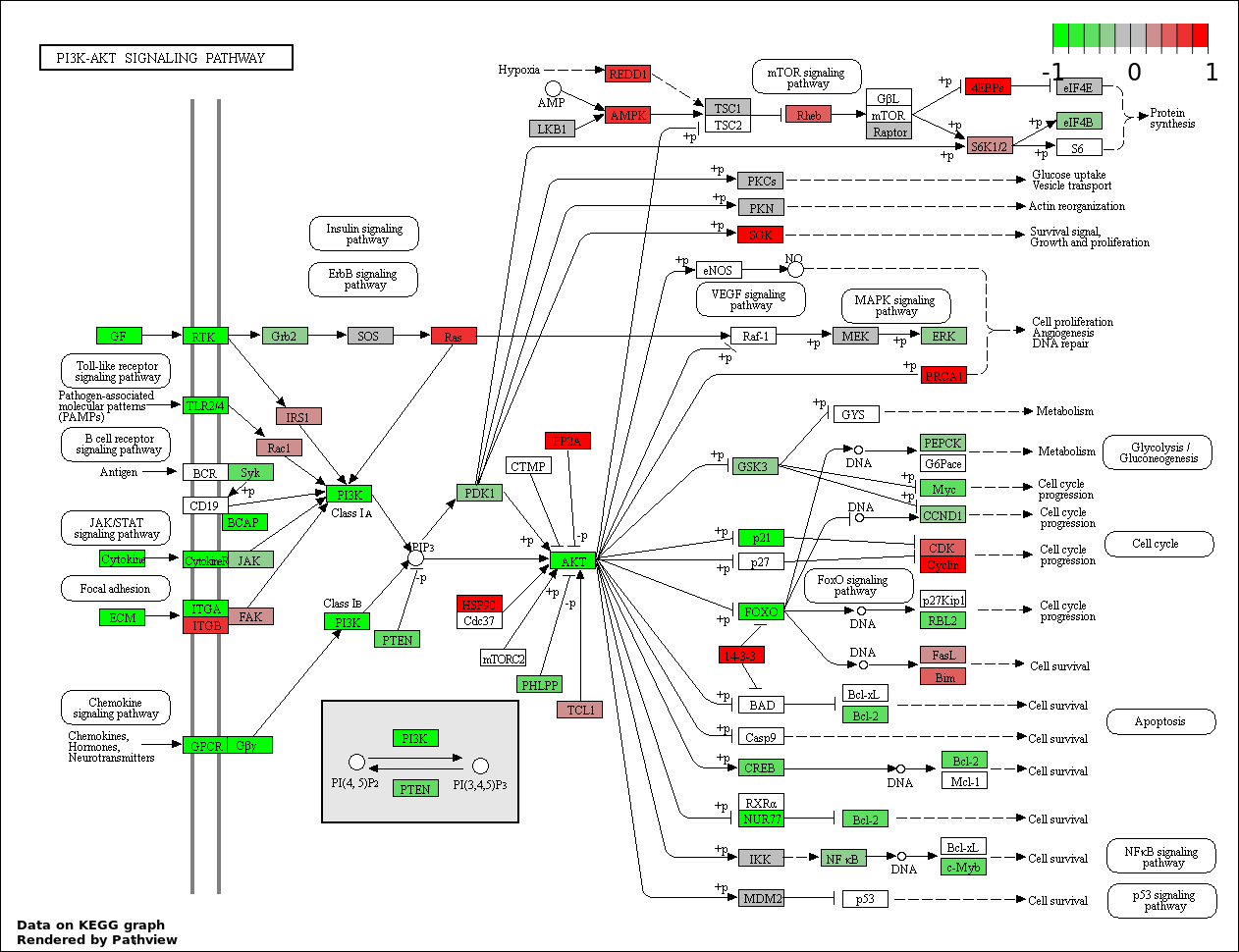
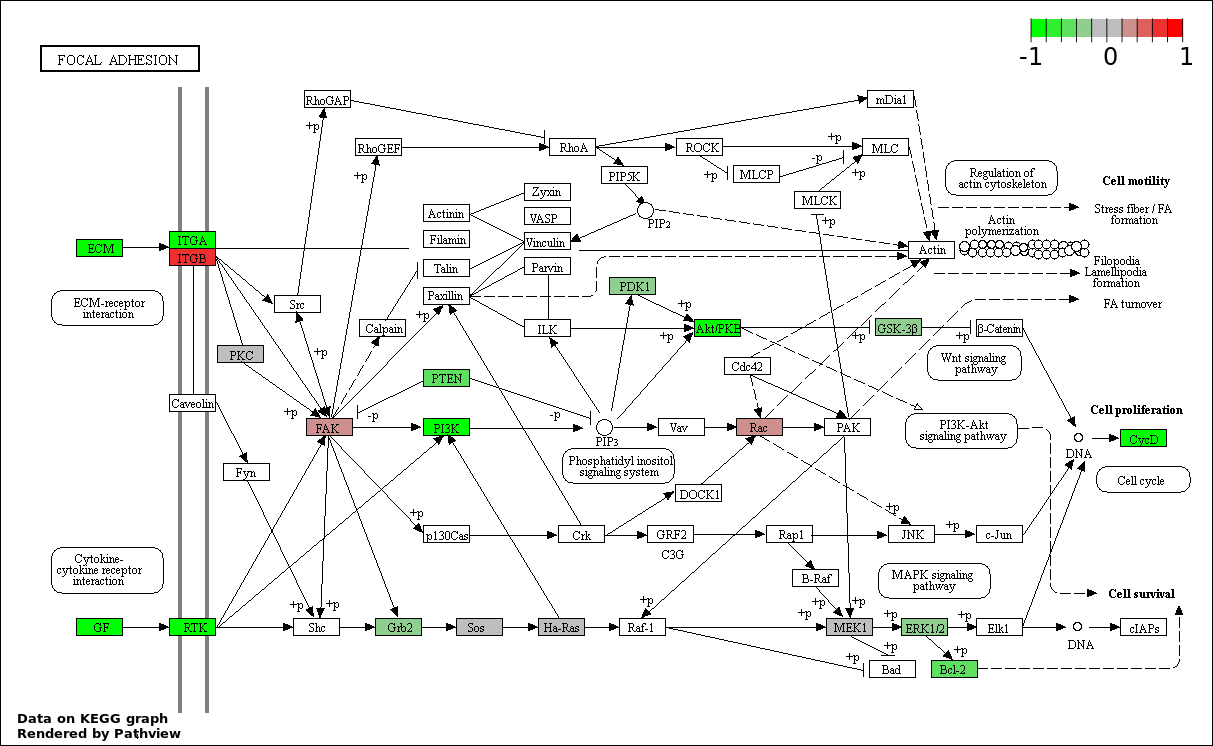
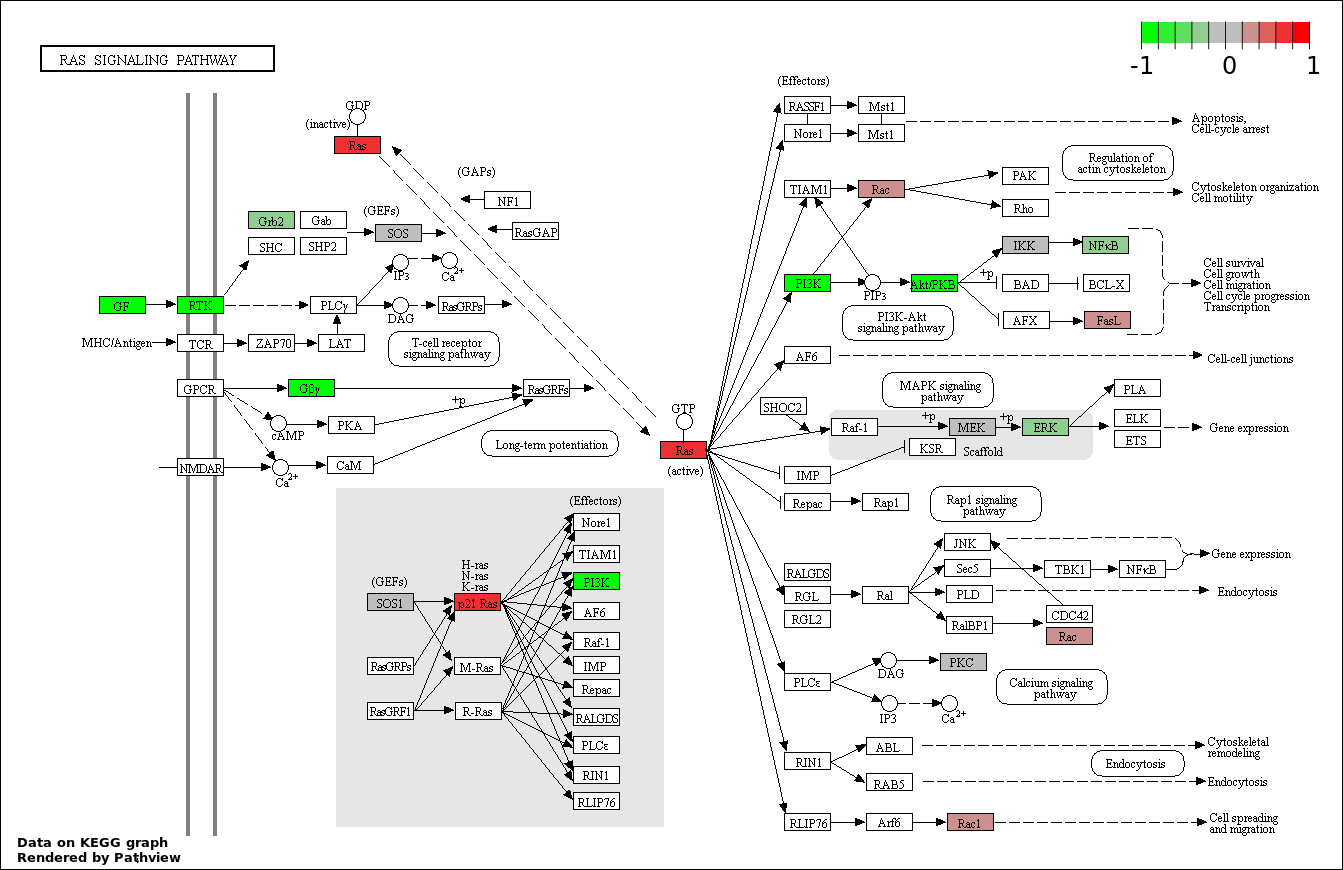
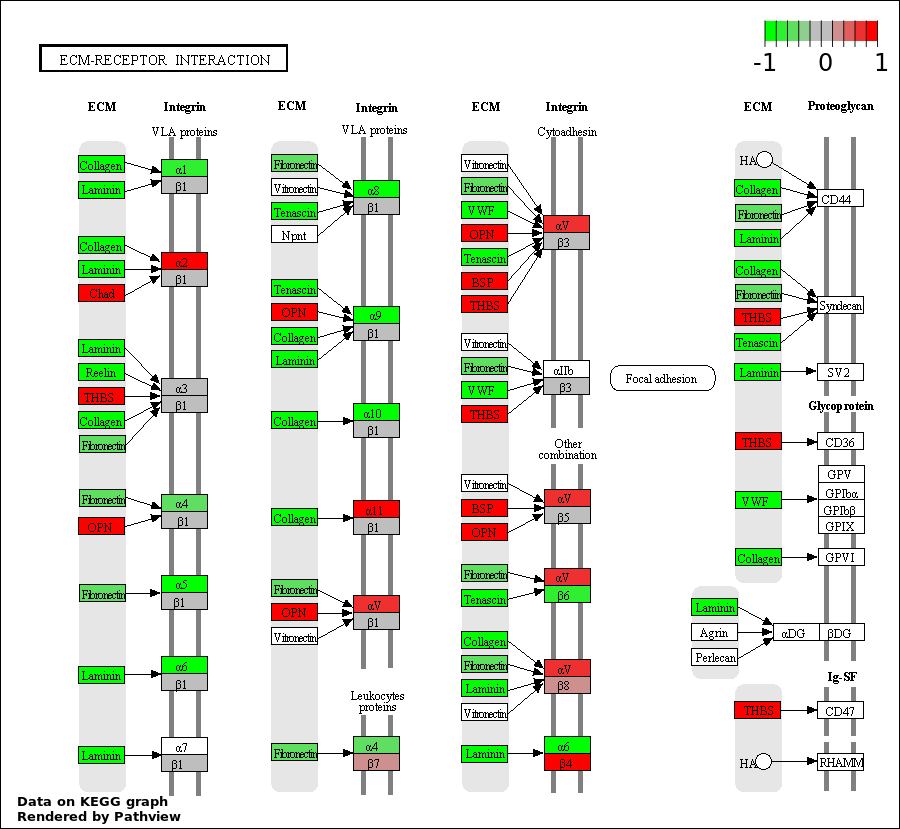
This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-let-7a-5p | AKT2 | -1.37 | 0 | 0.16 | 0.1135 | TargetScan | -0.12 | 0 | NA | |
| 2 | hsa-miR-29a-3p | AKT2 | 0.1 | 0.5732 | 0.16 | 0.1135 | MirTarget | -0.1 | 4.0E-5 | 24076586 | Furthermore a feed-back loop comprising of c-Myc miR-29 family and Akt2 were found in myeloid leukemogenesis |
| 3 | hsa-miR-106b-5p | AKT3 | 1.47 | 0 | -1.44 | 0 | miRNATAP | -0.16 | 0.00426 | NA | |
| 4 | hsa-miR-107 | AKT3 | 0.66 | 0 | -1.44 | 0 | PITA; miRanda | -0.26 | 0.0031 | NA | |
| 5 | hsa-miR-146b-5p | AKT3 | 1.09 | 1.0E-5 | -1.44 | 0 | miRNAWalker2 validate | -0.15 | 0.00189 | NA | |
| 6 | hsa-miR-15a-5p | AKT3 | 1.63 | 0 | -1.44 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.41 | 0 | NA | |
| 7 | hsa-miR-16-5p | AKT3 | 0.75 | 0 | -1.44 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.27 | 0.0001 | NA | |
| 8 | hsa-miR-17-5p | AKT3 | 2.07 | 0 | -1.44 | 0 | TargetScan; miRNATAP | -0.22 | 0 | NA | |
| 9 | hsa-miR-181a-5p | AKT3 | -0.38 | 0.05621 | -1.44 | 0 | miRNATAP | -0.23 | 9.0E-5 | NA | |
| 10 | hsa-miR-181b-5p | AKT3 | 0.67 | 0.00024 | -1.44 | 0 | miRNATAP | -0.37 | 0 | NA | |
| 11 | hsa-miR-20a-5p | AKT3 | 2.65 | 0 | -1.44 | 0 | miRNATAP | -0.24 | 0 | NA | |
| 12 | hsa-miR-22-3p | AKT3 | 1.43 | 0 | -1.44 | 0 | miRNATAP | -0.26 | 0.00109 | NA | |
| 13 | hsa-miR-29b-2-5p | AKT3 | 0.35 | 0.19484 | -1.44 | 0 | mirMAP | -0.18 | 2.0E-5 | NA | |
| 14 | hsa-miR-29b-3p | AKT3 | 3.11 | 0 | -1.44 | 0 | miRNATAP | -0.27 | 0 | 26512921 | MicroRNA 29B mir 29b regulates the Warburg effect in ovarian cancer by targeting AKT2 and AKT3 |
| 15 | hsa-miR-3065-5p | AKT3 | 0.65 | 0.09995 | -1.44 | 0 | mirMAP | -0.18 | 0 | NA | |
| 16 | hsa-miR-362-5p | AKT3 | 0.66 | 0.02433 | -1.44 | 0 | PITA; TargetScan; miRNATAP | -0.23 | 0 | NA | |
| 17 | hsa-miR-542-3p | AKT3 | 1.62 | 0 | -1.44 | 0 | miRanda | -0.19 | 1.0E-5 | NA | |
| 18 | hsa-miR-93-5p | AKT3 | 1.51 | 0 | -1.44 | 0 | miRNATAP | -0.25 | 0 | NA | |
| 19 | hsa-miR-335-3p | ANGPT1 | 1.51 | 0 | -2.86 | 0 | mirMAP | -0.33 | 1.0E-5 | NA | |
| 20 | hsa-miR-429 | ANGPT1 | 2.38 | 0 | -2.86 | 0 | miRanda | -0.25 | 0 | NA | |
| 21 | hsa-miR-590-3p | ANGPT1 | 0.84 | 0.00129 | -2.86 | 0 | PITA; miRanda; mirMAP | -0.4 | 0 | NA | |
| 22 | hsa-miR-125a-5p | ANGPT2 | -1.05 | 0 | -0.36 | 0.23492 | miRanda | -0.23 | 0.0005 | NA | |
| 23 | hsa-miR-135b-5p | ANGPT2 | 3.25 | 0 | -0.36 | 0.23492 | MirTarget | -0.21 | 0 | NA | |
| 24 | hsa-miR-142-5p | ANGPT2 | 1.3 | 0 | -0.36 | 0.23492 | MirTarget | -0.15 | 0.00403 | NA | |
| 25 | hsa-miR-186-5p | ANGPT2 | 0.85 | 0 | -0.36 | 0.23492 | MirTarget; mirMAP | -0.39 | 7.0E-5 | NA | |
| 26 | hsa-miR-34c-5p | ANGPT2 | -1 | 0.07244 | -0.36 | 0.23492 | miRanda | -0.16 | 0 | NA | |
| 27 | hsa-miR-374a-5p | ANGPT2 | -0.2 | 0.29808 | -0.36 | 0.23492 | mirMAP | -0.26 | 0.00171 | NA | |
| 28 | hsa-miR-374b-5p | ANGPT2 | 0.47 | 0.01092 | -0.36 | 0.23492 | mirMAP | -0.22 | 0.00511 | NA | |
| 29 | hsa-miR-429 | ANGPT2 | 2.38 | 0 | -0.36 | 0.23492 | miRanda | -0.22 | 0 | NA | |
| 30 | hsa-miR-664a-3p | ANGPT2 | 0.44 | 0.02142 | -0.36 | 0.23492 | mirMAP | -0.39 | 0 | NA | |
| 31 | hsa-let-7a-3p | ATF2 | 0.17 | 0.43183 | 0.01 | 0.93689 | MirTarget; mirMAP; miRNATAP | -0.13 | 0.0001 | NA | |
| 32 | hsa-miR-140-3p | ATF2 | -1.11 | 0 | 0.01 | 0.93689 | MirTarget; PITA; miRNATAP | -0.16 | 0.00016 | NA | |
| 33 | hsa-miR-15a-5p | ATF2 | 1.63 | 0 | 0.01 | 0.93689 | miRNAWalker2 validate | -0.19 | 0 | NA | |
| 34 | hsa-miR-181a-5p | ATF2 | -0.38 | 0.05621 | 0.01 | 0.93689 | mirMAP; miRNATAP | -0.15 | 0 | NA | |
| 35 | hsa-miR-222-3p | ATF2 | 0.03 | 0.88194 | 0.01 | 0.93689 | MirTarget | -0.13 | 0 | NA | |
| 36 | hsa-miR-26b-5p | ATF2 | 0.72 | 5.0E-5 | 0.01 | 0.93689 | MirTarget; miRNATAP | -0.12 | 0.00049 | 21901137 | Coordinated regulation of ATF2 by miR 26b in γ irradiated lung cancer cells; Concurrent analysis of time-series mRNA and microRNA profiles uncovered that expression of miR-26b was down regulated and its target activating transcription factor 2 ATF2 mRNA was up regulated in γ-irradiated H1299 cells; IR in miR-26b overexpressed H1299 cells could not induce expression of ATF2; From these results we concluded that IR-induced up-regulation of ATF2 was coordinately enhanced by suppression of miR-26b in lung cancer cells which may enhance the effect of IR in the MAPK signaling pathway |
| 37 | hsa-miR-30b-5p | ATF2 | 0.36 | 0.13803 | 0.01 | 0.93689 | mirMAP | -0.18 | 0 | NA | |
| 38 | hsa-miR-30c-5p | ATF2 | -0.33 | 0.1236 | 0.01 | 0.93689 | mirMAP | -0.26 | 0 | NA | |
| 39 | hsa-miR-374a-5p | ATF2 | -0.2 | 0.29808 | 0.01 | 0.93689 | mirMAP | -0.16 | 1.0E-5 | NA | |
| 40 | hsa-miR-15a-5p | BCL2 | 1.63 | 0 | -0.49 | 0.06421 | miRNAWalker2 validate; miRTarBase | -0.19 | 0.00401 | 26915294; 25594541; 18931683; 25623762; 22335947 | As a result transcript levels of the tumor-suppressive miR-15 and let-7 families increased which targeted and decreased the expression of the crucial prosurvival genes BCL-2 and BCL-XL respectively;MicroRNAs miRNAs encoded by the miR-15 cluster are known to induce G1 arrest and apoptosis by targeting G1 checkpoints and the anti-apoptotic B cell lymphoma 2 BCL-2 gene;MicroRNAs miRNAs are noncoding small RNAs that repress protein translation by targeting specific messenger RNAs miR-15a and miR-16-1 act as putative tumor suppressors by targeting the oncogene BCL2;miR 15a and miR 16 modulate drug resistance by targeting bcl 2 in human colon cancer cells; To investigate the reversal effect of targeted modulation of bcl-2 expression by miR-15a and miR-16 on drug resistance of human colon cancer cells;The expression of MiR-15a was significantly inhibited by Bcl-2 P < 0.05 |
| 41 | hsa-miR-21-5p | BCL2 | 4.38 | 0 | -0.49 | 0.06421 | miRNAWalker2 validate; miRTarBase | -0.2 | 1.0E-5 | 22964582; 21468550; 25994220; 25381586; 26555418; 23359184; 21376256 | Tumors harvested from these lungs have elevated levels of oncogenic miRNAs miR-21 and miR-155; are deficient for p53-regulated miRNAs; and have heightened expression of miR-34 target genes such as Met and Bcl-2;BCL-2 up-regulation could be achieved by miR-21 overexpression which prevented T24 cells from apoptosis induced by doxorubicin; Furthermore the miR-21 induced BCL-2 up-regulation could be cancelled by the PI3K inhibitor LY294002;Meanwhile miR-21 loss reduced STAT3 and Bcl-2 activation causing an increase in the apoptosis of tumour cells in CAC mice;Changes in the sensitivity of osteosarcoma cells to CDDP were examined after transfection with miR-21 mimics or anti-miR-21 or bcl-2 siRNA in combination with CDDP;The expression of Bax Bcl-2 and miR-21 in parental and paclitaxel-resistant cells was detected by RT-PCR and Western blotting;Resveratrol induces apoptosis of pancreatic cancers cells by inhibiting miR 21 regulation of BCL 2 expression; We also used Western blot to measure BCL-2 protein levels after down-regulation of miR-21 expression; Besides down-regulation of miR-21 expression can inhibit BCL-2 expression in PANC-1 CFPAC-1 and MIA Paca-2 cells; Over-expression of miR-21 expression can reverse down-regulation of BCL-2 expression and apoptosis induced by resveratrol; In this study we demonstrated that the effect of resveratrol on apoptosis is due to inhibiting miR-21 regulation of BCL-2 expression;Bcl 2 upregulation induced by miR 21 via a direct interaction is associated with apoptosis and chemoresistance in MIA PaCa 2 pancreatic cancer cells; However the roles and mechanisms of miRNA miR-21 in regulation of Bcl-2 in pancreatic cancer remain to be elucidated; Then luciferase activity was observed after miR-21 mimics and pRL-TK plasmids containing wild-type and mutant 3'UTRs of Bcl-2 mRNA were co-transfected; Cells transfected with miR-21 inhibitor revealed an opposite trend. There was a significant increase in luciferase activity in the cells transfected with the wild-type pRL-TK plasmid in contrast to those transfected with the mutant one indicating that miR-21 promotes Bcl-2 expression by binding directly to the 3'UTR of Bcl-2 mRNA; Upregulation of Bcl-2 directly induced by miR-21 is associated with apoptosis chemoresistance and proliferation of MIA PaCa-2 pancreatic cancer cells |
| 42 | hsa-miR-224-5p | BCL2 | 1.92 | 0 | -0.49 | 0.06421 | mirMAP | -0.14 | 0 | 24796455 | In addition the expressions of Bcl2 mRNA and protein were 1.05 ± 0.04 and 0.21 ± 0.03 in the miR-224 ASO group significantly lower than that in the control group 4.87 ± 0.96 and 0.88 ± 0.09 P < 0.01 |
| 43 | hsa-miR-24-2-5p | BCL2 | 1.44 | 0 | -0.49 | 0.06421 | miRNAWalker2 validate; miRTarBase | -0.16 | 0.00352 | NA | |
| 44 | hsa-miR-29a-5p | BCL2 | 1.9 | 0 | -0.49 | 0.06421 | mirMAP | -0.24 | 0 | 20041405 | Subsequent investigation characterized two antiapoptotic molecules Bcl-2 and Mcl-1 as direct targets of miR-29; Furthermore silencing of Bcl-2 and Mcl-1 phenocopied the proapoptotic effect of miR-29 whereas overexpression of these proteins attenuated the effect of miR-29 |
| 45 | hsa-miR-3065-5p | BCL2 | 0.65 | 0.09995 | -0.49 | 0.06421 | mirMAP | -0.12 | 0.00023 | NA | |
| 46 | hsa-miR-34a-5p | BCL2 | 1.41 | 0 | -0.49 | 0.06421 | miRNAWalker2 validate; miRTarBase | -0.19 | 0.00135 | 22964582; 24565525; 23155233; 24444609; 20687223; 22623155; 24988056; 18803879; 19714243; 25053345; 20433755; 21399894; 23862748 | Tumors harvested from these lungs have elevated levels of oncogenic miRNAs miR-21 and miR-155; are deficient for p53-regulated miRNAs; and have heightened expression of miR-34 target genes such as Met and Bcl-2;In vitro and in vivo experiments showed that miR-34a and DOX can be efficiently encapsulated into HA-CS NPs and delivered into tumor cells or tumor tissues and enhance anti-tumor effects of DOX by suppressing the expression of non-pump resistance and anti-apoptosis proto-oncogene Bcl-2;The miR-34a expression levels in cells after irradiation at 30 and 60 Gy were 0.17- and 18.7-times the BCL2 and caspase-9 expression levels respectively;Functional analyses further indicate that restoration of miR-34a inhibits B cell lymphoma-2 Bcl-2 protein expression to withdraw the survival advantage of these resistant NSCLC cells;Thus in PC3PR cells reduced expression of miR-34a confers paclitaxel resistance via up-regulating SIRT1 and Bcl2 expression; MiR-34a and its downstream targets SIRT1 and Bcl2 play important roles in the development of paclitaxel resistance all of which can be useful biomarkers and promising therapeutic targets for the drug resistance in hormone-refractory prostate cancer;MiR 34a inhibits proliferation and migration of breast cancer through down regulation of Bcl 2 and SIRT1; In this study we aimed to determine the effect of miR-34a on the growth of breast cancer and to investigate whether its effect is achieved by targeting Bcl-2 and SIRT1; Bcl-2 and SIRT1 as the targets of miR-34a were found to be in reverse correlation with ectopic expression of miR-34a;Target analysis indicated that micro RNA miR-34a directly regulates Bcl-2 and miR-34a overexpression decreased Bcl-2 protein level in gastric cancer cells; We also found that luteolin upregulates miR-34a expression and downregulates Bcl-2 expression; Based on these results we can draw the conclusion that luteolin partly decreases Bcl-2 expression through upregulating miR-34a expression;miR-34 targets Notch HMGA2 and Bcl-2 genes involved in the self-renewal and survival of cancer stem cells; Human gastric cancer cells were transfected with miR-34 mimics or infected with the lentiviral miR-34-MIF expression system and validated by miR-34 reporter assay using Bcl-2 3'UTR reporter; Human gastric cancer Kato III cells with miR-34 restoration reduced the expression of target genes Bcl-2 Notch and HMGA2; Bcl-2 3'UTR reporter assay showed that the transfected miR-34s were functional and confirmed that Bcl-2 is a direct target of miR-34; The mechanism of miR-34-mediated suppression of self-renewal appears to be related to the direct modulation of downstream targets Bcl-2 Notch and HMGA2 indicating that miR-34 may be involved in gastric cancer stem cell self-renewal/differentiation decision-making;Among the target proteins regulated by miR-34 are Notch pathway proteins and Bcl-2 suggesting the possibility of a role for miR-34 in the maintenance and survival of cancer stem cells; Our data support the view that miR-34 may be involved in pancreatic cancer stem cell self-renewal potentially via the direct modulation of downstream targets Bcl-2 and Notch implying that miR-34 may play an important role in pancreatic cancer stem cell self-renewal and/or cell fate determination;Manipulating miR-34a in prostate cancer cells confirms that this miRNA regulates BCL-2 and may in part regulate response to docetaxel;For instance miR-34a up-regulation corresponded with a down-regulation of BCL2 protein; Treating Par-4-overexpressing HT29 cells with a miR-34a antagomir functionally reversed both BCL2 down-regulation and apoptosis by 5-FU;Quantitative PCR and western analysis confirmed decreased expression of two genes BCL-2 and CCND1 in docetaxel-resistant cells which are both targeted by miR-34a;MicroRNA 34a targets Bcl 2 and sensitizes human hepatocellular carcinoma cells to sorafenib treatment; HCC tissues with lower miR-34a expression displayed higher expression of Bcl-2 protein than those with high expression of miR-34a; therefore an inverse correlation is evident between the miR-34a level and Bcl-2 expression; Bioinformatics and luciferase reporter assays revealed that miR-34a binds the 3'-UTR of the Bcl-2 mRNA and represses its translation; Western blotting analysis and qRT-PCR confirmed that Bcl-2 is inhibited by miR-34a overexpression; Functional analyses indicated that the restoration of miR-34a reduced cell viability promoted cell apoptosis and potentiated sorafenib-induced apoptosis and toxicity in HCC cell lines by inhibiting Bcl-2 expression |
| 47 | hsa-miR-450b-5p | BCL2 | 1.69 | 0 | -0.49 | 0.06421 | mirMAP | -0.2 | 0 | NA | |
| 48 | hsa-miR-452-5p | BCL2 | 0.64 | 0.04582 | -0.49 | 0.06421 | mirMAP | -0.12 | 0.00113 | NA | |
| 49 | hsa-miR-542-3p | BCL2 | 1.62 | 0 | -0.49 | 0.06421 | mirMAP | -0.19 | 3.0E-5 | NA | |
| 50 | hsa-miR-590-5p | BCL2 | 2.07 | 0 | -0.49 | 0.06421 | miRanda | -0.18 | 0.00054 | NA | |
| 51 | hsa-miR-96-5p | BCL2 | 3.04 | 0 | -0.49 | 0.06421 | miRNAWalker2 validate; TargetScan | -0.14 | 0.00193 | NA | |
| 52 | hsa-miR-181a-5p | BCL2L11 | -0.38 | 0.05621 | 0.41 | 0.00143 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.12 | 5.0E-5 | 23241956; 20841506 | Mechanistically inactivation of miR-181a elevated the expression of the proapoptotic molecule Bim which sensitized metastatic cells to anoikis;Furthermore we found that cell adhesion-up-regulated miR-181a contributes to FDC-mediated cell survival through Bim down-regulation implicating miR-181a as an upstream effector of the Bim-apoptosis signaling pathway; miR-181a inhibition and Bim upregulation significantly suppressed FDC-mediated protection against apoptosis in lymphoma cell lines and primary lymphoma cells |
| 53 | hsa-miR-195-5p | BCL2L11 | -1.02 | 5.0E-5 | 0.41 | 0.00143 | miRNAWalker2 validate | -0.1 | 5.0E-5 | NA | |
| 54 | hsa-miR-221-3p | BCL2L11 | -0.1 | 0.65445 | 0.41 | 0.00143 | miRNAWalker2 validate | -0.11 | 5.0E-5 | 26503209 | Knockdown of miR 221 promotes the cisplatin inducing apoptosis by targeting the BIM Bax/Bak axis in breast cancer |
| 55 | hsa-miR-30b-5p | BCL2L11 | 0.36 | 0.13803 | 0.41 | 0.00143 | miRNATAP | -0.13 | 0 | NA | |
| 56 | hsa-miR-30c-5p | BCL2L11 | -0.33 | 0.1236 | 0.41 | 0.00143 | miRNATAP | -0.13 | 0 | NA | |
| 57 | hsa-miR-30d-5p | BCL2L11 | -0.92 | 4.0E-5 | 0.41 | 0.00143 | miRNATAP | -0.13 | 0 | NA | |
| 58 | hsa-miR-10a-5p | BDNF | 0.79 | 0.00059 | -3.58 | 0 | MirTarget; miRNATAP | -0.28 | 0.00181 | NA | |
| 59 | hsa-miR-146b-5p | BDNF | 1.09 | 1.0E-5 | -3.58 | 0 | miRanda | -0.33 | 0.00012 | NA | |
| 60 | hsa-miR-148a-5p | BDNF | 1.46 | 0 | -3.58 | 0 | mirMAP | -0.24 | 0.00212 | NA | |
| 61 | hsa-miR-155-5p | BDNF | 0.81 | 0.00061 | -3.58 | 0 | miRNATAP | -0.23 | 0.00731 | NA | |
| 62 | hsa-miR-15a-5p | BDNF | 1.63 | 0 | -3.58 | 0 | MirTarget; miRNATAP | -0.3 | 0.00719 | 26581909 | MicroRNA 15a 5p suppresses cancer proliferation and division in human hepatocellular carcinoma by targeting BDNF; BDNF was then overexpressed in HepG2 and SNU-182 cells to evaluate its selective effect on miR-15a-5p in HCC modulation; MiR-15a-5p selectively and negatively regulated BDNF at both gene and protein levels in HCC cells; Forced overexpression of BDNF effectively reversed the tumor suppressive functions of miR-15a-5p on HCC proliferation and cell division in vitro; Our study demonstrated that miR-15a-5p is a tumor suppressor in HCC and its regulation is through BDNF in HCC |
| 63 | hsa-miR-182-5p | BDNF | 3.22 | 0 | -3.58 | 0 | MirTarget; miRNATAP | -0.49 | 0 | NA | |
| 64 | hsa-miR-210-3p | BDNF | 4.89 | 0 | -3.58 | 0 | miRNAWalker2 validate; miRTarBase | -0.22 | 0 | NA | |
| 65 | hsa-miR-29b-1-5p | BDNF | 1.71 | 0 | -3.58 | 0 | mirMAP | -0.25 | 0.00028 | NA | |
| 66 | hsa-miR-30e-5p | BDNF | 1.6 | 0 | -3.58 | 0 | miRNATAP | -0.44 | 9.0E-5 | NA | |
| 67 | hsa-miR-7-1-3p | BDNF | 2.61 | 0 | -3.58 | 0 | MirTarget | -0.3 | 0.00109 | NA | |
| 68 | hsa-miR-181a-5p | BRCA1 | -0.38 | 0.05621 | 1.29 | 0 | miRNAWalker2 validate | -0.39 | 0 | NA | |
| 69 | hsa-miR-199a-5p | BRCA1 | 1.31 | 0 | 1.29 | 0 | miRanda | -0.17 | 0.00042 | NA | |
| 70 | hsa-miR-199b-5p | BRCA1 | 2.14 | 0 | 1.29 | 0 | miRanda | -0.12 | 0.0033 | NA | |
| 71 | hsa-miR-320a | BRCA1 | -0.96 | 0 | 1.29 | 0 | miRanda | -0.26 | 8.0E-5 | NA | |
| 72 | hsa-miR-106a-5p | CCND1 | 1.39 | 6.0E-5 | -0.3 | 0.2554 | MirTarget; miRNATAP | -0.25 | 0 | NA | |
| 73 | hsa-miR-106b-5p | CCND1 | 1.47 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.26 | 1.0E-5 | NA | |
| 74 | hsa-miR-142-3p | CCND1 | 3.98 | 0 | -0.3 | 0.2554 | miRanda | -0.12 | 0.00053 | 23619912 | Transfection of miR-142-3p mimics in colon cancer cells downregulated cyclin D1 expression induced G1 phase cell cycle arrest and elevated the sensitivity of the cells to 5-fluorouracil |
| 75 | hsa-miR-15a-5p | CCND1 | 1.63 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.2 | 0.00193 | 22922827 | CCND1 has been found to be a target of miR-15a and miR-16-1 through analysis of complementary sequences between microRNAs and CCND1 mRNA; Moreover the transcription of CCND1 is suppressed by miR-15a and miR-16-1 via direct binding to the CCND1 3'-untranslated region 3'-UTR |
| 76 | hsa-miR-15b-5p | CCND1 | -1.26 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.3 | 0 | NA | |
| 77 | hsa-miR-16-1-3p | CCND1 | 1.5 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase | -0.25 | 3.0E-5 | 22922827; 18483394 | CCND1 has been found to be a target of miR-15a and miR-16-1 through analysis of complementary sequences between microRNAs and CCND1 mRNA; Moreover the transcription of CCND1 is suppressed by miR-15a and miR-16-1 via direct binding to the CCND1 3'-untranslated region 3'-UTR;Truncation in CCND1 mRNA alters miR 16 1 regulation in mantle cell lymphoma; Furthermore we demonstrated that this truncation alters miR-16-1 binding sites and through the use of reporter constructs we were able to show that miR-16-1 regulates CCND1 mRNA expression; This study introduces the role of miR-16-1 in the regulation of CCND1 in MCL |
| 78 | hsa-miR-16-5p | CCND1 | 0.75 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.37 | 0 | 23991964; 22922827; 18483394 | At the molecular level our results further revealed that cyclin D1 expression was negatively regulated by miR-16;CCND1 has been found to be a target of miR-15a and miR-16-1 through analysis of complementary sequences between microRNAs and CCND1 mRNA; Moreover the transcription of CCND1 is suppressed by miR-15a and miR-16-1 via direct binding to the CCND1 3'-untranslated region 3'-UTR;Truncation in CCND1 mRNA alters miR 16 1 regulation in mantle cell lymphoma; Furthermore we demonstrated that this truncation alters miR-16-1 binding sites and through the use of reporter constructs we were able to show that miR-16-1 regulates CCND1 mRNA expression; This study introduces the role of miR-16-1 in the regulation of CCND1 in MCL |
| 79 | hsa-miR-17-5p | CCND1 | 2.07 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; MirTarget; TargetScan; miRNATAP | -0.31 | 0 | 26431674 | Bioinformatics Prediction and In Vitro Analysis Revealed That miR 17 Targets Cyclin D1 mRNA in Triple Negative Breast Cancer Cells; In this study using bioinformatic analyses miR-17 was selected as it targets the 3'UTR of CCND1 gene with the highest score; After lentiviral transduction of miR-17 to the target cells gene expression analysis showed decreased expression of CCND1 gene |
| 80 | hsa-miR-186-5p | CCND1 | 0.85 | 0 | -0.3 | 0.2554 | mirMAP | -0.38 | 1.0E-5 | NA | |
| 81 | hsa-miR-193a-3p | CCND1 | 0.55 | 0.0319 | -0.3 | 0.2554 | MirTarget; PITA; miRanda | -0.19 | 0.00016 | NA | |
| 82 | hsa-miR-193b-3p | CCND1 | 1.1 | 0.00082 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.15 | 4.0E-5 | 27071318; 20655737; 20304954; 21893020; 26129688 | MicroRNA 193b inhibits the proliferation migration and invasion of gastric cancer cells via targeting cyclin D1; Further mechanism study indicated that CCND1 was a direct target of miR-193b in GC;CCND1 and ETS1 were revealed to be regulated by miR-193b directly;MicroRNA 193b represses cell proliferation and regulates cyclin D1 in melanoma; Overexpression of miR-193b in Malme-3M cells down-regulated CCND1 mRNA and protein by > or = 50%; A luciferase reporter assay confirmed that miR-193b directly regulates CCND1 by binding to the 3'untranslated region of CCND1 mRNA; These studies indicate that miR-193b represses cell proliferation and regulates CCND1 expression and suggest that dysregulation of miR-193b may play an important role in melanoma development;In a previous study we reported that miR-193b represses cell proliferation and regulates cyclin D1 in melanoma cells suggesting that miR-193b could act as a tumor suppressor;Epigenetically altered miR 193b targets cyclin D1 in prostate cancer; It has been suggested that miR-193b targets cyclin D1 in several malignancies; Here our aim was to determine if miR-193b targets cyclin D1 in prostate cancer; Furthermore the PC cell lines 22Rv1 and VCaP which express low levels of miR-193b and high levels of CCND1 showed significant growth retardation when treated with a CDK4/6 inhibitor; In contrast the inhibitor had no effect on the growth of PC-3 and DU145 cells with high miR-193b and low CCND1 expression; Taken together our data demonstrate that miR-193b targets cyclin D1 in prostate cancer |
| 83 | hsa-miR-19a-3p | CCND1 | 2.12 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.22 | 0 | 25985117 | Moreover miR-19a might play inhibitory roles in HCC malignancy via regulating Cyclin D1 expression |
| 84 | hsa-miR-19b-1-5p | CCND1 | 1.71 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase | -0.3 | 0 | NA | |
| 85 | hsa-miR-19b-3p | CCND1 | 2.11 | 0 | -0.3 | 0.2554 | miRNATAP | -0.18 | 0.0002 | NA | |
| 86 | hsa-miR-20a-5p | CCND1 | 2.65 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.24 | 0 | NA | |
| 87 | hsa-miR-20b-5p | CCND1 | 1.36 | 0.00261 | -0.3 | 0.2554 | MirTarget; miRNATAP | -0.18 | 0 | NA | |
| 88 | hsa-miR-365a-3p | CCND1 | 0.01 | 0.9536 | -0.3 | 0.2554 | miRNAWalker2 validate; miRTarBase | -0.19 | 0.00023 | NA | |
| 89 | hsa-miR-374a-5p | CCND1 | -0.2 | 0.29808 | -0.3 | 0.2554 | MirTarget | -0.31 | 1.0E-5 | 27191497 | microRNA 374a suppresses colon cancer progression by directly reducing CCND1 to inactivate the PI3K/AKT pathway; Furthermore luciferase reporter assays confirmed that miR-374a could directly reduce CCND1; We examined miR-374a levels by in situ hybridization and its correlation with CCND1 expression in CRC tumor tissues; High miR-374a expression with low level of CCND1 was protective factor in CRC; Together these findings indicate that miR-374a inactivates the PI3K/AKT axis by inhibiting CCND1 suppressing of colon cancer progression |
| 90 | hsa-miR-374b-5p | CCND1 | 0.47 | 0.01092 | -0.3 | 0.2554 | miRNAWalker2 validate; MirTarget | -0.26 | 0.00012 | NA | |
| 91 | hsa-miR-425-5p | CCND1 | 1.22 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate | -0.25 | 1.0E-5 | NA | |
| 92 | hsa-miR-589-3p | CCND1 | 1.34 | 2.0E-5 | -0.3 | 0.2554 | MirTarget | -0.13 | 0.00079 | NA | |
| 93 | hsa-miR-590-3p | CCND1 | 0.84 | 0.00129 | -0.3 | 0.2554 | mirMAP | -0.18 | 0.00086 | NA | |
| 94 | hsa-miR-769-3p | CCND1 | 0.45 | 0.07482 | -0.3 | 0.2554 | mirMAP | -0.14 | 0.00277 | NA | |
| 95 | hsa-miR-92a-3p | CCND1 | -0.14 | 0.49341 | -0.3 | 0.2554 | miRNAWalker2 validate | -0.34 | 0 | NA | |
| 96 | hsa-miR-93-5p | CCND1 | 1.51 | 0 | -0.3 | 0.2554 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.32 | 0 | NA | |
| 97 | hsa-miR-942-5p | CCND1 | -0.04 | 0.87063 | -0.3 | 0.2554 | MirTarget | -0.18 | 0.00098 | NA | |
| 98 | hsa-miR-106a-5p | CCND2 | 1.39 | 6.0E-5 | -1.64 | 0 | miRNATAP | -0.15 | 2.0E-5 | NA | |
| 99 | hsa-miR-106b-5p | CCND2 | 1.47 | 0 | -1.64 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.38 | 0 | NA | |
| 100 | hsa-miR-130b-5p | CCND2 | 1.54 | 0 | -1.64 | 0 | mirMAP | -0.27 | 0 | NA | |
| 101 | hsa-miR-141-3p | CCND2 | 3.37 | 0 | -1.64 | 0 | MirTarget; TargetScan | -0.24 | 0 | NA | |
| 102 | hsa-miR-151a-3p | CCND2 | 0.67 | 0.00028 | -1.64 | 0 | mirMAP | -0.28 | 0 | NA | |
| 103 | hsa-miR-15a-5p | CCND2 | 1.63 | 0 | -1.64 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.16 | 0.00712 | NA | |
| 104 | hsa-miR-17-5p | CCND2 | 2.07 | 0 | -1.64 | 0 | miRNAWalker2 validate; miRTarBase; TargetScan; miRNATAP | -0.37 | 0 | NA | |
| 105 | hsa-miR-182-5p | CCND2 | 3.22 | 0 | -1.64 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.37 | 0 | NA | |
| 106 | hsa-miR-183-5p | CCND2 | 2.39 | 0 | -1.64 | 0 | miRNATAP | -0.41 | 0 | NA | |
| 107 | hsa-miR-185-5p | CCND2 | 1.14 | 0 | -1.64 | 0 | MirTarget; miRNATAP | -0.24 | 0.00048 | NA | |
| 108 | hsa-miR-186-5p | CCND2 | 0.85 | 0 | -1.64 | 0 | mirMAP; miRNATAP | -0.36 | 1.0E-5 | NA | |
| 109 | hsa-miR-191-5p | CCND2 | 0.34 | 0.06681 | -1.64 | 0 | MirTarget | -0.2 | 0.00075 | NA | |
| 110 | hsa-miR-19a-3p | CCND2 | 2.12 | 0 | -1.64 | 0 | MirTarget; miRNATAP | -0.17 | 1.0E-5 | NA | |
| 111 | hsa-miR-19b-3p | CCND2 | 2.11 | 0 | -1.64 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.19 | 1.0E-5 | NA | |
| 112 | hsa-miR-200a-3p | CCND2 | 3.15 | 0 | -1.64 | 0 | MirTarget | -0.17 | 0 | NA | |
| 113 | hsa-miR-20a-5p | CCND2 | 2.65 | 0 | -1.64 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.28 | 0 | NA | |
| 114 | hsa-miR-21-3p | CCND2 | 2.54 | 0 | -1.64 | 0 | mirMAP | -0.21 | 0 | NA | |
| 115 | hsa-miR-2355-3p | CCND2 | 1.11 | 1.0E-5 | -1.64 | 0 | miRNATAP | -0.22 | 0 | NA | |
| 116 | hsa-miR-28-5p | CCND2 | 1.2 | 0 | -1.64 | 0 | miRanda | -0.29 | 0.00018 | NA | |
| 117 | hsa-miR-301a-3p | CCND2 | 2.7 | 0 | -1.64 | 0 | miRNAWalker2 validate | -0.27 | 0 | NA | |
| 118 | hsa-miR-320b | CCND2 | 0.23 | 0.37882 | -1.64 | 0 | mirMAP; miRNATAP | -0.12 | 0.00373 | NA | |
| 119 | hsa-miR-324-3p | CCND2 | -0.08 | 0.68923 | -1.64 | 0 | miRNAWalker2 validate | -0.28 | 0 | NA | |
| 120 | hsa-miR-331-5p | CCND2 | 0.58 | 0.00131 | -1.64 | 0 | miRNATAP | -0.28 | 1.0E-5 | NA | |
| 121 | hsa-miR-429 | CCND2 | 2.38 | 0 | -1.64 | 0 | miRNATAP | -0.21 | 0 | NA | |
| 122 | hsa-miR-450b-5p | CCND2 | 1.69 | 0 | -1.64 | 0 | MirTarget; PITA; miRNATAP | -0.15 | 9.0E-5 | NA | |
| 123 | hsa-miR-501-5p | CCND2 | 0.41 | 0.10435 | -1.64 | 0 | PITA; mirMAP; miRNATAP | -0.14 | 0.00086 | NA | |
| 124 | hsa-miR-503-5p | CCND2 | 1.97 | 0 | -1.64 | 0 | MirTarget | -0.13 | 5.0E-5 | 25860935 | We then identified two targets of miR-503 CCND2 and CCND3 |
| 125 | hsa-miR-550a-5p | CCND2 | 0.6 | 0.03148 | -1.64 | 0 | MirTarget | -0.22 | 0 | NA | |
| 126 | hsa-miR-589-3p | CCND2 | 1.34 | 2.0E-5 | -1.64 | 0 | mirMAP | -0.19 | 0 | NA | |
| 127 | hsa-miR-590-3p | CCND2 | 0.84 | 0.00129 | -1.64 | 0 | miRanda; mirMAP | -0.13 | 0.00821 | NA | |
| 128 | hsa-miR-590-5p | CCND2 | 2.07 | 0 | -1.64 | 0 | mirMAP | -0.26 | 0 | NA | |
| 129 | hsa-miR-660-5p | CCND2 | 2.05 | 0 | -1.64 | 0 | mirMAP | -0.15 | 0.00131 | NA | |
| 130 | hsa-miR-7-1-3p | CCND2 | 2.61 | 0 | -1.64 | 0 | mirMAP | -0.26 | 0 | NA | |
| 131 | hsa-miR-877-5p | CCND2 | -0.37 | 0.20671 | -1.64 | 0 | miRNAWalker2 validate | -0.15 | 6.0E-5 | NA | |
| 132 | hsa-miR-93-5p | CCND2 | 1.51 | 0 | -1.64 | 0 | miRNATAP | -0.41 | 0 | NA | |
| 133 | hsa-miR-96-5p | CCND2 | 3.04 | 0 | -1.64 | 0 | TargetScan; miRNATAP | -0.36 | 0 | NA | |
| 134 | hsa-miR-320b | CCND3 | 0.23 | 0.37882 | -0.86 | 1.0E-5 | miRanda | -0.11 | 0.00235 | NA | |
| 135 | hsa-miR-421 | CCND3 | 0.17 | 0.53528 | -0.86 | 1.0E-5 | PITA; miRanda | -0.12 | 0.00073 | NA | |
| 136 | hsa-miR-96-5p | CCND3 | 3.04 | 0 | -0.86 | 1.0E-5 | TargetScan | -0.12 | 0.00015 | NA | |
| 137 | hsa-miR-125b-5p | CCNE1 | -0.55 | 0.01072 | 3 | 0 | miRNAWalker2 validate | -0.21 | 0.00408 | NA | |
| 138 | hsa-miR-195-5p | CCNE1 | -1.02 | 5.0E-5 | 3 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.52 | 0 | 24402230 | Furthermore through qPCR and western blot assays we showed that overexpression of miR-195-5p reduced CCNE1 mRNA and protein levels respectively |
| 139 | hsa-miR-26a-5p | CCNE1 | -0.13 | 0.44003 | 3 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.7 | 0 | 22094936 | Cell cycle regulation and CCNE1 and CDC2 were the only significant overlapping pathway and genes differentially expressed between tumors with high and low levels of miR-26a and EZH2 respectively; Low mRNA levels of EZH2 CCNE1 and CDC2 and high levels of miR-26a are associated with favorable outcome on tamoxifen |
| 140 | hsa-miR-497-5p | CCNE1 | -0.05 | 0.78621 | 3 | 0 | MirTarget; miRNATAP | -0.5 | 0 | 25909221; 24112607; 24909281 | The effect of simultaneous overexpression of miR-497 and miR-34a on the inhibition of cell proliferation colony formation and tumor growth and the downregulation of cyclin E1 was stronger than the effect of each miRNA alone; The synergistic actions of miR-497 and miR-34a partly correlated with cyclin E1 levels; These results indicate cyclin E1 is downregulated by both miR-497 and miR-34a which synergistically retard the growth of human lung cancer cells;Western blot assays confirmed that overexpression of miR-497 reduced cyclin E1 protein levels; Inhibited cellular growth suppressed cellular migration and invasion and G1 cell cycle arrest were observed upon overexpression of miR-497 in cells possibly by targeting cyclin E1;miR 497 suppresses proliferation of human cervical carcinoma HeLa cells by targeting cyclin E1; Furthermore the target effect of miR-497 on the CCNE1 was identified by dual-luciferase reporter assay system qRT-PCR and Western blotting; Over-expressed miR-497 in HeLa cells could suppress cell proliferation by targeting CCNE1 |
| 141 | hsa-let-7a-3p | CCNE2 | 0.17 | 0.43183 | 2.15 | 0 | mirMAP | -0.22 | 0.00307 | NA | |
| 142 | hsa-let-7b-3p | CCNE2 | -1.82 | 0 | 2.15 | 0 | mirMAP | -0.49 | 0 | NA | |
| 143 | hsa-miR-126-3p | CCNE2 | 0.4 | 0.11564 | 2.15 | 0 | miRNAWalker2 validate | -0.15 | 0.00656 | NA | |
| 144 | hsa-miR-26a-5p | CCNE2 | -0.13 | 0.44003 | 2.15 | 0 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.51 | 0 | 24116110; 21901171 | The loss of miR 26a mediated post transcriptional regulation of cyclin E2 in pancreatic cancer cell proliferation and decreased patient survival; The in vitro and in vivo assays showed that overexpression of miR-26a resulted in cell cycle arrest inhibited cell proliferation and decreased tumor growth which was associated with cyclin E2 downregulation;We also show that enforced expression of miR-26a in AML cells is able to inhibit cell cycle progression by downregulating cyclin E2 expression |
| 145 | hsa-miR-26b-5p | CCNE2 | 0.72 | 5.0E-5 | 2.15 | 0 | miRNATAP | -0.22 | 0.00477 | NA | |
| 146 | hsa-miR-3065-5p | CCNE2 | 0.65 | 0.09995 | 2.15 | 0 | mirMAP | -0.2 | 0 | NA | |
| 147 | hsa-miR-30a-5p | CCNE2 | -0.92 | 0.00076 | 2.15 | 0 | miRNATAP | -0.49 | 0 | NA | |
| 148 | hsa-miR-30b-5p | CCNE2 | 0.36 | 0.13803 | 2.15 | 0 | miRNAWalker2 validate; miRTarBase | -0.2 | 0.0005 | 22384020 | A luciferase-based reporter assay demonstrated that miR-30b post-transcriptionally reduced 27% p = 0.005 of the gene expression by interacting with two binding sites in the 3'-UTR of CCNE2; The upregulation of miR-30b by trastuzumab may play a biological role in trastuzumab-induced cell growth inhibition by targeting CCNE2 |
| 149 | hsa-miR-30c-5p | CCNE2 | -0.33 | 0.1236 | 2.15 | 0 | miRNATAP | -0.44 | 0 | NA | |
| 150 | hsa-miR-30d-5p | CCNE2 | -0.92 | 4.0E-5 | 2.15 | 0 | miRNATAP | -0.35 | 0 | 25843294 | MicroRNA 30d 5p inhibits tumour cell proliferation and motility by directly targeting CCNE2 in non small cell lung cancer; In addition the re-introduction of CCNE2 expression antagonised the inhibitory effects of miR-30d-5p on the capacity of NSCLC cells for proliferation and motility |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 111 | 1618 | 4.651e-63 | 2.164e-59 |
| 2 | REGULATION OF PROTEIN MODIFICATION PROCESS | 109 | 1710 | 2.152e-58 | 3.337e-55 |
| 3 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 70 | 498 | 1.682e-58 | 3.337e-55 |
| 4 | REGULATION OF KINASE ACTIVITY | 78 | 776 | 2.65e-54 | 3.082e-51 |
| 5 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 108 | 1848 | 6.129e-54 | 5.704e-51 |
| 6 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 86 | 1036 | 1.602e-53 | 1.065e-50 |
| 7 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 86 | 1036 | 1.602e-53 | 1.065e-50 |
| 8 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 74 | 689 | 2.235e-53 | 1.3e-50 |
| 9 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 108 | 1929 | 4.454e-52 | 2.303e-49 |
| 10 | LOCOMOTION | 87 | 1114 | 5.163e-52 | 2.402e-49 |
| 11 | POSITIVE REGULATION OF CELL PROLIFERATION | 77 | 814 | 1.47e-51 | 6.22e-49 |
| 12 | PROTEIN PHOSPHORYLATION | 81 | 944 | 3.504e-51 | 1.359e-48 |
| 13 | EXTRACELLULAR STRUCTURE ORGANIZATION | 54 | 304 | 2.93e-50 | 1.049e-47 |
| 14 | REGULATION OF CELL PROLIFERATION | 95 | 1496 | 1.936e-49 | 6.436e-47 |
| 15 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 85 | 1135 | 3.356e-49 | 1.041e-46 |
| 16 | POSITIVE REGULATION OF KINASE ACTIVITY | 62 | 482 | 4.887e-49 | 1.421e-46 |
| 17 | REGULATION OF TRANSFERASE ACTIVITY | 79 | 946 | 6.775e-49 | 1.854e-46 |
| 18 | POSITIVE REGULATION OF CELL COMMUNICATION | 95 | 1532 | 1.576e-48 | 3.86e-46 |
| 19 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 94 | 1492 | 1.556e-48 | 3.86e-46 |
| 20 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 100 | 1791 | 2.416e-47 | 5.621e-45 |
| 21 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 40 | 138 | 2.249e-46 | 4.983e-44 |
| 22 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 95 | 1656 | 1.425e-45 | 3.014e-43 |
| 23 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 70 | 771 | 2.707e-45 | 5.475e-43 |
| 24 | CELL MOTILITY | 72 | 835 | 3.862e-45 | 7.189e-43 |
| 25 | LOCALIZATION OF CELL | 72 | 835 | 3.862e-45 | 7.189e-43 |
| 26 | POSITIVE REGULATION OF LOCOMOTION | 56 | 420 | 5.569e-45 | 9.966e-43 |
| 27 | RESPONSE TO ENDOGENOUS STIMULUS | 89 | 1450 | 1.231e-44 | 2.122e-42 |
| 28 | PHOSPHORYLATION | 83 | 1228 | 1.948e-44 | 3.237e-42 |
| 29 | REGULATION OF MAPK CASCADE | 65 | 660 | 4.575e-44 | 7.341e-42 |
| 30 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 63 | 616 | 1.224e-43 | 1.898e-41 |
| 31 | POSITIVE REGULATION OF MAPK CASCADE | 57 | 470 | 1.863e-43 | 2.797e-41 |
| 32 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 83 | 1275 | 3.511e-43 | 5.105e-41 |
| 33 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 89 | 1518 | 5.181e-43 | 7.305e-41 |
| 34 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 71 | 876 | 1.215e-42 | 1.663e-40 |
| 35 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 75 | 1008 | 1.266e-42 | 1.683e-40 |
| 36 | INTRACELLULAR SIGNAL TRANSDUCTION | 89 | 1572 | 8.8e-42 | 1.137e-39 |
| 37 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 54 | 470 | 7.863e-40 | 9.889e-38 |
| 38 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 88 | 1672 | 9.916e-39 | 1.214e-36 |
| 39 | RESPONSE TO GROWTH FACTOR | 53 | 475 | 2.094e-38 | 2.498e-36 |
| 40 | BIOLOGICAL ADHESION | 71 | 1032 | 6.626e-38 | 7.707e-36 |
| 41 | POSITIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 44 | 289 | 9.785e-38 | 1.11e-35 |
| 42 | RESPONSE TO EXTERNAL STIMULUS | 90 | 1821 | 1.467e-37 | 1.625e-35 |
| 43 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 79 | 1381 | 8.119e-37 | 8.786e-35 |
| 44 | REGULATION OF CELL DEATH | 81 | 1472 | 1.235e-36 | 1.306e-34 |
| 45 | TISSUE DEVELOPMENT | 81 | 1518 | 1.146e-35 | 1.185e-33 |
| 46 | PEPTIDYL TYROSINE MODIFICATION | 36 | 186 | 8.541e-35 | 8.64e-33 |
| 47 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 90 | 1977 | 9.696e-35 | 9.599e-33 |
| 48 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 77 | 1395 | 1.041e-34 | 1.009e-32 |
| 49 | INOSITOL LIPID MEDIATED SIGNALING | 31 | 124 | 9.197e-34 | 8.734e-32 |
| 50 | NEGATIVE REGULATION OF CELL DEATH | 62 | 872 | 1.345e-33 | 1.252e-31 |
| 51 | REGULATION OF MAP KINASE ACTIVITY | 42 | 319 | 2.545e-33 | 2.322e-31 |
| 52 | RESPONSE TO HORMONE | 62 | 893 | 5.248e-33 | 4.696e-31 |
| 53 | RESPONSE TO NITROGEN COMPOUND | 61 | 859 | 5.387e-33 | 4.73e-31 |
| 54 | TAXIS | 47 | 464 | 4.856e-32 | 4.184e-30 |
| 55 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 58 | 799 | 8.366e-32 | 7.078e-30 |
| 56 | POSITIVE REGULATION OF MAP KINASE ACTIVITY | 35 | 207 | 1.114e-31 | 9.253e-30 |
| 57 | CELL SUBSTRATE ADHESION | 32 | 164 | 4.26e-31 | 3.477e-29 |
| 58 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 66 | 1142 | 1.68e-30 | 1.348e-28 |
| 59 | CELLULAR RESPONSE TO NITROGEN COMPOUND | 47 | 505 | 2.24e-30 | 1.767e-28 |
| 60 | SIGNAL TRANSDUCTION BY PROTEIN PHOSPHORYLATION | 43 | 404 | 3.459e-30 | 2.682e-28 |
| 61 | REGULATION OF CELL ADHESION | 51 | 629 | 4.063e-30 | 3.099e-28 |
| 62 | VASCULATURE DEVELOPMENT | 45 | 469 | 1.221e-29 | 9.163e-28 |
| 63 | REGULATION OF PEPTIDYL TYROSINE PHOSPHORYLATION | 33 | 213 | 1.33e-28 | 9.823e-27 |
| 64 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 59 | 957 | 1.473e-28 | 1.071e-26 |
| 65 | LEUKOCYTE MIGRATION | 35 | 259 | 3.384e-28 | 2.423e-26 |
| 66 | POSITIVE REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 22 | 62 | 4.77e-28 | 3.363e-26 |
| 67 | REGULATION OF HYDROLASE ACTIVITY | 67 | 1327 | 1.563e-27 | 1.085e-25 |
| 68 | REGULATION OF CELL DIFFERENTIATION | 70 | 1492 | 6.074e-27 | 4.156e-25 |
| 69 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 52 | 788 | 1.968e-26 | 1.308e-24 |
| 70 | CIRCULATORY SYSTEM DEVELOPMENT | 52 | 788 | 1.968e-26 | 1.308e-24 |
| 71 | ANGIOGENESIS | 35 | 293 | 2.501e-26 | 1.639e-24 |
| 72 | LIPID PHOSPHORYLATION | 24 | 99 | 5.776e-26 | 3.733e-24 |
| 73 | POSITIVE REGULATION OF PEPTIDYL TYROSINE PHOSPHORYLATION | 28 | 162 | 9.555e-26 | 6.091e-24 |
| 74 | CELL DEVELOPMENT | 67 | 1426 | 9.96e-26 | 6.263e-24 |
| 75 | CELLULAR RESPONSE TO HORMONE STIMULUS | 44 | 552 | 1.264e-25 | 7.841e-24 |
| 76 | CELL MATRIX ADHESION | 25 | 119 | 2.555e-25 | 1.564e-23 |
| 77 | BLOOD VESSEL MORPHOGENESIS | 37 | 364 | 2.813e-25 | 1.7e-23 |
| 78 | PEPTIDYL AMINO ACID MODIFICATION | 52 | 841 | 4.147e-25 | 2.442e-23 |
| 79 | ORGAN MORPHOGENESIS | 52 | 841 | 4.147e-25 | 2.442e-23 |
| 80 | RESPONSE TO PEPTIDE | 38 | 404 | 1.008e-24 | 5.865e-23 |
| 81 | ACTIVATION OF PROTEIN KINASE ACTIVITY | 33 | 279 | 1.029e-24 | 5.914e-23 |
| 82 | IMMUNE SYSTEM PROCESS | 77 | 1984 | 1.394e-24 | 7.909e-23 |
| 83 | REGULATION OF IMMUNE SYSTEM PROCESS | 64 | 1403 | 8.516e-24 | 4.774e-22 |
| 84 | INTEGRIN MEDIATED SIGNALING PATHWAY | 21 | 82 | 2.129e-23 | 1.179e-21 |
| 85 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 40 | 513 | 6.029e-23 | 3.301e-21 |
| 86 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 54 | 1021 | 7.146e-23 | 3.866e-21 |
| 87 | CELLULAR RESPONSE TO PEPTIDE | 31 | 274 | 1.145e-22 | 6.125e-21 |
| 88 | POSITIVE REGULATION OF ERK1 AND ERK2 CASCADE | 26 | 172 | 2.071e-22 | 1.095e-20 |
| 89 | RESPONSE TO LIPID | 50 | 888 | 2.537e-22 | 1.326e-20 |
| 90 | PHOSPHATIDYLINOSITOL METABOLIC PROCESS | 27 | 193 | 2.566e-22 | 1.327e-20 |
| 91 | CELL PROLIFERATION | 44 | 672 | 3.509e-22 | 1.794e-20 |
| 92 | REGULATION OF ERK1 AND ERK2 CASCADE | 29 | 238 | 3.585e-22 | 1.813e-20 |
| 93 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 31 | 285 | 3.765e-22 | 1.884e-20 |
| 94 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 56 | 1152 | 5.521e-22 | 2.733e-20 |
| 95 | POSITIVE REGULATION OF HYDROLASE ACTIVITY | 50 | 905 | 5.815e-22 | 2.848e-20 |
| 96 | REGULATION OF LIPID KINASE ACTIVITY | 17 | 48 | 6.696e-22 | 3.246e-20 |
| 97 | PHOSPHATIDYLINOSITOL 3 PHOSPHATE BIOSYNTHETIC PROCESS | 17 | 49 | 1.016e-21 | 4.872e-20 |
| 98 | CELL DEATH | 52 | 1001 | 1.188e-21 | 5.642e-20 |
| 99 | RESPONSE TO WOUNDING | 40 | 563 | 1.84e-21 | 8.647e-20 |
| 100 | RESPONSE TO ABIOTIC STIMULUS | 52 | 1024 | 3.282e-21 | 1.527e-19 |
| 101 | POSITIVE REGULATION OF IMMUNE SYSTEM PROCESS | 48 | 867 | 4.144e-21 | 1.909e-19 |
| 102 | POSITIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 24 | 154 | 4.387e-21 | 2.001e-19 |
| 103 | CELL ACTIVATION | 39 | 568 | 2.086e-20 | 9.422e-19 |
| 104 | LIPID MODIFICATION | 26 | 210 | 3.802e-20 | 1.701e-18 |
| 105 | REGULATION OF VASCULATURE DEVELOPMENT | 27 | 233 | 4.026e-20 | 1.784e-18 |
| 106 | NEUROGENESIS | 59 | 1402 | 4.096e-20 | 1.798e-18 |
| 107 | PROTEIN AUTOPHOSPHORYLATION | 25 | 192 | 5.956e-20 | 2.59e-18 |
| 108 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 15 | 40 | 6.605e-20 | 2.846e-18 |
| 109 | EMBRYO DEVELOPMENT | 47 | 894 | 9.449e-20 | 4.033e-18 |
| 110 | POSITIVE REGULATION OF GENE EXPRESSION | 65 | 1733 | 1.223e-19 | 5.171e-18 |
| 111 | CELLULAR COMPONENT MORPHOGENESIS | 47 | 900 | 1.238e-19 | 5.191e-18 |
| 112 | POSITIVE REGULATION OF CELL ADHESION | 32 | 376 | 1.376e-19 | 5.717e-18 |
| 113 | WOUND HEALING | 35 | 470 | 1.745e-19 | 7.186e-18 |
| 114 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 66 | 1805 | 2.214e-19 | 9.037e-18 |
| 115 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 47 | 917 | 2.633e-19 | 1.065e-17 |
| 116 | REGULATION OF NEURON DEATH | 27 | 252 | 3.167e-19 | 1.27e-17 |
| 117 | REGULATION OF RESPONSE TO STRESS | 59 | 1468 | 3.767e-19 | 1.498e-17 |
| 118 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 44 | 823 | 9.102e-19 | 3.589e-17 |
| 119 | POSITIVE REGULATION OF CHEMOTAXIS | 20 | 120 | 2.396e-18 | 9.37e-17 |
| 120 | REGULATION OF PHOSPHOLIPID METABOLIC PROCESS | 16 | 61 | 2.783e-18 | 1.079e-16 |
| 121 | REGULATION OF CHEMOTAXIS | 23 | 180 | 3.004e-18 | 1.155e-16 |
| 122 | REGULATION OF LIPID METABOLIC PROCESS | 27 | 282 | 5.854e-18 | 2.233e-16 |
| 123 | SINGLE ORGANISM CELL ADHESION | 33 | 459 | 6.02e-18 | 2.277e-16 |
| 124 | REGULATION OF EPITHELIAL CELL MIGRATION | 22 | 166 | 7.543e-18 | 2.831e-16 |
| 125 | REGULATION OF RESPONSE TO EXTERNAL STIMULUS | 45 | 926 | 1.336e-17 | 4.973e-16 |
| 126 | REGULATION OF ENDOTHELIAL CELL MIGRATION | 19 | 114 | 1.738e-17 | 6.418e-16 |
| 127 | REGULATION OF CELL SUBSTRATE ADHESION | 22 | 173 | 1.863e-17 | 6.827e-16 |
| 128 | POSITIVE REGULATION OF RESPONSE TO EXTERNAL STIMULUS | 27 | 296 | 2.024e-17 | 7.359e-16 |
| 129 | GLYCEROPHOSPHOLIPID METABOLIC PROCESS | 27 | 297 | 2.206e-17 | 7.957e-16 |
| 130 | REGULATION OF IMMUNE RESPONSE | 43 | 858 | 2.596e-17 | 9.292e-16 |
| 131 | TUBE DEVELOPMENT | 35 | 552 | 2.809e-17 | 9.977e-16 |
| 132 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 54 | 1360 | 3.04e-17 | 1.071e-15 |
| 133 | PHOSPHATIDYLINOSITOL BIOSYNTHETIC PROCESS | 19 | 120 | 4.733e-17 | 1.656e-15 |
| 134 | REGULATION OF GTPASE ACTIVITY | 38 | 673 | 4.969e-17 | 1.725e-15 |
| 135 | RESPONSE TO STEROID HORMONE | 33 | 497 | 6.436e-17 | 2.218e-15 |
| 136 | PLATELET ACTIVATION | 20 | 142 | 7.41e-17 | 2.535e-15 |
| 137 | REGULATION OF TRANSPORT | 62 | 1804 | 7.944e-17 | 2.698e-15 |
| 138 | POSITIVE REGULATION OF TRANSPORT | 44 | 936 | 1.113e-16 | 3.751e-15 |
| 139 | REGULATION OF DEVELOPMENTAL GROWTH | 26 | 289 | 1.158e-16 | 3.875e-15 |
| 140 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 45 | 983 | 1.267e-16 | 4.209e-15 |
| 141 | NEURON PROJECTION DEVELOPMENT | 34 | 545 | 1.369e-16 | 4.518e-15 |
| 142 | EPITHELIUM DEVELOPMENT | 44 | 945 | 1.581e-16 | 5.18e-15 |
| 143 | FORMATION OF PRIMARY GERM LAYER | 18 | 110 | 1.766e-16 | 5.746e-15 |
| 144 | GLYCEROLIPID METABOLIC PROCESS | 28 | 356 | 2.383e-16 | 7.7e-15 |
| 145 | CELL SUBSTRATE JUNCTION ASSEMBLY | 13 | 41 | 2.712e-16 | 8.703e-15 |
| 146 | POSITIVE REGULATION OF CELL DIVISION | 19 | 132 | 2.983e-16 | 9.506e-15 |
| 147 | POSITIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 28 | 360 | 3.179e-16 | 1.006e-14 |
| 148 | POSITIVE REGULATION OF LIPID KINASE ACTIVITY | 12 | 32 | 3.658e-16 | 1.15e-14 |
| 149 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 40 | 801 | 4.492e-16 | 1.403e-14 |
| 150 | NEURON PROJECTION MORPHOGENESIS | 29 | 402 | 6.622e-16 | 2.054e-14 |
| 151 | REGULATION OF BODY FLUID LEVELS | 32 | 506 | 7.988e-16 | 2.461e-14 |
| 152 | CELL PROJECTION ORGANIZATION | 42 | 902 | 8.657e-16 | 2.65e-14 |
| 153 | REGULATION OF CELL CYCLE | 43 | 949 | 9.788e-16 | 2.977e-14 |
| 154 | RESPONSE TO ACID CHEMICAL | 26 | 319 | 1.276e-15 | 3.854e-14 |
| 155 | PHOSPHOLIPID METABOLIC PROCESS | 27 | 364 | 3.612e-15 | 1.084e-13 |
| 156 | SUBSTRATE ADHESION DEPENDENT CELL SPREADING | 12 | 38 | 4.143e-15 | 1.236e-13 |
| 157 | REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 19 | 152 | 4.341e-15 | 1.286e-13 |
| 158 | EMBRYONIC MORPHOGENESIS | 32 | 539 | 4.74e-15 | 1.396e-13 |
| 159 | POSITIVE REGULATION OF SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 12 | 39 | 5.927e-15 | 1.735e-13 |
| 160 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 40 | 872 | 7.692e-15 | 2.237e-13 |
| 161 | ACTIVATION OF MAPK ACTIVITY | 18 | 137 | 9.623e-15 | 2.781e-13 |
| 162 | HEAD DEVELOPMENT | 36 | 709 | 9.705e-15 | 2.787e-13 |
| 163 | RESPONSE TO CYTOKINE | 36 | 714 | 1.202e-14 | 3.431e-13 |
| 164 | CELL CHEMOTAXIS | 19 | 162 | 1.428e-14 | 4.053e-13 |
| 165 | POSITIVE REGULATION OF PHOSPHOLIPID METABOLIC PROCESS | 12 | 42 | 1.629e-14 | 4.593e-13 |
| 166 | REGULATION OF PROTEIN KINASE B SIGNALING | 17 | 121 | 1.741e-14 | 4.881e-13 |
| 167 | POSITIVE REGULATION OF EPITHELIAL CELL MIGRATION | 16 | 103 | 2.066e-14 | 5.756e-13 |
| 168 | NEURON DEVELOPMENT | 35 | 687 | 2.163e-14 | 5.992e-13 |
| 169 | UROGENITAL SYSTEM DEVELOPMENT | 24 | 299 | 2.38e-14 | 6.553e-13 |
| 170 | RESPONSE TO ALCOHOL | 26 | 362 | 2.596e-14 | 7.106e-13 |
| 171 | REGULATION OF NEURON APOPTOTIC PROCESS | 20 | 192 | 2.812e-14 | 7.652e-13 |
| 172 | POSITIVE REGULATION OF CELL CYCLE | 25 | 332 | 2.877e-14 | 7.782e-13 |
| 173 | CELL MORPHOGENESIS INVOLVED IN NEURON DIFFERENTIATION | 26 | 368 | 3.821e-14 | 1.028e-12 |
| 174 | NEGATIVE REGULATION OF NEURON DEATH | 19 | 171 | 3.89e-14 | 1.04e-12 |
| 175 | NEURON DIFFERENTIATION | 39 | 874 | 4.263e-14 | 1.133e-12 |
| 176 | REGULATION OF CELL MATRIX ADHESION | 15 | 90 | 4.67e-14 | 1.235e-12 |
| 177 | RESPONSE TO OXYGEN LEVELS | 24 | 311 | 5.676e-14 | 1.484e-12 |
| 178 | HEMOSTASIS | 24 | 311 | 5.676e-14 | 1.484e-12 |
| 179 | REPRODUCTIVE SYSTEM DEVELOPMENT | 27 | 408 | 5.82e-14 | 1.513e-12 |
| 180 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 31 | 552 | 5.906e-14 | 1.527e-12 |
| 181 | REGULATION OF PHOSPHOPROTEIN PHOSPHATASE ACTIVITY | 13 | 60 | 6.637e-14 | 1.706e-12 |
| 182 | REGULATION OF GROWTH | 33 | 633 | 6.859e-14 | 1.754e-12 |
| 183 | GASTRULATION | 18 | 155 | 8.616e-14 | 2.191e-12 |
| 184 | POSITIVE REGULATION OF VASCULATURE DEVELOPMENT | 17 | 133 | 8.664e-14 | 2.191e-12 |
| 185 | RESPONSE TO INSULIN | 20 | 205 | 9.858e-14 | 2.466e-12 |
| 186 | NEURON PROJECTION GUIDANCE | 20 | 205 | 9.858e-14 | 2.466e-12 |
| 187 | RESPONSE TO FIBROBLAST GROWTH FACTOR | 16 | 116 | 1.413e-13 | 3.515e-12 |
| 188 | POSITIVE REGULATION OF DNA METABOLIC PROCESS | 19 | 185 | 1.648e-13 | 4.079e-12 |
| 189 | IMMUNE SYSTEM DEVELOPMENT | 31 | 582 | 2.414e-13 | 5.944e-12 |
| 190 | REGULATION OF CELL DEVELOPMENT | 37 | 836 | 2.694e-13 | 6.597e-12 |
| 191 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 35 | 750 | 2.825e-13 | 6.881e-12 |
| 192 | POSITIVE REGULATION OF ENDOTHELIAL CELL MIGRATION | 13 | 67 | 3.049e-13 | 7.389e-12 |
| 193 | RESPONSE TO ESTROGEN | 20 | 218 | 3.162e-13 | 7.622e-12 |
| 194 | POSITIVE REGULATION OF INTRACELLULAR TRANSPORT | 25 | 370 | 3.333e-13 | 7.995e-12 |
| 195 | POSITIVE REGULATION OF PHOSPHOLIPASE ACTIVITY | 12 | 53 | 3.539e-13 | 8.402e-12 |
| 196 | POSITIVE REGULATION OF FIBROBLAST PROLIFERATION | 12 | 53 | 3.539e-13 | 8.402e-12 |
| 197 | POSITIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 29 | 514 | 3.668e-13 | 8.663e-12 |
| 198 | REGULATION OF NEURON DIFFERENTIATION | 30 | 554 | 3.993e-13 | 9.384e-12 |
| 199 | POSITIVE REGULATION OF DNA REPLICATION | 14 | 86 | 4.688e-13 | 1.096e-11 |
| 200 | REGULATION OF CELL PROJECTION ORGANIZATION | 30 | 558 | 4.802e-13 | 1.117e-11 |
| 201 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 31 | 602 | 5.877e-13 | 1.36e-11 |
| 202 | POSITIVE REGULATION OF CELL DEATH | 31 | 605 | 6.695e-13 | 1.542e-11 |
| 203 | POSITIVE REGULATION OF LIPID METABOLIC PROCESS | 16 | 128 | 6.769e-13 | 1.551e-11 |
| 204 | CELLULAR RESPONSE TO CYTOKINE STIMULUS | 31 | 606 | 6.99e-13 | 1.594e-11 |
| 205 | CELL JUNCTION ASSEMBLY | 16 | 129 | 7.654e-13 | 1.737e-11 |
| 206 | TISSUE MORPHOGENESIS | 29 | 533 | 9.067e-13 | 2.048e-11 |
| 207 | IMMUNE RESPONSE REGULATING CELL SURFACE RECEPTOR SIGNALING PATHWAY | 23 | 323 | 1.053e-12 | 2.368e-11 |
| 208 | FC RECEPTOR SIGNALING PATHWAY | 19 | 206 | 1.152e-12 | 2.576e-11 |
| 209 | POSITIVE REGULATION OF DEVELOPMENTAL GROWTH | 17 | 156 | 1.234e-12 | 2.747e-11 |
| 210 | GLAND DEVELOPMENT | 25 | 395 | 1.426e-12 | 3.159e-11 |
| 211 | MULTICELLULAR ORGANISM METABOLIC PROCESS | 14 | 93 | 1.433e-12 | 3.161e-11 |
| 212 | GLYCEROLIPID BIOSYNTHETIC PROCESS | 19 | 211 | 1.769e-12 | 3.882e-11 |
| 213 | CELL JUNCTION ORGANIZATION | 18 | 185 | 1.859e-12 | 4.061e-11 |
| 214 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 22 | 303 | 2.283e-12 | 4.964e-11 |
| 215 | REGULATION OF PROTEIN LOCALIZATION | 38 | 950 | 2.697e-12 | 5.837e-11 |
| 216 | MULTICELLULAR ORGANISMAL MACROMOLECULE METABOLIC PROCESS | 13 | 79 | 2.829e-12 | 6.094e-11 |
| 217 | REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 14 | 98 | 3.006e-12 | 6.446e-11 |
| 218 | REGULATION OF DNA METABOLIC PROCESS | 23 | 340 | 3.042e-12 | 6.492e-11 |
| 219 | MORPHOGENESIS OF A BRANCHING STRUCTURE | 17 | 167 | 3.764e-12 | 7.979e-11 |
| 220 | REGULATION OF ENDOTHELIAL CELL CHEMOTAXIS | 8 | 17 | 3.773e-12 | 7.979e-11 |
| 221 | REGULATION OF RESPONSE TO WOUNDING | 25 | 413 | 3.796e-12 | 7.993e-11 |
| 222 | REGULATION OF FIBROBLAST PROLIFERATION | 13 | 81 | 3.948e-12 | 8.201e-11 |
| 223 | REGULATION OF PHOSPHOLIPASE ACTIVITY | 12 | 64 | 3.924e-12 | 8.201e-11 |
| 224 | POSITIVE REGULATION OF PROTEIN KINASE B SIGNALING | 13 | 81 | 3.948e-12 | 8.201e-11 |
| 225 | POSITIVE REGULATION OF MITOTIC CELL CYCLE | 15 | 123 | 5.222e-12 | 1.08e-10 |
| 226 | RESPONSE TO ESTRADIOL | 16 | 146 | 5.305e-12 | 1.092e-10 |
| 227 | CELLULAR RESPONSE TO LIPID | 26 | 457 | 5.542e-12 | 1.136e-10 |
| 228 | POSITIVE REGULATION OF LIPASE ACTIVITY | 12 | 66 | 5.772e-12 | 1.178e-10 |
| 229 | REGULATION OF CELLULAR LOCALIZATION | 44 | 1277 | 6.041e-12 | 1.228e-10 |
| 230 | REGULATION OF CELL SIZE | 17 | 172 | 6.078e-12 | 1.23e-10 |
| 231 | FIBROBLAST GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 13 | 84 | 6.396e-12 | 1.288e-10 |
| 232 | REGULATION OF INTRACELLULAR TRANSPORT | 30 | 621 | 7.184e-12 | 1.441e-10 |
| 233 | APOPTOTIC SIGNALING PATHWAY | 21 | 289 | 7.288e-12 | 1.455e-10 |
| 234 | ACTIVATION OF IMMUNE RESPONSE | 25 | 427 | 7.851e-12 | 1.561e-10 |
| 235 | CELLULAR RESPONSE TO ACID CHEMICAL | 17 | 175 | 8.041e-12 | 1.592e-10 |
| 236 | POSITIVE REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 13 | 86 | 8.728e-12 | 1.721e-10 |
| 237 | LEUKOCYTE DIFFERENTIATION | 21 | 292 | 8.868e-12 | 1.741e-10 |
| 238 | CELL PART MORPHOGENESIS | 30 | 633 | 1.157e-11 | 2.261e-10 |
| 239 | ENDODERMAL CELL DIFFERENTIATION | 10 | 40 | 1.172e-11 | 2.282e-10 |
| 240 | PHOSPHOLIPID BIOSYNTHETIC PROCESS | 19 | 235 | 1.19e-11 | 2.308e-10 |
| 241 | REGULATION OF CELL DIVISION | 20 | 272 | 1.902e-11 | 3.672e-10 |
| 242 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 24 | 408 | 1.918e-11 | 3.688e-10 |
| 243 | RESPONSE TO AMINO ACID | 14 | 112 | 1.937e-11 | 3.71e-10 |
| 244 | POSITIVE REGULATION OF IMMUNE RESPONSE | 28 | 563 | 1.949e-11 | 3.716e-10 |
| 245 | CELLULAR RESPONSE TO VASCULAR ENDOTHELIAL GROWTH FACTOR STIMULUS | 9 | 30 | 2.072e-11 | 3.935e-10 |
| 246 | GROWTH | 24 | 410 | 2.124e-11 | 4.017e-10 |
| 247 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 29 | 609 | 2.351e-11 | 4.429e-10 |
| 248 | VASCULAR ENDOTHELIAL GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 12 | 74 | 2.384e-11 | 4.474e-10 |
| 249 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 28 | 573 | 2.941e-11 | 5.495e-10 |
| 250 | SINGLE ORGANISM BEHAVIOR | 23 | 384 | 3.605e-11 | 6.709e-10 |
| 251 | OSSIFICATION | 19 | 251 | 3.753e-11 | 6.956e-10 |
| 252 | POSITIVE REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 11 | 60 | 4.093e-11 | 7.558e-10 |
| 253 | FC EPSILON RECEPTOR SIGNALING PATHWAY | 15 | 142 | 4.251e-11 | 7.819e-10 |
| 254 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 25 | 465 | 4.91e-11 | 8.995e-10 |
| 255 | VASCULAR ENDOTHELIAL GROWTH FACTOR SIGNALING PATHWAY | 7 | 14 | 5.17e-11 | 9.434e-10 |
| 256 | ERBB SIGNALING PATHWAY | 12 | 79 | 5.306e-11 | 9.644e-10 |
| 257 | RESPIRATORY SYSTEM DEVELOPMENT | 17 | 197 | 5.354e-11 | 9.694e-10 |
| 258 | MYELOID LEUKOCYTE MIGRATION | 13 | 99 | 5.482e-11 | 9.848e-10 |
| 259 | POSITIVE REGULATION OF CELL SUBSTRATE ADHESION | 13 | 99 | 5.482e-11 | 9.848e-10 |
| 260 | RESPONSE TO DRUG | 24 | 431 | 5.978e-11 | 1.07e-09 |
| 261 | CELLULAR RESPONSE TO INSULIN STIMULUS | 15 | 146 | 6.344e-11 | 1.131e-09 |
| 262 | POSITIVE REGULATION OF CELL DEVELOPMENT | 25 | 472 | 6.745e-11 | 1.198e-09 |
| 263 | CELLULAR LIPID METABOLIC PROCESS | 35 | 913 | 7.123e-11 | 1.26e-09 |
| 264 | PLATELET DERIVED GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 9 | 34 | 7.317e-11 | 1.29e-09 |
| 265 | PEPTIDYL SERINE MODIFICATION | 15 | 148 | 7.713e-11 | 1.354e-09 |
| 266 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 22 | 363 | 7.859e-11 | 1.375e-09 |
| 267 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 24 | 437 | 7.944e-11 | 1.384e-09 |
| 268 | REGULATION OF SYNAPSE STRUCTURE OR ACTIVITY | 18 | 232 | 8.356e-11 | 1.451e-09 |
| 269 | HOMEOSTASIS OF NUMBER OF CELLS | 16 | 175 | 8.452e-11 | 1.462e-09 |
| 270 | RESPONSE TO EXTRACELLULAR STIMULUS | 24 | 441 | 9.576e-11 | 1.65e-09 |
| 271 | REGULATION OF LIPASE ACTIVITY | 12 | 83 | 9.661e-11 | 1.659e-09 |
| 272 | REGULATION OF CELLULAR RESPONSE TO STRESS | 30 | 691 | 9.92e-11 | 1.697e-09 |
| 273 | HOMEOSTATIC PROCESS | 43 | 1337 | 1.003e-10 | 1.709e-09 |
| 274 | REGULATION OF PROTEIN PHOSPHATASE TYPE 2A ACTIVITY | 8 | 24 | 1.069e-10 | 1.816e-09 |
| 275 | REGULATION OF PHOSPHATASE ACTIVITY | 14 | 128 | 1.204e-10 | 2.037e-09 |
| 276 | POSITIVE REGULATION OF GROWTH | 18 | 238 | 1.272e-10 | 2.144e-09 |
| 277 | ENDODERM FORMATION | 10 | 50 | 1.292e-10 | 2.171e-09 |
| 278 | PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 8 | 25 | 1.558e-10 | 2.607e-09 |
| 279 | REGULATION OF TYROSINE PHOSPHORYLATION OF STAT PROTEIN | 11 | 68 | 1.697e-10 | 2.821e-09 |
| 280 | POSITIVE REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 11 | 68 | 1.697e-10 | 2.821e-09 |
| 281 | REGULATION OF CELL CELL ADHESION | 22 | 380 | 1.886e-10 | 3.124e-09 |
| 282 | REGULATION OF DEPHOSPHORYLATION | 15 | 158 | 1.962e-10 | 3.237e-09 |
| 283 | RESPONSE TO ACTIVITY | 11 | 69 | 2e-10 | 3.289e-09 |
| 284 | NEGATIVE REGULATION OF NEURON APOPTOTIC PROCESS | 14 | 135 | 2.469e-10 | 4.046e-09 |
| 285 | REGULATION OF HEMOPOIESIS | 20 | 314 | 2.487e-10 | 4.06e-09 |
| 286 | REGULATION OF DNA REPLICATION | 15 | 161 | 2.561e-10 | 4.167e-09 |
| 287 | CELL CELL SIGNALING | 31 | 767 | 2.747e-10 | 4.452e-09 |
| 288 | ENDODERM DEVELOPMENT | 11 | 71 | 2.755e-10 | 4.452e-09 |
| 289 | REGULATION OF PHOSPHOLIPASE C ACTIVITY | 9 | 39 | 2.82e-10 | 4.525e-09 |
| 290 | ERBB2 SIGNALING PATHWAY | 9 | 39 | 2.82e-10 | 4.525e-09 |
| 291 | COGNITION | 18 | 251 | 3.031e-10 | 4.847e-09 |
| 292 | EPIDERMAL GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 10 | 55 | 3.51e-10 | 5.593e-09 |
| 293 | NEPHRON DEVELOPMENT | 13 | 115 | 3.718e-10 | 5.904e-09 |
| 294 | POSITIVE REGULATION OF STAT CASCADE | 11 | 73 | 3.757e-10 | 5.927e-09 |
| 295 | POSITIVE REGULATION OF JAK STAT CASCADE | 11 | 73 | 3.757e-10 | 5.927e-09 |
| 296 | REGULATION OF ORGANELLE ORGANIZATION | 39 | 1178 | 3.806e-10 | 5.983e-09 |
| 297 | LEUKOCYTE CHEMOTAXIS | 13 | 117 | 4.622e-10 | 7.24e-09 |
| 298 | REGULATION OF AXONOGENESIS | 15 | 168 | 4.67e-10 | 7.291e-09 |
| 299 | MORPHOGENESIS OF AN EPITHELIUM | 22 | 400 | 4.979e-10 | 7.748e-09 |
| 300 | REGULATION OF PEPTIDYL SERINE PHOSPHORYLATION | 13 | 118 | 5.145e-10 | 7.979e-09 |
| 301 | TELENCEPHALON DEVELOPMENT | 17 | 228 | 5.278e-10 | 8.159e-09 |
| 302 | NEGATIVE REGULATION OF CELL COMMUNICATION | 39 | 1192 | 5.347e-10 | 8.239e-09 |
| 303 | CELL CELL ADHESION | 27 | 608 | 5.704e-10 | 8.76e-09 |
| 304 | REGULATION OF CELL ACTIVATION | 24 | 484 | 6.304e-10 | 9.649e-09 |
| 305 | POSITIVE REGULATION OF ENDOTHELIAL CELL CHEMOTAXIS | 6 | 11 | 6.723e-10 | 1.026e-08 |
| 306 | CELLULAR RESPONSE TO EXTERNAL STIMULUS | 18 | 264 | 6.859e-10 | 1.043e-08 |
| 307 | POSITIVE REGULATION OF NEURON PROJECTION DEVELOPMENT | 17 | 232 | 6.902e-10 | 1.046e-08 |
| 308 | POSITIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 13 | 121 | 7.053e-10 | 1.065e-08 |
| 309 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 12 | 99 | 7.92e-10 | 1.193e-08 |
| 310 | POSITIVE REGULATION OF SMOOTH MUSCLE CELL MIGRATION | 8 | 30 | 8.046e-10 | 1.208e-08 |
| 311 | GLIOGENESIS | 15 | 175 | 8.275e-10 | 1.238e-08 |
| 312 | REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 20 | 337 | 8.579e-10 | 1.279e-08 |
| 313 | REGULATION OF TYROSINE PHOSPHORYLATION OF STAT3 PROTEIN | 9 | 44 | 9.002e-10 | 1.338e-08 |
| 314 | SENSORY ORGAN DEVELOPMENT | 24 | 493 | 9.107e-10 | 1.35e-08 |
| 315 | INSULIN RECEPTOR SIGNALING PATHWAY | 11 | 80 | 1.035e-09 | 1.529e-08 |
| 316 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 19 | 306 | 1.091e-09 | 1.606e-08 |
| 317 | SPROUTING ANGIOGENESIS | 9 | 45 | 1.115e-09 | 1.636e-08 |
| 318 | LYMPHOCYTE DIFFERENTIATION | 16 | 209 | 1.178e-09 | 1.724e-08 |
| 319 | CELLULAR RESPONSE TO STRESS | 45 | 1565 | 1.247e-09 | 1.819e-08 |
| 320 | RESPONSE TO MECHANICAL STIMULUS | 16 | 210 | 1.263e-09 | 1.837e-08 |
| 321 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 14 | 153 | 1.306e-09 | 1.887e-08 |
| 322 | PALLIUM DEVELOPMENT | 14 | 153 | 1.306e-09 | 1.887e-08 |
| 323 | HEMIDESMOSOME ASSEMBLY | 6 | 12 | 1.332e-09 | 1.919e-08 |
| 324 | AMEBOIDAL TYPE CELL MIGRATION | 14 | 154 | 1.423e-09 | 2.043e-08 |
| 325 | POSITIVE REGULATION OF CELL ACTIVATION | 19 | 311 | 1.429e-09 | 2.045e-08 |
| 326 | ODONTOGENESIS | 12 | 105 | 1.58e-09 | 2.255e-08 |
| 327 | REGULATION OF PROTEIN IMPORT INTO NUCLEUS TRANSLOCATION | 7 | 21 | 1.642e-09 | 2.337e-08 |
| 328 | CELL CYCLE PROCESS | 36 | 1081 | 1.718e-09 | 2.438e-08 |
| 329 | REGULATION OF CELL MORPHOGENESIS | 25 | 552 | 1.748e-09 | 2.473e-08 |
| 330 | BRANCHING MORPHOGENESIS OF AN EPITHELIAL TUBE | 13 | 131 | 1.898e-09 | 2.676e-08 |
| 331 | RESPONSE TO GLUCAGON | 9 | 48 | 2.051e-09 | 2.883e-08 |
| 332 | NEGATIVE REGULATION OF CELL CYCLE | 22 | 433 | 2.18e-09 | 3.055e-08 |
| 333 | BEHAVIOR | 24 | 516 | 2.246e-09 | 3.138e-08 |
| 334 | FOREBRAIN DEVELOPMENT | 20 | 357 | 2.322e-09 | 3.235e-08 |
| 335 | POSITIVE REGULATION OF PROTEIN IMPORT INTO NUCLEUS TRANSLOCATION | 6 | 13 | 2.452e-09 | 3.406e-08 |
| 336 | REGULATION OF SMOOTH MUSCLE CELL MIGRATION | 9 | 49 | 2.489e-09 | 3.447e-08 |
| 337 | CELL CYCLE | 40 | 1316 | 2.612e-09 | 3.606e-08 |
| 338 | REGULATION OF ADHERENS JUNCTION ORGANIZATION | 9 | 50 | 3.007e-09 | 4.137e-08 |
| 339 | GLIAL CELL DIFFERENTIATION | 13 | 136 | 3.014e-09 | 4.137e-08 |
| 340 | CONNECTIVE TISSUE DEVELOPMENT | 15 | 194 | 3.448e-09 | 4.718e-08 |
| 341 | REGULATION OF BLOOD VESSEL ENDOTHELIAL CELL MIGRATION | 9 | 51 | 3.618e-09 | 4.936e-08 |
| 342 | POSITIVE REGULATION OF CELLULAR COMPONENT BIOGENESIS | 21 | 406 | 3.763e-09 | 5.12e-08 |
| 343 | POSITIVE REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 8 | 36 | 3.934e-09 | 5.321e-08 |
| 344 | POSITIVE CHEMOTAXIS | 8 | 36 | 3.934e-09 | 5.321e-08 |
| 345 | REGULATION OF SYNAPTIC PLASTICITY | 13 | 140 | 4.304e-09 | 5.805e-08 |
| 346 | REGULATION OF CELLULAR COMPONENT BIOGENESIS | 29 | 767 | 4.959e-09 | 6.668e-08 |
| 347 | POSITIVE REGULATION OF TYROSINE PHOSPHORYLATION OF STAT3 PROTEIN | 8 | 37 | 4.972e-09 | 6.668e-08 |
| 348 | RESPONSE TO RADIATION | 21 | 413 | 5.088e-09 | 6.803e-08 |
| 349 | CELLULAR RESPONSE TO AMINO ACID STIMULUS | 9 | 53 | 5.172e-09 | 6.895e-08 |
| 350 | INFLAMMATORY RESPONSE | 22 | 454 | 5.194e-09 | 6.905e-08 |
| 351 | REGULATION OF LEUKOCYTE DIFFERENTIATION | 16 | 232 | 5.339e-09 | 7.077e-08 |
| 352 | REGULATION OF CELLULAR COMPONENT SIZE | 19 | 337 | 5.36e-09 | 7.085e-08 |
| 353 | MAMMARY GLAND DEVELOPMENT | 12 | 117 | 5.535e-09 | 7.295e-08 |
| 354 | REPRODUCTION | 39 | 1297 | 5.772e-09 | 7.587e-08 |
| 355 | REGULATION OF JAK STAT CASCADE | 13 | 144 | 6.075e-09 | 7.941e-08 |
| 356 | REGULATION OF STAT CASCADE | 13 | 144 | 6.075e-09 | 7.941e-08 |
| 357 | MESODERM DEVELOPMENT | 12 | 118 | 6.103e-09 | 7.954e-08 |
| 358 | CHEMICAL HOMEOSTASIS | 31 | 874 | 6.255e-09 | 8.13e-08 |
| 359 | LYMPHOCYTE ACTIVATION | 19 | 342 | 6.813e-09 | 8.83e-08 |
| 360 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 35 | 1087 | 7.1e-09 | 9.176e-08 |
| 361 | PEPTIDYL TYROSINE AUTOPHOSPHORYLATION | 8 | 39 | 7.778e-09 | 1.002e-07 |
| 362 | GLAND MORPHOGENESIS | 11 | 97 | 8.367e-09 | 1.075e-07 |
| 363 | MESODERMAL CELL DIFFERENTIATION | 7 | 26 | 8.871e-09 | 1.137e-07 |
| 364 | REGULATION OF MITOTIC CELL CYCLE | 22 | 468 | 9.012e-09 | 1.152e-07 |
| 365 | MEMORY | 11 | 98 | 9.336e-09 | 1.187e-07 |
| 366 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 11 | 98 | 9.336e-09 | 1.187e-07 |
| 367 | POSITIVE REGULATION OF CELL MATRIX ADHESION | 8 | 40 | 9.632e-09 | 1.221e-07 |
| 368 | POSITIVE REGULATION OF CELL CELL ADHESION | 16 | 243 | 1.034e-08 | 1.307e-07 |
| 369 | LIPID METABOLIC PROCESS | 36 | 1158 | 1.043e-08 | 1.316e-07 |
| 370 | REGULATION OF ANATOMICAL STRUCTURE SIZE | 22 | 472 | 1.051e-08 | 1.321e-07 |
| 371 | SKIN DEVELOPMENT | 15 | 211 | 1.082e-08 | 1.357e-07 |
| 372 | POSITIVE REGULATION OF VASCULAR ENDOTHELIAL GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 6 | 16 | 1.114e-08 | 1.393e-07 |
| 373 | REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 11 | 100 | 1.158e-08 | 1.444e-07 |
| 374 | RESPONSE TO KETONE | 14 | 182 | 1.244e-08 | 1.548e-07 |
| 375 | REGULATION OF PROTEIN IMPORT | 14 | 183 | 1.334e-08 | 1.656e-07 |
| 376 | REGULATION OF CELLULAR RESPONSE TO INSULIN STIMULUS | 9 | 59 | 1.386e-08 | 1.715e-07 |
| 377 | REGULATION OF GLUCOSE IMPORT | 9 | 60 | 1.615e-08 | 1.994e-07 |
| 378 | TUBE MORPHOGENESIS | 18 | 323 | 1.648e-08 | 2.028e-07 |
| 379 | REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 15 | 218 | 1.679e-08 | 2.062e-07 |
| 380 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 12 | 129 | 1.686e-08 | 2.064e-07 |
| 381 | REGULATION OF GLUCOSE IMPORT IN RESPONSE TO INSULIN STIMULUS | 6 | 17 | 1.705e-08 | 2.077e-07 |
| 382 | NEGATIVE REGULATION OF ANOIKIS | 6 | 17 | 1.705e-08 | 2.077e-07 |
| 383 | POSITIVE REGULATION OF PROTEIN IMPORT | 11 | 104 | 1.756e-08 | 2.133e-07 |
| 384 | REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 8 | 43 | 1.766e-08 | 2.134e-07 |
| 385 | REGULATION OF POLYSACCHARIDE METABOLIC PROCESS | 8 | 43 | 1.766e-08 | 2.134e-07 |
| 386 | REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 15 | 220 | 1.898e-08 | 2.288e-07 |
| 387 | CEREBRAL CORTEX DEVELOPMENT | 11 | 105 | 1.943e-08 | 2.336e-07 |
| 388 | CELL CYCLE PHASE TRANSITION | 16 | 255 | 2.043e-08 | 2.437e-07 |
| 389 | REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 7 | 29 | 2.048e-08 | 2.437e-07 |
| 390 | POSITIVE REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 7 | 29 | 2.048e-08 | 2.437e-07 |
| 391 | LEUKOCYTE CELL CELL ADHESION | 16 | 255 | 2.043e-08 | 2.437e-07 |
| 392 | REGENERATION | 13 | 161 | 2.342e-08 | 2.78e-07 |
| 393 | PLATELET DEGRANULATION | 11 | 107 | 2.371e-08 | 2.807e-07 |
| 394 | CYTOKINE MEDIATED SIGNALING PATHWAY | 21 | 452 | 2.445e-08 | 2.887e-07 |
| 395 | LIPID BIOSYNTHETIC PROCESS | 23 | 539 | 2.464e-08 | 2.903e-07 |
| 396 | TISSUE MIGRATION | 10 | 84 | 2.52e-08 | 2.961e-07 |
| 397 | POSITIVE REGULATION OF HEMOPOIESIS | 13 | 163 | 2.715e-08 | 3.182e-07 |
| 398 | SKELETAL SYSTEM DEVELOPMENT | 21 | 455 | 2.739e-08 | 3.202e-07 |
| 399 | LEUKOCYTE ACTIVATION | 20 | 414 | 2.819e-08 | 3.287e-07 |
| 400 | POSITIVE REGULATION OF LEUKOCYTE MIGRATION | 11 | 109 | 2.881e-08 | 3.351e-07 |
| 401 | RHYTHMIC PROCESS | 17 | 298 | 2.972e-08 | 3.449e-07 |
| 402 | ANTIGEN RECEPTOR MEDIATED SIGNALING PATHWAY | 14 | 195 | 2.99e-08 | 3.461e-07 |
| 403 | POSITIVE REGULATION OF LEUKOCYTE PROLIFERATION | 12 | 136 | 3.058e-08 | 3.523e-07 |
| 404 | POSITIVE REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 15 | 228 | 3.059e-08 | 3.523e-07 |
| 405 | CELLULAR RESPONSE TO ABIOTIC STIMULUS | 16 | 263 | 3.149e-08 | 3.618e-07 |
| 406 | REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 15 | 229 | 3.242e-08 | 3.716e-07 |
| 407 | MODULATION OF SYNAPTIC TRANSMISSION | 17 | 301 | 3.441e-08 | 3.934e-07 |
| 408 | CELL CYCLE G1 S PHASE TRANSITION | 11 | 111 | 3.486e-08 | 3.966e-07 |
| 409 | G1 S TRANSITION OF MITOTIC CELL CYCLE | 11 | 111 | 3.486e-08 | 3.966e-07 |
| 410 | POSITIVE REGULATION OF PEPTIDYL SERINE PHOSPHORYLATION | 10 | 88 | 3.967e-08 | 4.502e-07 |
| 411 | HEART DEVELOPMENT | 21 | 466 | 4.12e-08 | 4.664e-07 |
| 412 | POSITIVE REGULATION OF REACTIVE OXYGEN SPECIES BIOSYNTHETIC PROCESS | 8 | 48 | 4.387e-08 | 4.954e-07 |
| 413 | REGULATION OF GENERATION OF PRECURSOR METABOLITES AND ENERGY | 10 | 89 | 4.427e-08 | 4.988e-07 |
| 414 | POSITIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 13 | 171 | 4.803e-08 | 5.398e-07 |
| 415 | DEFENSE RESPONSE | 36 | 1231 | 4.945e-08 | 5.544e-07 |
| 416 | REGULATION OF CELL JUNCTION ASSEMBLY | 9 | 68 | 4.996e-08 | 5.588e-07 |
| 417 | CELLULAR RESPONSE TO GROWTH HORMONE STIMULUS | 6 | 20 | 5.198e-08 | 5.786e-07 |
| 418 | GLOMERULUS DEVELOPMENT | 8 | 49 | 5.194e-08 | 5.786e-07 |
| 419 | RAS PROTEIN SIGNAL TRANSDUCTION | 12 | 143 | 5.359e-08 | 5.951e-07 |
| 420 | POSITIVE REGULATION OF AXONOGENESIS | 9 | 69 | 5.692e-08 | 6.306e-07 |
| 421 | NEGATIVE REGULATION OF CELL PROJECTION ORGANIZATION | 12 | 144 | 5.791e-08 | 6.4e-07 |
| 422 | NEGATIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 11 | 117 | 6.039e-08 | 6.659e-07 |
| 423 | MITOCHONDRIAL MEMBRANE ORGANIZATION | 10 | 92 | 6.103e-08 | 6.713e-07 |
| 424 | RESPONSE TO OXIDATIVE STRESS | 18 | 352 | 6.117e-08 | 6.713e-07 |
| 425 | REGULATION OF DEFENSE RESPONSE | 27 | 759 | 6.177e-08 | 6.763e-07 |
| 426 | REGULATION OF MEMBRANE PERMEABILITY | 9 | 70 | 6.47e-08 | 7.067e-07 |
| 427 | RESPONSE TO INORGANIC SUBSTANCE | 21 | 479 | 6.57e-08 | 7.159e-07 |
| 428 | T CELL RECEPTOR SIGNALING PATHWAY | 12 | 146 | 6.749e-08 | 7.337e-07 |
| 429 | REGULATION OF CYTOPLASMIC TRANSPORT | 21 | 481 | 7.049e-08 | 7.645e-07 |
| 430 | RESPONSE TO LIGHT STIMULUS | 16 | 280 | 7.509e-08 | 8.126e-07 |
| 431 | FC GAMMA RECEPTOR SIGNALING PATHWAY | 10 | 95 | 8.314e-08 | 8.975e-07 |
| 432 | POSITIVE REGULATION OF CELL CYCLE PROCESS | 15 | 247 | 8.798e-08 | 9.476e-07 |
| 433 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 14 | 213 | 9.037e-08 | 9.711e-07 |
| 434 | MAMMARY GLAND EPITHELIUM DEVELOPMENT | 8 | 53 | 9.835e-08 | 1.052e-06 |
| 435 | REGULATION OF NITRIC OXIDE BIOSYNTHETIC PROCESS | 8 | 53 | 9.835e-08 | 1.052e-06 |
| 436 | REGULATION OF HOMEOSTATIC PROCESS | 20 | 447 | 9.884e-08 | 1.055e-06 |
| 437 | EMBRYONIC ORGAN DEVELOPMENT | 19 | 406 | 1.046e-07 | 1.114e-06 |
| 438 | REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 18 | 365 | 1.055e-07 | 1.121e-06 |
| 439 | CELL ADHESION MEDIATED BY INTEGRIN | 5 | 12 | 1.069e-07 | 1.133e-06 |
| 440 | DEVELOPMENT OF PRIMARY SEXUAL CHARACTERISTICS | 14 | 216 | 1.075e-07 | 1.137e-06 |
| 441 | REGULATION OF RAS PROTEIN SIGNAL TRANSDUCTION | 13 | 184 | 1.138e-07 | 1.201e-06 |
| 442 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 22 | 541 | 1.171e-07 | 1.23e-06 |
| 443 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 22 | 541 | 1.171e-07 | 1.23e-06 |
| 444 | POSITIVE REGULATION OF CARBOHYDRATE METABOLIC PROCESS | 9 | 75 | 1.192e-07 | 1.25e-06 |
| 445 | EPIDERMIS DEVELOPMENT | 15 | 253 | 1.204e-07 | 1.258e-06 |
| 446 | CELL CYCLE ARREST | 12 | 154 | 1.216e-07 | 1.269e-06 |
| 447 | LIMBIC SYSTEM DEVELOPMENT | 10 | 100 | 1.359e-07 | 1.414e-06 |
| 448 | DEVELOPMENTAL GROWTH | 17 | 333 | 1.478e-07 | 1.535e-06 |
| 449 | CELLULAR RESPONSE TO GLUCAGON STIMULUS | 7 | 38 | 1.524e-07 | 1.579e-06 |
| 450 | POSITIVE REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 15 | 258 | 1.552e-07 | 1.605e-06 |
| 451 | REGULATED EXOCYTOSIS | 14 | 224 | 1.683e-07 | 1.737e-06 |
| 452 | CELLULAR RESPONSE TO PROSTAGLANDIN STIMULUS | 6 | 24 | 1.741e-07 | 1.784e-06 |
| 453 | REGULATION OF POSITIVE CHEMOTAXIS | 6 | 24 | 1.741e-07 | 1.784e-06 |
| 454 | REGULATION OF ANOIKIS | 6 | 24 | 1.741e-07 | 1.784e-06 |
| 455 | RESPONSE TO NUTRIENT | 13 | 191 | 1.759e-07 | 1.798e-06 |
| 456 | ENDOTHELIAL CELL MIGRATION | 8 | 57 | 1.767e-07 | 1.803e-06 |
| 457 | POSITIVE REGULATION OF LEUKOCYTE DIFFERENTIATION | 11 | 131 | 1.934e-07 | 1.969e-06 |
| 458 | B CELL ACTIVATION | 11 | 132 | 2.09e-07 | 2.123e-06 |
| 459 | NEGATIVE REGULATION OF VASCULATURE DEVELOPMENT | 9 | 80 | 2.101e-07 | 2.13e-06 |
| 460 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 12 | 162 | 2.117e-07 | 2.141e-06 |
| 461 | NEGATIVE REGULATION OF CELL DEVELOPMENT | 16 | 303 | 2.206e-07 | 2.226e-06 |
| 462 | CELLULAR EXTRAVASATION | 6 | 25 | 2.27e-07 | 2.281e-06 |
| 463 | POSITIVE REGULATION OF BLOOD VESSEL ENDOTHELIAL CELL MIGRATION | 6 | 25 | 2.27e-07 | 2.281e-06 |
| 464 | CELL GROWTH | 11 | 135 | 2.627e-07 | 2.634e-06 |
| 465 | REGULATION OF HOMOTYPIC CELL CELL ADHESION | 16 | 307 | 2.633e-07 | 2.635e-06 |
| 466 | INSULIN LIKE GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 5 | 14 | 2.655e-07 | 2.651e-06 |
| 467 | RESPONSE TO ETHANOL | 11 | 136 | 2.831e-07 | 2.821e-06 |
| 468 | ORGAN REGENERATION | 9 | 83 | 2.895e-07 | 2.878e-06 |
| 469 | DEVELOPMENTAL PROGRAMMED CELL DEATH | 6 | 26 | 2.924e-07 | 2.901e-06 |
| 470 | PLACENTA DEVELOPMENT | 11 | 138 | 3.282e-07 | 3.25e-06 |
| 471 | REGULATION OF VASCULAR ENDOTHELIAL GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 6 | 27 | 3.725e-07 | 3.665e-06 |
| 472 | POSITIVE REGULATION OF MESENCHYMAL CELL PROLIFERATION | 6 | 27 | 3.725e-07 | 3.665e-06 |
| 473 | HETEROTYPIC CELL CELL ADHESION | 6 | 27 | 3.725e-07 | 3.665e-06 |
| 474 | TISSUE HOMEOSTASIS | 12 | 171 | 3.803e-07 | 3.733e-06 |
| 475 | CELL MIGRATION INVOLVED IN SPROUTING ANGIOGENESIS | 5 | 15 | 3.947e-07 | 3.86e-06 |
| 476 | REGULATION OF INNATE IMMUNE RESPONSE | 17 | 357 | 3.948e-07 | 3.86e-06 |
| 477 | REGULATION OF CARBOHYDRATE METABOLIC PROCESS | 12 | 172 | 4.05e-07 | 3.95e-06 |
| 478 | RESPONSE TO TOXIC SUBSTANCE | 14 | 241 | 4.108e-07 | 3.999e-06 |
| 479 | REGULATION OF LEUKOCYTE PROLIFERATION | 13 | 206 | 4.203e-07 | 4.083e-06 |
| 480 | REGULATION OF CARBOHYDRATE BIOSYNTHETIC PROCESS | 9 | 87 | 4.352e-07 | 4.218e-06 |
| 481 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 30 | 1004 | 4.751e-07 | 4.596e-06 |
| 482 | SECRETION | 22 | 588 | 4.847e-07 | 4.679e-06 |
| 483 | POSITIVE REGULATION OF CYTOPLASMIC TRANSPORT | 15 | 282 | 4.86e-07 | 4.682e-06 |
| 484 | REGULATION OF REACTIVE OXYGEN SPECIES BIOSYNTHETIC PROCESS | 8 | 65 | 5.01e-07 | 4.816e-06 |
| 485 | RESPONSE TO TRANSFORMING GROWTH FACTOR BETA | 11 | 144 | 5.04e-07 | 4.835e-06 |
| 486 | ORGANOPHOSPHATE BIOSYNTHETIC PROCESS | 19 | 450 | 5.069e-07 | 4.853e-06 |
| 487 | EXOCRINE SYSTEM DEVELOPMENT | 7 | 45 | 5.138e-07 | 4.909e-06 |
| 488 | RESPONSE TO CORTICOSTEROID | 12 | 176 | 5.185e-07 | 4.933e-06 |
| 489 | POSITIVE REGULATION OF DEFENSE RESPONSE | 17 | 364 | 5.177e-07 | 4.933e-06 |
| 490 | EPITHELIAL CELL PROLIFERATION | 9 | 89 | 5.293e-07 | 5.026e-06 |
| 491 | MESODERM MORPHOGENESIS | 8 | 66 | 5.648e-07 | 5.352e-06 |
| 492 | BRANCHING INVOLVED IN SALIVARY GLAND MORPHOGENESIS | 5 | 16 | 5.69e-07 | 5.382e-06 |
| 493 | POSITIVE REGULATION OF NEURON DEATH | 8 | 67 | 6.354e-07 | 5.997e-06 |
| 494 | POSITIVE REGULATION OF CYTOKINE PRODUCTION | 17 | 370 | 6.496e-07 | 6.119e-06 |
| 495 | NEGATIVE REGULATION OF TRANSPORT | 19 | 458 | 6.609e-07 | 6.213e-06 |
| 496 | DIGESTIVE SYSTEM DEVELOPMENT | 11 | 148 | 6.632e-07 | 6.221e-06 |
| 497 | REGULATION OF LEUKOCYTE MIGRATION | 11 | 149 | 7.093e-07 | 6.64e-06 |
| 498 | MULTICELLULAR ORGANISMAL RESPONSE TO STRESS | 8 | 68 | 7.135e-07 | 6.64e-06 |
| 499 | REGULATION OF NEUROLOGICAL SYSTEM PROCESS | 8 | 68 | 7.135e-07 | 6.64e-06 |
| 500 | REGULATION OF TOR SIGNALING | 8 | 68 | 7.135e-07 | 6.64e-06 |
| 501 | RESPONSE TO GROWTH HORMONE | 6 | 30 | 7.273e-07 | 6.755e-06 |
| 502 | REGULATION OF SYSTEM PROCESS | 20 | 507 | 7.298e-07 | 6.765e-06 |
| 503 | REGULATION OF MYELOID CELL DIFFERENTIATION | 12 | 183 | 7.864e-07 | 7.275e-06 |
| 504 | POSITIVE REGULATION OF GLYCOGEN METABOLIC PROCESS | 5 | 17 | 7.99e-07 | 7.377e-06 |
| 505 | INTRINSIC APOPTOTIC SIGNALING PATHWAY | 11 | 152 | 8.651e-07 | 7.971e-06 |
| 506 | HOMEOSTASIS OF NUMBER OF CELLS WITHIN A TISSUE | 6 | 31 | 8.937e-07 | 8.202e-06 |
| 507 | REGULATION OF PLATELET ACTIVATION | 6 | 31 | 8.937e-07 | 8.202e-06 |
| 508 | REGULATION OF CYTOKINE PRODUCTION | 21 | 563 | 9.401e-07 | 8.611e-06 |
| 509 | INTERSPECIES INTERACTION BETWEEN ORGANISMS | 23 | 662 | 9.47e-07 | 8.633e-06 |
| 510 | SYMBIOSIS ENCOMPASSING MUTUALISM THROUGH PARASITISM | 23 | 662 | 9.47e-07 | 8.633e-06 |
| 511 | T CELL DIFFERENTIATION | 10 | 123 | 9.481e-07 | 8.633e-06 |
| 512 | REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 17 | 381 | 9.73e-07 | 8.843e-06 |
| 513 | NEGATIVE REGULATION OF CELL ADHESION | 13 | 223 | 1.033e-06 | 9.37e-06 |
| 514 | CELLULAR RESPONSE TO EXTRACELLULAR STIMULUS | 12 | 188 | 1.047e-06 | 9.477e-06 |
| 515 | STAT CASCADE | 7 | 50 | 1.08e-06 | 9.74e-06 |
| 516 | JAK STAT CASCADE | 7 | 50 | 1.08e-06 | 9.74e-06 |
| 517 | SALIVARY GLAND DEVELOPMENT | 6 | 32 | 1.09e-06 | 9.791e-06 |
| 518 | REGULATION OF ACTIN CYTOSKELETON REORGANIZATION | 6 | 32 | 1.09e-06 | 9.791e-06 |
| 519 | POSITIVE REGULATION OF SYNAPTIC TRANSMISSION GLUTAMATERGIC | 5 | 18 | 1.097e-06 | 9.813e-06 |
| 520 | CELLULAR RESPONSE TO PROSTAGLANDIN E STIMULUS | 5 | 18 | 1.097e-06 | 9.813e-06 |
| 521 | BONE DEVELOPMENT | 11 | 156 | 1.119e-06 | 9.996e-06 |
| 522 | CELLULAR CHEMICAL HOMEOSTASIS | 21 | 570 | 1.145e-06 | 1.021e-05 |
| 523 | MESENCHYME DEVELOPMENT | 12 | 190 | 1.171e-06 | 1.04e-05 |
| 524 | PHAGOCYTOSIS | 12 | 190 | 1.171e-06 | 1.04e-05 |
| 525 | REGULATION OF WOUND HEALING | 10 | 126 | 1.184e-06 | 1.049e-05 |
| 526 | RESPONSE TO VITAMIN | 9 | 98 | 1.204e-06 | 1.065e-05 |
| 527 | POSITIVE REGULATION OF MITOTIC NUCLEAR DIVISION | 7 | 51 | 1.24e-06 | 1.083e-05 |
| 528 | NEGATIVE REGULATION OF NEURON DIFFERENTIATION | 12 | 191 | 1.237e-06 | 1.083e-05 |
| 529 | MUSCLE STRUCTURE DEVELOPMENT | 18 | 432 | 1.237e-06 | 1.083e-05 |
| 530 | HIPPOCAMPUS DEVELOPMENT | 8 | 73 | 1.239e-06 | 1.083e-05 |
| 531 | REGULATION OF ORGAN GROWTH | 8 | 73 | 1.239e-06 | 1.083e-05 |
| 532 | CELLULAR RESPONSE TO FATTY ACID | 7 | 51 | 1.24e-06 | 1.083e-05 |
| 533 | RESPONSE TO REACTIVE OXYGEN SPECIES | 12 | 191 | 1.237e-06 | 1.083e-05 |
| 534 | ORGANOPHOSPHATE METABOLIC PROCESS | 27 | 885 | 1.269e-06 | 1.106e-05 |
| 535 | SYSTEM PROCESS | 42 | 1785 | 1.278e-06 | 1.112e-05 |
| 536 | RESPONSE TO BIOTIC STIMULUS | 27 | 886 | 1.296e-06 | 1.125e-05 |
| 537 | SIGNAL TRANSDUCTION IN ABSENCE OF LIGAND | 6 | 33 | 1.32e-06 | 1.142e-05 |
| 538 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 6 | 33 | 1.32e-06 | 1.142e-05 |
| 539 | SEX DIFFERENTIATION | 14 | 266 | 1.338e-06 | 1.155e-05 |
| 540 | REGULATION OF CELL GROWTH | 17 | 391 | 1.387e-06 | 1.195e-05 |
| 541 | REGULATION OF PROTEIN TARGETING | 15 | 307 | 1.413e-06 | 1.215e-05 |
| 542 | REGULATION OF GLUCOSE TRANSPORT | 9 | 100 | 1.429e-06 | 1.227e-05 |
| 543 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 18 | 437 | 1.457e-06 | 1.248e-05 |
| 544 | KIDNEY VASCULATURE DEVELOPMENT | 5 | 19 | 1.475e-06 | 1.259e-05 |
| 545 | RENAL SYSTEM VASCULATURE DEVELOPMENT | 5 | 19 | 1.475e-06 | 1.259e-05 |
| 546 | REGULATION OF MESENCHYMAL CELL PROLIFERATION | 6 | 34 | 1.589e-06 | 1.349e-05 |
| 547 | RESPONSE TO PROSTAGLANDIN | 6 | 34 | 1.589e-06 | 1.349e-05 |
| 548 | PROTEIN KINASE B SIGNALING | 6 | 34 | 1.589e-06 | 1.349e-05 |
| 549 | SECRETION BY CELL | 19 | 486 | 1.598e-06 | 1.352e-05 |
| 550 | IMMUNE EFFECTOR PROCESS | 19 | 486 | 1.598e-06 | 1.352e-05 |
| 551 | GLIAL CELL DEVELOPMENT | 8 | 76 | 1.69e-06 | 1.427e-05 |
| 552 | NEGATIVE REGULATION OF RESPONSE TO EXTERNAL STIMULUS | 14 | 274 | 1.897e-06 | 1.599e-05 |
| 553 | REGULATION OF GLYCOGEN METABOLIC PROCESS | 6 | 35 | 1.9e-06 | 1.599e-05 |
| 554 | REGULATION OF NEURON PROJECTION REGENERATION | 5 | 20 | 1.95e-06 | 1.637e-05 |
| 555 | INTERACTION WITH HOST | 10 | 134 | 2.078e-06 | 1.742e-05 |
| 556 | REGULATION OF ION TRANSPORT | 21 | 592 | 2.086e-06 | 1.746e-05 |
| 557 | EPITHELIAL CELL DIFFERENTIATION | 19 | 495 | 2.091e-06 | 1.747e-05 |
| 558 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 30 | 1079 | 2.097e-06 | 1.748e-05 |
| 559 | REGULATION OF SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 14 | 278 | 2.247e-06 | 1.871e-05 |
| 560 | POSITIVE REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 6 | 36 | 2.26e-06 | 1.878e-05 |
| 561 | POSITIVE REGULATION OF MUSCLE TISSUE DEVELOPMENT | 7 | 56 | 2.373e-06 | 1.968e-05 |
| 562 | RESPONSE TO MOLECULE OF BACTERIAL ORIGIN | 15 | 321 | 2.451e-06 | 2.029e-05 |
| 563 | ACTIVATION OF INNATE IMMUNE RESPONSE | 12 | 204 | 2.467e-06 | 2.039e-05 |
| 564 | NEGATIVE REGULATION OF LIPID METABOLIC PROCESS | 8 | 80 | 2.503e-06 | 2.061e-05 |
| 565 | CELLULAR RESPONSE TO MECHANICAL STIMULUS | 8 | 80 | 2.503e-06 | 2.061e-05 |
| 566 | RESPONSE TO ALKALOID | 10 | 137 | 2.54e-06 | 2.084e-05 |
| 567 | POSITIVE REGULATION OF ADHERENS JUNCTION ORGANIZATION | 5 | 21 | 2.536e-06 | 2.084e-05 |
| 568 | GLUCOSE HOMEOSTASIS | 11 | 170 | 2.602e-06 | 2.127e-05 |
| 569 | CARBOHYDRATE HOMEOSTASIS | 11 | 170 | 2.602e-06 | 2.127e-05 |
| 570 | REGULATION OF METAL ION TRANSPORT | 15 | 325 | 2.853e-06 | 2.329e-05 |
| 571 | RESPONSE TO HYDROGEN PEROXIDE | 9 | 109 | 2.944e-06 | 2.399e-05 |
| 572 | EYE DEVELOPMENT | 15 | 326 | 2.962e-06 | 2.41e-05 |
| 573 | REGULATION OF MULTICELLULAR ORGANISMAL METABOLIC PROCESS | 6 | 38 | 3.145e-06 | 2.554e-05 |
| 574 | NEURON MIGRATION | 9 | 110 | 3.176e-06 | 2.57e-05 |
| 575 | POSITIVE REGULATION OF SYNAPTIC TRANSMISSION | 9 | 110 | 3.176e-06 | 2.57e-05 |
| 576 | POSITIVE REGULATION OF PROTEIN AUTOPHOSPHORYLATION | 5 | 22 | 3.253e-06 | 2.628e-05 |
| 577 | MITOTIC CELL CYCLE | 24 | 766 | 3.29e-06 | 2.653e-05 |
| 578 | REGULATION OF EPITHELIAL CELL APOPTOTIC PROCESS | 7 | 59 | 3.396e-06 | 2.734e-05 |
| 579 | ASTROCYTE DIFFERENTIATION | 6 | 39 | 3.684e-06 | 2.96e-05 |
| 580 | LEUKOCYTE HOMEOSTASIS | 7 | 60 | 3.81e-06 | 3.056e-05 |
| 581 | MITOCHONDRIAL TRANSPORT | 11 | 177 | 3.847e-06 | 3.081e-05 |
| 582 | NEGATIVE REGULATION OF PHOSPHORYLATION | 17 | 422 | 3.876e-06 | 3.099e-05 |
| 583 | EPHRIN RECEPTOR SIGNALING PATHWAY | 8 | 85 | 3.967e-06 | 3.166e-05 |
| 584 | CELLULAR MACROMOLECULE LOCALIZATION | 32 | 1234 | 4.026e-06 | 3.208e-05 |
| 585 | POSITIVE REGULATION OF CELLULAR RESPONSE TO INSULIN STIMULUS | 5 | 23 | 4.12e-06 | 3.266e-05 |
| 586 | POSITIVE REGULATION OF NEUROLOGICAL SYSTEM PROCESS | 5 | 23 | 4.12e-06 | 3.266e-05 |
| 587 | NEUROTROPHIN SIGNALING PATHWAY | 5 | 23 | 4.12e-06 | 3.266e-05 |
| 588 | SYNAPSE ORGANIZATION | 10 | 145 | 4.235e-06 | 3.351e-05 |
| 589 | OVARIAN FOLLICLE DEVELOPMENT | 7 | 61 | 4.264e-06 | 3.363e-05 |
| 590 | POSITIVE REGULATION OF STEM CELL PROLIFERATION | 7 | 61 | 4.264e-06 | 3.363e-05 |
| 591 | MOVEMENT IN ENVIRONMENT OF OTHER ORGANISM INVOLVED IN SYMBIOTIC INTERACTION | 8 | 87 | 4.728e-06 | 3.685e-05 |
| 592 | ENTRY INTO CELL OF OTHER ORGANISM INVOLVED IN SYMBIOTIC INTERACTION | 8 | 87 | 4.728e-06 | 3.685e-05 |
| 593 | VIRAL ENTRY INTO HOST CELL | 8 | 87 | 4.728e-06 | 3.685e-05 |
| 594 | ENTRY INTO OTHER ORGANISM INVOLVED IN SYMBIOTIC INTERACTION | 8 | 87 | 4.728e-06 | 3.685e-05 |
| 595 | MOVEMENT IN HOST ENVIRONMENT | 8 | 87 | 4.728e-06 | 3.685e-05 |
| 596 | ENTRY INTO HOST | 8 | 87 | 4.728e-06 | 3.685e-05 |
| 597 | ENTRY INTO HOST CELL | 8 | 87 | 4.728e-06 | 3.685e-05 |
| 598 | FOREBRAIN CELL MIGRATION | 7 | 62 | 4.763e-06 | 3.694e-05 |
| 599 | POSITIVE REGULATION OF NUCLEAR DIVISION | 7 | 62 | 4.763e-06 | 3.694e-05 |
| 600 | CELLULAR HOMEOSTASIS | 22 | 676 | 4.752e-06 | 3.694e-05 |
| 601 | CARTILAGE DEVELOPMENT | 10 | 147 | 4.788e-06 | 3.707e-05 |
| 602 | FEMALE SEX DIFFERENTIATION | 9 | 116 | 4.931e-06 | 3.811e-05 |
| 603 | POSITIVE REGULATION OF KIDNEY DEVELOPMENT | 6 | 41 | 4.986e-06 | 3.841e-05 |
| 604 | REGULATION OF CATABOLIC PROCESS | 23 | 731 | 4.984e-06 | 3.841e-05 |
| 605 | ION HOMEOSTASIS | 20 | 576 | 5.079e-06 | 3.906e-05 |
| 606 | RESPONSE TO TRANSITION METAL NANOPARTICLE | 10 | 148 | 5.086e-06 | 3.906e-05 |
| 607 | REGULATION OF COAGULATION | 8 | 88 | 5.153e-06 | 3.922e-05 |
| 608 | BLOOD VESSEL ENDOTHELIAL CELL MIGRATION | 5 | 24 | 5.159e-06 | 3.922e-05 |
| 609 | LEUKOCYTE PROLIFERATION | 8 | 88 | 5.153e-06 | 3.922e-05 |
| 610 | OVULATION CYCLE PROCESS | 8 | 88 | 5.153e-06 | 3.922e-05 |
| 611 | POSITIVE REGULATION OF CELL JUNCTION ASSEMBLY | 5 | 24 | 5.159e-06 | 3.922e-05 |
| 612 | NEGATIVE REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 8 | 88 | 5.153e-06 | 3.922e-05 |
| 613 | RESPONSE TO OSMOTIC STRESS | 7 | 63 | 5.31e-06 | 4.03e-05 |
| 614 | CELLULAR RESPONSE TO OXIDATIVE STRESS | 11 | 184 | 5.585e-06 | 4.232e-05 |
| 615 | B CELL DIFFERENTIATION | 8 | 89 | 5.609e-06 | 4.244e-05 |
| 616 | REGULATION OF PROTEIN STABILITY | 12 | 221 | 5.63e-06 | 4.253e-05 |
| 617 | POSITIVE REGULATION OF GLUCOSE TRANSPORT | 6 | 42 | 5.764e-06 | 4.347e-05 |
| 618 | MIDBRAIN DEVELOPMENT | 8 | 90 | 6.1e-06 | 4.593e-05 |
| 619 | POSITIVE REGULATION OF GLUCOSE IMPORT IN RESPONSE TO INSULIN STIMULUS | 4 | 12 | 6.233e-06 | 4.685e-05 |
| 620 | RESPONSE TO PROSTAGLANDIN E | 5 | 25 | 6.392e-06 | 4.782e-05 |
| 621 | REGULATION OF PEPTIDASE ACTIVITY | 16 | 392 | 6.38e-06 | 4.782e-05 |
| 622 | HISTONE PHOSPHORYLATION | 5 | 25 | 6.392e-06 | 4.782e-05 |
| 623 | CEREBRAL CORTEX CELL MIGRATION | 6 | 43 | 6.639e-06 | 4.958e-05 |
| 624 | REGULATION OF MULTICELLULAR ORGANISM GROWTH | 7 | 66 | 7.271e-06 | 5.422e-05 |
| 625 | SMALL GTPASE MEDIATED SIGNAL TRANSDUCTION | 15 | 352 | 7.492e-06 | 5.569e-05 |
| 626 | NEGATIVE REGULATION OF CELL PROLIFERATION | 21 | 643 | 7.489e-06 | 5.569e-05 |
| 627 | STEM CELL DIFFERENTIATION | 11 | 190 | 7.585e-06 | 5.629e-05 |
| 628 | NEGATIVE REGULATION OF LIPID BIOSYNTHETIC PROCESS | 6 | 44 | 7.618e-06 | 5.644e-05 |
| 629 | REPLACEMENT OSSIFICATION | 5 | 26 | 7.844e-06 | 5.784e-05 |
| 630 | NEGATIVE REGULATION OF LIPID TRANSPORT | 5 | 26 | 7.844e-06 | 5.784e-05 |
| 631 | ENDOCHONDRAL OSSIFICATION | 5 | 26 | 7.844e-06 | 5.784e-05 |
| 632 | EXOCYTOSIS | 14 | 310 | 7.89e-06 | 5.809e-05 |
| 633 | NEGATIVE REGULATION OF RESPONSE TO WOUNDING | 10 | 156 | 8.118e-06 | 5.967e-05 |
| 634 | REGULATION OF DNA BIOSYNTHETIC PROCESS | 8 | 94 | 8.441e-06 | 6.195e-05 |
| 635 | REGULATION OF ACTIN FILAMENT BASED PROCESS | 14 | 312 | 8.487e-06 | 6.219e-05 |
| 636 | ENDOCHONDRAL BONE MORPHOGENESIS | 6 | 45 | 8.711e-06 | 6.373e-05 |
| 637 | IMMUNE RESPONSE | 29 | 1100 | 8.786e-06 | 6.418e-05 |
| 638 | REGULATION OF CELL PROLIFERATION INVOLVED IN KIDNEY DEVELOPMENT | 4 | 13 | 8.926e-06 | 6.49e-05 |
| 639 | CELLULAR RESPONSE TO HEPATOCYTE GROWTH FACTOR STIMULUS | 4 | 13 | 8.926e-06 | 6.49e-05 |
| 640 | RESPONSE TO HEPATOCYTE GROWTH FACTOR | 4 | 13 | 8.926e-06 | 6.49e-05 |
| 641 | RESPONSE TO PURINE CONTAINING COMPOUND | 10 | 158 | 9.084e-06 | 6.594e-05 |
| 642 | MULTICELLULAR ORGANISMAL HOMEOSTASIS | 13 | 272 | 9.157e-06 | 6.636e-05 |
| 643 | EAR DEVELOPMENT | 11 | 195 | 9.702e-06 | 7.01e-05 |
| 644 | RESPONSE TO UV | 9 | 126 | 9.699e-06 | 7.01e-05 |
| 645 | REGULATION OF CYTOSKELETON ORGANIZATION | 18 | 502 | 9.961e-06 | 7.186e-05 |
| 646 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 19 | 554 | 1.051e-05 | 7.57e-05 |
| 647 | REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 8 | 97 | 1.066e-05 | 7.665e-05 |
| 648 | REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 11 | 197 | 1.068e-05 | 7.671e-05 |
| 649 | REGULATION OF LIPID BIOSYNTHETIC PROCESS | 9 | 128 | 1.102e-05 | 7.901e-05 |
| 650 | RESPONSE TO ANTIBIOTIC | 6 | 47 | 1.128e-05 | 8.061e-05 |
| 651 | POSITIVE REGULATION OF NEURON APOPTOTIC PROCESS | 6 | 47 | 1.128e-05 | 8.061e-05 |
| 652 | POSITIVE REGULATION OF RESPONSE TO WOUNDING | 10 | 162 | 1.132e-05 | 8.076e-05 |
| 653 | REGULATION OF SPROUTING ANGIOGENESIS | 5 | 28 | 1.151e-05 | 8.205e-05 |
| 654 | VESICLE MEDIATED TRANSPORT | 31 | 1239 | 1.182e-05 | 8.411e-05 |
| 655 | INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO DNA DAMAGE | 7 | 71 | 1.186e-05 | 8.412e-05 |
| 656 | SKIN EPIDERMIS DEVELOPMENT | 7 | 71 | 1.186e-05 | 8.412e-05 |
| 657 | ENDOCYTOSIS | 18 | 509 | 1.201e-05 | 8.503e-05 |
| 658 | POSITIVE REGULATION OF METANEPHROS DEVELOPMENT | 4 | 14 | 1.239e-05 | 8.696e-05 |
| 659 | POSITIVE REGULATION OF RHO PROTEIN SIGNAL TRANSDUCTION | 4 | 14 | 1.239e-05 | 8.696e-05 |
| 660 | POSITIVE REGULATION OF NITRIC OXIDE SYNTHASE BIOSYNTHETIC PROCESS | 4 | 14 | 1.239e-05 | 8.696e-05 |
| 661 | NEGATIVE REGULATION OF CATALYTIC ACTIVITY | 24 | 829 | 1.238e-05 | 8.696e-05 |
| 662 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 11 | 200 | 1.232e-05 | 8.696e-05 |
| 663 | POSITIVE REGULATION OF SPROUTING ANGIOGENESIS | 4 | 14 | 1.239e-05 | 8.696e-05 |
| 664 | SENSORY ORGAN MORPHOGENESIS | 12 | 239 | 1.244e-05 | 8.714e-05 |
| 665 | REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION TO PLASMA MEMBRANE | 6 | 48 | 1.277e-05 | 8.937e-05 |
| 666 | REGULATION OF ORGAN MORPHOGENESIS | 12 | 242 | 1.409e-05 | 9.843e-05 |
| 667 | POSITIVE REGULATION OF NF KAPPAB TRANSCRIPTION FACTOR ACTIVITY | 9 | 132 | 1.413e-05 | 9.857e-05 |
| 668 | REGULATION OF EXTENT OF CELL GROWTH | 8 | 101 | 1.436e-05 | 1e-04 |
| 669 | ANATOMICAL STRUCTURE HOMEOSTASIS | 13 | 285 | 1.505e-05 | 0.0001047 |
| 670 | MESENCHYMAL CELL DIFFERENTIATION | 9 | 134 | 1.595e-05 | 0.0001106 |
| 671 | MEMBRANE ORGANIZATION | 25 | 899 | 1.595e-05 | 0.0001106 |
| 672 | NEGATIVE REGULATION OF TOR SIGNALING | 5 | 30 | 1.64e-05 | 0.0001136 |
| 673 | POSITIVE REGULATION OF INNATE IMMUNE RESPONSE | 12 | 246 | 1.659e-05 | 0.0001145 |
| 674 | REGULATION OF MUSCLE ORGAN DEVELOPMENT | 8 | 103 | 1.659e-05 | 0.0001145 |
| 675 | NEGATIVE REGULATION OF DENDRITE MORPHOGENESIS | 4 | 15 | 1.675e-05 | 0.0001151 |
| 676 | JAK STAT CASCADE INVOLVED IN GROWTH HORMONE SIGNALING PATHWAY | 4 | 15 | 1.675e-05 | 0.0001151 |
| 677 | NEUROTROPHIN TRK RECEPTOR SIGNALING PATHWAY | 4 | 15 | 1.675e-05 | 0.0001151 |
| 678 | GRANULOCYTE MIGRATION | 7 | 75 | 1.706e-05 | 0.0001169 |
| 679 | ODONTOGENESIS OF DENTIN CONTAINING TOOTH | 7 | 75 | 1.706e-05 | 0.0001169 |
| 680 | DEVELOPMENTAL GROWTH INVOLVED IN MORPHOGENESIS | 8 | 104 | 1.78e-05 | 0.0001218 |
| 681 | RESPONSE TO NICOTINE | 6 | 51 | 1.824e-05 | 0.0001247 |
| 682 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 39 | 1784 | 1.838e-05 | 0.0001254 |
| 683 | NEGATIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 21 | 684 | 1.893e-05 | 0.0001289 |
| 684 | PROTEIN COMPLEX SUBUNIT ORGANIZATION | 35 | 1527 | 1.992e-05 | 0.0001355 |
| 685 | INNATE IMMUNE RESPONSE ACTIVATING CELL SURFACE RECEPTOR SIGNALING PATHWAY | 8 | 106 | 2.046e-05 | 0.0001388 |
| 686 | FAT CELL DIFFERENTIATION | 8 | 106 | 2.046e-05 | 0.0001388 |
| 687 | REGULATION OF T CELL DIFFERENTIATION | 8 | 107 | 2.191e-05 | 0.0001484 |
| 688 | POSITIVE REGULATION OF ACTIN CYTOSKELETON REORGANIZATION | 4 | 16 | 2.215e-05 | 0.0001494 |
| 689 | POSITIVE REGULATION OF TYROSINE PHOSPHORYLATION OF STAT5 PROTEIN | 4 | 16 | 2.215e-05 | 0.0001494 |
| 690 | REGULATION OF DEVELOPMENTAL PIGMENTATION | 4 | 16 | 2.215e-05 | 0.0001494 |
| 691 | REGULATION OF MORPHOGENESIS OF A BRANCHING STRUCTURE | 6 | 53 | 2.284e-05 | 0.0001536 |
| 692 | NEGATIVE REGULATION OF CELL SUBSTRATE ADHESION | 6 | 53 | 2.284e-05 | 0.0001536 |
| 693 | REGULATION OF CELL CYCLE ARREST | 8 | 108 | 2.345e-05 | 0.0001572 |
| 694 | REGULATION OF MYELOID LEUKOCYTE DIFFERENTIATION | 8 | 108 | 2.345e-05 | 0.0001572 |
| 695 | BONE MORPHOGENESIS | 7 | 79 | 2.402e-05 | 0.0001608 |
| 696 | DIVALENT INORGANIC CATION HOMEOSTASIS | 14 | 343 | 2.446e-05 | 0.0001635 |
| 697 | REGULATION OF OSSIFICATION | 10 | 178 | 2.563e-05 | 0.0001711 |
| 698 | CELLULAR RESPONSE TO OXYGEN LEVELS | 9 | 143 | 2.679e-05 | 0.0001786 |
| 699 | REGULATION OF MITOCHONDRION ORGANIZATION | 11 | 218 | 2.746e-05 | 0.0001825 |
| 700 | CELLULAR RESPONSE TO STEROID HORMONE STIMULUS | 11 | 218 | 2.746e-05 | 0.0001825 |
| 701 | REGULATION OF KIDNEY DEVELOPMENT | 6 | 55 | 2.833e-05 | 0.0001872 |
| 702 | POSITIVE REGULATION OF MYELOID CELL DIFFERENTIATION | 7 | 81 | 2.829e-05 | 0.0001872 |
| 703 | REGULATION OF RESPONSE TO CYTOKINE STIMULUS | 9 | 144 | 2.831e-05 | 0.0001872 |
| 704 | POSITIVE REGULATION OF LEUKOCYTE CHEMOTAXIS | 7 | 81 | 2.829e-05 | 0.0001872 |
| 705 | COLUMNAR CUBOIDAL EPITHELIAL CELL DIFFERENTIATION | 8 | 111 | 2.86e-05 | 0.0001888 |
| 706 | POSITIVE REGULATION OF NUCLEOTIDE CATABOLIC PROCESS | 4 | 17 | 2.872e-05 | 0.0001893 |
| 707 | RESPONSE TO IONIZING RADIATION | 9 | 145 | 2.99e-05 | 0.0001968 |
| 708 | REGULATION OF OSTEOBLAST DIFFERENTIATION | 8 | 112 | 3.052e-05 | 0.0002006 |
| 709 | NEGATIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 12 | 262 | 3.089e-05 | 0.0002027 |
| 710 | OVULATION CYCLE | 8 | 113 | 3.254e-05 | 0.000213 |
| 711 | REGULATION OF SYNAPSE ORGANIZATION | 8 | 113 | 3.254e-05 | 0.000213 |
| 712 | HAIR CYCLE | 7 | 83 | 3.317e-05 | 0.0002162 |
| 713 | MOLTING CYCLE | 7 | 83 | 3.317e-05 | 0.0002162 |
| 714 | RESPONSE TO FATTY ACID | 7 | 83 | 3.317e-05 | 0.0002162 |
| 715 | AGING | 12 | 264 | 3.327e-05 | 0.0002165 |
| 716 | REGULATION OF CELLULAR AMIDE METABOLIC PROCESS | 14 | 354 | 3.457e-05 | 0.0002246 |
| 717 | REGULATION OF EMBRYONIC DEVELOPMENT | 8 | 114 | 3.467e-05 | 0.000225 |
| 718 | RESPONSE TO TEMPERATURE STIMULUS | 9 | 148 | 3.514e-05 | 0.0002274 |
| 719 | MALE SEX DIFFERENTIATION | 9 | 148 | 3.514e-05 | 0.0002274 |
| 720 | REGULATION OF COLLATERAL SPROUTING | 4 | 18 | 3.661e-05 | 0.0002362 |
| 721 | RESPONSE TO PLATELET DERIVED GROWTH FACTOR | 4 | 18 | 3.661e-05 | 0.0002362 |
| 722 | CELLULAR RESPONSE TO ALCOHOL | 8 | 115 | 3.692e-05 | 0.000238 |
| 723 | PROTEIN COMPLEX BIOGENESIS | 28 | 1132 | 3.999e-05 | 0.000257 |
| 724 | PROTEIN COMPLEX ASSEMBLY | 28 | 1132 | 3.999e-05 | 0.000257 |
| 725 | REGULATION OF CELL CYCLE PROCESS | 18 | 558 | 4.042e-05 | 0.0002594 |
| 726 | NEUROMUSCULAR JUNCTION DEVELOPMENT | 5 | 36 | 4.116e-05 | 0.0002631 |
| 727 | REGULATION OF NUCLEOTIDE CATABOLIC PROCESS | 5 | 36 | 4.116e-05 | 0.0002631 |
| 728 | POSITIVE REGULATION OF GLUCOSE METABOLIC PROCESS | 5 | 36 | 4.116e-05 | 0.0002631 |
| 729 | POSITIVE REGULATION OF DNA BIOSYNTHETIC PROCESS | 6 | 59 | 4.247e-05 | 0.0002707 |
| 730 | VASCULOGENESIS | 6 | 59 | 4.247e-05 | 0.0002707 |
| 731 | TISSUE REMODELING | 7 | 87 | 4.501e-05 | 0.0002864 |
| 732 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 19 | 616 | 4.506e-05 | 0.0002864 |
| 733 | REGULATION OF NITRIC OXIDE SYNTHASE BIOSYNTHETIC PROCESS | 4 | 19 | 4.598e-05 | 0.0002911 |
| 734 | POSITIVE REGULATION OF CARDIAC MUSCLE CELL PROLIFERATION | 4 | 19 | 4.598e-05 | 0.0002911 |
| 735 | ASTROCYTE DEVELOPMENT | 4 | 19 | 4.598e-05 | 0.0002911 |
| 736 | CHONDROCYTE DIFFERENTIATION | 6 | 60 | 4.676e-05 | 0.0002956 |
| 737 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO CELL PERIPHERY | 5 | 37 | 4.717e-05 | 0.000297 |
| 738 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO PLASMA MEMBRANE | 5 | 37 | 4.717e-05 | 0.000297 |
| 739 | REGULATION OF PROTEIN AUTOPHOSPHORYLATION | 5 | 37 | 4.717e-05 | 0.000297 |
| 740 | REGULATION OF VESICLE MEDIATED TRANSPORT | 16 | 462 | 4.758e-05 | 0.0002992 |
| 741 | REGULATION OF STEM CELL PROLIFERATION | 7 | 88 | 4.845e-05 | 0.0003042 |
| 742 | REGULATION OF DENDRITE DEVELOPMENT | 8 | 120 | 5.008e-05 | 0.0003141 |
| 743 | RESPONSE TO TUMOR NECROSIS FACTOR | 11 | 233 | 5.04e-05 | 0.0003156 |
| 744 | RESPONSE TO HEAT | 7 | 89 | 5.211e-05 | 0.0003259 |
| 745 | PROTEIN LOCALIZATION TO NUCLEUS | 9 | 156 | 5.307e-05 | 0.0003314 |
| 746 | POSITIVE REGULATION OF ORGAN GROWTH | 5 | 38 | 5.384e-05 | 0.0003358 |
| 747 | PROTEIN LOCALIZATION | 38 | 1805 | 5.443e-05 | 0.000339 |
| 748 | REGULATION OF MUSCLE SYSTEM PROCESS | 10 | 195 | 5.566e-05 | 0.0003462 |
| 749 | MESONEPHROS DEVELOPMENT | 7 | 90 | 5.599e-05 | 0.0003478 |
| 750 | REGULATION OF TISSUE REMODELING | 6 | 62 | 5.64e-05 | 0.0003499 |
| 751 | POSITIVE REGULATION OF ION TRANSPORT | 11 | 236 | 5.657e-05 | 0.0003501 |
| 752 | NEGATIVE REGULATION OF GROWTH | 11 | 236 | 5.657e-05 | 0.0003501 |
| 753 | EMBRYONIC HEMOPOIESIS | 4 | 20 | 5.698e-05 | 0.0003512 |
| 754 | AMELOGENESIS | 4 | 20 | 5.698e-05 | 0.0003512 |
| 755 | REGULATION OF TYROSINE PHOSPHORYLATION OF STAT5 PROTEIN | 4 | 20 | 5.698e-05 | 0.0003512 |
| 756 | ENDOCRINE SYSTEM DEVELOPMENT | 8 | 123 | 5.97e-05 | 0.0003675 |
| 757 | REGULATION OF CELL ADHESION MEDIATED BY INTEGRIN | 5 | 39 | 6.122e-05 | 0.0003753 |
| 758 | CELLULAR RESPONSE TO NUTRIENT | 5 | 39 | 6.122e-05 | 0.0003753 |
| 759 | LONG TERM SYNAPTIC POTENTIATION | 5 | 39 | 6.122e-05 | 0.0003753 |
| 760 | REGULATION OF TRANSPORTER ACTIVITY | 10 | 198 | 6.327e-05 | 0.0003874 |
| 761 | REGULATION OF ENDOCYTOSIS | 10 | 199 | 6.6e-05 | 0.0004035 |
| 762 | ESTABLISHMENT OF PROTEIN LOCALIZATION | 32 | 1423 | 6.804e-05 | 0.0004149 |
| 763 | RESPONSE TO BACTERIUM | 17 | 528 | 6.802e-05 | 0.0004149 |
| 764 | REGULATION OF TRANSMEMBRANE TRANSPORT | 15 | 426 | 6.835e-05 | 0.0004163 |
| 765 | OSTEOBLAST DIFFERENTIATION | 8 | 126 | 7.082e-05 | 0.0004307 |
| 766 | NEGATIVE REGULATION OF AXONOGENESIS | 6 | 65 | 7.376e-05 | 0.000448 |
| 767 | CELLULAR RESPONSE TO BIOTIC STIMULUS | 9 | 163 | 7.456e-05 | 0.0004523 |
| 768 | SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 8 | 127 | 7.488e-05 | 0.0004537 |
| 769 | RESPONSE TO METAL ION | 13 | 333 | 7.521e-05 | 0.0004551 |
| 770 | REGULATION OF CYTOSOLIC CALCIUM ION CONCENTRATION | 10 | 203 | 7.792e-05 | 0.0004709 |
| 771 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 7 | 95 | 7.911e-05 | 0.0004774 |
| 772 | NEURAL NUCLEUS DEVELOPMENT | 6 | 66 | 8.041e-05 | 0.0004846 |
| 773 | MYELOID LEUKOCYTE DIFFERENTIATION | 7 | 96 | 8.455e-05 | 0.0005059 |
| 774 | SIGNAL TRANSDUCTION IN RESPONSE TO DNA DAMAGE | 7 | 96 | 8.455e-05 | 0.0005059 |
| 775 | REGULATION OF LEUKOCYTE CHEMOTAXIS | 7 | 96 | 8.455e-05 | 0.0005059 |
| 776 | SOMATIC STEM CELL DIVISION | 4 | 22 | 8.458e-05 | 0.0005059 |
| 777 | ACTIVATION OF PROTEIN KINASE B ACTIVITY | 4 | 22 | 8.458e-05 | 0.0005059 |
| 778 | ERK1 AND ERK2 CASCADE | 4 | 22 | 8.458e-05 | 0.0005059 |
| 779 | REGULATION OF ENDOTHELIAL CELL APOPTOTIC PROCESS | 5 | 42 | 8.809e-05 | 0.0005248 |
| 780 | REGULATION OF CELLULAR CARBOHYDRATE CATABOLIC PROCESS | 5 | 42 | 8.809e-05 | 0.0005248 |
| 781 | REGULATION OF CARBOHYDRATE CATABOLIC PROCESS | 5 | 42 | 8.809e-05 | 0.0005248 |
| 782 | POSITIVE REGULATION OF MITOCHONDRION ORGANIZATION | 9 | 167 | 8.984e-05 | 0.0005346 |
| 783 | LEARNING | 8 | 131 | 9.313e-05 | 0.0005527 |
| 784 | PROTEIN STABILIZATION | 8 | 131 | 9.313e-05 | 0.0005527 |
| 785 | REGULATION OF INFLAMMATORY RESPONSE | 12 | 294 | 9.359e-05 | 0.0005547 |
| 786 | RESPONSE TO CARBOHYDRATE | 9 | 168 | 9.405e-05 | 0.0005567 |
| 787 | ORGAN GROWTH | 6 | 68 | 9.513e-05 | 0.0005625 |
| 788 | NEGATIVE REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 7 | 98 | 9.636e-05 | 0.000569 |
| 789 | REGULATION OF LYMPHOCYTE DIFFERENTIATION | 8 | 132 | 9.823e-05 | 0.0005793 |
| 790 | RESPONSE TO ELECTRICAL STIMULUS | 5 | 43 | 9.881e-05 | 0.0005812 |
| 791 | MYELOID LEUKOCYTE MEDIATED IMMUNITY | 5 | 43 | 9.881e-05 | 0.0005812 |
| 792 | REGULATION OF CALCIUM ION TRANSPORT | 10 | 209 | 9.922e-05 | 0.0005829 |
| 793 | REGULATION OF PROTEOLYSIS | 20 | 711 | 0.0001007 | 0.0005909 |
| 794 | RESPONSE TO INCREASED OXYGEN LEVELS | 4 | 23 | 0.0001015 | 0.0005927 |
| 795 | LEUKOCYTE APOPTOTIC PROCESS | 4 | 23 | 0.0001015 | 0.0005927 |
| 796 | REGULATION OF METANEPHROS DEVELOPMENT | 4 | 23 | 0.0001015 | 0.0005927 |
| 797 | RESPONSE TO HYPEROXIA | 4 | 23 | 0.0001015 | 0.0005927 |
| 798 | NEGATIVE REGULATION OF CYTOKINE PRODUCTION | 10 | 211 | 0.0001073 | 0.0006258 |
| 799 | BRANCHING INVOLVED IN URETERIC BUD MORPHOGENESIS | 5 | 44 | 0.0001105 | 0.0006435 |
| 800 | POSITIVE REGULATION OF CATABOLIC PROCESS | 14 | 395 | 0.0001116 | 0.0006483 |
| 801 | HEART MORPHOGENESIS | 10 | 212 | 0.0001116 | 0.0006483 |
| 802 | CENTRAL NERVOUS SYSTEM NEURON DEVELOPMENT | 6 | 70 | 0.0001119 | 0.0006494 |
| 803 | STRIATED MUSCLE CELL DIFFERENTIATION | 9 | 173 | 0.0001176 | 0.0006816 |
| 804 | POSTTRANSCRIPTIONAL REGULATION OF GENE EXPRESSION | 15 | 448 | 0.0001195 | 0.0006914 |
| 805 | FOCAL ADHESION ASSEMBLY | 4 | 24 | 0.0001208 | 0.0006947 |
| 806 | POSITIVE REGULATION OF NUCLEOSIDE METABOLIC PROCESS | 4 | 24 | 0.0001208 | 0.0006947 |
| 807 | POSITIVE REGULATION OF RECEPTOR INTERNALIZATION | 4 | 24 | 0.0001208 | 0.0006947 |
| 808 | POSITIVE REGULATION OF ATP METABOLIC PROCESS | 4 | 24 | 0.0001208 | 0.0006947 |
| 809 | CELL SUBSTRATE ADHERENS JUNCTION ASSEMBLY | 4 | 24 | 0.0001208 | 0.0006947 |
| 810 | LUNG MORPHOGENESIS | 5 | 45 | 0.0001232 | 0.000707 |
| 811 | SUBSTANTIA NIGRA DEVELOPMENT | 5 | 45 | 0.0001232 | 0.000707 |
| 812 | ACTIN FILAMENT BASED PROCESS | 15 | 450 | 0.0001254 | 0.0007188 |
| 813 | CELLULAR RESPONSE TO RADIATION | 8 | 137 | 0.0001273 | 0.0007286 |
| 814 | REGULATION OF MUSCLE TISSUE DEVELOPMENT | 7 | 103 | 0.0001318 | 0.0007537 |
| 815 | PEPTIDYL THREONINE MODIFICATION | 5 | 46 | 0.000137 | 0.0007814 |
| 816 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 5 | 46 | 0.000137 | 0.0007814 |
| 817 | CELLULAR RESPONSE TO REACTIVE OXYGEN SPECIES | 7 | 104 | 0.0001401 | 0.0007978 |
| 818 | REGULATION OF CELL SHAPE | 8 | 139 | 0.0001408 | 0.0008008 |
| 819 | ASSOCIATIVE LEARNING | 6 | 73 | 0.0001415 | 0.0008017 |
| 820 | REGULATION OF PLASMA MEMBRANE ORGANIZATION | 6 | 73 | 0.0001415 | 0.0008017 |
| 821 | CELLULAR RESPONSE TO KETONE | 6 | 73 | 0.0001415 | 0.0008017 |
| 822 | DETECTION OF MECHANICAL STIMULUS INVOLVED IN SENSORY PERCEPTION | 4 | 25 | 0.0001426 | 0.0008071 |
| 823 | TYROSINE PHOSPHORYLATION OF STAT PROTEIN | 3 | 10 | 0.0001429 | 0.0008079 |
| 824 | POSITIVE REGULATION OF DEPHOSPHORYLATION | 5 | 47 | 0.000152 | 0.0008584 |
| 825 | REGULATION OF DENDRITE MORPHOGENESIS | 6 | 74 | 0.0001526 | 0.0008604 |
| 826 | REGULATION OF WNT SIGNALING PATHWAY | 12 | 310 | 0.0001538 | 0.0008663 |
| 827 | REGULATION OF GLUCOSE METABOLIC PROCESS | 7 | 106 | 0.0001578 | 0.0008878 |
| 828 | CELL DIVISION | 15 | 460 | 0.0001594 | 0.000896 |
| 829 | BIOMINERAL TISSUE DEVELOPMENT | 6 | 75 | 0.0001643 | 0.0009224 |
| 830 | CELLULAR RESPONSE TO VITAMIN | 4 | 26 | 0.0001671 | 0.0009367 |
| 831 | COLUMNAR CUBOIDAL EPITHELIAL CELL DEVELOPMENT | 5 | 48 | 0.0001682 | 0.0009418 |
| 832 | REGULATION OF CELLULAR RESPONSE TO HEAT | 6 | 76 | 0.0001768 | 0.0009877 |
| 833 | EXTRACELLULAR MATRIX DISASSEMBLY | 6 | 76 | 0.0001768 | 0.0009877 |
| 834 | REGULATION OF NUCLEOSIDE METABOLIC PROCESS | 5 | 49 | 0.0001857 | 0.001033 |
| 835 | POSITIVE REGULATION OF CHEMOKINE PRODUCTION | 5 | 49 | 0.0001857 | 0.001033 |
| 836 | REGULATION OF ATP METABOLIC PROCESS | 5 | 49 | 0.0001857 | 0.001033 |
| 837 | REGULATION OF RHO PROTEIN SIGNAL TRANSDUCTION | 7 | 109 | 0.0001877 | 0.001044 |
| 838 | NEUROLOGICAL SYSTEM PROCESS | 28 | 1242 | 0.0001941 | 0.001078 |
| 839 | POSITIVE REGULATION OF HEART GROWTH | 4 | 27 | 0.0001945 | 0.001079 |
| 840 | LYMPHOID PROGENITOR CELL DIFFERENTIATION | 3 | 11 | 0.0001949 | 0.00108 |
| 841 | EPITHELIAL CELL DEVELOPMENT | 9 | 186 | 0.0002031 | 0.001122 |
| 842 | REGULATION OF SYNAPTIC TRANSMISSION GLUTAMATERGIC | 5 | 50 | 0.0002045 | 0.001122 |
| 843 | LYMPHOCYTE HOMEOSTASIS | 5 | 50 | 0.0002045 | 0.001122 |
| 844 | REGULATION OF COENZYME METABOLIC PROCESS | 5 | 50 | 0.0002045 | 0.001122 |
| 845 | REGULATION OF RECEPTOR MEDIATED ENDOCYTOSIS | 6 | 78 | 0.0002041 | 0.001122 |
| 846 | RENAL TUBULE DEVELOPMENT | 6 | 78 | 0.0002041 | 0.001122 |
| 847 | REGULATION OF COFACTOR METABOLIC PROCESS | 5 | 50 | 0.0002045 | 0.001122 |
| 848 | POSITIVE REGULATION OF MYELOID LEUKOCYTE DIFFERENTIATION | 5 | 50 | 0.0002045 | 0.001122 |
| 849 | REGULATION OF CELL CYCLE G1 S PHASE TRANSITION | 8 | 147 | 0.0002069 | 0.001134 |
| 850 | POSITIVE REGULATION OF CELL GROWTH | 8 | 148 | 0.0002167 | 0.001186 |
| 851 | REGULATION OF SYNAPSE ASSEMBLY | 6 | 79 | 0.0002189 | 0.001197 |
| 852 | EAR MORPHOGENESIS | 7 | 112 | 0.0002221 | 0.001213 |
| 853 | REGULATION OF MACROPHAGE DERIVED FOAM CELL DIFFERENTIATION | 4 | 28 | 0.000225 | 0.001219 |
| 854 | NEGATIVE REGULATION OF DENDRITE DEVELOPMENT | 4 | 28 | 0.000225 | 0.001219 |
| 855 | POSITIVE REGULATION OF ACTIVATED T CELL PROLIFERATION | 4 | 28 | 0.000225 | 0.001219 |
| 856 | NEGATIVE REGULATION OF IMMUNE SYSTEM PROCESS | 13 | 372 | 0.0002249 | 0.001219 |
| 857 | MAMMARY GLAND DUCT MORPHOGENESIS | 4 | 28 | 0.000225 | 0.001219 |
| 858 | POSITIVE REGULATION OF PHOSPHATASE ACTIVITY | 4 | 28 | 0.000225 | 0.001219 |
| 859 | REGULATION OF INTERLEUKIN 12 PRODUCTION | 5 | 51 | 0.0002247 | 0.001219 |
| 860 | REGULATION OF CYTOKINE SECRETION | 8 | 149 | 0.0002269 | 0.001228 |
| 861 | POSITIVE REGULATION OF LYMPHOCYTE DIFFERENTIATION | 6 | 80 | 0.0002345 | 0.001265 |
| 862 | POSITIVE REGULATION OF INFLAMMATORY RESPONSE | 7 | 113 | 0.0002346 | 0.001265 |
| 863 | PROTEIN HETEROOLIGOMERIZATION | 7 | 113 | 0.0002346 | 0.001265 |
| 864 | RESPONSE TO ENDOPLASMIC RETICULUM STRESS | 10 | 233 | 0.0002404 | 0.001295 |
| 865 | ACTIVATION OF MAPKK ACTIVITY | 5 | 52 | 0.0002465 | 0.001326 |
| 866 | POSITIVE REGULATION OF ENDOCYTOSIS | 7 | 114 | 0.0002477 | 0.001331 |
| 867 | REGULATION OF CARDIAC MUSCLE CELL PROLIFERATION | 4 | 29 | 0.0002588 | 0.001379 |
| 868 | STEM CELL DIVISION | 4 | 29 | 0.0002588 | 0.001379 |
| 869 | RESPONSE TO PAIN | 4 | 29 | 0.0002588 | 0.001379 |
| 870 | POSITIVE REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 4 | 29 | 0.0002588 | 0.001379 |
| 871 | I KAPPAB PHOSPHORYLATION | 3 | 12 | 0.0002578 | 0.001379 |
| 872 | MAMMARY GLAND EPITHELIAL CELL PROLIFERATION | 3 | 12 | 0.0002578 | 0.001379 |
| 873 | ACTIVATION OF TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE ACTIVITY | 3 | 12 | 0.0002578 | 0.001379 |
| 874 | KIDNEY MORPHOGENESIS | 6 | 82 | 0.0002684 | 0.001429 |
| 875 | MESONEPHRIC TUBULE MORPHOGENESIS | 5 | 53 | 0.0002698 | 0.001435 |
| 876 | MUSCLE CELL DIFFERENTIATION | 10 | 237 | 0.0002755 | 0.001463 |
| 877 | CELL CYCLE CHECKPOINT | 9 | 194 | 0.0002777 | 0.001473 |
| 878 | POSITIVE REGULATION OF PEPTIDASE ACTIVITY | 8 | 154 | 0.000284 | 0.001505 |
| 879 | ESTABLISHMENT OF LOCALIZATION IN CELL | 34 | 1676 | 0.0002852 | 0.001509 |
| 880 | REGULATION OF ERBB SIGNALING PATHWAY | 6 | 83 | 0.0002867 | 0.001514 |
| 881 | EMBRYONIC PLACENTA DEVELOPMENT | 6 | 83 | 0.0002867 | 0.001514 |
| 882 | SENSORY PERCEPTION OF MECHANICAL STIMULUS | 8 | 155 | 0.0002967 | 0.00156 |
| 883 | PROTEIN IMPORT | 8 | 155 | 0.0002967 | 0.00156 |
| 884 | RESPONSE TO EPIDERMAL GROWTH FACTOR | 4 | 30 | 0.000296 | 0.00156 |
| 885 | NEGATIVE REGULATION OF MUSCLE CELL APOPTOTIC PROCESS | 4 | 30 | 0.000296 | 0.00156 |
| 886 | NEGATIVE REGULATION OF DEVELOPMENTAL GROWTH | 6 | 84 | 0.000306 | 0.001607 |
| 887 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY VIA DEATH DOMAIN RECEPTORS | 5 | 55 | 0.0003213 | 0.001686 |
| 888 | SINGLE ORGANISM CELLULAR LOCALIZATION | 22 | 898 | 0.0003218 | 0.001686 |
| 889 | TOLL LIKE RECEPTOR SIGNALING PATHWAY | 6 | 85 | 0.0003263 | 0.001708 |
| 890 | NEURONAL STEM CELL DIVISION | 3 | 13 | 0.0003325 | 0.001729 |
| 891 | BEHAVIORAL RESPONSE TO PAIN | 3 | 13 | 0.0003325 | 0.001729 |
| 892 | LEUKOCYTE TETHERING OR ROLLING | 3 | 13 | 0.0003325 | 0.001729 |
| 893 | LEUKOCYTE ADHESION TO VASCULAR ENDOTHELIAL CELL | 3 | 13 | 0.0003325 | 0.001729 |
| 894 | NEUROBLAST DIVISION | 3 | 13 | 0.0003325 | 0.001729 |
| 895 | REGULATION OF TRANSMISSION OF NERVE IMPULSE | 3 | 13 | 0.0003325 | 0.001729 |
| 896 | NEGATIVE REGULATION OF MITOTIC CELL CYCLE | 9 | 199 | 0.0003349 | 0.001739 |
| 897 | NEGATIVE REGULATION OF CELLULAR RESPONSE TO INSULIN STIMULUS | 4 | 31 | 0.000337 | 0.001748 |
| 898 | REGULATION OF PROTEIN SECRETION | 13 | 389 | 0.0003455 | 0.00179 |
| 899 | POSITIVE REGULATION OF B CELL ACTIVATION | 6 | 86 | 0.0003476 | 0.001799 |
| 900 | NEGATIVE REGULATION OF PEPTIDASE ACTIVITY | 10 | 245 | 0.0003586 | 0.001854 |
| 901 | SKELETAL SYSTEM MORPHOGENESIS | 9 | 201 | 0.0003604 | 0.001861 |
| 902 | POSITIVE REGULATION OF TUMOR NECROSIS FACTOR SUPERFAMILY CYTOKINE PRODUCTION | 5 | 57 | 0.00038 | 0.00196 |
| 903 | POSITIVE REGULATION OF MULTICELLULAR ORGANISM GROWTH | 4 | 32 | 0.0003819 | 0.001968 |
| 904 | NEGATIVE REGULATION OF CATABOLIC PROCESS | 9 | 203 | 0.0003875 | 0.001994 |
| 905 | EPITHELIAL CELL CELL ADHESION | 3 | 14 | 0.0004198 | 0.002144 |
| 906 | PROTEIN HETEROTRIMERIZATION | 3 | 14 | 0.0004198 | 0.002144 |
| 907 | REGULATION OF FIBROBLAST APOPTOTIC PROCESS | 3 | 14 | 0.0004198 | 0.002144 |
| 908 | REGULATION OF GLOMERULUS DEVELOPMENT | 3 | 14 | 0.0004198 | 0.002144 |
| 909 | T CELL MIGRATION | 3 | 14 | 0.0004198 | 0.002144 |
| 910 | POSITIVE REGULATION OF CELL SIZE | 3 | 14 | 0.0004198 | 0.002144 |
| 911 | BONE MATURATION | 3 | 14 | 0.0004198 | 0.002144 |
| 912 | NEGATIVE REGULATION OF KINASE ACTIVITY | 10 | 250 | 0.0004204 | 0.002145 |
| 913 | RESPONSE TO VITAMIN D | 4 | 33 | 0.0004309 | 0.002194 |
| 914 | REGULATION OF BONE RESORPTION | 4 | 33 | 0.0004309 | 0.002194 |
| 915 | KIDNEY EPITHELIUM DEVELOPMENT | 7 | 125 | 0.0004342 | 0.002208 |
| 916 | CELLULAR RESPONSE TO LIGHT STIMULUS | 6 | 91 | 0.0004713 | 0.002389 |
| 917 | ENSHEATHMENT OF NEURONS | 6 | 91 | 0.0004713 | 0.002389 |
| 918 | AXON ENSHEATHMENT | 6 | 91 | 0.0004713 | 0.002389 |
| 919 | WNT SIGNALING PATHWAY | 12 | 351 | 0.0004757 | 0.002408 |
| 920 | REGULATION OF MONOOXYGENASE ACTIVITY | 5 | 60 | 0.0004828 | 0.002439 |
| 921 | MATERNAL PROCESS INVOLVED IN FEMALE PREGNANCY | 5 | 60 | 0.0004828 | 0.002439 |
| 922 | T CELL HOMEOSTASIS | 4 | 34 | 0.0004843 | 0.002441 |
| 923 | ADHERENS JUNCTION ASSEMBLY | 4 | 34 | 0.0004843 | 0.002441 |
| 924 | INNER EAR MORPHOGENESIS | 6 | 92 | 0.0004996 | 0.002516 |
| 925 | REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION TO MITOCHONDRION | 7 | 128 | 0.0005008 | 0.002519 |
| 926 | REGULATION OF PROTEIN BINDING | 8 | 168 | 0.0005086 | 0.002553 |
| 927 | INTRACELLULAR RECEPTOR SIGNALING PATHWAY | 8 | 168 | 0.0005086 | 0.002553 |
| 928 | REGULATION OF NUCLEOTIDE METABOLIC PROCESS | 9 | 211 | 0.0005129 | 0.002572 |
| 929 | POSITIVE REGULATION OF SYNAPSE ASSEMBLY | 5 | 61 | 0.0005214 | 0.002597 |
| 930 | CELLULAR RESPONSE TO HYDROGEN PEROXIDE | 5 | 61 | 0.0005214 | 0.002597 |
| 931 | NEGATIVE REGULATION OF INTERLEUKIN 12 PRODUCTION | 3 | 15 | 0.0005205 | 0.002597 |
| 932 | RESPONSE TO VITAMIN E | 3 | 15 | 0.0005205 | 0.002597 |
| 933 | T CELL LINEAGE COMMITMENT | 3 | 15 | 0.0005205 | 0.002597 |
| 934 | T CELL APOPTOTIC PROCESS | 3 | 15 | 0.0005205 | 0.002597 |
| 935 | NUCLEAR IMPORT | 7 | 129 | 0.0005248 | 0.002612 |
| 936 | NEGATIVE REGULATION OF EPITHELIAL CELL APOPTOTIC PROCESS | 4 | 35 | 0.0005421 | 0.002686 |
| 937 | BONE REMODELING | 4 | 35 | 0.0005421 | 0.002686 |
| 938 | RESPONSE TO MINERALOCORTICOID | 4 | 35 | 0.0005421 | 0.002686 |
| 939 | RESPONSE TO IRON ION | 4 | 35 | 0.0005421 | 0.002686 |
| 940 | MULTI MULTICELLULAR ORGANISM PROCESS | 9 | 213 | 0.000549 | 0.002718 |
| 941 | NEGATIVE REGULATION OF CELL GROWTH | 8 | 170 | 0.0005501 | 0.00272 |
| 942 | NEGATIVE REGULATION OF CELL CYCLE PROCESS | 9 | 214 | 0.0005678 | 0.002805 |
| 943 | POSITIVE REGULATION OF T CELL PROLIFERATION | 6 | 95 | 0.0005928 | 0.002919 |
| 944 | ACTIVATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 6 | 95 | 0.0005928 | 0.002919 |
| 945 | REGULATION OF LIPID TRANSPORT | 6 | 95 | 0.0005928 | 0.002919 |
| 946 | REGULATION OF OSTEOCLAST DIFFERENTIATION | 5 | 63 | 0.0006053 | 0.002974 |
| 947 | T CELL SELECTION | 4 | 36 | 0.0006048 | 0.002974 |
| 948 | IN UTERO EMBRYONIC DEVELOPMENT | 11 | 311 | 0.0006162 | 0.003024 |
| 949 | CARDIOCYTE DIFFERENTIATION | 6 | 96 | 0.0006267 | 0.003073 |
| 950 | REGULATION OF GENE EXPRESSION BY GENETIC IMPRINTING | 3 | 16 | 0.0006356 | 0.003097 |
| 951 | POSITIVE REGULATION OF PHOSPHOPROTEIN PHOSPHATASE ACTIVITY | 3 | 16 | 0.0006356 | 0.003097 |
| 952 | POSITIVE REGULATION OF TRANSFORMING GROWTH FACTOR BETA PRODUCTION | 3 | 16 | 0.0006356 | 0.003097 |
| 953 | TOR SIGNALING | 3 | 16 | 0.0006356 | 0.003097 |
| 954 | RETINA VASCULATURE DEVELOPMENT IN CAMERA TYPE EYE | 3 | 16 | 0.0006356 | 0.003097 |
| 955 | RESPONSE TO UV B | 3 | 16 | 0.0006356 | 0.003097 |
| 956 | POSITIVE REGULATION OF PROTEOLYSIS | 12 | 363 | 0.0006399 | 0.003115 |
| 957 | SINGLE ORGANISM BIOSYNTHETIC PROCESS | 28 | 1340 | 0.0006538 | 0.003179 |
| 958 | POSITIVE REGULATION OF B CELL PROLIFERATION | 4 | 37 | 0.0006724 | 0.003255 |
| 959 | RESPONSE TO NERVE GROWTH FACTOR | 4 | 37 | 0.0006724 | 0.003255 |
| 960 | POSITIVE REGULATION OF PROTEIN TYROSINE KINASE ACTIVITY | 4 | 37 | 0.0006724 | 0.003255 |
| 961 | REGULATION OF RECEPTOR INTERNALIZATION | 4 | 37 | 0.0006724 | 0.003255 |
| 962 | CIRCULATORY SYSTEM PROCESS | 12 | 366 | 0.0006877 | 0.003326 |
| 963 | MYELOID LEUKOCYTE ACTIVATION | 6 | 98 | 0.000699 | 0.003374 |
| 964 | REGULATION OF CHEMOKINE PRODUCTION | 5 | 65 | 0.000699 | 0.003374 |
| 965 | COLLAGEN FIBRIL ORGANIZATION | 4 | 38 | 0.0007451 | 0.003589 |
| 966 | BONE MINERALIZATION | 4 | 38 | 0.0007451 | 0.003589 |
| 967 | CELLULAR RESPONSE TO UV | 5 | 66 | 0.0007497 | 0.003607 |
| 968 | CIRCADIAN RHYTHM | 7 | 137 | 0.0007509 | 0.003609 |
| 969 | BRANCH ELONGATION OF AN EPITHELIUM | 3 | 17 | 0.0007656 | 0.003665 |
| 970 | NEGATIVE REGULATION OF PLATELET ACTIVATION | 3 | 17 | 0.0007656 | 0.003665 |
| 971 | REGULATION OF DNA METHYLATION | 3 | 17 | 0.0007656 | 0.003665 |
| 972 | NEGATIVE REGULATION OF LIPID STORAGE | 3 | 17 | 0.0007656 | 0.003665 |
| 973 | REGULATION OF IMMUNE EFFECTOR PROCESS | 13 | 424 | 0.0007736 | 0.0037 |
| 974 | MITOTIC DNA INTEGRITY CHECKPOINT | 6 | 100 | 0.0007775 | 0.003714 |
| 975 | PROTEIN FOLDING | 9 | 224 | 0.0007867 | 0.003754 |
| 976 | RESPONSE TO ORGANOPHOSPHORUS | 7 | 139 | 0.0008179 | 0.003891 |
| 977 | CELL ACTIVATION INVOLVED IN IMMUNE RESPONSE | 7 | 139 | 0.0008179 | 0.003891 |
| 978 | MITOTIC CELL CYCLE CHECKPOINT | 7 | 139 | 0.0008179 | 0.003891 |
| 979 | REGULATION OF TUMOR NECROSIS FACTOR SUPERFAMILY CYTOKINE PRODUCTION | 6 | 101 | 0.0008193 | 0.003894 |
| 980 | TRABECULA MORPHOGENESIS | 4 | 39 | 0.0008233 | 0.003897 |
| 981 | REGULATION OF GRANULOCYTE CHEMOTAXIS | 4 | 39 | 0.0008233 | 0.003897 |
| 982 | PLATELET AGGREGATION | 4 | 39 | 0.0008233 | 0.003897 |
| 983 | NEGATIVE CHEMOTAXIS | 4 | 39 | 0.0008233 | 0.003897 |
| 984 | CARDIAC MUSCLE TISSUE DEVELOPMENT | 7 | 140 | 0.0008532 | 0.004034 |
| 985 | POSITIVE REGULATION OF INTERLEUKIN 6 PRODUCTION | 5 | 68 | 0.0008592 | 0.004059 |
| 986 | MUSCLE TISSUE DEVELOPMENT | 10 | 275 | 0.0008781 | 0.004144 |
| 987 | REGULATION OF MITOCHONDRIAL DEPOLARIZATION | 3 | 18 | 0.0009114 | 0.004236 |
| 988 | REGULATION OF CELL MATURATION | 3 | 18 | 0.0009114 | 0.004236 |
| 989 | MAMMARY GLAND MORPHOGENESIS | 4 | 40 | 0.0009071 | 0.004236 |
| 990 | INTRACELLULAR PROTEIN TRANSPORT | 19 | 781 | 0.0008999 | 0.004236 |
| 991 | NEGATIVE REGULATION OF RESPONSE TO REACTIVE OXYGEN SPECIES | 3 | 18 | 0.0009114 | 0.004236 |
| 992 | REGULATION OF ACTIVATED T CELL PROLIFERATION | 4 | 40 | 0.0009071 | 0.004236 |
| 993 | ORGAN MATURATION | 3 | 18 | 0.0009114 | 0.004236 |
| 994 | ENDOCRINE PANCREAS DEVELOPMENT | 4 | 40 | 0.0009071 | 0.004236 |
| 995 | RESPONSE TO CAFFEINE | 3 | 18 | 0.0009114 | 0.004236 |
| 996 | MAST CELL MEDIATED IMMUNITY | 3 | 18 | 0.0009114 | 0.004236 |
| 997 | NOTOCHORD DEVELOPMENT | 3 | 18 | 0.0009114 | 0.004236 |
| 998 | STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 6 | 103 | 0.0009079 | 0.004236 |
| 999 | LYMPHOCYTE APOPTOTIC PROCESS | 3 | 18 | 0.0009114 | 0.004236 |
| 1000 | MUSCLE CELL MIGRATION | 3 | 18 | 0.0009114 | 0.004236 |
| 1001 | CELLULAR RESPONSE TO DNA DAMAGE STIMULUS | 18 | 720 | 0.0009025 | 0.004236 |
| 1002 | REGULATION OF INTERLEUKIN 6 PRODUCTION | 6 | 104 | 0.0009549 | 0.004434 |
| 1003 | NEGATIVE REGULATION OF PROTEOLYSIS | 11 | 329 | 0.0009761 | 0.004528 |
| 1004 | I KAPPAB KINASE NF KAPPAB SIGNALING | 5 | 70 | 0.0009803 | 0.004539 |
| 1005 | EMBRYONIC ORGAN MORPHOGENESIS | 10 | 279 | 0.0009797 | 0.004539 |
| 1006 | REGULATION OF LIPID STORAGE | 4 | 41 | 0.0009967 | 0.004592 |
| 1007 | REGULATION OF MEMBRANE DEPOLARIZATION | 4 | 41 | 0.0009967 | 0.004592 |
| 1008 | CELLULAR RESPONSE TO ESTROGEN STIMULUS | 4 | 41 | 0.0009967 | 0.004592 |
| 1009 | MYELOID CELL ACTIVATION INVOLVED IN IMMUNE RESPONSE | 4 | 41 | 0.0009967 | 0.004592 |
| 1010 | MONOCYTE CHEMOTAXIS | 4 | 41 | 0.0009967 | 0.004592 |
| 1011 | REGULATION OF I KAPPAB KINASE NF KAPPAB SIGNALING | 9 | 233 | 0.001038 | 0.004779 |
| 1012 | POSITIVE REGULATION OF CALCIUM ION TRANSPORT | 6 | 106 | 0.001055 | 0.004849 |
| 1013 | REGULATION OF HYDROGEN PEROXIDE INDUCED CELL DEATH | 3 | 19 | 0.001074 | 0.004922 |
| 1014 | CARTILAGE DEVELOPMENT INVOLVED IN ENDOCHONDRAL BONE MORPHOGENESIS | 3 | 19 | 0.001074 | 0.004922 |
| 1015 | MACROPHAGE DIFFERENTIATION | 3 | 19 | 0.001074 | 0.004922 |
| 1016 | DETECTION OF MECHANICAL STIMULUS | 4 | 42 | 0.001092 | 0.004978 |
| 1017 | REGULATION OF BONE REMODELING | 4 | 42 | 0.001092 | 0.004978 |
| 1018 | NEGATIVE REGULATION OF CELL CYCLE PHASE TRANSITION | 7 | 146 | 0.001091 | 0.004978 |
| 1019 | DNA INTEGRITY CHECKPOINT | 7 | 146 | 0.001091 | 0.004978 |
| 1020 | REGULATION OF HEART GROWTH | 4 | 42 | 0.001092 | 0.004978 |
| 1021 | REGULATION OF BINDING | 10 | 283 | 0.001091 | 0.004978 |
| 1022 | MYELOID CELL DIFFERENTIATION | 8 | 189 | 0.001097 | 0.004993 |
| 1023 | REGULATION OF T CELL PROLIFERATION | 7 | 147 | 0.001135 | 0.005162 |
| 1024 | PANCREAS DEVELOPMENT | 5 | 73 | 0.001185 | 0.00538 |
| 1025 | G1 DNA DAMAGE CHECKPOINT | 5 | 73 | 0.001185 | 0.00538 |
| 1026 | REGULATION OF GLYCOPROTEIN METABOLIC PROCESS | 4 | 43 | 0.001194 | 0.005405 |
| 1027 | REGULATION OF MUSCLE CELL APOPTOTIC PROCESS | 4 | 43 | 0.001194 | 0.005405 |
| 1028 | REGULATION OF INSULIN RECEPTOR SIGNALING PATHWAY | 4 | 43 | 0.001194 | 0.005405 |
| 1029 | PATTERN RECOGNITION RECEPTOR SIGNALING PATHWAY | 6 | 109 | 0.001219 | 0.005511 |
| 1030 | MACROMOLECULAR COMPLEX ASSEMBLY | 28 | 1398 | 0.001245 | 0.005626 |
| 1031 | REGULATION OF SMOOTH MUSCLE CELL DIFFERENTIATION | 3 | 20 | 0.001253 | 0.005632 |
| 1032 | NEGATIVE REGULATION OF CELL CYCLE ARREST | 3 | 20 | 0.001253 | 0.005632 |
| 1033 | GENETIC IMPRINTING | 3 | 20 | 0.001253 | 0.005632 |
| 1034 | RESPONSE TO MUSCLE ACTIVITY | 3 | 20 | 0.001253 | 0.005632 |
| 1035 | LYMPH VESSEL DEVELOPMENT | 3 | 20 | 0.001253 | 0.005632 |
| 1036 | DEVELOPMENTAL MATURATION | 8 | 193 | 0.001254 | 0.005632 |
| 1037 | CARDIAC MUSCLE CELL DIFFERENTIATION | 5 | 74 | 0.00126 | 0.005653 |
| 1038 | AUTOPHAGY | 12 | 394 | 0.001297 | 0.005812 |
| 1039 | LABYRINTHINE LAYER DEVELOPMENT | 4 | 44 | 0.001303 | 0.005833 |
| 1040 | POSITIVE REGULATION OF CELLULAR AMIDE METABOLIC PROCESS | 6 | 111 | 0.001339 | 0.005983 |
| 1041 | POSITIVE REGULATION OF AUTOPHAGY | 5 | 75 | 0.001338 | 0.005983 |
| 1042 | ZYMOGEN ACTIVATION | 6 | 112 | 0.001402 | 0.006259 |
| 1043 | POSITIVE REGULATION OF PROTEIN COMPLEX ASSEMBLY | 8 | 197 | 0.001429 | 0.006374 |
| 1044 | B CELL HOMEOSTASIS | 3 | 21 | 0.00145 | 0.006445 |
| 1045 | MAST CELL ACTIVATION | 3 | 21 | 0.00145 | 0.006445 |
| 1046 | POSITIVE T CELL SELECTION | 3 | 21 | 0.00145 | 0.006445 |
| 1047 | POSITIVE REGULATION OF EXCITATORY POSTSYNAPTIC POTENTIAL | 3 | 21 | 0.00145 | 0.006445 |
| 1048 | REGULATION OF ALPHA BETA T CELL DIFFERENTIATION | 4 | 46 | 0.00154 | 0.006836 |
| 1049 | LYMPHOCYTE COSTIMULATION | 5 | 78 | 0.001595 | 0.007073 |
| 1050 | NEGATIVE REGULATION OF CELLULAR CATABOLIC PROCESS | 7 | 156 | 0.001599 | 0.007086 |
| 1051 | NEGATIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 6 | 115 | 0.001605 | 0.007106 |
| 1052 | NEGATIVE REGULATION OF TRANSFERASE ACTIVITY | 11 | 351 | 0.001636 | 0.007237 |
| 1053 | REGULATION OF AUTOPHAGY | 9 | 249 | 0.001644 | 0.007266 |
| 1054 | PROTEIN TARGETING | 12 | 406 | 0.001669 | 0.007339 |
| 1055 | THYMUS DEVELOPMENT | 4 | 47 | 0.001669 | 0.007339 |
| 1056 | POSITIVE REGULATION OF RECEPTOR MEDIATED ENDOCYTOSIS | 4 | 47 | 0.001669 | 0.007339 |
| 1057 | CELLULAR RESPONSE TO INTERLEUKIN 6 | 3 | 22 | 0.001666 | 0.007339 |
| 1058 | REGULATION OF RESPONSE TO INTERFERON GAMMA | 3 | 22 | 0.001666 | 0.007339 |
| 1059 | DETECTION OF ABIOTIC STIMULUS | 6 | 117 | 0.001753 | 0.0077 |
| 1060 | CELLULAR PROCESS INVOLVED IN REPRODUCTION IN MULTICELLULAR ORGANISM | 9 | 252 | 0.001785 | 0.007834 |
| 1061 | NUCLEAR TRANSPORT | 11 | 355 | 0.001789 | 0.007844 |
| 1062 | REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 4 | 48 | 0.001805 | 0.007886 |
| 1063 | RESPONSE TO AXON INJURY | 4 | 48 | 0.001805 | 0.007886 |
| 1064 | DIGESTIVE TRACT MORPHOGENESIS | 4 | 48 | 0.001805 | 0.007886 |
| 1065 | POSITIVE REGULATION OF WOUND HEALING | 4 | 48 | 0.001805 | 0.007886 |
| 1066 | MULTICELLULAR ORGANISM REPRODUCTION | 18 | 768 | 0.00185 | 0.008075 |
| 1067 | METANEPHROS DEVELOPMENT | 5 | 81 | 0.001885 | 0.008221 |
| 1068 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL METABOLIC PROCESS | 3 | 23 | 0.001901 | 0.008242 |
| 1069 | RESPONSE TO MAGNESIUM ION | 3 | 23 | 0.001901 | 0.008242 |
| 1070 | LYSOSOME LOCALIZATION | 3 | 23 | 0.001901 | 0.008242 |
| 1071 | REGULATION OF OSTEOBLAST PROLIFERATION | 3 | 23 | 0.001901 | 0.008242 |
| 1072 | TRABECULA FORMATION | 3 | 23 | 0.001901 | 0.008242 |
| 1073 | POSITIVE REGULATION OF COLLAGEN METABOLIC PROCESS | 3 | 23 | 0.001901 | 0.008242 |
| 1074 | REGULATION OF NEURONAL SYNAPTIC PLASTICITY | 4 | 49 | 0.001949 | 0.008428 |
| 1075 | REGULATION OF TOLL LIKE RECEPTOR SIGNALING PATHWAY | 4 | 49 | 0.001949 | 0.008428 |
| 1076 | REGULATION OF NITRIC OXIDE SYNTHASE ACTIVITY | 4 | 49 | 0.001949 | 0.008428 |
| 1077 | REGULATION OF CATION TRANSMEMBRANE TRANSPORT | 8 | 208 | 0.00201 | 0.008685 |
| 1078 | CYTOSKELETON ORGANIZATION | 19 | 838 | 0.002023 | 0.008731 |
| 1079 | REGULATION OF NUCLEAR DIVISION | 7 | 163 | 0.002053 | 0.008844 |
| 1080 | RESPONSE TO TOPOLOGICALLY INCORRECT PROTEIN | 7 | 163 | 0.002053 | 0.008844 |
| 1081 | REGULATION OF B CELL ACTIVATION | 6 | 121 | 0.002078 | 0.008945 |
| 1082 | RECEPTOR INTERNALIZATION | 4 | 50 | 0.002101 | 0.009025 |
| 1083 | VISUAL BEHAVIOR | 4 | 50 | 0.002101 | 0.009025 |
| 1084 | NEGATIVE REGULATION OF BLOOD VESSEL ENDOTHELIAL CELL MIGRATION | 3 | 24 | 0.002155 | 0.00919 |
| 1085 | REGULATION OF MYELOID CELL APOPTOTIC PROCESS | 3 | 24 | 0.002155 | 0.00919 |
| 1086 | POSITIVE REGULATION OF CELLULAR RESPONSE TO TRANSFORMING GROWTH FACTOR BETA STIMULUS | 3 | 24 | 0.002155 | 0.00919 |
| 1087 | AXON REGENERATION | 3 | 24 | 0.002155 | 0.00919 |
| 1088 | POSITIVE REGULATION OF OSTEOCLAST DIFFERENTIATION | 3 | 24 | 0.002155 | 0.00919 |
| 1089 | POSITIVE REGULATION OF EPITHELIAL CELL APOPTOTIC PROCESS | 3 | 24 | 0.002155 | 0.00919 |
| 1090 | POSITIVE REGULATION OF TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 3 | 24 | 0.002155 | 0.00919 |
| 1091 | REGULATION OF LONG TERM NEURONAL SYNAPTIC PLASTICITY | 3 | 24 | 0.002155 | 0.00919 |
| 1092 | LOCALIZATION WITHIN MEMBRANE | 6 | 122 | 0.002166 | 0.009231 |
| 1093 | POSITIVE REGULATION OF TRANSCRIPTION FACTOR IMPORT INTO NUCLEUS | 4 | 51 | 0.00226 | 0.009605 |
| 1094 | MECHANORECEPTOR DIFFERENTIATION | 4 | 51 | 0.00226 | 0.009605 |
| 1095 | HOMOTYPIC CELL CELL ADHESION | 4 | 51 | 0.00226 | 0.009605 |
| 1096 | CENTRAL NERVOUS SYSTEM NEURON DIFFERENTIATION | 7 | 166 | 0.002275 | 0.00966 |
| 1097 | POSITIVE REGULATION OF CELL CYCLE ARREST | 5 | 85 | 0.002331 | 0.009887 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | RECEPTOR BINDING | 89 | 1476 | 5.268e-44 | 4.894e-41 |
| 2 | KINASE ACTIVITY | 67 | 842 | 1.477e-39 | 6.86e-37 |
| 3 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 67 | 992 | 4.128e-35 | 1.278e-32 |
| 4 | PROTEIN KINASE ACTIVITY | 56 | 640 | 6.351e-35 | 1.475e-32 |
| 5 | PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 25 | 70 | 7.875e-32 | 1.463e-29 |
| 6 | PROTEIN TYROSINE KINASE ACTIVITY | 33 | 176 | 1.901e-31 | 2.944e-29 |
| 7 | GROWTH FACTOR BINDING | 28 | 123 | 2.609e-29 | 3.462e-27 |
| 8 | MOLECULAR FUNCTION REGULATOR | 65 | 1353 | 1.934e-25 | 2.246e-23 |
| 9 | GROWTH FACTOR ACTIVITY | 27 | 160 | 1.451e-24 | 1.498e-22 |
| 10 | GROWTH FACTOR RECEPTOR BINDING | 25 | 129 | 2.193e-24 | 2.037e-22 |
| 11 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE ACTIVITY | 20 | 64 | 2.709e-24 | 2.288e-22 |
| 12 | ENZYME BINDING | 70 | 1737 | 4.67e-23 | 3.615e-21 |
| 13 | PROTEIN COMPLEX BINDING | 52 | 935 | 5.448e-23 | 3.893e-21 |
| 14 | X1 PHOSPHATIDYLINOSITOL 3 KINASE ACTIVITY | 17 | 43 | 6.968e-23 | 4.624e-21 |
| 15 | TRANSMEMBRANE RECEPTOR PROTEIN KINASE ACTIVITY | 20 | 81 | 5.52e-22 | 3.419e-20 |
| 16 | PHOSPHATIDYLINOSITOL KINASE ACTIVITY | 17 | 51 | 2.265e-21 | 1.315e-19 |
| 17 | MACROMOLECULAR COMPLEX BINDING | 59 | 1399 | 3.691e-20 | 2.017e-18 |
| 18 | RAS GUANYL NUCLEOTIDE EXCHANGE FACTOR ACTIVITY | 26 | 228 | 3.115e-19 | 1.608e-17 |
| 19 | KINASE BINDING | 38 | 606 | 1.498e-18 | 7.322e-17 |
| 20 | SIGNAL TRANSDUCER ACTIVITY | 61 | 1731 | 4.68e-17 | 2.174e-15 |
| 21 | PLATELET DERIVED GROWTH FACTOR RECEPTOR BINDING | 10 | 15 | 5.259e-17 | 2.326e-15 |
| 22 | PLATELET DERIVED GROWTH FACTOR BINDING | 9 | 11 | 9.524e-17 | 4.022e-15 |
| 23 | GUANYL NUCLEOTIDE EXCHANGE FACTOR ACTIVITY | 26 | 303 | 3.672e-16 | 1.483e-14 |
| 24 | CELL ADHESION MOLECULE BINDING | 21 | 186 | 1.204e-15 | 4.661e-14 |
| 25 | INTEGRIN BINDING | 17 | 105 | 1.504e-15 | 5.589e-14 |
| 26 | COLLAGEN BINDING | 14 | 65 | 7.696e-15 | 2.75e-13 |
| 27 | EXTRACELLULAR MATRIX STRUCTURAL CONSTITUENT | 14 | 76 | 7.809e-14 | 2.687e-12 |
| 28 | PROTEIN PHOSPHATASE BINDING | 16 | 120 | 2.429e-13 | 8.06e-12 |
| 29 | PHOSPHATASE BINDING | 17 | 162 | 2.292e-12 | 7.341e-11 |
| 30 | ADENYL NUCLEOTIDE BINDING | 49 | 1514 | 2.862e-12 | 8.863e-11 |
| 31 | RIBONUCLEOTIDE BINDING | 55 | 1860 | 3.448e-12 | 1.033e-10 |
| 32 | INSULIN RECEPTOR SUBSTRATE BINDING | 7 | 11 | 5.111e-12 | 1.484e-10 |
| 33 | HEPARIN BINDING | 16 | 157 | 1.625e-11 | 4.574e-10 |
| 34 | KINASE REGULATOR ACTIVITY | 17 | 186 | 2.144e-11 | 5.857e-10 |
| 35 | PROTEIN PHOSPHATASE TYPE 2A REGULATOR ACTIVITY | 8 | 21 | 3.042e-11 | 8.075e-10 |
| 36 | NEUROTROPHIN RECEPTOR BINDING | 7 | 14 | 5.17e-11 | 1.334e-09 |
| 37 | INSULIN LIKE GROWTH FACTOR RECEPTOR BINDING | 7 | 15 | 9.605e-11 | 2.412e-09 |
| 38 | GLYCOSAMINOGLYCAN BINDING | 17 | 205 | 1.004e-10 | 2.455e-09 |
| 39 | CYTOKINE RECEPTOR BINDING | 19 | 271 | 1.404e-10 | 3.344e-09 |
| 40 | EXTRACELLULAR MATRIX BINDING | 10 | 51 | 1.592e-10 | 3.698e-09 |
| 41 | RECEPTOR ACTIVITY | 48 | 1649 | 2.012e-10 | 4.558e-09 |
| 42 | ENZYME REGULATOR ACTIVITY | 35 | 959 | 2.687e-10 | 5.944e-09 |
| 43 | FIBROBLAST GROWTH FACTOR RECEPTOR BINDING | 8 | 28 | 4.353e-10 | 9.191e-09 |
| 44 | FIBRONECTIN BINDING | 8 | 28 | 4.353e-10 | 9.191e-09 |
| 45 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 23 | 445 | 6.603e-10 | 1.363e-08 |
| 46 | SULFUR COMPOUND BINDING | 17 | 234 | 7.877e-10 | 1.585e-08 |
| 47 | IDENTICAL PROTEIN BINDING | 39 | 1209 | 8.02e-10 | 1.585e-08 |
| 48 | SIGNALING RECEPTOR ACTIVITY | 42 | 1393 | 1.228e-09 | 2.377e-08 |
| 49 | PHOSPHATIDYLINOSITOL 3 KINASE BINDING | 7 | 30 | 2.646e-08 | 5.017e-07 |
| 50 | CYTOKINE BINDING | 10 | 92 | 6.103e-08 | 1.134e-06 |
| 51 | PROTEIN DIMERIZATION ACTIVITY | 34 | 1149 | 9.312e-08 | 1.696e-06 |
| 52 | PROTEIN DOMAIN SPECIFIC BINDING | 23 | 624 | 3.407e-07 | 6.087e-06 |
| 53 | CHEMOATTRACTANT ACTIVITY | 6 | 27 | 3.725e-07 | 6.53e-06 |
| 54 | CYTOKINE RECEPTOR ACTIVITY | 9 | 89 | 5.293e-07 | 9.105e-06 |
| 55 | LAMININ BINDING | 6 | 30 | 7.273e-07 | 1.228e-05 |
| 56 | PROTEIN BINDING INVOLVED IN CELL ADHESION | 5 | 17 | 7.99e-07 | 1.326e-05 |
| 57 | VIRUS RECEPTOR ACTIVITY | 8 | 70 | 8.944e-07 | 1.458e-05 |
| 58 | PROTEIN HETERODIMERIZATION ACTIVITY | 19 | 468 | 9.132e-07 | 1.463e-05 |
| 59 | INSULIN RECEPTOR BINDING | 6 | 32 | 1.09e-06 | 1.716e-05 |
| 60 | HISTONE KINASE ACTIVITY | 5 | 19 | 1.475e-06 | 2.284e-05 |
| 61 | KINASE ACTIVATOR ACTIVITY | 7 | 62 | 4.763e-06 | 7.137e-05 |
| 62 | PHOSPHATASE REGULATOR ACTIVITY | 8 | 87 | 4.728e-06 | 7.137e-05 |
| 63 | EPHRIN RECEPTOR BINDING | 5 | 24 | 5.159e-06 | 7.607e-05 |
| 64 | CYCLIN DEPENDENT PROTEIN SERINE THREONINE KINASE REGULATOR ACTIVITY | 5 | 28 | 1.151e-05 | 0.0001671 |
| 65 | PROTEASE BINDING | 8 | 104 | 1.78e-05 | 0.0002544 |
| 66 | PHOSPHATIDYLINOSITOL PHOSPHATE KINASE ACTIVITY | 4 | 16 | 2.215e-05 | 0.0003117 |
| 67 | SODIUM CHANNEL REGULATOR ACTIVITY | 5 | 32 | 2.277e-05 | 0.0003158 |
| 68 | PROTEIN TYROSINE KINASE BINDING | 6 | 54 | 2.547e-05 | 0.0003479 |
| 69 | PROTEIN HOMODIMERIZATION ACTIVITY | 21 | 722 | 4.167e-05 | 0.0005611 |
| 70 | CYCLIN BINDING | 4 | 19 | 4.598e-05 | 0.0006102 |
| 71 | CHEMOKINE BINDING | 4 | 21 | 6.979e-05 | 0.0009132 |
| 72 | FIBROBLAST GROWTH FACTOR BINDING | 4 | 23 | 0.0001015 | 0.00131 |
| 73 | NON MEMBRANE SPANNING PROTEIN TYROSINE KINASE ACTIVITY | 5 | 46 | 0.000137 | 0.001744 |
| 74 | UBIQUITIN LIKE PROTEIN LIGASE BINDING | 11 | 264 | 0.000153 | 0.001921 |
| 75 | ION CHANNEL BINDING | 7 | 111 | 0.0002101 | 0.002602 |
| 76 | PROTEIN PHOSPHATASE 2A BINDING | 4 | 28 | 0.000225 | 0.00275 |
| 77 | CAMP RESPONSE ELEMENT BINDING | 3 | 13 | 0.0003325 | 0.004011 |
| 78 | STRUCTURAL MOLECULE ACTIVITY | 19 | 732 | 0.0004132 | 0.004921 |
| 79 | CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 4 | 34 | 0.0004843 | 0.005695 |
| 80 | CHANNEL REGULATOR ACTIVITY | 7 | 131 | 0.0005754 | 0.006682 |
| 81 | RECEPTOR SIGNALING PROTEIN ACTIVITY | 8 | 172 | 0.0005942 | 0.006815 |
| 82 | PEPTIDE HORMONE BINDING | 4 | 36 | 0.0006048 | 0.006852 |
| 83 | MHC CLASS II PROTEIN COMPLEX BINDING | 3 | 16 | 0.0006356 | 0.007114 |
| 84 | CYTOKINE ACTIVITY | 9 | 219 | 0.00067 | 0.007409 |
| 85 | CORECEPTOR ACTIVITY | 4 | 38 | 0.0007451 | 0.008144 |
| 86 | PROTEIN SERINE THREONINE TYROSINE KINASE ACTIVITY | 4 | 39 | 0.0008233 | 0.008893 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | EXTRACELLULAR MATRIX COMPONENT | 29 | 125 | 1.367e-30 | 7.981e-28 |
| 2 | EXTRACELLULAR MATRIX | 43 | 426 | 3.161e-29 | 9.231e-27 |
| 3 | PROTEIN COMPLEX INVOLVED IN CELL ADHESION | 18 | 30 | 1.636e-28 | 3.185e-26 |
| 4 | PROTEINACEOUS EXTRACELLULAR MATRIX | 38 | 356 | 9.833e-27 | 1.436e-24 |
| 5 | CELL SURFACE | 50 | 757 | 2.072e-25 | 2.42e-23 |
| 6 | BASEMENT MEMBRANE | 23 | 93 | 3.831e-25 | 3.729e-23 |
| 7 | RECEPTOR COMPLEX | 35 | 327 | 1.079e-24 | 9.002e-23 |
| 8 | PLASMA MEMBRANE RECEPTOR COMPLEX | 26 | 175 | 3.274e-22 | 2.39e-20 |
| 9 | CELL SUBSTRATE JUNCTION | 33 | 398 | 7.889e-20 | 5.119e-18 |
| 10 | COMPLEX OF COLLAGEN TRIMERS | 12 | 23 | 2.386e-18 | 1.393e-16 |
| 11 | SIDE OF MEMBRANE | 32 | 428 | 6.391e-18 | 3.393e-16 |
| 12 | PLASMA MEMBRANE PROTEIN COMPLEX | 34 | 510 | 1.832e-17 | 8.918e-16 |
| 13 | BASAL LAMINA | 11 | 21 | 6.099e-17 | 2.74e-15 |
| 14 | ANCHORING JUNCTION | 32 | 489 | 3.019e-16 | 1.259e-14 |
| 15 | INTRINSIC COMPONENT OF PLASMA MEMBRANE | 56 | 1649 | 6.492e-15 | 2.528e-13 |
| 16 | MEMBRANE PROTEIN COMPLEX | 43 | 1020 | 1.238e-14 | 4.517e-13 |
| 17 | MEMBRANE REGION | 45 | 1134 | 2.366e-14 | 8.128e-13 |
| 18 | COLLAGEN TRIMER | 15 | 88 | 3.298e-14 | 1.07e-12 |
| 19 | INTRACELLULAR VESICLE | 47 | 1259 | 5.355e-14 | 1.646e-12 |
| 20 | ENDOPLASMIC RETICULUM LUMEN | 20 | 201 | 6.769e-14 | 1.977e-12 |
| 21 | TRANSFERASE COMPLEX TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 21 | 237 | 1.564e-13 | 4.349e-12 |
| 22 | CELL JUNCTION | 44 | 1151 | 1.781e-13 | 4.728e-12 |
| 23 | PROTEIN PHOSPHATASE TYPE 2A COMPLEX | 9 | 20 | 2.672e-13 | 6.786e-12 |
| 24 | EXTRINSIC COMPONENT OF MEMBRANE | 21 | 252 | 5.206e-13 | 1.267e-11 |
| 25 | PROTEIN KINASE COMPLEX | 14 | 90 | 8.988e-13 | 2.1e-11 |
| 26 | EXTRACELLULAR SPACE | 47 | 1376 | 1.331e-12 | 2.989e-11 |
| 27 | PLASMA MEMBRANE REGION | 37 | 929 | 6.194e-12 | 1.34e-10 |
| 28 | CATALYTIC COMPLEX | 38 | 1038 | 3.73e-11 | 7.779e-10 |
| 29 | EXTERNAL SIDE OF PLASMA MEMBRANE | 18 | 238 | 1.272e-10 | 2.561e-09 |
| 30 | VESICLE LUMEN | 13 | 106 | 1.319e-10 | 2.568e-09 |
| 31 | MEMBRANE MICRODOMAIN | 19 | 288 | 3.945e-10 | 7.432e-09 |
| 32 | PLATELET ALPHA GRANULE | 11 | 75 | 5.075e-10 | 9.262e-09 |
| 33 | CYTOPLASMIC SIDE OF MEMBRANE | 15 | 170 | 5.514e-10 | 9.759e-09 |
| 34 | EXTRINSIC COMPONENT OF CYTOPLASMIC SIDE OF PLASMA MEMBRANE | 12 | 98 | 7.027e-10 | 1.207e-08 |
| 35 | PHOSPHATIDYLINOSITOL 3 KINASE COMPLEX | 7 | 20 | 1.105e-09 | 1.792e-08 |
| 36 | CYCLIN DEPENDENT PROTEIN KINASE HOLOENZYME COMPLEX | 8 | 31 | 1.074e-09 | 1.792e-08 |
| 37 | PHOSPHATASE COMPLEX | 9 | 48 | 2.051e-09 | 3.238e-08 |
| 38 | EXTRINSIC COMPONENT OF PLASMA MEMBRANE | 13 | 136 | 3.014e-09 | 4.633e-08 |
| 39 | PLATELET ALPHA GRANULE LUMEN | 9 | 55 | 7.282e-09 | 1.09e-07 |
| 40 | PERINUCLEAR REGION OF CYTOPLASM | 25 | 642 | 3.525e-08 | 5.146e-07 |
| 41 | CELL PROJECTION | 44 | 1786 | 1.885e-07 | 2.685e-06 |
| 42 | SECRETORY GRANULE LUMEN | 9 | 85 | 3.559e-07 | 4.949e-06 |
| 43 | CYTOPLASMIC VESICLE PART | 22 | 601 | 6.99e-07 | 9.493e-06 |
| 44 | HETEROTRIMERIC G PROTEIN COMPLEX | 6 | 32 | 1.09e-06 | 1.447e-05 |
| 45 | CELL LEADING EDGE | 16 | 350 | 1.496e-06 | 1.941e-05 |
| 46 | SECRETORY GRANULE | 16 | 352 | 1.611e-06 | 2.045e-05 |
| 47 | LEADING EDGE MEMBRANE | 10 | 134 | 2.078e-06 | 2.582e-05 |
| 48 | RUFFLE MEMBRANE | 8 | 80 | 2.503e-06 | 3.045e-05 |
| 49 | CELL PROJECTION MEMBRANE | 14 | 298 | 5.027e-06 | 5.991e-05 |
| 50 | BANDED COLLAGEN FIBRIL | 4 | 12 | 6.233e-06 | 7.28e-05 |
| 51 | TRANSFERASE COMPLEX | 22 | 703 | 8.813e-06 | 0.0001009 |
| 52 | VACUOLE | 30 | 1180 | 1.224e-05 | 0.0001374 |
| 53 | ENDOPLASMIC RETICULUM | 37 | 1631 | 1.396e-05 | 0.0001538 |
| 54 | PIGMENT GRANULE | 8 | 103 | 1.659e-05 | 0.0001794 |
| 55 | BASAL PART OF CELL | 6 | 51 | 1.824e-05 | 0.0001937 |
| 56 | BASOLATERAL PLASMA MEMBRANE | 11 | 211 | 2.031e-05 | 0.0002118 |
| 57 | SOMATODENDRITIC COMPARTMENT | 20 | 650 | 2.922e-05 | 0.0002994 |
| 58 | NEURON PROJECTION | 25 | 942 | 3.458e-05 | 0.0003482 |
| 59 | CELL PROJECTION PART | 25 | 946 | 3.706e-05 | 0.0003668 |
| 60 | PLASMA MEMBRANE RAFT | 7 | 86 | 4.177e-05 | 0.0004065 |
| 61 | NEURON PART | 30 | 1265 | 4.517e-05 | 0.0004325 |
| 62 | SECRETORY VESICLE | 16 | 461 | 4.638e-05 | 0.0004368 |
| 63 | RUFFLE | 9 | 156 | 5.307e-05 | 0.0004919 |
| 64 | LAMELLIPODIUM | 9 | 172 | 0.0001126 | 0.001027 |
| 65 | ENDOCYTIC VESICLE | 11 | 256 | 0.0001168 | 0.00105 |
| 66 | ENDOPLASMIC RETICULUM PART | 27 | 1163 | 0.0001589 | 0.001406 |
| 67 | BASAL PLASMA MEMBRANE | 4 | 33 | 0.0004309 | 0.003712 |
| 68 | DENDRITE | 14 | 451 | 0.0004322 | 0.003712 |
| 69 | MYELIN SHEATH | 8 | 168 | 0.0005086 | 0.004305 |
| 70 | SYNAPSE | 19 | 754 | 0.000592 | 0.004939 |
| 71 | ACTIN BASED CELL PROJECTION | 8 | 181 | 0.0008295 | 0.006823 |
| 72 | FILOPODIUM MEMBRANE | 3 | 18 | 0.0009114 | 0.007392 |
| 73 | LAMELLIPODIUM MEMBRANE | 3 | 19 | 0.001074 | 0.008378 |
| 74 | CELL BODY | 14 | 494 | 0.001048 | 0.008378 |
| 75 | ENDOSOME | 19 | 793 | 0.001076 | 0.008378 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04151_PI3K_AKT_signaling_pathway | 217 | 351 | 0 | 0 | |
| 2 | hsa04510_Focal_adhesion | 90 | 200 | 1.185e-128 | 1.066e-126 | |
| 3 | hsa04014_Ras_signaling_pathway | 77 | 236 | 1.197e-95 | 7.184e-94 | |
| 4 | hsa04512_ECM.receptor_interaction | 51 | 85 | 5.321e-80 | 2.395e-78 | |
| 5 | hsa04810_Regulation_of_actin_cytoskeleton | 57 | 214 | 4.52e-64 | 1.627e-62 | |
| 6 | hsa04722_Neurotrophin_signaling_pathway | 33 | 127 | 1.597e-36 | 4.79e-35 | |
| 7 | hsa04010_MAPK_signaling_pathway | 41 | 268 | 3.074e-35 | 7.904e-34 | |
| 8 | hsa04150_mTOR_signaling_pathway | 23 | 52 | 5.313e-32 | 1.196e-30 | |
| 9 | hsa04062_Chemokine_signaling_pathway | 34 | 189 | 9.64e-32 | 1.928e-30 | |
| 10 | hsa04012_ErbB_signaling_pathway | 25 | 87 | 4.497e-29 | 8.094e-28 | |
| 11 | hsa04910_Insulin_signaling_pathway | 28 | 138 | 8.416e-28 | 1.377e-26 | |
| 12 | hsa04630_Jak.STAT_signaling_pathway | 29 | 155 | 1.103e-27 | 1.655e-26 | |
| 13 | hsa04662_B_cell_receptor_signaling_pathway | 23 | 75 | 1.444e-27 | 2e-26 | |
| 14 | hsa04380_Osteoclast_differentiation | 23 | 128 | 1.019e-21 | 1.31e-20 | |
| 15 | hsa04640_Hematopoietic_cell_lineage | 20 | 88 | 3.384e-21 | 4.06e-20 | |
| 16 | hsa04370_VEGF_signaling_pathway | 19 | 76 | 4.707e-21 | 5.296e-20 | |
| 17 | hsa04664_Fc_epsilon_RI_signaling_pathway | 19 | 79 | 1.057e-20 | 1.118e-19 | |
| 18 | hsa04660_T_cell_receptor_signaling_pathway | 21 | 108 | 1.118e-20 | 1.118e-19 | |
| 19 | hsa04620_Toll.like_receptor_signaling_pathway | 20 | 102 | 8.016e-20 | 7.594e-19 | |
| 20 | hsa04960_Aldosterone.regulated_sodium_reabsorption | 14 | 42 | 8.304e-18 | 7.473e-17 | |
| 21 | hsa04914_Progesterone.mediated_oocyte_maturation | 17 | 87 | 5.387e-17 | 4.617e-16 | |
| 22 | hsa04650_Natural_killer_cell_mediated_cytotoxicity | 19 | 136 | 5.281e-16 | 4.321e-15 | |
| 23 | hsa04540_Gap_junction | 16 | 90 | 2.235e-15 | 1.749e-14 | |
| 24 | hsa04110_Cell_cycle | 18 | 128 | 2.83e-15 | 2.123e-14 | |
| 25 | hsa04114_Oocyte_meiosis | 17 | 114 | 6.262e-15 | 4.509e-14 | |
| 26 | hsa04210_Apoptosis | 15 | 89 | 3.929e-14 | 2.72e-13 | |
| 27 | hsa04390_Hippo_signaling_pathway | 18 | 154 | 7.688e-14 | 5.126e-13 | |
| 28 | hsa04666_Fc_gamma_R.mediated_phagocytosis | 15 | 95 | 1.073e-13 | 6.9e-13 | |
| 29 | hsa04115_p53_signaling_pathway | 13 | 69 | 4.557e-13 | 2.828e-12 | |
| 30 | hsa04974_Protein_digestion_and_absorption | 13 | 81 | 3.948e-12 | 2.369e-11 | |
| 31 | hsa04530_Tight_junction | 15 | 133 | 1.644e-11 | 9.544e-11 | |
| 32 | hsa04920_Adipocytokine_signaling_pathway | 11 | 68 | 1.697e-10 | 9.547e-10 | |
| 33 | hsa04730_Long.term_depression | 11 | 70 | 2.351e-10 | 1.282e-09 | |
| 34 | hsa04973_Carbohydrate_digestion_and_absorption | 9 | 44 | 9.002e-10 | 4.766e-09 | |
| 35 | hsa04360_Axon_guidance | 13 | 130 | 1.726e-09 | 8.875e-09 | |
| 36 | hsa04350_TGF.beta_signaling_pathway | 11 | 85 | 2.011e-09 | 1.005e-08 | |
| 37 | hsa04670_Leukocyte_transendothelial_migration | 12 | 117 | 5.535e-09 | 2.693e-08 | |
| 38 | hsa04144_Endocytosis | 15 | 203 | 6.407e-09 | 3.035e-08 | |
| 39 | hsa04916_Melanogenesis | 11 | 101 | 1.287e-08 | 5.941e-08 | |
| 40 | hsa04145_Phagosome | 13 | 156 | 1.603e-08 | 7.216e-08 | |
| 41 | hsa03015_mRNA_surveillance_pathway | 10 | 83 | 2.241e-08 | 9.838e-08 | |
| 42 | hsa04310_Wnt_signaling_pathway | 12 | 151 | 9.795e-08 | 4.198e-07 | |
| 43 | hsa04070_Phosphatidylinositol_signaling_system | 9 | 78 | 1.683e-07 | 7.046e-07 | |
| 44 | hsa04320_Dorso.ventral_axis_formation | 6 | 25 | 2.27e-07 | 9.285e-07 | |
| 45 | hsa04621_NOD.like_receptor_signaling_pathway | 8 | 59 | 2.327e-07 | 9.308e-07 | |
| 46 | hsa04912_GnRH_signaling_pathway | 9 | 101 | 1.554e-06 | 6.081e-06 | |
| 47 | hsa04520_Adherens_junction | 7 | 73 | 1.427e-05 | 5.464e-05 | |
| 48 | hsa04720_Long.term_potentiation | 6 | 70 | 0.0001119 | 0.0004197 | |
| 49 | hsa04514_Cell_adhesion_molecules_.CAMs. | 8 | 136 | 0.000121 | 0.0004444 | |
| 50 | hsa00562_Inositol_phosphate_metabolism | 5 | 57 | 0.00038 | 0.001368 | |
| 51 | hsa04623_Cytosolic_DNA.sensing_pathway | 4 | 56 | 0.003185 | 0.01124 | |
| 52 | hsa04962_Vasopressin.regulated_water_reabsorption | 3 | 44 | 0.01203 | 0.04165 | |
| 53 | hsa04020_Calcium_signaling_pathway | 6 | 177 | 0.01291 | 0.04384 | |
| 54 | hsa04672_Intestinal_immune_network_for_IgA_production | 3 | 49 | 0.01609 | 0.05364 | |
| 55 | hsa04622_RIG.I.like_receptor_signaling_pathway | 3 | 71 | 0.04203 | 0.1376 | |
| 56 | hsa04140_Regulation_of_autophagy | 2 | 34 | 0.05239 | 0.1673 | |
| 57 | hsa04612_Antigen_processing_and_presentation | 3 | 78 | 0.05299 | 0.1673 | |
| 58 | hsa04742_Taste_transduction | 2 | 52 | 0.1093 | 0.3372 | |
| 59 | hsa04141_Protein_processing_in_endoplasmic_reticulum | 4 | 168 | 0.1105 | 0.3372 | |
| 60 | hsa04270_Vascular_smooth_muscle_contraction | 3 | 116 | 0.1321 | 0.3962 | |
| 61 | hsa04610_Complement_and_coagulation_cascades | 2 | 69 | 0.1722 | 0.5082 | |
| 62 | hsa03320_PPAR_signaling_pathway | 2 | 70 | 0.1761 | 0.5113 | |
| 63 | hsa03013_RNA_transport | 3 | 152 | 0.2286 | 0.6428 | |
| 64 | hsa04972_Pancreatic_secretion | 2 | 101 | 0.2997 | 0.8052 | |
| 65 | hsa04120_Ubiquitin_mediated_proteolysis | 2 | 139 | 0.4464 | 1 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | TBX5-AS1 |
hsa-miR-130b-3p;hsa-miR-16-1-3p;hsa-miR-183-5p;hsa-miR-19b-1-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 10 | ITGA8 | Sponge network | -2.108 | 0 | -3.151 | 0 | 0.731 |
| 2 | TBX5-AS1 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 21 | PDGFRA | Sponge network | -2.108 | 0 | -0.92 | 0.00039 | 0.659 |
| 3 | LINC00702 |
hsa-miR-130b-3p;hsa-miR-16-1-3p;hsa-miR-183-5p;hsa-miR-19b-1-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 11 | ITGA8 | Sponge network | -2.856 | 0 | -3.151 | 0 | 0.657 |
| 4 | RP11-863P13.1 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-107;hsa-miR-30a-3p;hsa-miR-30b-3p;hsa-miR-30d-3p;hsa-miR-30e-3p | 10 | COL1A1 | Sponge network | 3.777 | 5.0E-5 | 2.421 | 0 | 0.617 |
| 5 | DNM3OS |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 11 | PDGFRA | Sponge network | 0.053 | 0.85755 | -0.92 | 0.00039 | 0.612 |
| 6 | MAGI2-AS3 |
hsa-miR-130b-3p;hsa-miR-16-1-3p;hsa-miR-183-5p;hsa-miR-19b-1-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-454-3p;hsa-miR-7-1-3p | 10 | ITGA8 | Sponge network | -1.892 | 0 | -3.151 | 0 | 0.607 |
| 7 | LINC00702 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-339-5p;hsa-miR-33a-5p;hsa-miR-34a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 29 | PDGFRA | Sponge network | -2.856 | 0 | -0.92 | 0.00039 | 0.589 |
| 8 | TBX5-AS1 |
hsa-miR-107;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-501-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 10 | FGF7 | Sponge network | -2.108 | 0 | -1.351 | 0 | 0.587 |
| 9 | AC109642.1 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-96-5p | 17 | PDGFRA | Sponge network | -2.791 | 0 | -0.92 | 0.00039 | 0.557 |
| 10 | GAS6-AS2 |
hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-5p | 12 | FGF2 | Sponge network | -1.761 | 0 | -2.545 | 0 | 0.556 |
| 11 | RP11-354E11.2 |
hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-185-3p;hsa-miR-18a-5p;hsa-miR-193b-5p;hsa-miR-590-3p;hsa-miR-671-5p;hsa-miR-9-5p;hsa-miR-940;hsa-miR-96-5p | 10 | GNG7 | Sponge network | -2.138 | 0 | -1.412 | 0 | 0.549 |
| 12 | TBX5-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-590-5p;hsa-miR-629-3p | 20 | PIK3R1 | Sponge network | -2.108 | 0 | -1.285 | 0 | 0.544 |
| 13 | MAGI2-AS3 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-429;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 24 | PDGFRA | Sponge network | -1.892 | 0 | -0.92 | 0.00039 | 0.534 |
| 14 | AC079630.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-942-5p | 16 | IL6R | Sponge network | -3.758 | 0 | -1.175 | 0 | 0.531 |
| 15 | AC109642.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-96-5p | 19 | PIK3R1 | Sponge network | -2.791 | 0 | -1.285 | 0 | 0.529 |
| 16 | LINC00702 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-93-3p | 18 | FGF2 | Sponge network | -2.856 | 0 | -2.545 | 0 | 0.527 |
| 17 | RP11-389C8.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-550a-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 24 | CCND2 | Sponge network | -2.039 | 0 | -1.641 | 0 | 0.52 |
| 18 | RP11-389C8.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 16 | PIK3R1 | Sponge network | -2.039 | 0 | -1.285 | 0 | 0.52 |
| 19 | MIR497HG |
hsa-miR-141-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-185-3p;hsa-miR-18a-5p;hsa-miR-3127-5p;hsa-miR-671-5p;hsa-miR-766-3p;hsa-miR-9-5p;hsa-miR-940;hsa-miR-96-5p | 11 | GNG7 | Sponge network | -2.142 | 0 | -1.412 | 0 | 0.519 |
| 20 | SH3RF3-AS1 |
hsa-miR-130b-3p;hsa-miR-16-1-3p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-26b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 10 | PDGFRA | Sponge network | -1.583 | 0 | -0.92 | 0.00039 | 0.516 |
| 21 | MAGI2-AS3 |
hsa-miR-107;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-16-2-3p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-429;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 10 | FGF7 | Sponge network | -1.892 | 0 | -1.351 | 0 | 0.514 |
| 22 | LINC00968 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p;hsa-miR-96-5p | 21 | PIK3R1 | Sponge network | -4.19 | 0 | -1.285 | 0 | 0.51 |
| 23 | CYP1B1-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-3p;hsa-miR-93-5p | 12 | COL4A3 | Sponge network | -1.073 | 0.00045 | -2.369 | 0 | 0.505 |
| 24 | LINC00702 |
hsa-miR-107;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 11 | FGF7 | Sponge network | -2.856 | 0 | -1.351 | 0 | 0.505 |
| 25 | AF131215.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-191-5p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-301a-3p;hsa-miR-324-3p;hsa-miR-550a-5p;hsa-miR-589-3p | 14 | CCND2 | Sponge network | -2.09 | 0 | -1.641 | 0 | 0.505 |
| 26 | TBX5-AS1 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-590-5p;hsa-miR-93-3p | 15 | FGF2 | Sponge network | -2.108 | 0 | -2.545 | 0 | 0.504 |
| 27 | LINC00968 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-590-5p;hsa-miR-93-3p | 12 | FGF2 | Sponge network | -4.19 | 0 | -2.545 | 0 | 0.504 |
| 28 | LINC00702 |
hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 19 | THBS1 | Sponge network | -2.856 | 0 | -0.931 | 0.0014 | 0.501 |
| 29 | MAGI2-AS3 |
hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-503-5p;hsa-miR-93-3p | 15 | FGF2 | Sponge network | -1.892 | 0 | -2.545 | 0 | 0.499 |
| 30 | RP11-720L2.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-93-5p | 13 | PIK3R1 | Sponge network | -2.305 | 0 | -1.285 | 0 | 0.496 |
| 31 | GAS6-AS2 |
hsa-miR-130b-3p;hsa-miR-16-1-3p;hsa-miR-183-5p;hsa-miR-19b-1-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 10 | ITGA8 | Sponge network | -1.761 | 0 | -3.151 | 0 | 0.495 |
| 32 | LINC00702 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p;hsa-miR-96-5p | 23 | PIK3R1 | Sponge network | -2.856 | 0 | -1.285 | 0 | 0.494 |
| 33 | AC079630.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-10a-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-3607-3p;hsa-miR-590-3p;hsa-miR-625-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 12 | COL4A3 | Sponge network | -3.758 | 0 | -2.369 | 0 | 0.493 |
| 34 | RP11-750H9.5 | hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-2355-3p;hsa-miR-301a-3p;hsa-miR-550a-5p;hsa-miR-877-5p | 10 | CCND2 | Sponge network | -1.959 | 0 | -1.641 | 0 | 0.493 |
| 35 | MAGI2-AS3 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-3p;hsa-miR-200b-5p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-429;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 14 | ITGA1 | Sponge network | -1.892 | 0 | -0.774 | 0.0005 | 0.493 |
| 36 | FENDRR |
hsa-miR-142-3p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-93-3p | 10 | FGF2 | Sponge network | -4.222 | 0 | -2.545 | 0 | 0.489 |
| 37 | PCED1B-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-877-5p;hsa-miR-93-5p;hsa-miR-96-5p | 18 | CCND2 | Sponge network | -0.672 | 0.02084 | -1.641 | 0 | 0.488 |
| 38 | MAGI2-AS3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 18 | PIK3R1 | Sponge network | -1.892 | 0 | -1.285 | 0 | 0.486 |
| 39 | LINC00968 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-942-5p | 19 | IL6R | Sponge network | -4.19 | 0 | -1.175 | 0 | 0.477 |
| 40 | AC109642.1 |
hsa-miR-141-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-193b-5p;hsa-miR-590-3p;hsa-miR-766-3p;hsa-miR-9-5p;hsa-miR-92a-3p;hsa-miR-96-5p | 10 | GNG7 | Sponge network | -2.791 | 0 | -1.412 | 0 | 0.473 |
| 41 | LINC00968 |
hsa-miR-141-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-185-3p;hsa-miR-18a-5p;hsa-miR-193b-5p;hsa-miR-590-3p;hsa-miR-671-5p;hsa-miR-9-5p;hsa-miR-96-5p | 10 | GNG7 | Sponge network | -4.19 | 0 | -1.412 | 0 | 0.472 |
| 42 | SH3RF3-AS1 |
hsa-miR-16-1-3p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-29b-1-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 11 | THBS1 | Sponge network | -1.583 | 0 | -0.931 | 0.0014 | 0.469 |
| 43 | RP11-399O19.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-7-1-3p;hsa-miR-93-5p | 18 | CCND2 | Sponge network | -0.873 | 0.00072 | -1.641 | 0 | 0.469 |
| 44 | RP11-354E11.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-93-3p | 11 | COL4A3 | Sponge network | -2.138 | 0 | -2.369 | 0 | 0.468 |
| 45 | AC109642.1 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-32-5p;hsa-miR-503-5p;hsa-miR-93-3p | 12 | FGF2 | Sponge network | -2.791 | 0 | -2.545 | 0 | 0.467 |
| 46 | MAGI2-AS3 |
hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 18 | THBS1 | Sponge network | -1.892 | 0 | -0.931 | 0.0014 | 0.461 |
| 47 | TBX5-AS1 |
hsa-miR-141-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-185-3p;hsa-miR-18a-5p;hsa-miR-193b-5p;hsa-miR-3127-5p;hsa-miR-766-3p;hsa-miR-9-5p;hsa-miR-92a-3p | 10 | GNG7 | Sponge network | -2.108 | 0 | -1.412 | 0 | 0.46 |
| 48 | AF131215.9 |
hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-590-3p | 13 | CCND2 | Sponge network | -1.808 | 0 | -1.641 | 0 | 0.46 |
| 49 | RP11-166D19.1 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-429;hsa-miR-454-3p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-96-5p | 21 | PDGFRA | Sponge network | -0.582 | 0.05253 | -0.92 | 0.00039 | 0.459 |
| 50 | LINC00472 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-93-5p | 14 | PIK3R1 | Sponge network | -2.952 | 0 | -1.285 | 0 | 0.458 |
| 51 | BZRAP1-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-550a-5p;hsa-miR-877-5p | 14 | CCND2 | Sponge network | -0.785 | 0.00723 | -1.641 | 0 | 0.456 |
| 52 | LINC00996 | hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-320b;hsa-miR-550a-5p | 10 | CCND2 | Sponge network | -1.372 | 0.00025 | -1.641 | 0 | 0.454 |
| 53 | GAS6-AS2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-429;hsa-miR-590-5p;hsa-miR-7-1-3p | 20 | PDGFRA | Sponge network | -1.761 | 0 | -0.92 | 0.00039 | 0.451 |
| 54 | FENDRR |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-335-3p;hsa-miR-3607-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-3p;hsa-miR-93-5p | 11 | COL4A3 | Sponge network | -4.222 | 0 | -2.369 | 0 | 0.45 |
| 55 | CTD-2013N24.2 |
hsa-miR-130b-3p;hsa-miR-16-1-3p;hsa-miR-183-5p;hsa-miR-19b-1-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-454-3p;hsa-miR-7-1-3p | 10 | ITGA8 | Sponge network | -1.745 | 0 | -3.151 | 0 | 0.448 |
| 56 | RP11-532F6.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-301a-3p | 13 | CCND2 | Sponge network | -2.028 | 0 | -1.641 | 0 | 0.445 |
| 57 | PCED1B-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-429;hsa-miR-93-5p;hsa-miR-96-5p | 11 | CSF1 | Sponge network | -0.672 | 0.02084 | -0.65 | 0.00787 | 0.444 |
| 58 | LINC00968 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-339-5p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-96-5p | 22 | PDGFRA | Sponge network | -4.19 | 0 | -0.92 | 0.00039 | 0.443 |
| 59 | LINC00968 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-590-3p;hsa-miR-625-5p;hsa-miR-93-3p;hsa-miR-93-5p | 11 | COL4A3 | Sponge network | -4.19 | 0 | -2.369 | 0 | 0.442 |
| 60 | CTD-2013N24.2 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-429 | 12 | FGF2 | Sponge network | -1.745 | 0 | -2.545 | 0 | 0.441 |
| 61 | FENDRR |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p;hsa-miR-96-5p | 17 | PIK3R1 | Sponge network | -4.222 | 0 | -1.285 | 0 | 0.438 |
| 62 | AC007743.1 |
hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-424-5p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p | 12 | PIK3R1 | Sponge network | -2.595 | 0 | -1.285 | 0 | 0.436 |
| 63 | LINC00702 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-450b-5p;hsa-miR-501-5p;hsa-miR-503-5p;hsa-miR-550a-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-660-5p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-5p;hsa-miR-96-5p | 34 | CCND2 | Sponge network | -2.856 | 0 | -1.641 | 0 | 0.433 |
| 64 | TBX5-AS1 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-362-5p;hsa-miR-450b-5p;hsa-miR-576-5p;hsa-miR-629-5p | 14 | IGF1 | Sponge network | -2.108 | 0 | -0.879 | 0.00545 | 0.429 |
| 65 | RP11-456K23.1 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-362-5p;hsa-miR-450b-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p | 15 | IGF1 | Sponge network | -1.488 | 0 | -0.879 | 0.00545 | 0.429 |
| 66 | RP11-389C8.2 |
hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-589-3p | 11 | FGF2 | Sponge network | -2.039 | 0 | -2.545 | 0 | 0.428 |
| 67 | CYP1B1-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-320b;hsa-miR-421;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-942-5p | 12 | IL6R | Sponge network | -1.073 | 0.00045 | -1.175 | 0 | 0.427 |
| 68 | LINC00472 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p | 10 | NTRK2 | Sponge network | -2.952 | 0 | -2.564 | 0 | 0.427 |
| 69 | RP11-166D19.1 |
hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 15 | THBS1 | Sponge network | -0.582 | 0.05253 | -0.931 | 0.0014 | 0.426 |
| 70 | MIR497HG |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 18 | PDGFRA | Sponge network | -2.142 | 0 | -0.92 | 0.00039 | 0.426 |
| 71 | TBX5-AS1 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 10 | ITGA1 | Sponge network | -2.108 | 0 | -0.774 | 0.0005 | 0.424 |
| 72 | RP11-389C8.2 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-3p;hsa-miR-7-1-3p | 12 | ITGA1 | Sponge network | -2.039 | 0 | -0.774 | 0.0005 | 0.42 |
| 73 | LINC00702 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-362-5p;hsa-miR-450b-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-629-5p | 16 | IGF1 | Sponge network | -2.856 | 0 | -0.879 | 0.00545 | 0.42 |
| 74 | AC003991.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-7-1-3p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | COL4A3 | Sponge network | -0.787 | 0.08132 | -2.369 | 0 | 0.418 |
| 75 | RP11-1024P17.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-629-3p | 15 | PIK3R1 | Sponge network | -2.062 | 0 | -1.285 | 0 | 0.418 |
| 76 | SH3RF3-AS1 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-503-5p;hsa-miR-590-5p | 13 | FGF2 | Sponge network | -1.583 | 0 | -2.545 | 0 | 0.417 |
| 77 | RP11-456K23.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p | 23 | PIK3R1 | Sponge network | -1.488 | 0 | -1.285 | 0 | 0.416 |
| 78 | WDFY3-AS2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-22-3p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-33a-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-942-5p | 19 | IL6R | Sponge network | -1.297 | 0 | -1.175 | 0 | 0.415 |
| 79 | RP11-354E11.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-96-5p | 17 | PIK3R1 | Sponge network | -2.138 | 0 | -1.285 | 0 | 0.41 |
| 80 | AC109642.1 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-629-5p | 12 | IGF1 | Sponge network | -2.791 | 0 | -0.879 | 0.00545 | 0.41 |
| 81 | CTD-2013N24.2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 22 | PDGFRA | Sponge network | -1.745 | 0 | -0.92 | 0.00039 | 0.41 |
| 82 | RP11-456K23.1 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 22 | PDGFRA | Sponge network | -1.488 | 0 | -0.92 | 0.00039 | 0.409 |
| 83 | RP11-401P9.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-590-5p;hsa-miR-93-5p | 14 | PIK3R1 | Sponge network | -3.04 | 0 | -1.285 | 0 | 0.408 |
| 84 | CTD-2013N24.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 18 | PIK3R1 | Sponge network | -1.745 | 0 | -1.285 | 0 | 0.407 |
| 85 | RP11-536K7.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-93-5p | 10 | PDGFRA | Sponge network | -1.239 | 5.0E-5 | -0.92 | 0.00039 | 0.407 |
| 86 | RP11-352D13.6 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-590-3p;hsa-miR-93-5p | 11 | IL6R | Sponge network | -4.634 | 0 | -1.175 | 0 | 0.406 |
| 87 | TBX5-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-324-3p;hsa-miR-450b-5p;hsa-miR-501-5p;hsa-miR-503-5p;hsa-miR-550a-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-877-5p | 26 | CCND2 | Sponge network | -2.108 | 0 | -1.641 | 0 | 0.404 |
| 88 | CTD-2269F5.1 |
hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-7-1-3p | 11 | THBS1 | Sponge network | -1.576 | 0.00334 | -0.931 | 0.0014 | 0.404 |
| 89 | AC011899.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-503-5p;hsa-miR-550a-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 19 | CCND2 | Sponge network | -2.611 | 0 | -1.641 | 0 | 0.404 |
| 90 | RP11-253E3.3 | hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-15a-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-200b-3p;hsa-miR-21-5p;hsa-miR-27b-5p;hsa-miR-29b-3p;hsa-miR-30e-5p;hsa-miR-660-5p | 11 | CDK6 | Sponge network | -2.404 | 0 | -0.635 | 0.01747 | 0.403 |
| 91 | RP11-401P9.4 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-590-5p;hsa-miR-93-3p | 10 | FGF2 | Sponge network | -3.04 | 0 | -2.545 | 0 | 0.402 |
| 92 | LINC00968 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-421;hsa-miR-590-3p;hsa-miR-590-5p | 14 | NTRK2 | Sponge network | -4.19 | 0 | -2.564 | 0 | 0.4 |
| 93 | TBX5-AS1 |
hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 15 | THBS1 | Sponge network | -2.108 | 0 | -0.931 | 0.0014 | 0.398 |
| 94 | LINC00968 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-96-5p | 27 | CCND2 | Sponge network | -4.19 | 0 | -1.641 | 0 | 0.396 |
| 95 | RP11-532F6.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-629-3p | 11 | PIK3R1 | Sponge network | -2.028 | 0 | -1.285 | 0 | 0.395 |
| 96 | AC109642.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-590-3p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | COL4A3 | Sponge network | -2.791 | 0 | -2.369 | 0 | 0.394 |
| 97 | RP11-720L2.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-93-5p | 12 | CCND2 | Sponge network | -2.305 | 0 | -1.641 | 0 | 0.394 |
| 98 | CTD-2013N24.2 |
hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 14 | THBS1 | Sponge network | -1.745 | 0 | -0.931 | 0.0014 | 0.393 |
| 99 | SH3RF3-AS1 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 10 | ITGA1 | Sponge network | -1.583 | 0 | -0.774 | 0.0005 | 0.393 |
| 100 | RP11-456K23.1 |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-589-3p;hsa-miR-590-5p | 14 | FGF2 | Sponge network | -1.488 | 0 | -2.545 | 0 | 0.393 |
| 101 | MIR497HG |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p;hsa-miR-96-5p | 19 | PIK3R1 | Sponge network | -2.142 | 0 | -1.285 | 0 | 0.391 |
| 102 | RP11-1024P17.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-33a-5p;hsa-miR-576-5p;hsa-miR-590-3p | 16 | IL6R | Sponge network | -2.062 | 0 | -1.175 | 0 | 0.389 |
| 103 | RP11-456K23.1 |
hsa-miR-106b-5p;hsa-miR-146b-5p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-29b-3p;hsa-miR-362-5p;hsa-miR-542-3p;hsa-miR-93-5p | 10 | AKT3 | Sponge network | -1.488 | 0 | -1.44 | 0 | 0.389 |
| 104 | LINC00702 |
hsa-miR-141-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-3127-5p;hsa-miR-590-3p;hsa-miR-671-5p;hsa-miR-766-3p;hsa-miR-92a-3p;hsa-miR-96-5p | 10 | GNG7 | Sponge network | -2.856 | 0 | -1.412 | 0 | 0.389 |
| 105 | RP11-720L2.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-93-5p | 10 | PDGFRA | Sponge network | -2.305 | 0 | -0.92 | 0.00039 | 0.388 |
| 106 | FENDRR |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-3p | 14 | NTRK2 | Sponge network | -4.222 | 0 | -2.564 | 0 | 0.387 |
| 107 | AC008268.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-93-5p | 10 | IL6R | Sponge network | -5.661 | 0 | -1.175 | 0 | 0.387 |
| 108 | RP11-476D10.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-590-3p | 10 | IL6R | Sponge network | -4.519 | 0 | -1.175 | 0 | 0.387 |
| 109 | MAGI2-AS3 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-362-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-629-5p | 15 | IGF1 | Sponge network | -1.892 | 0 | -0.879 | 0.00545 | 0.384 |
| 110 | LINC00702 |
hsa-miR-141-5p;hsa-miR-194-5p;hsa-miR-20a-3p;hsa-miR-29a-5p;hsa-miR-29b-1-5p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p | 11 | ITGA9 | Sponge network | -2.856 | 0 | -1.268 | 0.00011 | 0.383 |
| 111 | RP11-399O19.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-93-5p | 10 | PIK3R1 | Sponge network | -0.873 | 0.00072 | -1.285 | 0 | 0.38 |
| 112 | FGF14-AS2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-96-5p | 12 | PIK3R1 | Sponge network | -2.159 | 0 | -1.285 | 0 | 0.378 |
| 113 | MAGI2-AS3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-429;hsa-miR-501-5p;hsa-miR-503-5p;hsa-miR-590-3p;hsa-miR-660-5p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-5p | 27 | CCND2 | Sponge network | -1.892 | 0 | -1.641 | 0 | 0.378 |
| 114 | RP11-389C8.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-3p | 13 | NTRK2 | Sponge network | -2.039 | 0 | -2.564 | 0 | 0.377 |
| 115 | RP11-166D19.1 |
hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-93-3p | 11 | FGF2 | Sponge network | -0.582 | 0.05253 | -2.545 | 0 | 0.375 |
| 116 | HHIP-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-424-5p;hsa-miR-590-3p | 12 | PIK3R1 | Sponge network | -2.807 | 0 | -1.285 | 0 | 0.374 |
| 117 | SH3RF3-AS1 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-26b-5p;hsa-miR-29b-3p;hsa-miR-576-5p | 11 | IGF1 | Sponge network | -1.583 | 0 | -0.879 | 0.00545 | 0.374 |
| 118 | RP11-1024P17.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-429;hsa-miR-589-3p;hsa-miR-590-3p | 21 | CCND2 | Sponge network | -2.062 | 0 | -1.641 | 0 | 0.374 |
| 119 | LINC00092 |
hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-424-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-629-3p | 13 | PIK3R1 | Sponge network | -2.383 | 0 | -1.285 | 0 | 0.369 |
| 120 | LINC00702 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p | 17 | NTRK2 | Sponge network | -2.856 | 0 | -2.564 | 0 | 0.369 |
| 121 | LINC00702 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-29a-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-3p;hsa-miR-96-5p | 13 | FOXO3 | Sponge network | -2.856 | 0 | -0.895 | 0 | 0.369 |
| 122 | RP11-378A13.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 15 | PIK3R1 | Sponge network | -1.713 | 0 | -1.285 | 0 | 0.367 |
| 123 | RP11-1223D19.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-942-5p | 12 | IL6R | Sponge network | -0.862 | 0.05389 | -1.175 | 0 | 0.367 |
| 124 | RP11-1024P17.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-625-5p;hsa-miR-93-3p | 11 | COL4A3 | Sponge network | -2.062 | 0 | -2.369 | 0 | 0.366 |
| 125 | SH3RF3-AS1 |
hsa-miR-128-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-590-5p | 12 | PIK3R1 | Sponge network | -1.583 | 0 | -1.285 | 0 | 0.364 |
| 126 | RBPMS-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130a-3p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-93-5p | 11 | IL6R | Sponge network | -1.548 | 1.0E-5 | -1.175 | 0 | 0.364 |
| 127 | AC011899.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-590-3p | 12 | PIK3R1 | Sponge network | -2.611 | 0 | -1.285 | 0 | 0.364 |
| 128 | RP11-532F6.3 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-576-5p | 11 | PDGFRA | Sponge network | -2.028 | 0 | -0.92 | 0.00039 | 0.363 |
| 129 | RP11-389C8.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-942-5p | 16 | IL6R | Sponge network | -2.039 | 0 | -1.175 | 0 | 0.361 |
| 130 | RP11-1008C21.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-550a-5p;hsa-miR-590-3p | 17 | CCND2 | Sponge network | -1.826 | 3.0E-5 | -1.641 | 0 | 0.36 |
| 131 | AC007743.1 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-22-3p;hsa-miR-320b;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-942-5p | 14 | IL6R | Sponge network | -2.595 | 0 | -1.175 | 0 | 0.36 |
| 132 | RP11-284N8.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | CSF1 | Sponge network | -0.761 | 0.05061 | -0.65 | 0.00787 | 0.36 |
| 133 | RP11-284N8.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-183-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-331-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 20 | CCND2 | Sponge network | -0.761 | 0.05061 | -1.641 | 0 | 0.36 |
| 134 | AC003991.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130a-3p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-320b;hsa-miR-421;hsa-miR-576-5p;hsa-miR-93-5p | 13 | IL6R | Sponge network | -0.787 | 0.08132 | -1.175 | 0 | 0.359 |
| 135 | AC079630.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-424-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-93-5p | 12 | PIK3R1 | Sponge network | -3.758 | 0 | -1.285 | 0 | 0.359 |
| 136 | FENDRR |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-93-5p | 15 | IL6R | Sponge network | -4.222 | 0 | -1.175 | 0 | 0.358 |
| 137 | RP5-1042I8.7 |
hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p;hsa-miR-96-5p | 12 | PIK3R1 | Sponge network | -0.733 | 0.00018 | -1.285 | 0 | 0.358 |
| 138 | AC004947.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p | 13 | NTRK2 | Sponge network | -3.94 | 0 | -2.564 | 0 | 0.358 |
| 139 | AF131215.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-454-3p | 11 | NTRK2 | Sponge network | -2.09 | 0 | -2.564 | 0 | 0.358 |
| 140 | AC003090.1 |
hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p;hsa-miR-96-5p | 14 | PIK3R1 | Sponge network | -3.16 | 2.0E-5 | -1.285 | 0 | 0.358 |
| 141 | CTD-2269F5.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 10 | PDGFRA | Sponge network | -1.576 | 0.00334 | -0.92 | 0.00039 | 0.358 |
| 142 | RP11-352D13.6 |
hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-18a-5p;hsa-miR-193b-5p;hsa-miR-590-3p;hsa-miR-671-5p;hsa-miR-766-3p;hsa-miR-9-5p;hsa-miR-940;hsa-miR-96-5p | 10 | GNG7 | Sponge network | -4.634 | 0 | -1.412 | 0 | 0.357 |
| 143 | AC109642.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-942-5p | 15 | IL6R | Sponge network | -2.791 | 0 | -1.175 | 0 | 0.355 |
| 144 | GAS6-AS2 |
hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 13 | THBS1 | Sponge network | -1.761 | 0 | -0.931 | 0.0014 | 0.355 |
| 145 | AC003090.1 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 12 | IL6R | Sponge network | -3.16 | 2.0E-5 | -1.175 | 0 | 0.355 |
| 146 | RP11-354E11.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-576-5p;hsa-miR-590-3p | 12 | NTRK2 | Sponge network | -2.138 | 0 | -2.564 | 0 | 0.355 |
| 147 | RP11-389C8.2 |
hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-29a-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-93-5p | 10 | FLT1 | Sponge network | -2.039 | 0 | -0.854 | 4.0E-5 | 0.354 |
| 148 | LINC00702 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-3p;hsa-miR-200b-5p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 14 | ITGA1 | Sponge network | -2.856 | 0 | -0.774 | 0.0005 | 0.354 |
| 149 | GAS6-AS2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-590-5p | 15 | PIK3R1 | Sponge network | -1.761 | 0 | -1.285 | 0 | 0.354 |
| 150 | TBX5-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-942-5p | 17 | IL6R | Sponge network | -2.108 | 0 | -1.175 | 0 | 0.354 |
| 151 | CTD-2013N24.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-501-5p;hsa-miR-590-3p;hsa-miR-660-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 24 | CCND2 | Sponge network | -1.745 | 0 | -1.641 | 0 | 0.354 |
| 152 | AC011526.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | CCND2 | Sponge network | -2.783 | 0 | -1.641 | 0 | 0.352 |
| 153 | RP11-462G12.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-7-1-3p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | COL4A3 | Sponge network | -1.071 | 0.01175 | -2.369 | 0 | 0.352 |
| 154 | LINC00886 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-421;hsa-miR-590-5p;hsa-miR-708-3p;hsa-miR-93-5p;hsa-miR-942-5p | 11 | IL6R | Sponge network | -0.244 | 0.45423 | -1.175 | 0 | 0.35 |
| 155 | BAIAP2-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-320b;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-942-5p | 13 | IL6R | Sponge network | -0.182 | 0.51705 | -1.175 | 0 | 0.349 |
| 156 | CTD-2006K23.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-22-3p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-942-5p | 13 | IL6R | Sponge network | -2.286 | 0.05883 | -1.175 | 0 | 0.349 |
| 157 | PCED1B-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-93-5p;hsa-miR-96-5p | 13 | PDGFRA | Sponge network | -0.672 | 0.02084 | -0.92 | 0.00039 | 0.348 |
| 158 | RP11-389C8.2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-429;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 19 | PDGFRA | Sponge network | -2.039 | 0 | -0.92 | 0.00039 | 0.347 |
| 159 | LINC00961 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-424-5p;hsa-miR-589-3p;hsa-miR-96-5p | 12 | PIK3R1 | Sponge network | -2.724 | 0 | -1.285 | 0 | 0.346 |
| 160 | RP11-389C8.2 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-629-5p | 11 | IGF1 | Sponge network | -2.039 | 0 | -0.879 | 0.00545 | 0.345 |
| 161 | CTD-2003C8.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-550a-5p | 11 | CCND2 | Sponge network | -3.403 | 0 | -1.641 | 0 | 0.345 |
| 162 | AC109642.1 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-590-3p | 11 | NTRK2 | Sponge network | -2.791 | 0 | -2.564 | 0 | 0.344 |
| 163 | TBX5-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-7-1-3p;hsa-miR-93-3p | 10 | COL4A3 | Sponge network | -2.108 | 0 | -2.369 | 0 | 0.343 |
| 164 | RP11-677M14.3 | hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-22-3p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-708-3p;hsa-miR-942-5p | 11 | IL6R | Sponge network | -1.866 | 0 | -1.175 | 0 | 0.343 |
| 165 | RP11-23P13.6 | hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-20a-5p;hsa-miR-324-3p;hsa-miR-331-5p;hsa-miR-590-3p;hsa-miR-877-5p | 11 | CCND2 | Sponge network | -0.705 | 0.00072 | -1.641 | 0 | 0.34 |
| 166 | AC109642.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-503-5p;hsa-miR-550a-5p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-96-5p | 24 | CCND2 | Sponge network | -2.791 | 0 | -1.641 | 0 | 0.339 |
| 167 | AC011526.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | PIK3R1 | Sponge network | -2.783 | 0 | -1.285 | 0 | 0.338 |
| 168 | AC079630.4 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-182-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-3p | 11 | NTRK2 | Sponge network | -3.758 | 0 | -2.564 | 0 | 0.338 |
| 169 | RP11-1024P17.1 |
hsa-miR-142-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-589-3p;hsa-miR-93-3p | 13 | FGF2 | Sponge network | -2.062 | 0 | -2.545 | 0 | 0.338 |
| 170 | RP11-401P9.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-550a-5p;hsa-miR-590-5p;hsa-miR-93-5p | 19 | CCND2 | Sponge network | -3.04 | 0 | -1.641 | 0 | 0.337 |
| 171 | GAS6-AS2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-550a-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 23 | CCND2 | Sponge network | -1.761 | 0 | -1.641 | 0 | 0.337 |
| 172 | RP11-354E11.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-96-5p | 17 | CCND2 | Sponge network | -2.138 | 0 | -1.641 | 0 | 0.335 |
| 173 | MAGI2-AS3 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-335-3p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p | 16 | NTRK2 | Sponge network | -1.892 | 0 | -2.564 | 0 | 0.333 |
| 174 | LINC00261 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-10a-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-3p | 10 | COL4A3 | Sponge network | -2.566 | 0.00025 | -2.369 | 0 | 0.333 |
| 175 | AC004947.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-450b-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p | 15 | PIK3R1 | Sponge network | -3.94 | 0 | -1.285 | 0 | 0.332 |
| 176 | MIR497HG |
hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-590-5p;hsa-miR-671-5p;hsa-miR-7-1-3p | 13 | THBS1 | Sponge network | -2.142 | 0 | -0.931 | 0.0014 | 0.33 |
| 177 | WDFY3-AS2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-96-5p | 17 | PIK3R1 | Sponge network | -1.297 | 0 | -1.285 | 0 | 0.329 |
| 178 | CASC2 |
hsa-miR-130b-3p;hsa-miR-183-5p;hsa-miR-19b-1-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 10 | ITGA8 | Sponge network | -1.086 | 0 | -3.151 | 0 | 0.329 |
| 179 | SNHG18 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-20b-5p;hsa-miR-330-5p;hsa-miR-590-3p;hsa-miR-625-5p;hsa-miR-93-5p | 10 | COL4A3 | Sponge network | -1.073 | 0.00533 | -2.369 | 0 | 0.328 |
| 180 | RP11-1024P17.1 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-576-5p;hsa-miR-590-3p | 10 | ITGA1 | Sponge network | -2.062 | 0 | -0.774 | 0.0005 | 0.328 |
| 181 | AC109642.1 |
hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-32-5p;hsa-miR-590-3p;hsa-miR-92a-3p | 13 | THBS1 | Sponge network | -2.791 | 0 | -0.931 | 0.0014 | 0.328 |
| 182 | RP11-354E11.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-576-5p;hsa-miR-590-3p | 14 | IL6R | Sponge network | -2.138 | 0 | -1.175 | 0 | 0.327 |
| 183 | LINC00472 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-301a-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-550a-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 18 | CCND2 | Sponge network | -2.952 | 0 | -1.641 | 0 | 0.327 |
| 184 | AC144831.1 | hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-324-3p;hsa-miR-550a-5p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-5p | 14 | CCND2 | Sponge network | -2.063 | 0 | -1.641 | 0 | 0.327 |
| 185 | MAGI2-AS3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-429;hsa-miR-93-5p | 10 | CSF1 | Sponge network | -1.892 | 0 | -0.65 | 0.00787 | 0.326 |
| 186 | CASC2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p | 12 | NTRK2 | Sponge network | -1.086 | 0 | -2.564 | 0 | 0.326 |
| 187 | RP11-1024P17.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-301a-3p | 15 | PTEN | Sponge network | -2.062 | 0 | -0.419 | 0.00014 | 0.325 |
| 188 | MIR497HG |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-5p | 10 | NTRK2 | Sponge network | -2.142 | 0 | -2.564 | 0 | 0.325 |
| 189 | MIR497HG |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-625-5p;hsa-miR-7-1-3p;hsa-miR-93-3p;hsa-miR-93-5p | 11 | COL4A3 | Sponge network | -2.142 | 0 | -2.369 | 0 | 0.323 |
| 190 | CTD-2013N24.2 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-362-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p | 14 | IGF1 | Sponge network | -1.745 | 0 | -0.879 | 0.00545 | 0.323 |
| 191 | RP11-354E11.2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-96-5p | 16 | PDGFRA | Sponge network | -2.138 | 0 | -0.92 | 0.00039 | 0.323 |
| 192 | RP11-166D19.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-324-3p;hsa-miR-429;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-96-5p | 22 | CCND2 | Sponge network | -0.582 | 0.05253 | -1.641 | 0 | 0.322 |
| 193 | LINC00092 |
hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-503-5p;hsa-miR-589-3p;hsa-miR-590-3p | 14 | CCND2 | Sponge network | -2.383 | 0 | -1.641 | 0 | 0.321 |
| 194 | RP11-166D19.1 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-3p;hsa-miR-200b-5p;hsa-miR-200c-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-5p;hsa-miR-7-1-3p | 12 | ITGA1 | Sponge network | -0.582 | 0.05253 | -0.774 | 0.0005 | 0.321 |
| 195 | CTC-559E9.4 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-942-5p | 10 | IL6R | Sponge network | -0.737 | 0.0562 | -1.175 | 0 | 0.321 |
| 196 | MIR497HG |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-942-5p | 15 | IL6R | Sponge network | -2.142 | 0 | -1.175 | 0 | 0.321 |
| 197 | RP11-401P9.4 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-93-5p | 16 | PDGFRA | Sponge network | -3.04 | 0 | -0.92 | 0.00039 | 0.32 |
| 198 | LINC00702 |
hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-26b-3p;hsa-miR-576-5p;hsa-miR-7-1-3p | 10 | CREB5 | Sponge network | -2.856 | 0 | -0.937 | 0.00202 | 0.319 |
| 199 | RP11-166D19.1 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-629-5p | 10 | IGF1 | Sponge network | -0.582 | 0.05253 | -0.879 | 0.00545 | 0.319 |
| 200 | RP1-78O14.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-33a-5p | 10 | IL6R | Sponge network | -4.409 | 0 | -1.175 | 0 | 0.319 |
| 201 | NR2F1-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 17 | PIK3R1 | Sponge network | -0.427 | 0.1559 | -1.285 | 0 | 0.318 |
| 202 | NR2F1-AS1 |
hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-32-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-671-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 12 | THBS1 | Sponge network | -0.427 | 0.1559 | -0.931 | 0.0014 | 0.317 |
| 203 | RP5-1042I8.7 |
hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-20a-5p;hsa-miR-320b;hsa-miR-331-5p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-96-5p | 13 | CCND2 | Sponge network | -0.733 | 0.00018 | -1.641 | 0 | 0.316 |
| 204 | LINC00607 |
hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-582-5p;hsa-miR-590-5p;hsa-miR-629-3p | 10 | PIK3R1 | Sponge network | -2.277 | 0 | -1.285 | 0 | 0.316 |
| 205 | AF131215.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-454-3p | 11 | IL6R | Sponge network | -2.09 | 0 | -1.175 | 0 | 0.315 |
| 206 | AC011899.9 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-26b-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 15 | PDGFRA | Sponge network | -2.611 | 0 | -0.92 | 0.00039 | 0.315 |
| 207 | AC079630.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-331-5p;hsa-miR-550a-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-5p | 19 | CCND2 | Sponge network | -3.758 | 0 | -1.641 | 0 | 0.314 |
| 208 | RP5-1120P11.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-30a-3p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-330-3p;hsa-miR-338-3p | 10 | COL1A1 | Sponge network | 1.924 | 1.0E-5 | 2.421 | 0 | 0.314 |
| 209 | RP11-354E11.2 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-301a-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-629-5p | 11 | IGF1 | Sponge network | -2.138 | 0 | -0.879 | 0.00545 | 0.313 |
| 210 | LINC00702 |
hsa-miR-103a-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-425-5p;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-93-5p | 21 | PTEN | Sponge network | -2.856 | 0 | -0.419 | 0.00014 | 0.312 |
| 211 | RP11-736K20.5 |
hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-16-1-3p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 11 | PDGFRA | Sponge network | -1.84 | 0 | -0.92 | 0.00039 | 0.311 |
| 212 | RP11-399O19.9 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-429;hsa-miR-576-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 14 | PDGFRA | Sponge network | -0.873 | 0.00072 | -0.92 | 0.00039 | 0.31 |
| 213 | AC011899.9 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 11 | NTRK2 | Sponge network | -2.611 | 0 | -2.564 | 0 | 0.31 |
| 214 | RP11-352D13.6 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-3p | 10 | NTRK2 | Sponge network | -4.634 | 0 | -2.564 | 0 | 0.31 |
| 215 | PSMG3-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-421;hsa-miR-590-3p | 10 | IL6R | Sponge network | -0.522 | 0.09584 | -1.175 | 0 | 0.31 |
| 216 | AC093627.10 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-421;hsa-miR-590-5p;hsa-miR-942-5p | 11 | IL6R | Sponge network | -0.174 | 0.61136 | -1.175 | 0 | 0.309 |
| 217 | RP11-389C8.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 11 | COL4A3 | Sponge network | -2.039 | 0 | -2.369 | 0 | 0.308 |
| 218 | AF131215.9 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-590-3p | 11 | NTRK2 | Sponge network | -1.808 | 0 | -2.564 | 0 | 0.307 |
| 219 | DIO3OS |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-301a-3p;hsa-miR-324-3p;hsa-miR-331-5p;hsa-miR-550a-5p;hsa-miR-877-5p | 14 | CCND2 | Sponge network | -1.936 | 0.00085 | -1.641 | 0 | 0.307 |
| 220 | MAGI2-AS3 |
hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-26b-3p;hsa-miR-576-5p;hsa-miR-7-1-3p | 10 | CREB5 | Sponge network | -1.892 | 0 | -0.937 | 0.00202 | 0.305 |
| 221 | RP11-283G6.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-450b-5p;hsa-miR-93-5p;hsa-miR-96-5p | 12 | CCND2 | Sponge network | -3.669 | 1.0E-5 | -1.641 | 0 | 0.305 |
| 222 | CTC-366B18.4 | hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-320b;hsa-miR-590-3p;hsa-miR-93-5p | 10 | CCND2 | Sponge network | -0.652 | 0.01265 | -1.641 | 0 | 0.303 |
| 223 | RP11-389C8.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 16 | PTEN | Sponge network | -2.039 | 0 | -0.419 | 0.00014 | 0.303 |
| 224 | CTA-221G9.11 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130a-3p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-93-5p | 11 | IL6R | Sponge network | -0.645 | 0.06404 | -1.175 | 0 | 0.303 |
| 225 | RP11-462G12.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-582-5p;hsa-miR-93-5p | 11 | PIK3R1 | Sponge network | -1.071 | 0.01175 | -1.285 | 0 | 0.302 |
| 226 | FENDRR |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-331-5p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-550a-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 22 | CCND2 | Sponge network | -4.222 | 0 | -1.641 | 0 | 0.301 |
| 227 | RP11-378A13.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-503-5p;hsa-miR-550a-5p;hsa-miR-590-3p;hsa-miR-877-5p;hsa-miR-93-5p | 20 | CCND2 | Sponge network | -1.713 | 0 | -1.641 | 0 | 0.301 |
| 228 | BAIAP2-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-365a-3p;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-942-5p | 12 | CCND1 | Sponge network | -0.182 | 0.51705 | -0.296 | 0.2554 | 0.3 |
| 229 | RP11-378A13.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | COL4A3 | Sponge network | -1.713 | 0 | -2.369 | 0 | 0.3 |
| 230 | TBX5-AS1 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-5p | 14 | NTRK2 | Sponge network | -2.108 | 0 | -2.564 | 0 | 0.299 |
| 231 | RP4-647J21.1 | hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-183-5p;hsa-miR-200a-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-550a-5p;hsa-miR-93-5p | 10 | CCND2 | Sponge network | -0.153 | 0.73575 | -1.641 | 0 | 0.299 |
| 232 | RP11-378A13.1 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-3p | 14 | NTRK2 | Sponge network | -1.713 | 0 | -2.564 | 0 | 0.298 |
| 233 | CTD-2008P7.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130a-3p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-3p;hsa-miR-301a-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-942-5p | 15 | IL6R | Sponge network | -1.912 | 1.0E-5 | -1.175 | 0 | 0.298 |
| 234 | AC016735.2 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-96-5p | 10 | PIK3R1 | Sponge network | -2.711 | 0.00282 | -1.285 | 0 | 0.298 |
| 235 | RP11-401P9.4 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-5p | 13 | NTRK2 | Sponge network | -3.04 | 0 | -2.564 | 0 | 0.298 |
| 236 | LINC00922 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-324-3p;hsa-miR-501-5p;hsa-miR-7-1-3p;hsa-miR-877-5p | 14 | CCND2 | Sponge network | -0.842 | 0.11239 | -1.641 | 0 | 0.298 |
| 237 | RP11-88I21.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-629-3p;hsa-miR-93-5p;hsa-miR-96-5p | 11 | PIK3R1 | Sponge network | -8.789 | 0 | -1.285 | 0 | 0.297 |
| 238 | RP11-1008C21.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-629-3p;hsa-miR-93-5p | 13 | PIK3R1 | Sponge network | -1.249 | 0 | -1.285 | 0 | 0.297 |
| 239 | RP11-400K9.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-93-5p | 11 | PDGFRA | Sponge network | -1.193 | 0.00359 | -0.92 | 0.00039 | 0.297 |
| 240 | AC011899.9 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-200a-5p;hsa-miR-200b-3p;hsa-miR-200b-5p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-7-1-3p | 10 | ITGA1 | Sponge network | -2.611 | 0 | -0.774 | 0.0005 | 0.297 |
| 241 | RP11-476D10.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-550a-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-96-5p | 19 | CCND2 | Sponge network | -4.519 | 0 | -1.641 | 0 | 0.296 |
| 242 | RP5-839B4.8 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-320b;hsa-miR-93-5p | 10 | IL6R | Sponge network | -5.037 | 0 | -1.175 | 0 | 0.296 |
| 243 | RP11-166D19.1 |
hsa-miR-103a-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-425-5p;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 19 | PTEN | Sponge network | -0.582 | 0.05253 | -0.419 | 0.00014 | 0.295 |
| 244 | RP11-416I2.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-769-3p;hsa-miR-92a-3p;hsa-miR-93-5p;hsa-miR-942-5p | 13 | CCND1 | Sponge network | 3.177 | 1.0E-5 | -0.296 | 0.2554 | 0.294 |
| 245 | RP11-365O16.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-93-5p | 11 | PIK3R1 | Sponge network | -2.765 | 0.00017 | -1.285 | 0 | 0.293 |
| 246 | RP11-456K23.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-550a-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 23 | CCND2 | Sponge network | -1.488 | 0 | -1.641 | 0 | 0.292 |
| 247 | WDFY3-AS2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-425-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 15 | PTEN | Sponge network | -1.297 | 0 | -0.419 | 0.00014 | 0.292 |
| 248 | RP11-456K23.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-625-5p;hsa-miR-93-5p | 10 | COL4A3 | Sponge network | -1.488 | 0 | -2.369 | 0 | 0.292 |
| 249 | LINC00961 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-589-3p;hsa-miR-96-5p | 16 | CCND2 | Sponge network | -2.724 | 0 | -1.641 | 0 | 0.292 |
| 250 | CASC2 |
hsa-miR-106a-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p | 13 | PIK3R1 | Sponge network | -1.086 | 0 | -1.285 | 0 | 0.29 |
| 251 | RP11-400K9.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-93-5p | 12 | CCND2 | Sponge network | -1.193 | 0.00359 | -1.641 | 0 | 0.29 |
| 252 | AC004947.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-301a-3p;hsa-miR-450b-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-5p | 17 | CCND2 | Sponge network | -3.94 | 0 | -1.641 | 0 | 0.289 |
| 253 | AC011899.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-590-3p;hsa-miR-708-3p | 13 | IL6R | Sponge network | -2.611 | 0 | -1.175 | 0 | 0.287 |
| 254 | AC006129.1 | hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-877-5p | 11 | CCND2 | Sponge network | -1.587 | 0.00086 | -1.641 | 0 | 0.287 |
| 255 | LINC00968 |
hsa-miR-130b-3p;hsa-miR-15b-3p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-3p;hsa-miR-301a-3p;hsa-miR-450b-5p;hsa-miR-590-3p | 11 | IGF1 | Sponge network | -4.19 | 0 | -0.879 | 0.00545 | 0.287 |
| 256 | GAS6-AS2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-5p | 13 | NTRK2 | Sponge network | -1.761 | 0 | -2.564 | 0 | 0.287 |
| 257 | SH3RF3-AS1 |
hsa-miR-15a-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-21-3p;hsa-miR-503-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 12 | CCND2 | Sponge network | -1.583 | 0 | -1.641 | 0 | 0.286 |
| 258 | CTD-2013N24.2 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-186-5p;hsa-miR-200a-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 12 | ITGA1 | Sponge network | -1.745 | 0 | -0.774 | 0.0005 | 0.286 |
| 259 | TBX5-AS1 |
hsa-miR-103a-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-92a-3p | 19 | PTEN | Sponge network | -2.108 | 0 | -0.419 | 0.00014 | 0.286 |
| 260 | LINC00261 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-942-5p | 15 | IL6R | Sponge network | -2.566 | 0.00025 | -1.175 | 0 | 0.285 |
| 261 | LINC00619 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-576-5p;hsa-miR-7-1-3p | 13 | PDGFRA | Sponge network | -2.307 | 0.02217 | -0.92 | 0.00039 | 0.285 |
| 262 | AF131215.2 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-589-3p | 11 | PDGFRA | Sponge network | -2.09 | 0 | -0.92 | 0.00039 | 0.285 |
| 263 | RP11-284N8.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-22-5p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-93-5p;hsa-miR-96-5p | 11 | PIK3R1 | Sponge network | -0.761 | 0.05061 | -1.285 | 0 | 0.285 |
| 264 | HLA-F-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-335-5p;hsa-miR-93-5p;hsa-miR-96-5p | 11 | CSF1 | Sponge network | -0.495 | 0.12126 | -0.65 | 0.00787 | 0.284 |
| 265 | RP11-1008C21.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-22-3p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-93-5p;hsa-miR-942-5p | 14 | IL6R | Sponge network | -1.249 | 0 | -1.175 | 0 | 0.283 |
| 266 | LINC00968 |
hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-5p;hsa-miR-200b-5p;hsa-miR-20a-5p;hsa-miR-29b-1-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-671-5p | 14 | THBS1 | Sponge network | -4.19 | 0 | -0.931 | 0.0014 | 0.283 |
| 267 | AF131215.9 |
hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-590-3p | 10 | PIK3R1 | Sponge network | -1.808 | 0 | -1.285 | 0 | 0.282 |
| 268 | AC004947.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | COL4A3 | Sponge network | -3.94 | 0 | -2.369 | 0 | 0.282 |
| 269 | HHIP-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-22-3p;hsa-miR-33a-5p;hsa-miR-590-3p | 12 | IL6R | Sponge network | -2.807 | 0 | -1.175 | 0 | 0.282 |
| 270 | RP11-532F6.3 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-576-5p | 11 | NTRK2 | Sponge network | -2.028 | 0 | -2.564 | 0 | 0.281 |
| 271 | RP11-352D13.6 |
hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p;hsa-miR-96-5p | 13 | PIK3R1 | Sponge network | -4.634 | 0 | -1.285 | 0 | 0.28 |
| 272 | LINC00473 | hsa-let-7i-5p;hsa-miR-146b-5p;hsa-miR-150-5p;hsa-miR-155-5p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-342-3p;hsa-miR-342-5p;hsa-miR-500a-3p;hsa-miR-532-5p | 10 | IGF1R | Sponge network | -0.765 | 0.56027 | -0.179 | 0.43653 | 0.28 |
| 273 | LIPE-AS1 |
hsa-miR-106a-5p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-29b-3p;hsa-miR-590-5p;hsa-miR-93-5p | 11 | PIK3R1 | Sponge network | -0.734 | 0.00039 | -1.285 | 0 | 0.28 |
| 274 | LINC00092 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-3p;hsa-miR-301a-3p;hsa-miR-590-3p;hsa-miR-942-5p | 12 | IL6R | Sponge network | -2.383 | 0 | -1.175 | 0 | 0.28 |
| 275 | CTC-297N7.9 | hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-590-5p | 10 | NTRK2 | Sponge network | -1.562 | 6.0E-5 | -2.564 | 0 | 0.28 |
| 276 | RP11-476D10.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-590-3p;hsa-miR-96-5p | 10 | PIK3R1 | Sponge network | -4.519 | 0 | -1.285 | 0 | 0.279 |
| 277 | RP11-284N8.3 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p;hsa-miR-96-5p | 14 | PDGFRA | Sponge network | -0.761 | 0.05061 | -0.92 | 0.00039 | 0.278 |
| 278 | TBX5-AS1 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-182-5p;hsa-miR-29a-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-93-3p | 10 | FOXO3 | Sponge network | -2.108 | 0 | -0.895 | 0 | 0.278 |
| 279 | BDNF-AS |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 10 | PIK3R1 | Sponge network | -0.568 | 0.02011 | -1.285 | 0 | 0.277 |
| 280 | LINC00443 |
hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p | 10 | PIK3R1 | Sponge network | -3.704 | 0.0003 | -1.285 | 0 | 0.276 |
| 281 | LINC00702 |
hsa-miR-106b-5p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19b-3p;hsa-miR-200b-3p;hsa-miR-20a-5p;hsa-miR-28-5p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-501-5p;hsa-miR-505-5p;hsa-miR-590-3p;hsa-miR-93-5p | 14 | FLT1 | Sponge network | -2.856 | 0 | -0.854 | 4.0E-5 | 0.276 |
| 282 | MIR22HG |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-5p;hsa-miR-708-3p;hsa-miR-93-5p;hsa-miR-942-5p | 11 | IL6R | Sponge network | -1.704 | 0 | -1.175 | 0 | 0.273 |
| 283 | AC010226.4 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-93-5p | 10 | CCND2 | Sponge network | -1.081 | 2.0E-5 | -1.641 | 0 | 0.272 |
| 284 | RP11-1008C21.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-629-3p | 12 | PIK3R1 | Sponge network | -1.826 | 3.0E-5 | -1.285 | 0 | 0.272 |
| 285 | MIR497HG |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-15a-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-301a-3p;hsa-miR-324-3p;hsa-miR-503-5p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-877-5p;hsa-miR-93-5p;hsa-miR-96-5p | 27 | CCND2 | Sponge network | -2.142 | 0 | -1.641 | 0 | 0.271 |
| 286 | MIR497HG |
hsa-miR-142-3p;hsa-miR-15a-5p;hsa-miR-16-1-3p;hsa-miR-186-5p;hsa-miR-19b-1-5p;hsa-miR-200c-3p;hsa-miR-20a-3p;hsa-miR-21-3p;hsa-miR-21-5p;hsa-miR-503-5p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-93-3p | 13 | FGF2 | Sponge network | -2.142 | 0 | -2.545 | 0 | 0.271 |
| 287 | SNHG18 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 11 | PIK3R1 | Sponge network | -1.073 | 0.00533 | -1.285 | 0 | 0.27 |
| 288 | DNM3OS |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-183-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-191-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-501-5p;hsa-miR-550a-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-93-5p | 16 | CCND2 | Sponge network | 0.053 | 0.85755 | -1.641 | 0 | 0.27 |
| 289 | NR2F1-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-20a-5p;hsa-miR-29b-3p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 12 | PDGFRA | Sponge network | -0.427 | 0.1559 | -0.92 | 0.00039 | 0.27 |
| 290 | C1orf132 |
hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-182-5p;hsa-miR-335-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 10 | PIK3R1 | Sponge network | -0.86 | 0.02429 | -1.285 | 0 | 0.269 |
| 291 | C1orf132 |
hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-151a-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-21-3p;hsa-miR-2355-3p;hsa-miR-28-5p;hsa-miR-320b;hsa-miR-450b-5p;hsa-miR-503-5p;hsa-miR-550a-5p;hsa-miR-590-3p;hsa-miR-877-5p;hsa-miR-93-5p | 15 | CCND2 | Sponge network | -0.86 | 0.02429 | -1.641 | 0 | 0.269 |
| 292 | MAGI2-AS3 |
hsa-miR-103a-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-425-5p;hsa-miR-454-3p;hsa-miR-7-1-3p;hsa-miR-92a-3p;hsa-miR-93-5p | 20 | PTEN | Sponge network | -1.892 | 0 | -0.419 | 0.00014 | 0.268 |
| 293 | RP11-365O16.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-576-5p;hsa-miR-708-3p;hsa-miR-93-5p | 14 | IL6R | Sponge network | -2.765 | 0.00017 | -1.175 | 0 | 0.267 |
| 294 | AC004947.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-421;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-708-3p;hsa-miR-93-5p | 16 | IL6R | Sponge network | -3.94 | 0 | -1.175 | 0 | 0.265 |
| 295 | RP11-399O19.9 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-7-1-3p;hsa-miR-93-5p | 13 | PTEN | Sponge network | -0.873 | 0.00072 | -0.419 | 0.00014 | 0.264 |
| 296 | RP11-20J15.3 | hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-5p;hsa-miR-17-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-550a-5p;hsa-miR-877-5p | 11 | CCND2 | Sponge network | -1.709 | 0.0268 | -1.641 | 0 | 0.264 |
| 297 | LINC00702 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-942-5p | 19 | IL6R | Sponge network | -2.856 | 0 | -1.175 | 0 | 0.264 |
| 298 | RP1-78O14.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-629-3p | 10 | PIK3R1 | Sponge network | -4.409 | 0 | -1.285 | 0 | 0.264 |
| 299 | RP11-378A13.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-708-3p;hsa-miR-93-5p | 17 | IL6R | Sponge network | -1.713 | 0 | -1.175 | 0 | 0.263 |
| 300 | AC022182.3 |
hsa-miR-106b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-2355-3p;hsa-miR-877-5p;hsa-miR-93-5p | 10 | CCND2 | Sponge network | -0.559 | 0.20451 | -1.641 | 0 | 0.263 |
| 301 | RP11-536K7.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-877-5p;hsa-miR-93-5p | 13 | CCND2 | Sponge network | -1.239 | 5.0E-5 | -1.641 | 0 | 0.263 |
| 302 | RP5-839B4.8 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-629-3p;hsa-miR-93-5p | 11 | PIK3R1 | Sponge network | -5.037 | 0 | -1.285 | 0 | 0.262 |
| 303 | DIO3OS |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-21-5p;hsa-miR-424-5p;hsa-miR-629-3p | 11 | PIK3R1 | Sponge network | -1.936 | 0.00085 | -1.285 | 0 | 0.262 |
| 304 | RP11-389C8.2 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-429;hsa-miR-93-5p | 10 | CSF1 | Sponge network | -2.039 | 0 | -0.65 | 0.00787 | 0.26 |
| 305 | RP11-456K23.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-93-5p | 16 | IL6R | Sponge network | -1.488 | 0 | -1.175 | 0 | 0.26 |
| 306 | LINC00261 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-629-3p | 16 | PIK3R1 | Sponge network | -2.566 | 0.00025 | -1.285 | 0 | 0.259 |
| 307 | GAS6-AS2 |
hsa-miR-107;hsa-miR-141-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-21-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-5p;hsa-miR-7-1-3p | 11 | ITGA1 | Sponge network | -1.761 | 0 | -0.774 | 0.0005 | 0.258 |
| 308 | LINC00961 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-22-3p;hsa-miR-301a-3p;hsa-miR-576-5p | 11 | IL6R | Sponge network | -2.724 | 0 | -1.175 | 0 | 0.258 |
| 309 | LL22NC03-86G7.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-141-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-20b-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-93-5p;hsa-miR-96-5p | 11 | PDGFRA | Sponge network | -1.177 | 3.0E-5 | -0.92 | 0.00039 | 0.257 |
| 310 | RP11-166D19.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-429;hsa-miR-96-5p | 10 | CSF1 | Sponge network | -0.582 | 0.05253 | -0.65 | 0.00787 | 0.257 |
| 311 | RP11-401P9.4 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-33a-5p;hsa-miR-421;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-708-3p;hsa-miR-93-5p | 16 | IL6R | Sponge network | -3.04 | 0 | -1.175 | 0 | 0.256 |
| 312 | AC003991.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-142-3p;hsa-miR-17-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-19b-1-5p;hsa-miR-20a-5p;hsa-miR-20b-5p;hsa-miR-365a-3p;hsa-miR-425-5p;hsa-miR-769-3p;hsa-miR-92a-3p;hsa-miR-93-5p | 14 | CCND1 | Sponge network | -0.787 | 0.08132 | -0.296 | 0.2554 | 0.255 |
| 313 | LINC00619 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p | 10 | PIK3R1 | Sponge network | -2.307 | 0.02217 | -1.285 | 0 | 0.255 |
| 314 | DNM3OS |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-186-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-590-5p;hsa-miR-93-5p | 13 | PIK3R1 | Sponge network | 0.053 | 0.85755 | -1.285 | 0 | 0.255 |
| 315 | LIPE-AS1 |
hsa-miR-103a-3p;hsa-miR-106a-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-425-5p;hsa-miR-590-5p;hsa-miR-93-5p | 13 | PTEN | Sponge network | -0.734 | 0.00039 | -0.419 | 0.00014 | 0.254 |
| 316 | AC011526.1 |
hsa-let-7g-3p;hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200a-3p;hsa-miR-20a-5p;hsa-miR-421;hsa-miR-450b-5p;hsa-miR-93-5p;hsa-miR-96-5p | 12 | PDGFRA | Sponge network | -2.783 | 0 | -0.92 | 0.00039 | 0.254 |
| 317 | CTA-221G9.11 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-22-5p;hsa-miR-335-3p;hsa-miR-590-3p;hsa-miR-629-3p;hsa-miR-93-5p | 12 | PIK3R1 | Sponge network | -0.645 | 0.06404 | -1.285 | 0 | 0.254 |
| 318 | PCED1B-AS1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-128-3p;hsa-miR-1301-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-200c-3p;hsa-miR-20a-5p;hsa-miR-335-3p;hsa-miR-93-5p;hsa-miR-96-5p | 11 | PIK3R1 | Sponge network | -0.672 | 0.02084 | -1.285 | 0 | 0.254 |
| 319 | RP11-284N8.3 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-22-3p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-33a-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-93-5p;hsa-miR-942-5p | 16 | IL6R | Sponge network | -0.761 | 0.05061 | -1.175 | 0 | 0.253 |
| 320 | RP11-53M11.3 | hsa-miR-130b-5p;hsa-miR-141-3p;hsa-miR-182-5p;hsa-miR-183-5p;hsa-miR-191-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-7-1-3p | 10 | CCND2 | Sponge network | -2.058 | 0.01204 | -1.641 | 0 | 0.253 |
| 321 | LINC00968 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-107;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-20a-5p;hsa-miR-93-5p;hsa-miR-96-5p | 10 | CSF1 | Sponge network | -4.19 | 0 | -0.65 | 0.00787 | 0.253 |
| 322 | LINC00607 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-21-5p;hsa-miR-320b;hsa-miR-590-5p;hsa-miR-708-3p;hsa-miR-942-5p | 10 | IL6R | Sponge network | -2.277 | 0 | -1.175 | 0 | 0.253 |
| 323 | LINC00472 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-141-5p;hsa-miR-17-5p;hsa-miR-193b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-93-5p | 11 | IL6R | Sponge network | -2.952 | 0 | -1.175 | 0 | 0.253 |
| 324 | CTD-2013N24.2 |
hsa-miR-130b-3p;hsa-miR-130b-5p;hsa-miR-151a-5p;hsa-miR-15b-3p;hsa-miR-17-3p;hsa-miR-182-5p;hsa-miR-20a-3p;hsa-miR-21-5p;hsa-miR-29a-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p | 14 | NTRK2 | Sponge network | -1.745 | 0 | -2.564 | 0 | 0.252 |