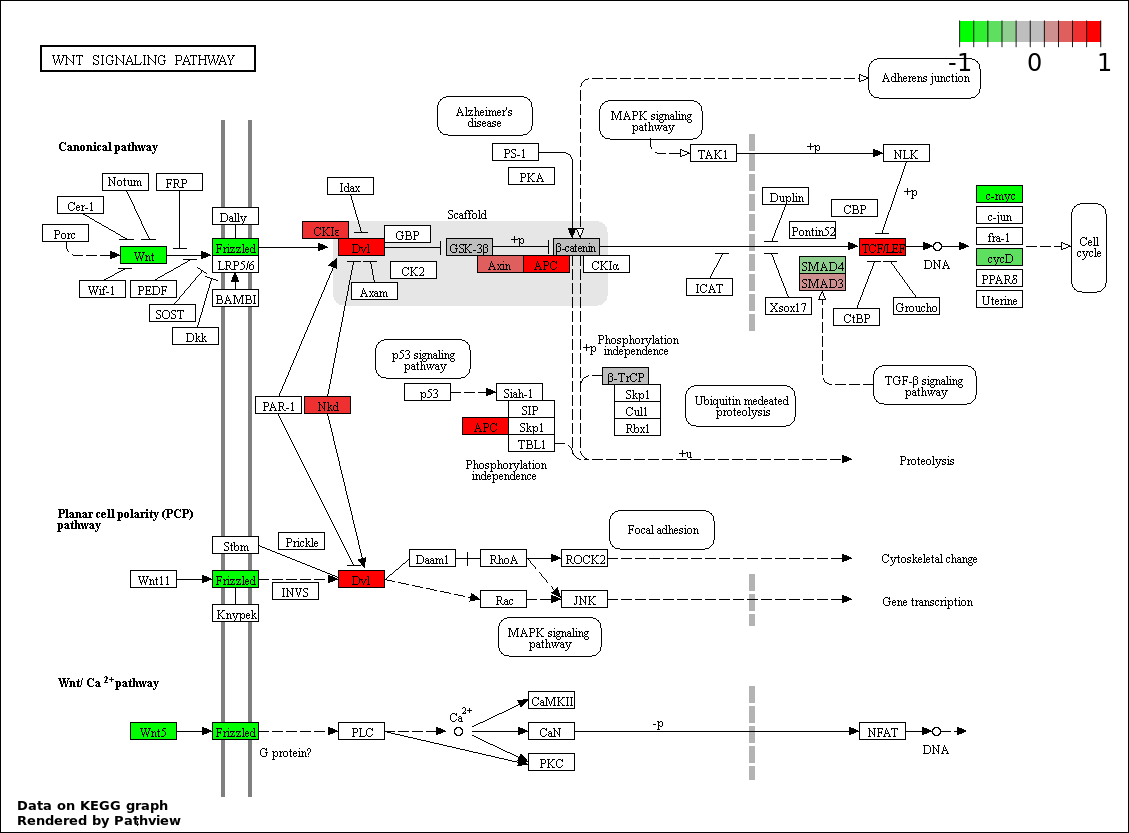
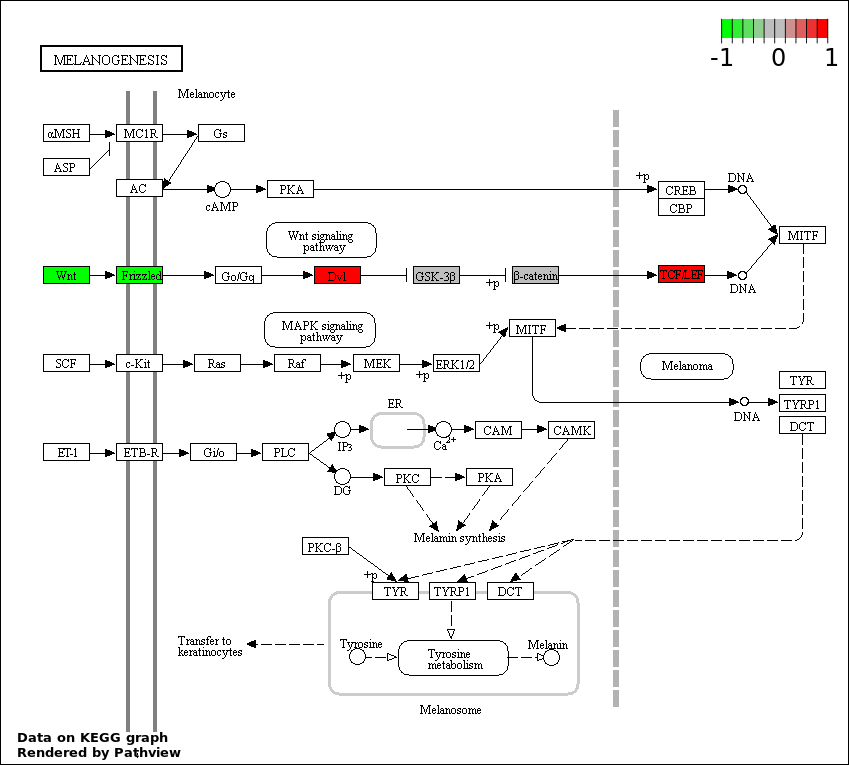
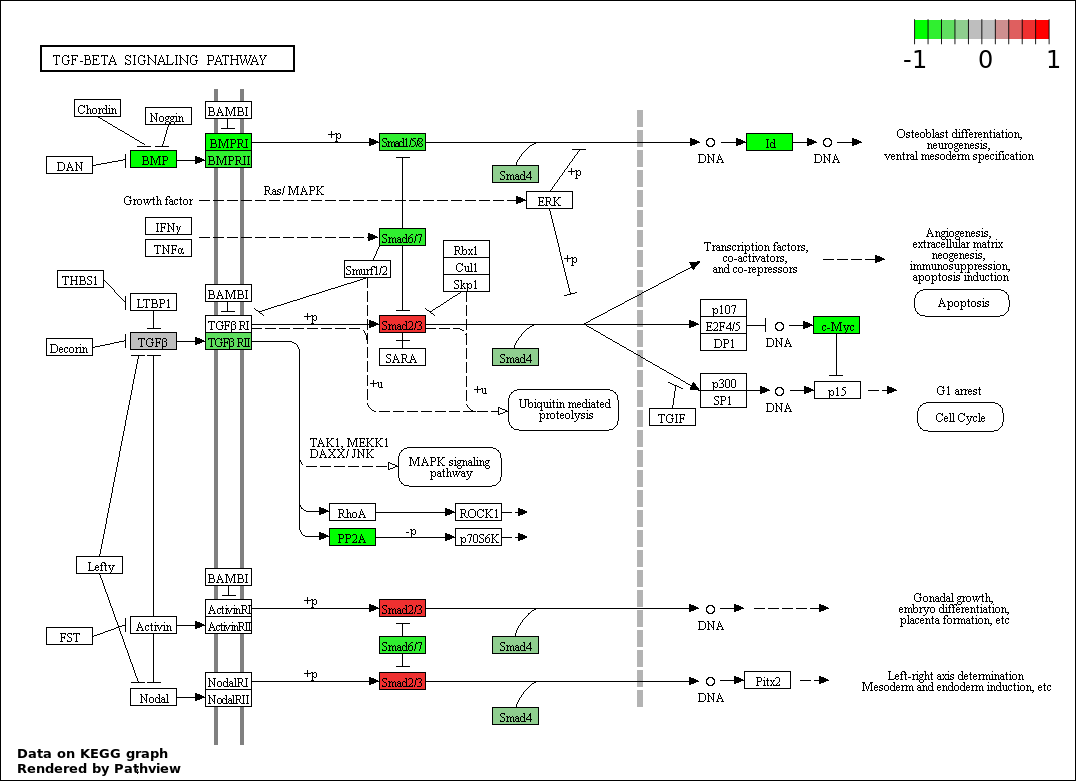
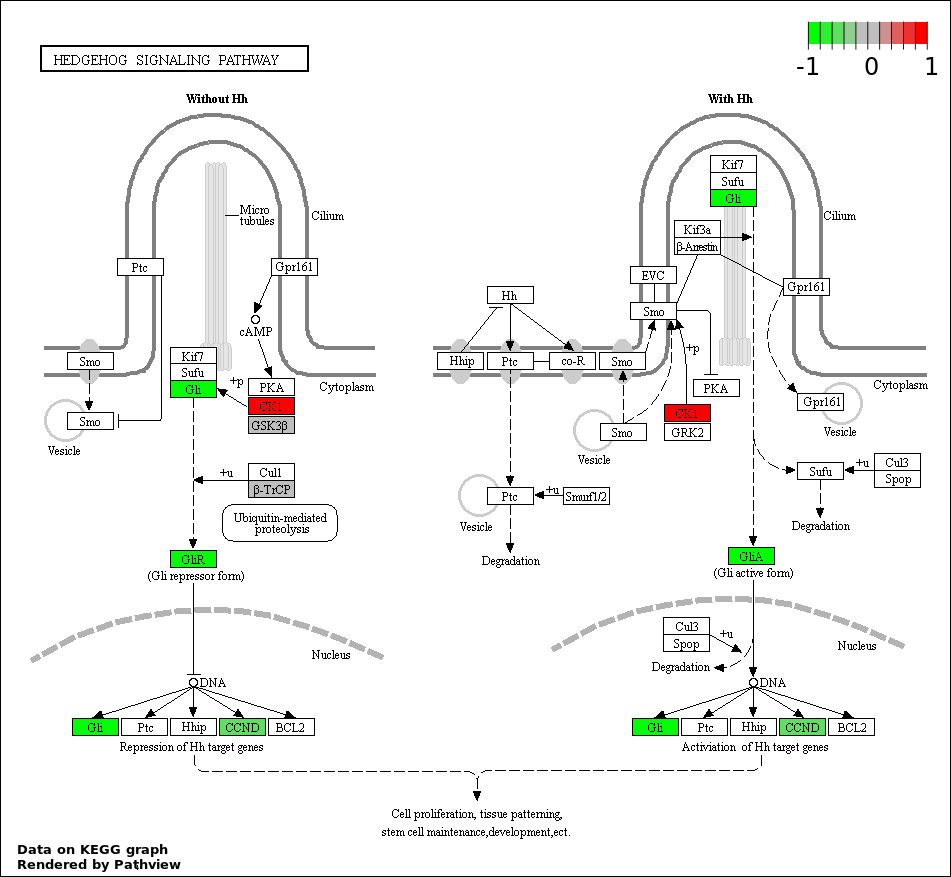
This regulatory network was inferred from the input dataset. The miRNAs and mRNAs are
presented as round and rectangle nodes respectively. The numerical value popped up upon mouse over the gene node is the log2 transformed fold-change of the gene expression between the two groups. All of the nodes are clickable, and the detailed information of the miRNAs/mRNAs and related cancer pathway will be displayed in another window. The edges between nodes are supported by both interactions (predicted or experimentally verified) and correlations learnt from cancer dataset. The numerical value popped up upon mouse over the edge is the correlation beat value (effect size) between the two nodes. The experimental evidences of the edges reported in previous cancer studies are highlighted by red/orange color. All of these information can be accessed by the "mouse-over" action. This network shows a full map of the miRNA-mRNA regulation of the input gene list(s), and the hub miRNAs (with the high network degree/betweenness centrality) would be the potential cancer drivers or tumor suppressors. The full result table can be accessed in the "Regulations" tab.
"miRNACancerMAP" is also a network visualization tool for users to draw their regulatory network by personal customization. Users can set the complexity of the network by limiting the number of nodes or edges. And the color of the nodes can be defined by different categories of the mRNAs and miRNAs, such as Gene-Ontology, pathway, and expression status. Users can also select to use network degree or network betweenness centrality to define the node size. And edges can be black or colored by the correlation. Purple edge means negative correlation (mostly found between miRNA and mRNA), and blue edge means positive correlation (found in PPI or miRNA-miRNA sponge effect). We can also add the protein-protein interactions (PPI) into the network. This result will show the cluster of genes regulated by some specific miRNAs. Additionally, miRNA-miRNA edges can be added by the "miRNA sponge" button, presenting some clusters of miRNAs that have the interactions via sponge effect.
miRNA-gene regulations
| Num | microRNA | Gene | miRNA log2FC | miRNA pvalue | Gene log2FC | Gene pvalue | Interaction | Correlation beta | Correlation P-value | PMID | Reported in cancer studies |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | hsa-miR-192-3p | ACTB | -0.64 | 0.00027 | -0.06 | 0.43313 | MirTarget | -0.11 | 0 | NA | |
| 2 | hsa-miR-92a-3p | ACTB | 0.21 | 0.13429 | -0.06 | 0.43313 | miRNAWalker2 validate | -0.1 | 6.0E-5 | NA | |
| 3 | hsa-miR-148a-5p | ACTG1 | -0.77 | 0 | -0.12 | 0.20938 | MirTarget; miRNATAP | -0.17 | 0 | NA | |
| 4 | hsa-miR-193b-3p | ACTG1 | -0.17 | 0.27202 | -0.12 | 0.20938 | miRNAWalker2 validate | -0.18 | 0 | NA | |
| 5 | hsa-miR-103a-3p | AMOT | 0.77 | 0 | -0.9 | 0.00079 | MirTarget | -0.45 | 0.00011 | NA | |
| 6 | hsa-miR-107 | AMOT | 0.24 | 0.01708 | -0.9 | 0.00079 | MirTarget; PITA; miRanda; miRNATAP | -0.41 | 0.00177 | NA | |
| 7 | hsa-miR-10b-3p | AMOT | 2.77 | 0 | -0.9 | 0.00079 | mirMAP | -0.14 | 0.00214 | NA | |
| 8 | hsa-miR-16-1-3p | AMOT | 0.39 | 0.00112 | -0.9 | 0.00079 | mirMAP | -0.31 | 0.00493 | NA | |
| 9 | hsa-miR-221-3p | AMOT | 1.12 | 0 | -0.9 | 0.00079 | miRNAWalker2 validate | -0.32 | 9.0E-5 | NA | |
| 10 | hsa-miR-30d-5p | AMOT | 0.72 | 0 | -0.9 | 0.00079 | mirMAP | -0.3 | 0.00241 | NA | |
| 11 | hsa-miR-330-5p | AMOT | 0.44 | 0.00533 | -0.9 | 0.00079 | miRanda | -0.28 | 0.00054 | NA | |
| 12 | hsa-miR-362-3p | AMOT | 0.81 | 0 | -0.9 | 0.00079 | miRanda | -0.29 | 0.00208 | NA | |
| 13 | hsa-miR-500a-5p | AMOT | 0.8 | 0 | -0.9 | 0.00079 | mirMAP | -0.29 | 0.00067 | NA | |
| 14 | hsa-miR-501-5p | AMOT | 1.15 | 0 | -0.9 | 0.00079 | miRNATAP | -0.29 | 3.0E-5 | NA | |
| 15 | hsa-miR-532-5p | AMOT | 1.03 | 0 | -0.9 | 0.00079 | MirTarget | -0.37 | 0.00017 | NA | |
| 16 | hsa-miR-618 | AMOT | 0.14 | 0.51715 | -0.9 | 0.00079 | mirMAP | -0.18 | 0.00497 | NA | |
| 17 | hsa-miR-766-3p | AMOT | 0.2 | 0.25723 | -0.9 | 0.00079 | MirTarget | -0.36 | 0 | NA | |
| 18 | hsa-miR-106b-5p | APC | 0.65 | 0 | -0.18 | 0.06792 | miRNAWalker2 validate; miRTarBase | -0.15 | 0.00024 | 23087084 | miR 106b downregulates adenomatous polyposis coli and promotes cell proliferation in human hepatocellular carcinoma; Moreover we demonstrated that miR-106b downregulates APC expression by directly targeting the 3'-untranslated region of APC messenger RNA; Taken together our results suggest that miR-106b plays an important role in promoting the proliferation of human hepatoma cells and presents a novel mechanism of micro RNA-mediated direct suppression of APC expression in cancer cells |
| 19 | hsa-miR-21-5p | APC | 1.51 | 0 | -0.18 | 0.06792 | miRNAWalker2 validate | -0.17 | 0 | 23773491; 24832083 | The prognostic significance of APC gene mutation and miR 21 expression in advanced stage colorectal cancer; The aim of this study was to analyse the association of APC gene mutation and miR-21 expression with clinical outcome in CRC patients; APC gene mutation and expression of APC and miR-21 were analysed by direct DNA sequencing and real-time reverse transcription polymerase chain reaction; APC gene expression was low in CRC and negatively correlated with miR-21 expression and gene mutation; In Taiwan downregulation of the APC gene in CRC correlated with gene mutation and miR-21 upregulation; APC mutation and miR-21 expression could be used to predict the clinical outcome of CRC especially in patients with advanced disease;MicroRNA 21 promotes tumour malignancy via increased nuclear translocation of β catenin and predicts poor outcome in APC mutated but not in APC wild type colorectal cancer; However in our preliminary data the prognostic value of miR-21 levels was observed only in adenomatous polyposis coli APC-mutated tumours not in APC-wild-type tumours; We enrolled 165 colorectal tumour to determine APC mutation miR-21 levels and nuclear β-catenin expression by direct sequencing real-time PCR and immunohistochemistry |
| 20 | hsa-let-7a-5p | APC2 | -0.33 | 0.00046 | 1.72 | 0 | TargetScan | -0.3 | 0.00538 | NA | |
| 21 | hsa-miR-125a-5p | APC2 | -0.91 | 0 | 1.72 | 0 | mirMAP | -0.2 | 0.0003 | NA | |
| 22 | hsa-miR-194-3p | APC2 | -0.77 | 3.0E-5 | 1.72 | 0 | mirMAP | -0.17 | 0.00111 | NA | |
| 23 | hsa-miR-199a-5p | APC2 | -1.99 | 0 | 1.72 | 0 | mirMAP | -0.1 | 0.00178 | NA | |
| 24 | hsa-miR-20a-3p | APC2 | -0.32 | 0.04679 | 1.72 | 0 | mirMAP | -0.25 | 7.0E-5 | NA | |
| 25 | hsa-miR-23b-5p | APC2 | -1.05 | 0 | 1.72 | 0 | mirMAP | -0.22 | 0.0013 | NA | |
| 26 | hsa-miR-28-5p | APC2 | -0.43 | 0 | 1.72 | 0 | mirMAP | -0.3 | 0.00527 | NA | |
| 27 | hsa-miR-505-5p | APC2 | -0.77 | 1.0E-5 | 1.72 | 0 | mirMAP | -0.15 | 0.00924 | NA | |
| 28 | hsa-miR-107 | AREG | 0.24 | 0.01708 | -1.67 | 0 | miRanda | -1.2 | 0 | NA | |
| 29 | hsa-miR-320a | AXIN1 | 0.33 | 0.02214 | 0.3 | 0.00968 | miRNAWalker2 validate | -0.1 | 0.00946 | NA | |
| 30 | hsa-miR-15b-5p | AXIN2 | 0.23 | 0.08248 | 0.11 | 0.75298 | miRTarBase; MirTarget; miRNATAP | -0.45 | 0.00081 | NA | |
| 31 | hsa-miR-16-2-3p | AXIN2 | -0.03 | 0.80516 | 0.11 | 0.75298 | mirMAP | -0.64 | 0 | NA | |
| 32 | hsa-miR-16-5p | AXIN2 | -0.4 | 0.0001 | 0.11 | 0.75298 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.49 | 0.00421 | NA | |
| 33 | hsa-miR-195-3p | AXIN2 | -1.09 | 0 | 0.11 | 0.75298 | mirMAP | -0.31 | 0.00083 | NA | |
| 34 | hsa-miR-195-5p | AXIN2 | -1.86 | 0 | 0.11 | 0.75298 | MirTarget; miRNATAP | -0.25 | 0.00506 | NA | |
| 35 | hsa-miR-26b-3p | AXIN2 | -1.26 | 0 | 0.11 | 0.75298 | MirTarget | -0.59 | 3.0E-5 | NA | |
| 36 | hsa-miR-326 | AXIN2 | -1.88 | 0 | 0.11 | 0.75298 | miRanda | -0.36 | 2.0E-5 | NA | |
| 37 | hsa-miR-338-5p | AXIN2 | -0.22 | 0.25239 | 0.11 | 0.75298 | miRNATAP | -0.34 | 0.00017 | NA | |
| 38 | hsa-miR-497-5p | AXIN2 | -1.41 | 0 | 0.11 | 0.75298 | MirTarget; miRNATAP | -0.29 | 0.00329 | NA | |
| 39 | hsa-miR-616-3p | AXIN2 | -0.7 | 2.0E-5 | 0.11 | 0.75298 | MirTarget | -0.42 | 8.0E-5 | NA | |
| 40 | hsa-let-7a-2-3p | BBC3 | -1.19 | 0 | 0.8 | 0 | MirTarget | -0.19 | 0 | NA | |
| 41 | hsa-let-7g-3p | BBC3 | -1.14 | 0 | 0.8 | 0 | MirTarget; miRNATAP | -0.18 | 0.0001 | NA | |
| 42 | hsa-miR-101-3p | BBC3 | -1.48 | 0 | 0.8 | 0 | miRNATAP | -0.32 | 0 | NA | |
| 43 | hsa-miR-125b-5p | BBC3 | -1.36 | 0 | 0.8 | 0 | miRNAWalker2 validate; miRTarBase | -0.25 | 0 | 25184537 | Thus far two of these target genes BBC3 and NEU1 that are tumor suppressor genes but not yet studied in PDAC appear to be functional targets of miR-125b since knockdown of miR125b caused their up regulation |
| 44 | hsa-miR-139-5p | BBC3 | -2.11 | 0 | 0.8 | 0 | miRNATAP | -0.23 | 0 | NA | |
| 45 | hsa-miR-140-5p | BBC3 | -0.22 | 0.01407 | 0.8 | 0 | miRNATAP | -0.25 | 0.0003 | NA | |
| 46 | hsa-miR-144-3p | BBC3 | -2.98 | 0 | 0.8 | 0 | miRNATAP | -0.11 | 0 | NA | |
| 47 | hsa-miR-27b-3p | BBC3 | -0.82 | 0 | 0.8 | 0 | miRNATAP | -0.2 | 0.00022 | NA | |
| 48 | hsa-miR-345-5p | BBC3 | -0.71 | 0 | 0.8 | 0 | miRNATAP | -0.17 | 4.0E-5 | NA | |
| 49 | hsa-miR-590-3p | BBC3 | -0.47 | 2.0E-5 | 0.8 | 0 | miRanda | -0.25 | 1.0E-5 | NA | |
| 50 | hsa-let-7b-5p | BIRC5 | -0.96 | 0 | 4.5 | 0 | miRNAWalker2 validate | -0.71 | 0 | NA | |
| 51 | hsa-miR-101-3p | BIRC5 | -1.48 | 0 | 4.5 | 0 | miRNAWalker2 validate | -1.33 | 0 | NA | |
| 52 | hsa-miR-10a-5p | BIRC5 | -1.48 | 0 | 4.5 | 0 | miRNAWalker2 validate | -0.71 | 0 | NA | |
| 53 | hsa-miR-203a-3p | BIRC5 | -1.34 | 9.0E-5 | 4.5 | 0 | miRTarBase | -0.13 | 0.00935 | 22713668; 27714672 | Luciferase assays were also performed to validate BIRC5 and LASP1 as miR-203 targets; Both miR-203 and BIRC5 siRNA signicantly inhibited cell proliferation in TNBC cells; Moreover up-regulated of BIRC5 and LASP1 was able to abrogate the effects induced by transfection with the miR-203 precursor;miR 203 is a predictive biomarker for colorectal cancer and its expression is associated with BIRC5; The purpose of this study was to explore the role of miR-203 in colorectal cancer CRC and evaluate the correlation between miR-203 and BIRC5; Finally miR-203 expression was negatively associated with that of BIRC5 r = -0.8150 P < 0.05 |
| 54 | hsa-miR-30c-5p | BIRC5 | -0.43 | 0.00016 | 4.5 | 0 | miRNAWalker2 validate | -0.45 | 0.00191 | NA | |
| 55 | hsa-miR-335-5p | BIRC5 | -1.61 | 0 | 4.5 | 0 | miRNAWalker2 validate; MirTarget | -0.5 | 0 | 23232114 | Genetic variation in a miR 335 binding site in BIRC5 alters susceptibility to lung cancer in Chinese Han populations; In support of the postulation that the 3' UTR SNP may directly affect miRNA-binding site reporter gene assays indicated BIRC5 was a direct target of miR-335 and the rs2239680 T>C change resulted in altered regulation of BIRC5 expression; Our findings defined a 3' UTR SNP in human BIRC5 oncogene that may increase individual susceptibility to lung cancer probably by attenuating the interaction between miR-335 and BIRC5 |
| 56 | hsa-miR-542-3p | BIRC5 | -1.31 | 0 | 4.5 | 0 | miRNAWalker2 validate; MirTarget; miRanda | -0.82 | 0 | NA | |
| 57 | hsa-miR-142-3p | BMP4 | -1.42 | 0 | 0.76 | 0.00257 | PITA; miRanda | -0.21 | 0.00115 | NA | |
| 58 | hsa-miR-130b-3p | BMP6 | 0.69 | 0.00011 | -1.04 | 0 | MirTarget | -0.26 | 0 | NA | |
| 59 | hsa-miR-362-3p | BMP6 | 0.81 | 0 | -1.04 | 0 | miRanda | -0.3 | 4.0E-5 | NA | |
| 60 | hsa-miR-454-3p | BMP6 | 0.67 | 0 | -1.04 | 0 | MirTarget | -0.22 | 0.00157 | NA | |
| 61 | hsa-miR-130b-5p | BMP7 | 0.17 | 0.33761 | 0.83 | 0.04887 | mirMAP | -0.32 | 0.00583 | NA | |
| 62 | hsa-miR-1976 | BMP7 | -0.43 | 0.00325 | 0.83 | 0.04887 | MirTarget | -0.45 | 0.00115 | NA | |
| 63 | hsa-miR-22-3p | BMP7 | -0.63 | 0 | 0.83 | 0.04887 | miRNAWalker2 validate; miRTarBase | -0.87 | 1.0E-5 | NA | |
| 64 | hsa-miR-30a-5p | BMP7 | -0.63 | 0.00011 | 0.83 | 0.04887 | mirMAP; miRNATAP | -0.35 | 0.00579 | NA | |
| 65 | hsa-miR-30e-5p | BMP7 | -0.63 | 0 | 0.83 | 0.04887 | mirMAP | -0.58 | 0.00444 | NA | |
| 66 | hsa-miR-34a-5p | BMP7 | 1.04 | 0 | 0.83 | 0.04887 | miRNAWalker2 validate | -0.45 | 0.00162 | NA | |
| 67 | hsa-miR-616-5p | BMP7 | 0.15 | 0.40284 | 0.83 | 0.04887 | mirMAP | -0.33 | 0.0036 | NA | |
| 68 | hsa-miR-194-3p | BMP8A | -0.77 | 3.0E-5 | 1.39 | 0 | MirTarget | -0.35 | 0 | NA | |
| 69 | hsa-miR-30b-3p | BMP8A | -0.44 | 0.00095 | 1.39 | 0 | MirTarget | -0.25 | 0.00222 | NA | |
| 70 | hsa-miR-30e-3p | BMP8A | -1.21 | 0 | 1.39 | 0 | MirTarget | -0.32 | 0.00032 | NA | |
| 71 | hsa-miR-455-5p | BMP8A | -0.27 | 0.05813 | 1.39 | 0 | MirTarget; miRanda | -0.22 | 0.00398 | NA | |
| 72 | hsa-miR-618 | BMP8B | 0.14 | 0.51715 | 0.28 | 0.41783 | mirMAP | -0.24 | 0.00328 | NA | |
| 73 | hsa-miR-140-5p | BMPR1A | -0.22 | 0.01407 | 0.07 | 0.48504 | miRanda | -0.15 | 0.00409 | NA | |
| 74 | hsa-miR-130b-3p | BMPR1B | 0.69 | 0.00011 | -3.31 | 0 | miRNATAP | -0.34 | 0.00049 | NA | |
| 75 | hsa-miR-140-3p | BMPR1B | 0.55 | 0 | -3.31 | 0 | MirTarget | -1.09 | 0 | NA | |
| 76 | hsa-miR-362-3p | BMPR1B | 0.81 | 0 | -3.31 | 0 | MirTarget; miRanda | -0.65 | 0 | NA | |
| 77 | hsa-miR-374a-5p | BMPR1B | 0.02 | 0.86978 | -3.31 | 0 | MirTarget | -0.96 | 0 | NA | |
| 78 | hsa-miR-374b-5p | BMPR1B | -0.31 | 0.00301 | -3.31 | 0 | MirTarget | -0.45 | 0.00882 | NA | |
| 79 | hsa-miR-106b-5p | BMPR2 | 0.65 | 0 | -0.74 | 0 | MirTarget; miRNATAP | -0.22 | 0 | NA | |
| 80 | hsa-miR-130b-3p | BMPR2 | 0.69 | 0.00011 | -0.74 | 0 | MirTarget; miRNATAP | -0.12 | 2.0E-5 | NA | |
| 81 | hsa-miR-148b-3p | BMPR2 | 0.27 | 0.00185 | -0.74 | 0 | mirMAP | -0.27 | 1.0E-5 | NA | |
| 82 | hsa-miR-17-5p | BMPR2 | 0.7 | 2.0E-5 | -0.74 | 0 | miRNAWalker2 validate; MirTarget; TargetScan; miRNATAP | -0.26 | 0 | NA | |
| 83 | hsa-miR-185-5p | BMPR2 | 0.48 | 0 | -0.74 | 0 | MirTarget | -0.25 | 0 | NA | |
| 84 | hsa-miR-19a-3p | BMPR2 | 1.02 | 0 | -0.74 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.17 | 0 | NA | |
| 85 | hsa-miR-19b-3p | BMPR2 | 0.6 | 0.00017 | -0.74 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.21 | 0 | NA | |
| 86 | hsa-miR-20a-5p | BMPR2 | 0.85 | 0 | -0.74 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.24 | 0 | NA | |
| 87 | hsa-miR-21-5p | BMPR2 | 1.51 | 0 | -0.74 | 0 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.28 | 0 | NA | |
| 88 | hsa-miR-25-3p | BMPR2 | 0.63 | 0 | -0.74 | 0 | miRNATAP | -0.17 | 0.00117 | NA | |
| 89 | hsa-miR-29a-5p | BMPR2 | -0.11 | 0.34962 | -0.74 | 0 | mirMAP | -0.15 | 0.00063 | NA | |
| 90 | hsa-miR-301a-3p | BMPR2 | 0.84 | 0 | -0.74 | 0 | MirTarget; miRNATAP | -0.11 | 0.00019 | NA | |
| 91 | hsa-miR-30d-5p | BMPR2 | 0.72 | 0 | -0.74 | 0 | mirMAP | -0.19 | 0 | NA | |
| 92 | hsa-miR-362-3p | BMPR2 | 0.81 | 0 | -0.74 | 0 | MirTarget; PITA; miRanda; miRNATAP | -0.18 | 0 | NA | |
| 93 | hsa-miR-455-5p | BMPR2 | -0.27 | 0.05813 | -0.74 | 0 | PITA | -0.13 | 0.00044 | NA | |
| 94 | hsa-miR-532-5p | BMPR2 | 1.03 | 0 | -0.74 | 0 | PITA | -0.29 | 0 | NA | |
| 95 | hsa-miR-590-5p | BMPR2 | -0.1 | 0.31003 | -0.74 | 0 | MirTarget; PITA; miRNATAP | -0.19 | 0.0002 | NA | |
| 96 | hsa-miR-618 | BMPR2 | 0.14 | 0.51715 | -0.74 | 0 | PITA; mirMAP | -0.1 | 5.0E-5 | NA | |
| 97 | hsa-miR-671-5p | BMPR2 | 0.84 | 0 | -0.74 | 0 | miRNATAP | -0.11 | 0.00171 | NA | |
| 98 | hsa-miR-769-5p | BMPR2 | 0.22 | 0.03334 | -0.74 | 0 | miRNATAP | -0.15 | 0.003 | NA | |
| 99 | hsa-miR-877-5p | BMPR2 | 1.36 | 0 | -0.74 | 0 | MirTarget | -0.15 | 0 | NA | |
| 100 | hsa-miR-92a-3p | BMPR2 | 0.21 | 0.13429 | -0.74 | 0 | miRNAWalker2 validate; miRNATAP | -0.24 | 0 | NA | |
| 101 | hsa-miR-93-3p | BMPR2 | 0.4 | 0.00131 | -0.74 | 0 | miRNAWalker2 validate | -0.19 | 0 | NA | |
| 102 | hsa-miR-93-5p | BMPR2 | 1.4 | 0 | -0.74 | 0 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.27 | 0 | NA | |
| 103 | hsa-miR-15b-5p | BTRC | 0.23 | 0.08248 | -0 | 0.97671 | MirTarget; miRNATAP | -0.12 | 0 | NA | |
| 104 | hsa-miR-16-5p | BTRC | -0.4 | 0.0001 | -0 | 0.97671 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.17 | 0 | NA | |
| 105 | hsa-miR-193a-3p | BTRC | -0.12 | 0.30939 | -0 | 0.97671 | MirTarget; miRanda | -0.15 | 0 | NA | |
| 106 | hsa-miR-193b-3p | BTRC | -0.17 | 0.27202 | -0 | 0.97671 | miRNAWalker2 validate | -0.16 | 0 | NA | |
| 107 | hsa-miR-361-3p | BTRC | 0.28 | 0.00549 | -0 | 0.97671 | MirTarget; PITA; miRNATAP | -0.1 | 0.00495 | NA | |
| 108 | hsa-miR-497-5p | BTRC | -1.41 | 0 | -0 | 0.97671 | MirTarget; miRNATAP | -0.11 | 0 | NA | |
| 109 | hsa-miR-505-3p | BTRC | -1.2 | 0 | -0 | 0.97671 | MirTarget | -0.13 | 0 | NA | |
| 110 | hsa-miR-106a-5p | CCND1 | -0.46 | 0.00972 | -0.9 | 1.0E-5 | MirTarget; miRNATAP | -0.26 | 0 | NA | |
| 111 | hsa-miR-106b-5p | CCND1 | 0.65 | 0 | -0.9 | 1.0E-5 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.43 | 0 | NA | |
| 112 | hsa-miR-1266-5p | CCND1 | 1.63 | 0 | -0.9 | 1.0E-5 | MirTarget | -0.23 | 0 | NA | |
| 113 | hsa-miR-15b-5p | CCND1 | 0.23 | 0.08248 | -0.9 | 1.0E-5 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.54 | 0 | NA | |
| 114 | hsa-miR-16-5p | CCND1 | -0.4 | 0.0001 | -0.9 | 1.0E-5 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.31 | 0.00178 | 23991964; 22922827; 18483394 | At the molecular level our results further revealed that cyclin D1 expression was negatively regulated by miR-16;CCND1 has been found to be a target of miR-15a and miR-16-1 through analysis of complementary sequences between microRNAs and CCND1 mRNA; Moreover the transcription of CCND1 is suppressed by miR-15a and miR-16-1 via direct binding to the CCND1 3'-untranslated region 3'-UTR;Truncation in CCND1 mRNA alters miR 16 1 regulation in mantle cell lymphoma; Furthermore we demonstrated that this truncation alters miR-16-1 binding sites and through the use of reporter constructs we were able to show that miR-16-1 regulates CCND1 mRNA expression; This study introduces the role of miR-16-1 in the regulation of CCND1 in MCL |
| 115 | hsa-miR-17-5p | CCND1 | 0.7 | 2.0E-5 | -0.9 | 1.0E-5 | miRNAWalker2 validate; MirTarget; TargetScan; miRNATAP | -0.34 | 0 | 26431674 | Bioinformatics Prediction and In Vitro Analysis Revealed That miR 17 Targets Cyclin D1 mRNA in Triple Negative Breast Cancer Cells; In this study using bioinformatic analyses miR-17 was selected as it targets the 3'UTR of CCND1 gene with the highest score; After lentiviral transduction of miR-17 to the target cells gene expression analysis showed decreased expression of CCND1 gene |
| 116 | hsa-miR-186-5p | CCND1 | -0.06 | 0.53529 | -0.9 | 1.0E-5 | mirMAP | -0.32 | 0.00286 | NA | |
| 117 | hsa-miR-19a-3p | CCND1 | 1.02 | 0 | -0.9 | 1.0E-5 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.28 | 0 | 25985117 | Moreover miR-19a might play inhibitory roles in HCC malignancy via regulating Cyclin D1 expression |
| 118 | hsa-miR-19b-1-5p | CCND1 | -0.28 | 0.07831 | -0.9 | 1.0E-5 | miRNAWalker2 validate; miRTarBase | -0.31 | 0 | NA | |
| 119 | hsa-miR-19b-3p | CCND1 | 0.6 | 0.00017 | -0.9 | 1.0E-5 | miRNATAP | -0.34 | 0 | NA | |
| 120 | hsa-miR-20a-5p | CCND1 | 0.85 | 0 | -0.9 | 1.0E-5 | miRNAWalker2 validate; miRTarBase; MirTarget; miRNATAP | -0.33 | 0 | NA | |
| 121 | hsa-miR-20b-5p | CCND1 | 0.46 | 0.02859 | -0.9 | 1.0E-5 | MirTarget; miRNATAP | -0.23 | 0 | NA | |
| 122 | hsa-miR-340-5p | CCND1 | -0 | 0.9685 | -0.9 | 1.0E-5 | mirMAP | -0.32 | 0.00013 | NA | |
| 123 | hsa-miR-425-5p | CCND1 | 0.59 | 2.0E-5 | -0.9 | 1.0E-5 | miRNAWalker2 validate | -0.39 | 0 | NA | |
| 124 | hsa-miR-503-5p | CCND1 | 0.19 | 0.26842 | -0.9 | 1.0E-5 | miRNAWalker2 validate; miRTarBase; MirTarget | -0.16 | 0.00815 | 26047605; 23731275 | MiR 503 inhibited cell proliferation of human breast cancer cells by suppressing CCND1 expression; Overexpression of miR-503 in breast cancer cell lines reduced cell proliferation through inducing G0/G1 cell cycle arrest by targeting CCND1;MicroRNA 503 suppresses proliferation and cell cycle progression of endometrioid endometrial cancer by negatively regulating cyclin D1; CCND1 has a binding sequence of miR-503 within its 3' untranslated region and was confirmed to be a direct target of miR-503 by the fluorescent reporter assays; Increasing the miR-503 level in EEC cells suppressed cell viability colon formation activity and cell-cycle progression and the inhibited oncogenic phenotypes induced by miR-503 were alleviated by ectopic expression of CCND1 without the untranslated region sequence; Collectively this study suggested that miR-503 plays a tumor-suppressor role by targeting CCND1; Abnormal suppression of miR-503 leads to an increase in the CCND1 level which may promote carcinogenesis and progression of EEC |
| 125 | hsa-miR-589-3p | CCND1 | 1.17 | 0 | -0.9 | 1.0E-5 | MirTarget | -0.18 | 0.00124 | NA | |
| 126 | hsa-miR-616-5p | CCND1 | 0.15 | 0.40284 | -0.9 | 1.0E-5 | mirMAP | -0.26 | 1.0E-5 | NA | |
| 127 | hsa-miR-7-1-3p | CCND1 | -0.57 | 2.0E-5 | -0.9 | 1.0E-5 | mirMAP | -0.26 | 0.00057 | NA | |
| 128 | hsa-miR-9-5p | CCND1 | 1.26 | 9.0E-5 | -0.9 | 1.0E-5 | miRNAWalker2 validate | -0.14 | 1.0E-5 | NA | |
| 129 | hsa-miR-92a-3p | CCND1 | 0.21 | 0.13429 | -0.9 | 1.0E-5 | miRNAWalker2 validate | -0.41 | 0 | NA | |
| 130 | hsa-miR-93-5p | CCND1 | 1.4 | 0 | -0.9 | 1.0E-5 | miRNAWalker2 validate; MirTarget; miRNATAP | -0.34 | 0 | NA | |
| 131 | hsa-miR-942-5p | CCND1 | 0.35 | 0.02833 | -0.9 | 1.0E-5 | MirTarget | -0.25 | 0.00012 | NA | |
| 132 | hsa-miR-130b-5p | CCND2 | 0.17 | 0.33761 | 0.36 | 0.03656 | mirMAP | -0.17 | 0.00018 | NA | |
| 133 | hsa-miR-20a-5p | CCND2 | 0.85 | 0 | 0.36 | 0.03656 | miRNAWalker2 validate; miRTarBase; miRNATAP | -0.16 | 0.00121 | NA | |
| 134 | hsa-miR-28-5p | CCND2 | -0.43 | 0 | 0.36 | 0.03656 | miRanda | -0.48 | 0 | NA | |
| 135 | hsa-miR-33a-3p | CCND2 | -0.68 | 1.0E-5 | 0.36 | 0.03656 | MirTarget | -0.26 | 0 | NA | |
| 136 | hsa-miR-3607-3p | CCND2 | -2.16 | 0 | 0.36 | 0.03656 | mirMAP | -0.12 | 0.0007 | NA | |
| 137 | hsa-miR-378a-3p | CCND2 | -1.19 | 0 | 0.36 | 0.03656 | miRNAWalker2 validate | -0.18 | 2.0E-5 | NA | |
| 138 | hsa-miR-548v | CCND2 | -0.27 | 0.17626 | 0.36 | 0.03656 | MirTarget | -0.15 | 0.00034 | NA | |
| 139 | hsa-miR-616-5p | CCND2 | 0.15 | 0.40284 | 0.36 | 0.03656 | mirMAP | -0.32 | 0 | NA | |
| 140 | hsa-miR-618 | CCND2 | 0.14 | 0.51715 | 0.36 | 0.03656 | mirMAP | -0.23 | 0 | NA | |
| 141 | hsa-miR-27b-3p | CCND3 | -0.82 | 0 | 0.08 | 0.47843 | miRNAWalker2 validate | -0.24 | 0 | NA | |
| 142 | hsa-miR-320a | CCND3 | 0.33 | 0.02214 | 0.08 | 0.47843 | miRanda | -0.12 | 0.00135 | NA | |
| 143 | hsa-miR-146b-5p | CDH1 | 0.42 | 0.04574 | -0.93 | 4.0E-5 | miRanda | -0.21 | 6.0E-5 | NA | |
| 144 | hsa-miR-185-5p | CDH1 | 0.48 | 0 | -0.93 | 4.0E-5 | MirTarget | -0.45 | 8.0E-5 | 27212161; 24025511 | We also found that ectopic expression of miR-185 reversed EMT via the up-regulation of E-cadherin and down-regulation of vimentin in epithelial and mesenchymal HCC cells;After transfection of miR-185 inhibitor into KYSE30 the expression of E-cadherin a downstream protein of Six1 was observed under confocal microscope; After transfection of miR-185 inhibitorthe expression level of E-cadherin decreased |
| 145 | hsa-let-7b-5p | CSNK1D | -0.96 | 0 | 0.47 | 0 | miRNAWalker2 validate | -0.19 | 0 | NA | |
| 146 | hsa-miR-29a-3p | CSNK1D | -0.86 | 0 | 0.47 | 0 | mirMAP | -0.14 | 0 | NA | |
| 147 | hsa-miR-29c-3p | CSNK1D | -1.44 | 0 | 0.47 | 0 | mirMAP | -0.15 | 0 | NA | |
| 148 | hsa-miR-148a-5p | CSNK1E | -0.77 | 0 | 0.61 | 0 | miRNATAP | -0.25 | 0 | NA | |
| 149 | hsa-miR-195-3p | CSNK1E | -1.09 | 0 | 0.61 | 0 | miRNATAP | -0.1 | 1.0E-5 | NA | |
| 150 | hsa-miR-140-5p | CTNNA1 | -0.22 | 0.01407 | 0.48 | 0 | miRanda | -0.1 | 0.00712 | NA |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CANONICAL WNT SIGNALING PATHWAY | 31 | 95 | 2.596e-46 | 1.208e-42 |
| 2 | WNT SIGNALING PATHWAY | 42 | 351 | 4.268e-43 | 9.93e-40 |
| 3 | EPITHELIUM DEVELOPMENT | 56 | 945 | 1.584e-41 | 2.457e-38 |
| 4 | ORGAN MORPHOGENESIS | 53 | 841 | 2.025e-40 | 2.356e-37 |
| 5 | TISSUE MORPHOGENESIS | 45 | 533 | 1.958e-39 | 1.822e-36 |
| 6 | TISSUE DEVELOPMENT | 63 | 1518 | 3.754e-38 | 2.911e-35 |
| 7 | REGULATION OF WNT SIGNALING PATHWAY | 37 | 310 | 9.888e-38 | 6.573e-35 |
| 8 | REGULATION OF CANONICAL WNT SIGNALING PATHWAY | 33 | 236 | 6.579e-36 | 3.827e-33 |
| 9 | NEUROGENESIS | 59 | 1402 | 1.191e-35 | 6.156e-33 |
| 10 | REGULATION OF PHOSPHORUS METABOLIC PROCESS | 61 | 1618 | 2.462e-34 | 1.145e-31 |
| 11 | REGULATION OF PROTEIN MODIFICATION PROCESS | 62 | 1710 | 5.021e-34 | 2.124e-31 |
| 12 | REGULATION OF MULTICELLULAR ORGANISMAL DEVELOPMENT | 61 | 1672 | 1.587e-33 | 6.154e-31 |
| 13 | TUBE DEVELOPMENT | 41 | 552 | 1.774e-33 | 6.348e-31 |
| 14 | POSITIVE REGULATION OF RESPONSE TO STIMULUS | 64 | 1929 | 4.617e-33 | 1.534e-30 |
| 15 | MORPHOGENESIS OF AN EPITHELIUM | 36 | 400 | 3.279e-32 | 1.017e-29 |
| 16 | REGULATION OF ANATOMICAL STRUCTURE MORPHOGENESIS | 49 | 1021 | 1.646e-31 | 4.788e-29 |
| 17 | POSITIVE REGULATION OF CELL COMMUNICATION | 57 | 1532 | 2.071e-31 | 5.668e-29 |
| 18 | EMBRYONIC MORPHOGENESIS | 39 | 539 | 2.469e-31 | 6.381e-29 |
| 19 | REGULATION OF CELL DIFFERENTIATION | 56 | 1492 | 5.69e-31 | 1.393e-28 |
| 20 | REGULATION OF CELLULAR PROTEIN LOCALIZATION | 39 | 552 | 6.129e-31 | 1.426e-28 |
| 21 | NEGATIVE REGULATION OF CELL COMMUNICATION | 51 | 1192 | 1.536e-30 | 3.404e-28 |
| 22 | POSITIVE REGULATION OF DEVELOPMENTAL PROCESS | 50 | 1142 | 2.42e-30 | 5.118e-28 |
| 23 | POSITIVE REGULATION OF CELL DIFFERENTIATION | 44 | 823 | 5.505e-30 | 1.114e-27 |
| 24 | TUBE MORPHOGENESIS | 32 | 323 | 6.727e-30 | 1.304e-27 |
| 25 | REGULATION OF ORGAN MORPHOGENESIS | 29 | 242 | 1.645e-29 | 3.061e-27 |
| 26 | POSITIVE REGULATION OF PROTEIN METABOLIC PROCESS | 54 | 1492 | 6.374e-29 | 1.141e-26 |
| 27 | NEURON DIFFERENTIATION | 44 | 874 | 6.787e-29 | 1.17e-26 |
| 28 | CELL DEVELOPMENT | 53 | 1426 | 7.101e-29 | 1.18e-26 |
| 29 | NEGATIVE REGULATION OF RESPONSE TO STIMULUS | 52 | 1360 | 7.528e-29 | 1.208e-26 |
| 30 | RESPONSE TO GROWTH FACTOR | 35 | 475 | 2.757e-28 | 4.277e-26 |
| 31 | SENSORY ORGAN DEVELOPMENT | 35 | 493 | 9.836e-28 | 1.476e-25 |
| 32 | POSITIVE REGULATION OF GENE EXPRESSION | 56 | 1733 | 1.236e-27 | 1.798e-25 |
| 33 | HIPPO SIGNALING | 15 | 27 | 4.042e-27 | 5.699e-25 |
| 34 | REGULATION OF PROTEIN LOCALIZATION | 43 | 950 | 2.511e-26 | 3.437e-24 |
| 35 | EMBRYO DEVELOPMENT | 42 | 894 | 2.616e-26 | 3.478e-24 |
| 36 | POSITIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 46 | 1135 | 2.942e-26 | 3.803e-24 |
| 37 | NEGATIVE REGULATION OF WNT SIGNALING PATHWAY | 25 | 197 | 4.185e-26 | 5.263e-24 |
| 38 | CELL FATE COMMITMENT | 26 | 227 | 5.976e-26 | 7.317e-24 |
| 39 | POSITIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 44 | 1036 | 7.507e-26 | 8.732e-24 |
| 40 | POSITIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 44 | 1036 | 7.507e-26 | 8.732e-24 |
| 41 | REGULATION OF CELL DEATH | 50 | 1472 | 2.902e-25 | 3.294e-23 |
| 42 | NEGATIVE REGULATION OF CANONICAL WNT SIGNALING PATHWAY | 23 | 162 | 3.255e-25 | 3.606e-23 |
| 43 | ANATOMICAL STRUCTURE FORMATION INVOLVED IN MORPHOGENESIS | 42 | 957 | 3.769e-25 | 4.078e-23 |
| 44 | POSITIVE REGULATION OF BIOSYNTHETIC PROCESS | 54 | 1805 | 7.096e-25 | 7.504e-23 |
| 45 | CELLULAR RESPONSE TO ORGANIC SUBSTANCE | 54 | 1848 | 2.202e-24 | 2.277e-22 |
| 46 | REGULATION OF CELLULAR LOCALIZATION | 46 | 1277 | 4.219e-24 | 4.267e-22 |
| 47 | NEGATIVE REGULATION OF DEVELOPMENTAL PROCESS | 38 | 801 | 7.883e-24 | 7.804e-22 |
| 48 | EMBRYO DEVELOPMENT ENDING IN BIRTH OR EGG HATCHING | 33 | 554 | 1.088e-23 | 1.055e-21 |
| 49 | REGULATION OF CELL DEVELOPMENT | 38 | 836 | 3.583e-23 | 3.402e-21 |
| 50 | REGULATION OF CELL PROLIFERATION | 48 | 1496 | 4.472e-23 | 4.162e-21 |
| 51 | EMBRYONIC ORGAN DEVELOPMENT | 29 | 406 | 5.265e-23 | 4.803e-21 |
| 52 | GROWTH | 29 | 410 | 6.939e-23 | 6.209e-21 |
| 53 | HEAD DEVELOPMENT | 35 | 709 | 1.877e-22 | 1.648e-20 |
| 54 | REGULATION OF EMBRYONIC DEVELOPMENT | 19 | 114 | 2.434e-22 | 2.097e-20 |
| 55 | POSITIVE REGULATION OF CELL DEVELOPMENT | 30 | 472 | 2.5e-22 | 2.115e-20 |
| 56 | CELL PROLIFERATION | 34 | 672 | 3.88e-22 | 3.167e-20 |
| 57 | TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 22 | 190 | 3.875e-22 | 3.167e-20 |
| 58 | BETA CATENIN DESTRUCTION COMPLEX DISASSEMBLY | 12 | 22 | 9.359e-22 | 7.508e-20 |
| 59 | POSITIVE REGULATION OF CELL DEATH | 32 | 605 | 2.138e-21 | 1.686e-19 |
| 60 | REGULATION OF OSSIFICATION | 21 | 178 | 2.452e-21 | 1.902e-19 |
| 61 | CELLULAR RESPONSE TO ENDOGENOUS STIMULUS | 39 | 1008 | 2.675e-21 | 2.041e-19 |
| 62 | CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 30 | 513 | 2.725e-21 | 2.045e-19 |
| 63 | GASTRULATION | 20 | 155 | 3.741e-21 | 2.763e-19 |
| 64 | CARDIOVASCULAR SYSTEM DEVELOPMENT | 35 | 788 | 5.84e-21 | 4.18e-19 |
| 65 | CIRCULATORY SYSTEM DEVELOPMENT | 35 | 788 | 5.84e-21 | 4.18e-19 |
| 66 | GLAND DEVELOPMENT | 27 | 395 | 6.253e-21 | 4.408e-19 |
| 67 | REGULATION OF STEM CELL DIFFERENTIATION | 18 | 113 | 7.769e-21 | 5.395e-19 |
| 68 | NON CANONICAL WNT SIGNALING PATHWAY | 19 | 140 | 1.446e-20 | 9.896e-19 |
| 69 | POSITIVE REGULATION OF STEM CELL DIFFERENTIATION | 14 | 50 | 3.573e-20 | 2.409e-18 |
| 70 | CARTILAGE DEVELOPMENT | 19 | 147 | 3.763e-20 | 2.501e-18 |
| 71 | CELLULAR COMPONENT MORPHOGENESIS | 36 | 900 | 4.5e-20 | 2.949e-18 |
| 72 | REGULATION OF INTRACELLULAR TRANSPORT | 31 | 621 | 5.411e-20 | 3.497e-18 |
| 73 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 43 | 1395 | 8.293e-20 | 5.286e-18 |
| 74 | EMBRYONIC ORGAN MORPHOGENESIS | 23 | 279 | 1.014e-19 | 6.374e-18 |
| 75 | BRANCHING MORPHOGENESIS OF AN EPITHELIAL TUBE | 18 | 131 | 1.249e-19 | 7.749e-18 |
| 76 | REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 21 | 218 | 1.798e-19 | 1.101e-17 |
| 77 | SMAD PROTEIN SIGNAL TRANSDUCTION | 14 | 56 | 2.145e-19 | 1.296e-17 |
| 78 | REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 21 | 220 | 2.178e-19 | 1.299e-17 |
| 79 | REGULATION OF OSTEOBLAST DIFFERENTIATION | 17 | 112 | 2.348e-19 | 1.383e-17 |
| 80 | REGULATION OF MAPK CASCADE | 31 | 660 | 3.134e-19 | 1.823e-17 |
| 81 | RESPONSE TO ENDOGENOUS STIMULUS | 43 | 1450 | 3.571e-19 | 2.038e-17 |
| 82 | CONNECTIVE TISSUE DEVELOPMENT | 20 | 194 | 3.591e-19 | 2.038e-17 |
| 83 | MORPHOGENESIS OF A BRANCHING STRUCTURE | 19 | 167 | 4.477e-19 | 2.51e-17 |
| 84 | ESTABLISHMENT OR MAINTENANCE OF CELL POLARITY | 18 | 141 | 4.892e-19 | 2.71e-17 |
| 85 | MAMMARY GLAND DEVELOPMENT | 17 | 117 | 5.097e-19 | 2.79e-17 |
| 86 | REGULATION OF PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 14 | 60 | 6.282e-19 | 3.399e-17 |
| 87 | POSITIVE REGULATION OF EPITHELIAL TO MESENCHYMAL TRANSITION | 12 | 34 | 7.466e-19 | 3.993e-17 |
| 88 | REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 20 | 207 | 1.317e-18 | 6.962e-17 |
| 89 | REGULATION OF CARTILAGE DEVELOPMENT | 14 | 63 | 1.334e-18 | 6.973e-17 |
| 90 | POSITIVE REGULATION OF PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 13 | 48 | 1.354e-18 | 6.999e-17 |
| 91 | POSITIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 38 | 1152 | 2.284e-18 | 1.168e-16 |
| 92 | REGULATION OF PROTEIN IMPORT | 19 | 183 | 2.591e-18 | 1.296e-16 |
| 93 | POSITIVE REGULATION OF OSSIFICATION | 15 | 84 | 2.584e-18 | 1.296e-16 |
| 94 | SKELETAL SYSTEM DEVELOPMENT | 26 | 455 | 3.111e-18 | 1.54e-16 |
| 95 | CELL JUNCTION ORGANIZATION | 19 | 185 | 3.189e-18 | 1.562e-16 |
| 96 | EYE DEVELOPMENT | 23 | 326 | 3.331e-18 | 1.598e-16 |
| 97 | REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 46 | 1784 | 3.331e-18 | 1.598e-16 |
| 98 | NEGATIVE REGULATION OF CELL DIFFERENTIATION | 29 | 609 | 3.737e-18 | 1.774e-16 |
| 99 | MESENCHYME DEVELOPMENT | 19 | 190 | 5.3e-18 | 2.491e-16 |
| 100 | NEGATIVE REGULATION OF MULTICELLULAR ORGANISMAL PROCESS | 35 | 983 | 6.611e-18 | 3.076e-16 |
| 101 | CENTRAL NERVOUS SYSTEM DEVELOPMENT | 33 | 872 | 1.192e-17 | 5.492e-16 |
| 102 | REGULATION OF TRANSFERASE ACTIVITY | 34 | 946 | 1.651e-17 | 7.531e-16 |
| 103 | EPITHELIAL CELL DIFFERENTIATION | 26 | 495 | 2.445e-17 | 1.104e-15 |
| 104 | POSITIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 23 | 360 | 2.99e-17 | 1.338e-15 |
| 105 | REGULATION OF KINASE ACTIVITY | 31 | 776 | 3.114e-17 | 1.38e-15 |
| 106 | REGULATION OF APOPTOTIC SIGNALING PATHWAY | 23 | 363 | 3.588e-17 | 1.575e-15 |
| 107 | PATTERN SPECIFICATION PROCESS | 24 | 418 | 6.463e-17 | 2.811e-15 |
| 108 | HEART DEVELOPMENT | 25 | 466 | 6.638e-17 | 2.86e-15 |
| 109 | DEVELOPMENTAL GROWTH | 22 | 333 | 7.759e-17 | 3.312e-15 |
| 110 | REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 30 | 750 | 1.076e-16 | 4.551e-15 |
| 111 | UROGENITAL SYSTEM DEVELOPMENT | 21 | 299 | 1.215e-16 | 5.093e-15 |
| 112 | MESODERM MORPHOGENESIS | 13 | 66 | 1.309e-16 | 5.44e-15 |
| 113 | MORPHOGENESIS OF EMBRYONIC EPITHELIUM | 16 | 134 | 1.488e-16 | 6.128e-15 |
| 114 | NEGATIVE REGULATION OF CELL PROLIFERATION | 28 | 643 | 1.554e-16 | 6.345e-15 |
| 115 | REGULATION OF EPITHELIAL TO MESENCHYMAL TRANSITION | 13 | 67 | 1.616e-16 | 6.54e-15 |
| 116 | FORMATION OF PRIMARY GERM LAYER | 15 | 110 | 1.799e-16 | 7.218e-15 |
| 117 | RESPIRATORY SYSTEM DEVELOPMENT | 18 | 197 | 2.125e-16 | 8.452e-15 |
| 118 | MAMMARY GLAND EPITHELIUM DEVELOPMENT | 12 | 53 | 3.297e-16 | 1.3e-14 |
| 119 | POSITIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 17 | 171 | 3.675e-16 | 1.437e-14 |
| 120 | SENSORY ORGAN MORPHOGENESIS | 19 | 239 | 3.958e-16 | 1.535e-14 |
| 121 | RESPONSE TO TRANSFORMING GROWTH FACTOR BETA | 16 | 144 | 4.788e-16 | 1.841e-14 |
| 122 | NEGATIVE REGULATION OF NITROGEN COMPOUND METABOLIC PROCESS | 40 | 1517 | 5.341e-16 | 1.988e-14 |
| 123 | RESPONSE TO BMP | 14 | 94 | 5.314e-16 | 1.988e-14 |
| 124 | CELLULAR RESPONSE TO BMP STIMULUS | 14 | 94 | 5.314e-16 | 1.988e-14 |
| 125 | MESODERM DEVELOPMENT | 15 | 118 | 5.317e-16 | 1.988e-14 |
| 126 | POSITIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 25 | 514 | 6.516e-16 | 2.406e-14 |
| 127 | REGULATION OF EPITHELIAL CELL PROLIFERATION | 20 | 285 | 7.013e-16 | 2.57e-14 |
| 128 | POSITIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 33 | 1004 | 7.464e-16 | 2.713e-14 |
| 129 | HEART MORPHOGENESIS | 18 | 212 | 7.834e-16 | 2.826e-14 |
| 130 | OSSIFICATION | 19 | 251 | 9.808e-16 | 3.51e-14 |
| 131 | POSITIVE REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 14 | 100 | 1.307e-15 | 4.642e-14 |
| 132 | STEM CELL DIFFERENTIATION | 17 | 190 | 2.181e-15 | 7.688e-14 |
| 133 | REGULATION OF PROTEIN TARGETING | 20 | 307 | 2.927e-15 | 1.024e-13 |
| 134 | POSITIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 16 | 162 | 3.181e-15 | 1.104e-13 |
| 135 | MESENCHYMAL CELL DIFFERENTIATION | 15 | 134 | 3.707e-15 | 1.278e-13 |
| 136 | NEGATIVE REGULATION OF GROWTH | 18 | 236 | 5.178e-15 | 1.771e-13 |
| 137 | LENS DEVELOPMENT IN CAMERA TYPE EYE | 12 | 66 | 5.694e-15 | 1.934e-13 |
| 138 | REGULATION OF TRANSPORT | 42 | 1804 | 6.146e-15 | 2.072e-13 |
| 139 | REGULATION OF STEM CELL PROLIFERATION | 13 | 88 | 6.875e-15 | 2.301e-13 |
| 140 | CELLULAR MACROMOLECULE LOCALIZATION | 35 | 1234 | 7.004e-15 | 2.328e-13 |
| 141 | MIDBRAIN DEVELOPMENT | 13 | 90 | 9.314e-15 | 3.052e-13 |
| 142 | MESONEPHROS DEVELOPMENT | 13 | 90 | 9.314e-15 | 3.052e-13 |
| 143 | REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 23 | 470 | 9.597e-15 | 3.123e-13 |
| 144 | REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 40 | 1656 | 1.02e-14 | 3.296e-13 |
| 145 | NEGATIVE REGULATION OF GENE EXPRESSION | 38 | 1493 | 1.13e-14 | 3.627e-13 |
| 146 | REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 21 | 381 | 1.545e-14 | 4.925e-13 |
| 147 | NEURAL TUBE DEVELOPMENT | 15 | 149 | 1.837e-14 | 5.815e-13 |
| 148 | POSITIVE REGULATION OF BIOMINERAL TISSUE DEVELOPMENT | 10 | 38 | 2.006e-14 | 6.264e-13 |
| 149 | MESENCHYME MORPHOGENESIS | 10 | 38 | 2.006e-14 | 6.264e-13 |
| 150 | PROTEIN COMPLEX SUBUNIT ORGANIZATION | 38 | 1527 | 2.314e-14 | 7.18e-13 |
| 151 | POSITIVE REGULATION OF WNT SIGNALING PATHWAY | 15 | 152 | 2.477e-14 | 7.632e-13 |
| 152 | GLAND MORPHOGENESIS | 13 | 97 | 2.549e-14 | 7.804e-13 |
| 153 | POSITIVE REGULATION OF MOLECULAR FUNCTION | 41 | 1791 | 2.587e-14 | 7.869e-13 |
| 154 | REGULATION OF CELL MORPHOGENESIS | 24 | 552 | 3.169e-14 | 9.574e-13 |
| 155 | KIDNEY EPITHELIUM DEVELOPMENT | 14 | 125 | 3.192e-14 | 9.582e-13 |
| 156 | EPITHELIAL TO MESENCHYMAL TRANSITION | 11 | 56 | 3.246e-14 | 9.681e-13 |
| 157 | REGULATION OF CATENIN IMPORT INTO NUCLEUS | 9 | 27 | 3.703e-14 | 1.097e-12 |
| 158 | CELL CYCLE | 35 | 1316 | 4.748e-14 | 1.398e-12 |
| 159 | NEGATIVE REGULATION OF PROTEIN METABOLIC PROCESS | 32 | 1087 | 4.835e-14 | 1.415e-12 |
| 160 | PROTEIN PHOSPHORYLATION | 30 | 944 | 4.919e-14 | 1.422e-12 |
| 161 | REGIONALIZATION | 19 | 311 | 4.908e-14 | 1.422e-12 |
| 162 | TUBE FORMATION | 14 | 129 | 4.981e-14 | 1.431e-12 |
| 163 | POSITIVE REGULATION OF CELL PROLIFERATION | 28 | 814 | 5.83e-14 | 1.654e-12 |
| 164 | INTRACELLULAR SIGNAL TRANSDUCTION | 38 | 1572 | 5.799e-14 | 1.654e-12 |
| 165 | REPRODUCTIVE SYSTEM DEVELOPMENT | 21 | 408 | 5.921e-14 | 1.67e-12 |
| 166 | SINGLE ORGANISM CELL ADHESION | 22 | 459 | 5.962e-14 | 1.671e-12 |
| 167 | ENZYME LINKED RECEPTOR PROTEIN SIGNALING PATHWAY | 26 | 689 | 6.379e-14 | 1.777e-12 |
| 168 | REGULATION OF CELL ADHESION | 25 | 629 | 6.53e-14 | 1.809e-12 |
| 169 | REGULATION OF ORGANELLE ORGANIZATION | 33 | 1178 | 7.149e-14 | 1.968e-12 |
| 170 | POSITIVE REGULATION OF INTRACELLULAR TRANSPORT | 20 | 370 | 1.001e-13 | 2.74e-12 |
| 171 | REGULATION OF ESTABLISHMENT OF PLANAR POLARITY | 13 | 110 | 1.357e-13 | 3.694e-12 |
| 172 | PALATE DEVELOPMENT | 12 | 85 | 1.381e-13 | 3.737e-12 |
| 173 | REGULATION OF CYTOPLASMIC TRANSPORT | 22 | 481 | 1.536e-13 | 4.131e-12 |
| 174 | POSITIVE REGULATION OF KINASE ACTIVITY | 22 | 482 | 1.602e-13 | 4.283e-12 |
| 175 | NEGATIVE REGULATION OF PHOSPHORUS METABOLIC PROCESS | 23 | 541 | 1.86e-13 | 4.917e-12 |
| 176 | NEGATIVE REGULATION OF PHOSPHATE METABOLIC PROCESS | 23 | 541 | 1.86e-13 | 4.917e-12 |
| 177 | REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 19 | 337 | 2.073e-13 | 5.442e-12 |
| 178 | DEVELOPMENTAL PROCESS INVOLVED IN REPRODUCTION | 24 | 602 | 2.082e-13 | 5.442e-12 |
| 179 | RHYTHMIC PROCESS | 18 | 298 | 2.916e-13 | 7.579e-12 |
| 180 | POSITIVE REGULATION OF TRANSFERASE ACTIVITY | 24 | 616 | 3.414e-13 | 8.777e-12 |
| 181 | NEGATIVE REGULATION OF PROTEIN MODIFICATION PROCESS | 24 | 616 | 3.414e-13 | 8.777e-12 |
| 182 | REGULATION OF CELL CYCLE | 29 | 949 | 3.781e-13 | 9.667e-12 |
| 183 | NEURON DEVELOPMENT | 25 | 687 | 4.672e-13 | 1.188e-11 |
| 184 | NEURAL TUBE FORMATION | 12 | 94 | 4.789e-13 | 1.211e-11 |
| 185 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 12 | 95 | 5.453e-13 | 1.372e-11 |
| 186 | REGULATION OF GROWTH | 24 | 633 | 6.119e-13 | 1.531e-11 |
| 187 | POSITIVE REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 9 | 36 | 7.104e-13 | 1.768e-11 |
| 188 | OSTEOBLAST DIFFERENTIATION | 13 | 126 | 8.058e-13 | 1.994e-11 |
| 189 | REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 19 | 365 | 8.556e-13 | 2.106e-11 |
| 190 | POSITIVE REGULATION OF MAPK CASCADE | 21 | 470 | 9.136e-13 | 2.226e-11 |
| 191 | ANTERIOR POSTERIOR PATTERN SPECIFICATION | 15 | 194 | 9.119e-13 | 2.226e-11 |
| 192 | REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 15 | 197 | 1.141e-12 | 2.764e-11 |
| 193 | REGULATION OF BINDING | 17 | 283 | 1.532e-12 | 3.675e-11 |
| 194 | SKELETAL SYSTEM MORPHOGENESIS | 15 | 201 | 1.528e-12 | 3.675e-11 |
| 195 | WNT SIGNALING PATHWAY CALCIUM MODULATING PATHWAY | 9 | 39 | 1.575e-12 | 3.758e-11 |
| 196 | ODONTOGENESIS | 12 | 105 | 1.85e-12 | 4.393e-11 |
| 197 | REGULATION OF HYDROLASE ACTIVITY | 33 | 1327 | 1.936e-12 | 4.574e-11 |
| 198 | MAMMARY GLAND MORPHOGENESIS | 9 | 40 | 2.022e-12 | 4.751e-11 |
| 199 | RESPONSE TO RETINOIC ACID | 12 | 107 | 2.326e-12 | 5.439e-11 |
| 200 | REGULATION OF SMAD PROTEIN IMPORT INTO NUCLEUS | 7 | 16 | 2.886e-12 | 6.583e-11 |
| 201 | STEM CELL PROLIFERATION | 10 | 60 | 2.859e-12 | 6.583e-11 |
| 202 | CHONDROCYTE DIFFERENTIATION | 10 | 60 | 2.859e-12 | 6.583e-11 |
| 203 | PARAXIAL MESODERM DEVELOPMENT | 7 | 16 | 2.886e-12 | 6.583e-11 |
| 204 | POSITIVE REGULATION OF OSTEOBLAST DIFFERENTIATION | 10 | 60 | 2.859e-12 | 6.583e-11 |
| 205 | POSITIVE REGULATION OF CATALYTIC ACTIVITY | 35 | 1518 | 2.997e-12 | 6.802e-11 |
| 206 | DEVELOPMENTAL INDUCTION | 8 | 27 | 3.061e-12 | 6.915e-11 |
| 207 | RESPONSE TO LIPID | 27 | 888 | 3.209e-12 | 7.214e-11 |
| 208 | PHOSPHATE CONTAINING COMPOUND METABOLIC PROCESS | 40 | 1977 | 3.241e-12 | 7.251e-11 |
| 209 | REGULATION OF MITOCHONDRIAL OUTER MEMBRANE PERMEABILIZATION INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 9 | 43 | 4.106e-12 | 9.141e-11 |
| 210 | PROTEIN LOCALIZATION | 38 | 1805 | 4.172e-12 | 9.244e-11 |
| 211 | CELL DIVISION | 20 | 460 | 5.539e-12 | 1.222e-10 |
| 212 | REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 8 | 29 | 5.859e-12 | 1.274e-10 |
| 213 | POSITIVE REGULATION OF PROTEIN INSERTION INTO MITOCHONDRIAL MEMBRANE INVOLVED IN APOPTOTIC SIGNALING PATHWAY | 8 | 29 | 5.859e-12 | 1.274e-10 |
| 214 | POSITIVE REGULATION OF CARTILAGE DEVELOPMENT | 8 | 29 | 5.859e-12 | 1.274e-10 |
| 215 | FOREBRAIN DEVELOPMENT | 18 | 357 | 6.126e-12 | 1.326e-10 |
| 216 | DIGESTIVE SYSTEM DEVELOPMENT | 13 | 148 | 6.438e-12 | 1.387e-10 |
| 217 | CELLULAR RESPONSE TO RETINOIC ACID | 10 | 65 | 6.617e-12 | 1.419e-10 |
| 218 | SEGMENTATION | 11 | 89 | 6.726e-12 | 1.436e-10 |
| 219 | IN UTERO EMBRYONIC DEVELOPMENT | 17 | 311 | 6.924e-12 | 1.471e-10 |
| 220 | PHOSPHORYLATION | 31 | 1228 | 7.309e-12 | 1.546e-10 |
| 221 | REGULATION OF CHONDROCYTE DIFFERENTIATION | 9 | 46 | 7.9e-12 | 1.663e-10 |
| 222 | POSITIVE REGULATION OF CANONICAL WNT SIGNALING PATHWAY | 12 | 119 | 8.376e-12 | 1.756e-10 |
| 223 | DORSAL VENTRAL PATTERN FORMATION | 11 | 91 | 8.627e-12 | 1.8e-10 |
| 224 | CELL CYCLE PROCESS | 29 | 1081 | 9.355e-12 | 1.943e-10 |
| 225 | REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 15 | 229 | 1.003e-11 | 2.074e-10 |
| 226 | CELL MORPHOGENESIS INVOLVED IN NEURON DIFFERENTIATION | 18 | 368 | 1.015e-11 | 2.089e-10 |
| 227 | POSITIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 13 | 154 | 1.07e-11 | 2.193e-10 |
| 228 | REGULATION OF ORGAN FORMATION | 8 | 32 | 1.414e-11 | 2.886e-10 |
| 229 | REGULATION OF DEPHOSPHORYLATION | 13 | 158 | 1.483e-11 | 3.013e-10 |
| 230 | EAR DEVELOPMENT | 14 | 195 | 1.494e-11 | 3.022e-10 |
| 231 | NEURON PROJECTION DEVELOPMENT | 21 | 545 | 1.507e-11 | 3.035e-10 |
| 232 | NEGATIVE REGULATION OF CELL CYCLE | 19 | 433 | 1.691e-11 | 3.391e-10 |
| 233 | DORSAL VENTRAL AXIS SPECIFICATION | 7 | 20 | 1.917e-11 | 3.828e-10 |
| 234 | LOCOMOTION | 29 | 1114 | 1.936e-11 | 3.85e-10 |
| 235 | CELL JUNCTION ASSEMBLY | 12 | 129 | 2.194e-11 | 4.345e-10 |
| 236 | REGULATION OF DEVELOPMENTAL GROWTH | 16 | 289 | 2.487e-11 | 4.884e-10 |
| 237 | POSITIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 16 | 289 | 2.487e-11 | 4.884e-10 |
| 238 | PROTEIN COMPLEX BIOGENESIS | 29 | 1132 | 2.847e-11 | 5.543e-10 |
| 239 | PROTEIN COMPLEX ASSEMBLY | 29 | 1132 | 2.847e-11 | 5.543e-10 |
| 240 | REGULATION OF BIOMINERAL TISSUE DEVELOPMENT | 10 | 75 | 2.911e-11 | 5.645e-10 |
| 241 | REPRODUCTION | 31 | 1297 | 2.943e-11 | 5.681e-10 |
| 242 | POSITIVE REGULATION OF STRESS ACTIVATED PROTEIN KINASE SIGNALING CASCADE | 12 | 135 | 3.762e-11 | 7.234e-10 |
| 243 | DEVELOPMENTAL GROWTH INVOLVED IN MORPHOGENESIS | 11 | 104 | 3.805e-11 | 7.286e-10 |
| 244 | POSITIVE REGULATION OF ORGANELLE ORGANIZATION | 21 | 573 | 3.831e-11 | 7.307e-10 |
| 245 | REGULATION OF CELLULAR COMPONENT MOVEMENT | 24 | 771 | 3.858e-11 | 7.327e-10 |
| 246 | POSITIVE REGULATION OF SMAD PROTEIN IMPORT INTO NUCLEUS | 6 | 12 | 4.081e-11 | 7.719e-10 |
| 247 | REGULATION OF KIDNEY DEVELOPMENT | 9 | 55 | 4.355e-11 | 8.204e-10 |
| 248 | REGULATION OF FAT CELL DIFFERENTIATION | 11 | 106 | 4.694e-11 | 8.807e-10 |
| 249 | NEGATIVE REGULATION OF MOLECULAR FUNCTION | 28 | 1079 | 4.99e-11 | 9.325e-10 |
| 250 | NEGATIVE REGULATION OF CELL DEVELOPMENT | 16 | 303 | 5.04e-11 | 9.381e-10 |
| 251 | POSITIVE REGULATION OF TRANSPORT | 26 | 936 | 6.388e-11 | 1.184e-09 |
| 252 | PATHWAY RESTRICTED SMAD PROTEIN PHOSPHORYLATION | 6 | 13 | 7.541e-11 | 1.392e-09 |
| 253 | NEGATIVE REGULATION OF CELL DEATH | 25 | 872 | 8.164e-11 | 1.501e-09 |
| 254 | NEGATIVE REGULATION OF DEVELOPMENTAL GROWTH | 10 | 84 | 9.262e-11 | 1.697e-09 |
| 255 | NEGATIVE REGULATION OF PHOSPHORYLATION | 18 | 422 | 9.618e-11 | 1.755e-09 |
| 256 | REGULATION OF MAP KINASE ACTIVITY | 16 | 319 | 1.081e-10 | 1.957e-09 |
| 257 | NEGATIVE REGULATION OF TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 23 | 740 | 1.078e-10 | 1.957e-09 |
| 258 | CELL CELL ADHESION | 21 | 608 | 1.144e-10 | 2.063e-09 |
| 259 | REGULATION OF ESTABLISHMENT OF PLANAR POLARITY INVOLVED IN NEURAL TUBE CLOSURE | 6 | 14 | 1.313e-10 | 2.359e-09 |
| 260 | RESPONSE TO OXYGEN CONTAINING COMPOUND | 31 | 1381 | 1.42e-10 | 2.541e-09 |
| 261 | ESTABLISHMENT OF CELL POLARITY | 10 | 88 | 1.484e-10 | 2.645e-09 |
| 262 | REGULATION OF CELL CELL ADHESION | 17 | 380 | 1.603e-10 | 2.846e-09 |
| 263 | POSITIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 18 | 437 | 1.695e-10 | 2.998e-09 |
| 264 | REGULATION OF CELLULAR RESPONSE TO STRESS | 22 | 691 | 1.843e-10 | 3.249e-09 |
| 265 | AXIS SPECIFICATION | 10 | 90 | 1.861e-10 | 3.268e-09 |
| 266 | POSITIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 11 | 121 | 1.997e-10 | 3.493e-09 |
| 267 | AXIS ELONGATION | 7 | 27 | 2.12e-10 | 3.694e-09 |
| 268 | INNER EAR MORPHOGENESIS | 10 | 92 | 2.322e-10 | 4.031e-09 |
| 269 | RESPONSE TO ORGANIC CYCLIC COMPOUND | 25 | 917 | 2.349e-10 | 4.064e-09 |
| 270 | CELL ACTIVATION | 20 | 568 | 2.386e-10 | 4.111e-09 |
| 271 | CELL DEATH | 26 | 1001 | 2.737e-10 | 4.7e-09 |
| 272 | MAMMARY GLAND DUCT MORPHOGENESIS | 7 | 28 | 2.812e-10 | 4.81e-09 |
| 273 | CELLULAR RESPONSE TO LIPID | 18 | 457 | 3.485e-10 | 5.94e-09 |
| 274 | REGULATION OF HEART MORPHOGENESIS | 7 | 29 | 3.688e-10 | 6.263e-09 |
| 275 | NEURON PROJECTION MORPHOGENESIS | 17 | 402 | 3.811e-10 | 6.448e-09 |
| 276 | REGULATION OF MEMBRANE PERMEABILITY | 9 | 70 | 4.126e-10 | 6.956e-09 |
| 277 | RESPONSE TO ABIOTIC STIMULUS | 26 | 1024 | 4.454e-10 | 7.482e-09 |
| 278 | RESPONSE TO EXTERNAL STIMULUS | 35 | 1821 | 4.67e-10 | 7.8e-09 |
| 279 | DIGESTIVE TRACT MORPHOGENESIS | 8 | 48 | 4.677e-10 | 7.8e-09 |
| 280 | MOVEMENT OF CELL OR SUBCELLULAR COMPONENT | 29 | 1275 | 4.739e-10 | 7.875e-09 |
| 281 | APPENDAGE DEVELOPMENT | 12 | 169 | 5.197e-10 | 8.575e-09 |
| 282 | LIMB DEVELOPMENT | 12 | 169 | 5.197e-10 | 8.575e-09 |
| 283 | CELL CYCLE PHASE TRANSITION | 14 | 255 | 5.28e-10 | 8.682e-09 |
| 284 | BRANCH ELONGATION OF AN EPITHELIUM | 6 | 17 | 5.332e-10 | 8.735e-09 |
| 285 | REGULATION OF ORGAN GROWTH | 9 | 73 | 6.066e-10 | 9.869e-09 |
| 286 | REGULATION OF NEURAL PRECURSOR CELL PROLIFERATION | 9 | 73 | 6.066e-10 | 9.869e-09 |
| 287 | REGULATION OF MUSCLE TISSUE DEVELOPMENT | 10 | 103 | 7.182e-10 | 1.16e-08 |
| 288 | REGULATION OF MUSCLE ORGAN DEVELOPMENT | 10 | 103 | 7.182e-10 | 1.16e-08 |
| 289 | NEGATIVE REGULATION OF NERVOUS SYSTEM DEVELOPMENT | 14 | 262 | 7.519e-10 | 1.211e-08 |
| 290 | MACROMOLECULAR COMPLEX ASSEMBLY | 30 | 1398 | 8.87e-10 | 1.423e-08 |
| 291 | CELL PROJECTION ORGANIZATION | 24 | 902 | 9.282e-10 | 1.484e-08 |
| 292 | REGULATION OF MORPHOGENESIS OF A BRANCHING STRUCTURE | 8 | 53 | 1.071e-09 | 1.707e-08 |
| 293 | MACROMOLECULAR COMPLEX DISASSEMBLY | 12 | 182 | 1.218e-09 | 1.935e-08 |
| 294 | REGULATION OF MESENCHYMAL CELL PROLIFERATION | 7 | 34 | 1.24e-09 | 1.962e-08 |
| 295 | POSITIVE REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 13 | 228 | 1.432e-09 | 2.258e-08 |
| 296 | REGULATION OF JUN KINASE ACTIVITY | 9 | 81 | 1.565e-09 | 2.46e-08 |
| 297 | EAR MORPHOGENESIS | 10 | 112 | 1.646e-09 | 2.578e-08 |
| 298 | KIDNEY MORPHOGENESIS | 9 | 82 | 1.749e-09 | 2.731e-08 |
| 299 | NEPHRON DEVELOPMENT | 10 | 115 | 2.134e-09 | 3.321e-08 |
| 300 | EMBRYONIC PATTERN SPECIFICATION | 8 | 58 | 2.258e-09 | 3.502e-08 |
| 301 | COCHLEA MORPHOGENESIS | 6 | 21 | 2.292e-09 | 3.542e-08 |
| 302 | ESTABLISHMENT OR MAINTENANCE OF APICAL BASAL CELL POLARITY | 7 | 37 | 2.337e-09 | 3.588e-08 |
| 303 | ESTABLISHMENT OR MAINTENANCE OF BIPOLAR CELL POLARITY | 7 | 37 | 2.337e-09 | 3.588e-08 |
| 304 | CARDIAC SEPTUM DEVELOPMENT | 9 | 85 | 2.42e-09 | 3.704e-08 |
| 305 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 11 | 153 | 2.477e-09 | 3.778e-08 |
| 306 | REGULATION OF RESPONSE TO STRESS | 30 | 1468 | 2.796e-09 | 4.252e-08 |
| 307 | POSITIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 23 | 876 | 2.83e-09 | 4.289e-08 |
| 308 | SIGNAL TRANSDUCTION BY PROTEIN PHOSPHORYLATION | 16 | 404 | 3.343e-09 | 5.051e-08 |
| 309 | POSITIVE REGULATION OF STEM CELL PROLIFERATION | 8 | 61 | 3.417e-09 | 5.145e-08 |
| 310 | CELLULAR RESPONSE TO ORGANIC CYCLIC COMPOUND | 17 | 465 | 3.455e-09 | 5.186e-08 |
| 311 | REGULATION OF JNK CASCADE | 11 | 159 | 3.72e-09 | 5.565e-08 |
| 312 | EMBRYONIC SKELETAL SYSTEM DEVELOPMENT | 10 | 122 | 3.808e-09 | 5.679e-08 |
| 313 | SOMITOGENESIS | 8 | 62 | 3.903e-09 | 5.802e-08 |
| 314 | VASCULATURE DEVELOPMENT | 17 | 469 | 3.927e-09 | 5.819e-08 |
| 315 | APICAL JUNCTION ASSEMBLY | 7 | 40 | 4.168e-09 | 6.156e-08 |
| 316 | POSITIVE REGULATION OF JUN KINASE ACTIVITY | 8 | 63 | 4.448e-09 | 6.55e-08 |
| 317 | MEMBRANE ORGANIZATION | 23 | 899 | 4.621e-09 | 6.782e-08 |
| 318 | MITOCHONDRIAL MEMBRANE ORGANIZATION | 9 | 92 | 4.927e-09 | 7.21e-08 |
| 319 | POSITIVE REGULATION OF MAP KINASE ACTIVITY | 12 | 207 | 5.26e-09 | 7.672e-08 |
| 320 | EMBRYONIC SKELETAL SYSTEM MORPHOGENESIS | 9 | 93 | 5.428e-09 | 7.868e-08 |
| 321 | NEGATIVE REGULATION OF CATALYTIC ACTIVITY | 22 | 829 | 5.422e-09 | 7.868e-08 |
| 322 | REGULATION OF PROTEIN PHOSPHATASE TYPE 2A ACTIVITY | 6 | 24 | 5.6e-09 | 8.092e-08 |
| 323 | PITUITARY GLAND DEVELOPMENT | 7 | 42 | 5.971e-09 | 8.574e-08 |
| 324 | MAMMARY GLAND EPITHELIAL CELL PROLIFERATION | 5 | 12 | 5.952e-09 | 8.574e-08 |
| 325 | REGULATION OF PHOSPHATASE ACTIVITY | 10 | 128 | 6.078e-09 | 8.702e-08 |
| 326 | SKIN DEVELOPMENT | 12 | 211 | 6.525e-09 | 9.313e-08 |
| 327 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO NUCLEUS | 10 | 129 | 6.555e-09 | 9.328e-08 |
| 328 | BETA CATENIN TCF COMPLEX ASSEMBLY | 7 | 43 | 7.096e-09 | 1.007e-07 |
| 329 | BRANCHING INVOLVED IN URETERIC BUD MORPHOGENESIS | 7 | 44 | 8.396e-09 | 1.187e-07 |
| 330 | MUSCLE STRUCTURE DEVELOPMENT | 16 | 432 | 8.647e-09 | 1.219e-07 |
| 331 | SEX DIFFERENTIATION | 13 | 266 | 9.203e-09 | 1.294e-07 |
| 332 | NEGATIVE REGULATION OF EMBRYONIC DEVELOPMENT | 6 | 26 | 9.484e-09 | 1.329e-07 |
| 333 | CELL PART MORPHOGENESIS | 19 | 633 | 9.62e-09 | 1.344e-07 |
| 334 | EYE MORPHOGENESIS | 10 | 136 | 1.093e-08 | 1.523e-07 |
| 335 | NEGATIVE REGULATION OF NUCLEOCYTOPLASMIC TRANSPORT | 8 | 71 | 1.173e-08 | 1.63e-07 |
| 336 | POSITIVE REGULATION OF MESENCHYMAL CELL PROLIFERATION | 6 | 27 | 1.213e-08 | 1.68e-07 |
| 337 | CELL CYCLE G2 M PHASE TRANSITION | 10 | 138 | 1.259e-08 | 1.738e-07 |
| 338 | POSITIVE REGULATION OF PROTEIN IMPORT | 9 | 104 | 1.466e-08 | 2.018e-07 |
| 339 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 5 | 14 | 1.49e-08 | 2.045e-07 |
| 340 | VENTRICULAR SEPTUM MORPHOGENESIS | 6 | 28 | 1.536e-08 | 2.097e-07 |
| 341 | DOPAMINERGIC NEURON DIFFERENTIATION | 6 | 28 | 1.536e-08 | 2.097e-07 |
| 342 | IMMUNE SYSTEM DEVELOPMENT | 18 | 582 | 1.555e-08 | 2.115e-07 |
| 343 | REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 7 | 48 | 1.581e-08 | 2.145e-07 |
| 344 | CELL CELL JUNCTION ASSEMBLY | 8 | 74 | 1.637e-08 | 2.214e-07 |
| 345 | ODONTOGENESIS OF DENTIN CONTAINING TOOTH | 8 | 75 | 1.823e-08 | 2.459e-07 |
| 346 | CARDIAC SEPTUM MORPHOGENESIS | 7 | 49 | 1.835e-08 | 2.468e-07 |
| 347 | CARDIAC CHAMBER DEVELOPMENT | 10 | 144 | 1.895e-08 | 2.541e-07 |
| 348 | STEM CELL DIVISION | 6 | 29 | 1.928e-08 | 2.577e-07 |
| 349 | DIENCEPHALON DEVELOPMENT | 8 | 77 | 2.251e-08 | 3.001e-07 |
| 350 | TAXIS | 16 | 464 | 2.353e-08 | 3.128e-07 |
| 351 | ESTABLISHMENT OR MAINTENANCE OF EPITHELIAL CELL APICAL BASAL POLARITY | 6 | 30 | 2.398e-08 | 3.178e-07 |
| 352 | RENAL TUBULE DEVELOPMENT | 8 | 78 | 2.496e-08 | 3.29e-07 |
| 353 | SOMITE DEVELOPMENT | 8 | 78 | 2.496e-08 | 3.29e-07 |
| 354 | BONE MORPHOGENESIS | 8 | 79 | 2.763e-08 | 3.631e-07 |
| 355 | WOUND HEALING | 16 | 470 | 2.812e-08 | 3.686e-07 |
| 356 | MESONEPHRIC TUBULE MORPHOGENESIS | 7 | 53 | 3.227e-08 | 4.195e-07 |
| 357 | ORGAN INDUCTION | 5 | 16 | 3.219e-08 | 4.195e-07 |
| 358 | POSITIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 7 | 53 | 3.227e-08 | 4.195e-07 |
| 359 | NEGATIVE REGULATION OF CELLULAR COMPONENT ORGANIZATION | 19 | 684 | 3.314e-08 | 4.296e-07 |
| 360 | BICELLULAR TIGHT JUNCTION ASSEMBLY | 6 | 32 | 3.623e-08 | 4.683e-07 |
| 361 | NEGATIVE REGULATION OF MITOTIC CELL CYCLE | 11 | 199 | 3.848e-08 | 4.959e-07 |
| 362 | NEGATIVE REGULATION OF CYTOPLASMIC TRANSPORT | 9 | 117 | 4.124e-08 | 5.301e-07 |
| 363 | CRANIAL SKELETAL SYSTEM DEVELOPMENT | 7 | 55 | 4.206e-08 | 5.392e-07 |
| 364 | POSITIVE REGULATION OF EMBRYONIC DEVELOPMENT | 6 | 33 | 4.406e-08 | 5.632e-07 |
| 365 | NEGATIVE REGULATION OF STEM CELL PROLIFERATION | 5 | 17 | 4.538e-08 | 5.773e-07 |
| 366 | REGULATION OF NEURON DIFFERENTIATION | 17 | 554 | 4.541e-08 | 5.773e-07 |
| 367 | RESPONSE TO ALCOHOL | 14 | 362 | 4.611e-08 | 5.846e-07 |
| 368 | POSITIVE REGULATION OF NEURON DIFFERENTIATION | 13 | 306 | 4.824e-08 | 6.099e-07 |
| 369 | EPIDERMIS DEVELOPMENT | 12 | 253 | 4.907e-08 | 6.187e-07 |
| 370 | BLOOD VESSEL MORPHOGENESIS | 14 | 364 | 4.937e-08 | 6.209e-07 |
| 371 | NEURON PROJECTION GUIDANCE | 11 | 205 | 5.217e-08 | 6.543e-07 |
| 372 | ORGAN FORMATION | 6 | 34 | 5.324e-08 | 6.659e-07 |
| 373 | LEUKOCYTE CELL CELL ADHESION | 12 | 255 | 5.349e-08 | 6.673e-07 |
| 374 | NEGATIVE REGULATION OF CELLULAR RESPONSE TO GROWTH FACTOR STIMULUS | 9 | 121 | 5.527e-08 | 6.876e-07 |
| 375 | RESPONSE TO WOUNDING | 17 | 563 | 5.73e-08 | 7.109e-07 |
| 376 | BIOLOGICAL ADHESION | 23 | 1032 | 5.94e-08 | 7.351e-07 |
| 377 | POSITIVE REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 12 | 258 | 6.081e-08 | 7.505e-07 |
| 378 | ENDOCRINE SYSTEM DEVELOPMENT | 9 | 123 | 6.373e-08 | 7.845e-07 |
| 379 | REGULATION OF CYTOSKELETON ORGANIZATION | 16 | 502 | 6.975e-08 | 8.563e-07 |
| 380 | EPITHELIAL CELL PROLIFERATION | 8 | 89 | 7.109e-08 | 8.705e-07 |
| 381 | POSITIVE REGULATION OF CELL ADHESION | 14 | 376 | 7.374e-08 | 9.005e-07 |
| 382 | NEGATIVE REGULATION OF MUSCLE ORGAN DEVELOPMENT | 6 | 36 | 7.634e-08 | 9.274e-07 |
| 383 | NEGATIVE REGULATION OF MUSCLE TISSUE DEVELOPMENT | 6 | 36 | 7.634e-08 | 9.274e-07 |
| 384 | POSITIVE REGULATION OF MITOCHONDRION ORGANIZATION | 10 | 167 | 7.756e-08 | 9.398e-07 |
| 385 | RESPONSE TO ACID CHEMICAL | 13 | 319 | 7.836e-08 | 9.471e-07 |
| 386 | REGULATION OF PROTEIN BINDING | 10 | 168 | 8.205e-08 | 9.89e-07 |
| 387 | EMBRYONIC DIGIT MORPHOGENESIS | 7 | 61 | 8.774e-08 | 1.055e-06 |
| 388 | REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION TO MITOCHONDRION | 9 | 128 | 9e-08 | 1.079e-06 |
| 389 | REGULATION OF PEPTIDYL THREONINE PHOSPHORYLATION | 6 | 37 | 9.066e-08 | 1.084e-06 |
| 390 | NEGATIVE REGULATION OF CELL GROWTH | 10 | 170 | 9.172e-08 | 1.094e-06 |
| 391 | RESPONSE TO ESTROGEN | 11 | 218 | 9.761e-08 | 1.162e-06 |
| 392 | NEPHRON EPITHELIUM DEVELOPMENT | 8 | 93 | 1.005e-07 | 1.192e-06 |
| 393 | SINGLE ORGANISM CELLULAR LOCALIZATION | 21 | 898 | 1.094e-07 | 1.295e-06 |
| 394 | TRACHEA DEVELOPMENT | 5 | 20 | 1.121e-07 | 1.32e-06 |
| 395 | BRANCHING INVOLVED IN MAMMARY GLAND DUCT MORPHOGENESIS | 5 | 20 | 1.121e-07 | 1.32e-06 |
| 396 | NEGATIVE REGULATION OF INTRACELLULAR PROTEIN TRANSPORT | 8 | 95 | 1.187e-07 | 1.395e-06 |
| 397 | CELLULAR RESPONSE TO ACID CHEMICAL | 10 | 175 | 1.204e-07 | 1.408e-06 |
| 398 | GLIOGENESIS | 10 | 175 | 1.204e-07 | 1.408e-06 |
| 399 | COCHLEA DEVELOPMENT | 6 | 39 | 1.26e-07 | 1.469e-06 |
| 400 | NEGATIVE REGULATION OF CELLULAR PROTEIN LOCALIZATION | 9 | 135 | 1.424e-07 | 1.656e-06 |
| 401 | NEGATIVE REGULATION OF ORGAN GROWTH | 5 | 21 | 1.464e-07 | 1.698e-06 |
| 402 | POSITIVE REGULATION OF CYTOPLASMIC TRANSPORT | 12 | 282 | 1.599e-07 | 1.851e-06 |
| 403 | REGULATION OF DNA METABOLIC PROCESS | 13 | 340 | 1.638e-07 | 1.891e-06 |
| 404 | NEGATIVE REGULATION OF FAT CELL DIFFERENTIATION | 6 | 41 | 1.719e-07 | 1.98e-06 |
| 405 | LYMPHOCYTE ACTIVATION | 13 | 342 | 1.752e-07 | 2.013e-06 |
| 406 | ACTIVIN RECEPTOR SIGNALING PATHWAY | 5 | 22 | 1.885e-07 | 2.155e-06 |
| 407 | SOMATIC STEM CELL DIVISION | 5 | 22 | 1.885e-07 | 2.155e-06 |
| 408 | CELL CELL SIGNALING | 19 | 767 | 1.978e-07 | 2.256e-06 |
| 409 | NEGATIVE REGULATION OF TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE SIGNALING PATHWAY | 8 | 102 | 2.067e-07 | 2.351e-06 |
| 410 | EPITHELIAL CELL DEVELOPMENT | 10 | 186 | 2.127e-07 | 2.414e-06 |
| 411 | REGULATION OF CIRCADIAN RHYTHM | 8 | 103 | 2.229e-07 | 2.524e-06 |
| 412 | NEURAL PRECURSOR CELL PROLIFERATION | 7 | 70 | 2.304e-07 | 2.602e-06 |
| 413 | LEUKOCYTE DIFFERENTIATION | 12 | 292 | 2.328e-07 | 2.62e-06 |
| 414 | NEGATIVE REGULATION OF INTRACELLULAR TRANSPORT | 9 | 143 | 2.331e-07 | 2.62e-06 |
| 415 | LEUKOCYTE ACTIVATION | 14 | 414 | 2.392e-07 | 2.681e-06 |
| 416 | REGULATION OF METANEPHROS DEVELOPMENT | 5 | 23 | 2.397e-07 | 2.681e-06 |
| 417 | CARDIAC CHAMBER MORPHOGENESIS | 8 | 104 | 2.403e-07 | 2.681e-06 |
| 418 | ENDODERM DEVELOPMENT | 7 | 71 | 2.543e-07 | 2.831e-06 |
| 419 | SYNAPSE ORGANIZATION | 9 | 145 | 2.624e-07 | 2.914e-06 |
| 420 | POSITIVE REGULATION OF LOCOMOTION | 14 | 420 | 2.847e-07 | 3.154e-06 |
| 421 | REGULATION OF CELL CYCLE PROCESS | 16 | 558 | 2.922e-07 | 3.229e-06 |
| 422 | ENDOCHONDRAL BONE MORPHOGENESIS | 6 | 45 | 3.052e-07 | 3.366e-06 |
| 423 | PANCREAS DEVELOPMENT | 7 | 73 | 3.085e-07 | 3.393e-06 |
| 424 | REGULATION OF PROTEOLYSIS | 18 | 711 | 3.128e-07 | 3.425e-06 |
| 425 | MALE SEX DIFFERENTIATION | 9 | 148 | 3.124e-07 | 3.425e-06 |
| 426 | REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY IN ABSENCE OF LIGAND | 6 | 46 | 3.493e-07 | 3.806e-06 |
| 427 | EMBRYONIC CRANIAL SKELETON MORPHOGENESIS | 6 | 46 | 3.493e-07 | 3.806e-06 |
| 428 | CELLULAR RESPONSE TO OXYGEN CONTAINING COMPOUND | 19 | 799 | 3.694e-07 | 4.016e-06 |
| 429 | RESPONSE TO STEROID HORMONE | 15 | 497 | 3.708e-07 | 4.022e-06 |
| 430 | LENS FIBER CELL DIFFERENTIATION | 5 | 25 | 3.748e-07 | 4.046e-06 |
| 431 | POSITIVE REGULATION OF PEPTIDYL THREONINE PHOSPHORYLATION | 5 | 25 | 3.748e-07 | 4.046e-06 |
| 432 | NEGATIVE REGULATION OF KINASE ACTIVITY | 11 | 250 | 3.866e-07 | 4.164e-06 |
| 433 | POSITIVE REGULATION OF NON CANONICAL WNT SIGNALING PATHWAY | 4 | 11 | 4.207e-07 | 4.521e-06 |
| 434 | CELL CYCLE ARREST | 9 | 154 | 4.375e-07 | 4.691e-06 |
| 435 | REGULATION OF BMP SIGNALING PATHWAY | 7 | 77 | 4.462e-07 | 4.773e-06 |
| 436 | RESPONSE TO HORMONE | 20 | 893 | 4.534e-07 | 4.828e-06 |
| 437 | ANTERIOR POSTERIOR AXIS SPECIFICATION | 6 | 48 | 4.531e-07 | 4.828e-06 |
| 438 | OVULATION CYCLE | 8 | 113 | 4.561e-07 | 4.845e-06 |
| 439 | NEGATIVE REGULATION OF INTRACELLULAR SIGNAL TRANSDUCTION | 14 | 437 | 4.589e-07 | 4.864e-06 |
| 440 | NEGATIVE REGULATION OF CARTILAGE DEVELOPMENT | 5 | 26 | 4.617e-07 | 4.883e-06 |
| 441 | BONE DEVELOPMENT | 9 | 156 | 4.88e-07 | 5.148e-06 |
| 442 | NEGATIVE REGULATION OF EPITHELIAL CELL PROLIFERATION | 8 | 115 | 5.218e-07 | 5.493e-06 |
| 443 | RESPONSE TO LITHIUM ION | 5 | 27 | 5.639e-07 | 5.923e-06 |
| 444 | NEGATIVE REGULATION OF CELL MORPHOGENESIS INVOLVED IN DIFFERENTIATION | 8 | 117 | 5.956e-07 | 6.241e-06 |
| 445 | ANATOMICAL STRUCTURE REGRESSION | 4 | 12 | 6.281e-07 | 6.524e-06 |
| 446 | REGULATION OF MACROPHAGE CYTOKINE PRODUCTION | 4 | 12 | 6.281e-07 | 6.524e-06 |
| 447 | TRACHEA MORPHOGENESIS | 4 | 12 | 6.281e-07 | 6.524e-06 |
| 448 | LYMPHOCYTE DIFFERENTIATION | 10 | 209 | 6.239e-07 | 6.524e-06 |
| 449 | METANEPHROS DEVELOPMENT | 7 | 81 | 6.323e-07 | 6.553e-06 |
| 450 | REGULATION OF DNA REPLICATION | 9 | 161 | 6.366e-07 | 6.582e-06 |
| 451 | POSITIVE REGULATION OF PROTEIN CATABOLIC PROCESS | 11 | 263 | 6.389e-07 | 6.591e-06 |
| 452 | CELL FATE COMMITMENT INVOLVED IN FORMATION OF PRIMARY GERM LAYER | 5 | 28 | 6.831e-07 | 7.032e-06 |
| 453 | REGULATION OF PROTEIN CATABOLIC PROCESS | 13 | 393 | 8.478e-07 | 8.708e-06 |
| 454 | T CELL DIFFERENTIATION | 8 | 123 | 8.725e-07 | 8.942e-06 |
| 455 | REGULATION OF CELL DIVISION | 11 | 272 | 8.897e-07 | 9.099e-06 |
| 456 | GDP METABOLIC PROCESS | 4 | 13 | 9.03e-07 | 9.215e-06 |
| 457 | POSITIVE REGULATION OF CELL CYCLE | 12 | 332 | 9.107e-07 | 9.272e-06 |
| 458 | REGULATION OF MITOCHONDRION ORGANIZATION | 10 | 218 | 9.168e-07 | 9.314e-06 |
| 459 | MITOTIC CELL CYCLE | 18 | 766 | 9.202e-07 | 9.329e-06 |
| 460 | VENTRICULAR SEPTUM DEVELOPMENT | 6 | 54 | 9.255e-07 | 9.362e-06 |
| 461 | REGULATION OF PROTEIN STABILITY | 10 | 221 | 1.038e-06 | 1.045e-05 |
| 462 | REGULATION OF MITOTIC CELL CYCLE | 14 | 468 | 1.037e-06 | 1.045e-05 |
| 463 | NEGATIVE REGULATION OF PROTEIN SERINE THREONINE KINASE ACTIVITY | 8 | 126 | 1.048e-06 | 1.053e-05 |
| 464 | POSITIVE REGULATION OF BINDING | 8 | 127 | 1.113e-06 | 1.115e-05 |
| 465 | OVULATION CYCLE PROCESS | 7 | 88 | 1.114e-06 | 1.115e-05 |
| 466 | NEGATIVE REGULATION OF CELL ADHESION | 10 | 223 | 1.127e-06 | 1.125e-05 |
| 467 | EMBRYONIC SKELETAL JOINT DEVELOPMENT | 4 | 14 | 1.258e-06 | 1.254e-05 |
| 468 | CIRCADIAN REGULATION OF GENE EXPRESSION | 6 | 57 | 1.281e-06 | 1.271e-05 |
| 469 | REGULATION OF CYTOKINE PRODUCTION INVOLVED IN IMMUNE RESPONSE | 6 | 57 | 1.281e-06 | 1.271e-05 |
| 470 | ENDOCARDIAL CUSHION DEVELOPMENT | 5 | 32 | 1.373e-06 | 1.359e-05 |
| 471 | MITOCHONDRIAL TRANSPORT | 9 | 177 | 1.406e-06 | 1.389e-05 |
| 472 | VASCULOGENESIS | 6 | 59 | 1.575e-06 | 1.553e-05 |
| 473 | EMBRYONIC AXIS SPECIFICATION | 5 | 33 | 1.61e-06 | 1.584e-05 |
| 474 | NEGATIVE REGULATION OF TRANSFERASE ACTIVITY | 12 | 351 | 1.629e-06 | 1.599e-05 |
| 475 | REGULATION OF PHOSPHOPROTEIN PHOSPHATASE ACTIVITY | 6 | 60 | 1.741e-06 | 1.706e-05 |
| 476 | BRAIN MORPHOGENESIS | 5 | 34 | 1.878e-06 | 1.836e-05 |
| 477 | TRANSCRIPTION FROM RNA POLYMERASE II PROMOTER | 17 | 724 | 1.926e-06 | 1.878e-05 |
| 478 | CIRCADIAN RHYTHM | 8 | 137 | 1.972e-06 | 1.92e-05 |
| 479 | POSITIVE REGULATION OF DNA METABOLIC PROCESS | 9 | 185 | 2.027e-06 | 1.969e-05 |
| 480 | PROTEIN LOCALIZATION TO SYNAPSE | 4 | 16 | 2.266e-06 | 2.192e-05 |
| 481 | APOPTOTIC PROCESS INVOLVED IN MORPHOGENESIS | 4 | 16 | 2.266e-06 | 2.192e-05 |
| 482 | POSITIVE REGULATION OF PROTEOLYSIS | 12 | 363 | 2.308e-06 | 2.228e-05 |
| 483 | POSITIVE REGULATION OF CELL CELL ADHESION | 10 | 243 | 2.442e-06 | 2.353e-05 |
| 484 | REGULATION OF TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 7 | 99 | 2.473e-06 | 2.372e-05 |
| 485 | REGULATION OF CELLULAR RESPONSE TO TRANSFORMING GROWTH FACTOR BETA STIMULUS | 7 | 99 | 2.473e-06 | 2.372e-05 |
| 486 | HEAD MORPHOGENESIS | 5 | 36 | 2.52e-06 | 2.413e-05 |
| 487 | PROTEIN DEPHOSPHORYLATION | 9 | 190 | 2.526e-06 | 2.413e-05 |
| 488 | POSITIVE REGULATION OF CELL PROJECTION ORGANIZATION | 11 | 303 | 2.542e-06 | 2.424e-05 |
| 489 | EPIDERMAL CELL DIFFERENTIATION | 8 | 142 | 2.581e-06 | 2.456e-05 |
| 490 | NEGATIVE REGULATION OF NEURON DIFFERENTIATION | 9 | 191 | 2.637e-06 | 2.504e-05 |
| 491 | CAMERA TYPE EYE MORPHOGENESIS | 7 | 101 | 2.828e-06 | 2.68e-05 |
| 492 | CELLULAR RESPONSE TO LITHIUM ION | 4 | 17 | 2.95e-06 | 2.784e-05 |
| 493 | ESTABLISHMENT OF TISSUE POLARITY | 4 | 17 | 2.95e-06 | 2.784e-05 |
| 494 | NEGATIVE REGULATION OF MAPK CASCADE | 8 | 145 | 3.017e-06 | 2.842e-05 |
| 495 | NEGATIVE REGULATION OF CELLULAR RESPONSE TO TRANSFORMING GROWTH FACTOR BETA STIMULUS | 6 | 66 | 3.068e-06 | 2.878e-05 |
| 496 | NEGATIVE REGULATION OF TRANSFORMING GROWTH FACTOR BETA RECEPTOR SIGNALING PATHWAY | 6 | 66 | 3.068e-06 | 2.878e-05 |
| 497 | CELL MOTILITY | 18 | 835 | 3.117e-06 | 2.912e-05 |
| 498 | LOCALIZATION OF CELL | 18 | 835 | 3.117e-06 | 2.912e-05 |
| 499 | RESPONSE TO ESTRADIOL | 8 | 146 | 3.176e-06 | 2.956e-05 |
| 500 | CELLULAR COMPONENT DISASSEMBLY | 14 | 515 | 3.172e-06 | 2.956e-05 |
| 501 | POSITIVE REGULATION OF EPITHELIAL CELL MIGRATION | 7 | 103 | 3.226e-06 | 2.996e-05 |
| 502 | NEURON FATE COMMITMENT | 6 | 67 | 3.353e-06 | 3.107e-05 |
| 503 | REGULATION OF ACTIN FILAMENT BASED PROCESS | 11 | 312 | 3.368e-06 | 3.116e-05 |
| 504 | KIDNEY MESENCHYME DEVELOPMENT | 4 | 18 | 3.775e-06 | 3.485e-05 |
| 505 | NEGATIVE REGULATION OF OSSIFICATION | 6 | 69 | 3.987e-06 | 3.667e-05 |
| 506 | SYNAPSE ASSEMBLY | 6 | 69 | 3.987e-06 | 3.667e-05 |
| 507 | REGULATION OF CELLULAR COMPONENT BIOGENESIS | 17 | 767 | 4.172e-06 | 3.829e-05 |
| 508 | NEGATIVE REGULATION OF OSTEOBLAST DIFFERENTIATION | 5 | 40 | 4.314e-06 | 3.951e-05 |
| 509 | SKIN EPIDERMIS DEVELOPMENT | 6 | 71 | 4.717e-06 | 4.312e-05 |
| 510 | REGULATION OF NON CANONICAL WNT SIGNALING PATHWAY | 4 | 19 | 4.759e-06 | 4.325e-05 |
| 511 | ENTRAINMENT OF CIRCADIAN CLOCK BY PHOTOPERIOD | 4 | 19 | 4.759e-06 | 4.325e-05 |
| 512 | POSITIVE REGULATION OF CHONDROCYTE DIFFERENTIATION | 4 | 19 | 4.759e-06 | 4.325e-05 |
| 513 | POSITIVE REGULATION OF KIDNEY DEVELOPMENT | 5 | 41 | 4.889e-06 | 4.417e-05 |
| 514 | LUNG ALVEOLUS DEVELOPMENT | 5 | 41 | 4.889e-06 | 4.417e-05 |
| 515 | RECEPTOR CLUSTERING | 5 | 41 | 4.889e-06 | 4.417e-05 |
| 516 | NEGATIVE REGULATION OF LOCOMOTION | 10 | 263 | 4.935e-06 | 4.45e-05 |
| 517 | REGULATION OF CELL GROWTH | 12 | 391 | 4.944e-06 | 4.45e-05 |
| 518 | COLUMNAR CUBOIDAL EPITHELIAL CELL DIFFERENTIATION | 7 | 111 | 5.312e-06 | 4.771e-05 |
| 519 | NEGATIVE REGULATION OF ESTABLISHMENT OF PROTEIN LOCALIZATION | 9 | 209 | 5.504e-06 | 4.934e-05 |
| 520 | POSTTRANSCRIPTIONAL GENE SILENCING | 5 | 42 | 5.523e-06 | 4.942e-05 |
| 521 | POSITIVE REGULATION OF PROTEIN BINDING | 6 | 73 | 5.551e-06 | 4.957e-05 |
| 522 | TONGUE DEVELOPMENT | 4 | 20 | 5.92e-06 | 5.257e-05 |
| 523 | GMP METABOLIC PROCESS | 4 | 20 | 5.92e-06 | 5.257e-05 |
| 524 | NEGATIVE REGULATION OF CHONDROCYTE DIFFERENTIATION | 4 | 20 | 5.92e-06 | 5.257e-05 |
| 525 | NEGATIVE REGULATION OF STEM CELL DIFFERENTIATION | 5 | 43 | 6.219e-06 | 5.512e-05 |
| 526 | BODY MORPHOGENESIS | 5 | 44 | 6.983e-06 | 6.177e-05 |
| 527 | FEMALE SEX DIFFERENTIATION | 7 | 116 | 7.111e-06 | 6.279e-05 |
| 528 | DEVELOPMENT OF PRIMARY SEXUAL CHARACTERISTICS | 9 | 216 | 7.185e-06 | 6.332e-05 |
| 529 | POSITIVE REGULATION OF CELLULAR COMPONENT BIOGENESIS | 12 | 406 | 7.24e-06 | 6.368e-05 |
| 530 | APOPTOTIC PROCESS INVOLVED IN DEVELOPMENT | 4 | 21 | 7.279e-06 | 6.378e-05 |
| 531 | NEGATIVE REGULATION OF NEURAL PRECURSOR CELL PROLIFERATION | 4 | 21 | 7.279e-06 | 6.378e-05 |
| 532 | LUNG MORPHOGENESIS | 5 | 45 | 7.817e-06 | 6.812e-05 |
| 533 | THYMOCYTE AGGREGATION | 5 | 45 | 7.817e-06 | 6.812e-05 |
| 534 | T CELL DIFFERENTIATION IN THYMUS | 5 | 45 | 7.817e-06 | 6.812e-05 |
| 535 | REGULATION OF PEPTIDYL SERINE PHOSPHORYLATION | 7 | 118 | 7.96e-06 | 6.923e-05 |
| 536 | REGULATION OF EPITHELIAL CELL MIGRATION | 8 | 166 | 8.21e-06 | 7.114e-05 |
| 537 | CENTRAL NERVOUS SYSTEM NEURON DIFFERENTIATION | 8 | 166 | 8.21e-06 | 7.114e-05 |
| 538 | ENDOCARDIAL CUSHION MORPHOGENESIS | 4 | 22 | 8.854e-06 | 7.658e-05 |
| 539 | POSITIVE REGULATION OF HYDROLASE ACTIVITY | 18 | 905 | 9.443e-06 | 8.152e-05 |
| 540 | REGULATION OF CATABOLIC PROCESS | 16 | 731 | 9.603e-06 | 8.275e-05 |
| 541 | REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 7 | 122 | 9.912e-06 | 8.51e-05 |
| 542 | LOCALIZATION WITHIN MEMBRANE | 7 | 122 | 9.912e-06 | 8.51e-05 |
| 543 | POSITIVE REGULATION OF MULTICELLULAR ORGANISMAL METABOLIC PROCESS | 4 | 23 | 1.067e-05 | 9.125e-05 |
| 544 | POSITIVE REGULATION OF COLLAGEN METABOLIC PROCESS | 4 | 23 | 1.067e-05 | 9.125e-05 |
| 545 | REGULATION OF CELL SUBSTRATE ADHESION | 8 | 173 | 1.11e-05 | 9.464e-05 |
| 546 | TELENCEPHALON DEVELOPMENT | 9 | 228 | 1.11e-05 | 9.464e-05 |
| 547 | HAIR CYCLE | 6 | 83 | 1.171e-05 | 9.945e-05 |
| 548 | MOLTING CYCLE | 6 | 83 | 1.171e-05 | 9.945e-05 |
| 549 | POSITIVE REGULATION OF MUSCLE CELL DIFFERENTIATION | 6 | 84 | 1.255e-05 | 0.0001064 |
| 550 | PHOTOPERIODISM | 4 | 24 | 1.274e-05 | 0.0001073 |
| 551 | REGULATION OF LEUKOCYTE DIFFERENTIATION | 9 | 232 | 1.275e-05 | 0.0001073 |
| 552 | CARDIAC EPITHELIAL TO MESENCHYMAL TRANSITION | 4 | 24 | 1.274e-05 | 0.0001073 |
| 553 | POSITIVE REGULATION OF NEURON PROJECTION DEVELOPMENT | 9 | 232 | 1.275e-05 | 0.0001073 |
| 554 | RESPONSE TO DRUG | 12 | 431 | 1.318e-05 | 0.0001106 |
| 555 | REGULATION OF VASCULATURE DEVELOPMENT | 9 | 233 | 1.32e-05 | 0.0001106 |
| 556 | FACE DEVELOPMENT | 5 | 50 | 1.323e-05 | 0.0001107 |
| 557 | CYTOSKELETON ORGANIZATION | 17 | 838 | 1.33e-05 | 0.0001111 |
| 558 | REGULATION OF REPRODUCTIVE PROCESS | 7 | 129 | 1.428e-05 | 0.0001191 |
| 559 | POSITIVE REGULATION OF DNA REPLICATION | 6 | 86 | 1.437e-05 | 0.0001196 |
| 560 | POSITIVE REGULATION OF FAT CELL DIFFERENTIATION | 5 | 51 | 1.46e-05 | 0.0001213 |
| 561 | REGULATION OF CHEMOTAXIS | 8 | 180 | 1.482e-05 | 0.0001229 |
| 562 | FOREBRAIN REGIONALIZATION | 4 | 25 | 1.51e-05 | 0.0001249 |
| 563 | MUSCLE CELL DIFFERENTIATION | 9 | 237 | 1.511e-05 | 0.0001249 |
| 564 | POSITIVE REGULATION OF GROWTH | 9 | 238 | 1.562e-05 | 0.0001289 |
| 565 | PROTEIN STABILIZATION | 7 | 131 | 1.579e-05 | 0.00013 |
| 566 | POSITIVE REGULATION OF PEPTIDYL SERINE PHOSPHORYLATION | 6 | 88 | 1.64e-05 | 0.0001349 |
| 567 | POSITIVE REGULATION OF CELL DIVISION | 7 | 132 | 1.659e-05 | 0.0001359 |
| 568 | MAINTENANCE OF CELL NUMBER | 7 | 132 | 1.659e-05 | 0.0001359 |
| 569 | DEVELOPMENTAL PROGRAMMED CELL DEATH | 4 | 26 | 1.776e-05 | 0.0001447 |
| 570 | ENTRAINMENT OF CIRCADIAN CLOCK | 4 | 26 | 1.776e-05 | 0.0001447 |
| 571 | REGULATION OF CELL FATE COMMITMENT | 4 | 26 | 1.776e-05 | 0.0001447 |
| 572 | REGULATION OF MICROTUBULE BASED PROCESS | 9 | 243 | 1.842e-05 | 0.0001499 |
| 573 | ENDOTHELIUM DEVELOPMENT | 6 | 90 | 1.866e-05 | 0.000151 |
| 574 | REGULATION OF CELL MATRIX ADHESION | 6 | 90 | 1.866e-05 | 0.000151 |
| 575 | REGULATION OF GLIOGENESIS | 6 | 90 | 1.866e-05 | 0.000151 |
| 576 | PROTEIN LOCALIZATION TO MEMBRANE | 11 | 376 | 1.95e-05 | 0.0001575 |
| 577 | HINDBRAIN DEVELOPMENT | 7 | 137 | 2.111e-05 | 0.0001702 |
| 578 | RESPONSE TO OXYGEN LEVELS | 10 | 311 | 2.122e-05 | 0.0001708 |
| 579 | CELLULAR RESPONSE TO STRESS | 24 | 1565 | 2.206e-05 | 0.0001772 |
| 580 | NEGATIVE REGULATION OF CELL CELL ADHESION | 7 | 138 | 2.213e-05 | 0.0001775 |
| 581 | POSITIVE REGULATION OF MUSCLE TISSUE DEVELOPMENT | 5 | 56 | 2.317e-05 | 0.0001852 |
| 582 | OUTFLOW TRACT MORPHOGENESIS | 5 | 56 | 2.317e-05 | 0.0001852 |
| 583 | GASTRULATION WITH MOUTH FORMING SECOND | 4 | 28 | 2.409e-05 | 0.000191 |
| 584 | NEGATIVE REGULATION OF TRANSPORT | 12 | 458 | 2.401e-05 | 0.000191 |
| 585 | REGULATION OF NEUROBLAST PROLIFERATION | 4 | 28 | 2.409e-05 | 0.000191 |
| 586 | MORPHOGENESIS OF A POLARIZED EPITHELIUM | 4 | 28 | 2.409e-05 | 0.000191 |
| 587 | METANEPHROS MORPHOGENESIS | 4 | 28 | 2.409e-05 | 0.000191 |
| 588 | CARDIAC MUSCLE TISSUE DEVELOPMENT | 7 | 140 | 2.428e-05 | 0.0001921 |
| 589 | REGULATION OF CARDIAC MUSCLE CELL PROLIFERATION | 4 | 29 | 2.782e-05 | 0.0002186 |
| 590 | POSITIVE REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 4 | 29 | 2.782e-05 | 0.0002186 |
| 591 | REGULATION OF OLIGODENDROCYTE DIFFERENTIATION | 4 | 29 | 2.782e-05 | 0.0002186 |
| 592 | NEUROBLAST PROLIFERATION | 4 | 29 | 2.782e-05 | 0.0002186 |
| 593 | REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 6 | 97 | 2.861e-05 | 0.0002245 |
| 594 | POSITIVE REGULATION OF CATABOLIC PROCESS | 11 | 395 | 3.065e-05 | 0.0002401 |
| 595 | REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 6 | 100 | 3.4e-05 | 0.0002659 |
| 596 | PEPTIDYL SERINE MODIFICATION | 7 | 148 | 3.47e-05 | 0.0002709 |
| 597 | REGULATION OF PRODUCTION OF MOLECULAR MEDIATOR OF IMMUNE RESPONSE | 6 | 101 | 3.597e-05 | 0.0002803 |
| 598 | REGULATION OF IMMUNE SYSTEM PROCESS | 22 | 1403 | 3.684e-05 | 0.0002867 |
| 599 | REGULATION OF CYTOKINE PRODUCTION | 13 | 563 | 4.044e-05 | 0.0003141 |
| 600 | SALIVARY GLAND DEVELOPMENT | 4 | 32 | 4.152e-05 | 0.0003215 |
| 601 | POSITIVE REGULATION OF BMP SIGNALING PATHWAY | 4 | 32 | 4.152e-05 | 0.0003215 |
| 602 | NEGATIVE REGULATION OF CARDIAC MUSCLE TISSUE GROWTH | 3 | 12 | 4.68e-05 | 0.0003597 |
| 603 | REGULATION OF T CELL APOPTOTIC PROCESS | 4 | 33 | 4.703e-05 | 0.0003597 |
| 604 | REGULATION OF THYMOCYTE APOPTOTIC PROCESS | 3 | 12 | 4.68e-05 | 0.0003597 |
| 605 | EMBRYONIC DIGESTIVE TRACT DEVELOPMENT | 4 | 33 | 4.703e-05 | 0.0003597 |
| 606 | HEART FORMATION | 3 | 12 | 4.68e-05 | 0.0003597 |
| 607 | CARDIAC VENTRICLE DEVELOPMENT | 6 | 106 | 4.723e-05 | 0.0003597 |
| 608 | FAT CELL DIFFERENTIATION | 6 | 106 | 4.723e-05 | 0.0003597 |
| 609 | EMBRYONIC EYE MORPHOGENESIS | 4 | 33 | 4.703e-05 | 0.0003597 |
| 610 | NEGATIVE REGULATION OF HEART GROWTH | 3 | 12 | 4.68e-05 | 0.0003597 |
| 611 | NEGATIVE REGULATION OF RESPONSE TO EXTERNAL STIMULUS | 9 | 274 | 4.717e-05 | 0.0003597 |
| 612 | ORGANELLE LOCALIZATION | 11 | 415 | 4.797e-05 | 0.0003647 |
| 613 | MUSCLE TISSUE DEVELOPMENT | 9 | 275 | 4.852e-05 | 0.0003674 |
| 614 | POSITIVE REGULATION OF DEVELOPMENTAL GROWTH | 7 | 156 | 4.856e-05 | 0.0003674 |
| 615 | PROTEIN LOCALIZATION TO NUCLEUS | 7 | 156 | 4.856e-05 | 0.0003674 |
| 616 | REGULATION OF CYSTEINE TYPE ENDOPEPTIDASE ACTIVITY | 8 | 213 | 4.942e-05 | 0.0003733 |
| 617 | NEURAL NUCLEUS DEVELOPMENT | 5 | 66 | 5.163e-05 | 0.0003894 |
| 618 | HORMONE MEDIATED SIGNALING PATHWAY | 7 | 158 | 5.265e-05 | 0.0003964 |
| 619 | LUNG EPITHELIUM DEVELOPMENT | 4 | 34 | 5.305e-05 | 0.0003988 |
| 620 | ACTIVATION OF PROTEIN KINASE ACTIVITY | 9 | 279 | 5.425e-05 | 0.0004071 |
| 621 | MICROTUBULE CYTOSKELETON ORGANIZATION | 10 | 348 | 5.492e-05 | 0.0004115 |
| 622 | POSITIVE REGULATION OF NEURON DEATH | 5 | 67 | 5.552e-05 | 0.0004147 |
| 623 | POSITIVE REGULATION OF ENDOTHELIAL CELL MIGRATION | 5 | 67 | 5.552e-05 | 0.0004147 |
| 624 | REGENERATION | 7 | 161 | 5.931e-05 | 0.0004423 |
| 625 | NEURONAL STEM CELL DIVISION | 3 | 13 | 6.057e-05 | 0.0004438 |
| 626 | NEGATIVE REGULATION OF OLIGODENDROCYTE DIFFERENTIATION | 3 | 13 | 6.057e-05 | 0.0004438 |
| 627 | TYPE B PANCREATIC CELL DEVELOPMENT | 3 | 13 | 6.057e-05 | 0.0004438 |
| 628 | POSITIVE REGULATION OF STEROID BIOSYNTHETIC PROCESS | 3 | 13 | 6.057e-05 | 0.0004438 |
| 629 | REGULATION OF CELL PROLIFERATION INVOLVED IN KIDNEY DEVELOPMENT | 3 | 13 | 6.057e-05 | 0.0004438 |
| 630 | NEUROBLAST DIVISION | 3 | 13 | 6.057e-05 | 0.0004438 |
| 631 | MESENCHYMAL CELL PROLIFERATION | 3 | 13 | 6.057e-05 | 0.0004438 |
| 632 | REGULATION OF CELL FATE SPECIFICATION | 3 | 13 | 6.057e-05 | 0.0004438 |
| 633 | POSITIVE REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 5 | 68 | 5.964e-05 | 0.0004438 |
| 634 | MESODERMAL CELL FATE COMMITMENT | 3 | 13 | 6.057e-05 | 0.0004438 |
| 635 | POSITIVE REGULATION OF CARDIAC MUSCLE CELL DIFFERENTIATION | 3 | 13 | 6.057e-05 | 0.0004438 |
| 636 | REGULATION OF GTPASE ACTIVITY | 14 | 673 | 6.249e-05 | 0.0004572 |
| 637 | REGULATION OF NUCLEAR DIVISION | 7 | 163 | 6.413e-05 | 0.0004684 |
| 638 | DEPHOSPHORYLATION | 9 | 286 | 6.564e-05 | 0.0004788 |
| 639 | ESTABLISHMENT OF LOCALIZATION IN CELL | 24 | 1676 | 6.624e-05 | 0.0004824 |
| 640 | RESPONSE TO NITROGEN COMPOUND | 16 | 859 | 6.773e-05 | 0.0004924 |
| 641 | MUSCLE ORGAN MORPHOGENESIS | 5 | 70 | 6.858e-05 | 0.0004978 |
| 642 | REGULATION OF ENDOTHELIAL CELL MIGRATION | 6 | 114 | 7.097e-05 | 0.0005144 |
| 643 | CELL FATE SPECIFICATION | 5 | 71 | 7.341e-05 | 0.0005313 |
| 644 | HINDLIMB MORPHOGENESIS | 4 | 37 | 7.449e-05 | 0.0005373 |
| 645 | NEGATIVE REGULATION OF EPITHELIAL CELL DIFFERENTIATION | 4 | 37 | 7.449e-05 | 0.0005373 |
| 646 | RENAL VESICLE DEVELOPMENT | 3 | 14 | 7.675e-05 | 0.0005494 |
| 647 | METANEPHRIC MESENCHYME DEVELOPMENT | 3 | 14 | 7.675e-05 | 0.0005494 |
| 648 | CONVERGENT EXTENSION | 3 | 14 | 7.675e-05 | 0.0005494 |
| 649 | CRANIOFACIAL SUTURE MORPHOGENESIS | 3 | 14 | 7.675e-05 | 0.0005494 |
| 650 | RESPONSE TO LAMINAR FLUID SHEAR STRESS | 3 | 14 | 7.675e-05 | 0.0005494 |
| 651 | ENERGY RESERVE METABOLIC PROCESS | 5 | 72 | 7.851e-05 | 0.0005612 |
| 652 | ANGIOGENESIS | 9 | 293 | 7.899e-05 | 0.0005637 |
| 653 | REGULATION OF GENE EXPRESSION EPIGENETIC | 8 | 229 | 8.213e-05 | 0.0005852 |
| 654 | REGULATION OF MULTICELLULAR ORGANISMAL METABOLIC PROCESS | 4 | 38 | 8.286e-05 | 0.0005895 |
| 655 | NEGATIVE REGULATION OF MAP KINASE ACTIVITY | 5 | 73 | 8.388e-05 | 0.0005931 |
| 656 | EMBRYONIC HEART TUBE DEVELOPMENT | 5 | 73 | 8.388e-05 | 0.0005931 |
| 657 | HIPPOCAMPUS DEVELOPMENT | 5 | 73 | 8.388e-05 | 0.0005931 |
| 658 | REGULATION OF PLASMA MEMBRANE ORGANIZATION | 5 | 73 | 8.388e-05 | 0.0005931 |
| 659 | REGULATION OF AXON GUIDANCE | 4 | 39 | 9.189e-05 | 0.0006479 |
| 660 | GLANDULAR EPITHELIAL CELL DIFFERENTIATION | 4 | 39 | 9.189e-05 | 0.0006479 |
| 661 | POSITIVE REGULATION OF ENDOTHELIAL CELL DIFFERENTIATION | 3 | 15 | 9.551e-05 | 0.0006653 |
| 662 | MESENCHYMAL TO EPITHELIAL TRANSITION | 3 | 15 | 9.551e-05 | 0.0006653 |
| 663 | REGULATION OF EPITHELIAL CELL DIFFERENTIATION INVOLVED IN KIDNEY DEVELOPMENT | 3 | 15 | 9.551e-05 | 0.0006653 |
| 664 | ENDOCARDIAL CUSHION FORMATION | 3 | 15 | 9.551e-05 | 0.0006653 |
| 665 | REGULATION OF MESODERM DEVELOPMENT | 3 | 15 | 9.551e-05 | 0.0006653 |
| 666 | REGULATION OF CELL PROLIFERATION INVOLVED IN HEART MORPHOGENESIS | 3 | 15 | 9.551e-05 | 0.0006653 |
| 667 | POSITIVE REGULATION OF T CELL APOPTOTIC PROCESS | 3 | 15 | 9.551e-05 | 0.0006653 |
| 668 | NEURAL CREST CELL DIFFERENTIATION | 5 | 75 | 9.545e-05 | 0.0006653 |
| 669 | POSITIVE REGULATION OF CELL MATRIX ADHESION | 4 | 40 | 0.0001016 | 0.0007068 |
| 670 | NEGATIVE REGULATION OF DEPHOSPHORYLATION | 5 | 77 | 0.0001082 | 0.0007515 |
| 671 | PROSTATE GLAND DEVELOPMENT | 4 | 41 | 0.0001121 | 0.0007772 |
| 672 | ESTABLISHMENT OR MAINTENANCE OF MONOPOLAR CELL POLARITY | 3 | 16 | 0.000117 | 0.0008091 |
| 673 | CELL SURFACE RECEPTOR SIGNALING PATHWAY INVOLVED IN HEART DEVELOPMENT | 3 | 16 | 0.000117 | 0.0008091 |
| 674 | NEGATIVE REGULATION OF PROTEIN BINDING | 5 | 79 | 0.0001222 | 0.0008439 |
| 675 | POSITIVE REGULATION OF NEURAL PRECURSOR CELL PROLIFERATION | 4 | 42 | 0.0001233 | 0.0008462 |
| 676 | EPITHELIAL CELL MORPHOGENESIS | 4 | 42 | 0.0001233 | 0.0008462 |
| 677 | REGULATION OF HEART GROWTH | 4 | 42 | 0.0001233 | 0.0008462 |
| 678 | GENITALIA DEVELOPMENT | 4 | 42 | 0.0001233 | 0.0008462 |
| 679 | REGULATION OF PROTEASOMAL PROTEIN CATABOLIC PROCESS | 7 | 181 | 0.0001236 | 0.000847 |
| 680 | NEGATIVE REGULATION OF VASCULATURE DEVELOPMENT | 5 | 80 | 0.0001298 | 0.0008879 |
| 681 | REGULATION OF HEMOPOIESIS | 9 | 314 | 0.0001335 | 0.0009121 |
| 682 | IMMUNE SYSTEM PROCESS | 26 | 1984 | 0.0001354 | 0.0009211 |
| 683 | MORPHOGENESIS OF AN EPITHELIAL SHEET | 4 | 43 | 0.0001353 | 0.0009211 |
| 684 | REGULATION OF POLYSACCHARIDE METABOLIC PROCESS | 4 | 43 | 0.0001353 | 0.0009211 |
| 685 | REGULATION OF PROTEIN SECRETION | 10 | 389 | 0.0001376 | 0.0009335 |
| 686 | REGULATION OF FIBROBLAST PROLIFERATION | 5 | 81 | 0.0001376 | 0.0009335 |
| 687 | POSITIVE REGULATION OF CELL CYCLE PROCESS | 8 | 247 | 0.0001386 | 0.0009389 |
| 688 | NEGATIVE REGULATION OF KIDNEY DEVELOPMENT | 3 | 17 | 0.0001415 | 0.0009485 |
| 689 | POSITIVE REGULATION OF EXTRACELLULAR MATRIX ORGANIZATION | 3 | 17 | 0.0001415 | 0.0009485 |
| 690 | DORSAL VENTRAL NEURAL TUBE PATTERNING | 3 | 17 | 0.0001415 | 0.0009485 |
| 691 | REGULATION OF STEM CELL POPULATION MAINTENANCE | 3 | 17 | 0.0001415 | 0.0009485 |
| 692 | MAMMARY GLAND ALVEOLUS DEVELOPMENT | 3 | 17 | 0.0001415 | 0.0009485 |
| 693 | MAMMARY GLAND LOBULE DEVELOPMENT | 3 | 17 | 0.0001415 | 0.0009485 |
| 694 | NEGATIVE REGULATION OF CARDIAC MUSCLE TISSUE DEVELOPMENT | 3 | 17 | 0.0001415 | 0.0009485 |
| 695 | LABYRINTHINE LAYER DEVELOPMENT | 4 | 44 | 0.0001481 | 0.0009918 |
| 696 | POSITIVE REGULATION OF LEUKOCYTE DIFFERENTIATION | 6 | 131 | 0.0001529 | 0.001021 |
| 697 | NEGATIVE REGULATION OF BINDING | 6 | 131 | 0.0001529 | 0.001021 |
| 698 | EMBRYONIC PLACENTA DEVELOPMENT | 5 | 83 | 0.0001545 | 0.00103 |
| 699 | REGULATION OF CELL PROJECTION ORGANIZATION | 12 | 558 | 0.0001581 | 0.001052 |
| 700 | REGULATION OF NEURON DEATH | 8 | 252 | 0.000159 | 0.001057 |
| 701 | REGULATION OF LYMPHOCYTE DIFFERENTIATION | 6 | 132 | 0.0001594 | 0.001058 |
| 702 | REGULATION OF RESPONSE TO EXTERNAL STIMULUS | 16 | 926 | 0.0001616 | 0.001068 |
| 703 | SUBSTANTIA NIGRA DEVELOPMENT | 4 | 45 | 0.0001618 | 0.001068 |
| 704 | EXOCRINE SYSTEM DEVELOPMENT | 4 | 45 | 0.0001618 | 0.001068 |
| 705 | REGULATION OF ALCOHOL BIOSYNTHETIC PROCESS | 4 | 45 | 0.0001618 | 0.001068 |
| 706 | NEGATIVE REGULATION OF HYDROLASE ACTIVITY | 10 | 397 | 0.0001623 | 0.00107 |
| 707 | GLANDULAR EPITHELIAL CELL DEVELOPMENT | 3 | 18 | 0.000169 | 0.001106 |
| 708 | PERICARDIUM DEVELOPMENT | 3 | 18 | 0.000169 | 0.001106 |
| 709 | OSTEOBLAST DEVELOPMENT | 3 | 18 | 0.000169 | 0.001106 |
| 710 | POST ANAL TAIL MORPHOGENESIS | 3 | 18 | 0.000169 | 0.001106 |
| 711 | NOTOCHORD DEVELOPMENT | 3 | 18 | 0.000169 | 0.001106 |
| 712 | RESPONSE TO INORGANIC SUBSTANCE | 11 | 479 | 0.0001706 | 0.001115 |
| 713 | GUANOSINE CONTAINING COMPOUND METABOLIC PROCESS | 4 | 46 | 0.0001764 | 0.001151 |
| 714 | POSITIVE REGULATION OF CELLULAR PROTEIN CATABOLIC PROCESS | 7 | 192 | 0.000178 | 0.001158 |
| 715 | REGULATION OF NEURON APOPTOTIC PROCESS | 7 | 192 | 0.000178 | 0.001158 |
| 716 | CELL GROWTH | 6 | 135 | 0.0001802 | 0.001171 |
| 717 | NEGATIVE REGULATION OF SEQUENCE SPECIFIC DNA BINDING TRANSCRIPTION FACTOR ACTIVITY | 6 | 136 | 0.0001876 | 0.001216 |
| 718 | GLIAL CELL DIFFERENTIATION | 6 | 136 | 0.0001876 | 0.001216 |
| 719 | POSITIVE REGULATION OF NEURON APOPTOTIC PROCESS | 4 | 47 | 0.0001919 | 0.001242 |
| 720 | TISSUE REMODELING | 5 | 87 | 0.0001928 | 0.001246 |
| 721 | REGULATION OF CARDIAC MUSCLE CELL DIFFERENTIATION | 3 | 19 | 0.0001998 | 0.001288 |
| 722 | ENTEROENDOCRINE CELL DIFFERENTIATION | 3 | 19 | 0.0001998 | 0.001288 |
| 723 | REGULATION OF NEURON PROJECTION DEVELOPMENT | 10 | 408 | 0.0002023 | 0.001302 |
| 724 | PLACENTA DEVELOPMENT | 6 | 138 | 0.0002031 | 0.001305 |
| 725 | COLUMNAR CUBOIDAL EPITHELIAL CELL DEVELOPMENT | 4 | 48 | 0.0002084 | 0.001336 |
| 726 | POSITIVE REGULATION OF ACTIN FILAMENT BUNDLE ASSEMBLY | 4 | 48 | 0.0002084 | 0.001336 |
| 727 | CELLULAR RESPONSE TO ABIOTIC STIMULUS | 8 | 263 | 0.0002128 | 0.001362 |
| 728 | CELLULAR RESPONSE TO EXTERNAL STIMULUS | 8 | 264 | 0.0002183 | 0.001393 |
| 729 | AGING | 8 | 264 | 0.0002183 | 0.001393 |
| 730 | RESPONSE TO RADIATION | 10 | 413 | 0.0002231 | 0.001422 |
| 731 | REGULATION OF STEROID BIOSYNTHETIC PROCESS | 4 | 49 | 0.0002258 | 0.001438 |
| 732 | NEGATIVE REGULATION OF APOPTOTIC SIGNALING PATHWAY | 7 | 200 | 0.0002286 | 0.001453 |
| 733 | POSITIVE REGULATION OF LYMPHOCYTE APOPTOTIC PROCESS | 3 | 20 | 0.000234 | 0.001486 |
| 734 | MULTICELLULAR ORGANISM REPRODUCTION | 14 | 768 | 0.0002482 | 0.001573 |
| 735 | SECRETION | 12 | 588 | 0.0002557 | 0.001619 |
| 736 | POSITIVE REGULATION OF IMMUNE SYSTEM PROCESS | 15 | 867 | 0.000259 | 0.001637 |
| 737 | PROTEASOMAL PROTEIN CATABOLIC PROCESS | 8 | 271 | 0.0002605 | 0.001645 |
| 738 | REGULATION OF DNA BINDING | 5 | 93 | 0.0002634 | 0.001661 |
| 739 | HOMOTYPIC CELL CELL ADHESION | 4 | 51 | 0.0002639 | 0.001662 |
| 740 | REGULATION OF GENE SILENCING BY RNA | 3 | 21 | 0.0002718 | 0.0017 |
| 741 | CHONDROCYTE DEVELOPMENT | 3 | 21 | 0.0002718 | 0.0017 |
| 742 | ECTODERM DEVELOPMENT | 3 | 21 | 0.0002718 | 0.0017 |
| 743 | POSITIVE REGULATION OF EXCITATORY POSTSYNAPTIC POTENTIAL | 3 | 21 | 0.0002718 | 0.0017 |
| 744 | REGULATION OF POSTTRANSCRIPTIONAL GENE SILENCING | 3 | 21 | 0.0002718 | 0.0017 |
| 745 | REGULATION OF CELLULAR PROTEIN CATABOLIC PROCESS | 8 | 274 | 0.0002806 | 0.001752 |
| 746 | REGULATION OF TRANSCRIPTION FACTOR IMPORT INTO NUCLEUS | 5 | 95 | 0.0002908 | 0.001814 |
| 747 | REGULATION OF PROTEASOMAL UBIQUITIN DEPENDENT PROTEIN CATABOLIC PROCESS | 6 | 148 | 0.0002963 | 0.001843 |
| 748 | POSITIVE REGULATION OF CELL GROWTH | 6 | 148 | 0.0002963 | 0.001843 |
| 749 | MYELOID LEUKOCYTE DIFFERENTIATION | 5 | 96 | 0.0003053 | 0.001897 |
| 750 | POSITIVE REGULATION OF FIBROBLAST PROLIFERATION | 4 | 53 | 0.0003063 | 0.001898 |
| 751 | NEGATIVE REGULATION OF CELL SUBSTRATE ADHESION | 4 | 53 | 0.0003063 | 0.001898 |
| 752 | NEGATIVE REGULATION OF CYTOKINE PRODUCTION INVOLVED IN IMMUNE RESPONSE | 3 | 22 | 0.0003133 | 0.001931 |
| 753 | DSRNA FRAGMENTATION | 3 | 22 | 0.0003133 | 0.001931 |
| 754 | EMBRYONIC PLACENTA MORPHOGENESIS | 3 | 22 | 0.0003133 | 0.001931 |
| 755 | PRODUCTION OF SMALL RNA INVOLVED IN GENE SILENCING BY RNA | 3 | 22 | 0.0003133 | 0.001931 |
| 756 | POSITIVE REGULATION OF PROTEIN SECRETION | 7 | 211 | 0.0003165 | 0.001948 |
| 757 | NEGATIVE REGULATION OF REPRODUCTIVE PROCESS | 4 | 54 | 0.0003293 | 0.002021 |
| 758 | REGULATION OF LYMPHOCYTE APOPTOTIC PROCESS | 4 | 54 | 0.0003293 | 0.002021 |
| 759 | POSITIVE REGULATION OF PROTEASOMAL PROTEIN CATABOLIC PROCESS | 5 | 98 | 0.0003359 | 0.002054 |
| 760 | NEGATIVE REGULATION OF EXTRINSIC APOPTOTIC SIGNALING PATHWAY | 5 | 98 | 0.0003359 | 0.002054 |
| 761 | REGULATION OF ENDOTHELIAL CELL PROLIFERATION | 5 | 98 | 0.0003359 | 0.002054 |
| 762 | REGULATION OF MUSCLE CELL DIFFERENTIATION | 6 | 152 | 0.0003419 | 0.002088 |
| 763 | PALLIUM DEVELOPMENT | 6 | 153 | 0.0003541 | 0.002156 |
| 764 | TRANSCRIPTION INITIATION FROM RNA POLYMERASE II PROMOTER | 6 | 153 | 0.0003541 | 0.002156 |
| 765 | POSITIVE REGULATION OF ALCOHOL BIOSYNTHETIC PROCESS | 3 | 23 | 0.0003587 | 0.002176 |
| 766 | SCF DEPENDENT PROTEASOMAL UBIQUITIN DEPENDENT PROTEIN CATABOLIC PROCESS | 3 | 23 | 0.0003587 | 0.002176 |
| 767 | POSITIVE REGULATION OF STEROID METABOLIC PROCESS | 3 | 23 | 0.0003587 | 0.002176 |
| 768 | LIMBIC SYSTEM DEVELOPMENT | 5 | 100 | 0.0003688 | 0.002235 |
| 769 | KERATINOCYTE DIFFERENTIATION | 5 | 101 | 0.0003862 | 0.002334 |
| 770 | REGULATION OF EXTENT OF CELL GROWTH | 5 | 101 | 0.0003862 | 0.002334 |
| 771 | CELLULAR RESPONSE TO INORGANIC SUBSTANCE | 6 | 156 | 0.0003927 | 0.00237 |
| 772 | EMBRYONIC CAMERA TYPE EYE MORPHOGENESIS | 3 | 24 | 0.0004082 | 0.00245 |
| 773 | RESPONSE TO STEROL | 3 | 24 | 0.0004082 | 0.00245 |
| 774 | REGULATION OF ODONTOGENESIS | 3 | 24 | 0.0004082 | 0.00245 |
| 775 | ENDOTHELIAL CELL PROLIFERATION | 3 | 24 | 0.0004082 | 0.00245 |
| 776 | POSTTRANSCRIPTIONAL REGULATION OF GENE EXPRESSION | 10 | 448 | 0.0004247 | 0.002547 |
| 777 | CELLULAR GLUCAN METABOLIC PROCESS | 4 | 58 | 0.0004335 | 0.002593 |
| 778 | GLUCAN METABOLIC PROCESS | 4 | 58 | 0.0004335 | 0.002593 |
| 779 | ACTIN FILAMENT BASED PROCESS | 10 | 450 | 0.0004398 | 0.002627 |
| 780 | REGULATION OF EPITHELIAL CELL APOPTOTIC PROCESS | 4 | 59 | 0.0004629 | 0.00274 |
| 781 | EPITHELIAL TUBE BRANCHING INVOLVED IN LUNG MORPHOGENESIS | 3 | 25 | 0.0004618 | 0.00274 |
| 782 | LUNG CELL DIFFERENTIATION | 3 | 25 | 0.0004618 | 0.00274 |
| 783 | REGULATION OF TRANSFORMING GROWTH FACTOR BETA PRODUCTION | 3 | 25 | 0.0004618 | 0.00274 |
| 784 | REGULATION OF FIBROBLAST GROWTH FACTOR RECEPTOR SIGNALING PATHWAY | 3 | 25 | 0.0004618 | 0.00274 |
| 785 | EPITHELIAL CELL APOPTOTIC PROCESS | 3 | 25 | 0.0004618 | 0.00274 |
| 786 | REGULATION OF GLIAL CELL DIFFERENTIATION | 4 | 59 | 0.0004629 | 0.00274 |
| 787 | NEGATIVE REGULATION OF CELL DIVISION | 4 | 60 | 0.0004936 | 0.002915 |
| 788 | POSITIVE REGULATION OF SMOOTH MUSCLE CELL PROLIFERATION | 4 | 60 | 0.0004936 | 0.002915 |
| 789 | POSITIVE REGULATION OF HEMOPOIESIS | 6 | 163 | 0.0004958 | 0.002924 |
| 790 | REGULATION OF T CELL DIFFERENTIATION | 5 | 107 | 0.0005035 | 0.002965 |
| 791 | NEGATIVE REGULATION OF GLIAL CELL DIFFERENTIATION | 3 | 26 | 0.0005197 | 0.003045 |
| 792 | REGULATION OF MESONEPHROS DEVELOPMENT | 3 | 26 | 0.0005197 | 0.003045 |
| 793 | MESODERMAL CELL DIFFERENTIATION | 3 | 26 | 0.0005197 | 0.003045 |
| 794 | REGULATION OF HORMONE METABOLIC PROCESS | 3 | 26 | 0.0005197 | 0.003045 |
| 795 | OVARIAN FOLLICLE DEVELOPMENT | 4 | 61 | 0.0005258 | 0.003074 |
| 796 | REGULATION OF CELL CYCLE ARREST | 5 | 108 | 0.0005254 | 0.003074 |
| 797 | POSITIVE REGULATION OF NUCLEAR DIVISION | 4 | 62 | 0.0005594 | 0.003262 |
| 798 | EMBRYONIC HEART TUBE MORPHOGENESIS | 4 | 62 | 0.0005594 | 0.003262 |
| 799 | CELLULAR RESPONSE TO HORMONE STIMULUS | 11 | 552 | 0.000568 | 0.003308 |
| 800 | REGULATION OF ENDOTHELIAL CELL DIFFERENTIATION | 3 | 27 | 0.000582 | 0.003381 |
| 801 | NEGATIVE REGULATION OF AXON GUIDANCE | 3 | 27 | 0.000582 | 0.003381 |
| 802 | NEGATIVE REGULATION OF PROTEIN COMPLEX DISASSEMBLY | 6 | 170 | 0.0006188 | 0.00359 |
| 803 | POSITIVE REGULATION OF PRODUCTION OF MOLECULAR MEDIATOR OF IMMUNE RESPONSE | 4 | 64 | 0.0006312 | 0.003653 |
| 804 | RIBONUCLEOSIDE DIPHOSPHATE METABOLIC PROCESS | 4 | 64 | 0.0006312 | 0.003653 |
| 805 | POSITIVE REGULATION OF LEUKOCYTE APOPTOTIC PROCESS | 3 | 28 | 0.000649 | 0.003751 |
| 806 | REGULATION OF CELL SIZE | 6 | 172 | 0.000658 | 0.003799 |
| 807 | REGULATION OF PEPTIDASE ACTIVITY | 9 | 392 | 0.0006775 | 0.003906 |
| 808 | CELLULAR RESPONSE TO ALCOHOL | 5 | 115 | 0.0006993 | 0.004027 |
| 809 | SOMATIC STEM CELL POPULATION MAINTENANCE | 4 | 66 | 0.0007093 | 0.004075 |
| 810 | FOREBRAIN GENERATION OF NEURONS | 4 | 66 | 0.0007093 | 0.004075 |
| 811 | EPIDERMIS MORPHOGENESIS | 3 | 29 | 0.0007207 | 0.004119 |
| 812 | NEGATIVE REGULATION OF PRODUCTION OF MOLECULAR MEDIATOR OF IMMUNE RESPONSE | 3 | 29 | 0.0007207 | 0.004119 |
| 813 | REGULATION OF EXTRACELLULAR MATRIX ORGANIZATION | 3 | 29 | 0.0007207 | 0.004119 |
| 814 | EMBRYONIC HINDLIMB MORPHOGENESIS | 3 | 29 | 0.0007207 | 0.004119 |
| 815 | RESPONSE TO CORTICOSTEROID | 6 | 176 | 0.0007421 | 0.004237 |
| 816 | SPECIFICATION OF SYMMETRY | 5 | 117 | 0.0007561 | 0.004311 |
| 817 | REGULATION OF CELL ACTIVATION | 10 | 484 | 0.0007718 | 0.004395 |
| 818 | REGULATION OF MICROTUBULE POLYMERIZATION OR DEPOLYMERIZATION | 6 | 178 | 0.0007871 | 0.004477 |
| 819 | PROTEIN LOCALIZATION TO CYTOSKELETON | 3 | 30 | 0.0007972 | 0.004496 |
| 820 | SECRETION BY CELL | 10 | 486 | 0.0007964 | 0.004496 |
| 821 | SMOOTH MUSCLE CELL DIFFERENTIATION | 3 | 30 | 0.0007972 | 0.004496 |
| 822 | POSITIVE REGULATION OF CYTOKINE PRODUCTION INVOLVED IN IMMUNE RESPONSE | 3 | 30 | 0.0007972 | 0.004496 |
| 823 | REGULATION OF CELL JUNCTION ASSEMBLY | 4 | 68 | 0.0007941 | 0.004496 |
| 824 | ORGAN GROWTH | 4 | 68 | 0.0007941 | 0.004496 |
| 825 | MODULATION OF EXCITATORY POSTSYNAPTIC POTENTIAL | 3 | 30 | 0.0007972 | 0.004496 |
| 826 | POSITIVE REGULATION OF AXONOGENESIS | 4 | 69 | 0.000839 | 0.004726 |
| 827 | POSITIVE REGULATION OF CHEMOTAXIS | 5 | 120 | 0.0008476 | 0.004769 |
| 828 | PROTEIN TARGETING | 9 | 406 | 0.0008683 | 0.004879 |
| 829 | RESPONSE TO KETONE | 6 | 182 | 0.0008835 | 0.004959 |
| 830 | ADHERENS JUNCTION ORGANIZATION | 4 | 71 | 0.0009342 | 0.005237 |
| 831 | NEGATIVE REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 3 | 32 | 0.0009653 | 0.005398 |
| 832 | PATTERNING OF BLOOD VESSELS | 3 | 32 | 0.0009653 | 0.005398 |
| 833 | ENDOTHELIAL CELL DIFFERENTIATION | 4 | 72 | 0.0009845 | 0.005499 |
| 834 | RESPONSE TO METAL ION | 8 | 333 | 0.00101 | 0.005637 |
| 835 | STEROID HORMONE MEDIATED SIGNALING PATHWAY | 5 | 125 | 0.001018 | 0.005674 |
| 836 | RESPONSE TO VITAMIN D | 3 | 33 | 0.001057 | 0.005884 |
| 837 | CELLULAR RESPONSE TO NITROGEN COMPOUND | 10 | 505 | 0.001065 | 0.005919 |
| 838 | MYELOID CELL DIFFERENTIATION | 6 | 189 | 0.001074 | 0.005963 |
| 839 | REGULATION OF CELLULAR COMPONENT SIZE | 8 | 337 | 0.001091 | 0.006042 |
| 840 | REGULATION OF STEROID METABOLIC PROCESS | 4 | 74 | 0.001091 | 0.006042 |
| 841 | RESPONSE TO FLUID SHEAR STRESS | 3 | 34 | 0.001154 | 0.006371 |
| 842 | NEGATIVE REGULATION OF MITOTIC NUCLEAR DIVISION | 3 | 34 | 0.001154 | 0.006371 |
| 843 | NEURAL TUBE PATTERNING | 3 | 34 | 0.001154 | 0.006371 |
| 844 | REGULATION OF MEMBRANE POTENTIAL | 8 | 343 | 0.001221 | 0.006729 |
| 845 | CELL CYCLE CHECKPOINT | 6 | 194 | 0.001228 | 0.006762 |
| 846 | NEGATIVE REGULATION OF CELLULAR AMIDE METABOLIC PROCESS | 5 | 131 | 0.001255 | 0.006872 |
| 847 | BONE REMODELING | 3 | 35 | 0.001257 | 0.006872 |
| 848 | REGULATION OF GLYCOGEN METABOLIC PROCESS | 3 | 35 | 0.001257 | 0.006872 |
| 849 | REGULATION OF GASTRULATION | 3 | 35 | 0.001257 | 0.006872 |
| 850 | EMBRYONIC CAMERA TYPE EYE DEVELOPMENT | 3 | 35 | 0.001257 | 0.006872 |
| 851 | RESPONSE TO IRON ION | 3 | 35 | 0.001257 | 0.006872 |
| 852 | REGULATION OF ACTIN FILAMENT BUNDLE ASSEMBLY | 4 | 77 | 0.001265 | 0.006909 |
| 853 | CELLULAR PROTEIN COMPLEX ASSEMBLY | 8 | 346 | 0.00129 | 0.007036 |
| 854 | MICROTUBULE BASED PROCESS | 10 | 522 | 0.001364 | 0.007394 |
| 855 | POSITIVE REGULATION OF PROTEIN ACETYLATION | 3 | 36 | 0.001365 | 0.007394 |
| 856 | REGULATION OF TRANSCRIPTION REGULATORY REGION DNA BINDING | 3 | 36 | 0.001365 | 0.007394 |
| 857 | POSITIVE REGULATION OF CYCLIN DEPENDENT PROTEIN KINASE ACTIVITY | 3 | 36 | 0.001365 | 0.007394 |
| 858 | CORTICAL CYTOSKELETON ORGANIZATION | 3 | 36 | 0.001365 | 0.007394 |
| 859 | POSITIVE CHEMOTAXIS | 3 | 36 | 0.001365 | 0.007394 |
| 860 | DENDRITE DEVELOPMENT | 4 | 79 | 0.001392 | 0.00752 |
| 861 | REGULATION OF LEUKOCYTE APOPTOTIC PROCESS | 4 | 79 | 0.001392 | 0.00752 |
| 862 | POLYSACCHARIDE METABOLIC PROCESS | 4 | 80 | 0.001458 | 0.007852 |
| 863 | POSITIVE REGULATION OF LYMPHOCYTE DIFFERENTIATION | 4 | 80 | 0.001458 | 0.007852 |
| 864 | CELLULAR RESPONSE TO MECHANICAL STIMULUS | 4 | 80 | 0.001458 | 0.007852 |
| 865 | NEGATIVE REGULATION OF GLIOGENESIS | 3 | 37 | 0.001479 | 0.007928 |
| 866 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO CELL PERIPHERY | 3 | 37 | 0.001479 | 0.007928 |
| 867 | GLIAL CELL MIGRATION | 3 | 37 | 0.001479 | 0.007928 |
| 868 | POSITIVE REGULATION OF PROTEIN LOCALIZATION TO PLASMA MEMBRANE | 3 | 37 | 0.001479 | 0.007928 |
| 869 | DNA TEMPLATED TRANSCRIPTION INITIATION | 6 | 202 | 0.001509 | 0.008082 |
| 870 | RESPONSE TO EXTRACELLULAR STIMULUS | 9 | 441 | 0.001542 | 0.008246 |
| 871 | MUSCLE ORGAN DEVELOPMENT | 7 | 277 | 0.001568 | 0.008375 |
| 872 | NEGATIVE REGULATION OF AXON EXTENSION | 3 | 38 | 0.001599 | 0.008509 |
| 873 | CELLULAR RESPONSE TO DSRNA | 3 | 38 | 0.001599 | 0.008509 |
| 874 | REGULATION OF CHROMOSOME ORGANIZATION | 7 | 278 | 0.0016 | 0.008509 |
| 875 | NEGATIVE REGULATION OF LEUKOCYTE DIFFERENTIATION | 4 | 82 | 0.001597 | 0.008509 |
| 876 | MITOTIC CELL CYCLE CHECKPOINT | 5 | 139 | 0.001632 | 0.008657 |
| 877 | GENE SILENCING BY RNA | 5 | 139 | 0.001632 | 0.008657 |
| 878 | RESPONSE TO LIGHT STIMULUS | 7 | 280 | 0.001667 | 0.008833 |
| 879 | ORGAN REGENERATION | 4 | 83 | 0.00167 | 0.008842 |
| 880 | REGULATION OF HOMEOSTATIC PROCESS | 9 | 447 | 0.001691 | 0.008941 |
| 881 | NEGATIVE REGULATION OF TRANSCRIPTION FACTOR IMPORT INTO NUCLEUS | 3 | 39 | 0.001724 | 0.009075 |
| 882 | NEGATIVE REGULATION OF PEPTIDYL TYROSINE PHOSPHORYLATION | 3 | 39 | 0.001724 | 0.009075 |
| 883 | REGULATION OF CELL ADHESION MEDIATED BY INTEGRIN | 3 | 39 | 0.001724 | 0.009075 |
| 884 | PLATELET AGGREGATION | 3 | 39 | 0.001724 | 0.009075 |
| 885 | NUCLEOSIDE DIPHOSPHATE METABOLIC PROCESS | 4 | 84 | 0.001746 | 0.009178 |
| 886 | PLATELET ACTIVATION | 5 | 142 | 0.001792 | 0.009413 |
| 887 | POSITIVE REGULATION OF CHROMATIN MODIFICATION | 4 | 85 | 0.001823 | 0.009565 |
| 888 | RESPONSE TO MECHANICAL STIMULUS | 6 | 210 | 0.001837 | 0.009627 |
| 889 | REGULATION OF MEIOTIC CELL CYCLE | 3 | 40 | 0.001856 | 0.009702 |
| 890 | ENDOCRINE PANCREAS DEVELOPMENT | 3 | 40 | 0.001856 | 0.009702 |
| 891 | NEGATIVE REGULATION OF CELL PROJECTION ORGANIZATION | 5 | 144 | 0.001906 | 0.009952 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | BETA CATENIN BINDING | 18 | 84 | 2.528e-23 | 2.349e-20 |
| 2 | FRIZZLED BINDING | 14 | 36 | 1.553e-22 | 7.211e-20 |
| 3 | TRANSFORMING GROWTH FACTOR BETA RECEPTOR BINDING | 14 | 50 | 3.573e-20 | 1.106e-17 |
| 4 | ENZYME BINDING | 47 | 1737 | 1.736e-19 | 4.032e-17 |
| 5 | PROTEIN DOMAIN SPECIFIC BINDING | 30 | 624 | 6.835e-19 | 1.27e-16 |
| 6 | RECEPTOR BINDING | 42 | 1476 | 4.841e-18 | 7.496e-16 |
| 7 | KINASE BINDING | 28 | 606 | 3.388e-17 | 3.934e-15 |
| 8 | WNT ACTIVATED RECEPTOR ACTIVITY | 10 | 22 | 2.977e-17 | 3.934e-15 |
| 9 | WNT PROTEIN BINDING | 10 | 31 | 1.95e-15 | 2.013e-13 |
| 10 | I SMAD BINDING | 7 | 11 | 8.539e-14 | 7.933e-12 |
| 11 | SMAD BINDING | 11 | 72 | 6.071e-13 | 5.127e-11 |
| 12 | GAMMA CATENIN BINDING | 6 | 12 | 4.081e-11 | 3.159e-09 |
| 13 | G PROTEIN COUPLED RECEPTOR BINDING | 15 | 259 | 5.772e-11 | 4.125e-09 |
| 14 | PDZ DOMAIN BINDING | 10 | 90 | 1.861e-10 | 1.235e-08 |
| 15 | GROWTH FACTOR ACTIVITY | 12 | 160 | 2.757e-10 | 1.708e-08 |
| 16 | UBIQUITIN LIKE PROTEIN LIGASE BINDING | 14 | 264 | 8.302e-10 | 4.782e-08 |
| 17 | CYTOKINE ACTIVITY | 13 | 219 | 8.75e-10 | 4.782e-08 |
| 18 | CYTOKINE RECEPTOR BINDING | 14 | 271 | 1.166e-09 | 6.02e-08 |
| 19 | PROTEIN PHOSPHATASE TYPE 2A REGULATOR ACTIVITY | 6 | 21 | 2.292e-09 | 1.12e-07 |
| 20 | IONOTROPIC GLUTAMATE RECEPTOR BINDING | 6 | 23 | 4.221e-09 | 1.961e-07 |
| 21 | PROTEIN DIMERIZATION ACTIVITY | 26 | 1149 | 5.029e-09 | 2.225e-07 |
| 22 | MOLECULAR FUNCTION REGULATOR | 28 | 1353 | 8.149e-09 | 3.43e-07 |
| 23 | RECEPTOR SIGNALING PROTEIN ACTIVITY | 11 | 172 | 8.491e-09 | 3.43e-07 |
| 24 | RECEPTOR SERINE THREONINE KINASE BINDING | 5 | 15 | 2.224e-08 | 8.609e-07 |
| 25 | RECEPTOR AGONIST ACTIVITY | 5 | 16 | 3.219e-08 | 1.196e-06 |
| 26 | MACROMOLECULAR COMPLEX BINDING | 27 | 1399 | 6.778e-08 | 2.422e-06 |
| 27 | GLUTAMATE RECEPTOR BINDING | 6 | 37 | 9.066e-08 | 3.119e-06 |
| 28 | PROTEIN HETERODIMERIZATION ACTIVITY | 15 | 468 | 1.721e-07 | 5.71e-06 |
| 29 | KINASE ACTIVITY | 20 | 842 | 1.783e-07 | 5.712e-06 |
| 30 | PROTEIN COMPLEX BINDING | 21 | 935 | 2.143e-07 | 6.637e-06 |
| 31 | IDENTICAL PROTEIN BINDING | 24 | 1209 | 2.511e-07 | 7.524e-06 |
| 32 | SIGNAL TRANSDUCER ACTIVITY | 29 | 1731 | 4.088e-07 | 1.187e-05 |
| 33 | PROTEIN SERINE THREONINE KINASE ACTIVITY | 14 | 445 | 5.701e-07 | 1.605e-05 |
| 34 | GUANYLATE KINASE ACTIVITY | 4 | 12 | 6.281e-07 | 1.716e-05 |
| 35 | PHOSPHATASE BINDING | 9 | 162 | 6.706e-07 | 1.763e-05 |
| 36 | CADHERIN BINDING | 5 | 28 | 6.831e-07 | 1.763e-05 |
| 37 | TRANSCRIPTION FACTOR BINDING | 15 | 524 | 7.225e-07 | 1.814e-05 |
| 38 | ARMADILLO REPEAT DOMAIN BINDING | 4 | 13 | 9.03e-07 | 2.208e-05 |
| 39 | RECEPTOR ACTIVATOR ACTIVITY | 5 | 32 | 1.373e-06 | 3.27e-05 |
| 40 | PROTEIN KINASE ACTIVITY | 16 | 640 | 1.777e-06 | 4.126e-05 |
| 41 | REGULATORY REGION NUCLEIC ACID BINDING | 18 | 818 | 2.337e-06 | 5.169e-05 |
| 42 | TRANSFERASE ACTIVITY TRANSFERRING PHOSPHORUS CONTAINING GROUPS | 20 | 992 | 2.306e-06 | 5.169e-05 |
| 43 | PROTEIN SERINE THREONINE PHOSPHATASE ACTIVITY | 6 | 64 | 2.557e-06 | 5.523e-05 |
| 44 | TRANSMEMBRANE RECEPTOR PROTEIN SERINE THREONINE KINASE ACTIVITY | 4 | 17 | 2.95e-06 | 6.228e-05 |
| 45 | RECEPTOR REGULATOR ACTIVITY | 5 | 45 | 7.817e-06 | 0.0001614 |
| 46 | R SMAD BINDING | 4 | 23 | 1.067e-05 | 0.0002154 |
| 47 | NUCLEIC ACID BINDING TRANSCRIPTION FACTOR ACTIVITY | 21 | 1199 | 1.111e-05 | 0.0002196 |
| 48 | NUCLEOTIDE KINASE ACTIVITY | 4 | 24 | 1.274e-05 | 0.0002466 |
| 49 | PHOSPHATASE REGULATOR ACTIVITY | 6 | 87 | 1.536e-05 | 0.0002912 |
| 50 | CORE PROMOTER PROXIMAL REGION DNA BINDING | 11 | 371 | 1.723e-05 | 0.0003201 |
| 51 | PROTEIN C TERMINUS BINDING | 8 | 186 | 1.878e-05 | 0.0003355 |
| 52 | CELL ADHESION MOLECULE BINDING | 8 | 186 | 1.878e-05 | 0.0003355 |
| 53 | RECEPTOR SIGNALING PROTEIN SERINE THREONINE KINASE ACTIVITY | 6 | 92 | 2.116e-05 | 0.0003709 |
| 54 | TRANSCRIPTION FACTOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 10 | 328 | 3.338e-05 | 0.0005742 |
| 55 | PROTEIN HOMODIMERIZATION ACTIVITY | 15 | 722 | 3.408e-05 | 0.0005756 |
| 56 | GLYCOPROTEIN BINDING | 6 | 101 | 3.597e-05 | 0.0005967 |
| 57 | RNA POLYMERASE II TRANSCRIPTION FACTOR BINDING | 6 | 104 | 4.243e-05 | 0.0006915 |
| 58 | CO SMAD BINDING | 3 | 12 | 4.68e-05 | 0.0007496 |
| 59 | ION CHANNEL BINDING | 6 | 111 | 6.116e-05 | 0.0009629 |
| 60 | ENZYME REGULATOR ACTIVITY | 17 | 959 | 7.258e-05 | 0.001124 |
| 61 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II CORE PROMOTER PROXIMAL REGION SEQUENCE SPECIFIC BINDING | 8 | 226 | 7.491e-05 | 0.001141 |
| 62 | PROTEIN PHOSPHATASE BINDING | 6 | 120 | 9.436e-05 | 0.001414 |
| 63 | PHOSPHOTRANSFERASE ACTIVITY PHOSPHATE GROUP AS ACCEPTOR | 4 | 40 | 0.0001016 | 0.001499 |
| 64 | PHOSPHOPROTEIN PHOSPHATASE ACTIVITY | 7 | 178 | 0.0001114 | 0.001617 |
| 65 | RNA POLYMERASE II TRANSCRIPTION FACTOR ACTIVITY SEQUENCE SPECIFIC DNA BINDING | 13 | 629 | 0.0001237 | 0.001768 |
| 66 | NUCLEOBASE CONTAINING COMPOUND KINASE ACTIVITY | 4 | 48 | 0.0002084 | 0.002933 |
| 67 | TRANSCRIPTION FACTOR ACTIVITY PROTEIN BINDING | 12 | 588 | 0.0002557 | 0.003546 |
| 68 | CHROMATIN BINDING | 10 | 435 | 0.000337 | 0.004604 |
| 69 | PROTEIN PHOSPHORYLATED AMINO ACID BINDING | 3 | 24 | 0.0004082 | 0.005495 |
| 70 | TRANSCRIPTION COREPRESSOR ACTIVITY | 7 | 221 | 0.0004183 | 0.005551 |
| 71 | PHOSPHOPROTEIN BINDING | 4 | 60 | 0.0004936 | 0.006459 |
| 72 | TRANSCRIPTIONAL ACTIVATOR ACTIVITY RNA POLYMERASE II TRANSCRIPTION REGULATORY REGION SEQUENCE SPECIFIC BINDING | 8 | 315 | 0.0007058 | 0.009106 |
| Num | GO | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|
| 1 | CELL JUNCTION | 38 | 1151 | 2.218e-18 | 1.296e-15 |
| 2 | WNT SIGNALOSOME | 8 | 11 | 2.468e-16 | 7.206e-14 |
| 3 | APICAL JUNCTION COMPLEX | 15 | 128 | 1.847e-15 | 3.596e-13 |
| 4 | CELL CELL JUNCTION | 21 | 383 | 1.713e-14 | 2.501e-12 |
| 5 | ANCHORING JUNCTION | 23 | 489 | 2.225e-14 | 2.599e-12 |
| 6 | PHOSPHATASE COMPLEX | 10 | 48 | 2.637e-13 | 2.567e-11 |
| 7 | PROTEIN PHOSPHATASE TYPE 2A COMPLEX | 7 | 20 | 1.917e-11 | 1.599e-09 |
| 8 | CYTOPLASMIC VESICLE PART | 21 | 601 | 9.247e-11 | 6.751e-09 |
| 9 | BETA CATENIN DESTRUCTION COMPLEX | 6 | 14 | 1.313e-10 | 8.521e-09 |
| 10 | EXTRACELLULAR MATRIX | 17 | 426 | 9.229e-10 | 5.39e-08 |
| 11 | CELL CELL ADHERENS JUNCTION | 8 | 54 | 1.251e-09 | 6.641e-08 |
| 12 | NEURON PROJECTION | 24 | 942 | 2.191e-09 | 1.066e-07 |
| 13 | CELL CORTEX | 13 | 238 | 2.413e-09 | 1.084e-07 |
| 14 | CYTOSKELETON | 35 | 1967 | 3.622e-09 | 1.511e-07 |
| 15 | TRANSCRIPTION FACTOR COMPLEX | 14 | 298 | 3.965e-09 | 1.544e-07 |
| 16 | CELL PROJECTION | 33 | 1786 | 4.607e-09 | 1.682e-07 |
| 17 | INTRACELLULAR VESICLE | 27 | 1259 | 7.542e-09 | 2.591e-07 |
| 18 | NEURON PART | 27 | 1265 | 8.341e-09 | 2.706e-07 |
| 19 | VESICLE MEMBRANE | 17 | 512 | 1.44e-08 | 4.426e-07 |
| 20 | LATERAL PLASMA MEMBRANE | 7 | 50 | 2.123e-08 | 5.904e-07 |
| 21 | CELL SUBSTRATE JUNCTION | 15 | 398 | 2.076e-08 | 5.904e-07 |
| 22 | CYTOPLASMIC REGION | 13 | 287 | 2.27e-08 | 6.026e-07 |
| 23 | ENDOCYTIC VESICLE MEMBRANE | 10 | 152 | 3.178e-08 | 7.424e-07 |
| 24 | CELL SURFACE | 20 | 757 | 3.166e-08 | 7.424e-07 |
| 25 | CELL LEADING EDGE | 14 | 350 | 3.031e-08 | 7.424e-07 |
| 26 | EXTRACELLULAR SPACE | 27 | 1376 | 4.824e-08 | 1.084e-06 |
| 27 | ENDOCYTIC VESICLE | 12 | 256 | 5.584e-08 | 1.208e-06 |
| 28 | LAMELLIPODIUM | 10 | 172 | 1.024e-07 | 2.135e-06 |
| 29 | PLASMA MEMBRANE REGION | 21 | 929 | 1.926e-07 | 3.879e-06 |
| 30 | AXON | 14 | 418 | 2.688e-07 | 5.224e-06 |
| 31 | PROTEINACEOUS EXTRACELLULAR MATRIX | 13 | 356 | 2.773e-07 | 5.224e-06 |
| 32 | MEMBRANE REGION | 23 | 1134 | 3.202e-07 | 5.844e-06 |
| 33 | CELL BODY | 15 | 494 | 3.434e-07 | 6.076e-06 |
| 34 | SOMATODENDRITIC COMPARTMENT | 17 | 650 | 4.386e-07 | 7.534e-06 |
| 35 | PLASMA MEMBRANE PROTEIN COMPLEX | 15 | 510 | 5.14e-07 | 8.577e-06 |
| 36 | SYNAPSE | 18 | 754 | 7.335e-07 | 1.19e-05 |
| 37 | CELL PROJECTION PART | 20 | 946 | 1.114e-06 | 1.758e-05 |
| 38 | CYTOSKELETAL PART | 25 | 1436 | 1.542e-06 | 2.329e-05 |
| 39 | MEMBRANE MICRODOMAIN | 11 | 288 | 1.555e-06 | 2.329e-05 |
| 40 | GOLGI LUMEN | 7 | 94 | 1.744e-06 | 2.546e-05 |
| 41 | MICROTUBULE CYTOSKELETON | 21 | 1068 | 1.85e-06 | 2.635e-05 |
| 42 | EXCITATORY SYNAPSE | 9 | 197 | 3.399e-06 | 4.726e-05 |
| 43 | POSTSYNAPSE | 12 | 378 | 3.501e-06 | 4.755e-05 |
| 44 | SYNAPSE PART | 15 | 610 | 4.688e-06 | 6.222e-05 |
| 45 | CHROMOSOME | 18 | 880 | 6.443e-06 | 8.362e-05 |
| 46 | INTERCALATED DISC | 5 | 51 | 1.46e-05 | 0.0001814 |
| 47 | PLASMA MEMBRANE RAFT | 6 | 86 | 1.437e-05 | 0.0001814 |
| 48 | NEURONAL POSTSYNAPTIC DENSITY | 5 | 53 | 1.766e-05 | 0.0002149 |
| 49 | NUCLEAR CHROMOSOME | 13 | 523 | 1.883e-05 | 0.0002244 |
| 50 | EXTRINSIC COMPONENT OF PLASMA MEMBRANE | 7 | 136 | 2.013e-05 | 0.0002351 |
| 51 | DENDRITE | 12 | 451 | 2.064e-05 | 0.0002364 |
| 52 | CYTOPLASMIC MICROTUBULE | 5 | 57 | 2.527e-05 | 0.0002838 |
| 53 | PERINUCLEAR REGION OF CYTOPLASM | 14 | 642 | 3.759e-05 | 0.0004142 |
| 54 | CELL CELL CONTACT ZONE | 5 | 64 | 4.448e-05 | 0.000481 |
| 55 | BASOLATERAL PLASMA MEMBRANE | 8 | 211 | 4.623e-05 | 0.0004909 |
| 56 | CLATHRIN COATED ENDOCYTIC VESICLE | 5 | 65 | 4.795e-05 | 0.0005001 |
| 57 | APICAL PART OF CELL | 10 | 361 | 7.452e-05 | 0.0007548 |
| 58 | NUCLEAR CHROMATIN | 9 | 291 | 7.497e-05 | 0.0007548 |
| 59 | CHROMATIN | 11 | 441 | 8.261e-05 | 0.0008177 |
| 60 | ACTIN CYTOSKELETON | 11 | 444 | 8.773e-05 | 0.0008539 |
| 61 | CELL CORTEX PART | 6 | 119 | 9.009e-05 | 0.0008625 |
| 62 | PLASMA MEMBRANE RECEPTOR COMPLEX | 7 | 175 | 0.0001002 | 0.0009437 |
| 63 | MICROTUBULE ORGANIZING CENTER | 13 | 623 | 0.0001125 | 0.001043 |
| 64 | CORTICAL CYTOSKELETON | 5 | 81 | 0.0001376 | 0.001256 |
| 65 | EXTRINSIC COMPONENT OF MEMBRANE | 8 | 252 | 0.000159 | 0.001429 |
| 66 | CATALYTIC COMPLEX | 17 | 1038 | 0.0001883 | 0.001666 |
| 67 | IONOTROPIC GLUTAMATE RECEPTOR COMPLEX | 4 | 47 | 0.0001919 | 0.001673 |
| 68 | CLATHRIN COATED ENDOCYTIC VESICLE MEMBRANE | 4 | 48 | 0.0002084 | 0.00179 |
| 69 | PLATELET ALPHA GRANULE LUMEN | 4 | 55 | 0.0003534 | 0.002991 |
| 70 | RNA POLYMERASE II TRANSCRIPTION FACTOR COMPLEX | 5 | 101 | 0.0003862 | 0.003222 |
| 71 | CLATHRIN COATED VESICLE | 6 | 157 | 0.0004063 | 0.003342 |
| 72 | CORTICAL ACTIN CYTOSKELETON | 4 | 58 | 0.0004335 | 0.003516 |
| 73 | MYELIN SHEATH | 6 | 168 | 0.0005815 | 0.00464 |
| 74 | COATED VESICLE | 7 | 234 | 0.000588 | 0.00464 |
| 75 | SUPRAMOLECULAR FIBER | 12 | 670 | 0.0008166 | 0.006359 |
| 76 | MICROTUBULE | 9 | 405 | 0.0008533 | 0.006557 |
| 77 | ACTIN FILAMENT | 4 | 70 | 0.0008857 | 0.006631 |
| 78 | CYCLIN DEPENDENT PROTEIN KINASE HOLOENZYME COMPLEX | 3 | 31 | 0.0008787 | 0.006631 |
| 79 | RECEPTOR COMPLEX | 8 | 327 | 0.000899 | 0.006646 |
| 80 | HISTONE METHYLTRANSFERASE COMPLEX | 4 | 71 | 0.0009342 | 0.006819 |
| 81 | SPINDLE POLE | 5 | 126 | 0.001055 | 0.007608 |
| 82 | NUCLEAR TRANSCRIPTION FACTOR COMPLEX | 5 | 127 | 0.001093 | 0.007786 |
| 83 | PLATELET ALPHA GRANULE | 4 | 75 | 0.001147 | 0.00807 |
| 84 | GOLGI APPARATUS | 19 | 1445 | 0.001185 | 0.008242 |
| 85 | MEMBRANE PROTEIN COMPLEX | 15 | 1020 | 0.001379 | 0.009472 |
| 86 | DENDRITIC SHAFT | 3 | 37 | 0.001479 | 0.009928 |
| 87 | ENDOPLASMIC RETICULUM LUMEN | 6 | 201 | 0.001472 | 0.009928 |
Over-represented Pathway
| Num | Pathway | Pathview | Overlap | Size | P Value | Adj. P Value |
|---|---|---|---|---|---|---|
| 1 | hsa04390_Hippo_signaling_pathway | 122 | 154 | 3.197e-289 | 5.754e-287 | |
| 2 | hsa04310_Wnt_signaling_pathway | 46 | 151 | 7.67e-68 | 6.903e-66 | |
| 3 | hsa04916_Melanogenesis | 28 | 101 | 1.709e-39 | 1.025e-37 | |
| 4 | hsa04350_TGF.beta_signaling_pathway | 24 | 85 | 4.244e-34 | 1.91e-32 | |
| 5 | hsa04340_Hedgehog_signaling_pathway | 20 | 56 | 6.518e-31 | 2.346e-29 | |
| 6 | hsa04530_Tight_junction | 20 | 133 | 1.56e-22 | 4.681e-21 | |
| 7 | hsa04520_Adherens_junction | 16 | 73 | 5.24e-21 | 1.347e-19 | |
| 8 | hsa04110_Cell_cycle | 16 | 128 | 7.041e-17 | 1.584e-15 | |
| 9 | hsa04114_Oocyte_meiosis | 13 | 114 | 2.174e-13 | 4.348e-12 | |
| 10 | hsa04151_PI3K_AKT_signaling_pathway | 18 | 351 | 4.617e-12 | 8.311e-11 | |
| 11 | hsa04144_Endocytosis | 11 | 203 | 4.719e-08 | 7.722e-07 | |
| 12 | hsa04510_Focal_adhesion | 10 | 200 | 4.165e-07 | 6.247e-06 | |
| 13 | hsa03015_mRNA_surveillance_pathway | 7 | 83 | 7.475e-07 | 1.035e-05 | |
| 14 | hsa04115_p53_signaling_pathway | 6 | 69 | 3.987e-06 | 5.127e-05 | |
| 15 | hsa04710_Circadian_rhythm_._mammal | 4 | 23 | 1.067e-05 | 0.000128 | |
| 16 | hsa04722_Neurotrophin_signaling_pathway | 7 | 127 | 1.29e-05 | 0.0001451 | |
| 17 | hsa04810_Regulation_of_actin_cytoskeleton | 8 | 214 | 5.108e-05 | 0.0005408 | |
| 18 | hsa04670_Leukocyte_transendothelial_migration | 6 | 117 | 8.2e-05 | 0.00082 | |
| 19 | hsa04910_Insulin_signaling_pathway | 6 | 138 | 0.0002031 | 0.001924 | |
| 20 | hsa04010_MAPK_signaling_pathway | 7 | 268 | 0.001298 | 0.01168 | |
| 21 | hsa04330_Notch_signaling_pathway | 3 | 47 | 0.002953 | 0.02531 | |
| 22 | hsa04720_Long.term_potentiation | 3 | 70 | 0.009007 | 0.07049 | |
| 23 | hsa04730_Long.term_depression | 3 | 70 | 0.009007 | 0.07049 | |
| 24 | hsa04630_Jak.STAT_signaling_pathway | 4 | 155 | 0.01505 | 0.1129 | |
| 25 | hsa04012_ErbB_signaling_pathway | 3 | 87 | 0.01619 | 0.1166 | |
| 26 | hsa04270_Vascular_smooth_muscle_contraction | 3 | 116 | 0.03412 | 0.2362 | |
| 27 | hsa04380_Osteoclast_differentiation | 3 | 128 | 0.04363 | 0.2909 | |
| 28 | hsa04660_T_cell_receptor_signaling_pathway | 2 | 108 | 0.1409 | 0.906 | |
| 29 | hsa04120_Ubiquitin_mediated_proteolysis | 2 | 139 | 0.2081 | 1 | |
| 30 | hsa04145_Phagosome | 2 | 156 | 0.2463 | 1 | |
| 31 | hsa04062_Chemokine_signaling_pathway | 2 | 189 | 0.3207 | 1 | |
| 32 | hsa04014_Ras_signaling_pathway | 2 | 236 | 0.4231 | 1 |
lncRNA-mediated sponge
| Num | lncRNA | miRNAs | miRNAs count | Gene | Sponge regulatory network | lncRNA log2FC | lncRNA pvalue | Gene log2FC | Gene pvalue | lncRNA-gene Pearson correlation |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | LINC00982 | hsa-miR-15b-5p;hsa-miR-186-5p;hsa-miR-192-3p;hsa-miR-193b-3p;hsa-miR-193b-5p;hsa-miR-194-5p;hsa-miR-197-3p;hsa-miR-224-3p;hsa-miR-365a-3p;hsa-miR-550a-5p | 10 | WTIP | Sponge network | -0.089 | 0.85759 | -0.083 | 0.6902 | 0.506 |
| 2 | HCG11 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-93-5p | 10 | TGFBR2 | Sponge network | -0.781 | 0 | -0.523 | 3.0E-5 | 0.472 |
| 3 | HCG11 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-362-3p;hsa-miR-455-5p;hsa-miR-618;hsa-miR-877-5p;hsa-miR-93-5p | 16 | BMPR2 | Sponge network | -0.781 | 0 | -0.736 | 0 | 0.47 |
| 4 | GAS5 | hsa-let-7a-2-3p;hsa-let-7g-3p;hsa-miR-101-3p;hsa-miR-125b-5p;hsa-miR-139-5p;hsa-miR-140-5p;hsa-miR-144-3p;hsa-miR-27b-3p;hsa-miR-345-5p;hsa-miR-590-3p | 10 | BBC3 | Sponge network | 1.966 | 0 | 0.805 | 0 | 0.454 |
| 5 | RP11-12A2.3 |
hsa-miR-1301-3p;hsa-miR-148b-3p;hsa-miR-148b-5p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-30d-5p;hsa-miR-338-3p;hsa-miR-877-5p;hsa-miR-93-3p | 13 | DLG2 | Sponge network | -4.779 | 0 | -1.947 | 0 | 0.452 |
| 6 | MAGI2-AS3 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-92a-3p;hsa-miR-93-5p | 11 | TGFBR2 | Sponge network | -1.801 | 0 | -0.523 | 3.0E-5 | 0.428 |
| 7 | MAGI2-AS3 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-30d-5p;hsa-miR-362-3p;hsa-miR-532-5p;hsa-miR-769-5p;hsa-miR-877-5p;hsa-miR-92a-3p;hsa-miR-93-3p;hsa-miR-93-5p | 18 | BMPR2 | Sponge network | -1.801 | 0 | -0.736 | 0 | 0.408 |
| 8 | RP11-175O19.4 | hsa-miR-193b-3p;hsa-miR-195-3p;hsa-miR-22-3p;hsa-miR-28-5p;hsa-miR-30c-5p;hsa-miR-30e-5p;hsa-miR-328-3p;hsa-miR-378a-3p;hsa-miR-455-3p;hsa-miR-505-5p | 10 | YWHAZ | Sponge network | 0.743 | 0 | 0.521 | 0 | 0.405 |
| 9 | RP11-166D19.1 |
hsa-miR-106b-5p;hsa-miR-17-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-324-3p;hsa-miR-330-5p;hsa-miR-484;hsa-miR-589-3p;hsa-miR-590-5p;hsa-miR-629-3p;hsa-miR-92a-3p;hsa-miR-940 | 15 | FZD4 | Sponge network | -0.244 | 0.28835 | -0.346 | 0.01097 | 0.384 |
| 10 | RP11-54O7.3 | hsa-miR-107;hsa-miR-140-3p;hsa-miR-15b-5p;hsa-miR-192-3p;hsa-miR-193b-3p;hsa-miR-193b-5p;hsa-miR-194-5p;hsa-miR-224-3p;hsa-miR-550a-5p;hsa-miR-589-5p | 10 | WTIP | Sponge network | -3.012 | 0 | -0.083 | 0.6902 | 0.377 |
| 11 | LDLRAD4-AS1 |
hsa-miR-1301-3p;hsa-miR-148b-3p;hsa-miR-148b-5p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-3127-5p;hsa-miR-421;hsa-miR-93-3p | 12 | DLG2 | Sponge network | -3.366 | 0 | -1.947 | 0 | 0.374 |
| 12 | LINC00238 | hsa-miR-1301-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-338-3p;hsa-miR-93-3p | 10 | DLG2 | Sponge network | -4.997 | 0 | -1.947 | 0 | 0.37 |
| 13 | RP11-12A2.3 |
hsa-miR-106b-5p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-30d-5p;hsa-miR-671-5p;hsa-miR-877-5p;hsa-miR-92a-3p;hsa-miR-93-3p;hsa-miR-93-5p | 15 | BMPR2 | Sponge network | -4.779 | 0 | -0.736 | 0 | 0.362 |
| 14 | MAGI2-AS3 |
hsa-miR-106b-5p;hsa-miR-1266-5p;hsa-miR-15b-5p;hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-589-3p;hsa-miR-616-5p;hsa-miR-9-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 13 | CCND1 | Sponge network | -1.801 | 0 | -0.902 | 1.0E-5 | 0.359 |
| 15 | SOCS2-AS1 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-21-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-532-5p;hsa-miR-671-5p;hsa-miR-93-3p;hsa-miR-93-5p | 12 | BMPR2 | Sponge network | -0.984 | 3.0E-5 | -0.736 | 0 | 0.355 |
| 16 | RP11-166D19.1 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-185-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-532-5p;hsa-miR-590-5p;hsa-miR-618;hsa-miR-877-5p;hsa-miR-92a-3p;hsa-miR-93-3p | 17 | BMPR2 | Sponge network | -0.244 | 0.28835 | -0.736 | 0 | 0.348 |
| 17 | SOCS2-AS1 |
hsa-miR-103a-3p;hsa-miR-140-3p;hsa-miR-146b-5p;hsa-miR-151a-3p;hsa-miR-16-1-3p;hsa-miR-17-5p;hsa-miR-181b-5p;hsa-miR-185-5p;hsa-miR-301a-3p;hsa-miR-425-5p;hsa-miR-532-5p;hsa-miR-93-5p | 12 | TEAD1 | Sponge network | -0.984 | 3.0E-5 | -0.192 | 0.10455 | 0.336 |
| 18 | CASC2 | hsa-miR-130b-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-362-3p;hsa-miR-532-5p;hsa-miR-877-5p;hsa-miR-92a-3p;hsa-miR-93-5p | 12 | BMPR2 | Sponge network | -0.596 | 0.00187 | -0.736 | 0 | 0.327 |
| 19 | RP11-166D19.1 |
hsa-miR-106b-5p;hsa-miR-130b-3p;hsa-miR-17-5p;hsa-miR-18a-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-301a-3p;hsa-miR-590-5p;hsa-miR-92a-3p | 11 | TGFBR2 | Sponge network | -0.244 | 0.28835 | -0.523 | 3.0E-5 | 0.324 |
| 20 | RP11-12A2.3 |
hsa-miR-103a-3p;hsa-miR-151a-3p;hsa-miR-17-5p;hsa-miR-181a-5p;hsa-miR-181b-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-301a-3p;hsa-miR-425-5p;hsa-miR-93-5p | 10 | TEAD1 | Sponge network | -4.779 | 0 | -0.192 | 0.10455 | 0.323 |
| 21 | MAGI2-AS3 |
hsa-miR-1301-3p;hsa-miR-148b-3p;hsa-miR-15b-3p;hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-30d-5p;hsa-miR-3127-5p;hsa-miR-769-3p;hsa-miR-769-5p;hsa-miR-877-5p;hsa-miR-93-3p | 14 | DLG2 | Sponge network | -1.801 | 0 | -1.947 | 0 | 0.298 |
| 22 | RP11-289F5.1 | hsa-miR-1301-3p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-182-5p;hsa-miR-185-5p;hsa-miR-19a-3p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-25-3p;hsa-miR-769-3p | 10 | DLG2 | Sponge network | -6.122 | 0 | -1.947 | 0 | 0.268 |
| 23 | LDLRAD4-AS1 |
hsa-miR-106b-5p;hsa-miR-148b-3p;hsa-miR-17-5p;hsa-miR-19a-3p;hsa-miR-20a-5p;hsa-miR-21-5p;hsa-miR-25-3p;hsa-miR-301a-3p;hsa-miR-93-3p;hsa-miR-93-5p | 10 | BMPR2 | Sponge network | -3.366 | 0 | -0.736 | 0 | 0.263 |
| 24 | RP11-166D19.1 |
hsa-miR-106a-5p;hsa-miR-106b-5p;hsa-miR-1266-5p;hsa-miR-15b-5p;hsa-miR-16-5p;hsa-miR-17-5p;hsa-miR-186-5p;hsa-miR-19a-3p;hsa-miR-19b-1-5p;hsa-miR-19b-3p;hsa-miR-20a-5p;hsa-miR-425-5p;hsa-miR-589-3p;hsa-miR-616-5p;hsa-miR-92a-3p;hsa-miR-942-5p | 16 | CCND1 | Sponge network | -0.244 | 0.28835 | -0.902 | 1.0E-5 | 0.253 |