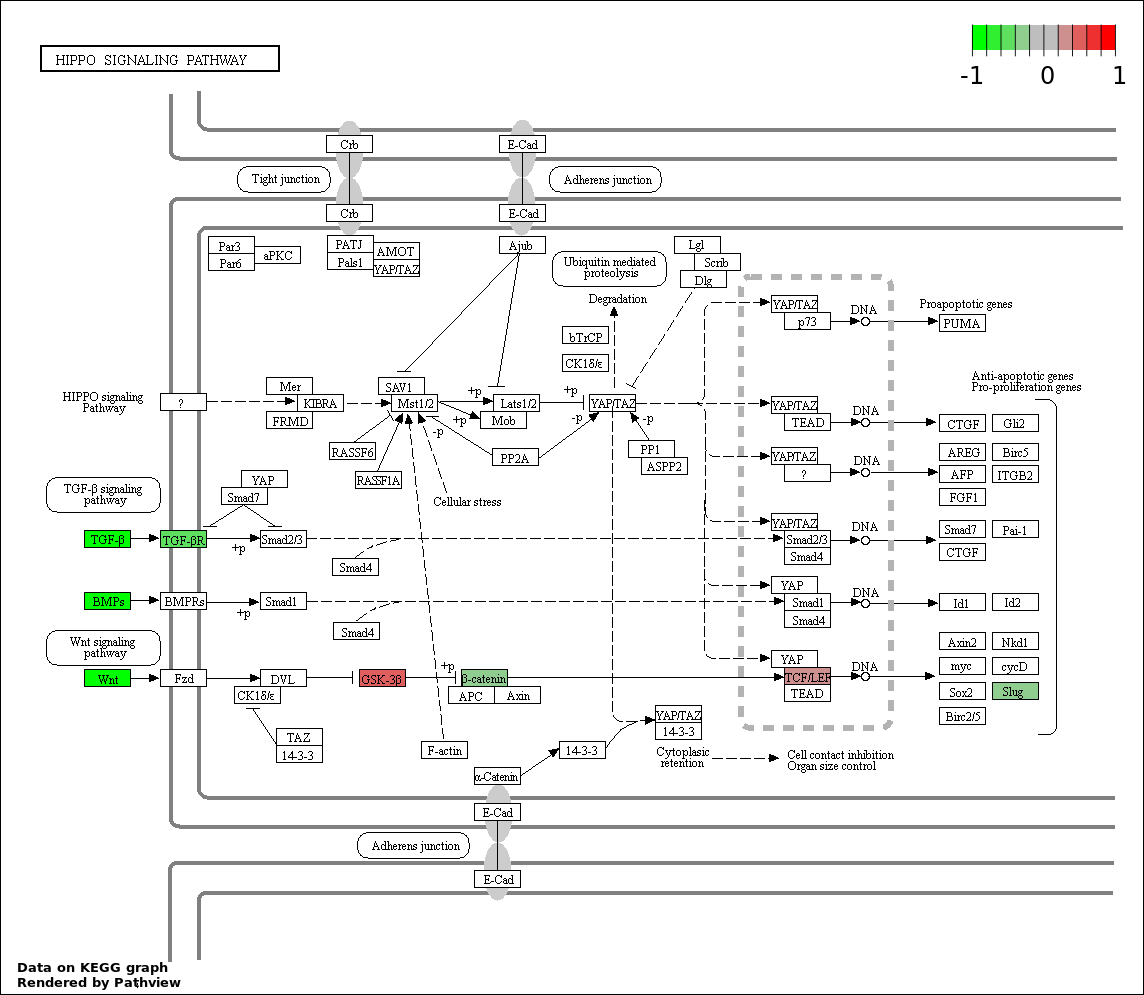
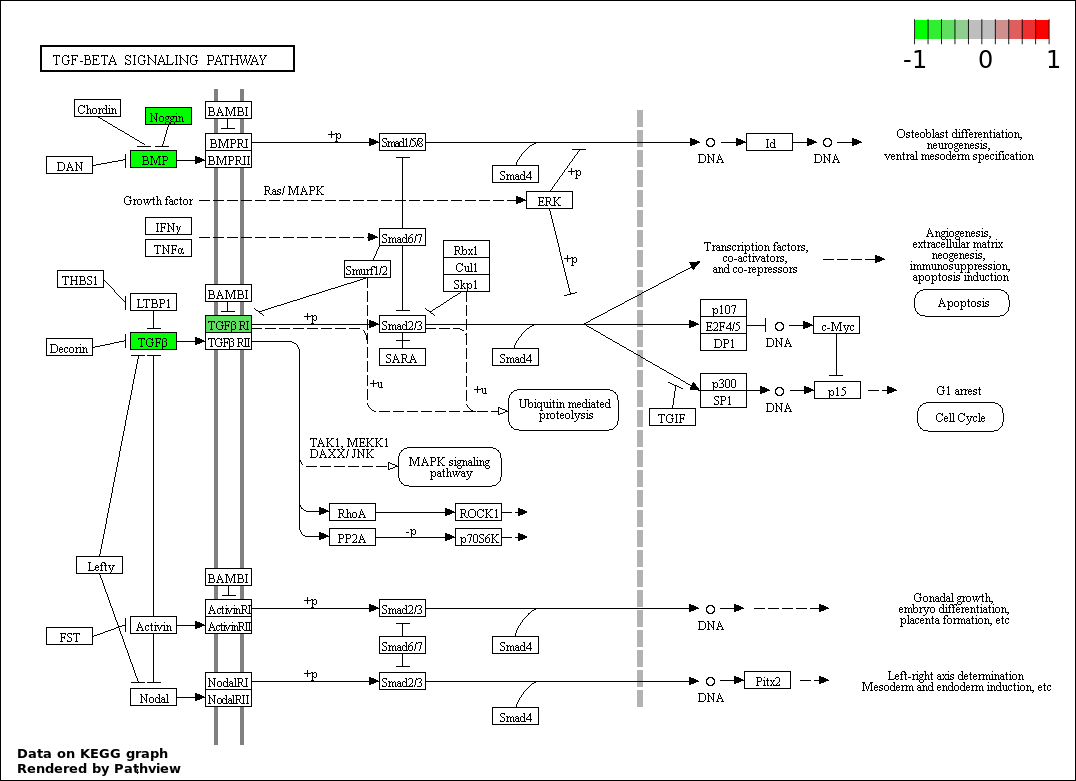
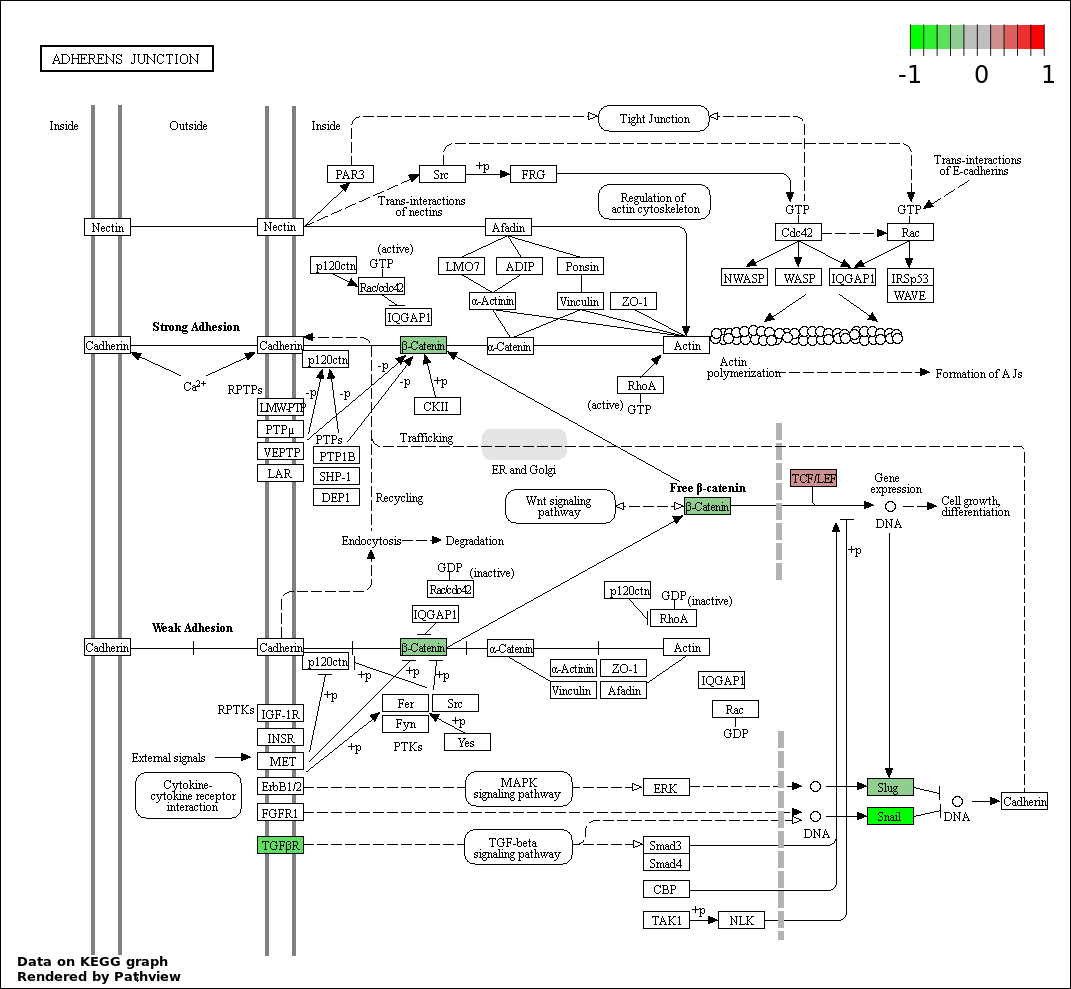
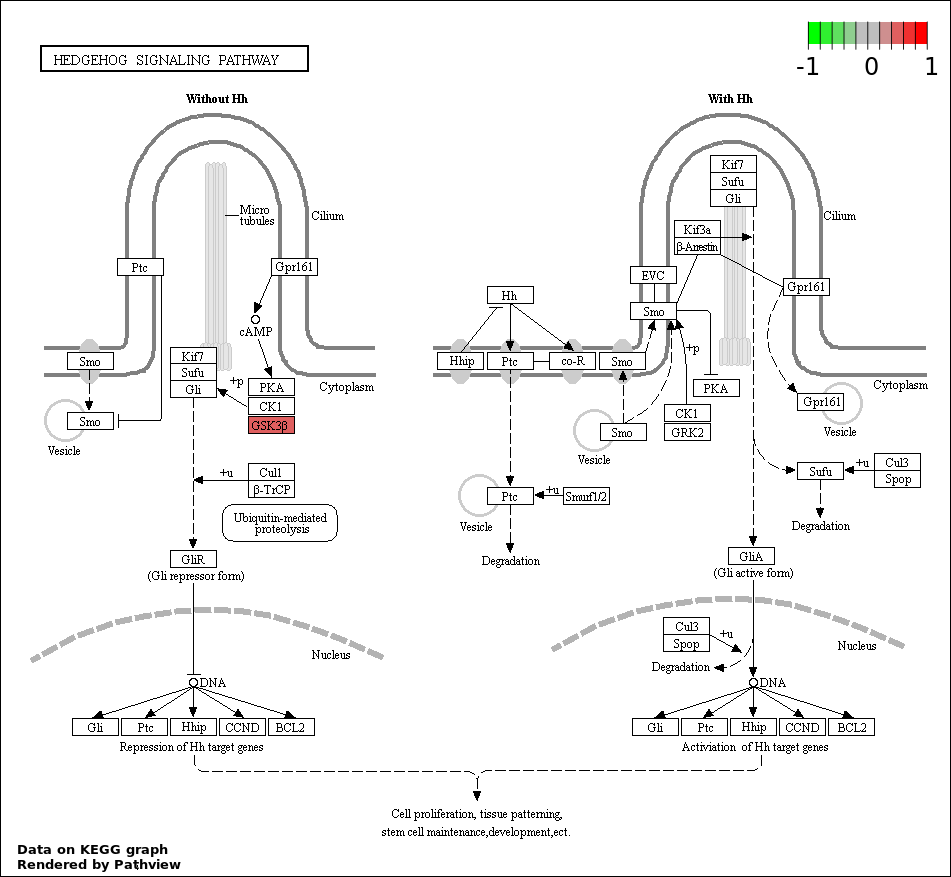
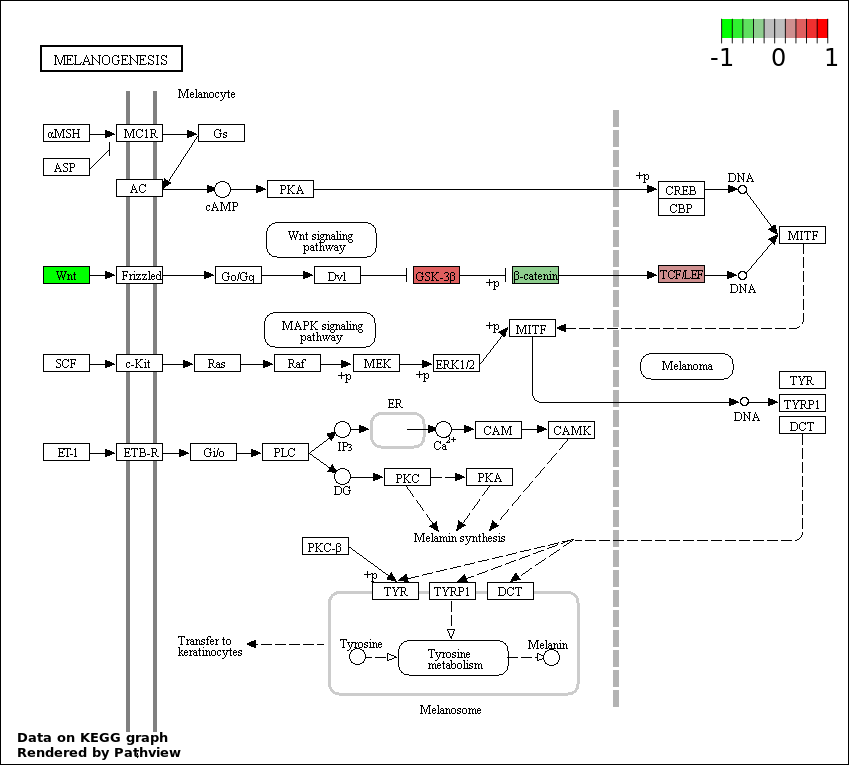
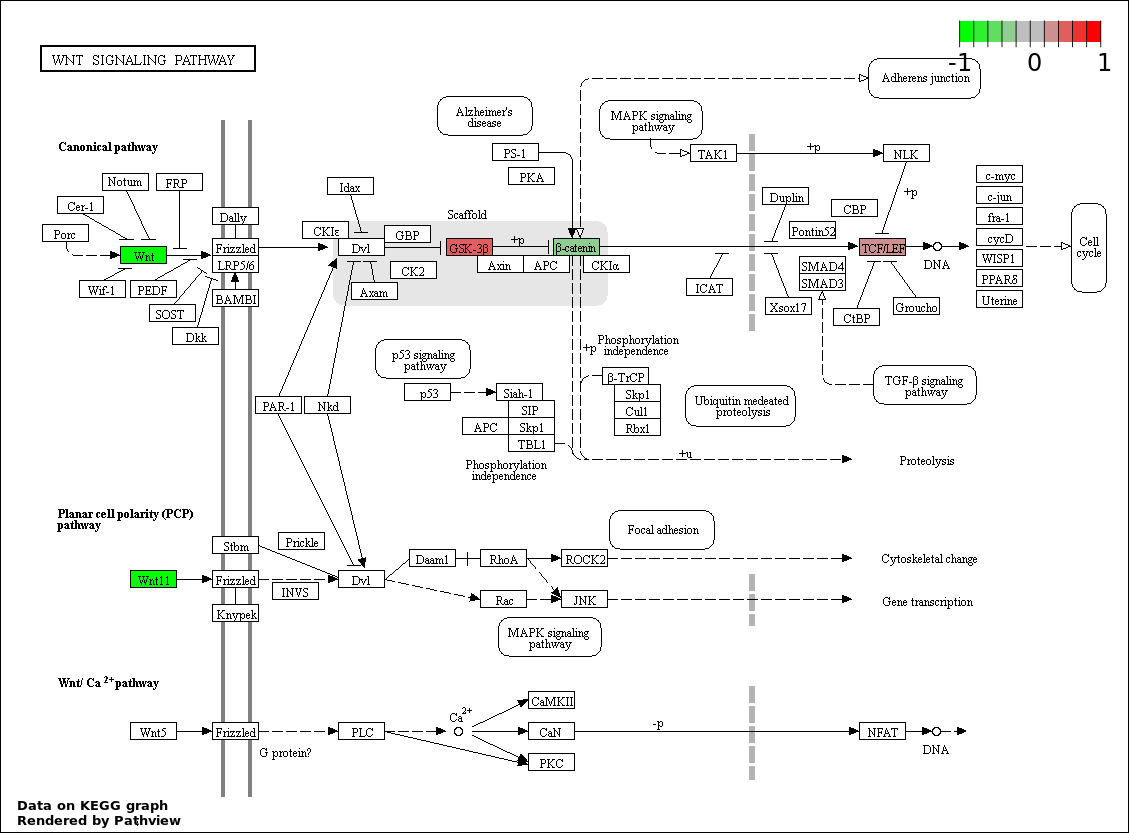
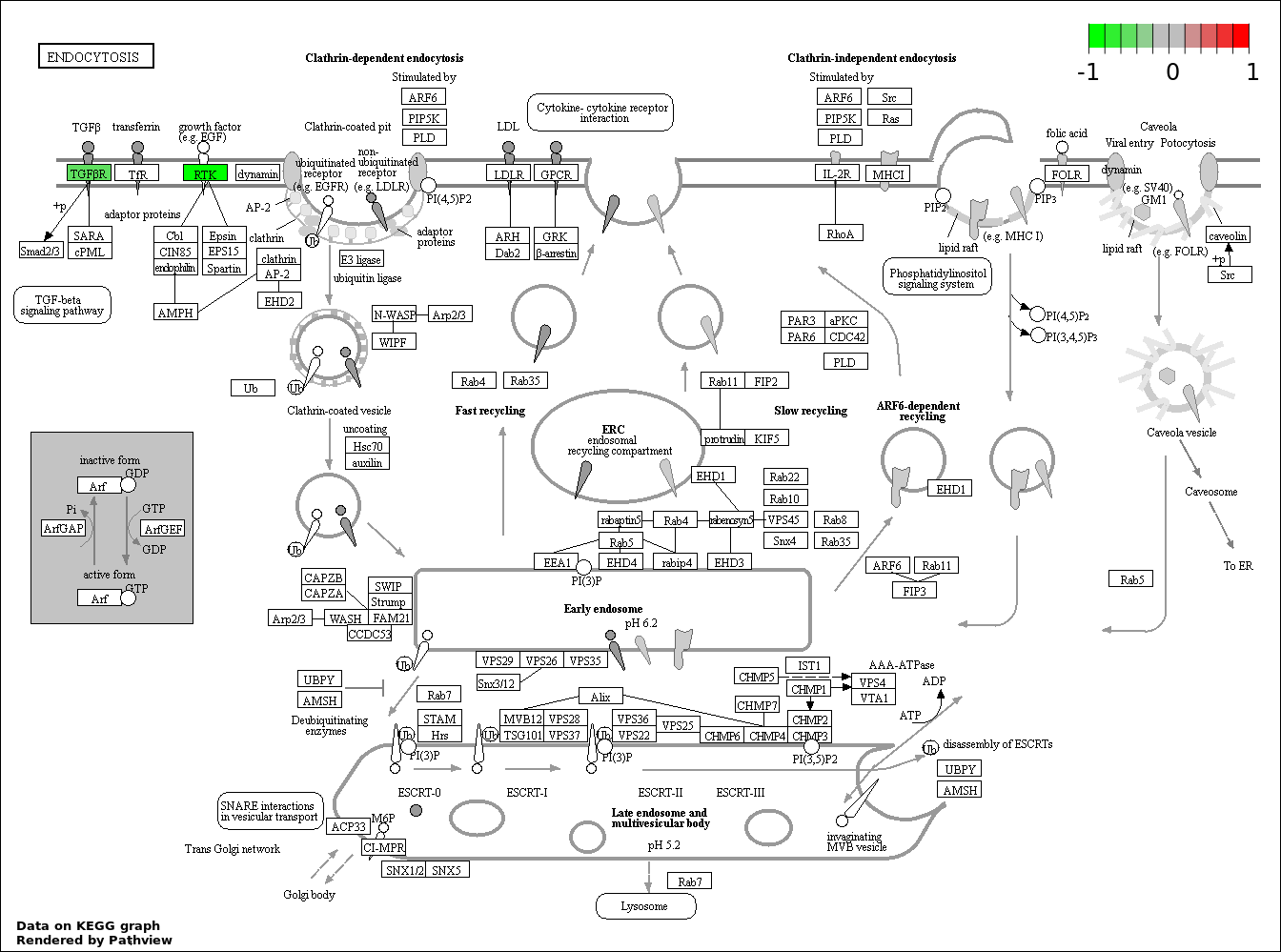
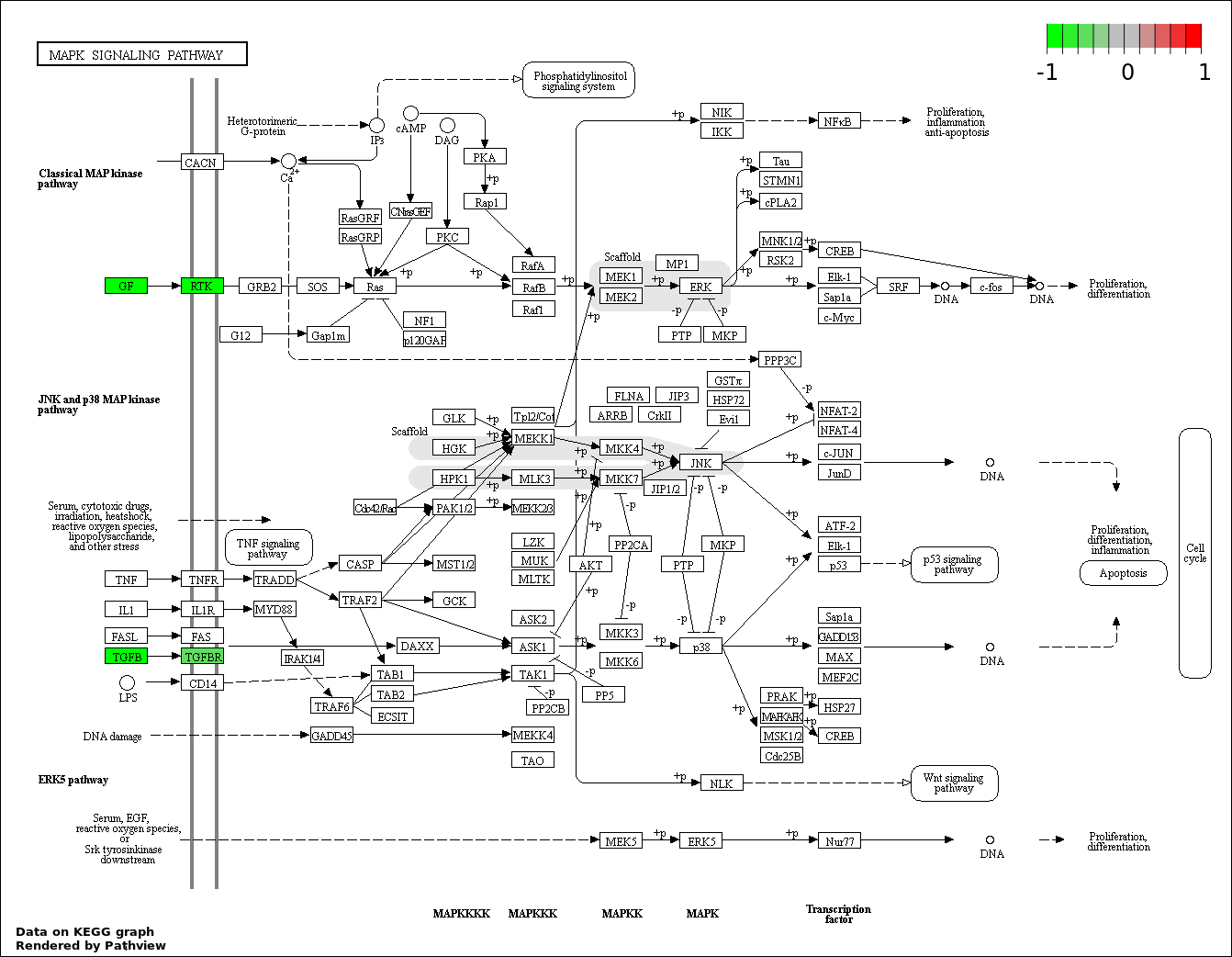
This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-106a-5p | BMP2 | 2.49 | 0 | -1.46 | 0.00276 | miRNATAP | -0.14 | 0.03534 | NA | |
| 2 | hsa-miR-17-5p | BMP2 | 3.27 | 0 | -1.46 | 0.00276 | TargetScan | -0.18 | 0.01393 | NA | |
| 3 | hsa-miR-20a-5p | BMP2 | 3.16 | 0 | -1.46 | 0.00276 | miRNATAP | -0.17 | 0.01894 | NA | |
| 4 | hsa-miR-3065-5p | BMP2 | 2.75 | 0 | -1.46 | 0.00276 | mirMAP | -0.27 | 0 | NA | |
| 5 | hsa-miR-335-3p | BMP2 | 3.09 | 0 | -1.46 | 0.00276 | mirMAP | -0.12 | 0.02404 | NA | |
| 6 | hsa-miR-106a-5p | BMP7 | 2.49 | 0 | -1.78 | 0.02876 | mirMAP | -0.21 | 0.04978 | NA | |
| 7 | hsa-miR-17-5p | BMP7 | 3.27 | 0 | -1.78 | 0.02876 | mirMAP | -0.3 | 0.01237 | NA | |
| 8 | hsa-miR-20b-5p | BMP7 | 2.08 | 1.0E-5 | -1.78 | 0.02876 | mirMAP | -0.17 | 0.04205 | NA | |
| 9 | hsa-miR-338-3p | BMP7 | 0.45 | 0.14458 | -1.78 | 0.02876 | miRanda | -0.41 | 0.00118 | NA | |
| 10 | hsa-miR-342-3p | BMP7 | 1.49 | 0 | -1.78 | 0.02876 | miRNAWalker2 validate; miRTarBase; miRanda | -0.56 | 4.0E-5 | NA | |
| 11 | hsa-miR-362-3p | BMP7 | 2.08 | 0 | -1.78 | 0.02876 | miRanda | -0.31 | 0.0207 | NA | |
| 12 | hsa-miR-450b-5p | BMP7 | 1.69 | 0 | -1.78 | 0.02876 | mirMAP | -0.31 | 0.03248 | NA | |
| 13 | hsa-miR-28-5p | CTNNB1 | 0.23 | 0.07429 | -0.22 | 0.11826 | miRanda | -0.18 | 0.00083 | NA | |
| 14 | hsa-miR-15a-5p | EPB41L5 | 2.35 | 0 | -0.61 | 0.00186 | MirTarget | -0.11 | 0.00453 | NA | |
| 15 | hsa-miR-26b-5p | EPB41L5 | 0.89 | 0 | -0.61 | 0.00186 | mirMAP | -0.1 | 0.04021 | NA | |
| 16 | hsa-miR-139-5p | FAM83D | -1.53 | 0 | 0.49 | 0.09144 | miRanda | -0.13 | 0.01377 | NA | |
| 17 | hsa-miR-429 | FAM83D | 4.49 | 0 | 0.49 | 0.09144 | miRanda | -0.13 | 0 | NA | |
| 18 | hsa-let-7a-3p | FGFR2 | 1.42 | 0 | -1.11 | 0.01288 | miRNATAP | -0.4 | 0.00015 | NA | |
| 19 | hsa-let-7b-3p | FGFR2 | 0.87 | 1.0E-5 | -1.11 | 0.01288 | mirMAP; miRNATAP | -0.31 | 0.00423 | NA | |
| 20 | hsa-miR-107 | FGFR2 | 1.31 | 0 | -1.11 | 0.01288 | PITA; miRanda | -0.4 | 0.00027 | NA | |
| 21 | hsa-miR-126-5p | FGFR2 | 0.9 | 2.0E-5 | -1.11 | 0.01288 | mirMAP; miRNATAP | -0.4 | 7.0E-5 | NA | |
| 22 | hsa-miR-146b-5p | FGFR2 | 1.76 | 0 | -1.11 | 0.01288 | miRanda | -0.15 | 0.02282 | NA | |
| 23 | hsa-miR-186-5p | FGFR2 | 1.47 | 0 | -1.11 | 0.01288 | miRNAWalker2 validate | -0.47 | 3.0E-5 | NA | |
| 24 | hsa-miR-193a-3p | FGFR2 | 1.6 | 0 | -1.11 | 0.01288 | miRanda | -0.18 | 0.0066 | NA | |
| 25 | hsa-miR-19b-1-5p | FGFR2 | 2.58 | 0 | -1.11 | 0.01288 | miRNAWalker2 validate; miRTarBase | -0.43 | 0 | NA | |
| 26 | hsa-miR-324-5p | FGFR2 | 2.96 | 0 | -1.11 | 0.01288 | miRanda | -0.25 | 0.00013 | NA | |
| 27 | hsa-miR-338-3p | FGFR2 | 0.45 | 0.14458 | -1.11 | 0.01288 | miRanda; miRNATAP | -0.26 | 0.00014 | NA | |
| 28 | hsa-miR-423-3p | FGFR2 | 2.58 | 0 | -1.11 | 0.01288 | PITA; miRNATAP | -0.23 | 0.00331 | NA | |
| 29 | hsa-miR-590-3p | FGFR2 | 2.59 | 0 | -1.11 | 0.01288 | miRanda | -0.31 | 6.0E-5 | NA | |
| 30 | hsa-miR-628-5p | FGFR2 | 0.8 | 2.0E-5 | -1.11 | 0.01288 | PITA; miRNATAP | -0.57 | 0 | NA | |
| 31 | hsa-miR-130b-3p | FOXF2 | 3.54 | 0 | -1.01 | 0.00891 | miRNATAP | -0.11 | 0.03859 | NA | |
| 32 | hsa-miR-139-5p | GSK3B | -1.53 | 0 | 0.41 | 0.00051 | mirMAP | -0.1 | 0 | NA | |
| 33 | hsa-let-7a-3p | HGF | 1.42 | 0 | -2.97 | 0 | mirMAP | -0.54 | 9.0E-5 | NA | |
| 34 | hsa-let-7b-3p | HGF | 0.87 | 1.0E-5 | -2.97 | 0 | mirMAP | -0.57 | 6.0E-5 | NA | |
| 35 | hsa-let-7f-1-3p | HGF | 1.55 | 0 | -2.97 | 0 | mirMAP | -0.34 | 0.00407 | NA | |
| 36 | hsa-miR-126-5p | HGF | 0.9 | 2.0E-5 | -2.97 | 0 | mirMAP | -0.3 | 0.02302 | NA | |
| 37 | hsa-miR-129-5p | HGF | -0.52 | 0.22258 | -2.97 | 0 | mirMAP | -0.18 | 0.00828 | NA | |
| 38 | hsa-miR-141-3p | HGF | 5.02 | 0 | -2.97 | 0 | MirTarget; TargetScan | -0.68 | 0 | NA | |
| 39 | hsa-miR-181a-5p | HGF | 2.3 | 0 | -2.97 | 0 | mirMAP | -0.73 | 0 | NA | |
| 40 | hsa-miR-181b-5p | HGF | 2.49 | 0 | -2.97 | 0 | mirMAP | -0.71 | 0 | NA | |
| 41 | hsa-miR-181c-5p | HGF | 1.59 | 0 | -2.97 | 0 | mirMAP | -0.66 | 0 | NA | |
| 42 | hsa-miR-200a-3p | HGF | 4.59 | 0 | -2.97 | 0 | MirTarget | -0.57 | 0 | NA | |
| 43 | hsa-miR-203a-3p | HGF | 3.12 | 0 | -2.97 | 0 | MirTarget | -0.31 | 0 | NA | |
| 44 | hsa-miR-224-3p | HGF | 1.88 | 6.0E-5 | -2.97 | 0 | mirMAP | -0.43 | 0 | NA | |
| 45 | hsa-miR-26b-5p | HGF | 0.89 | 0 | -2.97 | 0 | MirTarget; miRNATAP | -0.86 | 0 | NA | |
| 46 | hsa-miR-30b-5p | HGF | 0.8 | 0.00013 | -2.97 | 0 | mirMAP | -0.75 | 0 | NA | |
| 47 | hsa-miR-30c-5p | HGF | 0.78 | 0.00029 | -2.97 | 0 | mirMAP | -0.87 | 0 | NA | |
| 48 | hsa-miR-30d-5p | HGF | 0.68 | 0.00271 | -2.97 | 0 | mirMAP | -1.14 | 0 | NA | |
| 49 | hsa-miR-30e-5p | HGF | 1.24 | 0 | -2.97 | 0 | mirMAP | -1.43 | 0 | NA | |
| 50 | hsa-miR-335-3p | HGF | 3.09 | 0 | -2.97 | 0 | mirMAP | -0.3 | 0 | NA | |
| 51 | hsa-miR-33a-3p | HGF | 1.73 | 0 | -2.97 | 0 | mirMAP | -0.71 | 0 | NA | |
| 52 | hsa-miR-34c-5p | HGF | 2.07 | 0 | -2.97 | 0 | miRanda | -0.18 | 0.0199 | NA | |
| 53 | hsa-miR-421 | HGF | 1.18 | 1.0E-5 | -2.97 | 0 | miRanda | -0.42 | 7.0E-5 | NA | |
| 54 | hsa-miR-429 | HGF | 4.49 | 0 | -2.97 | 0 | miRanda | -0.52 | 0 | NA | |
| 55 | hsa-miR-582-5p | HGF | 0.61 | 0.03299 | -2.97 | 0 | PITA | -0.42 | 2.0E-5 | NA | |
| 56 | hsa-miR-590-3p | HGF | 2.59 | 0 | -2.97 | 0 | miRanda; mirMAP | -0.64 | 0 | NA | |
| 57 | hsa-miR-590-5p | HGF | 3.18 | 0 | -2.97 | 0 | miRanda | -0.71 | 0 | NA | |
| 58 | hsa-miR-651-5p | HGF | 2.33 | 0 | -2.97 | 0 | MirTarget | -0.88 | 0 | NA | |
| 59 | hsa-miR-7-1-3p | HGF | 1.85 | 0 | -2.97 | 0 | mirMAP | -0.58 | 0 | NA | |
| 60 | hsa-miR-944 | HGF | 2.91 | 0 | -2.97 | 0 | mirMAP | -0.41 | 0 | NA | |
| 61 | hsa-miR-106b-3p | HIF1A | 1.74 | 0 | -0.09 | 0.69445 | miRNAWalker2 validate | -0.12 | 0.01114 | NA | |
| 62 | hsa-miR-106b-5p | HIF1A | 2.47 | 0 | -0.09 | 0.69445 | MirTarget | -0.23 | 0 | NA | |
| 63 | hsa-miR-107 | HIF1A | 1.31 | 0 | -0.09 | 0.69445 | miRNAWalker2 validate; miRTarBase; miRanda | -0.12 | 0.02648 | NA | |
| 64 | hsa-miR-151a-3p | HIF1A | 1.24 | 0 | -0.09 | 0.69445 | MirTarget | -0.17 | 0.00034 | NA | |
| 65 | hsa-miR-20a-5p | HIF1A | 3.16 | 0 | -0.09 | 0.69445 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.12 | 0.0006 | 22901144 | Correlation analysis showed that the key miRNAs miR-20a and miR-20b negatively correlated with the target proteins VEGF-A and HIF-1alpha |
| 66 | hsa-miR-28-5p | HIF1A | 0.23 | 0.07429 | -0.09 | 0.69445 | miRanda | -0.41 | 0 | NA | |
| 67 | hsa-miR-330-3p | HIF1A | 0.88 | 0.00074 | -0.09 | 0.69445 | MirTarget; PITA | -0.1 | 0.01705 | NA | |
| 68 | hsa-miR-361-5p | HIF1A | 0.97 | 0 | -0.09 | 0.69445 | miRanda | -0.24 | 0.00023 | NA | |
| 69 | hsa-miR-660-5p | HIF1A | 1.83 | 0 | -0.09 | 0.69445 | MirTarget | -0.14 | 0.0012 | NA | |
| 70 | hsa-miR-93-5p | HIF1A | 3.04 | 0 | -0.09 | 0.69445 | MirTarget | -0.19 | 0 | NA | |
| 71 | hsa-let-7e-5p | HMGA2 | 0.86 | 6.0E-5 | 3.21 | 0.00037 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.73 | 0.00033 | NA | |
| 72 | hsa-let-7f-5p | HMGA2 | 0.5 | 0.00356 | 3.21 | 0.00037 | miRNAWalker2 validate; MirTarget | -1.1 | 2.0E-5 | NA | |
| 73 | hsa-let-7g-5p | HMGA2 | 1.2 | 0 | 3.21 | 0.00037 | miRNAWalker2 validate; miRTarBase; MirTarget | -1.43 | 0 | 21472347; 18308936 | Furthermore K-Ras and HMGA2 are well known as targets of let-7g; In this study we evaluated the potential role of precursor pre-let-7g in lung cancer cell metastasis focusing on the two targets of let-7g HMGA2 and K-Ras;In let-7g expressing tumors reductions in Ras family and HMGA2 protein levels were detected; Ectopic expression of K-RasG12D largely rescued let-7g mediated tumor suppression whereas ectopic expression of HMGA2 was less effective |
| 74 | hsa-miR-106b-5p | HMGA2 | 2.47 | 0 | 3.21 | 0.00037 | miRNATAP | -0.37 | 0.02815 | NA | |
| 75 | hsa-miR-141-5p | HMGA2 | 3.81 | 0 | 3.21 | 0.00037 | mirMAP | -0.39 | 5.0E-5 | NA | |
| 76 | hsa-miR-195-3p | HMGA2 | -0.66 | 0.01421 | 3.21 | 0.00037 | mirMAP | -0.35 | 0.03597 | NA | |
| 77 | hsa-miR-20b-5p | HMGA2 | 2.08 | 1.0E-5 | 3.21 | 0.00037 | miRNATAP | -0.3 | 0.00097 | NA | |
| 78 | hsa-miR-23b-3p | HMGA2 | -0.25 | 0.1502 | 3.21 | 0.00037 | mirMAP | -0.87 | 0.00068 | NA | |
| 79 | hsa-miR-29b-2-5p | HMGA2 | 0.91 | 0.00667 | 3.21 | 0.00037 | mirMAP | -1.28 | 0 | NA | |
| 80 | hsa-miR-30b-5p | HMGA2 | 0.8 | 0.00013 | 3.21 | 0.00037 | mirMAP | -0.73 | 0.0005 | NA | |
| 81 | hsa-miR-30c-5p | HMGA2 | 0.78 | 0.00029 | 3.21 | 0.00037 | mirMAP | -1.07 | 0 | NA | |
| 82 | hsa-miR-30d-3p | HMGA2 | 0.98 | 4.0E-5 | 3.21 | 0.00037 | MirTarget | -0.82 | 1.0E-5 | NA | |
| 83 | hsa-miR-30d-5p | HMGA2 | 0.68 | 0.00271 | 3.21 | 0.00037 | mirMAP | -1.48 | 0 | NA | |
| 84 | hsa-miR-30e-3p | HMGA2 | -0.22 | 0.23538 | 3.21 | 0.00037 | MirTarget | -1.74 | 0 | NA | |
| 85 | hsa-miR-30e-5p | HMGA2 | 1.24 | 0 | 3.21 | 0.00037 | mirMAP | -1.45 | 0 | NA | |
| 86 | hsa-miR-320a | HMGA2 | 0.44 | 0.03902 | 3.21 | 0.00037 | mirMAP | -0.6 | 0.00404 | NA | |
| 87 | hsa-miR-326 | HMGA2 | 0.72 | 0.06103 | 3.21 | 0.00037 | miRNATAP | -0.23 | 0.0476 | NA | |
| 88 | hsa-miR-330-3p | HMGA2 | 0.88 | 0.00074 | 3.21 | 0.00037 | mirMAP | -0.73 | 1.0E-5 | NA | |
| 89 | hsa-miR-34a-5p | HMGA2 | 1.9 | 0 | 3.21 | 0.00037 | miRNAWalker2 validate | -0.57 | 0.00093 | 18803879 | miR-34 targets Notch HMGA2 and Bcl-2 genes involved in the self-renewal and survival of cancer stem cells; Human gastric cancer Kato III cells with miR-34 restoration reduced the expression of target genes Bcl-2 Notch and HMGA2; The mechanism of miR-34-mediated suppression of self-renewal appears to be related to the direct modulation of downstream targets Bcl-2 Notch and HMGA2 indicating that miR-34 may be involved in gastric cancer stem cell self-renewal/differentiation decision-making |
| 90 | hsa-miR-361-5p | HMGA2 | 0.97 | 0 | 3.21 | 0.00037 | PITA; miRanda | -0.6 | 0.02366 | NA | |
| 91 | hsa-miR-497-5p | HMGA2 | 0.03 | 0.91815 | 3.21 | 0.00037 | MirTarget | -0.5 | 0.00468 | NA | |
| 92 | hsa-miR-508-3p | HMGA2 | 1.05 | 0.03939 | 3.21 | 0.00037 | PITA; miRNATAP | -0.25 | 0.00497 | NA | |
| 93 | hsa-miR-532-5p | HMGA2 | 1.56 | 0 | 3.21 | 0.00037 | mirMAP | -0.41 | 0.02799 | NA | |
| 94 | hsa-miR-664a-3p | HMGA2 | 0.63 | 0.0052 | 3.21 | 0.00037 | mirMAP | -1.44 | 0 | NA | |
| 95 | hsa-miR-664a-5p | HMGA2 | 0.51 | 0.03293 | 3.21 | 0.00037 | MirTarget | -1.31 | 0 | NA | |
| 96 | hsa-miR-93-3p | HMGA2 | 2.85 | 0 | 3.21 | 0.00037 | mirMAP | -0.28 | 0.04937 | NA | |
| 97 | hsa-miR-93-5p | HMGA2 | 3.04 | 0 | 3.21 | 0.00037 | miRNATAP | -0.34 | 0.0164 | NA | |
| 98 | hsa-miR-98-5p | HMGA2 | 1.11 | 0 | 3.21 | 0.00037 | miRNAWalker2 validate; miRTarBase; MirTarget | -1.25 | 0 | 17222355; 23700794 | Here we report that HMGA2 expression in head and neck squamous cell carcinoma HNSCC cells is regulated in part by miRNA-98 miR-98;MicroRNA 98 sensitizes cisplatin resistant human lung adenocarcinoma cells by up regulation of HMGA2; MiR-98 a member in the let-7 family acts as a negative regulator in the expression of HMGA2 high mobility group A2 oncogene and it has been shown to have a nearly 3-fold decrease in A549/DDP cells; We found that elevated expression of miR-98 led to a higher sensitivity of A549/DDP cells to cisplatin and the protein level of HMGA2 was clearly up-regulated in both A549/DDP and A549 cells by miR-98; We for the first time demonstrated that the expression of miR-98 increases cells spontaneous apoptosis and sensitizes cells to cisplatin at least in part via HMGA2 up-regulation |
| 99 | hsa-miR-26b-5p | LEF1 | 0.89 | 0 | 0.33 | 0.28979 | miRNATAP | -0.27 | 0.00054 | 24785257 | miR26b is downregulated in several cancers and tumors and miR26b directly targets the lymphoid enhancer factor 1 Lef13'UTR and inhibits endogenous Lef1 expression; We report that miR26b expression is associated with human colon cancer through the regulation of LEF1 expression in colon cancer cells; Analyses of multiple colon cancer cell lines revealed an inverse correlation between miR26b and LEF1 expression; Normal human colon cells express low levels of LEF1 and high levels of miR26b; however human colon cancer cells have decreased miR26b expression and increased LEF1 expression; We demonstrate that miR26b expression is a potent inhibitor of colon cancer cell proliferation and significantly decreases LEF1 expression; Analyses of human colon cancer databases also demonstrated a link between miR26b and LEF1 expression |
| 100 | hsa-miR-320a | LEF1 | 0.44 | 0.03902 | 0.33 | 0.28979 | miRanda; mirMAP | -0.15 | 0.03893 | NA | |
| 101 | hsa-miR-320b | LEF1 | 1.56 | 0 | 0.33 | 0.28979 | miRanda; mirMAP | -0.17 | 0.00218 | NA | |
| 102 | hsa-miR-33a-3p | LEF1 | 1.73 | 0 | 0.33 | 0.28979 | miRNATAP | -0.16 | 0.00117 | NA | |
| 103 | hsa-miR-1226-3p | LIMS1 | 1.38 | 0.00021 | -0.39 | 0.14814 | miRNAWalker2 validate | -0.11 | 0.00444 | NA | |
| 104 | hsa-miR-148b-5p | LIMS1 | 2.18 | 0 | -0.39 | 0.14814 | MirTarget | -0.21 | 1.0E-5 | NA | |
| 105 | hsa-miR-182-5p | LIMS1 | 3.54 | 0 | -0.39 | 0.14814 | MirTarget; miRNATAP | -0.12 | 0.00035 | NA | |
| 106 | hsa-miR-186-5p | LIMS1 | 1.47 | 0 | -0.39 | 0.14814 | mirMAP | -0.31 | 0 | NA | |
| 107 | hsa-miR-192-5p | LIMS1 | 2.69 | 0 | -0.39 | 0.14814 | miRNAWalker2 validate; MirTarget | -0.11 | 0.00394 | NA | |
| 108 | hsa-miR-29b-3p | LIMS1 | 1.66 | 0 | -0.39 | 0.14814 | MirTarget; miRNATAP | -0.24 | 0 | NA | |
| 109 | hsa-miR-29c-3p | LIMS1 | -0.01 | 0.971 | -0.39 | 0.14814 | MirTarget; miRNATAP | -0.21 | 3.0E-5 | NA | |
| 110 | hsa-miR-30b-5p | LIMS1 | 0.8 | 0.00013 | -0.39 | 0.14814 | MirTarget | -0.31 | 0 | NA | |
| 111 | hsa-miR-30c-5p | LIMS1 | 0.78 | 0.00029 | -0.39 | 0.14814 | MirTarget | -0.38 | 0 | NA | |
| 112 | hsa-miR-30d-3p | LIMS1 | 0.98 | 4.0E-5 | -0.39 | 0.14814 | mirMAP | -0.19 | 0.00052 | NA | |
| 113 | hsa-miR-30d-5p | LIMS1 | 0.68 | 0.00271 | -0.39 | 0.14814 | MirTarget | -0.2 | 0.00039 | NA | |
| 114 | hsa-miR-30e-5p | LIMS1 | 1.24 | 0 | -0.39 | 0.14814 | MirTarget | -0.34 | 0 | NA | |
| 115 | hsa-miR-32-3p | LIMS1 | 2.02 | 0 | -0.39 | 0.14814 | mirMAP | -0.11 | 0.00763 | NA | |
| 116 | hsa-miR-374a-3p | LIMS1 | 0.53 | 0.00193 | -0.39 | 0.14814 | mirMAP | -0.16 | 0.0377 | NA | |
| 117 | hsa-miR-429 | LIMS1 | 4.49 | 0 | -0.39 | 0.14814 | miRanda; miRNATAP | -0.12 | 0 | NA | |
| 118 | hsa-miR-505-3p | LIMS1 | 1.45 | 0 | -0.39 | 0.14814 | miRNAWalker2 validate; MirTarget | -0.11 | 0.04472 | NA | |
| 119 | hsa-miR-576-5p | LIMS1 | 2.2 | 0 | -0.39 | 0.14814 | mirMAP | -0.21 | 4.0E-5 | NA | |
| 120 | hsa-miR-590-3p | LIMS1 | 2.59 | 0 | -0.39 | 0.14814 | miRNAWalker2 validate; miRanda | -0.13 | 0.00616 | NA | |
| 121 | hsa-miR-96-5p | LIMS1 | 4.89 | 0 | -0.39 | 0.14814 | TargetScan; miRNATAP | -0.13 | 1.0E-5 | NA | |
| 122 | hsa-miR-192-5p | LOXL2 | 2.69 | 0 | 0.75 | 0.01695 | miRNAWalker2 validate | -0.18 | 6.0E-5 | NA | |
| 123 | hsa-miR-28-5p | LOXL2 | 0.23 | 0.07429 | 0.75 | 0.01695 | miRanda | -0.46 | 0.00018 | NA | |
| 124 | hsa-miR-361-5p | LOXL2 | 0.97 | 0 | 0.75 | 0.01695 | miRanda | -0.54 | 0 | NA | |
| 125 | hsa-let-7a-5p | LOXL3 | 0.62 | 3.0E-5 | -0.97 | 0.00081 | miRNATAP | -0.68 | 0 | NA | |
| 126 | hsa-let-7b-5p | LOXL3 | 0.6 | 0.0014 | -0.97 | 0.00081 | miRNATAP | -0.36 | 0 | NA | |
| 127 | hsa-let-7d-5p | LOXL3 | 1.33 | 0 | -0.97 | 0.00081 | miRNATAP | -0.3 | 0.00011 | NA | |
| 128 | hsa-let-7e-5p | LOXL3 | 0.86 | 6.0E-5 | -0.97 | 0.00081 | miRNATAP | -0.36 | 0 | NA | |
| 129 | hsa-let-7f-5p | LOXL3 | 0.5 | 0.00356 | -0.97 | 0.00081 | miRNATAP | -0.48 | 0 | NA | |
| 130 | hsa-let-7g-5p | LOXL3 | 1.2 | 0 | -0.97 | 0.00081 | miRNATAP | -0.59 | 0 | NA | |
| 131 | hsa-miR-125a-5p | LOXL3 | 0.6 | 0.01023 | -0.97 | 0.00081 | PITA; miRanda | -0.24 | 6.0E-5 | NA | |
| 132 | hsa-miR-2110 | LOXL3 | 1.17 | 6.0E-5 | -0.97 | 0.00081 | miRNATAP | -0.36 | 0 | NA | |
| 133 | hsa-miR-320a | LOXL3 | 0.44 | 0.03902 | -0.97 | 0.00081 | miRanda | -0.38 | 0 | NA | |
| 134 | hsa-miR-320b | LOXL3 | 1.56 | 0 | -0.97 | 0.00081 | miRanda | -0.24 | 0 | NA | |
| 135 | hsa-miR-34a-5p | LOXL3 | 1.9 | 0 | -0.97 | 0.00081 | MirTarget; miRNATAP | -0.4 | 0 | NA | |
| 136 | hsa-miR-942-5p | LOXL3 | 2.35 | 0 | -0.97 | 0.00081 | MirTarget | -0.18 | 0.00013 | NA | |
| 137 | hsa-miR-101-3p | NOG | 0.52 | 0.00376 | -1.05 | 0.08272 | miRNATAP | -0.35 | 0.03349 | NA | |
| 138 | hsa-miR-148b-3p | NOG | 1.98 | 0 | -1.05 | 0.08272 | MirTarget | -0.71 | 0 | NA | |
| 139 | hsa-miR-186-5p | NOG | 1.47 | 0 | -1.05 | 0.08272 | MirTarget; miRNATAP | -0.57 | 0.00017 | NA | |
| 140 | hsa-miR-200b-3p | NOG | 3.78 | 0 | -1.05 | 0.08272 | MirTarget; TargetScan | -0.31 | 0 | NA | |
| 141 | hsa-miR-200c-3p | NOG | 3.5 | 0 | -1.05 | 0.08272 | MirTarget; miRNATAP | -0.38 | 0 | NA | |
| 142 | hsa-miR-26b-3p | NOG | 2.18 | 0 | -1.05 | 0.08272 | MirTarget | -0.37 | 0.00098 | NA | |
| 143 | hsa-miR-3607-3p | NOG | 2.69 | 0 | -1.05 | 0.08272 | MirTarget; miRNATAP | -0.24 | 0.00366 | NA | |
| 144 | hsa-miR-429 | NOG | 4.49 | 0 | -1.05 | 0.08272 | MirTarget; PITA; miRanda; miRNATAP | -0.26 | 0 | NA | |
| 145 | hsa-miR-582-3p | NOG | 0.01 | 0.97713 | -1.05 | 0.08272 | PITA; miRNATAP | -0.49 | 0 | NA | |
| 146 | hsa-miR-101-3p | NOTCH1 | 0.52 | 0.00376 | -0.53 | 0.03181 | MirTarget | -0.14 | 0.02945 | NA | |
| 147 | hsa-miR-18a-3p | NOTCH1 | 2.15 | 0 | -0.53 | 0.03181 | MirTarget | -0.11 | 0.00101 | NA | |
| 148 | hsa-miR-28-5p | NOTCH1 | 0.23 | 0.07429 | -0.53 | 0.03181 | miRanda | -0.37 | 9.0E-5 | NA | |
| 149 | hsa-miR-30b-5p | NOTCH1 | 0.8 | 0.00013 | -0.53 | 0.03181 | miRNATAP | -0.12 | 0.03899 | NA | |
| 150 | hsa-miR-32-5p | NOTCH1 | 2.34 | 0 | -0.53 | 0.03181 | miRNATAP | -0.16 | 0.0004 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | EPITHELIAL TO MESENCHYMAL TRANSITION | 26 | 56 | 4.06e-70 | 1.889e-66 |
| 2 | MESENCHYMAL CELL DIFFERENTIATION | 26 | 134 | 2.279e-58 | 5.301e-55 |
| 3 | MESENCHYME DEVELOPMENT | 26 | 190 | 4.46e-54 | 5.188e-51 |
| 4 | STEM CELL DIFFERENTIATION | 26 | 190 | 4.46e-54 | 5.188e-51 |
| 5 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 26 | 513 | 2.31e-42 | 2.149e-39 |
| 6 | CELLULAR COMPONENT MORPHOGENESIS | 26 | 900 | 6.796e-36 | 5.27e-33 |
| 7 | CELL DEVELOPMENT | 26 | 1426 | 1.224e-30 | 8.137e-28 |
| 8 | TISSUE DEVELOPMENT | 26 | 1518 | 6.307e-30 | 3.668e-27 |
| 9 | TISSUE MORPHOGENESIS | 19 | 533 | 4.942e-25 | 2.555e-22 |
| 10 | REGULATION OF STEM CELL DIFFERENTIATION | 13 | 113 | 2.868e-23 | 1.335e-20 |
| 11 | RESPONSE TO GROWTH FACTOR | 17 | 475 | 4.706e-22 | 1.991e-19 |
| 12 | MESENCHYME MORPHOGENESIS | 10 | 38 | 8.739e-22 | 3.388e-19 |
| 13 | GLAND DEVELOPMENT | 16 | 395 | 1.758e-21 | 6.292e-19 |
| 14 | MORPHOGENESIS OF AN EPITHELIUM | 16 | 400 | 2.153e-21 | 6.679e-19 |
| 15 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 20 | 1021 | 2.077e-21 | 6.679e-19 |
| 16 | ORGAN MORPHOGENESIS | 19 | 841 | 2.906e-21 | 8.45e-19 |
| 17 | REGULATION OF OSSIFICATION | 13 | 178 | 1.326e-20 | 3.629e-18 |
| 18 | POSITIVE REGULATION OF STEM CELL DIFFERENTIATION | 10 | 50 | 1.882e-20 | 4.7e-18 |
| 19 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 20 | 1142 | 1.919e-20 | 4.7e-18 |
| 20 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 19 | 957 | 3.333e-20 | 7.753e-18 |
| 21 | REGULATION OF CELL DIFFERENTIATION | 21 | 1492 | 8.522e-20 | 1.888e-17 |
| 22 | EMBRYONIC ORGAN DEVELOPMENT | 15 | 406 | 1.998e-19 | 4.226e-17 |
| 23 | REGULATION OF CARTILAGE DEVELOPMENT | 10 | 63 | 2.32e-19 | 4.693e-17 |
| 24 | REGULATION OF ORGAN MORPHOGENESIS | 13 | 242 | 7.801e-19 | 1.512e-16 |
| 25 | CARDIAC EPITHELIAL TO MESENCHYMAL TRANSITION | 8 | 24 | 1.789e-18 | 3.33e-16 |
| 26 | NEGATIVE REGULATION OF GENE EXPRESSION | 20 | 1493 | 3.817e-18 | 6.831e-16 |
| 27 | REGULATION OF CELL PROLIFERATION | 20 | 1496 | 3.971e-18 | 6.843e-16 |
| 28 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 17 | 823 | 5.224e-18 | 8.682e-16 |
| 29 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 13 | 285 | 6.698e-18 | 1.054e-15 |
| 30 | REGULATION OF CELL DEVELOPMENT | 17 | 836 | 6.798e-18 | 1.054e-15 |
| 31 | HEART MORPHOGENESIS | 12 | 212 | 1.247e-17 | 1.872e-15 |
| 32 | TUBE DEVELOPMENT | 15 | 552 | 1.989e-17 | 2.892e-15 |
| 33 | EMBRYO DEVELOPMENT | 17 | 894 | 2.095e-17 | 2.953e-15 |
| 34 | LOCOMOTION | 18 | 1114 | 2.383e-17 | 3.262e-15 |
| 35 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 20 | 1672 | 3.526e-17 | 4.557e-15 |
| 36 | TUBE MORPHOGENESIS | 13 | 323 | 3.442e-17 | 4.557e-15 |
| 37 | POSITIVE REGULATION OF EPITHELIAL TO MESENCHYMAL TRANSITION | 8 | 34 | 4.382e-17 | 5.51e-15 |
| 38 | EPITHELIUM DEVELOPMENT | 17 | 945 | 5.302e-17 | 6.493e-15 |
| 39 | SKELETAL SYSTEM DEVELOPMENT | 14 | 455 | 6.144e-17 | 7.33e-15 |
| 40 | POSITIVE REGULATION OF GENE EXPRESSION | 20 | 1733 | 7.112e-17 | 8.273e-15 |
| 41 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 16 | 771 | 7.568e-17 | 8.589e-15 |
| 42 | REGULATION OF EPITHELIAL TO MESENCHYMAL TRANSITION | 9 | 67 | 9.073e-17 | 1.005e-14 |
| 43 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 17 | 983 | 1.025e-16 | 1.109e-14 |
| 44 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 16 | 801 | 1.382e-16 | 1.461e-14 |
| 45 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 17 | 1008 | 1.559e-16 | 1.612e-14 |
| 46 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 19 | 1517 | 1.851e-16 | 1.858e-14 |
| 47 | SENSORY ORGAN DEVELOPMENT | 14 | 493 | 1.877e-16 | 1.858e-14 |
| 48 | CELL MOTILITY | 16 | 835 | 2.659e-16 | 2.475e-14 |
| 49 | LOCALIZATION OF CELL | 16 | 835 | 2.659e-16 | 2.475e-14 |
| 50 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 18 | 1275 | 2.582e-16 | 2.475e-14 |
| 51 | CONNECTIVE TISSUE DEVELOPMENT | 11 | 194 | 3.659e-16 | 3.338e-14 |
| 52 | ENDOCARDIAL CUSHION MORPHOGENESIS | 7 | 22 | 4.367e-16 | 3.908e-14 |
| 53 | EMBRYONIC MORPHOGENESIS | 14 | 539 | 6.478e-16 | 5.687e-14 |
| 54 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 18 | 1360 | 8.033e-16 | 6.922e-14 |
| 55 | POSITIVE REGULATION OF LOCOMOTION | 13 | 420 | 1.043e-15 | 8.826e-14 |
| 56 | REGULATION OF STEM CELL PROLIFERATION | 9 | 88 | 1.193e-15 | 9.913e-14 |
| 57 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 18 | 1395 | 1.255e-15 | 1.024e-13 |
| 58 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 17 | 1152 | 1.439e-15 | 1.154e-13 |
| 59 | CARTILAGE DEVELOPMENT | 10 | 147 | 1.621e-15 | 1.278e-13 |
| 60 | NEGATIVE REGULATION OF CARTILAGE DEVELOPMENT | 7 | 26 | 1.679e-15 | 1.302e-13 |
| 61 | REGULATION OF MORPHOGENESIS OF A BRANCHING STRUCTURE | 8 | 53 | 2.107e-15 | 1.607e-13 |
| 62 | NEGATIVE REGULATION OF CELL COMMUNICATION | 17 | 1192 | 2.535e-15 | 1.902e-13 |
| 63 | REGULATION OF CELL DEATH | 18 | 1472 | 3.218e-15 | 2.376e-13 |
| 64 | REGULATION OF HEART MORPHOGENESIS | 7 | 29 | 3.973e-15 | 2.771e-13 |
| 65 | HEART DEVELOPMENT | 13 | 466 | 3.99e-15 | 2.771e-13 |
| 66 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 15 | 788 | 3.873e-15 | 2.771e-13 |
| 67 | CIRCULATORY SYSTEM DEVELOPMENT | 15 | 788 | 3.873e-15 | 2.771e-13 |
| 68 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 10 | 162 | 4.363e-15 | 2.986e-13 |
| 69 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 19 | 1805 | 4.618e-15 | 3.114e-13 |
| 70 | POSITIVE REGULATION OF CELL DEVELOPMENT | 13 | 472 | 4.705e-15 | 3.127e-13 |
| 71 | POSITIVE REGULATION OF CELL PROLIFERATION | 15 | 814 | 6.243e-15 | 4.091e-13 |
| 72 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 19 | 1848 | 7.126e-15 | 4.605e-13 |
| 73 | NEGATIVE REGULATION OF CELL PROLIFERATION | 14 | 643 | 7.416e-15 | 4.727e-13 |
| 74 | ENDOCARDIAL CUSHION DEVELOPMENT | 7 | 32 | 8.547e-15 | 5.374e-13 |
| 75 | FORMATION OF PRIMARY GERM LAYER | 9 | 110 | 9.533e-15 | 5.914e-13 |
| 76 | CELL PROLIFERATION | 14 | 672 | 1.361e-14 | 8.224e-13 |
| 77 | MESODERM MORPHOGENESIS | 8 | 66 | 1.351e-14 | 8.224e-13 |
| 78 | NEGATIVE REGULATION OF CELL DEATH | 15 | 872 | 1.714e-14 | 1.023e-12 |
| 79 | REGULATION OF BINDING | 11 | 283 | 2.4e-14 | 1.414e-12 |
| 80 | KIDNEY EPITHELIUM DEVELOPMENT | 9 | 125 | 3.103e-14 | 1.805e-12 |
| 81 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 13 | 554 | 3.679e-14 | 2.113e-12 |
| 82 | UROGENITAL SYSTEM DEVELOPMENT | 11 | 299 | 4.393e-14 | 2.469e-12 |
| 83 | PATTERN SPECIFICATION PROCESS | 12 | 418 | 4.404e-14 | 2.469e-12 |
| 84 | RESPONSE TO ENDOGENOUS STIMULUS | 17 | 1450 | 6.437e-14 | 3.566e-12 |
| 85 | IN UTERO EMBRYONIC DEVELOPMENT | 11 | 311 | 6.765e-14 | 3.703e-12 |
| 86 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 18 | 1784 | 9.177e-14 | 4.965e-12 |
| 87 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 13 | 609 | 1.233e-13 | 6.594e-12 |
| 88 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 15 | 1004 | 1.344e-13 | 7.106e-12 |
| 89 | DIGESTIVE SYSTEM DEVELOPMENT | 9 | 148 | 1.461e-13 | 7.637e-12 |
| 90 | REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 11 | 337 | 1.629e-13 | 8.424e-12 |
| 91 | MESONEPHROS DEVELOPMENT | 8 | 90 | 1.788e-13 | 9.145e-12 |
| 92 | REGULATION OF CANONICAL WNT SIGNALING PATHWAY | 10 | 236 | 1.947e-13 | 9.846e-12 |
| 93 | POSITIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 9 | 154 | 2.1e-13 | 1.051e-11 |
| 94 | GASTRULATION | 9 | 155 | 2.228e-13 | 1.103e-11 |
| 95 | EPITHELIAL CELL DIFFERENTIATION | 12 | 495 | 3.264e-13 | 1.599e-11 |
| 96 | OSSIFICATION | 10 | 251 | 3.608e-13 | 1.749e-11 |
| 97 | POSITIVE REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 8 | 100 | 4.259e-13 | 2.043e-11 |
| 98 | MORPHOGENESIS OF A BRANCHING STRUCTURE | 9 | 167 | 4.393e-13 | 2.086e-11 |
| 99 | REGULATION OF MUSCLE TISSUE DEVELOPMENT | 8 | 103 | 5.429e-13 | 2.526e-11 |
| 100 | REGULATION OF MUSCLE ORGAN DEVELOPMENT | 8 | 103 | 5.429e-13 | 2.526e-11 |
| 101 | NEUROGENESIS | 16 | 1402 | 8.494e-13 | 3.913e-11 |
| 102 | EMBRYONIC ORGAN MORPHOGENESIS | 10 | 279 | 1.037e-12 | 4.729e-11 |
| 103 | REGULATION OF OSTEOBLAST DIFFERENTIATION | 8 | 112 | 1.078e-12 | 4.869e-11 |
| 104 | REPRODUCTIVE SYSTEM DEVELOPMENT | 11 | 408 | 1.307e-12 | 5.849e-11 |
| 105 | NEGATIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 13 | 740 | 1.464e-12 | 6.487e-11 |
| 106 | MESODERM DEVELOPMENT | 8 | 118 | 1.65e-12 | 7.244e-11 |
| 107 | REGULATION OF WNT SIGNALING PATHWAY | 10 | 310 | 2.954e-12 | 1.285e-10 |
| 108 | REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 9 | 207 | 3.072e-12 | 1.311e-10 |
| 109 | REGIONALIZATION | 10 | 311 | 3.05e-12 | 1.311e-10 |
| 110 | PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 5 | 13 | 3.154e-12 | 1.334e-10 |
| 111 | SKIN EPIDERMIS DEVELOPMENT | 7 | 71 | 3.269e-12 | 1.358e-10 |
| 112 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 12 | 602 | 3.261e-12 | 1.358e-10 |
| 113 | BRANCHING MORPHOGENESIS OF AN EPITHELIAL TUBE | 8 | 131 | 3.862e-12 | 1.59e-10 |
| 114 | REGULATION OF CELL ADHESION | 12 | 629 | 5.449e-12 | 2.224e-10 |
| 115 | REGULATION OF GROWTH | 12 | 633 | 5.868e-12 | 2.374e-10 |
| 116 | CELL FATE COMMITMENT | 9 | 227 | 7.047e-12 | 2.827e-10 |
| 117 | REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 9 | 229 | 7.625e-12 | 3.032e-10 |
| 118 | HAIR CYCLE | 7 | 83 | 1.01e-11 | 3.951e-10 |
| 119 | MOLTING CYCLE | 7 | 83 | 1.01e-11 | 3.951e-10 |
| 120 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 16 | 1656 | 1.087e-11 | 4.213e-10 |
| 121 | PALATE DEVELOPMENT | 7 | 85 | 1.199e-11 | 4.574e-10 |
| 122 | NEURON DIFFERENTIATION | 13 | 874 | 1.191e-11 | 4.574e-10 |
| 123 | BLOOD VESSEL MORPHOGENESIS | 10 | 364 | 1.446e-11 | 5.469e-10 |
| 124 | NEGATIVE REGULATION OF CANONICAL WNT SIGNALING PATHWAY | 8 | 162 | 2.149e-11 | 8.065e-10 |
| 125 | GLAND MORPHOGENESIS | 7 | 97 | 3.09e-11 | 1.132e-09 |
| 126 | REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 6 | 48 | 3.068e-11 | 1.132e-09 |
| 127 | DIGESTIVE TRACT MORPHOGENESIS | 6 | 48 | 3.068e-11 | 1.132e-09 |
| 128 | REGULATION OF CELL MORPHOGENESIS | 11 | 552 | 3.405e-11 | 1.238e-09 |
| 129 | GROWTH | 10 | 410 | 4.659e-11 | 1.68e-09 |
| 130 | REGULATION OF CELLULAR COMPONENT BIOGENESIS | 12 | 767 | 5.475e-11 | 1.96e-09 |
| 131 | POSITIVE REGULATION OF CELL COMMUNICATION | 15 | 1532 | 5.911e-11 | 2.099e-09 |
| 132 | ANGIOGENESIS | 9 | 293 | 6.907e-11 | 2.435e-09 |
| 133 | TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 8 | 190 | 7.723e-11 | 2.702e-09 |
| 134 | COLUMNAR CUBOIDAL EPITHELIAL CELL DIFFERENTIATION | 7 | 111 | 8.075e-11 | 2.804e-09 |
| 135 | RESPONSE TO ABIOTIC STIMULUS | 13 | 1024 | 8.606e-11 | 2.966e-09 |
| 136 | ANTERIOR POSTERIOR PATTERN SPECIFICATION | 8 | 194 | 9.123e-11 | 3.099e-09 |
| 137 | POSITIVE REGULATION OF CELL DEATH | 11 | 605 | 9.072e-11 | 3.099e-09 |
| 138 | NOTCH SIGNALING PATHWAY | 7 | 114 | 9.759e-11 | 3.29e-09 |
| 139 | NEGATIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 7 | 115 | 1.038e-10 | 3.402e-09 |
| 140 | RESPIRATORY SYSTEM DEVELOPMENT | 8 | 197 | 1.031e-10 | 3.402e-09 |
| 141 | NEPHRON DEVELOPMENT | 7 | 115 | 1.038e-10 | 3.402e-09 |
| 142 | NEGATIVE REGULATION OF WNT SIGNALING PATHWAY | 8 | 197 | 1.031e-10 | 3.402e-09 |
| 143 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 16 | 1929 | 1.097e-10 | 3.569e-09 |
| 144 | RESPONSE TO OXYGEN LEVELS | 9 | 311 | 1.173e-10 | 3.791e-09 |
| 145 | SKELETAL SYSTEM MORPHOGENESIS | 8 | 201 | 1.211e-10 | 3.886e-09 |
| 146 | REGULATION OF PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 6 | 60 | 1.239e-10 | 3.948e-09 |
| 147 | POSITIVE REGULATION OF STEM CELL PROLIFERATION | 6 | 61 | 1.373e-10 | 4.346e-09 |
| 148 | VASCULATURE DEVELOPMENT | 10 | 469 | 1.735e-10 | 5.454e-09 |
| 149 | SKIN DEVELOPMENT | 8 | 211 | 1.783e-10 | 5.569e-09 |
| 150 | EPIDERMIS MORPHOGENESIS | 5 | 29 | 2.87e-10 | 8.902e-09 |
| 151 | RESPONSE TO LIPID | 12 | 888 | 2.963e-10 | 9.071e-09 |
| 152 | NEGATIVE REGULATION OF OSSIFICATION | 6 | 69 | 2.944e-10 | 9.071e-09 |
| 153 | ENDODERM DEVELOPMENT | 6 | 71 | 3.511e-10 | 1.068e-08 |
| 154 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 11 | 689 | 3.613e-10 | 1.092e-08 |
| 155 | FOREBRAIN DEVELOPMENT | 9 | 357 | 3.978e-10 | 1.194e-08 |
| 156 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 9 | 363 | 4.608e-10 | 1.375e-08 |
| 157 | HEAD DEVELOPMENT | 11 | 709 | 4.891e-10 | 1.45e-08 |
| 158 | RESPONSE TO TRANSFORMING GROWTH FACTOR BETA | 7 | 144 | 5.081e-10 | 1.496e-08 |
| 159 | RENAL TUBULE DEVELOPMENT | 6 | 78 | 6.259e-10 | 1.832e-08 |
| 160 | EPITHELIAL TO MESENCHYMAL TRANSITION INVOLVED IN ENDOCARDIAL CUSHION FORMATION | 4 | 11 | 7.357e-10 | 2.14e-08 |
| 161 | REGULATION OF MUSCLE CELL DIFFERENTIATION | 7 | 152 | 7.428e-10 | 2.147e-08 |
| 162 | POSITIVE REGULATION OF OSSIFICATION | 6 | 84 | 9.854e-10 | 2.83e-08 |
| 163 | POSITIVE REGULATION OF CELLULAR COMPONENT BIOGENESIS | 9 | 406 | 1.234e-09 | 3.522e-08 |
| 164 | REGULATION OF PROTEIN BINDING | 7 | 168 | 1.497e-09 | 4.247e-08 |
| 165 | APPENDAGE DEVELOPMENT | 7 | 169 | 1.56e-09 | 4.373e-08 |
| 166 | LIMB DEVELOPMENT | 7 | 169 | 1.56e-09 | 4.373e-08 |
| 167 | MESENCHYMAL CELL PROLIFERATION | 4 | 13 | 1.591e-09 | 4.407e-08 |
| 168 | REPRODUCTION | 13 | 1297 | 1.588e-09 | 4.407e-08 |
| 169 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 12 | 1036 | 1.724e-09 | 4.718e-08 |
| 170 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 12 | 1036 | 1.724e-09 | 4.718e-08 |
| 171 | PROSTATE GLAND DEVELOPMENT | 5 | 41 | 1.792e-09 | 4.876e-08 |
| 172 | NEPHRON EPITHELIUM DEVELOPMENT | 6 | 93 | 1.834e-09 | 4.96e-08 |
| 173 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 14 | 1618 | 1.856e-09 | 4.992e-08 |
| 174 | MUSCLE STRUCTURE DEVELOPMENT | 9 | 432 | 2.125e-09 | 5.684e-08 |
| 175 | EXOCRINE SYSTEM DEVELOPMENT | 5 | 45 | 2.911e-09 | 7.741e-08 |
| 176 | ENDOCARDIAL CUSHION FORMATION | 4 | 15 | 3.032e-09 | 7.972e-08 |
| 177 | REGULATION OF CELL PROLIFERATION INVOLVED IN HEART MORPHOGENESIS | 4 | 15 | 3.032e-09 | 7.972e-08 |
| 178 | NEGATIVE REGULATION OF CELL DEVELOPMENT | 8 | 303 | 3.12e-09 | 8.155e-08 |
| 179 | REGULATION OF CHONDROCYTE DIFFERENTIATION | 5 | 46 | 3.264e-09 | 8.484e-08 |
| 180 | CARDIAC CHAMBER MORPHOGENESIS | 6 | 104 | 3.616e-09 | 9.295e-08 |
| 181 | DEVELOPMENTAL GROWTH INVOLVED IN MORPHOGENESIS | 6 | 104 | 3.616e-09 | 9.295e-08 |
| 182 | REGULATION OF PROTEIN MODIFICATION PROCESS | 14 | 1710 | 3.82e-09 | 9.742e-08 |
| 183 | ODONTOGENESIS | 6 | 105 | 3.831e-09 | 9.742e-08 |
| 184 | CARDIAC VENTRICLE DEVELOPMENT | 6 | 106 | 4.058e-09 | 1.018e-07 |
| 185 | CELL SURFACE RECEPTOR SIGNALING PATHWAY INVOLVED IN HEART DEVELOPMENT | 4 | 16 | 4.04e-09 | 1.018e-07 |
| 186 | POSITIVE REGULATION OF PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 5 | 48 | 4.07e-09 | 1.018e-07 |
| 187 | EAR DEVELOPMENT | 7 | 195 | 4.228e-09 | 1.052e-07 |
| 188 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 11 | 872 | 4.306e-09 | 1.066e-07 |
| 189 | REGULATION OF MAPK CASCADE | 10 | 660 | 4.714e-09 | 1.161e-07 |
| 190 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 12 | 1135 | 4.846e-09 | 1.187e-07 |
| 191 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 7 | 200 | 5.041e-09 | 1.228e-07 |
| 192 | EYE DEVELOPMENT | 8 | 326 | 5.535e-09 | 1.341e-07 |
| 193 | DEVELOPMENTAL GROWTH | 8 | 333 | 6.535e-09 | 1.575e-07 |
| 194 | MESONEPHRIC TUBULE MORPHOGENESIS | 5 | 53 | 6.791e-09 | 1.629e-07 |
| 195 | CARDIAC MUSCLE TISSUE MORPHOGENESIS | 5 | 54 | 7.477e-09 | 1.784e-07 |
| 196 | REGULATION OF KIDNEY DEVELOPMENT | 5 | 55 | 8.218e-09 | 1.951e-07 |
| 197 | RESPONSE TO EXTERNAL STIMULUS | 14 | 1821 | 8.641e-09 | 2.041e-07 |
| 198 | POSITIVE REGULATION OF MUSCLE TISSUE DEVELOPMENT | 5 | 56 | 9.015e-09 | 2.119e-07 |
| 199 | EMBRYONIC SKELETAL SYSTEM DEVELOPMENT | 6 | 122 | 9.484e-09 | 2.217e-07 |
| 200 | ENDOCRINE SYSTEM DEVELOPMENT | 6 | 123 | 9.961e-09 | 2.318e-07 |
| 201 | REGULATION OF CELL CYCLE | 11 | 949 | 1.039e-08 | 2.405e-07 |
| 202 | NEGATIVE REGULATION OF CHONDROCYTE DIFFERENTIATION | 4 | 20 | 1.072e-08 | 2.468e-07 |
| 203 | OSTEOBLAST DIFFERENTIATION | 6 | 126 | 1.152e-08 | 2.64e-07 |
| 204 | POSITIVE REGULATION OF BINDING | 6 | 127 | 1.208e-08 | 2.754e-07 |
| 205 | STEM CELL PROLIFERATION | 5 | 60 | 1.284e-08 | 2.915e-07 |
| 206 | REGULATION OF REPRODUCTIVE PROCESS | 6 | 129 | 1.327e-08 | 2.996e-07 |
| 207 | CARDIAC VENTRICLE MORPHOGENESIS | 5 | 62 | 1.519e-08 | 3.415e-07 |
| 208 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 10 | 750 | 1.596e-08 | 3.57e-07 |
| 209 | SOMATIC STEM CELL DIVISION | 4 | 22 | 1.615e-08 | 3.596e-07 |
| 210 | MUSCLE CELL DIFFERENTIATION | 7 | 237 | 1.631e-08 | 3.613e-07 |
| 211 | NEUROEPITHELIAL CELL DIFFERENTIATION | 5 | 63 | 1.649e-08 | 3.636e-07 |
| 212 | MORPHOGENESIS OF EMBRYONIC EPITHELIUM | 6 | 134 | 1.667e-08 | 3.658e-07 |
| 213 | POSITIVE REGULATION OF GROWTH | 7 | 238 | 1.679e-08 | 3.667e-07 |
| 214 | SENSORY ORGAN MORPHOGENESIS | 7 | 239 | 1.728e-08 | 3.757e-07 |
| 215 | PROSTATE GLAND MORPHOGENESIS | 4 | 23 | 1.953e-08 | 4.227e-07 |
| 216 | CARDIAC MUSCLE TISSUE DEVELOPMENT | 6 | 140 | 2.167e-08 | 4.669e-07 |
| 217 | ORGAN GROWTH | 5 | 68 | 2.435e-08 | 5.22e-07 |
| 218 | CARDIAC CHAMBER DEVELOPMENT | 6 | 144 | 2.565e-08 | 5.451e-07 |
| 219 | EPIDERMIS DEVELOPMENT | 7 | 253 | 2.556e-08 | 5.451e-07 |
| 220 | IMMUNE SYSTEM DEVELOPMENT | 9 | 582 | 2.825e-08 | 5.948e-07 |
| 221 | MUSCLE ORGAN MORPHOGENESIS | 5 | 70 | 2.822e-08 | 5.948e-07 |
| 222 | NEGATIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 7 | 262 | 3.25e-08 | 6.811e-07 |
| 223 | AMEBOIDAL TYPE CELL MIGRATION | 6 | 154 | 3.832e-08 | 7.978e-07 |
| 224 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER INVOLVED IN CELLULAR RESPONSE TO CHEMICAL STIMULUS | 4 | 27 | 3.858e-08 | 7.978e-07 |
| 225 | AXIS ELONGATION | 4 | 27 | 3.858e-08 | 7.978e-07 |
| 226 | REGULATION OF BIOMINERAL TISSUE DEVELOPMENT | 5 | 75 | 4.006e-08 | 8.248e-07 |
| 227 | REGULATION OF NEUROBLAST PROLIFERATION | 4 | 28 | 4.497e-08 | 9.218e-07 |
| 228 | MUSCLE TISSUE DEVELOPMENT | 7 | 275 | 4.529e-08 | 9.243e-07 |
| 229 | SOMITE DEVELOPMENT | 5 | 78 | 4.887e-08 | 9.931e-07 |
| 230 | REGULATION OF CARDIAC MUSCLE CELL PROLIFERATION | 4 | 29 | 5.212e-08 | 1.041e-06 |
| 231 | REGULATION OF EXTRACELLULAR MATRIX ORGANIZATION | 4 | 29 | 5.212e-08 | 1.041e-06 |
| 232 | STEM CELL DIVISION | 4 | 29 | 5.212e-08 | 1.041e-06 |
| 233 | POSITIVE REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 4 | 29 | 5.212e-08 | 1.041e-06 |
| 234 | REGULATION OF EPITHELIAL CELL MIGRATION | 6 | 166 | 5.992e-08 | 1.191e-06 |
| 235 | KIDNEY MORPHOGENESIS | 5 | 82 | 6.295e-08 | 1.246e-06 |
| 236 | ORGAN REGENERATION | 5 | 83 | 6.692e-08 | 1.314e-06 |
| 237 | EMBRYONIC PLACENTA DEVELOPMENT | 5 | 83 | 6.692e-08 | 1.314e-06 |
| 238 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 10 | 876 | 6.919e-08 | 1.353e-06 |
| 239 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 13 | 1791 | 7.784e-08 | 1.515e-06 |
| 240 | SALIVARY GLAND DEVELOPMENT | 4 | 32 | 7.87e-08 | 1.519e-06 |
| 241 | REGULATION OF ORGAN FORMATION | 4 | 32 | 7.87e-08 | 1.519e-06 |
| 242 | GLIOGENESIS | 6 | 175 | 8.2e-08 | 1.577e-06 |
| 243 | TISSUE REMODELING | 5 | 87 | 8.487e-08 | 1.625e-06 |
| 244 | ORGAN FORMATION | 4 | 34 | 1.013e-07 | 1.916e-06 |
| 245 | REGULATION OF MESENCHYMAL CELL PROLIFERATION | 4 | 34 | 1.013e-07 | 1.916e-06 |
| 246 | HEART VALVE DEVELOPMENT | 4 | 34 | 1.013e-07 | 1.916e-06 |
| 247 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 12 | 1492 | 1.025e-07 | 1.931e-06 |
| 248 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 10 | 917 | 1.062e-07 | 1.993e-06 |
| 249 | CELL JUNCTION ORGANIZATION | 6 | 185 | 1.14e-07 | 2.13e-06 |
| 250 | EMBRYONIC SKELETAL SYSTEM MORPHOGENESIS | 5 | 93 | 1.187e-07 | 2.21e-06 |
| 251 | REGULATION OF CELLULAR RESPONSE TO STRESS | 9 | 691 | 1.228e-07 | 2.276e-06 |
| 252 | RESPONSE TO BMP | 5 | 94 | 1.253e-07 | 2.305e-06 |
| 253 | CELLULAR RESPONSE TO BMP STIMULUS | 5 | 94 | 1.253e-07 | 2.305e-06 |
| 254 | NEGATIVE REGULATION OF MUSCLE ORGAN DEVELOPMENT | 4 | 36 | 1.285e-07 | 2.335e-06 |
| 255 | NEGATIVE REGULATION OF MUSCLE TISSUE DEVELOPMENT | 4 | 36 | 1.285e-07 | 2.335e-06 |
| 256 | HEAD MORPHOGENESIS | 4 | 36 | 1.285e-07 | 2.335e-06 |
| 257 | CANONICAL WNT SIGNALING PATHWAY | 5 | 95 | 1.322e-07 | 2.393e-06 |
| 258 | PROTEIN PHOSPHORYLATION | 10 | 944 | 1.393e-07 | 2.513e-06 |
| 259 | NEGATIVE REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 4 | 37 | 1.439e-07 | 2.585e-06 |
| 260 | RESPONSE TO STEROID HORMONE | 8 | 497 | 1.448e-07 | 2.592e-06 |
| 261 | CAMERA TYPE EYE MORPHOGENESIS | 5 | 101 | 1.797e-07 | 3.204e-06 |
| 262 | NEGATIVE REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 5 | 102 | 1.888e-07 | 3.353e-06 |
| 263 | POSITIVE REGULATION OF EPITHELIAL CELL MIGRATION | 5 | 103 | 1.983e-07 | 3.508e-06 |
| 264 | REGULATION OF HEART GROWTH | 4 | 42 | 2.428e-07 | 4.28e-06 |
| 265 | BODY MORPHOGENESIS | 4 | 44 | 2.94e-07 | 5.162e-06 |
| 266 | REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 6 | 218 | 3.004e-07 | 5.255e-06 |
| 267 | REGULATION OF EXTRACELLULAR MATRIX ASSEMBLY | 3 | 11 | 3.196e-07 | 5.549e-06 |
| 268 | EPITHELIAL CELL DIFFERENTIATION INVOLVED IN PROSTATE GLAND DEVELOPMENT | 3 | 11 | 3.196e-07 | 5.549e-06 |
| 269 | LUNG MORPHOGENESIS | 4 | 45 | 3.224e-07 | 5.555e-06 |
| 270 | VENTRICULAR CARDIAC MUSCLE TISSUE DEVELOPMENT | 4 | 45 | 3.224e-07 | 5.555e-06 |
| 271 | NEGATIVE REGULATION OF CELL ADHESION | 6 | 223 | 3.433e-07 | 5.894e-06 |
| 272 | REGULATION OF CYTOKINE PRODUCTION | 8 | 563 | 3.747e-07 | 6.389e-06 |
| 273 | MAMMARY GLAND DEVELOPMENT | 5 | 117 | 3.748e-07 | 6.389e-06 |
| 274 | POSITIVE REGULATION OF CELL ADHESION | 7 | 376 | 3.795e-07 | 6.444e-06 |
| 275 | REGULATION OF CELL CELL ADHESION | 7 | 380 | 4.075e-07 | 6.895e-06 |
| 276 | REGULATION OF MESENCHYMAL CELL APOPTOTIC PROCESS | 3 | 12 | 4.257e-07 | 7.178e-06 |
| 277 | NEGATIVE REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 5 | 121 | 4.432e-07 | 7.445e-06 |
| 278 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 11 | 1381 | 4.8e-07 | 8.034e-06 |
| 279 | FACE DEVELOPMENT | 4 | 50 | 4.961e-07 | 8.274e-06 |
| 280 | ARTERY MORPHOGENESIS | 4 | 51 | 5.378e-07 | 8.938e-06 |
| 281 | CARDIOBLAST DIFFERENTIATION | 3 | 13 | 5.53e-07 | 9.157e-06 |
| 282 | HEPATICOBILIARY SYSTEM DEVELOPMENT | 5 | 128 | 5.862e-07 | 9.672e-06 |
| 283 | TUBE FORMATION | 5 | 129 | 6.093e-07 | 1.002e-05 |
| 284 | NEGATIVE REGULATION OF BINDING | 5 | 131 | 6.577e-07 | 1.078e-05 |
| 285 | MAINTENANCE OF CELL NUMBER | 5 | 132 | 6.83e-07 | 1.115e-05 |
| 286 | EMBRYONIC SKELETAL JOINT DEVELOPMENT | 3 | 14 | 7.032e-07 | 1.144e-05 |
| 287 | NEGATIVE REGULATION OF MUSCLE CELL DIFFERENTIATION | 4 | 55 | 7.314e-07 | 1.182e-05 |
| 288 | CRANIAL SKELETAL SYSTEM DEVELOPMENT | 4 | 55 | 7.314e-07 | 1.182e-05 |
| 289 | CELL GROWTH | 5 | 135 | 7.635e-07 | 1.229e-05 |
| 290 | SMAD PROTEIN SIGNAL TRANSDUCTION | 4 | 56 | 7.87e-07 | 1.258e-05 |
| 291 | GLIAL CELL DIFFERENTIATION | 5 | 136 | 7.919e-07 | 1.258e-05 |
| 292 | OUTFLOW TRACT MORPHOGENESIS | 4 | 56 | 7.87e-07 | 1.258e-05 |
| 293 | EYE MORPHOGENESIS | 5 | 136 | 7.919e-07 | 1.258e-05 |
| 294 | NEGATIVE REGULATION OF CELL CELL ADHESION | 5 | 138 | 8.513e-07 | 1.343e-05 |
| 295 | PLACENTA DEVELOPMENT | 5 | 138 | 8.513e-07 | 1.343e-05 |
| 296 | MORPHOGENESIS OF AN EPITHELIAL FOLD | 3 | 15 | 8.782e-07 | 1.376e-05 |
| 297 | STRIATED MUSCLE CELL PROLIFERATION | 3 | 15 | 8.782e-07 | 1.376e-05 |
| 298 | EMBRYONIC PATTERN SPECIFICATION | 4 | 58 | 9.075e-07 | 1.417e-05 |
| 299 | REGULATION OF GLIAL CELL DIFFERENTIATION | 4 | 59 | 9.727e-07 | 1.514e-05 |
| 300 | NEGATIVE REGULATION OF CELL CYCLE | 7 | 433 | 9.787e-07 | 1.518e-05 |
| 301 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 7 | 437 | 1.041e-06 | 1.599e-05 |
| 302 | CHONDROCYTE DIFFERENTIATION | 4 | 60 | 1.041e-06 | 1.599e-05 |
| 303 | POSITIVE REGULATION OF OSTEOBLAST DIFFERENTIATION | 4 | 60 | 1.041e-06 | 1.599e-05 |
| 304 | RESPONSE TO HORMONE | 9 | 893 | 1.065e-06 | 1.631e-05 |
| 305 | MORPHOGENESIS OF AN ENDOTHELIUM | 3 | 16 | 1.08e-06 | 1.642e-05 |
| 306 | PARAXIAL MESODERM DEVELOPMENT | 3 | 16 | 1.08e-06 | 1.642e-05 |
| 307 | RESPONSE TO ESTRADIOL | 5 | 146 | 1.125e-06 | 1.705e-05 |
| 308 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 11 | 1518 | 1.234e-06 | 1.864e-05 |
| 309 | BRANCH ELONGATION OF AN EPITHELIUM | 3 | 17 | 1.31e-06 | 1.954e-05 |
| 310 | NEGATIVE REGULATION OF ALCOHOL BIOSYNTHETIC PROCESS | 3 | 17 | 1.31e-06 | 1.954e-05 |
| 311 | NEGATIVE REGULATION OF STEM CELL PROLIFERATION | 3 | 17 | 1.31e-06 | 1.954e-05 |
| 312 | POSITIVE REGULATION OF EXTRACELLULAR MATRIX ORGANIZATION | 3 | 17 | 1.31e-06 | 1.954e-05 |
| 313 | CELLULAR RESPONSE TO LIPID | 7 | 457 | 1.403e-06 | 2.085e-05 |
| 314 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 7 | 465 | 1.575e-06 | 2.305e-05 |
| 315 | NEPHRON TUBULE FORMATION | 3 | 18 | 1.571e-06 | 2.305e-05 |
| 316 | NOTOCHORD DEVELOPMENT | 3 | 18 | 1.571e-06 | 2.305e-05 |
| 317 | PHOSPHORYLATION | 10 | 1228 | 1.575e-06 | 2.305e-05 |
| 318 | PROTEIN LOCALIZATION TO NUCLEUS | 5 | 156 | 1.56e-06 | 2.305e-05 |
| 319 | NEURON FATE COMMITMENT | 4 | 67 | 1.627e-06 | 2.373e-05 |
| 320 | POSITIVE REGULATION OF MAPK CASCADE | 7 | 470 | 1.691e-06 | 2.458e-05 |
| 321 | REGULATION OF TRANSFERASE ACTIVITY | 9 | 946 | 1.72e-06 | 2.494e-05 |
| 322 | REGULATION OF CELL JUNCTION ASSEMBLY | 4 | 68 | 1.727e-06 | 2.495e-05 |
| 323 | REGULATION OF PROTEIN LOCALIZATION | 9 | 950 | 1.782e-06 | 2.566e-05 |
| 324 | REGENERATION | 5 | 161 | 1.822e-06 | 2.617e-05 |
| 325 | REGULATION OF CARDIAC MUSCLE CELL DIFFERENTIATION | 3 | 19 | 1.864e-06 | 2.652e-05 |
| 326 | MUSCLE CELL PROLIFERATION | 3 | 19 | 1.864e-06 | 2.652e-05 |
| 327 | POSITIVE REGULATION OF CARDIAC MUSCLE CELL PROLIFERATION | 3 | 19 | 1.864e-06 | 2.652e-05 |
| 328 | CELL FATE SPECIFICATION | 4 | 71 | 2.055e-06 | 2.915e-05 |
| 329 | EXTRACELLULAR STRUCTURE ORGANIZATION | 6 | 304 | 2.092e-06 | 2.959e-05 |
| 330 | TRACHEA DEVELOPMENT | 3 | 20 | 2.191e-06 | 3.089e-05 |
| 331 | EMBRYONIC HEART TUBE DEVELOPMENT | 4 | 73 | 2.298e-06 | 3.21e-05 |
| 332 | REGULATION OF ORGAN GROWTH | 4 | 73 | 2.298e-06 | 3.21e-05 |
| 333 | REGULATION OF NEURAL PRECURSOR CELL PROLIFERATION | 4 | 73 | 2.298e-06 | 3.21e-05 |
| 334 | ARTERY DEVELOPMENT | 4 | 75 | 2.561e-06 | 3.526e-05 |
| 335 | POSITIVE REGULATION OF NEUROBLAST PROLIFERATION | 3 | 21 | 2.554e-06 | 3.526e-05 |
| 336 | LEFT RIGHT PATTERN FORMATION | 3 | 21 | 2.554e-06 | 3.526e-05 |
| 337 | ODONTOGENESIS OF DENTIN CONTAINING TOOTH | 4 | 75 | 2.561e-06 | 3.526e-05 |
| 338 | NEURAL CREST CELL DIFFERENTIATION | 4 | 75 | 2.561e-06 | 3.526e-05 |
| 339 | REGULATION OF MAP KINASE ACTIVITY | 6 | 319 | 2.763e-06 | 3.793e-05 |
| 340 | DIENCEPHALON DEVELOPMENT | 4 | 77 | 2.847e-06 | 3.896e-05 |
| 341 | POSITIVE REGULATION OF MESONEPHROS DEVELOPMENT | 3 | 22 | 2.955e-06 | 4.032e-05 |
| 342 | NEGATIVE REGULATION OF PROTEIN BINDING | 4 | 79 | 3.155e-06 | 4.292e-05 |
| 343 | CELLULAR RESPONSE TO MECHANICAL STIMULUS | 4 | 80 | 3.318e-06 | 4.501e-05 |
| 344 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER IN RESPONSE TO STRESS | 3 | 23 | 3.395e-06 | 4.592e-05 |
| 345 | REGULATION OF PROTEIN IMPORT | 5 | 183 | 3.417e-06 | 4.609e-05 |
| 346 | POSITIVE REGULATION OF CELL CYCLE | 6 | 332 | 3.479e-06 | 4.679e-05 |
| 347 | EPITHELIAL CELL DEVELOPMENT | 5 | 186 | 3.7e-06 | 4.962e-05 |
| 348 | NEGATIVE REGULATION OF STEROID METABOLIC PROCESS | 3 | 24 | 3.877e-06 | 5.168e-05 |
| 349 | NEGATIVE REGULATION OF MYOBLAST DIFFERENTIATION | 3 | 24 | 3.877e-06 | 5.168e-05 |
| 350 | MYELOID CELL DIFFERENTIATION | 5 | 189 | 4.002e-06 | 5.32e-05 |
| 351 | REGULATION OF KINASE ACTIVITY | 8 | 776 | 4.156e-06 | 5.509e-05 |
| 352 | REGULATION OF STRIATED MUSCLE CELL DIFFERENTIATION | 4 | 85 | 4.229e-06 | 5.59e-05 |
| 353 | EPITHELIAL TUBE BRANCHING INVOLVED IN LUNG MORPHOGENESIS | 3 | 25 | 4.401e-06 | 5.737e-05 |
| 354 | LUNG CELL DIFFERENTIATION | 3 | 25 | 4.401e-06 | 5.737e-05 |
| 355 | REGULATION OF TRANSFORMING GROWTH FACTOR BETA PRODUCTION | 3 | 25 | 4.401e-06 | 5.737e-05 |
| 356 | REGULATION OF FIBROBLAST GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 3 | 25 | 4.401e-06 | 5.737e-05 |
| 357 | POSITIVE REGULATION OF PEPTIDYL THREONINE PHOSPHORYLATION | 3 | 25 | 4.401e-06 | 5.737e-05 |
| 358 | DEVELOPMENTAL MATURATION | 5 | 193 | 4.433e-06 | 5.761e-05 |
| 359 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 7 | 552 | 4.895e-06 | 6.345e-05 |
| 360 | HEART TRABECULA MORPHOGENESIS | 3 | 26 | 4.971e-06 | 6.355e-05 |
| 361 | NEGATIVE REGULATION OF GLIAL CELL DIFFERENTIATION | 3 | 26 | 4.971e-06 | 6.355e-05 |
| 362 | REGULATION OF MESONEPHROS DEVELOPMENT | 3 | 26 | 4.971e-06 | 6.355e-05 |
| 363 | NEGATIVE REGULATION OF EMBRYONIC DEVELOPMENT | 3 | 26 | 4.971e-06 | 6.355e-05 |
| 364 | HEART GROWTH | 3 | 26 | 4.971e-06 | 6.355e-05 |
| 365 | REGULATION OF NEURON DIFFERENTIATION | 7 | 554 | 5.013e-06 | 6.39e-05 |
| 366 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 9 | 1079 | 5.081e-06 | 6.459e-05 |
| 367 | PROTEIN LOCALIZATION TO ORGANELLE | 7 | 556 | 5.133e-06 | 6.508e-05 |
| 368 | REGULATION OF CELL CYCLE PROCESS | 7 | 558 | 5.255e-06 | 6.645e-05 |
| 369 | REGULATION OF GLIOGENESIS | 4 | 90 | 5.314e-06 | 6.7e-05 |
| 370 | POSITIVE REGULATION OF HEART GROWTH | 3 | 27 | 5.588e-06 | 6.952e-05 |
| 371 | DEVELOPMENTAL INDUCTION | 3 | 27 | 5.588e-06 | 6.952e-05 |
| 372 | REGULATION OF ASTROCYTE DIFFERENTIATION | 3 | 27 | 5.588e-06 | 6.952e-05 |
| 373 | RESPONSE TO LITHIUM ION | 3 | 27 | 5.588e-06 | 6.952e-05 |
| 374 | POSITIVE REGULATION OF MESENCHYMAL CELL PROLIFERATION | 3 | 27 | 5.588e-06 | 6.952e-05 |
| 375 | RESPONSE TO ALCOHOL | 6 | 362 | 5.718e-06 | 7.095e-05 |
| 376 | REGULATION OF DNA BINDING | 4 | 93 | 6.056e-06 | 7.494e-05 |
| 377 | VENTRICULAR SEPTUM MORPHOGENESIS | 3 | 28 | 6.253e-06 | 7.717e-05 |
| 378 | CELL MORPHOGENESIS INVOLVED IN NEURON DIFFERENTIATION | 6 | 368 | 6.282e-06 | 7.733e-05 |
| 379 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 4 | 95 | 6.591e-06 | 8.092e-05 |
| 380 | PROTEIN LOCALIZATION | 11 | 1805 | 6.746e-06 | 8.26e-05 |
| 381 | MYELOID LEUKOCYTE DIFFERENTIATION | 4 | 96 | 6.872e-06 | 8.348e-05 |
| 382 | CARDIOCYTE DIFFERENTIATION | 4 | 96 | 6.872e-06 | 8.348e-05 |
| 383 | NEGATIVE REGULATION OF CYTOKINE PRODUCTION | 5 | 211 | 6.845e-06 | 8.348e-05 |
| 384 | POSITIVE REGULATION OF CARTILAGE DEVELOPMENT | 3 | 29 | 6.968e-06 | 8.444e-05 |
| 385 | SMOOTH MUSCLE CELL DIFFERENTIATION | 3 | 30 | 7.736e-06 | 9.349e-05 |
| 386 | REGULATION OF RESPONSE TO STRESS | 10 | 1468 | 7.857e-06 | 9.471e-05 |
| 387 | RESPONSE TO ESTROGEN | 5 | 218 | 8.021e-06 | 9.644e-05 |
| 388 | REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 4 | 100 | 8.083e-06 | 9.693e-05 |
| 389 | REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 5 | 220 | 8.385e-06 | 0.0001003 |
| 390 | REGULATION OF CELL GROWTH | 6 | 391 | 8.882e-06 | 0.000106 |
| 391 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER IN RESPONSE TO HYPOXIA | 3 | 32 | 9.434e-06 | 0.0001123 |
| 392 | TELENCEPHALON DEVELOPMENT | 5 | 228 | 9.97e-06 | 0.0001183 |
| 393 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 7 | 616 | 1.003e-05 | 0.0001185 |
| 394 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 7 | 616 | 1.003e-05 | 0.0001185 |
| 395 | EMBRYONIC DIGESTIVE TRACT DEVELOPMENT | 3 | 33 | 1.037e-05 | 0.0001219 |
| 396 | REGULATION OF ORGANELLE ORGANIZATION | 9 | 1178 | 1.037e-05 | 0.0001219 |
| 397 | NEURON PROJECTION MORPHOGENESIS | 6 | 402 | 1.04e-05 | 0.0001219 |
| 398 | REGULATION OF VASCULATURE DEVELOPMENT | 5 | 233 | 1.107e-05 | 0.0001295 |
| 399 | NEGATIVE REGULATION OF GROWTH | 5 | 236 | 1.178e-05 | 0.0001374 |
| 400 | HAIR CELL DIFFERENTIATION | 3 | 35 | 1.242e-05 | 0.0001444 |
| 401 | REGULATION OF TRANSCRIPTION REGULATORY REGION DNA BINDING | 3 | 36 | 1.353e-05 | 0.000157 |
| 402 | REGULATION OF EMBRYONIC DEVELOPMENT | 4 | 114 | 1.359e-05 | 0.0001573 |
| 403 | RESPONSE TO FIBROBLAST GROWTH FACTOR | 4 | 116 | 1.455e-05 | 0.000168 |
| 404 | NEGATIVE REGULATION OF GLIOGENESIS | 3 | 37 | 1.472e-05 | 0.0001687 |
| 405 | MYOBLAST DIFFERENTIATION | 3 | 37 | 1.472e-05 | 0.0001687 |
| 406 | REGULATION OF PEPTIDYL THREONINE PHOSPHORYLATION | 3 | 37 | 1.472e-05 | 0.0001687 |
| 407 | NEGATIVE REGULATION OF CYTOPLASMIC TRANSPORT | 4 | 117 | 1.505e-05 | 0.0001721 |
| 408 | COLLAGEN FIBRIL ORGANIZATION | 3 | 38 | 1.596e-05 | 0.0001807 |
| 409 | BONE MINERALIZATION | 3 | 38 | 1.596e-05 | 0.0001807 |
| 410 | POSITIVE REGULATION OF BIOMINERAL TISSUE DEVELOPMENT | 3 | 38 | 1.596e-05 | 0.0001807 |
| 411 | POSITIVE REGULATION OF ORGAN GROWTH | 3 | 38 | 1.596e-05 | 0.0001807 |
| 412 | CYTOKINE PRODUCTION | 4 | 120 | 1.664e-05 | 0.0001879 |
| 413 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 6 | 437 | 1.67e-05 | 0.0001882 |
| 414 | WNT SIGNALING PATHWAY CALCIUM MODULATING PATHWAY | 3 | 39 | 1.728e-05 | 0.0001928 |
| 415 | ANATOMICAL STRUCTURE MATURATION | 3 | 39 | 1.728e-05 | 0.0001928 |
| 416 | TRABECULA MORPHOGENESIS | 3 | 39 | 1.728e-05 | 0.0001928 |
| 417 | GLANDULAR EPITHELIAL CELL DIFFERENTIATION | 3 | 39 | 1.728e-05 | 0.0001928 |
| 418 | REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 4 | 122 | 1.776e-05 | 0.0001977 |
| 419 | POSITIVE REGULATION OF CELL MATRIX ADHESION | 3 | 40 | 1.866e-05 | 0.0002073 |
| 420 | NEGATIVE REGULATION OF LOCOMOTION | 5 | 263 | 1.986e-05 | 0.000219 |
| 421 | REGULATION OF CELLULAR LOCALIZATION | 9 | 1277 | 1.986e-05 | 0.000219 |
| 422 | CELLULAR RESPONSE TO ABIOTIC STIMULUS | 5 | 263 | 1.986e-05 | 0.000219 |
| 423 | POSITIVE REGULATION OF KIDNEY DEVELOPMENT | 3 | 41 | 2.012e-05 | 0.0002213 |
| 424 | AGING | 5 | 264 | 2.023e-05 | 0.000222 |
| 425 | PITUITARY GLAND DEVELOPMENT | 3 | 42 | 2.165e-05 | 0.0002365 |
| 426 | POSITIVE REGULATION OF NEURAL PRECURSOR CELL PROLIFERATION | 3 | 42 | 2.165e-05 | 0.0002365 |
| 427 | CELL DIVISION | 6 | 460 | 2.231e-05 | 0.0002431 |
| 428 | NEGATIVE REGULATION OF STEM CELL DIFFERENTIATION | 3 | 43 | 2.325e-05 | 0.0002528 |
| 429 | LABYRINTHINE LAYER DEVELOPMENT | 3 | 44 | 2.493e-05 | 0.0002692 |
| 430 | NEGATIVE REGULATION OF LIPID BIOSYNTHETIC PROCESS | 3 | 44 | 2.493e-05 | 0.0002692 |
| 431 | BRANCHING INVOLVED IN URETERIC BUD MORPHOGENESIS | 3 | 44 | 2.493e-05 | 0.0002692 |
| 432 | WOUND HEALING | 6 | 470 | 2.518e-05 | 0.0002706 |
| 433 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 6 | 470 | 2.518e-05 | 0.0002706 |
| 434 | MUSCLE ORGAN DEVELOPMENT | 5 | 277 | 2.548e-05 | 0.0002731 |
| 435 | ACTIVATION OF PROTEIN KINASE ACTIVITY | 5 | 279 | 2.637e-05 | 0.0002821 |
| 436 | NEGATIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 4 | 135 | 2.645e-05 | 0.0002823 |
| 437 | CELL DEATH | 8 | 1001 | 2.668e-05 | 0.0002829 |
| 438 | SPROUTING ANGIOGENESIS | 3 | 45 | 2.669e-05 | 0.0002829 |
| 439 | REGULATION OF ALCOHOL BIOSYNTHETIC PROCESS | 3 | 45 | 2.669e-05 | 0.0002829 |
| 440 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 7 | 724 | 2.851e-05 | 0.000301 |
| 441 | EMBRYONIC CRANIAL SKELETON MORPHOGENESIS | 3 | 46 | 2.853e-05 | 0.000301 |
| 442 | REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 4 | 138 | 2.883e-05 | 0.0003035 |
| 443 | POSITIVE REGULATION OF KINASE ACTIVITY | 6 | 482 | 2.902e-05 | 0.0003048 |
| 444 | POSITIVE REGULATION OF NEURON APOPTOTIC PROCESS | 3 | 47 | 3.045e-05 | 0.0003191 |
| 445 | REGULATION OF DEVELOPMENTAL GROWTH | 5 | 289 | 3.121e-05 | 0.0003263 |
| 446 | REGULATION OF MYOBLAST DIFFERENTIATION | 3 | 48 | 3.245e-05 | 0.0003385 |
| 447 | CELLULAR RESPONSE TO OXYGEN LEVELS | 4 | 143 | 3.315e-05 | 0.0003443 |
| 448 | NEGATIVE REGULATION OF INTRACELLULAR TRANSPORT | 4 | 143 | 3.315e-05 | 0.0003443 |
| 449 | REGULATION OF STEROID BIOSYNTHETIC PROCESS | 3 | 49 | 3.453e-05 | 0.0003571 |
| 450 | CARDIAC SEPTUM MORPHOGENESIS | 3 | 49 | 3.453e-05 | 0.0003571 |
| 451 | ENDODERM FORMATION | 3 | 50 | 3.671e-05 | 0.0003787 |
| 452 | POSITIVE REGULATION OF CELL GROWTH | 4 | 148 | 3.792e-05 | 0.0003904 |
| 453 | NEGATIVE REGULATION OF RESPONSE TO DNA DAMAGE STIMULUS | 3 | 52 | 4.132e-05 | 0.0004235 |
| 454 | CELL CELL SIGNALING | 7 | 767 | 4.123e-05 | 0.0004235 |
| 455 | MULTICELLULAR ORGANISM REPRODUCTION | 7 | 768 | 4.157e-05 | 0.0004249 |
| 456 | REGULATION OF PROTEIN TARGETING | 5 | 307 | 4.164e-05 | 0.0004249 |
| 457 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 4 | 153 | 4.318e-05 | 0.0004387 |
| 458 | PALLIUM DEVELOPMENT | 4 | 153 | 4.318e-05 | 0.0004387 |
| 459 | MAMMARY GLAND EPITHELIUM DEVELOPMENT | 3 | 53 | 4.376e-05 | 0.0004436 |
| 460 | REGULATION OF HEMOPOIESIS | 5 | 314 | 4.636e-05 | 0.0004669 |
| 461 | VENTRICULAR SEPTUM DEVELOPMENT | 3 | 54 | 4.629e-05 | 0.0004669 |
| 462 | CELL CYCLE PROCESS | 8 | 1081 | 4.628e-05 | 0.0004669 |
| 463 | BONE DEVELOPMENT | 4 | 156 | 4.658e-05 | 0.0004671 |
| 464 | POSITIVE REGULATION OF DEVELOPMENTAL GROWTH | 4 | 156 | 4.658e-05 | 0.0004671 |
| 465 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 8 | 1087 | 4.814e-05 | 0.0004817 |
| 466 | RESPONSE TO ACID CHEMICAL | 5 | 319 | 4.997e-05 | 0.000499 |
| 467 | POSITIVE REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 3 | 57 | 5.447e-05 | 0.0005427 |
| 468 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 6 | 541 | 5.53e-05 | 0.0005487 |
| 469 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 6 | 541 | 5.53e-05 | 0.0005487 |
| 470 | NEURON PROJECTION DEVELOPMENT | 6 | 545 | 5.762e-05 | 0.0005704 |
| 471 | POSITIVE REGULATION OF DNA BIOSYNTHETIC PROCESS | 3 | 59 | 6.041e-05 | 0.0005968 |
| 472 | RESPONSE TO METAL ION | 5 | 333 | 6.126e-05 | 0.0006039 |
| 473 | CELLULAR RESPONSE TO HORMONE STIMULUS | 6 | 552 | 6.184e-05 | 0.0006084 |
| 474 | POSITIVE REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 3 | 60 | 6.354e-05 | 0.0006224 |
| 475 | OLIGODENDROCYTE DIFFERENTIATION | 3 | 60 | 6.354e-05 | 0.0006224 |
| 476 | EMBRYONIC DIGIT MORPHOGENESIS | 3 | 61 | 6.677e-05 | 0.0006513 |
| 477 | REGULATION OF VIRAL TRANSCRIPTION | 3 | 61 | 6.677e-05 | 0.0006513 |
| 478 | REGULATION OF DNA METABOLIC PROCESS | 5 | 340 | 6.76e-05 | 0.0006581 |
| 479 | RESPONSE TO WOUNDING | 6 | 563 | 6.899e-05 | 0.0006702 |
| 480 | REGULATION OF CELL SUBSTRATE ADHESION | 4 | 173 | 6.967e-05 | 0.0006753 |
| 481 | SOMITOGENESIS | 3 | 62 | 7.01e-05 | 0.0006767 |
| 482 | POSITIVE REGULATION OF PHOSPHATIDYLINOSITOL 3 KINASE SIGNALING | 3 | 62 | 7.01e-05 | 0.0006767 |
| 483 | HOMEOSTASIS OF NUMBER OF CELLS | 4 | 175 | 7.284e-05 | 0.0007017 |
| 484 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 6 | 573 | 7.605e-05 | 0.0007311 |
| 485 | WNT SIGNALING PATHWAY | 5 | 351 | 7.858e-05 | 0.0007539 |
| 486 | REGULATION OF RESPONSE TO OXIDATIVE STRESS | 3 | 65 | 8.075e-05 | 0.0007731 |
| 487 | PROTEIN COMPLEX SUBUNIT ORGANIZATION | 9 | 1527 | 8.15e-05 | 0.0007787 |
| 488 | NEGATIVE REGULATION OF CELLULAR RESPONSE TO TRANSFORMING GROWTH FACTOR BETA STIMULUS | 3 | 66 | 8.453e-05 | 0.000801 |
| 489 | SOMATIC STEM CELL POPULATION MAINTENANCE | 3 | 66 | 8.453e-05 | 0.000801 |
| 490 | NEGATIVE REGULATION OF TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 3 | 66 | 8.453e-05 | 0.000801 |
| 491 | LENS DEVELOPMENT IN CAMERA TYPE EYE | 3 | 66 | 8.453e-05 | 0.000801 |
| 492 | POSITIVE REGULATION OF NEURON DEATH | 3 | 67 | 8.841e-05 | 0.0008258 |
| 493 | ENDOCARDIUM DEVELOPMENT | 2 | 11 | 8.874e-05 | 0.0008258 |
| 494 | SECRETION | 6 | 588 | 8.771e-05 | 0.0008258 |
| 495 | RESPONSE TO FOLLICLE STIMULATING HORMONE | 2 | 11 | 8.874e-05 | 0.0008258 |
| 496 | REGULATION OF DNA TEMPLATED TRANSCRIPTION IN RESPONSE TO STRESS | 3 | 67 | 8.841e-05 | 0.0008258 |
| 497 | PROSTATE GLAND GROWTH | 2 | 11 | 8.874e-05 | 0.0008258 |
| 498 | VENTRICULAR TRABECULA MYOCARDIUM MORPHOGENESIS | 2 | 11 | 8.874e-05 | 0.0008258 |
| 499 | PROSTATE GLANDULAR ACINUS DEVELOPMENT | 2 | 11 | 8.874e-05 | 0.0008258 |
| 500 | ENDOTHELIAL TUBE MORPHOGENESIS | 2 | 11 | 8.874e-05 | 0.0008258 |
| 501 | POSITIVE REGULATION OF DNA METABOLIC PROCESS | 4 | 185 | 9.035e-05 | 0.0008391 |
| 502 | POSITIVE REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 3 | 68 | 9.241e-05 | 0.0008566 |
| 503 | RESPONSE TO ACTIVITY | 3 | 69 | 9.653e-05 | 0.0008929 |
| 504 | POSITIVE REGULATION OF CYTOKINE PRODUCTION | 5 | 370 | 0.0001007 | 0.0009299 |
| 505 | NEGATIVE REGULATION OF NEURON DIFFERENTIATION | 4 | 191 | 0.0001022 | 0.0009418 |
| 506 | NEGATIVE REGULATION OF IMMUNE SYSTEM PROCESS | 5 | 372 | 0.0001033 | 0.00095 |
| 507 | CARDIAC LEFT VENTRICLE MORPHOGENESIS | 2 | 12 | 0.0001064 | 0.0009558 |
| 508 | REGULATION OF TIMING OF CELL DIFFERENTIATION | 2 | 12 | 0.0001064 | 0.0009558 |
| 509 | CARTILAGE MORPHOGENESIS | 2 | 12 | 0.0001064 | 0.0009558 |
| 510 | REGULATION OF VITAMIN METABOLIC PROCESS | 2 | 12 | 0.0001064 | 0.0009558 |
| 511 | REGULATION OF NEURON APOPTOTIC PROCESS | 4 | 192 | 0.0001043 | 0.0009558 |
| 512 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 10 | 1977 | 0.0001045 | 0.0009558 |
| 513 | REGULATION OF MACROPHAGE CYTOKINE PRODUCTION | 2 | 12 | 0.0001064 | 0.0009558 |
| 514 | NEGATIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 3 | 71 | 0.0001051 | 0.0009558 |
| 515 | POSITIVE REGULATION OF SMAD PROTEIN IMPORT INTO NUCLEUS | 2 | 12 | 0.0001064 | 0.0009558 |
| 516 | REGULATION OF DEVELOPMENT HETEROCHRONIC | 2 | 12 | 0.0001064 | 0.0009558 |
| 517 | POSITIVE REGULATION OF ASTROCYTE DIFFERENTIATION | 2 | 12 | 0.0001064 | 0.0009558 |
| 518 | NEGATIVE REGULATION OF RECEPTOR BINDING | 2 | 12 | 0.0001064 | 0.0009558 |
| 519 | IMMUNE SYSTEM PROCESS | 10 | 1984 | 0.0001077 | 0.0009654 |
| 520 | ENDOTHELIAL CELL DIFFERENTIATION | 3 | 72 | 0.0001096 | 0.0009807 |
| 521 | POSITIVE REGULATION OF PROTEIN BINDING | 3 | 73 | 0.0001142 | 0.00102 |
| 522 | REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 5 | 381 | 0.0001156 | 0.00103 |
| 523 | CELLULAR MACROMOLECULE LOCALIZATION | 8 | 1234 | 0.0001178 | 0.001048 |
| 524 | REGULATION OF STEROID METABOLIC PROCESS | 3 | 74 | 0.0001189 | 0.001056 |
| 525 | NEGATIVE REGULATION OF MITOTIC CELL CYCLE | 4 | 199 | 0.0001198 | 0.001061 |
| 526 | NEURONAL STEM CELL DIVISION | 2 | 13 | 0.0001256 | 0.001089 |
| 527 | NEGATIVE REGULATION OF OLIGODENDROCYTE DIFFERENTIATION | 2 | 13 | 0.0001256 | 0.001089 |
| 528 | LEFT RIGHT AXIS SPECIFICATION | 2 | 13 | 0.0001256 | 0.001089 |
| 529 | CELLULAR RESPONSE TO HEPATOCYTE GROWTH FACTOR STIMULUS | 2 | 13 | 0.0001256 | 0.001089 |
| 530 | HEART VALVE FORMATION | 2 | 13 | 0.0001256 | 0.001089 |
| 531 | RESPONSE TO HEPATOCYTE GROWTH FACTOR | 2 | 13 | 0.0001256 | 0.001089 |
| 532 | NEUROBLAST DIVISION | 2 | 13 | 0.0001256 | 0.001089 |
| 533 | GLIAL CELL FATE COMMITMENT | 2 | 13 | 0.0001256 | 0.001089 |
| 534 | BIOMINERAL TISSUE DEVELOPMENT | 3 | 75 | 0.0001238 | 0.001089 |
| 535 | POSITIVE REGULATION OF CARDIAC MUSCLE CELL DIFFERENTIATION | 2 | 13 | 0.0001256 | 0.001089 |
| 536 | REGULATION OF BICELLULAR TIGHT JUNCTION ASSEMBLY | 2 | 13 | 0.0001256 | 0.001089 |
| 537 | NEGATIVE REGULATION OF DNA DAMAGE RESPONSE SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 2 | 13 | 0.0001256 | 0.001089 |
| 538 | CELL PART MORPHOGENESIS | 6 | 633 | 0.0001315 | 0.001137 |
| 539 | REGULATION OF BMP SIGNALING PATHWAY | 3 | 77 | 0.0001338 | 0.001155 |
| 540 | BONE MORPHOGENESIS | 3 | 79 | 0.0001444 | 0.001244 |
| 541 | LYMPHOCYTE DIFFERENTIATION | 4 | 209 | 0.0001446 | 0.001244 |
| 542 | CELL MIGRATION INVOLVED IN HEART DEVELOPMENT | 2 | 14 | 0.0001465 | 0.001246 |
| 543 | SPECIFICATION OF ORGAN IDENTITY | 2 | 14 | 0.0001465 | 0.001246 |
| 544 | CELL MIGRATION INVOLVED IN GASTRULATION | 2 | 14 | 0.0001465 | 0.001246 |
| 545 | REGULATION OF GLOMERULUS DEVELOPMENT | 2 | 14 | 0.0001465 | 0.001246 |
| 546 | POSITIVE REGULATION OF TRANSCRIPTION REGULATORY REGION DNA BINDING | 2 | 14 | 0.0001465 | 0.001246 |
| 547 | REGULATION OF EXTRACELLULAR MATRIX DISASSEMBLY | 2 | 14 | 0.0001465 | 0.001246 |
| 548 | RESPONSE TO MECHANICAL STIMULUS | 4 | 210 | 0.0001473 | 0.001251 |
| 549 | NEGATIVE REGULATION OF LIPID METABOLIC PROCESS | 3 | 80 | 0.0001499 | 0.001271 |
| 550 | RESPONSE TO PEPTIDE | 5 | 404 | 0.0001521 | 0.001284 |
| 551 | SIGNAL TRANSDUCTION BY PROTEIN PHOSPHORYLATION | 5 | 404 | 0.0001521 | 0.001284 |
| 552 | METANEPHROS DEVELOPMENT | 3 | 81 | 0.0001556 | 0.001311 |
| 553 | POSITIVE REGULATION OF HOMEOSTATIC PROCESS | 4 | 216 | 0.0001641 | 0.001381 |
| 554 | POSITIVE REGULATION OF ENDOTHELIAL CELL DIFFERENTIATION | 2 | 15 | 0.0001689 | 0.001413 |
| 555 | OTIC VESICLE DEVELOPMENT | 2 | 15 | 0.0001689 | 0.001413 |
| 556 | EPITHELIAL CELL FATE COMMITMENT | 2 | 15 | 0.0001689 | 0.001413 |
| 557 | CELLULAR RESPONSE TO STEROID HORMONE STIMULUS | 4 | 218 | 0.0001701 | 0.001421 |
| 558 | POSITIVE REGULATION OF MUSCLE CELL DIFFERENTIATION | 3 | 84 | 0.0001733 | 0.001442 |
| 559 | TISSUE MIGRATION | 3 | 84 | 0.0001733 | 0.001442 |
| 560 | CARDIAC SEPTUM DEVELOPMENT | 3 | 85 | 0.0001795 | 0.001489 |
| 561 | POSITIVE REGULATION OF CHROMATIN MODIFICATION | 3 | 85 | 0.0001795 | 0.001489 |
| 562 | CELL CYCLE | 8 | 1316 | 0.0001844 | 0.001526 |
| 563 | NEGATIVE REGULATION OF PHOSPHORYLATION | 5 | 422 | 0.0001863 | 0.00154 |
| 564 | REGULATION OF SMAD PROTEIN IMPORT INTO NUCLEUS | 2 | 16 | 0.0001928 | 0.00158 |
| 565 | ORGAN INDUCTION | 2 | 16 | 0.0001928 | 0.00158 |
| 566 | APOPTOTIC PROCESS INVOLVED IN MORPHOGENESIS | 2 | 16 | 0.0001928 | 0.00158 |
| 567 | ATRIOVENTRICULAR VALVE MORPHOGENESIS | 2 | 16 | 0.0001928 | 0.00158 |
| 568 | BRANCHING INVOLVED IN SALIVARY GLAND MORPHOGENESIS | 2 | 16 | 0.0001928 | 0.00158 |
| 569 | REGULATION OF HYDROLASE ACTIVITY | 8 | 1327 | 0.0001953 | 0.001597 |
| 570 | NEGATIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 6 | 684 | 0.0002006 | 0.001638 |
| 571 | NEURON DEVELOPMENT | 6 | 687 | 0.0002054 | 0.001667 |
| 572 | SEGMENTATION | 3 | 89 | 0.0002057 | 0.001667 |
| 573 | RESPONSE TO DRUG | 5 | 431 | 0.0002055 | 0.001667 |
| 574 | EPITHELIAL CELL PROLIFERATION | 3 | 89 | 0.0002057 | 0.001667 |
| 575 | AXIS SPECIFICATION | 3 | 90 | 0.0002126 | 0.001711 |
| 576 | NEGATIVE REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY | 3 | 90 | 0.0002126 | 0.001711 |
| 577 | ENDOTHELIUM DEVELOPMENT | 3 | 90 | 0.0002126 | 0.001711 |
| 578 | REGULATION OF CELL MATRIX ADHESION | 3 | 90 | 0.0002126 | 0.001711 |
| 579 | NEGATIVE REGULATION OF KIDNEY DEVELOPMENT | 2 | 17 | 0.0002184 | 0.00174 |
| 580 | CELLULAR RESPONSE TO LITHIUM ION | 2 | 17 | 0.0002184 | 0.00174 |
| 581 | REGULATION OF RECEPTOR BINDING | 2 | 17 | 0.0002184 | 0.00174 |
| 582 | NEGATIVE REGULATION OF ANOIKIS | 2 | 17 | 0.0002184 | 0.00174 |
| 583 | REGULATION OF STEM CELL POPULATION MAINTENANCE | 2 | 17 | 0.0002184 | 0.00174 |
| 584 | NEGATIVE REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 2 | 17 | 0.0002184 | 0.00174 |
| 585 | REGULATION OF DNA BIOSYNTHETIC PROCESS | 3 | 94 | 0.0002417 | 0.001922 |
| 586 | REGULATION OF HORMONE BIOSYNTHETIC PROCESS | 2 | 18 | 0.0002455 | 0.001946 |
| 587 | PERICARDIUM DEVELOPMENT | 2 | 18 | 0.0002455 | 0.001946 |
| 588 | NEGATIVE REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 3 | 95 | 0.0002494 | 0.001973 |
| 589 | INFLAMMATORY RESPONSE | 5 | 454 | 0.0002615 | 0.002066 |
| 590 | BIOLOGICAL ADHESION | 7 | 1032 | 0.0002643 | 0.002084 |
| 591 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 3 | 98 | 0.0002733 | 0.00214 |
| 592 | NEGATIVE REGULATION OF TRANSPORT | 5 | 458 | 0.0002723 | 0.00214 |
| 593 | POSITIVE REGULATION OF CELL CYCLE PROCESS | 4 | 247 | 0.0002741 | 0.00214 |
| 594 | ATRIOVENTRICULAR VALVE DEVELOPMENT | 2 | 19 | 0.0002741 | 0.00214 |
| 595 | POSITIVE REGULATION OF CHONDROCYTE DIFFERENTIATION | 2 | 19 | 0.0002741 | 0.00214 |
| 596 | REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 3 | 98 | 0.0002733 | 0.00214 |
| 597 | SINGLE ORGANISM CELL ADHESION | 5 | 459 | 0.0002751 | 0.002144 |
| 598 | EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 3 | 99 | 0.0002816 | 0.00218 |
| 599 | REGULATION OF TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 3 | 99 | 0.0002816 | 0.00218 |
| 600 | REGULATION OF CELLULAR RESPONSE TO TRANSFORMING GROWTH FACTOR BETA STIMULUS | 3 | 99 | 0.0002816 | 0.00218 |
| 601 | POSITIVE REGULATION OF CELL SUBSTRATE ADHESION | 3 | 99 | 0.0002816 | 0.00218 |
| 602 | MODIFICATION OF MORPHOLOGY OR PHYSIOLOGY OF OTHER ORGANISM | 3 | 100 | 0.0002901 | 0.002235 |
| 603 | TAXIS | 5 | 464 | 0.0002892 | 0.002235 |
| 604 | LIMBIC SYSTEM DEVELOPMENT | 3 | 100 | 0.0002901 | 0.002235 |
| 605 | REGULATION OF TRANSPORT | 9 | 1804 | 0.0002926 | 0.002251 |
| 606 | REGULATION OF NEURON DEATH | 4 | 252 | 0.0002958 | 0.002271 |
| 607 | NEGATIVE REGULATION OF BIOMINERAL TISSUE DEVELOPMENT | 2 | 20 | 0.0003044 | 0.002322 |
| 608 | REGULATION OF SMOOTH MUSCLE CELL DIFFERENTIATION | 2 | 20 | 0.0003044 | 0.002322 |
| 609 | TONGUE DEVELOPMENT | 2 | 20 | 0.0003044 | 0.002322 |
| 610 | MYELOID DENDRITIC CELL DIFFERENTIATION | 2 | 20 | 0.0003044 | 0.002322 |
| 611 | LEUKOCYTE CELL CELL ADHESION | 4 | 255 | 0.0003094 | 0.002356 |
| 612 | RESPONSE TO INORGANIC SUBSTANCE | 5 | 479 | 0.0003348 | 0.002531 |
| 613 | ECTODERM DEVELOPMENT | 2 | 21 | 0.0003361 | 0.002531 |
| 614 | CEREBRAL CORTEX DEVELOPMENT | 3 | 105 | 0.0003348 | 0.002531 |
| 615 | CELL AGGREGATION | 2 | 21 | 0.0003361 | 0.002531 |
| 616 | APOPTOTIC PROCESS INVOLVED IN DEVELOPMENT | 2 | 21 | 0.0003361 | 0.002531 |
| 617 | POSITIVE REGULATION OF ADHERENS JUNCTION ORGANIZATION | 2 | 21 | 0.0003361 | 0.002531 |
| 618 | CARTILAGE CONDENSATION | 2 | 21 | 0.0003361 | 0.002531 |
| 619 | REGULATION OF CYTOPLASMIC TRANSPORT | 5 | 481 | 0.0003413 | 0.002565 |
| 620 | REGULATION OF FAT CELL DIFFERENTIATION | 3 | 106 | 0.0003443 | 0.00258 |
| 621 | FAT CELL DIFFERENTIATION | 3 | 106 | 0.0003443 | 0.00258 |
| 622 | CELLULAR RESPONSE TO EXTERNAL STIMULUS | 4 | 264 | 0.0003529 | 0.00264 |
| 623 | PLATELET DEGRANULATION | 3 | 107 | 0.0003539 | 0.002643 |
| 624 | SECRETION BY CELL | 5 | 486 | 0.0003579 | 0.002669 |
| 625 | SEX DIFFERENTIATION | 4 | 266 | 0.0003632 | 0.002704 |
| 626 | NEGATIVE REGULATION OF CYTOKINE PRODUCTION INVOLVED IN IMMUNE RESPONSE | 2 | 22 | 0.0003694 | 0.002729 |
| 627 | BETA CATENIN DESTRUCTION COMPLEX DISASSEMBLY | 2 | 22 | 0.0003694 | 0.002729 |
| 628 | MODULATION OF TRANSCRIPTION IN OTHER ORGANISM INVOLVED IN SYMBIOTIC INTERACTION | 2 | 22 | 0.0003694 | 0.002729 |
| 629 | PROTEIN LOCALIZATION TO CELL SURFACE | 2 | 22 | 0.0003694 | 0.002729 |
| 630 | EMBRYONIC PLACENTA MORPHOGENESIS | 2 | 22 | 0.0003694 | 0.002729 |
| 631 | REGULATION OF CELL DIVISION | 4 | 272 | 0.0003952 | 0.002914 |
| 632 | TRANSMEMBRANE RECEPTOR PROTEIN TYROSINE KINASE SIGNALING PATHWAY | 5 | 498 | 0.0004002 | 0.002947 |
| 633 | REGULATION OF OSTEOBLAST PROLIFERATION | 2 | 23 | 0.0004043 | 0.002965 |
| 634 | EAR MORPHOGENESIS | 3 | 112 | 0.0004046 | 0.002965 |
| 635 | REGULATION OF SUPEROXIDE METABOLIC PROCESS | 2 | 23 | 0.0004043 | 0.002965 |
| 636 | NEGATIVE REGULATION OF BLOOD VESSEL ENDOTHELIAL CELL MIGRATION | 2 | 24 | 0.0004407 | 0.003199 |
| 637 | EMBRYONIC CAMERA TYPE EYE MORPHOGENESIS | 2 | 24 | 0.0004407 | 0.003199 |
| 638 | RESPONSE TO STEROL | 2 | 24 | 0.0004407 | 0.003199 |
| 639 | POSITIVE REGULATION OF CELL JUNCTION ASSEMBLY | 2 | 24 | 0.0004407 | 0.003199 |
| 640 | REGULATION OF ODONTOGENESIS | 2 | 24 | 0.0004407 | 0.003199 |
| 641 | REGULATION OF ANOIKIS | 2 | 24 | 0.0004407 | 0.003199 |
| 642 | REGULATION OF LIPID METABOLIC PROCESS | 4 | 282 | 0.0004529 | 0.003282 |
| 643 | NEGATIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 3 | 117 | 0.0004598 | 0.003328 |
| 644 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 6 | 799 | 0.0004625 | 0.003337 |
| 645 | POSITIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 5 | 514 | 0.0004625 | 0.003337 |
| 646 | REGULATION OF PEPTIDYL SERINE PHOSPHORYLATION | 3 | 118 | 0.0004714 | 0.003396 |
| 647 | CELLULAR RESPONSE TO EPIDERMAL GROWTH FACTOR STIMULUS | 2 | 25 | 0.0004786 | 0.003442 |
| 648 | POSITIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 3 | 121 | 0.0005073 | 0.003643 |
| 649 | DEVELOPMENTAL PROGRAMMED CELL DEATH | 2 | 26 | 0.0005181 | 0.003669 |
| 650 | NEGATIVE REGULATION OF STRIATED MUSCLE CELL DIFFERENTIATION | 2 | 26 | 0.0005181 | 0.003669 |
| 651 | LEUKOCYTE DIFFERENTIATION | 4 | 292 | 0.0005164 | 0.003669 |
| 652 | MESODERMAL CELL DIFFERENTIATION | 2 | 26 | 0.0005181 | 0.003669 |
| 653 | REGULATION OF HORMONE METABOLIC PROCESS | 2 | 26 | 0.0005181 | 0.003669 |
| 654 | REGULATION OF P38MAPK CASCADE | 2 | 26 | 0.0005181 | 0.003669 |
| 655 | MYELOID DENDRITIC CELL ACTIVATION | 2 | 26 | 0.0005181 | 0.003669 |
| 656 | POSITIVE REGULATION OF VASCULAR ENDOTHELIAL GROWTH FACTOR PRODUCTION | 2 | 26 | 0.0005181 | 0.003669 |
| 657 | REGULATION OF CELL FATE COMMITMENT | 2 | 26 | 0.0005181 | 0.003669 |
| 658 | T CELL DIFFERENTIATION | 3 | 123 | 0.0005322 | 0.003763 |
| 659 | REGULATION OF ENDOTHELIAL CELL DIFFERENTIATION | 2 | 27 | 0.0005591 | 0.003918 |
| 660 | NEGATIVE REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO DNA DAMAGE | 2 | 27 | 0.0005591 | 0.003918 |
| 661 | INFLAMMATORY RESPONSE TO ANTIGENIC STIMULUS | 2 | 27 | 0.0005591 | 0.003918 |
| 662 | STEROID HORMONE MEDIATED SIGNALING PATHWAY | 3 | 125 | 0.0005578 | 0.003918 |
| 663 | NEGATIVE REGULATION OF SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 2 | 27 | 0.0005591 | 0.003918 |
| 664 | SUBSTRATE DEPENDENT CELL MIGRATION | 2 | 27 | 0.0005591 | 0.003918 |
| 665 | NEGATIVE REGULATION OF CATALYTIC ACTIVITY | 6 | 829 | 0.0005624 | 0.003935 |
| 666 | CHROMATIN MODIFICATION | 5 | 539 | 0.0005742 | 0.004012 |
| 667 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 4 | 303 | 0.0005934 | 0.00414 |
| 668 | REGULATION OF DNA DAMAGE RESPONSE SIGNAL TRANSDUCTION BY P53 CLASS MEDIATOR | 2 | 28 | 0.0006016 | 0.004153 |
| 669 | REGULATION OF LIPID BIOSYNTHETIC PROCESS | 3 | 128 | 0.0005978 | 0.004153 |
| 670 | AUDITORY RECEPTOR CELL DIFFERENTIATION | 2 | 28 | 0.0006016 | 0.004153 |
| 671 | DOPAMINERGIC NEURON DIFFERENTIATION | 2 | 28 | 0.0006016 | 0.004153 |
| 672 | RESPONSE TO GONADOTROPIN | 2 | 28 | 0.0006016 | 0.004153 |
| 673 | MAMMARY GLAND DUCT MORPHOGENESIS | 2 | 28 | 0.0006016 | 0.004153 |
| 674 | METANEPHROS MORPHOGENESIS | 2 | 28 | 0.0006016 | 0.004153 |
| 675 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 3 | 129 | 0.0006115 | 0.004203 |
| 676 | NUCLEAR IMPORT | 3 | 129 | 0.0006115 | 0.004203 |
| 677 | CELL JUNCTION ASSEMBLY | 3 | 129 | 0.0006115 | 0.004203 |
| 678 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 4 | 306 | 0.0006158 | 0.004226 |
| 679 | NEGATIVE REGULATION OF PRODUCTION OF MOLECULAR MEDIATOR OF IMMUNE RESPONSE | 2 | 29 | 0.0006457 | 0.004405 |
| 680 | EMBRYONIC HINDLIMB MORPHOGENESIS | 2 | 29 | 0.0006457 | 0.004405 |
| 681 | REGULATION OF OLIGODENDROCYTE DIFFERENTIATION | 2 | 29 | 0.0006457 | 0.004405 |
| 682 | NEUROBLAST PROLIFERATION | 2 | 29 | 0.0006457 | 0.004405 |
| 683 | POSITIVE REGULATION OF CELL DIVISION | 3 | 132 | 0.0006538 | 0.004454 |
| 684 | RESPONSE TO NITROGEN COMPOUND | 6 | 859 | 0.0006785 | 0.004615 |
| 685 | PROTEIN EXPORT FROM NUCLEUS | 2 | 30 | 0.0006913 | 0.004675 |
| 686 | PROTEIN LOCALIZATION TO CYTOSKELETON | 2 | 30 | 0.0006913 | 0.004675 |
| 687 | RESPONSE TO EPIDERMAL GROWTH FACTOR | 2 | 30 | 0.0006913 | 0.004675 |
| 688 | NEGATIVE REGULATION OF MUSCLE CELL APOPTOTIC PROCESS | 2 | 30 | 0.0006913 | 0.004675 |
| 689 | HINDBRAIN DEVELOPMENT | 3 | 137 | 0.0007284 | 0.004912 |
| 690 | CELL ACTIVATION | 5 | 568 | 0.000728 | 0.004912 |
| 691 | HYALURONAN METABOLIC PROCESS | 2 | 31 | 0.0007383 | 0.004965 |
| 692 | REGULATION OF VASCULAR ENDOTHELIAL GROWTH FACTOR PRODUCTION | 2 | 31 | 0.0007383 | 0.004965 |
| 693 | NON CANONICAL WNT SIGNALING PATHWAY | 3 | 140 | 0.0007757 | 0.005208 |
| 694 | POSITIVE REGULATION OF GLIAL CELL DIFFERENTIATION | 2 | 32 | 0.0007869 | 0.005246 |
| 695 | POSITIVE REGULATION OF BMP SIGNALING PATHWAY | 2 | 32 | 0.0007869 | 0.005246 |
| 696 | PATTERNING OF BLOOD VESSELS | 2 | 32 | 0.0007869 | 0.005246 |
| 697 | POSITIVE REGULATION OF EPIDERMIS DEVELOPMENT | 2 | 32 | 0.0007869 | 0.005246 |
| 698 | BLOOD VESSEL REMODELING | 2 | 32 | 0.0007869 | 0.005246 |
| 699 | RESPONSE TO BIOTIC STIMULUS | 6 | 886 | 0.0007983 | 0.005314 |
| 700 | EPIDERMAL CELL DIFFERENTIATION | 3 | 142 | 0.0008083 | 0.005373 |
| 701 | MULTI ORGANISM REPRODUCTIVE PROCESS | 6 | 891 | 0.0008221 | 0.005457 |
| 702 | DENDRITIC CELL DIFFERENTIATION | 2 | 33 | 0.000837 | 0.005509 |
| 703 | EMBRYONIC AXIS SPECIFICATION | 2 | 33 | 0.000837 | 0.005509 |
| 704 | REGULATION OF MYELINATION | 2 | 33 | 0.000837 | 0.005509 |
| 705 | REGULATION OF T CELL APOPTOTIC PROCESS | 2 | 33 | 0.000837 | 0.005509 |
| 706 | RESPONSE TO VITAMIN D | 2 | 33 | 0.000837 | 0.005509 |
| 707 | EMBRYONIC EYE MORPHOGENESIS | 2 | 33 | 0.000837 | 0.005509 |
| 708 | REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY | 3 | 145 | 0.0008587 | 0.005636 |
| 709 | REGULATION OF RESPONSE TO DNA DAMAGE STIMULUS | 3 | 145 | 0.0008587 | 0.005636 |
| 710 | CELL PROJECTION ORGANIZATION | 6 | 902 | 0.0008766 | 0.005745 |
| 711 | LUNG EPITHELIUM DEVELOPMENT | 2 | 34 | 0.0008886 | 0.005799 |
| 712 | PROTEIN KINASE B SIGNALING | 2 | 34 | 0.0008886 | 0.005799 |
| 713 | REGULATION OF INTRINSIC APOPTOTIC SIGNALING PATHWAY IN RESPONSE TO DNA DAMAGE | 2 | 34 | 0.0008886 | 0.005799 |
| 714 | GAMETE GENERATION | 5 | 595 | 0.0008974 | 0.005848 |
| 715 | PEPTIDYL SERINE MODIFICATION | 3 | 148 | 0.0009112 | 0.005921 |
| 716 | MALE SEX DIFFERENTIATION | 3 | 148 | 0.0009112 | 0.005921 |
| 717 | NEURAL TUBE DEVELOPMENT | 3 | 149 | 0.0009291 | 0.006029 |
| 718 | LYMPHOCYTE ACTIVATION | 4 | 342 | 0.0009328 | 0.006045 |
| 719 | NEGATIVE REGULATION OF RESPONSE TO OXIDATIVE STRESS | 2 | 35 | 0.0009417 | 0.006061 |
| 720 | BONE REMODELING | 2 | 35 | 0.0009417 | 0.006061 |
| 721 | NEGATIVE REGULATION OF CELLULAR RESPONSE TO OXIDATIVE STRESS | 2 | 35 | 0.0009417 | 0.006061 |
| 722 | EMBRYONIC CAMERA TYPE EYE DEVELOPMENT | 2 | 35 | 0.0009417 | 0.006061 |
| 723 | REGULATION OF RESPONSE TO REACTIVE OXYGEN SPECIES | 2 | 35 | 0.0009417 | 0.006061 |
| 724 | POSITIVE REGULATION OF CHROMOSOME ORGANIZATION | 3 | 150 | 0.0009473 | 0.006079 |
| 725 | CHROMATIN REMODELING | 3 | 150 | 0.0009473 | 0.006079 |
| 726 | REGULATION OF REACTIVE OXYGEN SPECIES METABOLIC PROCESS | 3 | 152 | 0.0009842 | 0.006291 |
| 727 | REGULATION OF CHROMATIN ORGANIZATION | 3 | 152 | 0.0009842 | 0.006291 |
| 728 | POSITIVE REGULATION OF WNT SIGNALING PATHWAY | 3 | 152 | 0.0009842 | 0.006291 |
| 729 | CELL CELL ADHESION | 5 | 608 | 0.0009887 | 0.006311 |
| 730 | POSITIVE REGULATION OF PROTEIN ACETYLATION | 2 | 36 | 0.0009963 | 0.006351 |
| 731 | CELL CYCLE ARREST | 3 | 154 | 0.001022 | 0.006506 |
| 732 | PROTEIN IMPORT | 3 | 155 | 0.001041 | 0.006619 |
| 733 | HINDLIMB MORPHOGENESIS | 2 | 37 | 0.001052 | 0.006672 |
| 734 | GLIAL CELL MIGRATION | 2 | 37 | 0.001052 | 0.006672 |
| 735 | CELLULAR RESPONSE TO INORGANIC SUBSTANCE | 3 | 156 | 0.001061 | 0.006716 |
| 736 | NUCLEAR TRANSPORT | 4 | 355 | 0.001071 | 0.006772 |
| 737 | REGULATION OF INTRACELLULAR TRANSPORT | 5 | 621 | 0.001087 | 0.006862 |
| 738 | HORMONE MEDIATED SIGNALING PATHWAY | 3 | 158 | 0.001101 | 0.00693 |
| 739 | REGULATION OF DEPHOSPHORYLATION | 3 | 158 | 0.001101 | 0.00693 |
| 740 | CYTOKINE SECRETION | 2 | 38 | 0.00111 | 0.00698 |
| 741 | POSITIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 4 | 360 | 0.001128 | 0.007084 |
| 742 | POSITIVE REGULATION OF VIRAL TRANSCRIPTION | 2 | 39 | 0.001169 | 0.007292 |
| 743 | NEGATIVE REGULATION OF TRANSCRIPTION FACTOR IMPORT INTO NUCLEUS | 2 | 39 | 0.001169 | 0.007292 |
| 744 | REGULATION OF CELL ADHESION MEDIATED BY INTEGRIN | 2 | 39 | 0.001169 | 0.007292 |
| 745 | ASTROCYTE DIFFERENTIATION | 2 | 39 | 0.001169 | 0.007292 |
| 746 | NEGATIVE REGULATION OF ENDOTHELIAL CELL MIGRATION | 2 | 39 | 0.001169 | 0.007292 |
| 747 | CELL CHEMOTAXIS | 3 | 162 | 0.001183 | 0.007367 |
| 748 | POSITIVE REGULATION OF HEMOPOIESIS | 3 | 163 | 0.001204 | 0.007479 |
| 749 | CELLULAR RESPONSE TO BIOTIC STIMULUS | 3 | 163 | 0.001204 | 0.007479 |
| 750 | MAMMARY GLAND MORPHOGENESIS | 2 | 40 | 0.00123 | 0.007608 |
| 751 | ENDODERMAL CELL DIFFERENTIATION | 2 | 40 | 0.00123 | 0.007608 |
| 752 | NEGATIVE REGULATION OF OSTEOBLAST DIFFERENTIATION | 2 | 40 | 0.00123 | 0.007608 |
| 753 | POSITIVE REGULATION OF DNA BINDING | 2 | 42 | 0.001355 | 0.008352 |
| 754 | NEGATIVE REGULATION OF BMP SIGNALING PATHWAY | 2 | 42 | 0.001355 | 0.008352 |
| 755 | GENITALIA DEVELOPMENT | 2 | 42 | 0.001355 | 0.008352 |
| 756 | NEGATIVE REGULATION OF CELL GROWTH | 3 | 170 | 0.001359 | 0.008362 |
| 757 | POSITIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 3 | 171 | 0.001382 | 0.008493 |
| 758 | REGULATION OF CARBOHYDRATE METABOLIC PROCESS | 3 | 172 | 0.001405 | 0.008625 |
| 759 | REGULATION OF MUSCLE CELL APOPTOTIC PROCESS | 2 | 43 | 0.00142 | 0.008683 |
| 760 | REGULATION OF POLYSACCHARIDE METABOLIC PROCESS | 2 | 43 | 0.00142 | 0.008683 |
| 761 | BETA CATENIN TCF COMPLEX ASSEMBLY | 2 | 43 | 0.00142 | 0.008683 |
| 762 | STRIATED MUSCLE CELL DIFFERENTIATION | 3 | 173 | 0.001429 | 0.008723 |
| 763 | CHROMATIN ORGANIZATION | 5 | 663 | 0.001454 | 0.008868 |
| 764 | CELLULAR RESPONSE TO ACID CHEMICAL | 3 | 175 | 0.001476 | 0.008992 |
| 765 | RESPONSE TO CORTICOSTEROID | 3 | 176 | 0.001501 | 0.009128 |
| 766 | NEGATIVE REGULATION OF DNA BINDING | 2 | 46 | 0.001624 | 0.009813 |
| 767 | PEPTIDYL THREONINE MODIFICATION | 2 | 46 | 0.001624 | 0.009813 |
| 768 | MACROMOLECULAR COMPLEX ASSEMBLY | 7 | 1398 | 0.001622 | 0.009813 |
| 769 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 2 | 46 | 0.001624 | 0.009813 |
| 770 | REGULATION OF OXIDATIVE STRESS INDUCED CELL DEATH | 2 | 46 | 0.001624 | 0.009813 |
| 771 | RESPONSE TO KETONE | 3 | 182 | 0.001652 | 0.009969 |
| 772 | REGULATION OF IMMUNE SYSTEM PROCESS | 7 | 1403 | 0.001656 | 0.009983 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR BINDING | 6 | 50 | 3.966e-11 | 3.685e-08 |
| 2 | MACROMOLECULAR COMPLEX BINDING | 13 | 1399 | 3.996e-09 | 1.856e-06 |
| 3 | RECEPTOR BINDING | 13 | 1476 | 7.654e-09 | 2.37e-06 |
| 4 | SMAD BINDING | 5 | 72 | 3.256e-08 | 7.563e-06 |
| 5 | CHROMATIN BINDING | 8 | 435 | 5.204e-08 | 9.669e-06 |
| 6 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II DISTAL ENHANCER SEQUENCE SPECIFIC BINDING | 5 | 90 | 1.007e-07 | 1.559e-05 |
| 7 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 7 | 328 | 1.506e-07 | 1.999e-05 |
| 8 | TRANSCRIPTION FACTOR BINDING | 8 | 524 | 2.17e-07 | 2.52e-05 |
| 9 | REGULATORY REGION NUCLEIC ACID BINDING | 9 | 818 | 5.114e-07 | 5.279e-05 |
| 10 | RNA POLYMERASE II TRANSCRIPTION FACTOR ACTIVITY SEQUENCE SPECIFIC DNA BINDING | 8 | 629 | 8.663e-07 | 8.047e-05 |
| 11 | CYTOKINE RECEPTOR BINDING | 6 | 271 | 1.074e-06 | 9.068e-05 |
| 12 | NUCLEIC ACID BINDING TRANSCRIPTION FACTOR ACTIVITY | 10 | 1199 | 1.267e-06 | 9.81e-05 |
| 13 | PROTEIN HETERODIMERIZATION ACTIVITY | 7 | 468 | 1.643e-06 | 0.0001173 |
| 14 | GROWTH FACTOR ACTIVITY | 5 | 160 | 1.767e-06 | 0.0001173 |
| 15 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION REGULATORY REGION SEQUENCE SPECIFIC BINDING | 6 | 315 | 2.569e-06 | 0.0001591 |
| 16 | SEQUENCE SPECIFIC DNA BINDING | 9 | 1037 | 3.67e-06 | 0.0002131 |
| 17 | PROTEIN DIMERIZATION ACTIVITY | 9 | 1149 | 8.477e-06 | 0.0004632 |
| 18 | KINASE BINDING | 7 | 606 | 9.018e-06 | 0.0004654 |
| 19 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 5 | 226 | 9.553e-06 | 0.0004671 |
| 20 | PROTEIN KINASE ACTIVITY | 7 | 640 | 1.286e-05 | 0.0005975 |
| 21 | ENZYME BINDING | 10 | 1737 | 3.447e-05 | 0.001525 |
| 22 | DOUBLE STRANDED DNA BINDING | 7 | 764 | 4.021e-05 | 0.001698 |
| 23 | CORE PROMOTER BINDING | 4 | 152 | 4.209e-05 | 0.0017 |
| 24 | KINASE ACTIVITY | 7 | 842 | 7.449e-05 | 0.002883 |
| 25 | I SMAD BINDING | 2 | 11 | 8.874e-05 | 0.003298 |
| 26 | BETA CATENIN BINDING | 3 | 84 | 0.0001733 | 0.004878 |
| 27 | TRANSMEMBRANE RECEPTOR PROTEIN KINASE ACTIVITY | 3 | 81 | 0.0001556 | 0.004878 |
| 28 | C2H2 ZINC FINGER DOMAIN BINDING | 2 | 14 | 0.0001465 | 0.004878 |
| 29 | PROTEIN COMPLEX BINDING | 7 | 935 | 0.0001436 | 0.004878 |
| 30 | RECEPTOR SERINE THREONINE KINASE BINDING | 2 | 15 | 0.0001689 | 0.004878 |
| 31 | PROTEIN KINASE A CATALYTIC SUBUNIT BINDING | 2 | 15 | 0.0001689 | 0.004878 |
| 32 | OXIDOREDUCTASE ACTIVITY ACTING ON THE CH NH2 GROUP OF DONORS OXYGEN AS ACCEPTOR | 2 | 15 | 0.0001689 | 0.004878 |
| 33 | CYTOKINE ACTIVITY | 4 | 219 | 0.0001731 | 0.004878 |
| 34 | TRANSFORMING GROWTH FACTOR BETA BINDING | 2 | 16 | 0.0001928 | 0.005269 |
| 35 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 7 | 992 | 0.0002072 | 0.0055 |
| 36 | TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE ACTIVITY | 2 | 17 | 0.0002184 | 0.005635 |
| 37 | CYTOKINE BINDING | 3 | 92 | 0.0002268 | 0.005695 |
| 38 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 5 | 445 | 0.0002384 | 0.005827 |
| 39 | OXIDOREDUCTASE ACTIVITY ACTING ON THE CH NH2 GROUP OF DONORS | 2 | 19 | 0.0002741 | 0.00653 |
| 40 | DNA BINDING BENDING | 2 | 20 | 0.0003044 | 0.007069 |
| 41 | RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 3 | 104 | 0.0003255 | 0.007376 |
| 42 | TRANSCRIPTIONAL REPRESSOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 3 | 105 | 0.0003348 | 0.007406 |
| 43 | IONOTROPIC GLUTAMATE RECEPTOR BINDING | 2 | 23 | 0.0004043 | 0.008536 |
| 44 | FIBROBLAST GROWTH FACTOR BINDING | 2 | 23 | 0.0004043 | 0.008536 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | EXTRACELLULAR MATRIX | 7 | 426 | 8.777e-07 | 0.0005126 |
| 2 | TRANSCRIPTION FACTOR COMPLEX | 6 | 298 | 1.864e-06 | 0.0005443 |
| 3 | EXTRACELLULAR SPACE | 10 | 1376 | 4.406e-06 | 0.0008577 |
| 4 | NUCLEAR TRANSCRIPTION FACTOR COMPLEX | 4 | 127 | 2.08e-05 | 0.003037 |
| 5 | PLATELET ALPHA GRANULE LUMEN | 3 | 55 | 4.892e-05 | 0.005714 |
| 6 | WNT SIGNALOSOME | 2 | 11 | 8.874e-05 | 0.008637 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04390_Hippo_signaling_pathway | 10 | 154 | 2.605e-15 | 4.69e-13 | |
| 2 | hsa04350_TGF.beta_signaling_pathway | 6 | 85 | 1.059e-09 | 9.534e-08 | |
| 3 | hsa04520_Adherens_junction | 5 | 73 | 3.493e-08 | 2.096e-06 | |
| 4 | hsa04340_Hedgehog_signaling_pathway | 4 | 56 | 7.87e-07 | 3.542e-05 | |
| 5 | hsa04916_Melanogenesis | 4 | 101 | 8.409e-06 | 0.0003027 | |
| 6 | hsa04310_Wnt_signaling_pathway | 4 | 151 | 4.101e-05 | 0.00123 | |
| 7 | hsa04144_Endocytosis | 4 | 203 | 0.0001293 | 0.003325 | |
| 8 | hsa04010_MAPK_signaling_pathway | 4 | 268 | 0.0003736 | 0.008406 | |
| 9 | hsa04110_Cell_cycle | 3 | 128 | 0.0005978 | 0.01076 | |
| 10 | hsa04380_Osteoclast_differentiation | 3 | 128 | 0.0005978 | 0.01076 | |
| 11 | hsa04330_Notch_signaling_pathway | 2 | 47 | 0.001695 | 0.02773 | |
| 12 | hsa04510_Focal_adhesion | 3 | 200 | 0.002162 | 0.03243 | |
| 13 | hsa04151_PI3K_AKT_signaling_pathway | 3 | 351 | 0.01033 | 0.1431 | |
| 14 | hsa04014_Ras_signaling_pathway | 2 | 236 | 0.03742 | 0.449 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | RP11-166D19.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 21 | HGF | Sponge network | -3.855 | 0 | -2.968 | 0 | 0.679 |
| 2 | MAGI2-AS3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 24 | HGF | Sponge network | -2.414 | 0 | -2.968 | 0 | 0.664 |
| 3 | LINC00152 |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-361-5p;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-98-5p | 18 | HMGA2 | Sponge network | 0.948 | 0.00462 | 3.207 | 0.00037 | 0.647 |
| 4 | MIR4435-1HG |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-361-5p;hsa-miR-664a-3p;hsa-miR-664a-5p;hsa-miR-98-5p | 13 | HMGA2 | Sponge network | 1.256 | 5.0E-5 | 3.207 | 0.00037 | 0.612 |
| 5 | MIR143HG |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-34c-5p;hsa-miR-429;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-944 | 25 | HGF | Sponge network | -4.237 | 0 | -2.968 | 0 | 0.608 |
| 6 | LINC00702 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-651-5p;hsa-miR-7-1-3p | 24 | HGF | Sponge network | -2.704 | 0 | -2.968 | 0 | 0.604 |
| 7 | RP11-166D19.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-942-5p | 11 | LOXL3 | Sponge network | -3.855 | 0 | -0.968 | 0.00081 | 0.583 |
| 8 | HAND2-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-34c-5p;hsa-miR-421;hsa-miR-429;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 26 | HGF | Sponge network | -5.605 | 0 | -2.968 | 0 | 0.579 |
| 9 | DNM3OS |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-34c-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 17 | HGF | Sponge network | -2.298 | 1.0E-5 | -2.968 | 0 | 0.575 |
| 10 | CTC-296K1.3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-582-5p | 10 | HGF | Sponge network | -5.677 | 0 | -2.968 | 0 | 0.575 |
| 11 | ADAMTS9-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-34c-5p;hsa-miR-421;hsa-miR-429;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 23 | HGF | Sponge network | -7.614 | 0 | -2.968 | 0 | 0.574 |
| 12 | LUCAT1 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-93-3p | 10 | HMGA2 | Sponge network | 0.475 | 0.56236 | 3.207 | 0.00037 | 0.558 |
| 13 | RP11-439E19.10 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-340-5p;hsa-miR-34c-5p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 19 | TGFBR3 | Sponge network | -0.548 | 0.31987 | -2.591 | 0 | 0.553 |
| 14 | FENDRR |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-32-3p;hsa-miR-34c-5p;hsa-miR-3607-3p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-455-5p;hsa-miR-484;hsa-miR-590-3p;hsa-miR-7-1-3p | 31 | TGFBR3 | Sponge network | -4.793 | 0 | -2.591 | 0 | 0.553 |
| 15 | ACTA2-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-590-3p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 18 | HGF | Sponge network | -3.838 | 0 | -2.968 | 0 | 0.552 |
| 16 | LINC00704 |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-532-5p;hsa-miR-664a-3p;hsa-miR-664a-5p;hsa-miR-93-5p | 12 | HMGA2 | Sponge network | 0.568 | 0.53191 | 3.207 | 0.00037 | 0.54 |
| 17 | GAS6-AS2 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-5p;hsa-miR-651-5p;hsa-miR-7-1-3p | 16 | HGF | Sponge network | -2.655 | 0 | -2.968 | 0 | 0.528 |
| 18 | AGAP11 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-424-5p;hsa-miR-455-5p | 13 | TGFBR3 | Sponge network | -0.015 | 0.97979 | -2.591 | 0 | 0.527 |
| 19 | RP11-774O3.3 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-625-5p;hsa-miR-7-1-3p | 21 | TGFBR3 | Sponge network | -1.032 | 0.00307 | -2.591 | 0 | 0.526 |
| 20 | RP11-175K6.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-421;hsa-miR-590-5p;hsa-miR-651-5p | 11 | HGF | Sponge network | -2.386 | 0 | -2.968 | 0 | 0.523 |
| 21 | RP11-166D19.1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-335-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-944 | 28 | TGFB2 | Sponge network | -3.855 | 0 | -1.199 | 0.00331 | 0.52 |
| 22 | MAGI2-AS3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-942-5p | 10 | LOXL3 | Sponge network | -2.414 | 0 | -0.968 | 0.00081 | 0.52 |
| 23 | PWAR6 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-340-5p;hsa-miR-34c-5p;hsa-miR-409-3p;hsa-miR-450b-5p;hsa-miR-455-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 29 | TGFBR3 | Sponge network | -2.542 | 0 | -2.591 | 0 | 0.519 |
| 24 | RP11-805I24.3 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-32-3p;hsa-miR-340-5p;hsa-miR-34c-5p;hsa-miR-3607-3p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-455-5p;hsa-miR-589-3p;hsa-miR-627-5p;hsa-miR-7-1-3p | 30 | TGFBR3 | Sponge network | -5.815 | 0 | -2.591 | 0 | 0.515 |
| 25 | NR2F1-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 19 | HGF | Sponge network | -1.881 | 0 | -2.968 | 0 | 0.513 |
| 26 | DNM3OS |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b | 10 | LOXL3 | Sponge network | -2.298 | 1.0E-5 | -0.968 | 0.00081 | 0.512 |
| 27 | RP11-386M24.6 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-340-5p;hsa-miR-424-5p | 12 | TGFBR3 | Sponge network | 1.004 | 0.46474 | -2.591 | 0 | 0.509 |
| 28 | RP11-253E3.3 |
hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-532-5p;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-93-5p;hsa-miR-98-5p | 17 | HMGA2 | Sponge network | -0.01 | 0.98061 | 3.207 | 0.00037 | 0.508 |
| 29 | RP11-440I14.2 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-340-5p | 12 | TGFBR3 | Sponge network | -2.401 | 0.02461 | -2.591 | 0 | 0.506 |
| 30 | RP4-601P9.2 | hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p | 12 | TGFBR3 | Sponge network | -0.891 | 0.45058 | -2.591 | 0 | 0.502 |
| 31 | LINC00520 |
hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-195-3p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-320a;hsa-miR-497-5p;hsa-miR-664a-3p;hsa-miR-93-5p;hsa-miR-98-5p | 10 | HMGA2 | Sponge network | 1.687 | 0.08566 | 3.207 | 0.00037 | 0.495 |
| 32 | VIM-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b | 10 | LOXL3 | Sponge network | -1.424 | 0.00627 | -0.968 | 0.00081 | 0.493 |
| 33 | RP11-145A3.1 |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-532-5p;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-98-5p | 12 | HMGA2 | Sponge network | 1.041 | 0.23373 | 3.207 | 0.00037 | 0.492 |
| 34 | RP11-175K6.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-942-5p | 10 | LOXL3 | Sponge network | -2.386 | 0 | -0.968 | 0.00081 | 0.492 |
| 35 | RP11-180N14.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-141-3p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-7-1-3p;hsa-miR-944 | 10 | HGF | Sponge network | -4.46 | 0 | -2.968 | 0 | 0.489 |
| 36 | AC005682.5 |
hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-326;hsa-miR-34a-5p;hsa-miR-664a-5p;hsa-miR-93-5p;hsa-miR-98-5p | 13 | HMGA2 | Sponge network | -0.787 | 0.08468 | 3.207 | 0.00037 | 0.489 |
| 37 | NR2F1-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-34a-5p;hsa-miR-942-5p | 12 | LOXL3 | Sponge network | -1.881 | 0 | -0.968 | 0.00081 | 0.488 |
| 38 | VIM-AS1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-944 | 12 | TGFB2 | Sponge network | -1.424 | 0.00627 | -1.199 | 0.00331 | 0.484 |
| 39 | LINC00663 | hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-186-5p;hsa-miR-193a-3p;hsa-miR-21-5p;hsa-miR-424-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 10 | TGFBR3 | Sponge network | -0.813 | 0.00071 | -2.591 | 0 | 0.478 |
| 40 | RP11-253E3.3 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-3200-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-98-5p | 15 | TGFBR1 | Sponge network | -0.01 | 0.98061 | -0.532 | 0.01115 | 0.478 |
| 41 | TBX5-AS1 |
hsa-let-7a-3p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-421;hsa-miR-590-5p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 13 | HGF | Sponge network | -2.557 | 2.0E-5 | -2.968 | 0 | 0.478 |
| 42 | RP11-389G6.3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-590-3p;hsa-miR-944 | 14 | HGF | Sponge network | -7.573 | 0 | -2.968 | 0 | 0.477 |
| 43 | WDFY3-AS2 |
hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-3607-3p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-455-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 27 | TGFBR3 | Sponge network | -1.607 | 0 | -2.591 | 0 | 0.476 |
| 44 | MEG3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-34a-5p;hsa-miR-942-5p | 11 | LOXL3 | Sponge network | -2.367 | 0 | -0.968 | 0.00081 | 0.475 |
| 45 | C1orf132 | hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-340-5p;hsa-miR-34c-5p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-7-1-3p | 18 | TGFBR3 | Sponge network | -1.519 | 0.00245 | -2.591 | 0 | 0.475 |
| 46 | SNHG14 |
hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-340-5p;hsa-miR-34c-5p;hsa-miR-361-3p;hsa-miR-409-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-455-5p;hsa-miR-484;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-625-5p;hsa-miR-627-5p;hsa-miR-7-1-3p | 33 | TGFBR3 | Sponge network | -2.055 | 0 | -2.591 | 0 | 0.471 |
| 47 | ALDH1L1-AS2 | hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-424-5p;hsa-miR-590-3p | 12 | TGFBR3 | Sponge network | -1.944 | 0.08573 | -2.591 | 0 | 0.471 |
| 48 | MAGI2-AS3 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-335-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-362-3p;hsa-miR-362-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-500a-5p;hsa-miR-500b-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-944 | 29 | TGFB2 | Sponge network | -2.414 | 0 | -1.199 | 0.00331 | 0.468 |
| 49 | LINC00702 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-34a-5p;hsa-miR-942-5p | 12 | LOXL3 | Sponge network | -2.704 | 0 | -0.968 | 0.00081 | 0.466 |
| 50 | HAND2-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-34a-5p;hsa-miR-942-5p | 12 | LOXL3 | Sponge network | -5.605 | 0 | -0.968 | 0.00081 | 0.465 |
| 51 | CTD-2171N6.1 | hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-664a-5p | 14 | HMGA2 | Sponge network | 4.988 | 0 | 3.207 | 0.00037 | 0.465 |
| 52 | VIM-AS1 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-320a;hsa-miR-532-5p;hsa-miR-93-5p | 13 | HMGA2 | Sponge network | -1.424 | 0.00627 | 3.207 | 0.00037 | 0.464 |
| 53 | RP11-863P13.3 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-93-3p;hsa-miR-93-5p | 11 | HMGA2 | Sponge network | 0.637 | 0.3984 | 3.207 | 0.00037 | 0.462 |
| 54 | MEG3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-141-3p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-5p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 14 | HGF | Sponge network | -2.367 | 0 | -2.968 | 0 | 0.462 |
| 55 | RP11-594N15.3 | hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-340-5p;hsa-miR-590-3p | 14 | TGFBR3 | Sponge network | -0.887 | 0.13282 | -2.591 | 0 | 0.46 |
| 56 | RP11-54O7.3 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-16-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-455-5p;hsa-miR-589-3p;hsa-miR-625-5p;hsa-miR-627-5p | 20 | TGFBR3 | Sponge network | -2.059 | 0.00162 | -2.591 | 0 | 0.46 |
| 57 | LINC00473 |
hsa-let-7b-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-590-3p;hsa-miR-651-5p;hsa-miR-7-1-3p | 10 | HGF | Sponge network | -5.53 | 0 | -2.968 | 0 | 0.458 |
| 58 | FENDRR |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-34c-5p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-944 | 19 | HGF | Sponge network | -4.793 | 0 | -2.968 | 0 | 0.458 |
| 59 | RP11-166D19.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-944;hsa-miR-96-5p | 32 | TGFBR1 | Sponge network | -3.855 | 0 | -0.532 | 0.01115 | 0.457 |
| 60 | LINC00702 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-362-3p;hsa-miR-362-5p;hsa-miR-454-3p;hsa-miR-500a-5p;hsa-miR-500b-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p | 26 | TGFB2 | Sponge network | -2.704 | 0 | -1.199 | 0.00331 | 0.457 |
| 61 | BVES-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-429 | 11 | HGF | Sponge network | -4.161 | 1.0E-5 | -2.968 | 0 | 0.455 |
| 62 | LINC00327 |
hsa-let-7a-3p;hsa-miR-126-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-944 | 14 | HGF | Sponge network | -1.951 | 0.01135 | -2.968 | 0 | 0.453 |
| 63 | RASSF8-AS1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-944 | 18 | TGFB2 | Sponge network | -0.877 | 0.00508 | -1.199 | 0.00331 | 0.452 |
| 64 | RP11-326C3.11 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-34a-5p;hsa-miR-942-5p | 10 | LOXL3 | Sponge network | -1.196 | 3.0E-5 | -0.968 | 0.00081 | 0.452 |
| 65 | LINC01082 | hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-29a-5p;hsa-miR-32-3p;hsa-miR-484 | 10 | TGFBR3 | Sponge network | -6.023 | 0 | -2.591 | 0 | 0.45 |
| 66 | CTC-510F12.2 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-181a-5p;hsa-miR-185-3p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-34c-5p;hsa-miR-361-3p;hsa-miR-484 | 12 | TGFBR3 | Sponge network | -1.759 | 0 | -2.591 | 0 | 0.45 |
| 67 | RP11-25K19.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b | 10 | LOXL3 | Sponge network | -1.478 | 0.00829 | -0.968 | 0.00081 | 0.449 |
| 68 | FAM225B | hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-34a-5p;hsa-miR-664a-3p;hsa-miR-664a-5p | 16 | HMGA2 | Sponge network | 0.864 | 0.07672 | 3.207 | 0.00037 | 0.446 |
| 69 | ZFHX4-AS1 |
hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-944 | 19 | HGF | Sponge network | -2.966 | 0.00743 | -2.968 | 0 | 0.445 |
| 70 | RP11-145A3.1 |
hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-944;hsa-miR-98-5p | 18 | TGFBR1 | Sponge network | 1.041 | 0.23373 | -0.532 | 0.01115 | 0.445 |
| 71 | LINC00654 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 20 | HGF | Sponge network | -1.448 | 0.00044 | -2.968 | 0 | 0.444 |
| 72 | RP11-145A3.1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-944 | 13 | TGFB2 | Sponge network | 1.041 | 0.23373 | -1.199 | 0.00331 | 0.443 |
| 73 | MAGI2-AS3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-500a-5p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-769-5p;hsa-miR-944;hsa-miR-98-5p | 32 | TGFBR1 | Sponge network | -2.414 | 0 | -0.532 | 0.01115 | 0.443 |
| 74 | RASSF8-AS1 |
hsa-miR-141-3p;hsa-miR-200a-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-944 | 13 | HGF | Sponge network | -0.877 | 0.00508 | -2.968 | 0 | 0.443 |
| 75 | AF131217.1 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-339-5p;hsa-miR-340-5p;hsa-miR-34c-5p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-627-5p;hsa-miR-7-1-3p | 30 | TGFBR3 | Sponge network | -5.31 | 0 | -2.591 | 0 | 0.442 |
| 76 | AC002480.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-3200-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-664a-3p;hsa-miR-98-5p | 18 | TGFBR1 | Sponge network | -1.522 | 0.03484 | -0.532 | 0.01115 | 0.442 |
| 77 | GAS6-AS2 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-429;hsa-miR-590-5p;hsa-miR-664a-3p | 20 | TGFBR1 | Sponge network | -2.655 | 0 | -0.532 | 0.01115 | 0.441 |
| 78 | RP11-863P13.1 | hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-93-3p | 10 | HMGA2 | Sponge network | 1.292 | 0.24187 | 3.207 | 0.00037 | 0.439 |
| 79 | HOTTIP | hsa-let-7i-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-340-5p;hsa-miR-34c-5p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-455-5p;hsa-miR-484;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-627-5p | 26 | TGFBR3 | Sponge network | -1.941 | 0.00419 | -2.591 | 0 | 0.439 |
| 80 | RP11-867G23.10 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-34a-5p;hsa-miR-942-5p | 10 | LOXL3 | Sponge network | -5.684 | 0 | -0.968 | 0.00081 | 0.438 |
| 81 | RP6-91H8.3 |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-361-5p;hsa-miR-664a-5p;hsa-miR-98-5p | 15 | HMGA2 | Sponge network | 0.534 | 0.59239 | 3.207 | 0.00037 | 0.437 |
| 82 | MIR143HG |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-34a-5p;hsa-miR-942-5p | 12 | LOXL3 | Sponge network | -4.237 | 0 | -0.968 | 0.00081 | 0.437 |
| 83 | LINC00702 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-454-3p;hsa-miR-500a-5p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-96-5p | 28 | TGFBR1 | Sponge network | -2.704 | 0 | -0.532 | 0.01115 | 0.434 |
| 84 | RBPMS-AS1 | hsa-let-7i-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-424-5p;hsa-miR-7-1-3p | 12 | TGFBR3 | Sponge network | -0.791 | 0.03849 | -2.591 | 0 | 0.432 |
| 85 | RP11-499E18.1 | hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-664a-5p | 11 | HMGA2 | Sponge network | -0.083 | 0.80148 | 3.207 | 0.00037 | 0.432 |
| 86 | PDZRN3-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-942-5p | 10 | LOXL3 | Sponge network | -5.049 | 1.0E-5 | -0.968 | 0.00081 | 0.431 |
| 87 | NR2F1-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-362-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-500b-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-944 | 26 | TGFB2 | Sponge network | -1.881 | 0 | -1.199 | 0.00331 | 0.431 |
| 88 | RP11-789C17.1 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-340-5p;hsa-miR-409-3p;hsa-miR-424-5p;hsa-miR-455-5p | 12 | TGFBR3 | Sponge network | -0.395 | 0.5787 | -2.591 | 0 | 0.431 |
| 89 | RP11-693J15.4 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-582-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 14 | HGF | Sponge network | -3.319 | 0.00281 | -2.968 | 0 | 0.429 |
| 90 | RP11-253E3.3 |
hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p | 10 | TGFB2 | Sponge network | -0.01 | 0.98061 | -1.199 | 0.00331 | 0.427 |
| 91 | RP11-531A24.5 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-339-5p;hsa-miR-3607-3p;hsa-miR-625-5p;hsa-miR-627-5p;hsa-miR-7-1-3p | 25 | TGFBR3 | Sponge network | -1.752 | 0 | -2.591 | 0 | 0.427 |
| 92 | GAS6-AS2 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-5p | 14 | TGFB2 | Sponge network | -2.655 | 0 | -1.199 | 0.00331 | 0.427 |
| 93 | LINC00883 |
hsa-miR-1226-3p;hsa-miR-148b-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-192-5p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-30d-3p;hsa-miR-505-3p;hsa-miR-576-5p | 10 | LIMS1 | Sponge network | -0.614 | 0.0511 | -0.388 | 0.14814 | 0.426 |
| 94 | ACTA2-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-34a-5p;hsa-miR-942-5p | 10 | LOXL3 | Sponge network | -3.838 | 0 | -0.968 | 0.00081 | 0.424 |
| 95 | LINC00672 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-34c-5p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-590-3p | 14 | TGFBR3 | Sponge network | -1.068 | 0.00048 | -2.591 | 0 | 0.424 |
| 96 | NDUFA6-AS1 | hsa-let-7i-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-424-5p;hsa-miR-627-5p | 11 | TGFBR3 | Sponge network | -0.691 | 0.00063 | -2.591 | 0 | 0.422 |
| 97 | PGM5-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-7-1-3p | 15 | HGF | Sponge network | -11.072 | 0 | -2.968 | 0 | 0.422 |
| 98 | FAM225A | hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-664a-5p | 13 | HMGA2 | Sponge network | 1.173 | 0.062 | 3.207 | 0.00037 | 0.419 |
| 99 | RP11-284N8.3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-582-5p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-944 | 12 | HGF | Sponge network | -1.414 | 0.007 | -2.968 | 0 | 0.418 |
| 100 | DNM3OS |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-5p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-944 | 23 | TGFB2 | Sponge network | -2.298 | 1.0E-5 | -1.199 | 0.00331 | 0.418 |
| 101 | RP11-887P2.5 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-32-3p;hsa-miR-339-5p;hsa-miR-340-5p;hsa-miR-34c-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 24 | TGFBR3 | Sponge network | -6.751 | 0 | -2.591 | 0 | 0.412 |
| 102 | RP5-1159O4.2 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-340-5p;hsa-miR-455-5p | 10 | TGFBR3 | Sponge network | -0.035 | 0.94377 | -2.591 | 0 | 0.412 |
| 103 | ADAMTS9-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-34a-5p;hsa-miR-942-5p | 11 | LOXL3 | Sponge network | -7.614 | 0 | -0.968 | 0.00081 | 0.408 |
| 104 | RP11-890B15.2 |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-326;hsa-miR-330-3p;hsa-miR-532-5p | 11 | HMGA2 | Sponge network | 0.169 | 0.82185 | 3.207 | 0.00037 | 0.408 |
| 105 | AC002480.5 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-331-3p | 11 | TGFBR1 | Sponge network | -2.128 | 0.02032 | -0.532 | 0.01115 | 0.406 |
| 106 | RP11-88I18.3 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-222-5p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-3607-3p;hsa-miR-450b-5p;hsa-miR-455-5p;hsa-miR-484 | 16 | TGFBR3 | Sponge network | -7.88 | 0 | -2.591 | 0 | 0.405 |
| 107 | LINC00883 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-576-5p;hsa-miR-664a-3p | 15 | TGFBR1 | Sponge network | -0.614 | 0.0511 | -0.532 | 0.01115 | 0.405 |
| 108 | RP11-326C3.11 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-330-5p | 11 | TGFB2 | Sponge network | -1.196 | 3.0E-5 | -1.199 | 0.00331 | 0.404 |
| 109 | RP11-426C22.4 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-374a-5p | 14 | TGFBR1 | Sponge network | -0.045 | 0.93351 | -0.532 | 0.01115 | 0.404 |
| 110 | LINC00473 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-34a-5p;hsa-miR-942-5p | 12 | LOXL3 | Sponge network | -5.53 | 0 | -0.968 | 0.00081 | 0.404 |
| 111 | RP11-417E7.1 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p | 11 | TGFBR1 | Sponge network | 1.513 | 0.0271 | -0.532 | 0.01115 | 0.404 |
| 112 | RP4-735C1.4 | hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-16-5p;hsa-miR-181b-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-455-5p | 11 | TGFBR3 | Sponge network | -2.719 | 0.06688 | -2.591 | 0 | 0.403 |
| 113 | LINC00865 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-339-5p;hsa-miR-34c-5p;hsa-miR-3607-3p;hsa-miR-409-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-7-1-3p | 28 | TGFBR3 | Sponge network | -2.379 | 0 | -2.591 | 0 | 0.401 |
| 114 | RP11-456K23.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b | 10 | LOXL3 | Sponge network | -1.962 | 1.0E-5 | -0.968 | 0.00081 | 0.4 |
| 115 | RP11-311F12.1 | hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-326;hsa-miR-34a-5p;hsa-miR-532-5p;hsa-miR-664a-5p;hsa-miR-98-5p | 12 | HMGA2 | Sponge network | 0.733 | 0.47246 | 3.207 | 0.00037 | 0.398 |
| 116 | RP11-456K23.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-590-5p | 14 | HGF | Sponge network | -1.962 | 1.0E-5 | -2.968 | 0 | 0.398 |
| 117 | RP1-65J11.1 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-32-3p;hsa-miR-339-5p;hsa-miR-3607-3p;hsa-miR-455-5p;hsa-miR-7-1-3p | 23 | TGFBR3 | Sponge network | -5.989 | 0 | -2.591 | 0 | 0.398 |
| 118 | RP11-750H9.5 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-374a-5p | 13 | TGFBR1 | Sponge network | -1.207 | 0.04802 | -0.532 | 0.01115 | 0.397 |
| 119 | RP11-963H4.3 | hsa-let-7b-5p;hsa-miR-16-5p;hsa-miR-181b-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-590-5p | 13 | TGFBR3 | Sponge network | -3.737 | 0.00111 | -2.591 | 0 | 0.397 |
| 120 | ST3GAL6-AS1 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-330-3p | 11 | TGFBR1 | Sponge network | -0.653 | 0.51876 | -0.532 | 0.01115 | 0.397 |
| 121 | RP11-81H14.2 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-582-5p;hsa-miR-7-1-3p;hsa-miR-944 | 17 | HGF | Sponge network | -2.322 | 0.00014 | -2.968 | 0 | 0.396 |
| 122 | EMX2OS | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-34c-5p;hsa-miR-424-5p;hsa-miR-455-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-627-5p;hsa-miR-7-1-3p | 23 | TGFBR3 | Sponge network | -2.206 | 0.03389 | -2.591 | 0 | 0.395 |
| 123 | ST3GAL6-AS1 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-330-3p | 10 | HMGA2 | Sponge network | -0.653 | 0.51876 | 3.207 | 0.00037 | 0.395 |
| 124 | RP11-346C20.3 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-409-3p;hsa-miR-484;hsa-miR-590-5p | 18 | TGFBR3 | Sponge network | -0.334 | 0.27675 | -2.591 | 0 | 0.395 |
| 125 | RP11-356I2.4 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-664a-3p | 15 | TGFBR1 | Sponge network | -0.68 | 0.02565 | -0.532 | 0.01115 | 0.395 |
| 126 | RASSF8-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-429;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-769-5p;hsa-miR-944 | 22 | TGFBR1 | Sponge network | -0.877 | 0.00508 | -0.532 | 0.01115 | 0.394 |
| 127 | RP13-514E23.1 |
hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-32-3p;hsa-miR-340-5p;hsa-miR-409-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-484;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-625-5p;hsa-miR-7-1-3p | 27 | TGFBR3 | Sponge network | -2.158 | 0.0006 | -2.591 | 0 | 0.393 |
| 128 | CTB-61M7.2 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-769-5p;hsa-miR-944;hsa-miR-96-5p;hsa-miR-98-5p | 15 | TGFBR1 | Sponge network | -1.52 | 0.16098 | -0.532 | 0.01115 | 0.392 |
| 129 | CTB-12O2.1 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p | 11 | TGFBR3 | Sponge network | 0.106 | 0.91025 | -2.591 | 0 | 0.392 |
| 130 | RP11-35G9.5 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-454-3p;hsa-miR-589-3p | 14 | TGFB2 | Sponge network | -0.537 | 0.0574 | -1.199 | 0.00331 | 0.392 |
| 131 | DNM3OS |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-944;hsa-miR-96-5p | 27 | TGFBR1 | Sponge network | -2.298 | 1.0E-5 | -0.532 | 0.01115 | 0.391 |
| 132 | RP11-307B6.3 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-424-5p;hsa-miR-589-3p | 18 | TGFBR3 | Sponge network | -5.373 | 4.0E-5 | -2.591 | 0 | 0.391 |
| 133 | RP11-384P7.7 |
hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 15 | HGF | Sponge network | -3.649 | 3.0E-5 | -2.968 | 0 | 0.39 |
| 134 | SH3BP5-AS1 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-340-5p;hsa-miR-424-5p | 11 | TGFBR3 | Sponge network | -0.031 | 0.9308 | -2.591 | 0 | 0.389 |
| 135 | LINC00152 |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-2110;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-98-5p | 10 | TGFBR1 | Sponge network | 0.948 | 0.00462 | -0.532 | 0.01115 | 0.388 |
| 136 | RP11-214O1.2 | hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-5p;hsa-miR-29b-2-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-98-5p | 11 | HMGA2 | Sponge network | 0.855 | 0.28125 | 3.207 | 0.00037 | 0.388 |
| 137 | RP11-867G23.10 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-330-5p;hsa-miR-361-5p;hsa-miR-500a-5p;hsa-miR-500b-5p;hsa-miR-589-3p | 10 | TGFB2 | Sponge network | -5.684 | 0 | -1.199 | 0.00331 | 0.388 |
| 138 | RP11-401O9.4 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-3200-3p;hsa-miR-330-3p;hsa-miR-590-5p | 12 | TGFBR1 | Sponge network | -0.359 | 0.78702 | -0.532 | 0.01115 | 0.388 |
| 139 | FRMD6-AS2 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-944 | 11 | HGF | Sponge network | -6.455 | 0 | -2.968 | 0 | 0.388 |
| 140 | LINC00565 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-944 | 10 | TGFB2 | Sponge network | -1.493 | 0.05998 | -1.199 | 0.00331 | 0.387 |
| 141 | RP4-800J21.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-34a-5p;hsa-miR-942-5p | 10 | LOXL3 | Sponge network | -2.473 | 0.00577 | -0.968 | 0.00081 | 0.387 |
| 142 | RP1-151F17.2 |
hsa-miR-1226-3p;hsa-miR-148b-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-192-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-96-5p | 13 | LIMS1 | Sponge network | -1.606 | 0 | -0.388 | 0.14814 | 0.386 |
| 143 | MIR143HG |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-331-5p;hsa-miR-335-3p;hsa-miR-335-5p;hsa-miR-33a-3p;hsa-miR-362-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-500a-5p;hsa-miR-500b-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-944 | 29 | TGFB2 | Sponge network | -4.237 | 0 | -1.199 | 0.00331 | 0.386 |
| 144 | LINC00996 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-34a-5p | 10 | LOXL3 | Sponge network | -1.208 | 0.03775 | -0.968 | 0.00081 | 0.386 |
| 145 | TMEM51-AS1 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-340-5p;hsa-miR-409-3p;hsa-miR-424-5p;hsa-miR-627-5p | 11 | TGFBR3 | Sponge network | -0.057 | 0.90024 | -2.591 | 0 | 0.381 |
| 146 | LINC00163 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-362-3p;hsa-miR-589-3p | 15 | TGFB2 | Sponge network | -4.312 | 0.00014 | -1.199 | 0.00331 | 0.38 |
| 147 | BDNF-AS | hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-340-5p;hsa-miR-361-3p;hsa-miR-409-3p;hsa-miR-484;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-7-1-3p | 22 | TGFBR3 | Sponge network | -0.773 | 0.01202 | -2.591 | 0 | 0.379 |
| 148 | RP11-720L2.4 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-2110 | 13 | TGFBR1 | Sponge network | -3.686 | 2.0E-5 | -0.532 | 0.01115 | 0.379 |
| 149 | BVES-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-942-5p | 11 | LOXL3 | Sponge network | -4.161 | 1.0E-5 | -0.968 | 0.00081 | 0.379 |
| 150 | CTB-176F20.3 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-340-5p;hsa-miR-34c-5p | 11 | TGFBR3 | Sponge network | 0.79 | 0.14954 | -2.591 | 0 | 0.378 |
| 151 | RP1-151F17.2 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-5p;hsa-miR-335-3p;hsa-miR-335-5p;hsa-miR-33a-3p;hsa-miR-374a-5p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-590-5p | 18 | TGFB2 | Sponge network | -1.606 | 0 | -1.199 | 0.00331 | 0.377 |
| 152 | LINC00163 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-589-3p;hsa-miR-664a-3p | 17 | TGFBR1 | Sponge network | -4.312 | 0.00014 | -0.532 | 0.01115 | 0.376 |
| 153 | RP11-532F6.3 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-590-3p | 11 | TGFB2 | Sponge network | -1.772 | 1.0E-5 | -1.199 | 0.00331 | 0.375 |
| 154 | LUCAT1 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-429;hsa-miR-96-5p | 11 | TGFBR1 | Sponge network | 0.475 | 0.56236 | -0.532 | 0.01115 | 0.373 |
| 155 | VIM-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-944 | 17 | TGFBR1 | Sponge network | -1.424 | 0.00627 | -0.532 | 0.01115 | 0.371 |
| 156 | APCDD1L-AS1 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-3200-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-664a-3p;hsa-miR-96-5p | 24 | TGFBR1 | Sponge network | -2.022 | 0.00702 | -0.532 | 0.01115 | 0.371 |
| 157 | RP11-356I2.4 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-330-3p;hsa-miR-34a-5p;hsa-miR-664a-3p;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-93-5p | 15 | HMGA2 | Sponge network | -0.68 | 0.02565 | 3.207 | 0.00037 | 0.369 |
| 158 | MIR497HG |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-32-3p;hsa-miR-339-5p;hsa-miR-34c-5p;hsa-miR-3607-3p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-625-5p;hsa-miR-7-1-3p | 29 | TGFBR3 | Sponge network | -3.802 | 0 | -2.591 | 0 | 0.367 |
| 159 | RP4-794H19.1 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b | 13 | TGFBR1 | Sponge network | -0.296 | 0.38999 | -0.532 | 0.01115 | 0.367 |
| 160 | LINC00641 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-361-3p;hsa-miR-409-3p;hsa-miR-424-5p;hsa-miR-484;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-627-5p;hsa-miR-7-1-3p | 27 | TGFBR3 | Sponge network | -1.851 | 0 | -2.591 | 0 | 0.366 |
| 161 | RP11-1006G14.4 | hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p;hsa-miR-7-1-3p | 16 | TGFBR3 | Sponge network | -0.747 | 1.0E-5 | -2.591 | 0 | 0.365 |
| 162 | APCDD1L-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-335-3p;hsa-miR-589-3p | 18 | TGFB2 | Sponge network | -2.022 | 0.00702 | -1.199 | 0.00331 | 0.365 |
| 163 | RP11-701P16.5 | hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-34a-5p;hsa-miR-93-3p;hsa-miR-93-5p | 11 | HMGA2 | Sponge network | -0.53 | 0.58778 | 3.207 | 0.00037 | 0.363 |
| 164 | RP11-359B12.2 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-582-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 19 | HGF | Sponge network | -1.336 | 0 | -2.968 | 0 | 0.363 |
| 165 | RP11-401O9.3 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p | 12 | TGFBR1 | Sponge network | -0.262 | 0.83673 | -0.532 | 0.01115 | 0.363 |
| 166 | LINC00242 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-326;hsa-miR-330-3p;hsa-miR-34a-5p;hsa-miR-93-3p | 11 | HMGA2 | Sponge network | -0.58 | 0.1728 | 3.207 | 0.00037 | 0.363 |
| 167 | AC009948.5 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-532-5p;hsa-miR-664a-3p;hsa-miR-664a-5p | 16 | HMGA2 | Sponge network | -0.099 | 0.70721 | 3.207 | 0.00037 | 0.363 |
| 168 | RP11-887P2.5 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-181b-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30e-5p;hsa-miR-33a-3p;hsa-miR-34c-5p;hsa-miR-421;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-944 | 17 | HGF | Sponge network | -6.751 | 0 | -2.968 | 0 | 0.362 |
| 169 | RP11-359E10.1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-330-5p;hsa-miR-331-5p;hsa-miR-335-3p;hsa-miR-361-5p | 12 | TGFB2 | Sponge network | -1.216 | 0.0438 | -1.199 | 0.00331 | 0.362 |
| 170 | RP11-1049A21.2 |
hsa-miR-126-5p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-32-3p;hsa-miR-500a-5p;hsa-miR-664a-3p;hsa-miR-769-5p;hsa-miR-96-5p | 10 | TGFBR1 | Sponge network | -4.749 | 0.00016 | -0.532 | 0.01115 | 0.362 |
| 171 | AC002480.3 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-200a-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-324-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-454-3p | 14 | TGFB2 | Sponge network | -1.522 | 0.03484 | -1.199 | 0.00331 | 0.361 |
| 172 | RP11-16N11.2 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-16-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-424-5p;hsa-miR-455-5p | 13 | TGFBR3 | Sponge network | -0.498 | 0.19566 | -2.591 | 0 | 0.361 |
| 173 | PGM5-AS1 |
hsa-miR-16-2-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-3607-3p;hsa-miR-450b-5p;hsa-miR-484;hsa-miR-7-1-3p | 16 | TGFBR3 | Sponge network | -11.072 | 0 | -2.591 | 0 | 0.36 |
| 174 | LINC00242 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-362-5p;hsa-miR-500b-5p;hsa-miR-590-5p | 17 | TGFB2 | Sponge network | -0.58 | 0.1728 | -1.199 | 0.00331 | 0.359 |
| 175 | RP11-150O12.3 | hsa-let-7b-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-32-3p;hsa-miR-339-5p;hsa-miR-34c-5p;hsa-miR-7-1-3p | 14 | TGFBR3 | Sponge network | -2.41 | 0.03971 | -2.591 | 0 | 0.359 |
| 176 | ZFHX4-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-769-5p;hsa-miR-944 | 22 | TGFBR1 | Sponge network | -2.966 | 0.00743 | -0.532 | 0.01115 | 0.359 |
| 177 | PWRN1 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-340-5p;hsa-miR-424-5p;hsa-miR-627-5p | 14 | TGFBR3 | Sponge network | -1.872 | 0.13697 | -2.591 | 0 | 0.357 |
| 178 | NR2F1-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-3200-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-944;hsa-miR-98-5p | 30 | TGFBR1 | Sponge network | -1.881 | 0 | -0.532 | 0.01115 | 0.357 |
| 179 | MIR497HG |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-34c-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 14 | HGF | Sponge network | -3.802 | 0 | -2.968 | 0 | 0.356 |
| 180 | RP11-120J1.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-651-5p | 11 | HGF | Sponge network | -6.067 | 0 | -2.968 | 0 | 0.356 |
| 181 | LOXL1-AS1 |
hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-340-5p;hsa-miR-424-5p;hsa-miR-484;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-625-5p;hsa-miR-627-5p;hsa-miR-7-1-3p | 25 | TGFBR3 | Sponge network | -0.968 | 0.00099 | -2.591 | 0 | 0.356 |
| 182 | RP4-647J21.1 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-301a-3p;hsa-miR-32-3p | 11 | TGFBR1 | Sponge network | -0.501 | 0.33476 | -0.532 | 0.01115 | 0.356 |
| 183 | LINC00861 |
hsa-let-7a-3p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-944 | 12 | HGF | Sponge network | -1.254 | 0.02528 | -2.968 | 0 | 0.355 |
| 184 | RP11-244O19.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-590-3p;hsa-miR-769-5p | 18 | TGFBR1 | Sponge network | -0.869 | 0.00571 | -0.532 | 0.01115 | 0.354 |
| 185 | LINC00883 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-33a-3p | 14 | TGFB2 | Sponge network | -0.614 | 0.0511 | -1.199 | 0.00331 | 0.354 |
| 186 | RP11-400N13.3 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-664a-3p | 10 | TGFBR1 | Sponge network | 5.002 | 0.00017 | -0.532 | 0.01115 | 0.353 |
| 187 | RP11-326C3.11 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-331-3p | 14 | TGFBR1 | Sponge network | -1.196 | 3.0E-5 | -0.532 | 0.01115 | 0.353 |
| 188 | RP11-426C22.4 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-374a-5p | 13 | TGFB2 | Sponge network | -0.045 | 0.93351 | -1.199 | 0.00331 | 0.35 |
| 189 | DIO3OS |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-34c-5p;hsa-miR-582-5p | 12 | HGF | Sponge network | -3.619 | 0 | -2.968 | 0 | 0.349 |
| 190 | SERHL | hsa-let-7i-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-424-5p;hsa-miR-625-5p;hsa-miR-627-5p | 10 | TGFBR3 | Sponge network | -0.636 | 0.05212 | -2.591 | 0 | 0.348 |
| 191 | RP11-356J5.12 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-335-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-362-3p;hsa-miR-374a-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p | 25 | TGFB2 | Sponge network | -2.015 | 0 | -1.199 | 0.00331 | 0.347 |
| 192 | RP11-44N12.5 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-32-3p;hsa-miR-576-5p;hsa-miR-944 | 11 | TGFBR1 | Sponge network | -0.426 | 0.67852 | -0.532 | 0.01115 | 0.346 |
| 193 | HCG11 |
hsa-miR-1226-3p;hsa-miR-148b-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-30d-3p;hsa-miR-374a-3p;hsa-miR-576-5p;hsa-miR-590-3p | 10 | LIMS1 | Sponge network | -1.194 | 0.00331 | -0.388 | 0.14814 | 0.346 |
| 194 | RP11-140I16.3 | hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-185-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p | 10 | TGFBR3 | Sponge network | -0.674 | 0.08468 | -2.591 | 0 | 0.346 |
| 195 | RP11-531A24.5 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-651-5p;hsa-miR-7-1-3p;hsa-miR-944 | 19 | HGF | Sponge network | -1.752 | 0 | -2.968 | 0 | 0.345 |
| 196 | LINC00883 |
hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-34a-5p;hsa-miR-532-5p;hsa-miR-664a-3p;hsa-miR-93-3p | 10 | HMGA2 | Sponge network | -0.614 | 0.0511 | 3.207 | 0.00037 | 0.344 |
| 197 | LINC00327 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-362-3p;hsa-miR-500b-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-944 | 20 | TGFB2 | Sponge network | -1.951 | 0.01135 | -1.199 | 0.00331 | 0.343 |
| 198 | AC002066.1 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-664a-3p;hsa-miR-98-5p | 12 | TGFBR1 | Sponge network | -1.314 | 0.04594 | -0.532 | 0.01115 | 0.341 |
| 199 | RP11-536K7.3 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-374a-5p;hsa-miR-590-3p | 12 | TGFB2 | Sponge network | -0.673 | 0.17143 | -1.199 | 0.00331 | 0.34 |
| 200 | MIR22HG |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-32-3p;hsa-miR-424-5p;hsa-miR-484;hsa-miR-590-5p;hsa-miR-625-5p;hsa-miR-627-5p | 22 | TGFBR3 | Sponge network | -1.994 | 0 | -2.591 | 0 | 0.34 |
| 201 | RP11-426C22.5 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-589-3p | 13 | TGFB2 | Sponge network | -0.559 | 0.08048 | -1.199 | 0.00331 | 0.34 |
| 202 | AC093627.8 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-331-3p | 11 | TGFBR1 | Sponge network | -5.744 | 0 | -0.532 | 0.01115 | 0.34 |
| 203 | RP11-13K12.5 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-3607-3p;hsa-miR-424-5p;hsa-miR-589-3p;hsa-miR-625-5p;hsa-miR-7-1-3p | 24 | TGFBR3 | Sponge network | -4.086 | 0 | -2.591 | 0 | 0.339 |
| 204 | MIR143HG |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-34c-5p;hsa-miR-3607-3p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-455-5p;hsa-miR-484;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-625-5p;hsa-miR-627-5p;hsa-miR-7-1-3p | 33 | TGFBR3 | Sponge network | -4.237 | 0 | -2.591 | 0 | 0.339 |
| 205 | AC104654.2 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-324-5p;hsa-miR-944 | 10 | TGFB2 | Sponge network | -3.012 | 0.00731 | -1.199 | 0.00331 | 0.339 |
| 206 | CTD-2554C21.2 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-34c-5p;hsa-miR-424-5p | 22 | TGFBR3 | Sponge network | -2.822 | 0.00245 | -2.591 | 0 | 0.339 |
| 207 | TP73-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-7-1-3p | 15 | HGF | Sponge network | -1.97 | 0 | -2.968 | 0 | 0.339 |
| 208 | AF131215.2 | hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-455-5p;hsa-miR-589-3p;hsa-miR-7-1-3p | 16 | TGFBR3 | Sponge network | -1.455 | 0.00692 | -2.591 | 0 | 0.336 |
| 209 | RP11-175K6.1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-5p;hsa-miR-374a-5p;hsa-miR-590-5p | 12 | TGFB2 | Sponge network | -2.386 | 0 | -1.199 | 0.00331 | 0.335 |
| 210 | RP11-696N14.1 | hsa-let-7i-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-338-3p;hsa-miR-340-5p;hsa-miR-409-3p | 10 | TGFBR3 | Sponge network | -0.09 | 0.83006 | -2.591 | 0 | 0.335 |
| 211 | RP11-536O18.1 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-330-3p | 12 | TGFBR1 | Sponge network | -5.37 | 4.0E-5 | -0.532 | 0.01115 | 0.334 |
| 212 | RP11-6O2.3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-590-5p;hsa-miR-944 | 11 | HGF | Sponge network | -4.533 | 0 | -2.968 | 0 | 0.334 |
| 213 | RP11-536K7.3 |
hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-200a-3p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-582-5p;hsa-miR-590-3p | 10 | HGF | Sponge network | -0.673 | 0.17143 | -2.968 | 0 | 0.332 |
| 214 | DENND5B-AS1 | hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-320a;hsa-miR-326;hsa-miR-664a-5p;hsa-miR-93-5p | 13 | HMGA2 | Sponge network | -0.041 | 0.91492 | 3.207 | 0.00037 | 0.331 |
| 215 | RP11-448G15.3 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15b-5p;hsa-miR-193b-3p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-34c-5p;hsa-miR-424-5p;hsa-miR-484 | 10 | TGFBR3 | Sponge network | -0.654 | 0.05236 | -2.591 | 0 | 0.331 |
| 216 | RP11-20J15.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-361-5p | 15 | TGFBR1 | Sponge network | -5.104 | 8.0E-5 | -0.532 | 0.01115 | 0.331 |
| 217 | AC005682.5 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-330-5p;hsa-miR-335-5p;hsa-miR-33a-3p;hsa-miR-362-5p;hsa-miR-454-3p | 14 | TGFB2 | Sponge network | -0.787 | 0.08468 | -1.199 | 0.00331 | 0.33 |
| 218 | RP11-62F24.2 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-32-3p;hsa-miR-589-3p | 12 | TGFBR1 | Sponge network | -4.034 | 0.00034 | -0.532 | 0.01115 | 0.33 |
| 219 | RP4-798P15.3 |
hsa-miR-148b-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-32-3p;hsa-miR-374a-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-96-5p | 13 | LIMS1 | Sponge network | -0.958 | 0.06952 | -0.388 | 0.14814 | 0.328 |
| 220 | RP11-456K23.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-429;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p | 23 | TGFBR1 | Sponge network | -1.962 | 1.0E-5 | -0.532 | 0.01115 | 0.328 |
| 221 | MIR4435-1HG |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-361-5p;hsa-miR-429;hsa-miR-664a-3p;hsa-miR-769-5p;hsa-miR-98-5p | 12 | TGFBR1 | Sponge network | 1.256 | 5.0E-5 | -0.532 | 0.01115 | 0.327 |
| 222 | MEG3 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-331-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-944 | 22 | TGFB2 | Sponge network | -2.367 | 0 | -1.199 | 0.00331 | 0.327 |
| 223 | ACTA2-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-362-5p;hsa-miR-454-3p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-944 | 21 | TGFB2 | Sponge network | -3.838 | 0 | -1.199 | 0.00331 | 0.327 |
| 224 | RP11-20J15.3 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-361-5p | 12 | TGFB2 | Sponge network | -5.104 | 8.0E-5 | -1.199 | 0.00331 | 0.327 |
| 225 | LINC01010 | hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-664a-3p;hsa-miR-93-5p | 12 | HMGA2 | Sponge network | 0.747 | 0.44797 | 3.207 | 0.00037 | 0.326 |
| 226 | APCDD1L-AS1 |
hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-7-1-3p | 10 | HGF | Sponge network | -2.022 | 0.00702 | -2.968 | 0 | 0.326 |
| 227 | RP11-412D9.4 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-330-3p;hsa-miR-590-3p | 10 | TGFB2 | Sponge network | -0.793 | 0.00544 | -1.199 | 0.00331 | 0.325 |
| 228 | RP5-1172A22.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-361-5p | 16 | TGFBR1 | Sponge network | -3.373 | 0.00572 | -0.532 | 0.01115 | 0.324 |
| 229 | ADAMTS9-AS1 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-34c-5p;hsa-miR-3607-3p;hsa-miR-361-3p;hsa-miR-455-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-625-5p;hsa-miR-627-5p;hsa-miR-7-1-3p | 30 | TGFBR3 | Sponge network | -7.614 | 0 | -2.591 | 0 | 0.324 |
| 230 | USP3-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-651-5p;hsa-miR-944 | 13 | HGF | Sponge network | -1.454 | 0 | -2.968 | 0 | 0.324 |
| 231 | LINC00365 | hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-340-5p;hsa-miR-484;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-627-5p;hsa-miR-7-1-3p | 19 | TGFBR3 | Sponge network | -1.81 | 0.03876 | -2.591 | 0 | 0.323 |
| 232 | MIR143HG |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-3200-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-500a-5p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-769-5p;hsa-miR-944;hsa-miR-98-5p | 32 | TGFBR1 | Sponge network | -4.237 | 0 | -0.532 | 0.01115 | 0.323 |
| 233 | RP11-120J1.1 |
hsa-let-7b-5p;hsa-miR-16-2-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p | 13 | TGFBR3 | Sponge network | -6.067 | 0 | -2.591 | 0 | 0.323 |
| 234 | RP11-25K19.1 |
hsa-miR-130b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-944 | 10 | TGFB2 | Sponge network | -1.478 | 0.00829 | -1.199 | 0.00331 | 0.322 |
| 235 | RP11-244O19.1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29c-3p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-590-3p | 12 | TGFB2 | Sponge network | -0.869 | 0.00571 | -1.199 | 0.00331 | 0.322 |
| 236 | RP11-815I9.4 |
hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-98-5p | 16 | TGFBR1 | Sponge network | -0.665 | 0.01108 | -0.532 | 0.01115 | 0.321 |
| 237 | PRKAG2-AS1 |
hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-141-3p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-944 | 11 | HGF | Sponge network | -2.45 | 4.0E-5 | -2.968 | 0 | 0.32 |
| 238 | LINC00840 | hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-627-5p;hsa-miR-7-1-3p | 19 | TGFBR3 | Sponge network | -1.001 | 0.27483 | -2.591 | 0 | 0.32 |
| 239 | BVES-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-429 | 11 | TGFB2 | Sponge network | -4.161 | 1.0E-5 | -1.199 | 0.00331 | 0.32 |
| 240 | AC104654.2 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-944 | 12 | TGFBR1 | Sponge network | -3.012 | 0.00731 | -0.532 | 0.01115 | 0.319 |
| 241 | RP11-175K6.1 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-484;hsa-miR-590-5p | 23 | TGFBR3 | Sponge network | -2.386 | 0 | -2.591 | 0 | 0.318 |
| 242 | AC005682.5 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-454-3p;hsa-miR-769-5p;hsa-miR-98-5p | 15 | TGFBR1 | Sponge network | -0.787 | 0.08468 | -0.532 | 0.01115 | 0.318 |
| 243 | RP11-835E18.5 | hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-32-3p;hsa-miR-576-5p | 10 | TGFBR1 | Sponge network | -0.943 | 0.00219 | -0.532 | 0.01115 | 0.318 |
| 244 | RP11-1008C21.2 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-320b;hsa-miR-330-5p;hsa-miR-361-5p | 11 | TGFB2 | Sponge network | -0.999 | 0.0012 | -1.199 | 0.00331 | 0.317 |
| 245 | AC002480.3 |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-320a;hsa-miR-664a-3p;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-93-5p;hsa-miR-98-5p | 13 | HMGA2 | Sponge network | -1.522 | 0.03484 | 3.207 | 0.00037 | 0.317 |
| 246 | RP11-805I24.3 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-34c-5p;hsa-miR-421;hsa-miR-429;hsa-miR-7-1-3p;hsa-miR-944 | 18 | HGF | Sponge network | -5.815 | 0 | -2.968 | 0 | 0.317 |
| 247 | RP5-899E9.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-98-5p | 14 | TGFBR1 | Sponge network | -1.271 | 0.10391 | -0.532 | 0.01115 | 0.317 |
| 248 | CTD-2013N24.2 |
hsa-miR-1226-3p;hsa-miR-148b-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-192-5p;hsa-miR-29b-3p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-32-3p;hsa-miR-374a-3p;hsa-miR-429;hsa-miR-576-5p;hsa-miR-590-3p | 16 | LIMS1 | Sponge network | -1.002 | 1.0E-5 | -0.388 | 0.14814 | 0.316 |
| 249 | AC009948.5 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-200b-3p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-664a-3p;hsa-miR-769-5p | 11 | TGFBR1 | Sponge network | -0.099 | 0.70721 | -0.532 | 0.01115 | 0.314 |
| 250 | AC007743.1 | hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-424-5p;hsa-miR-455-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 17 | TGFBR3 | Sponge network | -1.004 | 0.05376 | -2.591 | 0 | 0.314 |
| 251 | RP11-736K20.5 |
hsa-let-7b-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-27a-3p;hsa-miR-34c-5p;hsa-miR-589-3p;hsa-miR-590-3p | 15 | TGFBR3 | Sponge network | -1.492 | 8.0E-5 | -2.591 | 0 | 0.314 |
| 252 | C20orf166-AS1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-362-3p;hsa-miR-429;hsa-miR-454-3p | 16 | TGFB2 | Sponge network | -6.333 | 0 | -1.199 | 0.00331 | 0.313 |
| 253 | LINC00536 | hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-330-3p;hsa-miR-664a-3p;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | HMGA2 | Sponge network | 1.763 | 0.05616 | 3.207 | 0.00037 | 0.313 |
| 254 | RP11-863P13.3 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-454-3p | 12 | TGFBR1 | Sponge network | 0.637 | 0.3984 | -0.532 | 0.01115 | 0.312 |
| 255 | C4A-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-301a-3p;hsa-miR-330-5p;hsa-miR-361-5p;hsa-miR-589-3p;hsa-miR-590-5p | 11 | TGFB2 | Sponge network | -1.76 | 0.00265 | -1.199 | 0.00331 | 0.311 |
| 256 | RP11-457M11.5 | hsa-let-7i-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-340-5p;hsa-miR-424-5p | 10 | TGFBR3 | Sponge network | 0.041 | 0.93432 | -2.591 | 0 | 0.311 |
| 257 | RP11-434B12.1 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-340-5p;hsa-miR-361-3p;hsa-miR-409-3p;hsa-miR-590-3p;hsa-miR-7-1-3p | 14 | TGFBR3 | Sponge network | -1.086 | 0.00017 | -2.591 | 0 | 0.31 |
| 258 | RP1-65J11.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30c-5p;hsa-miR-30e-5p;hsa-miR-421;hsa-miR-7-1-3p;hsa-miR-944 | 15 | HGF | Sponge network | -5.989 | 0 | -2.968 | 0 | 0.309 |
| 259 | HAND2-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-362-3p;hsa-miR-362-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-500a-5p;hsa-miR-500b-5p;hsa-miR-590-3p;hsa-miR-944 | 28 | TGFB2 | Sponge network | -5.605 | 0 | -1.199 | 0.00331 | 0.309 |
| 260 | RP11-552M11.8 | hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-338-3p;hsa-miR-339-5p;hsa-miR-340-5p;hsa-miR-409-3p;hsa-miR-7-1-3p | 22 | TGFBR3 | Sponge network | -1.455 | 0.00021 | -2.591 | 0 | 0.309 |
| 261 | LINC00707 | hsa-let-7g-5p;hsa-miR-195-3p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-326;hsa-miR-664a-3p;hsa-miR-98-5p | 11 | HMGA2 | Sponge network | 0.011 | 0.99212 | 3.207 | 0.00037 | 0.309 |
| 262 | CTC-429P9.1 | hsa-let-7i-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-29a-5p | 10 | TGFBR3 | Sponge network | -0.978 | 0.0029 | -2.591 | 0 | 0.308 |
| 263 | LINC00242 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-590-5p;hsa-miR-96-5p | 17 | TGFBR1 | Sponge network | -0.58 | 0.1728 | -0.532 | 0.01115 | 0.308 |
| 264 | RP11-693J15.4 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-34a-5p;hsa-miR-942-5p | 10 | LOXL3 | Sponge network | -3.319 | 0.00281 | -0.968 | 0.00081 | 0.308 |
| 265 | RP11-35G9.5 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-454-3p;hsa-miR-589-3p;hsa-miR-769-5p;hsa-miR-96-5p | 15 | TGFBR1 | Sponge network | -0.537 | 0.0574 | -0.532 | 0.01115 | 0.306 |
| 266 | ZNF582-AS1 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-409-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-590-5p;hsa-miR-625-5p;hsa-miR-7-1-3p | 27 | TGFBR3 | Sponge network | -1.7 | 0.00019 | -2.591 | 0 | 0.305 |
| 267 | RP11-867G23.10 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-2110;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-500a-5p;hsa-miR-589-3p;hsa-miR-769-5p | 15 | TGFBR1 | Sponge network | -5.684 | 0 | -0.532 | 0.01115 | 0.305 |
| 268 | RP5-1172A22.1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-361-5p | 12 | TGFB2 | Sponge network | -3.373 | 0.00572 | -1.199 | 0.00331 | 0.305 |
| 269 | AC079767.4 |
hsa-miR-129-5p;hsa-miR-141-3p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-33a-3p;hsa-miR-944 | 10 | HGF | Sponge network | 0.594 | 0.61846 | -2.968 | 0 | 0.303 |
| 270 | AC013271.3 | hsa-let-7b-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-27a-3p;hsa-miR-340-5p;hsa-miR-484;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-627-5p;hsa-miR-7-1-3p | 19 | TGFBR3 | Sponge network | -0.291 | 0.40736 | -2.591 | 0 | 0.302 |
| 271 | RP11-378A13.1 |
hsa-let-7i-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-27a-3p;hsa-miR-32-3p;hsa-miR-339-5p;hsa-miR-409-3p;hsa-miR-424-5p;hsa-miR-590-3p | 19 | TGFBR3 | Sponge network | -0.977 | 8.0E-5 | -2.591 | 0 | 0.302 |
| 272 | RP11-426C22.5 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-589-3p;hsa-miR-664a-3p | 18 | TGFBR1 | Sponge network | -0.559 | 0.08048 | -0.532 | 0.01115 | 0.301 |
| 273 | RP1-170O19.14 | hsa-let-7i-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-627-5p | 12 | TGFBR3 | Sponge network | -1.119 | 0.055 | -2.591 | 0 | 0.301 |
| 274 | USP3-AS1 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-340-5p;hsa-miR-455-5p;hsa-miR-589-3p;hsa-miR-625-5p;hsa-miR-627-5p | 25 | TGFBR3 | Sponge network | -1.454 | 0 | -2.591 | 0 | 0.3 |
| 275 | FLG-AS1 |
hsa-miR-107;hsa-miR-148b-3p;hsa-miR-182-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-330-3p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-532-5p | 11 | SNAI2 | Sponge network | -1.812 | 0.00363 | -0.28 | 0.48988 | 0.3 |
| 276 | RP11-678G14.3 | hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-424-5p;hsa-miR-484;hsa-miR-625-5p;hsa-miR-7-1-3p | 20 | TGFBR3 | Sponge network | -4.141 | 0.0002 | -2.591 | 0 | 0.299 |
| 277 | CTD-2013N24.2 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-330-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-362-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-500a-5p;hsa-miR-500b-5p;hsa-miR-589-3p;hsa-miR-590-3p | 21 | TGFB2 | Sponge network | -1.002 | 1.0E-5 | -1.199 | 0.00331 | 0.299 |
| 278 | AP001347.6 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-3p | 17 | TGFBR3 | Sponge network | -1.554 | 0.00016 | -2.591 | 0 | 0.299 |
| 279 | RP5-899E9.1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-330-5p | 10 | TGFB2 | Sponge network | -1.271 | 0.10391 | -1.199 | 0.00331 | 0.299 |
| 280 | RP11-344E13.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-454-3p;hsa-miR-98-5p | 10 | TGFBR1 | Sponge network | 0.233 | 0.66682 | -0.532 | 0.01115 | 0.298 |
| 281 | CTC-429P9.5 | hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-29a-5p;hsa-miR-590-3p | 12 | TGFBR3 | Sponge network | -1.199 | 0.00202 | -2.591 | 0 | 0.297 |
| 282 | RP1-151F17.2 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-148b-3p;hsa-miR-182-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-330-3p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-96-5p | 11 | SNAI2 | Sponge network | -1.606 | 0 | -0.28 | 0.48988 | 0.297 |
| 283 | RP4-607I7.1 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | HMGA2 | Sponge network | -1.496 | 0.07885 | 3.207 | 0.00037 | 0.296 |
| 284 | APCDD1L-AS1 |
hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-532-5p;hsa-miR-664a-3p;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-93-5p | 13 | HMGA2 | Sponge network | -2.022 | 0.00702 | 3.207 | 0.00037 | 0.295 |
| 285 | RP11-91K9.1 | hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-361-5p;hsa-miR-664a-3p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | HMGA2 | Sponge network | 2.717 | 0.00521 | 3.207 | 0.00037 | 0.295 |
| 286 | RP4-639F20.1 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-222-5p;hsa-miR-29a-5p | 11 | TGFBR3 | Sponge network | -1.492 | 1.0E-5 | -2.591 | 0 | 0.295 |
| 287 | MIAT |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-34a-5p;hsa-miR-664a-5p;hsa-miR-93-5p;hsa-miR-98-5p | 13 | HMGA2 | Sponge network | 0.258 | 0.5834 | 3.207 | 0.00037 | 0.294 |
| 288 | RASSF8-AS1 |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-361-5p;hsa-miR-664a-3p;hsa-miR-93-5p | 17 | HMGA2 | Sponge network | -0.877 | 0.00508 | 3.207 | 0.00037 | 0.292 |
| 289 | RP11-384P7.7 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-3200-3p;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-429;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-944;hsa-miR-96-5p;hsa-miR-98-5p | 25 | TGFBR1 | Sponge network | -3.649 | 3.0E-5 | -0.532 | 0.01115 | 0.291 |
| 290 | RP1-151F17.2 |
hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-93-3p | 14 | HMGA2 | Sponge network | -1.606 | 0 | 3.207 | 0.00037 | 0.29 |
| 291 | RP6-91H8.3 |
hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-361-5p;hsa-miR-769-5p;hsa-miR-98-5p | 11 | TGFBR1 | Sponge network | 0.534 | 0.59239 | -0.532 | 0.01115 | 0.29 |
| 292 | MIAT |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-miR-200b-3p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-944;hsa-miR-98-5p | 10 | TGFBR1 | Sponge network | 0.258 | 0.5834 | -0.532 | 0.01115 | 0.29 |
| 293 | CTD-2013N24.2 |
hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-7-1-3p | 15 | HGF | Sponge network | -1.002 | 1.0E-5 | -2.968 | 0 | 0.289 |
| 294 | CTC-296K1.3 |
hsa-let-7b-5p;hsa-miR-16-2-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-29a-5p;hsa-miR-625-5p | 15 | TGFBR3 | Sponge network | -5.677 | 0 | -2.591 | 0 | 0.289 |
| 295 | HAND2-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-3200-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-500a-5p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-769-5p;hsa-miR-944;hsa-miR-98-5p | 32 | TGFBR1 | Sponge network | -5.605 | 0 | -0.532 | 0.01115 | 0.287 |
| 296 | RP11-1134I14.8 | hsa-let-7i-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-455-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-7-1-3p | 24 | TGFBR3 | Sponge network | -0.743 | 0.25713 | -2.591 | 0 | 0.287 |
| 297 | ZFHX4-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-324-5p;hsa-miR-330-3p;hsa-miR-331-5p;hsa-miR-33a-3p;hsa-miR-429;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-944 | 16 | TGFB2 | Sponge network | -2.966 | 0.00743 | -1.199 | 0.00331 | 0.286 |
| 298 | LINC00526 |
hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-2110;hsa-miR-664a-3p;hsa-miR-98-5p | 10 | TGFBR1 | Sponge network | -0.696 | 0.0183 | -0.532 | 0.01115 | 0.286 |
| 299 | CTD-2589M5.4 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-424-5p;hsa-miR-590-5p | 15 | TGFBR3 | Sponge network | -7.181 | 0 | -2.591 | 0 | 0.286 |
| 300 | AF131217.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30c-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-34c-5p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-7-1-3p;hsa-miR-944 | 19 | HGF | Sponge network | -5.31 | 0 | -2.968 | 0 | 0.285 |
| 301 | ZNF667-AS1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-34c-5p;hsa-miR-590-5p;hsa-miR-7-1-3p | 13 | HGF | Sponge network | -1.846 | 0.00013 | -2.968 | 0 | 0.284 |
| 302 | CTC-378H22.2 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-16-2-3p;hsa-miR-2110;hsa-miR-331-3p | 10 | TGFBR1 | Sponge network | -1.776 | 0.01283 | -0.532 | 0.01115 | 0.284 |
| 303 | C4A-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-3200-3p;hsa-miR-361-5p;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-769-5p;hsa-miR-96-5p | 19 | TGFBR1 | Sponge network | -1.76 | 0.00265 | -0.532 | 0.01115 | 0.284 |
| 304 | CTC-296K1.3 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-330-5p;hsa-miR-454-3p | 11 | TGFB2 | Sponge network | -5.677 | 0 | -1.199 | 0.00331 | 0.283 |
| 305 | RP11-368L12.1 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-29a-5p;hsa-miR-424-5p;hsa-miR-7-1-3p | 11 | TGFBR3 | Sponge network | -2.049 | 0.14175 | -2.591 | 0 | 0.282 |
| 306 | PART1 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-3607-3p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-455-5p;hsa-miR-484;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-625-5p;hsa-miR-627-5p;hsa-miR-7-1-3p | 33 | TGFBR3 | Sponge network | -3.06 | 7.0E-5 | -2.591 | 0 | 0.281 |
| 307 | HAR1A |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320b;hsa-miR-942-5p | 10 | LOXL3 | Sponge network | -2.085 | 0 | -0.968 | 0.00081 | 0.281 |
| 308 | RP13-1039J1.4 | hsa-let-7b-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-27a-3p;hsa-miR-589-3p;hsa-miR-625-5p;hsa-miR-627-5p | 15 | TGFBR3 | Sponge network | -0.399 | 0.3348 | -2.591 | 0 | 0.281 |
| 309 | RP3-439F8.1 |
hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-326;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | HMGA2 | Sponge network | -0.149 | 0.78031 | 3.207 | 0.00037 | 0.28 |
| 310 | AP000473.5 | hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-454-3p | 13 | TGFBR1 | Sponge network | -5.021 | 3.0E-5 | -0.532 | 0.01115 | 0.279 |
| 311 | RP11-400K9.4 |
hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-3607-3p;hsa-miR-7-1-3p | 18 | TGFBR3 | Sponge network | -1.449 | 0.00424 | -2.591 | 0 | 0.279 |
| 312 | GAS6-AS2 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-508-3p;hsa-miR-664a-3p;hsa-miR-664a-5p | 10 | HMGA2 | Sponge network | -2.655 | 0 | 3.207 | 0.00037 | 0.279 |
| 313 | RP11-531A24.5 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-107;hsa-miR-126-5p;hsa-miR-146b-5p;hsa-miR-186-5p;hsa-miR-193a-3p;hsa-miR-19b-1-5p;hsa-miR-324-5p;hsa-miR-338-3p;hsa-miR-628-5p | 11 | FGFR2 | Sponge network | -1.752 | 0 | -1.105 | 0.01288 | 0.279 |
| 314 | RP11-398K22.12 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-326;hsa-miR-34a-5p;hsa-miR-532-5p;hsa-miR-664a-3p;hsa-miR-664a-5p | 14 | HMGA2 | Sponge network | -0.361 | 0.28295 | 3.207 | 0.00037 | 0.279 |
| 315 | RP11-1024P17.1 |
hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-27a-3p;hsa-miR-424-5p;hsa-miR-484;hsa-miR-589-3p;hsa-miR-590-3p | 14 | TGFBR3 | Sponge network | -1.552 | 0 | -2.591 | 0 | 0.278 |
| 316 | RP11-389G6.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-125a-5p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b | 10 | LOXL3 | Sponge network | -7.573 | 0 | -0.968 | 0.00081 | 0.278 |
| 317 | LINC00565 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-576-5p;hsa-miR-664a-3p;hsa-miR-944 | 16 | TGFBR1 | Sponge network | -1.493 | 0.05998 | -0.532 | 0.01115 | 0.278 |
| 318 | RP11-356J5.12 |
hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-320a;hsa-miR-326;hsa-miR-361-5p;hsa-miR-508-3p;hsa-miR-98-5p | 14 | HMGA2 | Sponge network | -2.015 | 0 | 3.207 | 0.00037 | 0.277 |
| 319 | AC093627.10 |
hsa-let-7a-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-421;hsa-miR-429;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 17 | HGF | Sponge network | -2.338 | 8.0E-5 | -2.968 | 0 | 0.277 |
| 320 | RP4-639F20.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-96-5p | 16 | TGFBR1 | Sponge network | -1.492 | 1.0E-5 | -0.532 | 0.01115 | 0.277 |
| 321 | LINC00702 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-148b-3p;hsa-miR-182-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-532-5p;hsa-miR-96-5p | 11 | SNAI2 | Sponge network | -2.704 | 0 | -0.28 | 0.48988 | 0.277 |
| 322 | RP11-158K1.3 | hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-186-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-424-5p;hsa-miR-7-1-3p | 14 | TGFBR3 | Sponge network | -0.454 | 0.19049 | -2.591 | 0 | 0.276 |
| 323 | PCED1B-AS1 |
hsa-miR-107;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-335-3p;hsa-miR-429 | 11 | TGFB2 | Sponge network | -0.575 | 0.17488 | -1.199 | 0.00331 | 0.275 |
| 324 | RP11-753H16.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b | 12 | TGFBR1 | Sponge network | -5.702 | 0 | -0.532 | 0.01115 | 0.275 |
| 325 | RP11-360F5.1 |
hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-149-5p;hsa-miR-200a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-324-5p;hsa-miR-33a-3p;hsa-miR-429 | 10 | TGFB2 | Sponge network | -0.286 | 0.77753 | -1.199 | 0.00331 | 0.274 |
| 326 | LINC00327 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-3200-3p;hsa-miR-361-5p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-944;hsa-miR-96-5p;hsa-miR-98-5p | 23 | TGFBR1 | Sponge network | -1.951 | 0.01135 | -0.532 | 0.01115 | 0.273 |
| 327 | FENDRR |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-107;hsa-miR-126-5p;hsa-miR-146b-5p;hsa-miR-186-5p;hsa-miR-193a-3p;hsa-miR-19b-1-5p;hsa-miR-324-5p;hsa-miR-423-3p;hsa-miR-590-3p;hsa-miR-628-5p | 12 | FGFR2 | Sponge network | -4.793 | 0 | -1.105 | 0.01288 | 0.273 |
| 328 | RP11-389G6.3 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-3607-3p;hsa-miR-590-3p | 18 | TGFBR3 | Sponge network | -7.573 | 0 | -2.591 | 0 | 0.273 |
| 329 | FLG-AS1 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-3200-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-500a-5p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-769-5p | 26 | TGFBR1 | Sponge network | -1.812 | 0.00363 | -0.532 | 0.01115 | 0.272 |
| 330 | WDFY3-AS2 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-let-7f-1-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-26b-5p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p;hsa-miR-944 | 10 | HGF | Sponge network | -1.607 | 0 | -2.968 | 0 | 0.271 |
| 331 | LINC00578 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-664a-3p | 16 | TGFBR1 | Sponge network | -1.091 | 0.13719 | -0.532 | 0.01115 | 0.27 |
| 332 | C20orf166-AS1 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-339-5p;hsa-miR-3607-3p;hsa-miR-361-3p;hsa-miR-455-5p;hsa-miR-484;hsa-miR-625-5p | 23 | TGFBR3 | Sponge network | -6.333 | 0 | -2.591 | 0 | 0.27 |
| 333 | A1BG-AS1 | hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-589-3p | 10 | TGFBR1 | Sponge network | -0.309 | 0.61914 | -0.532 | 0.01115 | 0.27 |
| 334 | RP11-359B12.2 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-429;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-96-5p | 22 | TGFBR1 | Sponge network | -1.336 | 0 | -0.532 | 0.01115 | 0.27 |
| 335 | RP11-532F6.3 |
hsa-miR-1226-3p;hsa-miR-148b-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-192-5p;hsa-miR-30d-3p;hsa-miR-32-3p;hsa-miR-374a-3p;hsa-miR-505-3p;hsa-miR-576-5p;hsa-miR-590-3p | 11 | LIMS1 | Sponge network | -1.772 | 1.0E-5 | -0.388 | 0.14814 | 0.269 |
| 336 | LINC00667 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-186-5p;hsa-miR-320a;hsa-miR-330-3p;hsa-miR-454-3p;hsa-miR-664a-3p | 11 | TGFBR1 | Sponge network | -0.697 | 0.01067 | -0.532 | 0.01115 | 0.269 |
| 337 | RP11-411K7.1 |
hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-195-3p;hsa-miR-20b-5p;hsa-miR-29b-2-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-3p;hsa-miR-30e-5p;hsa-miR-326;hsa-miR-330-3p;hsa-miR-34a-5p;hsa-miR-532-5p;hsa-miR-664a-3p;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-93-5p | 19 | HMGA2 | Sponge network | -0.894 | 0.3797 | 3.207 | 0.00037 | 0.269 |
| 338 | RP11-401P9.4 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148a-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-29b-3p;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-331-5p;hsa-miR-33a-3p;hsa-miR-361-5p;hsa-miR-374a-5p;hsa-miR-454-3p;hsa-miR-590-5p;hsa-miR-944 | 18 | TGFB2 | Sponge network | -2.738 | 0 | -1.199 | 0.00331 | 0.269 |
| 339 | CTD-2013N24.2 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-3200-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-374a-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-500a-5p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p | 26 | TGFBR1 | Sponge network | -1.002 | 1.0E-5 | -0.532 | 0.01115 | 0.269 |
| 340 | DIO3OS |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-339-5p;hsa-miR-34c-5p;hsa-miR-3607-3p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-455-5p;hsa-miR-484;hsa-miR-589-3p;hsa-miR-625-5p;hsa-miR-627-5p | 26 | TGFBR3 | Sponge network | -3.619 | 0 | -2.591 | 0 | 0.268 |
| 341 | ACTA2-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-769-5p;hsa-miR-944;hsa-miR-96-5p | 25 | TGFBR1 | Sponge network | -3.838 | 0 | -0.532 | 0.01115 | 0.267 |
| 342 | RP11-532F6.3 |
hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-576-5p;hsa-miR-590-3p | 16 | TGFBR1 | Sponge network | -1.772 | 1.0E-5 | -0.532 | 0.01115 | 0.267 |
| 343 | AF131215.9 | hsa-let-7i-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-424-5p;hsa-miR-589-3p | 13 | TGFBR3 | Sponge network | -0.75 | 0.09642 | -2.591 | 0 | 0.266 |
| 344 | LINC00654 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-429;hsa-miR-454-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-769-5p;hsa-miR-944;hsa-miR-96-5p;hsa-miR-98-5p | 27 | TGFBR1 | Sponge network | -1.448 | 0.00044 | -0.532 | 0.01115 | 0.266 |
| 345 | PCED1B-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-3200-3p;hsa-miR-429;hsa-miR-769-5p;hsa-miR-96-5p;hsa-miR-98-5p | 17 | TGFBR1 | Sponge network | -0.575 | 0.17488 | -0.532 | 0.01115 | 0.265 |
| 346 | RP11-145A3.1 |
hsa-miR-126-5p;hsa-miR-129-5p;hsa-miR-181c-5p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-944 | 10 | HGF | Sponge network | 1.041 | 0.23373 | -2.968 | 0 | 0.265 |
| 347 | RP11-25K19.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-429;hsa-miR-944 | 18 | TGFBR1 | Sponge network | -1.478 | 0.00829 | -0.532 | 0.01115 | 0.265 |
| 348 | NR2F1-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-148b-3p;hsa-miR-182-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-330-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-429;hsa-miR-532-5p | 11 | SNAI2 | Sponge network | -1.881 | 0 | -0.28 | 0.48988 | 0.265 |
| 349 | PDZRN3-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-361-5p;hsa-miR-500a-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-590-5p | 23 | TGFBR1 | Sponge network | -5.049 | 1.0E-5 | -0.532 | 0.01115 | 0.265 |
| 350 | RP11-554A11.4 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-32-3p;hsa-miR-3200-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-769-5p | 15 | TGFBR1 | Sponge network | -3.989 | 0 | -0.532 | 0.01115 | 0.264 |
| 351 | BZRAP1-AS1 |
hsa-let-7a-3p;hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181b-5p;hsa-miR-200a-3p;hsa-miR-224-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-944 | 10 | HGF | Sponge network | -2.343 | 0 | -2.968 | 0 | 0.264 |
| 352 | LINC00702 |
hsa-miR-1226-3p;hsa-miR-148b-5p;hsa-miR-182-5p;hsa-miR-186-5p;hsa-miR-192-5p;hsa-miR-29b-3p;hsa-miR-29c-3p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-3p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-374a-3p;hsa-miR-505-3p;hsa-miR-576-5p;hsa-miR-590-3p;hsa-miR-96-5p | 17 | LIMS1 | Sponge network | -2.704 | 0 | -0.388 | 0.14814 | 0.263 |
| 353 | AC079767.4 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-944 | 11 | TGFBR1 | Sponge network | 0.594 | 0.61846 | -0.532 | 0.01115 | 0.262 |
| 354 | LINC00473 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-330-3p;hsa-miR-361-5p;hsa-miR-454-3p;hsa-miR-590-3p;hsa-miR-664a-3p;hsa-miR-769-5p | 22 | TGFBR1 | Sponge network | -5.53 | 0 | -0.532 | 0.01115 | 0.261 |
| 355 | RP11-359E10.1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-32-3p;hsa-miR-361-5p;hsa-miR-576-5p | 17 | TGFBR1 | Sponge network | -1.216 | 0.0438 | -0.532 | 0.01115 | 0.261 |
| 356 | CTD-2647L4.4 | hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-590-5p | 11 | TGFBR3 | Sponge network | -0.968 | 0.00075 | -2.591 | 0 | 0.261 |
| 357 | BVES-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-320a;hsa-miR-320b;hsa-miR-331-3p;hsa-miR-361-5p;hsa-miR-429;hsa-miR-576-5p;hsa-miR-664a-3p;hsa-miR-98-5p | 21 | TGFBR1 | Sponge network | -4.161 | 1.0E-5 | -0.532 | 0.01115 | 0.26 |
| 358 | TBX5-AS1 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-3200-3p;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-374a-5p;hsa-miR-576-5p;hsa-miR-590-5p;hsa-miR-664a-3p;hsa-miR-769-5p;hsa-miR-944 | 23 | TGFBR1 | Sponge network | -2.557 | 2.0E-5 | -0.532 | 0.01115 | 0.26 |
| 359 | RP11-356J5.12 |
hsa-let-7f-1-3p;hsa-miR-126-5p;hsa-miR-141-3p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-200a-3p;hsa-miR-26b-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 13 | HGF | Sponge network | -2.015 | 0 | -2.968 | 0 | 0.259 |
| 360 | CASC2 |
hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-34c-5p;hsa-miR-409-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-590-3p;hsa-miR-625-5p | 20 | TGFBR3 | Sponge network | -0.802 | 0.00027 | -2.591 | 0 | 0.259 |
| 361 | CTC-296K1.4 | hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-29a-5p;hsa-miR-424-5p;hsa-miR-590-5p | 12 | TGFBR3 | Sponge network | -6.559 | 0 | -2.591 | 0 | 0.258 |
| 362 | RP11-359E19.2 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-223-3p;hsa-miR-27a-3p;hsa-miR-29a-5p | 11 | TGFBR3 | Sponge network | -4.608 | 0.00038 | -2.591 | 0 | 0.258 |
| 363 | AC009228.1 | hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p | 14 | TGFBR3 | Sponge network | -5.764 | 0 | -2.591 | 0 | 0.257 |
| 364 | CTD-2554C21.3 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-339-5p;hsa-miR-34c-5p;hsa-miR-361-3p;hsa-miR-455-5p;hsa-miR-484;hsa-miR-7-1-3p | 21 | TGFBR3 | Sponge network | -2.359 | 0.00283 | -2.591 | 0 | 0.257 |
| 365 | AC020571.3 |
hsa-let-7a-5p;hsa-let-7d-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-374a-5p;hsa-miR-944;hsa-miR-96-5p | 14 | TGFBR1 | Sponge network | -0.862 | 0.18422 | -0.532 | 0.01115 | 0.257 |
| 366 | RP11-809C18.3 | hsa-let-7i-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-339-5p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-627-5p | 16 | TGFBR3 | Sponge network | -4.139 | 6.0E-5 | -2.591 | 0 | 0.257 |
| 367 | HLA-F-AS1 |
hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-330-3p;hsa-miR-34a-5p;hsa-miR-532-5p;hsa-miR-664a-3p;hsa-miR-93-5p | 10 | HMGA2 | Sponge network | -0.277 | 0.54585 | 3.207 | 0.00037 | 0.256 |
| 368 | LINC00854 | hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-34a-5p;hsa-miR-664a-5p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | HMGA2 | Sponge network | -0.431 | 0.09016 | 3.207 | 0.00037 | 0.256 |
| 369 | FRMD6-AS2 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-126-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-301a-3p;hsa-miR-944;hsa-miR-96-5p | 15 | TGFBR1 | Sponge network | -6.455 | 0 | -0.532 | 0.01115 | 0.256 |
| 370 | AF131217.1 |
hsa-let-7a-3p;hsa-let-7b-3p;hsa-miR-107;hsa-miR-126-5p;hsa-miR-146b-5p;hsa-miR-186-5p;hsa-miR-193a-3p;hsa-miR-19b-1-5p;hsa-miR-338-3p;hsa-miR-590-3p;hsa-miR-628-5p | 11 | FGFR2 | Sponge network | -5.31 | 0 | -1.105 | 0.01288 | 0.255 |
| 371 | RP11-325F22.2 | hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-331-3p;hsa-miR-96-5p | 11 | TGFBR1 | Sponge network | -1.801 | 0.01297 | -0.532 | 0.01115 | 0.255 |
| 372 | AP001046.5 | hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-330-3p;hsa-miR-331-3p;hsa-miR-96-5p | 12 | TGFBR1 | Sponge network | -0.882 | 0.0083 | -0.532 | 0.01115 | 0.255 |
| 373 | BZRAP1-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-130b-3p;hsa-miR-141-3p;hsa-miR-148b-3p;hsa-miR-149-5p;hsa-miR-186-5p;hsa-miR-200a-3p;hsa-miR-301a-3p;hsa-miR-330-5p;hsa-miR-335-3p;hsa-miR-335-5p;hsa-miR-361-5p;hsa-miR-454-3p;hsa-miR-944 | 15 | TGFB2 | Sponge network | -2.343 | 0 | -1.199 | 0.00331 | 0.255 |
| 374 | CTD-3064M3.3 | hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-339-5p;hsa-miR-361-3p;hsa-miR-424-5p;hsa-miR-625-5p | 16 | TGFBR3 | Sponge network | -1.26 | 0.00058 | -2.591 | 0 | 0.254 |
| 375 | AC010226.4 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-222-5p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-424-5p;hsa-miR-590-3p | 13 | TGFBR3 | Sponge network | -1.506 | 0 | -2.591 | 0 | 0.254 |
| 376 | APCDD1L-AS1 |
hsa-miR-107;hsa-miR-126-5p;hsa-miR-148b-3p;hsa-miR-182-5p;hsa-miR-200b-3p;hsa-miR-200c-3p;hsa-miR-330-3p;hsa-miR-335-3p;hsa-miR-532-5p;hsa-miR-96-5p | 10 | SNAI2 | Sponge network | -2.022 | 0.00702 | -0.28 | 0.48988 | 0.253 |
| 377 | RP1-151F17.2 |
hsa-miR-126-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-26b-5p;hsa-miR-30b-5p;hsa-miR-30c-5p;hsa-miR-30d-5p;hsa-miR-30e-5p;hsa-miR-335-3p;hsa-miR-33a-3p;hsa-miR-421;hsa-miR-590-3p;hsa-miR-590-5p;hsa-miR-7-1-3p | 15 | HGF | Sponge network | -1.606 | 0 | -2.968 | 0 | 0.253 |
| 378 | MAGI2-AS3 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-181c-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-193a-3p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-32-3p;hsa-miR-339-5p;hsa-miR-3607-3p;hsa-miR-424-5p;hsa-miR-450b-5p;hsa-miR-455-5p;hsa-miR-589-3p;hsa-miR-590-3p;hsa-miR-625-5p;hsa-miR-7-1-3p | 28 | TGFBR3 | Sponge network | -2.414 | 0 | -2.591 | 0 | 0.252 |
| 379 | RP11-35G9.3 | hsa-let-7d-5p;hsa-let-7g-5p;hsa-miR-128-3p;hsa-miR-186-5p;hsa-miR-200b-3p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-320a;hsa-miR-320b;hsa-miR-429;hsa-miR-576-5p;hsa-miR-664a-3p;hsa-miR-769-5p | 13 | TGFBR1 | Sponge network | -0.328 | 0.16721 | -0.532 | 0.01115 | 0.252 |
| 380 | A2M-AS1 |
hsa-miR-15a-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-186-5p;hsa-miR-18a-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-23a-3p;hsa-miR-27a-3p;hsa-miR-29a-5p;hsa-miR-3607-3p;hsa-miR-484;hsa-miR-590-3p;hsa-miR-7-1-3p | 14 | TGFBR3 | Sponge network | -1.953 | 0.00771 | -2.591 | 0 | 0.252 |
| 381 | CTC-297N7.9 |
hsa-let-7b-5p;hsa-miR-15a-5p;hsa-miR-15b-5p;hsa-miR-16-2-3p;hsa-miR-16-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-185-3p;hsa-miR-18a-5p;hsa-miR-193b-3p;hsa-miR-21-5p;hsa-miR-23a-3p;hsa-miR-34c-5p;hsa-miR-590-5p | 14 | TGFBR3 | Sponge network | -2.896 | 1.0E-5 | -2.591 | 0 | 0.252 |
| 382 | CTC-296K1.3 |
hsa-let-7a-5p;hsa-let-7b-5p;hsa-let-7d-5p;hsa-let-7e-5p;hsa-let-7f-5p;hsa-let-7g-5p;hsa-miR-101-3p;hsa-miR-130b-3p;hsa-miR-16-2-3p;hsa-miR-181c-5p;hsa-miR-186-5p;hsa-miR-2110;hsa-miR-301a-3p;hsa-miR-454-3p | 14 | TGFBR1 | Sponge network | -5.677 | 0 | -0.532 | 0.01115 | 0.252 |
| 383 | RP11-1100L3.8 | hsa-let-7b-5p;hsa-let-7i-5p;hsa-miR-185-3p;hsa-miR-186-5p;hsa-miR-18a-3p;hsa-miR-18a-5p;hsa-miR-21-5p;hsa-miR-222-5p;hsa-miR-455-5p;hsa-miR-625-5p | 10 | TGFBR3 | Sponge network | -2.719 | 0.00019 | -2.591 | 0 | 0.252 |
| 384 | PCED1B-AS1 |
hsa-let-7e-5p;hsa-miR-106b-5p;hsa-miR-141-5p;hsa-miR-20b-5p;hsa-miR-23b-3p;hsa-miR-29b-2-5p;hsa-miR-30d-3p;hsa-miR-30e-3p;hsa-miR-532-5p;hsa-miR-93-3p;hsa-miR-93-5p;hsa-miR-98-5p | 12 | HMGA2 | Sponge network | -0.575 | 0.17488 | 3.207 | 0.00037 | 0.251 |