRNA 34a suppresses the breast cancer stem cell like characteristics by downregulating Notch1 pathway; In this study we verified that miR-34a directly and functionally targeted Notch1 in MCF-7 cells; We reported that miR-34a negatively regulated cell proliferation migration and invasion and breast cancer stem cell propagation by downregulating Notch1; The expression of miR-34a was negatively correlated with tumor stages metastasis and Notch1 expression in breast cancer tissues; Furthermore overexpression of miR-34a increased chemosensitivity of breast cancer cells to paclitaxel PTX by downregulating the Notch1 pathway; Taken together our results indicate that miR-34a inhibited breast cancer stemness and increased the chemosensitivity to PTX partially by downregulating the Notch1 pathway suggesting that miR-34a/Notch1 play an important role in regulating breast cancer stem cells;MicroRNA 34a suppresses invasion through downregulation of Notch1 and Jagged1 in cervical carcinoma and choriocarcinoma cells; Computational miRNA target prediction suggested that Notch1 and Jagged1 were targets of miR-34a; By using functional assays miR-34a was demonstrated to bind to the 3' untranslated regions of Notch1 and Jagged1; Forced expression of miR-34a altered the expression of Notch1 and Jagged1 protein as well as Notch signaling as shown by the response of Hairy Enhancer of Split-1 protein to these treatments using western blot analysis;We showed that miR-34a as a tumor suppressor could separately reduce the stemness of BCSCs and activate the cytotoxic susceptibility of BCSCs to natural killer NK cells in vitro via down regulating the expression of Notch1 signaling molecules;Mechanistically miR-34a sequesters Notch1 mRNA to generate a sharp threshold response where a bimodal Notch signal specifies the choice between self-renewal and differentiation;Most importantly BR-DIM intervention in PCa patients prior to radical prostatectomy showed re-expression of miR-34a which was consistent with decreased expression of AR PSA and Notch-1 in PCa tissue specimens;We also found that reexpression of miR-34a and miR-200b by transfection led to reduced expression of Notch-1 resulting in the inhibition of osteosarcoma cell proliferation invasion and angiogenesis;Rhamnetin and cirsiliol induce radiosensitization and inhibition of epithelial mesenchymal transition EMT by miR 34a mediated suppression of Notch 1 expression in non small cell lung cancer cell lines; Indeed rhamnetin and cirsiliol increased the expression of tumor-suppressive microRNA miR-34a in a p53-dependent manner leading to inhibition of Notch-1 expression;MicroRNA 34a targets notch1 and inhibits cell proliferation in glioblastoma multiforme; Also we identified notch1 as a direct target gene of miR-34a; Knockdown of notch1 showed similar cellular functions as overexpression of miR-34a both in vitro and in vivo; Collectively our findings show that miR-34a is downregulated in GBM cells and inhibits GBM growth by targeting notch1;The re-expression of miR-34 led to a marked reduction in the expression of its target gene Notch-1;Inactivation of AR and Notch 1 signaling by miR 34a attenuates prostate cancer aggressiveness; We found that over-expression of miR-34a led to reduced expression of AR PSA and Notch-1; These findings suggest that the loss of miR-34a is directly linked with up-regulation of AR and Notch-1 both of which are highly expressed in PCa and thus finding innovative approaches by which miR-34a expression could be up-regulated will have a huge impact on the treatment of PCa especially for the treatment of mCRPC;miR 34a may regulate sensitivity of breast cancer cells to adriamycin via targeting Notch1; To explore the influence of miR-34a on Notch1 expression in breast cancer cells and to explore the role of miR-34a in the sensitivity of breast cancer cells to Adriamycin ADR; The expression levels of Notch1 mRNA and protein in MCF-7/ADR cells transfected with miR-34a mimics were significantly up-regulated; On the contrary the expressions of Notch1 mRNA and protein in MCF-7 cells transfected with miR-34a inhibitor were down-regulated; miR-34a negatively regulates the expression of Notch1 at both mRNA and protein levels;MicroRNA 34a modulates chemosensitivity of breast cancer cells to adriamycin by targeting Notch1; The association of miR-34a and Notch1 was analyzed by dual-luciferase reporter assay and Notch1-siRNA technology; Real-time PCR assay was performed to test the expression of miR-34a and Notch1 in 38 selective breast cancer tissue samples; MiR-34a mimic could inhibit the luciferase activity of the construct containing wild-type 3' UTR of Notch1 in MCF-7/ADR cells; Further there was an inverse association between Notch1 and miR-34a expression in breast cancer; Dysregulation of miR-34a plays critical roles in the acquired ADR resistance of breast cancer at least in part via targeting Notch1 |
| 97 | hsa-miR-103a-3p | NOTCH2 | 0.77 | 0 | -0.38 | 0.00411 | MirTarget | -0.2 | 0.00055 | NA | |
| 98 | hsa-miR-107 | NOTCH2 | 0.24 | 0.01708 | -0.38 | 0.00411 | miRNAWalker2 validate; MirTarget; miRanda; miRNATAP | -0.21 | 0.001 | NA | |
| 99 | hsa-miR-146b-5p | NOTCH2 | 0.42 | 0.04574 | -0.38 | 0.00411 | miRanda | -0.14 | 1.0E-5 | NA | |
| 100 | hsa-miR-148a-5p | NOTCH2 | -0.77 | 0 | -0.38 | 0.00411 | MirTarget | -0.12 | 0.00236 | NA | |
| 101 | hsa-miR-15a-5p | NOTCH2 | 0.35 | 0.00077 | -0.38 | 0.00411 | MirTarget | -0.21 | 0.00076 | NA | |
| 102 | hsa-miR-16-1-3p | NOTCH2 | 0.39 | 0.00112 | -0.38 | 0.00411 | mirMAP | -0.17 | 0.00213 | NA | |
| 103 | hsa-miR-17-5p | NOTCH2 | 0.7 | 2.0E-5 | -0.38 | 0.00411 | miRNAWalker2 validate | -0.11 | 0.00704 | NA | |
| 104 | hsa-miR-29b-3p | NOTCH2 | -0.35 | 0.01214 | -0.38 | 0.00411 | MirTarget | -0.17 | 0.00018 | NA | |
| 105 | hsa-miR-34a-5p | NOTCH2 | 1.04 | 0 | -0.38 | 0.00411 | miRNAWalker2 validate; miRNATAP | -0.21 | 0 | NA | |
| 106 | hsa-miR-374a-5p | NOTCH2 | 0.02 | 0.86978 | -0.38 | 0.00411 | mirMAP | -0.18 | 0.00666 | NA | |
| 107 | hsa-miR-324-5p | NOTCH4 | 0.37 | 0.00592 | 0.46 | 0.00012 | miRanda | -0.18 | 4.0E-5 | NA | |
| 108 | hsa-miR-421 | NOTCH4 | 0.94 | 0 | 0.46 | 0.00012 | miRanda | -0.14 | 3.0E-5 | NA | |
| 109 | hsa-miR-330-5p | NUMB | 0.44 | 0.00533 | -0.37 | 0 | miRanda | -0.1 | 1.0E-5 | NA | |
| 110 | hsa-miR-339-5p | NUMB | 0.28 | 0.03557 | -0.37 | 0 | miRanda | -0.13 | 0 | NA | |
| 111 | hsa-miR-589-5p | NUMB | 1.19 | 0 | -0.37 | 0 | miRNATAP | -0.11 | 0.00016 | NA | |
| 112 | hsa-miR-590-5p | NUMB | -0.1 | 0.31003 | -0.37 | 0 | miRanda | -0.11 | 0.00174 | NA | |
| 113 | hsa-miR-122-5p | NUMBL | -1.24 | 0 | 0.39 | 0.00366 | miRNAWalker2 validate; miRTarBase | -0.21 | 0 | NA | |
| 114 | hsa-miR-194-3p | NUMBL | -0.77 | 3.0E-5 | 0.39 | 0.00366 | MirTarget | -0.29 | 0 | NA | |
| 115 | hsa-miR-374b-5p | NUMBL | -0.31 | 0.00301 | 0.39 | 0.00366 | miRNATAP | -0.17 | 0.00846 | NA | |
| 116 | hsa-miR-30c-5p | PSEN2 | -0.43 | 0.00016 | 0.39 | 0.00069 | miRNATAP | -0.19 | 0.00011 | NA | |
| 117 | hsa-miR-421 | PSEN2 | 0.94 | 0 | 0.39 | 0.00069 | miRanda | -0.11 | 0.00061 | NA | |
| 118 | hsa-miR-576-5p | RBPJ | -0.38 | 0.00471 | 0.08 | 0.47404 | mirMAP | -0.16 | 2.0E-5 | NA | |
| 119 | hsa-miR-616-5p | RBPJ | 0.15 | 0.40284 | 0.08 | 0.47404 | MirTarget | -0.14 | 0 | NA | |
| 120 | hsa-miR-664a-3p | RBPJ | 0.49 | 0.00073 | 0.08 | 0.47404 | mirMAP | -0.15 | 1.0E-5 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | NOTCH SIGNALING PATHWAY | 18 | 114 | 1.916e-33 | 8.917e-30 |
| 2 | NOTCH RECEPTOR PROCESSING | 8 | 16 | 1.588e-19 | 3.695e-16 |
| 3 | REGULATION OF NOTCH SIGNALING PATHWAY | 9 | 67 | 5.778e-16 | 8.962e-13 |
| 4 | EPITHELIUM DEVELOPMENT | 17 | 945 | 3.597e-15 | 4.184e-12 |
| 5 | POSITIVE REGULATION OF NOTCH SIGNALING PATHWAY | 7 | 34 | 5.42e-14 | 5.044e-11 |
| 6 | MORPHOGENESIS OF AN EPITHELIUM | 12 | 400 | 3.483e-13 | 2.701e-10 |
| 7 | TISSUE DEVELOPMENT | 18 | 1518 | 5.036e-13 | 3.347e-10 |
| 8 | EMBRYO DEVELOPMENT | 15 | 894 | 7.784e-13 | 4.527e-10 |
| 9 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 15 | 957 | 2.074e-12 | 1.072e-09 |
| 10 | ORGAN MORPHOGENESIS | 14 | 841 | 6.615e-12 | 3.078e-09 |
| 11 | TISSUE MORPHOGENESIS | 12 | 533 | 1.01e-11 | 4.274e-09 |
| 12 | TUBE DEVELOPMENT | 12 | 552 | 1.519e-11 | 5.889e-09 |
| 13 | LYMPHOCYTE DIFFERENTIATION | 9 | 209 | 2.066e-11 | 7.394e-09 |
| 14 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 13 | 750 | 2.891e-11 | 8.968e-09 |
| 15 | CELL SURFACE RECEPTOR SIGNALING PATHWAY INVOLVED IN HEART DEVELOPMENT | 5 | 16 | 2.752e-11 | 8.968e-09 |
| 16 | MORPHOGENESIS OF AN EPITHELIAL SHEET | 6 | 43 | 4.857e-11 | 1.412e-08 |
| 17 | HEART DEVELOPMENT | 11 | 466 | 5.435e-11 | 1.488e-08 |
| 18 | POSITIVE REGULATION OF GENE EXPRESSION | 17 | 1733 | 6.621e-11 | 1.621e-08 |
| 19 | B CELL DIFFERENTIATION | 7 | 89 | 6.551e-11 | 1.621e-08 |
| 20 | POSITIVE REGULATION OF CELL COMMUNICATION | 16 | 1532 | 1.295e-10 | 3.013e-08 |
| 21 | EMBRYONIC MORPHOGENESIS | 11 | 539 | 2.558e-10 | 5.667e-08 |
| 22 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 17 | 1929 | 3.582e-10 | 7.576e-08 |
| 23 | LEUKOCYTE DIFFERENTIATION | 9 | 292 | 4.055e-10 | 8.203e-08 |
| 24 | PATTERN SPECIFICATION PROCESS | 10 | 418 | 4.282e-10 | 8.301e-08 |
| 25 | EAR DEVELOPMENT | 8 | 195 | 4.604e-10 | 8.498e-08 |
| 26 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 16 | 1672 | 4.749e-10 | 8.498e-08 |
| 27 | IMMUNE SYSTEM DEVELOPMENT | 11 | 582 | 5.759e-10 | 9.924e-08 |
| 28 | HEART MORPHOGENESIS | 8 | 212 | 8.935e-10 | 1.414e-07 |
| 29 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 12 | 788 | 9.116e-10 | 1.414e-07 |
| 30 | CIRCULATORY SYSTEM DEVELOPMENT | 12 | 788 | 9.116e-10 | 1.414e-07 |
| 31 | TUBE MORPHOGENESIS | 9 | 323 | 9.865e-10 | 1.481e-07 |
| 32 | B CELL ACTIVATION | 7 | 132 | 1.071e-09 | 1.557e-07 |
| 33 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 16 | 1805 | 1.467e-09 | 2.069e-07 |
| 34 | CELL FATE COMMITMENT | 8 | 227 | 1.534e-09 | 2.099e-07 |
| 35 | LYMPHOCYTE ACTIVATION | 9 | 342 | 1.629e-09 | 2.166e-07 |
| 36 | SENSORY ORGAN DEVELOPMENT | 10 | 493 | 2.108e-09 | 2.725e-07 |
| 37 | MATURE B CELL DIFFERENTIATION INVOLVED IN IMMUNE RESPONSE | 4 | 13 | 3.343e-09 | 4.204e-07 |
| 38 | SEGMENTATION | 6 | 89 | 4.406e-09 | 5.396e-07 |
| 39 | ENDOTHELIUM DEVELOPMENT | 6 | 90 | 4.716e-09 | 5.627e-07 |
| 40 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 11 | 724 | 5.667e-09 | 6.592e-07 |
| 41 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 10 | 554 | 6.444e-09 | 7.139e-07 |
| 42 | EPITHELIAL CELL FATE COMMITMENT | 4 | 15 | 6.368e-09 | 7.139e-07 |
| 43 | LEUKOCYTE ACTIVATION | 9 | 414 | 8.619e-09 | 9.159e-07 |
| 44 | REGULATION OF BINDING | 8 | 283 | 8.661e-09 | 9.159e-07 |
| 45 | CARDIAC CHAMBER MORPHOGENESIS | 6 | 104 | 1.132e-08 | 1.145e-06 |
| 46 | MATURE B CELL DIFFERENTIATION | 4 | 17 | 1.108e-08 | 1.145e-06 |
| 47 | REGULATION OF CELL DIFFERENTIATION | 14 | 1492 | 1.232e-08 | 1.22e-06 |
| 48 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 12 | 1004 | 1.396e-08 | 1.353e-06 |
| 49 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 12 | 1021 | 1.683e-08 | 1.598e-06 |
| 50 | REGIONALIZATION | 8 | 311 | 1.806e-08 | 1.681e-06 |
| 51 | REGULATION OF EMBRYONIC DEVELOPMENT | 6 | 114 | 1.969e-08 | 1.796e-06 |
| 52 | REGULATION OF CELL DEVELOPMENT | 11 | 836 | 2.506e-08 | 2.242e-06 |
| 53 | POSITIVE REGULATION OF BINDING | 6 | 127 | 3.763e-08 | 3.304e-06 |
| 54 | SOMITOGENESIS | 5 | 62 | 3.878e-08 | 3.341e-06 |
| 55 | EPITHELIAL CELL DIFFERENTIATION | 9 | 495 | 4.025e-08 | 3.405e-06 |
| 56 | NEUROGENESIS | 13 | 1402 | 5.837e-08 | 4.765e-06 |
| 57 | NEURON FATE COMMITMENT | 5 | 67 | 5.756e-08 | 4.765e-06 |
| 58 | SENSORY ORGAN MORPHOGENESIS | 7 | 239 | 6.564e-08 | 5.266e-06 |
| 59 | EPIDERMAL CELL DIFFERENTIATION | 6 | 142 | 7.328e-08 | 5.78e-06 |
| 60 | SKIN EPIDERMIS DEVELOPMENT | 5 | 71 | 7.727e-08 | 5.992e-06 |
| 61 | CARDIAC CHAMBER DEVELOPMENT | 6 | 144 | 7.965e-08 | 6.075e-06 |
| 62 | ENDOTHELIAL CELL DIFFERENTIATION | 5 | 72 | 8.294e-08 | 6.225e-06 |
| 63 | AUDITORY RECEPTOR CELL DIFFERENTIATION | 4 | 28 | 9.419e-08 | 6.957e-06 |
| 64 | EPIDERMIS DEVELOPMENT | 7 | 253 | 9.682e-08 | 7.039e-06 |
| 65 | REGULATION OF NEURON DIFFERENTIATION | 9 | 554 | 1.053e-07 | 7.538e-06 |
| 66 | TRANSCRIPTION INITIATION FROM RNA POLYMERASE II PROMOTER | 6 | 153 | 1.142e-07 | 8.051e-06 |
| 67 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 14 | 1784 | 1.184e-07 | 8.225e-06 |
| 68 | SOMITE DEVELOPMENT | 5 | 78 | 1.243e-07 | 8.446e-06 |
| 69 | REGULATION OF CELL PROLIFERATION | 13 | 1496 | 1.253e-07 | 8.446e-06 |
| 70 | CELL ACTIVATION | 9 | 568 | 1.302e-07 | 8.653e-06 |
| 71 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER IN RESPONSE TO HYPOXIA | 4 | 32 | 1.647e-07 | 1.079e-05 |
| 72 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 10 | 801 | 2.071e-07 | 1.339e-05 |
| 73 | HAIR CELL DIFFERENTIATION | 4 | 35 | 2.391e-07 | 1.503e-05 |
| 74 | B CELL ACTIVATION INVOLVED IN IMMUNE RESPONSE | 4 | 35 | 2.391e-07 | 1.503e-05 |
| 75 | RESPONSE TO OXYGEN LEVELS | 7 | 311 | 3.923e-07 | 2.434e-05 |
| 76 | ANTERIOR POSTERIOR PATTERN SPECIFICATION | 6 | 194 | 4.638e-07 | 2.803e-05 |
| 77 | NEURON DIFFERENTIATION | 10 | 874 | 4.633e-07 | 2.803e-05 |
| 78 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 12 | 1395 | 5.132e-07 | 3.061e-05 |
| 79 | CELL FATE DETERMINATION | 4 | 43 | 5.586e-07 | 3.249e-05 |
| 80 | VENTRICULAR TRABECULA MYOCARDIUM MORPHOGENESIS | 3 | 11 | 5.517e-07 | 3.249e-05 |
| 81 | DNA TEMPLATED TRANSCRIPTION INITIATION | 6 | 202 | 5.879e-07 | 3.377e-05 |
| 82 | REGULATION OF TIMING OF CELL DIFFERENTIATION | 3 | 12 | 7.348e-07 | 4.07e-05 |
| 83 | COLUMNAR CUBOIDAL EPITHELIAL CELL DIFFERENTIATION | 5 | 111 | 7.283e-07 | 4.07e-05 |
| 84 | REGULATION OF DEVELOPMENT HETEROCHRONIC | 3 | 12 | 7.348e-07 | 4.07e-05 |
| 85 | SKIN DEVELOPMENT | 6 | 211 | 7.587e-07 | 4.153e-05 |
| 86 | COVALENT CHROMATIN MODIFICATION | 7 | 345 | 7.876e-07 | 4.261e-05 |
| 87 | CARDIAC SEPTUM MORPHOGENESIS | 4 | 49 | 9.528e-07 | 5.096e-05 |
| 88 | ARTERY MORPHOGENESIS | 4 | 51 | 1.121e-06 | 5.863e-05 |
| 89 | MECHANORECEPTOR DIFFERENTIATION | 4 | 51 | 1.121e-06 | 5.863e-05 |
| 90 | REGULATION OF ESTABLISHMENT OF PLANAR POLARITY INVOLVED IN NEURAL TUBE CLOSURE | 3 | 14 | 1.213e-06 | 6.203e-05 |
| 91 | NEGATIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 9 | 740 | 1.2e-06 | 6.203e-05 |
| 92 | REGULATION OF CANONICAL WNT SIGNALING PATHWAY | 6 | 236 | 1.457e-06 | 7.369e-05 |
| 93 | N TERMINAL PROTEIN AMINO ACID ACETYLATION | 3 | 15 | 1.515e-06 | 7.58e-05 |
| 94 | TUBE FORMATION | 5 | 129 | 1.534e-06 | 7.592e-05 |
| 95 | OUTFLOW TRACT MORPHOGENESIS | 4 | 56 | 1.639e-06 | 8.029e-05 |
| 96 | REGULATION OF ORGAN MORPHOGENESIS | 6 | 242 | 1.686e-06 | 8.17e-05 |
| 97 | MAINTENANCE OF CELL NUMBER | 5 | 132 | 1.718e-06 | 8.241e-05 |
| 98 | MORPHOGENESIS OF EMBRYONIC EPITHELIUM | 5 | 134 | 1.85e-06 | 8.754e-05 |
| 99 | MORPHOGENESIS OF AN ENDOTHELIUM | 3 | 16 | 1.863e-06 | 8.754e-05 |
| 100 | MEMBRANE PROTEIN INTRACELLULAR DOMAIN PROTEOLYSIS | 3 | 17 | 2.259e-06 | 0.0001051 |
| 101 | EMBRYONIC DIGIT MORPHOGENESIS | 4 | 61 | 2.317e-06 | 0.0001067 |
| 102 | CARDIAC VENTRICLE MORPHOGENESIS | 4 | 62 | 2.474e-06 | 0.0001128 |
| 103 | NEUROEPITHELIAL CELL DIFFERENTIATION | 4 | 63 | 2.639e-06 | 0.0001192 |
| 104 | REGULATION OF PROTEIN ACETYLATION | 4 | 64 | 2.811e-06 | 0.0001258 |
| 105 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 9 | 823 | 2.885e-06 | 0.0001279 |
| 106 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 8 | 609 | 2.997e-06 | 0.0001315 |
| 107 | NEURAL TUBE DEVELOPMENT | 5 | 149 | 3.118e-06 | 0.0001356 |
| 108 | IMMUNE SYSTEM PROCESS | 13 | 1984 | 3.208e-06 | 0.0001371 |
| 109 | NEURONAL STEM CELL POPULATION MAINTENANCE | 3 | 19 | 3.213e-06 | 0.0001371 |
| 110 | REGULATION OF DNA TEMPLATED TRANSCRIPTION IN RESPONSE TO STRESS | 4 | 67 | 3.381e-06 | 0.000143 |
| 111 | REGULATION OF MUSCLE CELL DIFFERENTIATION | 5 | 152 | 3.439e-06 | 0.0001441 |
| 112 | MUSCLE TISSUE DEVELOPMENT | 6 | 275 | 3.529e-06 | 0.0001453 |
| 113 | MUSCLE STRUCTURE DEVELOPMENT | 7 | 432 | 3.509e-06 | 0.0001453 |
| 114 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 7 | 437 | 3.785e-06 | 0.0001545 |
| 115 | COCHLEA MORPHOGENESIS | 3 | 21 | 4.4e-06 | 0.000178 |
| 116 | NEGATIVE REGULATION OF CANONICAL WNT SIGNALING PATHWAY | 5 | 162 | 4.697e-06 | 0.0001884 |
| 117 | BETA CATENIN DESTRUCTION COMPLEX DISASSEMBLY | 3 | 22 | 5.09e-06 | 0.0001957 |
| 118 | MEMBRANE PROTEIN ECTODOMAIN PROTEOLYSIS | 3 | 22 | 5.09e-06 | 0.0001957 |
| 119 | SOMATIC STEM CELL DIVISION | 3 | 22 | 5.09e-06 | 0.0001957 |
| 120 | ANGIOGENESIS | 6 | 293 | 5.081e-06 | 0.0001957 |
| 121 | AORTA MORPHOGENESIS | 3 | 22 | 5.09e-06 | 0.0001957 |
| 122 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 10 | 1142 | 5.214e-06 | 0.0001973 |
| 123 | CELL DEVELOPMENT | 11 | 1426 | 5.201e-06 | 0.0001973 |
| 124 | ARTERY DEVELOPMENT | 4 | 75 | 5.315e-06 | 0.0001994 |
| 125 | RHYTHMIC PROCESS | 6 | 298 | 5.599e-06 | 0.0002084 |
| 126 | POSITIVE REGULATION OF CELL DEVELOPMENT | 7 | 472 | 6.277e-06 | 0.0002318 |
| 127 | REGULATION OF WNT SIGNALING PATHWAY | 6 | 310 | 7.018e-06 | 0.0002571 |
| 128 | HAIR CYCLE | 4 | 83 | 7.966e-06 | 0.0002873 |
| 129 | MOLTING CYCLE | 4 | 83 | 7.966e-06 | 0.0002873 |
| 130 | HEART TRABECULA MORPHOGENESIS | 3 | 26 | 8.557e-06 | 0.0003039 |
| 131 | N TERMINAL PROTEIN AMINO ACID MODIFICATION | 3 | 26 | 8.557e-06 | 0.0003039 |
| 132 | CARDIAC SEPTUM DEVELOPMENT | 4 | 85 | 8.758e-06 | 0.0003064 |
| 133 | REGULATION OF STRIATED MUSCLE CELL DIFFERENTIATION | 4 | 85 | 8.758e-06 | 0.0003064 |
| 134 | TISSUE REMODELING | 4 | 87 | 9.607e-06 | 0.0003315 |
| 135 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER INVOLVED IN CELLULAR RESPONSE TO CHEMICAL STIMULUS | 3 | 27 | 9.617e-06 | 0.0003315 |
| 136 | NEGATIVE REGULATION OF NEURON DIFFERENTIATION | 5 | 191 | 1.047e-05 | 0.0003581 |
| 137 | NEGATIVE REGULATION OF NOTCH SIGNALING PATHWAY | 3 | 28 | 1.076e-05 | 0.0003616 |
| 138 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 7 | 513 | 1.08e-05 | 0.0003616 |
| 139 | REGULATION OF SPROUTING ANGIOGENESIS | 3 | 28 | 1.076e-05 | 0.0003616 |
| 140 | STEM CELL DIVISION | 3 | 29 | 1.199e-05 | 0.0003984 |
| 141 | NEGATIVE REGULATION OF WNT SIGNALING PATHWAY | 5 | 197 | 1.216e-05 | 0.0004013 |
| 142 | NEURAL TUBE FORMATION | 4 | 94 | 1.306e-05 | 0.000428 |
| 143 | CARDIOCYTE DIFFERENTIATION | 4 | 96 | 1.42e-05 | 0.000462 |
| 144 | CARDIAC ATRIUM DEVELOPMENT | 3 | 31 | 1.472e-05 | 0.0004755 |
| 145 | CHROMATIN MODIFICATION | 7 | 539 | 1.488e-05 | 0.0004776 |
| 146 | LYMPHOCYTE ACTIVATION INVOLVED IN IMMUNE RESPONSE | 4 | 98 | 1.541e-05 | 0.000491 |
| 147 | ENDOCARDIAL CUSHION DEVELOPMENT | 3 | 32 | 1.622e-05 | 0.00051 |
| 148 | BLOOD VESSEL REMODELING | 3 | 32 | 1.622e-05 | 0.00051 |
| 149 | BLOOD VESSEL MORPHOGENESIS | 6 | 364 | 1.748e-05 | 0.0005458 |
| 150 | HEART VALVE DEVELOPMENT | 3 | 34 | 1.953e-05 | 0.0006058 |
| 151 | CARDIAC VENTRICLE DEVELOPMENT | 4 | 106 | 2.101e-05 | 0.0006473 |
| 152 | MEMBRANE PROTEIN PROTEOLYSIS | 3 | 35 | 2.134e-05 | 0.0006532 |
| 153 | POSITIVE REGULATION OF PROTEIN ACETYLATION | 3 | 36 | 2.325e-05 | 0.0007072 |
| 154 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 10 | 1360 | 2.42e-05 | 0.0007313 |
| 155 | MYOBLAST DIFFERENTIATION | 3 | 37 | 2.528e-05 | 0.000754 |
| 156 | NEGATIVE REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 3 | 37 | 2.528e-05 | 0.000754 |
| 157 | TRABECULA MORPHOGENESIS | 3 | 39 | 2.967e-05 | 0.0008738 |
| 158 | COCHLEA DEVELOPMENT | 3 | 39 | 2.967e-05 | 0.0008738 |
| 159 | POSITIVE REGULATION OF GROWTH | 5 | 238 | 3.02e-05 | 0.0008837 |
| 160 | EMBRYONIC ORGAN DEVELOPMENT | 6 | 406 | 3.23e-05 | 0.0009393 |
| 161 | AORTA DEVELOPMENT | 3 | 41 | 3.454e-05 | 0.0009981 |
| 162 | REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 4 | 122 | 3.65e-05 | 0.001048 |
| 163 | T CELL DIFFERENTIATION | 4 | 123 | 3.769e-05 | 0.001076 |
| 164 | BETA CATENIN TCF COMPLEX ASSEMBLY | 3 | 43 | 3.99e-05 | 0.001132 |
| 165 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 11 | 1791 | 4.5e-05 | 0.001269 |
| 166 | PROTEIN ACETYLATION | 4 | 129 | 4.541e-05 | 0.001273 |
| 167 | VENTRICULAR CARDIAC MUSCLE TISSUE DEVELOPMENT | 3 | 45 | 4.578e-05 | 0.001276 |
| 168 | NEGATIVE REGULATION OF CELL PROLIFERATION | 7 | 643 | 4.609e-05 | 0.001277 |
| 169 | NEGATIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 5 | 262 | 4.775e-05 | 0.001309 |
| 170 | REGULATION OF CELL DEATH | 10 | 1472 | 4.783e-05 | 0.001309 |
| 171 | HISTONE H3 ACETYLATION | 3 | 46 | 4.892e-05 | 0.001331 |
| 172 | CHROMATIN ORGANIZATION | 7 | 663 | 5.596e-05 | 0.001511 |
| 173 | NEGATIVE REGULATION OF CELL COMMUNICATION | 9 | 1192 | 5.618e-05 | 0.001511 |
| 174 | CELL ACTIVATION INVOLVED IN IMMUNE RESPONSE | 4 | 139 | 6.076e-05 | 0.001625 |
| 175 | MUSCLE ORGAN DEVELOPMENT | 5 | 277 | 6.218e-05 | 0.001652 |
| 176 | CARDIAC MUSCLE TISSUE DEVELOPMENT | 4 | 140 | 6.248e-05 | 0.001652 |
| 177 | EMBRYONIC ORGAN MORPHOGENESIS | 5 | 279 | 6.434e-05 | 0.001691 |
| 178 | CELLULAR RESPONSE TO OXYGEN LEVELS | 4 | 143 | 6.786e-05 | 0.001774 |
| 179 | VASCULATURE DEVELOPMENT | 6 | 469 | 7.205e-05 | 0.001873 |
| 180 | CARDIAC MUSCLE TISSUE MORPHOGENESIS | 3 | 54 | 7.927e-05 | 0.002049 |
| 181 | CHROMATIN REMODELING | 4 | 150 | 8.169e-05 | 0.0021 |
| 182 | NEGATIVE REGULATION OF MUSCLE CELL DIFFERENTIATION | 3 | 55 | 8.376e-05 | 0.002141 |
| 183 | IMMUNE EFFECTOR PROCESS | 6 | 486 | 8.768e-05 | 0.002229 |
| 184 | POSITIVE REGULATION OF MUSCLE TISSUE DEVELOPMENT | 3 | 56 | 8.841e-05 | 0.002236 |
| 185 | NEGATIVE REGULATION OF CELL DEVELOPMENT | 5 | 303 | 9.496e-05 | 0.002354 |
| 186 | POSITIVE REGULATION OF DEVELOPMENTAL GROWTH | 4 | 156 | 9.51e-05 | 0.002354 |
| 187 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 8 | 983 | 9.494e-05 | 0.002354 |
| 188 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 5 | 303 | 9.496e-05 | 0.002354 |
| 189 | PROTEIN ACYLATION | 4 | 159 | 0.0001024 | 0.00252 |
| 190 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 4 | 162 | 0.00011 | 0.002695 |
| 191 | REGULATION OF VIRAL TRANSCRIPTION | 3 | 61 | 0.0001142 | 0.002782 |
| 192 | REGULATION OF PROTEIN BINDING | 4 | 168 | 0.0001266 | 0.002979 |
| 193 | LOOP OF HENLE DEVELOPMENT | 2 | 11 | 0.0001268 | 0.002979 |
| 194 | ENDOCARDIUM DEVELOPMENT | 2 | 11 | 0.0001268 | 0.002979 |
| 195 | NEGATIVE REGULATION OF SPROUTING ANGIOGENESIS | 2 | 11 | 0.0001268 | 0.002979 |
| 196 | POSITIVE REGULATION OF RECEPTOR BIOSYNTHETIC PROCESS | 2 | 11 | 0.0001268 | 0.002979 |
| 197 | EPITHELIAL TO MESENCHYMAL TRANSITION INVOLVED IN ENDOCARDIAL CUSHION FORMATION | 2 | 11 | 0.0001268 | 0.002979 |
| 198 | ENDOTHELIAL TUBE MORPHOGENESIS | 2 | 11 | 0.0001268 | 0.002979 |
| 199 | APPENDAGE DEVELOPMENT | 4 | 169 | 0.0001295 | 0.003014 |
| 200 | LIMB DEVELOPMENT | 4 | 169 | 0.0001295 | 0.003014 |
| 201 | DEVELOPMENTAL GROWTH | 5 | 333 | 0.0001478 | 0.003422 |
| 202 | CARDIAC LEFT VENTRICLE MORPHOGENESIS | 2 | 12 | 0.000152 | 0.00345 |
| 203 | AMYLOID PRECURSOR PROTEIN METABOLIC PROCESS | 2 | 12 | 0.000152 | 0.00345 |
| 204 | DISTAL TUBULE DEVELOPMENT | 2 | 12 | 0.000152 | 0.00345 |
| 205 | LATERAL VENTRICLE DEVELOPMENT | 2 | 12 | 0.000152 | 0.00345 |
| 206 | MUSCLE ORGAN MORPHOGENESIS | 3 | 70 | 0.000172 | 0.003886 |
| 207 | NEURONAL STEM CELL DIVISION | 2 | 13 | 0.0001794 | 0.003902 |
| 208 | EYELID DEVELOPMENT IN CAMERA TYPE EYE | 2 | 13 | 0.0001794 | 0.003902 |
| 209 | LEFT RIGHT AXIS SPECIFICATION | 2 | 13 | 0.0001794 | 0.003902 |
| 210 | CARDIOBLAST DIFFERENTIATION | 2 | 13 | 0.0001794 | 0.003902 |
| 211 | NEGATIVE REGULATION OF EPIDERMAL CELL DIFFERENTIATION | 2 | 13 | 0.0001794 | 0.003902 |
| 212 | NEUROBLAST DIVISION | 2 | 13 | 0.0001794 | 0.003902 |
| 213 | CELL FATE SPECIFICATION | 3 | 71 | 0.0001794 | 0.003902 |
| 214 | POSITIVE REGULATION OF GLUCONEOGENESIS | 2 | 13 | 0.0001794 | 0.003902 |
| 215 | EPITHELIAL CELL DEVELOPMENT | 4 | 186 | 0.0001872 | 0.004033 |
| 216 | REGULATION OF CELL PROJECTION ORGANIZATION | 6 | 558 | 0.0001865 | 0.004033 |
| 217 | POSITIVE REGULATION OF PROTEIN BINDING | 3 | 73 | 0.0001949 | 0.004178 |
| 218 | REGULATION OF CYTOKINE PRODUCTION | 6 | 563 | 0.0001958 | 0.004179 |
| 219 | POSITIVE REGULATION OF CELL PROLIFERATION | 7 | 814 | 0.0002014 | 0.004278 |
| 220 | FOREBRAIN DEVELOPMENT | 5 | 357 | 0.0002043 | 0.004322 |
| 221 | IMMUNE RESPONSE | 8 | 1100 | 0.0002069 | 0.004357 |
| 222 | CONVERGENT EXTENSION | 2 | 14 | 0.0002091 | 0.004384 |
| 223 | ODONTOGENESIS OF DENTIN CONTAINING TOOTH | 3 | 75 | 0.0002111 | 0.004405 |
| 224 | POSITIVE REGULATION OF GENE EXPRESSION EPIGENETIC | 3 | 78 | 0.0002371 | 0.004835 |
| 225 | RESPONSE TO MURAMYL DIPEPTIDE | 2 | 15 | 0.0002411 | 0.004835 |
| 226 | ENDOCARDIAL CUSHION FORMATION | 2 | 15 | 0.0002411 | 0.004835 |
| 227 | RENAL TUBULE DEVELOPMENT | 3 | 78 | 0.0002371 | 0.004835 |
| 228 | ECTODERMAL PLACODE DEVELOPMENT | 2 | 15 | 0.0002411 | 0.004835 |
| 229 | REGULATION OF CELL PROLIFERATION INVOLVED IN HEART MORPHOGENESIS | 2 | 15 | 0.0002411 | 0.004835 |
| 230 | RESPIRATORY SYSTEM DEVELOPMENT | 4 | 197 | 0.0002333 | 0.004835 |
| 231 | ECTODERMAL PLACODE MORPHOGENESIS | 2 | 15 | 0.0002411 | 0.004835 |
| 232 | ECTODERMAL PLACODE FORMATION | 2 | 15 | 0.0002411 | 0.004835 |
| 233 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 8 | 1135 | 0.0002564 | 0.00512 |
| 234 | CARDIAC RIGHT VENTRICLE MORPHOGENESIS | 2 | 16 | 0.0002753 | 0.005427 |
| 235 | NEGATIVE REGULATION OF EPIDERMIS DEVELOPMENT | 2 | 16 | 0.0002753 | 0.005427 |
| 236 | APOPTOTIC PROCESS INVOLVED IN MORPHOGENESIS | 2 | 16 | 0.0002753 | 0.005427 |
| 237 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 8 | 1152 | 0.0002837 | 0.00557 |
| 238 | POSITIVE REGULATION OF CHROMATIN MODIFICATION | 3 | 85 | 0.0003056 | 0.005949 |
| 239 | NEGATIVE REGULATION OF CELL DEATH | 7 | 872 | 0.0003068 | 0.005949 |
| 240 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 7 | 872 | 0.0003068 | 0.005949 |
| 241 | REGULATION OF CHROMATIN BINDING | 2 | 17 | 0.0003117 | 0.005979 |
| 242 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 9 | 1492 | 0.0003122 | 0.005979 |
| 243 | SEGMENT SPECIFICATION | 2 | 17 | 0.0003117 | 0.005979 |
| 244 | NEGATIVE REGULATION OF GENE EXPRESSION | 9 | 1493 | 0.0003138 | 0.005984 |
| 245 | NEGATIVE REGULATION OF MYOTUBE DIFFERENTIATION | 2 | 18 | 0.0003503 | 0.006625 |
| 246 | PERICARDIUM DEVELOPMENT | 2 | 18 | 0.0003503 | 0.006625 |
| 247 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 9 | 1517 | 0.0003534 | 0.006658 |
| 248 | REGULATION OF PROTEIN STABILITY | 4 | 221 | 0.0003612 | 0.006777 |
| 249 | CELLULAR COMPONENT MORPHOGENESIS | 7 | 900 | 0.0003717 | 0.006947 |
| 250 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 5 | 408 | 0.0003782 | 0.00704 |
| 251 | GROWTH | 5 | 410 | 0.0003868 | 0.007142 |
| 252 | INNER EAR MORPHOGENESIS | 3 | 92 | 0.0003857 | 0.007142 |
| 253 | REGULATION OF CELL MIGRATION INVOLVED IN SPROUTING ANGIOGENESIS | 2 | 19 | 0.0003911 | 0.007165 |
| 254 | POSITIVE REGULATION OF CARDIAC MUSCLE CELL PROLIFERATION | 2 | 19 | 0.0003911 | 0.007165 |
| 255 | NEPHRON EPITHELIUM DEVELOPMENT | 3 | 93 | 0.0003981 | 0.007236 |
| 256 | REGULATION OF DNA BINDING | 3 | 93 | 0.0003981 | 0.007236 |
| 257 | REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 4 | 229 | 0.0004132 | 0.007452 |
| 258 | REGULATION OF GENE EXPRESSION EPIGENETIC | 4 | 229 | 0.0004132 | 0.007452 |
| 259 | CANONICAL WNT SIGNALING PATHWAY | 3 | 95 | 0.0004238 | 0.007613 |
| 260 | TONGUE DEVELOPMENT | 2 | 20 | 0.0004342 | 0.00774 |
| 261 | POSITIVE REGULATION OF NEURON PROJECTION DEVELOPMENT | 4 | 232 | 0.000434 | 0.00774 |
| 262 | INTERSPECIES INTERACTION BETWEEN ORGANISMS | 6 | 662 | 0.0004666 | 0.008255 |
| 263 | SYMBIOSIS ENCOMPASSING MUTUALISM THROUGH PARASITISM | 6 | 662 | 0.0004666 | 0.008255 |
| 264 | MUSCLE CELL DIFFERENTIATION | 4 | 237 | 0.0004704 | 0.008291 |
| 265 | LEFT RIGHT PATTERN FORMATION | 2 | 21 | 0.0004794 | 0.008354 |
| 266 | REGULATION OF RECEPTOR BIOSYNTHETIC PROCESS | 2 | 21 | 0.0004794 | 0.008354 |
| 267 | APOPTOTIC PROCESS INVOLVED IN DEVELOPMENT | 2 | 21 | 0.0004794 | 0.008354 |
| 268 | POSITIVE REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 3 | 100 | 0.0004925 | 0.00855 |
| 269 | KERATINOCYTE DIFFERENTIATION | 3 | 101 | 0.000507 | 0.008737 |
| 270 | CELL PROLIFERATION | 6 | 672 | 0.0005052 | 0.008737 |
| 271 | ENDOCARDIAL CUSHION MORPHOGENESIS | 2 | 22 | 0.0005268 | 0.008946 |
| 272 | REGULATION OF ANDROGEN RECEPTOR SIGNALING PATHWAY | 2 | 22 | 0.0005268 | 0.008946 |
| 273 | MODULATION OF TRANSCRIPTION IN OTHER ORGANISM INVOLVED IN SYMBIOTIC INTERACTION | 2 | 22 | 0.0005268 | 0.008946 |
| 274 | HISTONE H3 DEACETYLATION | 2 | 22 | 0.0005268 | 0.008946 |
| 275 | REGULATION OF MUSCLE TISSUE DEVELOPMENT | 3 | 103 | 0.0005369 | 0.009052 |
| 276 | REGULATION OF MUSCLE ORGAN DEVELOPMENT | 3 | 103 | 0.0005369 | 0.009052 |
| 277 | ODONTOGENESIS | 3 | 105 | 0.0005679 | 0.00954 |
| 278 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL METABOLIC PROCESS | 2 | 23 | 0.0005764 | 0.009579 |
| 279 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER IN RESPONSE TO STRESS | 2 | 23 | 0.0005764 | 0.009579 |
| 280 | POSITIVE REGULATION OF COLLAGEN METABOLIC PROCESS | 2 | 23 | 0.0005764 | 0.009579 |
| 281 | SPINAL CORD DEVELOPMENT | 3 | 106 | 0.0005839 | 0.009668 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | NOTCH BINDING | 8 | 18 | 5.389e-19 | 5.006e-16 |
| 2 | TRANSCRIPTION FACTOR BINDING | 10 | 524 | 3.784e-09 | 1.758e-06 |
| 3 | TRANSCRIPTION FACTOR ACTIVITY PROTEIN BINDING | 10 | 588 | 1.136e-08 | 2.458e-06 |
| 4 | CHROMATIN BINDING | 9 | 435 | 1.323e-08 | 2.458e-06 |
| 5 | RECEPTOR BINDING | 14 | 1476 | 1.073e-08 | 2.458e-06 |
| 6 | RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 5 | 104 | 5.265e-07 | 8.153e-05 |
| 7 | CORE PROMOTER BINDING | 5 | 152 | 3.439e-06 | 0.0003993 |
| 8 | PEPTIDE N ACETYLTRANSFERASE ACTIVITY | 4 | 67 | 3.381e-06 | 0.0003993 |
| 9 | TRANSCRIPTION COACTIVATOR ACTIVITY | 6 | 296 | 5.387e-06 | 0.000556 |
| 10 | CHROMATIN DNA BINDING | 4 | 80 | 6.878e-06 | 0.000639 |
| 11 | BETA CATENIN BINDING | 4 | 84 | 8.355e-06 | 0.0006872 |
| 12 | RNA POLYMERASE II REPRESSING TRANSCRIPTION FACTOR BINDING | 3 | 27 | 9.617e-06 | 0.0006872 |
| 13 | N ACETYLTRANSFERASE ACTIVITY | 4 | 87 | 9.607e-06 | 0.0006872 |
| 14 | NF KAPPAB BINDING | 3 | 30 | 1.331e-05 | 0.000883 |
| 15 | HISTONE DEACETYLASE BINDING | 4 | 105 | 2.023e-05 | 0.001106 |
| 16 | ACETYLTRANSFERASE ACTIVITY | 4 | 103 | 1.876e-05 | 0.001106 |
| 17 | N ACYLTRANSFERASE ACTIVITY | 4 | 104 | 1.949e-05 | 0.001106 |
| 18 | FRIZZLED BINDING | 3 | 36 | 2.325e-05 | 0.0012 |
| 19 | RAC GTPASE BINDING | 3 | 39 | 2.967e-05 | 0.001417 |
| 20 | MACROMOLECULAR COMPLEX BINDING | 10 | 1399 | 3.091e-05 | 0.001417 |
| 21 | STRUCTURE SPECIFIC DNA BINDING | 4 | 118 | 3.203e-05 | 0.001417 |
| 22 | ENZYME BINDING | 11 | 1737 | 3.386e-05 | 0.00143 |
| 23 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 4 | 133 | 5.116e-05 | 0.002066 |
| 24 | CALCIUM ION BINDING | 7 | 697 | 7.67e-05 | 0.002969 |
| 25 | ACTIVATING TRANSCRIPTION FACTOR BINDING | 3 | 57 | 9.322e-05 | 0.003331 |
| 26 | REPRESSING TRANSCRIPTION FACTOR BINDING | 3 | 57 | 9.322e-05 | 0.003331 |
| 27 | HISTONE DEACETYLASE ACTIVITY H3 K14 SPECIFIC | 2 | 12 | 0.000152 | 0.005229 |
| 28 | REGULATORY REGION NUCLEIC ACID BINDING | 7 | 818 | 0.0002075 | 0.006885 |
| 29 | RHO GTPASE BINDING | 3 | 78 | 0.0002371 | 0.007318 |
| 30 | PEROXISOME PROLIFERATOR ACTIVATED RECEPTOR BINDING | 2 | 15 | 0.0002411 | 0.007318 |
| 31 | CORE PROMOTER PROXIMAL REGION DNA BINDING | 5 | 371 | 0.0002442 | 0.007318 |
| 32 | ZINC ION BINDING | 8 | 1155 | 0.0002888 | 0.008384 |
| 33 | TRANSFERASE ACTIVITY TRANSFERRING ACYL GROUPS OTHER THAN AMINO ACYL GROUPS | 4 | 211 | 0.000303 | 0.008395 |
| 34 | TRANSCRIPTIONAL REPRESSOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 3 | 86 | 0.0003163 | 0.008395 |
| 35 | NAD DEPENDENT PROTEIN DEACETYLASE ACTIVITY | 2 | 17 | 0.0003117 | 0.008395 |
| 36 | LIGASE ACTIVITY | 5 | 406 | 0.0003698 | 0.009543 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CHROMATIN | 8 | 441 | 2.651e-07 | 7.741e-05 |
| 2 | NUCLEAR CHROMATIN | 7 | 291 | 2.504e-07 | 7.741e-05 |
| 3 | APICAL PART OF CELL | 7 | 361 | 1.066e-06 | 0.0002076 |
| 4 | CHROMOSOME | 9 | 880 | 4.988e-06 | 0.0005883 |
| 5 | TRANSCRIPTIONAL REPRESSOR COMPLEX | 4 | 74 | 5.037e-06 | 0.0005883 |
| 6 | NUCLEAR CHROMOSOME | 7 | 523 | 1.225e-05 | 0.001192 |
| 7 | ACETYLTRANSFERASE COMPLEX | 4 | 100 | 1.669e-05 | 0.001392 |
| 8 | NUCLEOPLASM PART | 7 | 708 | 8.462e-05 | 0.004942 |
| 9 | GOLGI MEMBRANE | 7 | 703 | 8.094e-05 | 0.004942 |
| 10 | APICAL PLASMA MEMBRANE | 5 | 292 | 7.978e-05 | 0.004942 |
| 11 | HISTONE DEACETYLASE COMPLEX | 3 | 61 | 0.0001142 | 0.006062 |
| 12 | WNT SIGNALOSOME | 2 | 11 | 0.0001268 | 0.00617 |
| 13 | SIN3 COMPLEX | 2 | 13 | 0.0001794 | 0.008061 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04330_Notch_signaling_pathway | 31 | 47 | 5.891e-88 | 1.06e-85 | |
| 2 | hsa04310_Wnt_signaling_pathway | 6 | 151 | 1.056e-07 | 9.506e-06 | |
| 3 | hsa04916_Melanogenesis | 5 | 101 | 4.55e-07 | 2.73e-05 | |
| 4 | hsa04320_Dorso.ventral_axis_formation | 3 | 25 | 7.578e-06 | 0.000341 | |
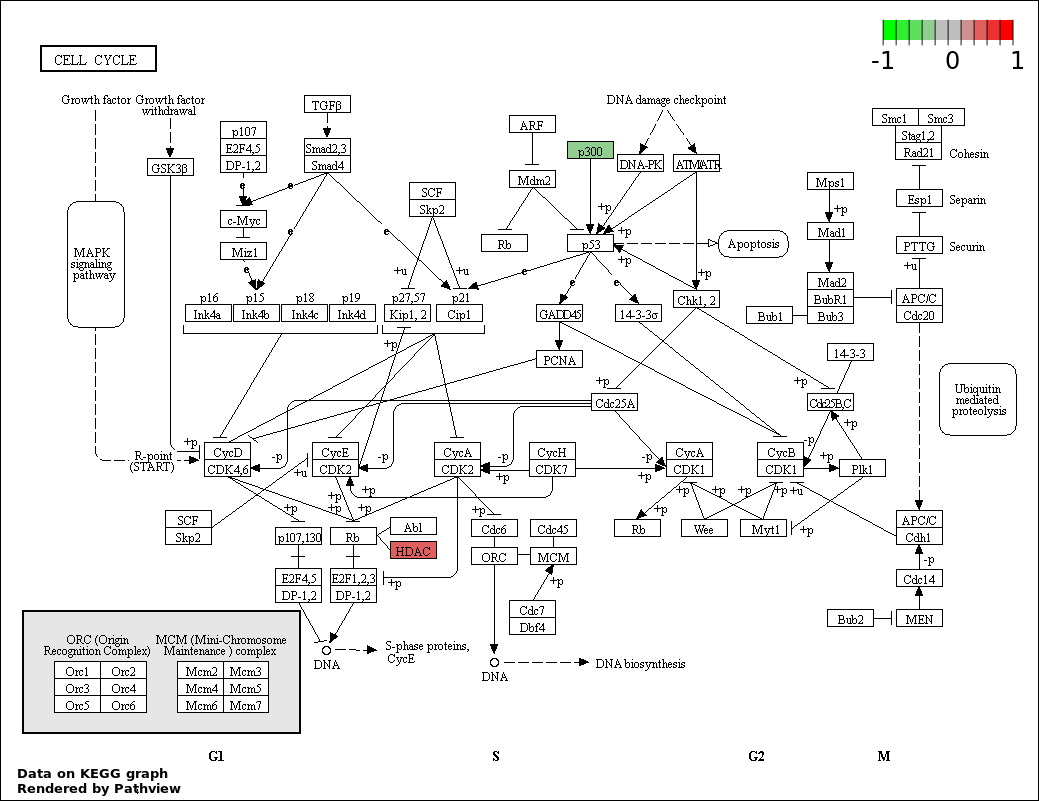
| 5 | hsa04110_Cell_cycle | 4 | 128 | 4.405e-05 | 0.001586 | |
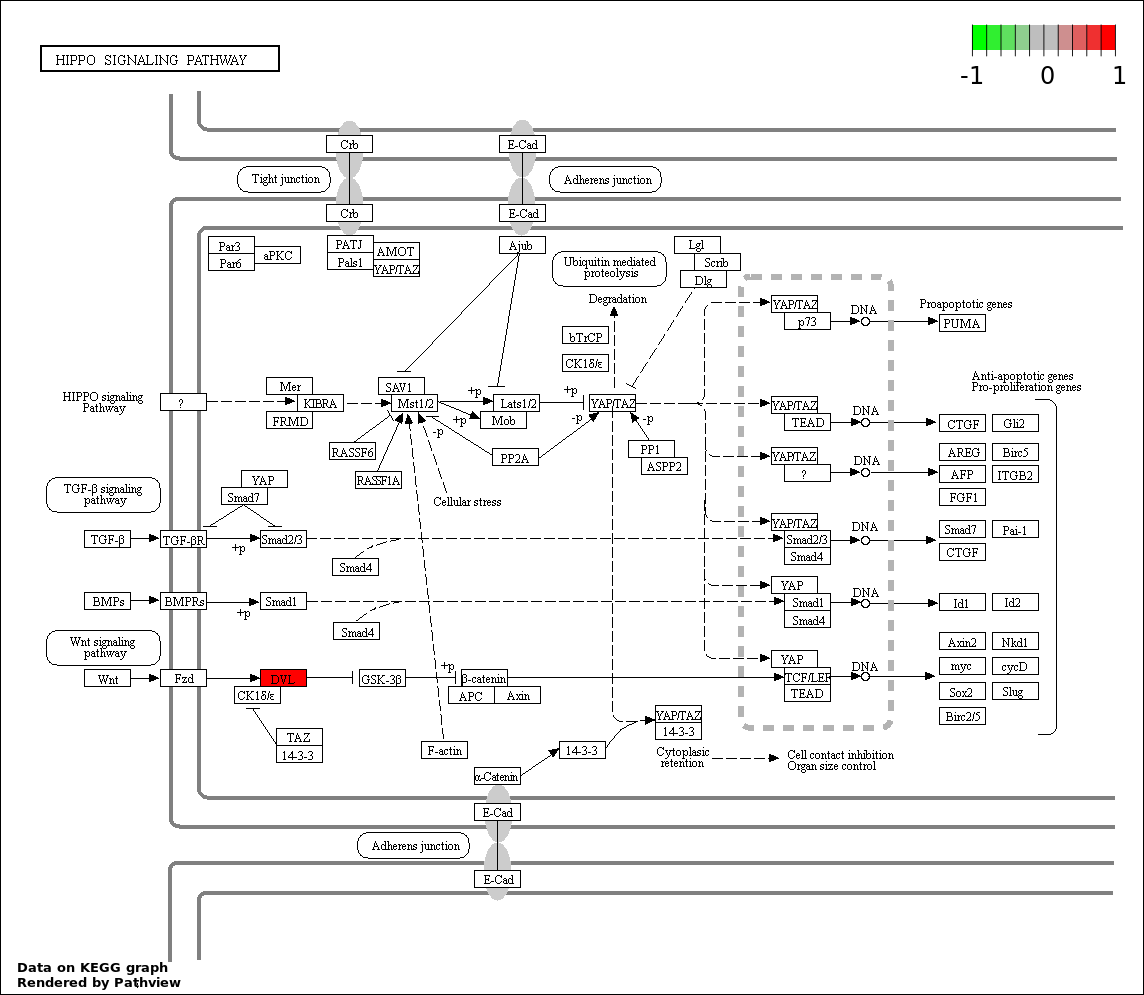
| 6 | hsa04390_Hippo_signaling_pathway | 3 | 154 | 0.001718 | 0.05154 | |
| 7 | hsa00514_Other_types_of_O.glycan_biosynthesis | 2 | 46 | 0.002306 | 0.05931 | |
| 8 | hsa04720_Long.term_potentiation | 2 | 70 | 0.005259 | 0.1141 | |
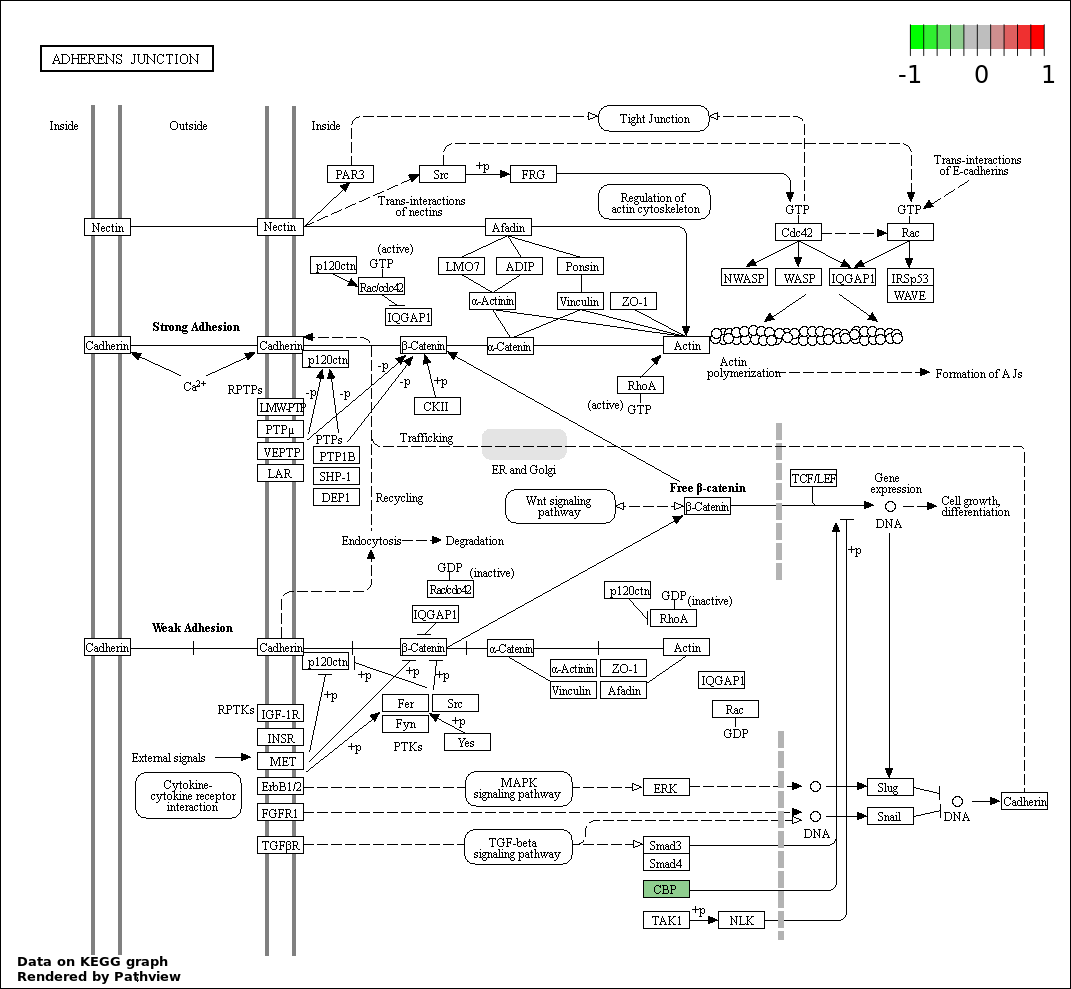
| 9 | hsa04520_Adherens_junction | 2 | 73 | 0.005706 | 0.1141 | |
| 10 | hsa04350_TGF.beta_signaling_pathway | 2 | 85 | 0.007662 | 0.1379 | |
| 11 | hsa04630_Jak.STAT_signaling_pathway | 2 | 155 | 0.02396 | 0.392 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | TPRG1-AS1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-125a-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-92a-3p;hsa-miR-92b-3p;hsa-miR-93-5p | 14 | KAT2B | Sponge network | -0.756 | 0.03021 | -0.939 | 0 | 0.573 |
| 2 | RP11-12A2.3 | hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-92a-3p;hsa-miR-92b-3p;hsa-miR-93-5p | 13 | KAT2B | Sponge network | -4.779 | 0 | -0.939 | 0 | 0.539 |
| 3 | AC004862.6 | hsa-miR-106b-5p;hsa-miR-125a-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-93-5p | 11 | KAT2B | Sponge network | -2.202 | 0.00081 | -0.939 | 0 | 0.489 |
| 4 | LINC00238 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-125a-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-342-3p | 13 | KAT2B | Sponge network | -4.997 | 0 | -0.939 | 0 | 0.478 |
| 5 | AC005550.3 | hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-181d-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-20a-5p;hsa-miR-92a-3p;hsa-miR-92b-3p | 11 | KAT2B | Sponge network | -2.571 | 0.00132 | -0.939 | 0 | 0.44 |
| 6 | RP11-166D19.1 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-590-5p;hsa-miR-92a-3p | 10 | KAT2B | Sponge network | -0.244 | 0.28835 | -0.939 | 0 | 0.254 |